Abstract
Rheumatoid arthritis (RA) is a joint disorder and is considered an important public health concern nowadays. So, identifying novel biomarkers and treatment modalities is urgently needed to improve the health standard of RA patients. Factors involved in RA pathogenesis are genetic/epigenetic modification, environment, and lifestyle. In the case of epigenetic modification, the expression deregulation of microRNAs and the role of histone deacetylase (HDAC) in RA is an important aspect that needs to be addressed. The present study is designed to evaluate the expression pattern of microRNAs related to the HDAC family. Five microRNAs, miR-92a-3p, miR-455-3p, miR-222, miR-140, and miR-146a related to the HDAC family were selected for the present study. Real-time polymerase chain reaction was used to estimate the level of expression of the above-mentioned microRNAs in 150 patients of RA versus 150 controls. Oxidative stress level and histone deacetylation status were measured using the enzyme-linked immunosorbent assay. Statistical analysis showed significant downregulation (P < .0001) of selected microRNAs in RA patients versus controls. Significantly raised level of HDAC (P < .0001) and 8-hydroxy-2’-deoxyguanosine (P < .0001) was observed in patients versus controls. A good diagnostic potential of selected microRNAs in RA was shown by the receiver operating curve analysis. The current study showed a significant role of deregulated expression of the above-mentioned microRNAs in RA initiation and can act as an excellent diagnostic marker for this disease.
Keywords: diagnosis, histone deacetylation, microRNA, oxidative stress, rheumatoid arthritis
1. Introduction
MicroRNAs (miRNAs) are evolutionary conserved small (21–25 nt), endogenous, non-coding RNAs that regulate the translation of messenger RNA through complementary mRNA–mRNA interaction.[1] They function as a post-transcriptional regulator of their target mRNA through either translational repression or mRNA degradation.[2] Approximately 30% of fundamental genes involved in basic cellular processes, for example, proliferation, survival, differentiation, invasion, and apoptosis, have been identified to be regulated by miRNAs.[3–6] Deregulation expression of miRNAs has been reported in various pathological conditions such as cardiovascular and neurodegenerative diseases, diabetes, cancers, and autoimmune diseases.[7–10] The literature review suggests that miRNA regulation is an important aspect of the mammalian immune system. The dysregulation of miRNAs is significantly implicated in the pathophysiology of both immunity and autoimmunity. Among the most prevalent autoimmune diseases is rheumatoid arthritis (RA).
RA is a chronic, inflammatory autoimmune disease that affects the joints.[11,12] It is characterized by increasing symmetric inflammation of affected joints causing cartilage weakening and destruction, bone erosion, and impairment.[13] Nodule growth under the skin, swollen and tender joints, weight loss, fever, and weariness, are also some of the common symptoms of RA.[14,15] RA is prevalent in 1% of the global population and presents at any age. Still, usually, the onset of the disease is from the age of 35 to 60 years. It is more prevalent in women than in men.[15–17] Although the cause of RA is unknown, genetically predisposed individuals are more likely to develop the disease because of a confluence of genetic variations, epigenetic changes, and environmental factors brought on by stochastic events such as injuries or infections.[13] Risk factors associated with the development of RA include smoking, exposure to ultraviolet radiation, obesity, infections, drugs, sex hormones, and changes in the microbiome of the mouth, stomach, and lungs.[18–22] The change in the expression profile of various miRNAs has been reported in different autoimmune diseases including RA. Several RA-associated miRNAs have been identified in previous studies. Studies on deacetylation-linked miRNAs in RA still need to be explored. The identification of miRNAs involved in RA not only helps understand the mechanisms behind this pathogenesis but also acts as potential biomarkers and novel targets for treating patients with RA.
Recently, a specific sub-group of miRNAs has been identified as Epi-miRNAs. Epi-miRNAs can target major enzymatic regulators or effectors of epigenetic machinery such as DNA methyl transferase, histone deacetylase (HDAC), and poly-comb disease.[23] HDAC removes the acetyl group from histones and induces gene silencing and abnormalities in several physiological processes.[24] The present study is designed for the investigation of the expression deregulation of miRNAs that target the HDAC in RA patients. The miRNAs included in this study are miR-92a-3p, miR-455-3p, miR-222, miR-140, and miR-146a.
MiR-92a-3p is located at q31.313.[25] It has oncogenic roles in multiple diseases such as cancer.[26,27] The overexpression of miR-92a-3p in lymphocytes has been associated with autoimmune disorders but the underlying mechanisms are yet unknown.[28] MiR-455-3p encoded by COL27A1 gene is present on chromosome number 9 (9q32).[29,30] This miRNA has been associated with many human diseases, for example, adipogenesis, chondrogenesis, preeclampsia, and several types of cancers.[31] MiR-222 is present on the X chromosome (Xp11.3) in a cluster form along with miR-221.[32] miR-221/222 plays an important role as an oncogene and is involved in the severity of different inflammatory diseases such as RA.[33,34] MiR-140 located on chromosome 16 (16q22.1) has multiple target sites and is downregulated in several human malignancies.[35,36] It is also involved in various other pathological conditions such as adipogenesis, chondrogenic differentiation, and skeleton development. MiR-146a is present on human chromosomes 5, 10, and 19.[37,38] Several studies have shown its involvement in inflammation, differentiation, and role in adaptive and innate immunity.[12,39,40]
Furthermore, histone deacetylation and oxidative stress levels will also be assessed in patients of RA compared to controls to elucidate the diagnostic significance of miRNAs in RA patients.
2. Materials and methods
2.1. Sample collection
The present study was conducted after getting ethical approval from the ethical review boards of both the Traditional Chinese Medical Hospital of Huzhou and the associated hospital. One hundred and fifty RA patients’ blood samples who fulfilled the American Rheumatology Criteria 1987, were collected from the hospitals. The same number of healthy individuals were recruited from the above-mentioned hospitals. Patients and control samples were collected during the years 2018 to 2022. For the collection of samples from patients, a non-probability purposive sampling technique was used. The clinical details of all the individuals selected in the current study were also collected. The individuals chosen in the study had no previous disease history of RA or other joint disorders and were diagnosed with different subtypes of RA, with a duration of symptoms of <12 months. Furthermore, individuals who showed no history of other diseases including hypertension, diabetes, impaired liver functioning, current or chronic diarrhea, proteinuria, lung disease, heart failure, and any chronic inflammatory conditions were also engaged in the present study.
Patients included in this study were receiving 2 disease-modifying antirheumatic drugs (DMARDs). First is a conventional synthetic DMARD in which patients receive the dosage of methotrexate (10 mg tablet, 15 mg, and 20 mg 1 tablet weekly for 4 weeks), duration is 3–6 months initially and then continue for life if remission achieved specific doses. Second is biological DMARDs, in which patients receive etanercept (50 mg subcutaneously weekly for 3–6 months), adalimumab (40 mg subcutaneously weekly for 3–6 months), and infliximab (3 mg IV at 0, 2, and 6 weeks).
Control samples were collected from people who visited these hospitals for regular medical checkups. The control samples were also selected based on criteria of age groups and sex-matched individuals having no earlier family or genetic history of joint diseases including RA. Written consent was taken from each individual before the collection of samples. Performa was filled for each participating individual including the required data about ethnicity, gender, age, dietary consumption, and other habits. The histopathological and demographic parameters of patients having RA patients along with controls are provided in Table 1.
Table 1.
Detailed characteristics of participants included in the study cohort.
| Parameter | Patients | Controls |
|---|---|---|
| Gender | ||
| Male | 58 | 63 |
| Female | 92 | 87 |
| Age | ||
| ˂50 | 124 | 132 |
| ≥50 | 26 | 18 |
| Anti-CCP | ||
| Positive | 84 | |
| Negative | 66 | |
| ESR | ||
| ˂31 | 57 | |
| ≥31 | 93 | |
| CRP | ||
| ˂14 | 69 | |
| ≥14 | 81 | |
| Treatment | ||
| Methotrexate | 114 | |
| Biologics | 36 |
anti-CCP = anti–cyclic citrullinated peptide, CRP = C-reactive protein, ESR = erythrocyte sedimentation rate.
2.2. miRNA identification and primer designing
The detailed sequences of miRNAs; miR-92a-3p, miR-455-3p, miR-222, miR-140, miR-146a, and U6 endogenous control were retrieved from the miRBase database. Specific primers for selected miRNAs were designed through Primer Quest of Integrated DNA Technology (IDT) software and confirmed by NCBI Primer Blast and UCSC Insilico polymerase chain reaction (PCR).
2.3. cDNA synthesis and quantitative PCR
Blood samples of RA patients and controls were subjected to RNA extraction manually through the Trizol method (Guanidinium thiocyanate-phenol-chloroform extraction). RNA concentration was quantified via a nanodrop spectrophotometer (ND-1000, Thermo Scientific, DE). The RNA extracted from the blood samples proceeded to cDNA synthesis using a high capacity cDNA reverse transcription kit (Invitrogen, CA) and stored at −20°C. The presence of cDNA in samples was confirmed by the amplification of U6 as an endogenous control.
Quantitative real-time PCR was used for the expression analysis of miR-92a-3p, miR-455-3p, miR-222, miR-140, and miR-146a in the whole blood of RA patients’ samples and their controls. Furthermore, U6 was used as an internal control. For the execution of the PCR StepOne PCR System (Applied Biosystem, MA) was used. The relative expression of selected miRNAs was evaluated by performing 2-ΔΔCT analysis method.
2.4. HDAC level
HDAC level was estimated in patients having RA and control samples following a kit specific to measure histone deacetylation (Abcam, Cambridge, UK). Firstly, the cell lysate of blood and control samples was prepared using a lysis buffer. Then HDAC activity was later assessed at regular intervals employing the fluorescence reader (AMP Diagnostics, Graz, Austria).
2.5. Measurement of oxidative stress
Oxidative stress in RA patients and control samples was determined by calculating the concentration of 8-hydroxy-2’-deoxyguanosine (8-OHdG). As per the method outlined, the experiment was started with the preparation of the cell lysate of blood and control samples followed by measurement of 8-OHdG levels. The experiment was performed using an 8-hydroxy-2-deoxyguanosine kit (Abcam). The absorbance of 8-OHdG was assessed via a plate reader (AMP Diagnostics, Graz, Austria), and the calibration curves attained depicted the calibration of the provided samples.
2.6. Statistical analysis
GraphPad Prism and SPSS software were used for performing suitable statistical analysis. The association of relative expression of miR-92a-3p, miR-455-3p, miR-222, miR-140, and miR-146a with the clinical parameters including gender, age, and smoking status was assessed by applying student t test. Diagnostic values of miRNAs were estimated using the receiver operating curve analysis by calculating the area under the curve. Spearman correlation was also performed between miRNA-miRNA correlation and miRNA to HDAC and 8-OHdG levels. The statistical significance was set at an established P-value; P ≤ .05.
3. Results
3.1. Expression analysis of miRNAs in RA patients and controls
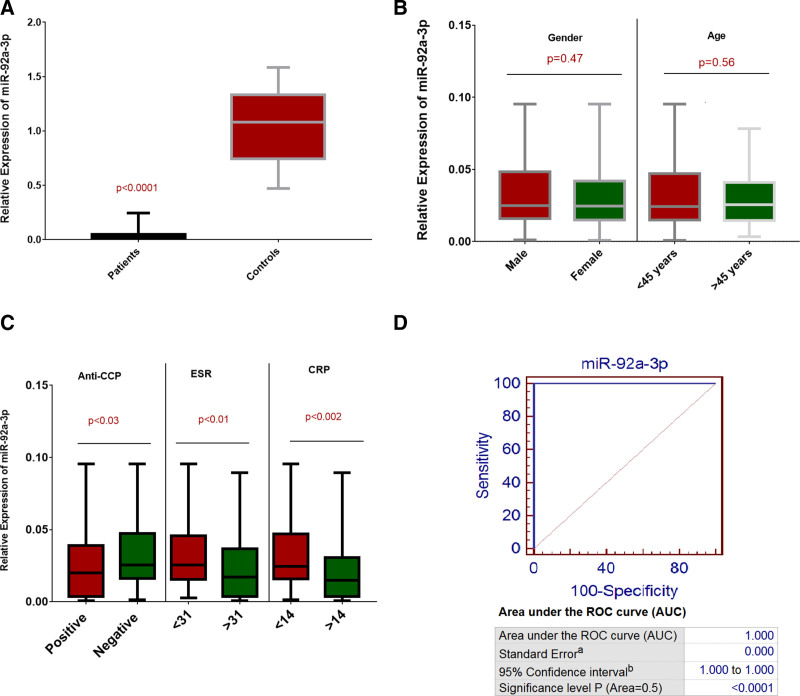
The relative expression of miRNA-92-3p was evaluated in RA patients and controls using real-time PCR. The expression of miRNA-92-3p was significantly downregulated (P < .0001) in patients in comparison to controls, as indicated in Figure 1A. Downregulated expression of miRNA-92-3p was observed among RA patients having negative anti–cyclic citrullinated peptide (CCP; P < .03) in comparison to those with positive anti-CCP, with erythrocyte sedimentation rate (ESR) > 31 (P < .01) in comparison to ESR < 31 and with C-reactive protein (CRP) > 14 (P < .002) versus CRP < 14, as shown in Figure 1C. Furthermore, downregulation of miRNA-92-3p was also observed among different age (P = .56) and gender (P = .47)-based categories of RA patients. Statistically non-significant results were observed (Fig. 1B).
Figure 1.
Expression analysis of miR-92a-3p. Relative expression of miR-92a-3p in (A) RA patients compared to controls, (B) male patients compared to female patients and patients with age group <45 yr compared to >45 yr, (C) anti–CCP-positive patients compared anti–CCP-negative patients; ESR < 31 compared to > 31 and CRP < 14 compared to CRP > 14. (D) ROC analysis of miR-92a-3p in RA patients. AUC = area under the curve, CCP = cyclic citrullinated peptide, CRP = C-reactive protein, ESR = erythrocyte sedimentation rate, RA = rheumatoid arthritis, ROC = receiver operating curve.
Expression analysis of the second selected miRNA-455-3p was investigated among samples of patients having RA and controls. A statistically significant reduced expression (P < .0001) of the selected miRNA was observed among RA patients in comparison to controls (Fig. 2A). Statistically significant reduced expression of miRNA-455-3p was noticed in patients of RA with negative anti-CCP (P < .01) in comparison to positive anti-CCP, with ESR > 31 (P < .04) versus ESR < 31 and with CRP > 14 (P < .01) versus CRP < 14, as shown in Figure 2C. Furthermore, miRNA-455-3p showed significantly decreased expression in different age (P < .008) and gender (P < .02)-based groups of RA patients as represented in Figure 2B.
Figure 2.
Expression analysis of miR-455-3p. Relative expression of miR-455-3p in (A) RA patients compared to controls, (B) male patients compared to female patients and patients with age group <45 yr compared to >45 yr, (C) anti–CCP-positive patients compared anti–CCP-negative patients; ESR < 31 compared to >31 and CRP < 14 compared to CRP > 14. (D) ROC analysis of miR-92a-3p in RA patients. AUC = area under the curve, CCP = cyclic citrullinated peptide, CRP = C-reactive protein, ESR = erythrocyte sedimentation rate, RA = rheumatoid arthritis, ROC = receiver operating curve.
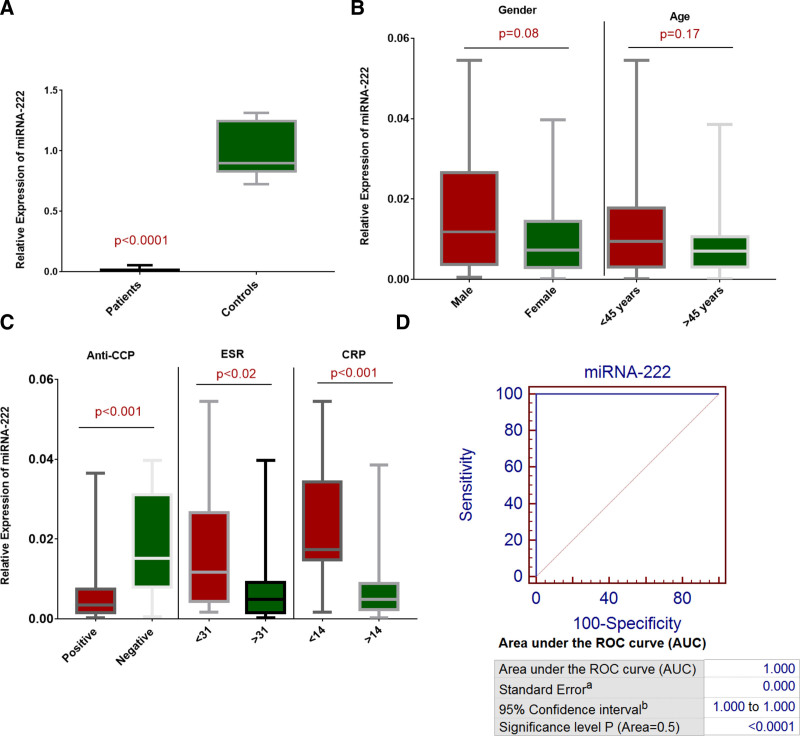
The third selected miRNA, miRNA-222 expression level was assessed among RA patients and controls. miRNA-222 showed significant downregulation (P < .0001) among patients versus controls as indicated in Figure 3A. Statically significant downregulation of miRNA-222 was observed among RA patients having positive anti-CCP (P < .001) versus negative anti-CCP, with ESR > 31 (P < .02) in comparison to ESR < 31 and with CRP > 14 (P < .001) versus CRP < 14, as shown in Figure 3C. Furthermore, miRNA-222 also showed downregulation in different genders (P = .08) and age groups (P = .17) of RA patients. The results obtained were found statistically non-significant (Fig. 3B).
Figure 3.
Expression analysis of miR-222. Relative expression of miR-222 in (A) RA patients compared to controls, (B) male patients compared to female patients and patients with age group <45 yr compared to >45 yr, (C) anti–CCP-positive patients compared anti–CCP-negative patients; ESR < 31 compared to >31 and CRP < 14 compared to CRP > 14. (D) ROC analysis of miR-92a-3p in RA patients. AUC = area under the curve, CCP = cyclic citrullinated peptide, CRP = C-reactive protein, ESR = erythrocyte sedimentation rate, RA = rheumatoid arthritis, ROC = receiver operating curve.
The expression level of the third selected miRNA, miRNA-140 was assessed among RA patients and controls. miRNA-140 (P < .0001) indicated a statistically significant downregulation among RA patients in comparison to controls as indicated in Figure 4A. Further analysis showed strong downregulation of miRNA-140 in RA patients with positive anti-CCP (P < .007) compared to negative anti-CCP, with ESR > 31 (P < .0001) versus ESR < 31 and with CRP > 14 (P < .001) versus CRP < 14, as indicated in Figure 4C. Furthermore, downregulated expression of the same miRNA-140 was found in different age groups (P = .29) and gender (P = .45) of RA patients. Thus, the findings were statistically non-significant (Fig. 4B).
Figure 4.
Expression analysis of miR-140. Relative expression of miR-140 in (A) RA patients compared to controls, (B) male patients compared to female patients and patients with age group <45 yr compared to >45 yr, (C) anti–CCP-positive patients compared anti–CCP-negative patients; ESR < 31 compared to > 31 and CRP < 14 compared to CRP > 14. (D) ROC analysis of miR-92a-3p in RA patients. AUC = area under the curve, CCP = cyclic citrullinated peptide, CRP = C-reactive protein, ESR = erythrocyte sedimentation rate, RA = rheumatoid arthritis, ROC = receiver operating curve.
The expression level of the fourth selected miRNA, miRNA-146a was measured by performing real-time PCR. Significantly reduced expression of miRNA-146a was observed among RA patients (P < .0001) versus controls as indicated in Figure 5A. Downregulated expression of miRNA-146a was observed in RA patients having positive anti-CCP (P < .001) in comparison to negative anti-CCP, with ESR > 31(P < .001) in comparison to ESR < 31 and with CRP > 14 (P < .001) in comparison to CRP < 14, as shown in Figure 5C. Further analysis indicated the downregulation of miRNA-146a in different gender (P < .002) and age groups (P < .03) of RA patients, as shown in Figure 5B.
Figure 5.
Expression analysis of miR-146a. Relative expression of miR-146a in (A) RA patients compared to controls, (B) male patients compared to female patients and patients with age group <45 yr compared to >45 yr, (C) anti–CCP-positive patients compared anti–CCP-negative patients; ESR < 31 compared to > 31 and CRP < 14 compared to CRP > 14. (D) ROC analysis of miR-92a-3p in RA patients. AUC = area under the curve, CCP = cyclic citrullinated peptide, CRP = C-reactive protein, ESR = erythrocyte sedimentation rate, RA = rheumatoid arthritis, ROC = receiver operating curve.
3.2. Association of expression level of selected miRNAs with treatment modalities
In the present study, patients having RA were categorized into 2 groups depending upon the nature of treatments such as 76% of patients receiving methotrexate treatment and 14% biologics treatment as listed in Table 1. Relative expression of the aforementioned miRNAs showed a correlation with treatment types received by RA patients. Significantly upregulated expression of miRNA-92a-3p (P < .0001), miRNA-455-3p (P < .01), miRNA-222 (P < .007), miRNA-140 (P < .03), and miRNA-146a (P < .0001) was observed among the RA patients given biologics treatment in comparison to those receiving methotrexate, as illustrated in Figure 6.
Figure 6.
Association of selected miRNAs with different treatment modalities used in RA patient treatments. Level of significance P < .05. miRNAs = microRNA, RA = rheumatoid arthritis.
3.3. Spearman correlation of selected miRNAs
Correlation between the selected miRNAs, miR-92a-3p, miR-455-3p, miR-222, miR-140, and miR-146a HDAC level, and 8-OHdG level in RA patients was assessed by performing spearman correlation analysis (Table 2). A significantly positive correlation was observed between miR-92a-3p versus miR-455-3p (r = 0.419; P < .001), miR-92a-3p versus miR-140 (r = 0.682; P < .0001), miR-455-3p versus miR-140 (r = 0.558; P < .0003), miR-455-3p versus miR-222 (r = 0.402; P < .001), miR-140 versus miR-146a (r = 0.305; P < .04), HDAC versus 8-OHdG (r = 0.514; P < .001) in RA patients. However, a strongly negative correlation was observed between HDAC versus miR-92a-3p (r = –0.527; P < .0004), HDAC versus miR-455-3p (r = –0.441; P < .001), HDAC versus miR-222 (r = –0.417; P < .001), HDAC versus miR-140 (r = –0.311; P < .03), and HDAC versus miR-146a (r = –0.386; P < .02) in RA patients (Table 2). Negative correlation was also observed between 8-OHdG versus miR-92a-3p (r = –0.413; P < .001), 8-OHdG versus miR-455-3p (r = –0.373; P < .02), 8-OHdG versus miR-222 (r = –0.305; P < .03), 8-OHdG versus miR-140 (r = –0.338; P < .03), and 8-OHdG versus miR-146a (r = –0.315; P < .03) in RA patients, as shown in Table 2.
Table 2.
Correlation analysis of microRNAs and parameters in RA patients.
| Parameters | miR-455-3p | miR-140 | miR-146a | miR-222 | HDAC | 8-OHdG |
|---|---|---|---|---|---|---|
| miR-92a-3p | r = 0.419** | r = 0.685*** | r = 0.493** | r = 0.207 | r = –0.527*** | r = −0.413** |
| miR-455-3p | r = 0.558*** | r = 0.268 | r = 0.402** | r = –0.441** | r = –0.373* | |
| miR-140 | r = 0.305* | r = 0.143 | r = –0.31* | r = –0.338* | ||
| miR-146a | r = 0.146 | r = –0.386* | r = 0–.315* | |||
| miR-222 | r = –0.417** | r = –0.305** | ||||
| HDAC | r = 0.54** |
r = spearman correlation coefficient; level of significance P < .05.
HDAC = histone deacetylase, 8-OHdG = 8-hydroxy-2’-deoxyguanosine.
P < .05,
P < .01, and
P < .001.
3.4. Diagnostic potential of selected miRNAs in RA patients
The diagnostic significance of selected miRNAs, miR-92a-3p, miR-455-3p, miR-222, miR-140, and miR-146a was assessed by performing receiver operating curve analysis. 100% specificity and sensitivity were observed for miR-92a-3p (P < .0001) in RA patients, as shown in Figure 1D. 98% specificity and sensitivity were found in the case of miR-455-3p (P < .0001) in RA patients (Fig. 2D). 100% specificity and sensitivity were observed for miR-222 (P < .0001) in RA patients as shown in Figure 3D. However, 100 % specificity and sensitivity were calculated for miR-140 (P < .0001) (Fig. 4D). 99% specificity and sensitivity were observed in the case of miR-146a (P < .0001) as shown in Figure 5D.
3.5. Measurement of deacetylation status and oxidative stress among RA patients and controls
In the current study design, the deacetylation level was assessed in RA patients and controls as shown in Figure 7A. A significantly increased level of histone deacetylation was noticed in RA (P < .0001) patients versus controls as illustrated in Figure 7A.
Figure 7.
(A) Measurement of HDAC level in RA patients compared to controls. (B) Measurement of 8-OHdG level in RA patients compared to controls. Level of significance P < .05. 8-OHdG = 8-hydroxyguanosine, HDAC = histone deacetylation, RA = rheumatoid arthritis.
Oxidative stress was estimated by measuring the 8-OHdG in patients versus controls as demonstrated in Figure 7B. A significantly increased level of 8-OHdG was observed among RA patients (P < .0001) versus controls as shown in Figure 7B.
4. Discussion
MiRNAs are usually small non-coding RNA fragments. These play a role in the generation of a reduced final output of proteins either by suppressing or destabilizing the translation mechanism. This protein dysregulation is involved in different physiological functions and ultimately results in accelerating the mechanism of pathogenesis.[41] The critical functioning of miRNA deregulation in different diseases has been reported extensively in different populations.[8–10,42,43] However, very few studies have been reported regarding the involvement of miRNAs in RA.[11,13] RA is a joint disorder worldwide with an increased ratio of systemic complications and severe health problems.[44] Literature has reported that the isolation and identification of new genetic parameters connected with different type of RA might potentially serve as an important tool in detecting potential biomarkers for early-stage disease diagnosis and therapeutic strategies.[14,16,45,46]
Present study results showed a significantly decreased expression of miR-92a-3p, miR-455-3p, miR-222, miR-140, and miR-146a in RA patients versus controls. Furthermore, the relative mRNA expression levels of the selected miRNAs were also assessed in various clinical and demographic parameters. Significant downregulation of these selected miRNAs was observed in RA patients with positive anti-CCP in comparison to negative anti-CCP, with ESR > 31 versus ESR < 31, and with CRP > 14 versus CRP < 14. Peng et al[47] have reported a decreased expression of miR-140 in synovial fluids and tissues of RA patients as compared to controls. miR-140 has been reported to be involved in osteoarthritis (OA) by regulating the expression of genes critical in the pathogenesis of this disease. The miR-140, suppresses the gene that plays a critical role in OA by targeting genes such as Smad3, ADAMTS-5, and HDAC4. Previous study has reported that miR-140 was involved in regulating, developing, and maintaining homeostasis and its downregulation is a contributing factor leading to age-related arthritis development.[44] The miR-140 knockout rat model has indicated the generation of the rat phenotype with early-stage OA, which establishes the fact that miR-140 is critical in maintaining cartilage homeostasis.[18,21]
miR-146a is a member of the miRNAs family that plays a dual role. It positively affects the disease activity by stimulating the inflammatory responses and negatively affects by suppression of joint destruction.[14–16] A significantly increased expression of miR-146a has been observed in synovial fluid/tissue of RA patients and significantly downregulated expression has been observed in plasma/serum.[3,4] Previous studies have reported a significantly increased expression of miR-146a, among the synovial fluids of patients having RA, however, reduced expression has been reported in OA.[6–10] Ramasamy et al[43] have reported a strong expression of miR-146a among macrophages of RA thereby initiating anti-inflammatory and proinflammatory symptoms accordingly. Furthermore, previous studies have reported that peripheral blood mononuclear cells and synovial tissues can frequently secrete miRNAs including miR-146a, which ultimately resulted in increased expression of miR-146a in synovial fluid and decreased expression in the serum of RA patients.[15,42]
miR-455-3p has a regulatory function in most tissue/organs and its upregulation has been reported for cartilage development.[22] In the present study, significant downregulation of miR-455-3p was observed in RA patients compared to controls. The reason behind the expression deregulation of miR-455-3p and the enhanced risk of RA needs yet to be explored. However, previous studies have indicated the expression dysregulation of miR-455-3p to suppress the genes associated with DNA methylation such as Smad3 and SOX6, and increased risk of bone/muscle disorder.[19,22] Kim et al[20] have reported that miR-455-3p expression dysregulation results in inhibiting DNMT3A expression. Hence this miRNA plays a significant role in the regulation of DNA methylation of cartilage-associated genes and the PI3K-Akt signaling cascade. miR-455-3p inhibits chondrogenic differentiation degeneration through modification of DNA methylation. This ultimately suppresses the degeneration of cartilage during chondrogenic differentiation.[19] Hu et al[22] have reported that the downregulation of miR-455-3p results in the upregulation of P21-activated kinases (PAKs), which ultimately leads to inhibition of tansfroming growth factor-β/Smad signaling in chondrocytes and increased risk of OA. Another study by Ito et al[23] has reported that miR-455-3p also targets the hypoxia-inducible factor-2α (HIF-2α) and the absence of this factor increased the frequency of cartilage degeneration/arthritis.
miR-92a-3p has been identified as the regulator of chondrogenic differentiation and bone growth/maturation. Studies have reported the expression deregulation of miR-92a-3p and enhanced risk of OA and arthritis. The mechanism behind this expression variation of miR-92a-3p and disease risk is still unknown. However, studies have shown that expression deregulation of miRNA-92a-3p leads to a change in the regulation of related genes/pathways and an increased risk of arthritis. Mao et al[31] have suggested that Interleukin-1β suppresses the expression of miR-92a-3p and plays a critical role in the regulation of ADAMTS-4/5 in OA pathogenesis and chondrogenesis. Another study has reported that suppression of miR-92a-3p enhanced the expression of its related gene ADAMTS-4/5 and was found associated with reduced aggrecan expression.[32] This expression variation cascade ultimately results in cartilage degradation/erosion and an increased risk of arthritis.[33] Mao et al[31] have reported the role of miR-92a-3p in disease severity by targeting the HDAC2 and showed that histone hyperacetylation resulted in upregulated expression of cartilage matrix.
miR-222 has the regulatory function for RA and it regulates different proteins which control angiogenesis and inflammation. In the present study, significant downregulation of miR-222 was found associated with an increased risk of RA. Zhuo et al[36] have suggested a significantly upregulated expression of miR-222 in RA patients compared to controls. Song et al[30] have reported that downregulation of miR-222 might have an association with cartilage destruction via targeting HDAC-4 and alteration of matrix metalloproteinases (MMP)-13 levels. Pandis et al[48] have reported that the deregulation of miR-222 resulted in the inhibition of proinflammatory cytokines, cell invasion, and apoptosis by targeting the MMP-3, MMP-9, and vascular endothelial growth factor.
Previous studies have reported that initiation/progression of arthritis is controlled by other factors, such as expression variation and activation of HDAC has also shown significant contribution in this process. Studies have shown that HDAC inhibitors can inhibit/control arthritis in animal models[14,15,49] by reducing the expression of matrix-degrading enzymes in chondrocytes and fibroblasts.[11,13,50] Furthermore, previous studies have found that HDAC regulation may be related to miRNAs.[31,50] To correlate the role of our selected miRNAs and HDAC in RA patients, measurement of HDAC level was measured among RA patients compared to controls. A significantly increased HDAC status was observed in RA patients in comparison to controls in the present study. Studies have reported that miRNAs have specific HDAC targets.[31,50,51] miRNA-222 targets HDAC4 and regulates MMP-13 which leads to the increased degradation of chondrocytes and increased risk of arthritis pathogenesis.[51] Mao et al[31] have reported that HDAC2 is targeted by miR-92a-3p and HDAC4 is targeted by miR-140 which leads to hyperacetylation of matrix protein and regulates the cartilage development and homeostasis. Chen et al[50] have reported that miR-455-3p directly targets the HDAC2/8 and increases histone acetylation and probability of cartilage degeneration. The findings of the current study suggested that a decreased expression of selected miRNA may result in enhanced regulation of HDAC and increased histone deacetylation. This increased histone deacetylation activates the different proteases and increases the risk of cartilage degradation and pathogenesis.
Oxidative stress was estimated by measuring the 8-OHdG level among patients of RA and controls. A significantly raised level of 8-OHdG was noticed among RA patients in comparison to controls. Further analysis showed that increased reactive oxygen species (ROS) levels showed a significant negative correlation with the deregulated expression levels of selected miRNAs. Ito et al[23] have suggested that miR-455-3p and miRNA-140 regulate the expression of HIF1α. Singhal et al[52] have indicated that hypoxia or generation of ROS is regulated with activation of HIF2α. Ramasamy et al[43] have reported that expression variation of miR-222 leads to the deregulation of peroxisome proliferator-activated-1α and the overproduction of ROS which ultimately leads to cartilage matrix degradation.
In conclusion, our study demonstrates the important impact that the deregulation of miRNAs (miR-92a-3p, miR-455-3p, miR-222, miR-140, and miR-146a) was found associated with RA initiation and pathogenesis. Further analysis revealed that alteration with increased levels of HDAC and oxidative stress with concomitant low expression of selected miRNAs have prognostic/diagnostic implications in RA. This also paved the way for further analysis of the functional consequences of the deregulation of miRNAs to fully understand the underlying mechanism underpinnings of the development of RA. Even more importantly, the value of prognostic markers should be tested both prospectively and for their predictive value in terms of therapy efficacy. A larger study cohort with a screening of a large number of miRNAs will be helpful to further clarify the mechanism involved in RA pathogenesis.
Acknowledgments
The authors are thankful to patients and staff of collaborating hospitals, for their contribution to this research.
Author contributions
Conceptualization: Zengxin Li, Mengchang Wang, Ishrat Mahjabeen.
Data curation: Zengxin Li, Wen Zhao, Mengchang Wang.
Formal analysis: Zengxin Li, Wen Zhao, Mengchang Wang, Muhammad Zahid Hussain.
Investigation: Zengxin Li, Wen Zhao.
Funding acquisition: Wen Zhao.
Validation: Mengchang Wang.
Visualization: Mengchang Wang.
Writing – original draft: Mengchang Wang, Ishrat Mahjabeen.
Writing – review & editing: Mengchang Wang, Ishrat Mahjabeen.
Methodology: Muhammad Zahid Hussain.
Resources: Muhammad Zahid Hussain.
Software: Ishrat Mahjabeen.
Abbreviations:
- 8-OHdG
- 8-hydroxy-2’-deoxyguanosine
- anti-CCP
- anti–cyclic citrullinated peptide
- AUC
- area under the curve
- CRP
- C-reactive protein
- ESR
- erythrocyte sedimentation rate
- HDAC
- histone deacetylase
- HIF-2α
- hypoxia-inducible factor-2α
- miRNA
- microRNA
- MMPs
- matrix metalloproteinases
- OA
- osteoarthritis
- PAKs =
- P21-activated kinases
- RA
- rheumatoid arthritis
- ROC
- receiver operating curve
- ROS
- reactive oxygen species.
ZL and WZ contributed equally to this work.
All data generated or analyzed during this study are included in this published article [and its supplementary information files].
How to cite this article: Li Z, Zhao W, Wang M, Hussain MZ, Mahjabeen I. Role of microRNAs deregulation in initiation of rheumatoid arthritis: A retrospective observational study. Medicine 2024;103:3(e36595).
The authors have no funding and conflicts of interest to disclose.
Informed consent was taken from all the participants of the present study.
The present study was conducted after getting ethical approval from the ethical review boards of both the Traditional Chinese Medical Hospital of Huzhou and the associated hospital.
Contributor Information
Zengxin Li, Email: 13843275615@163.com.
Wen Zhao, Email: zhaowen147@hotmail.com.
Muhammad Zahid Hussain, Email: zahidrajpoot@icloud.com.
Ishrat Mahjabeen, Email: ishrat.mahjabeen@comsats.edu.pk.
References
- [1].Zhang Y, Wang Z, Gemeinhart RA. Progress in microRNA delivery. J Control Release. 2013;172:962–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [2].Catalanotto C, Cogoni C, Zardo G. MicroRNA in control of gene expression: an overview of nuclear functions. Int J Mol Sci. 2016;17:1712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [3].Ambros V. The functions of animal microRNAs. Nature. 2004;431:350–5. [DOI] [PubMed] [Google Scholar]
- [4].Berezikov E, Guryev V, van de Belt J, et al. Phylogenetic shadowing and computational identification of human microRNA genes. Cell. 2005;120:21–4. [DOI] [PubMed] [Google Scholar]
- [5].Friedman RC, Farh KKH, Burge CB, et al. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009;19:92–105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].Jakob P, Landmesser U. Role of microRNAs in stem/progenitor cells and cardiovascular repair. Cardiovasc Res. 2012;93:614–22. [DOI] [PubMed] [Google Scholar]
- [7].Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857–66. [DOI] [PubMed] [Google Scholar]
- [8].Guay C, Roggli E, Nesca V, et al. Diabetes mellitus, a microRNA-related disease? Transl Res. 2011;157:253–64. [DOI] [PubMed] [Google Scholar]
- [9].Miller BH, Wahlestedt C. MicroRNA dysregulation in psychiatric disease. Brain Res. 2010;1338:89–99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [10].Williams AH, Liu N, Van Rooij E, et al. MicroRNA control of muscle development and disease. Curr Opin Cell Biol. 2009;21:461–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [11].Lin YJ, Anzaghe M, Schülke S. Update on the pathomechanism, diagnosis, and treatment options for rheumatoid arthritis. Cells. 2020;9:880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [12].Murata K, Yoshitomi H, Tanida S, et al. Plasma and synovial fluid microRNAs as potential biomarkers of rheumatoid arthritis and osteoarthritis. Arthritis Res Ther. 2010;12:R86–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [13].Smolen JS, Aletaha D, McInnes IB. Rheumatoid arthritis. Lancet. 2016;388:2023–38. [DOI] [PubMed] [Google Scholar]
- [14].Bullock J, Rizvi SA, Saleh AM, et al. Rheumatoid arthritis: a brief overview of the treatment. Med Princ Pract. 2018;27:501–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [15].McInnes IB, Schett G. Pathogenetic insights from the treatment of rheumatoid arthritis. Lancet. 2017;389:2328–37. [DOI] [PubMed] [Google Scholar]
- [16].Hameed K, Gibson T, Kadir M, et al. The prevalence of rheumatoid arthritis in affluent and poor urban communities of Pakistan. Br J Rheumatol. 1995;34:252–6. [DOI] [PubMed] [Google Scholar]
- [17].Farooqi A, Gibson T. Prevalence of the major rheumatic disorders in the adult population of North Pakistan. Br J Rheumatol. 1998;37:491–5. [DOI] [PubMed] [Google Scholar]
- [18].Myasoedova E, Crowson CS, Kremers HM, et al. Is the incidence of rheumatoid arthritis rising? Results from Olmsted County, Minnesota, 1955–2007. Arthritis Rheum. 2010;62:1576–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [19].Sun H, Zhao X, Zhang C, et al. MiR-455-3p inhibits the degenerate process of chondrogenic differentiation through modification of DNA methylation. Cell Death Dis. 2018;9:537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [20].Kim KI, Park YS, Im GI. Changes in the epigenetic status of the SOX-9 promoter in human osteoarthritic cartilage. J Bone Miner Res. 2013;28:1050–60. [DOI] [PubMed] [Google Scholar]
- [21].Littlejohn EA, Monrad SU. Early diagnosis and treatment of rheumatoid arthritis. Prim Care. 2018;45:237–55. [DOI] [PubMed] [Google Scholar]
- [22].Hu S, Zhao X, Mao G, et al. MicroRNA-455-3p promotes TGF-β signaling and inhibits osteoarthritis development by directly targeting PAK2. Exp Mol Med. 2019;51:1–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [23].Ito Y, Matsuzaki T, Ayabe F, et al. Both microRNA-455-5p and-3p repress hypoxia-inducible factor-2α expression and coordinately regulate cartilage homeostasis. Nat Commun. 2021;12:4148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [24].Zhou C, Zhao L, Wang K, et al. MicroRNA-146a inhibits NF-κB activation and pro-inflammatory cytokine production by regulating IRAK1 expression in THP-1 cells. Exp Ther Med. 2019;18:3078–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [25].Liu C, Zhang Y, Chen H, et al. Function analysis of rs9589207 polymorphism in miR-92a in gastric cancer. Tumour Biol. 2016;37:4439–44. [DOI] [PubMed] [Google Scholar]
- [26].Zhao JY, Wang F, Li Y, et al. Five miRNAs considered as molecular targets for predicting esophageal cancer. Med Sci Monit. 2015;21:3222–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [27].Ren P, Gong F, Zhang Y, et al. MicroRNA-92a promotes growth, metastasis, and chemoresistance in non-small cell lung cancer cells by targeting PTEN. Tumour Biol. 2016;37:3215–25. [DOI] [PubMed] [Google Scholar]
- [28].Xiao C, Srinivasan L, Calado DP, et al. Lymphoproliferative disease and autoimmunity in mice with increased miR-17-92 expression in lymphocytes. Nat Immunol. 2008;9:405–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].Kumar S, Reddy PH. A new discovery of MicroRNA-455-3p in Alzheimer’s disease. J Alzheimer's Dis. 2019;72:S117–30. [DOI] [PubMed] [Google Scholar]
- [30].Song J, Jin EH, Kim D, et al. MicroRNA-222 regulates MMP-13 via targeting HDAC-4 during osteoarthritis pathogenesis. BBA Clin. 2015;3:79–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [31].Mao G, Zhang Z, Huang Z, et al. MicroRNA-92a-3p regulates the expression of cartilage-specific genes by directly targeting histone deacetylase 2 in chondrogenesis and degradation. Osteoarthritis Cartilage. 2017;25:521–32. [DOI] [PubMed] [Google Scholar]
- [32].Ning G, Liu X, Dai M, et al. MicroRNA-92a upholds Bmp signaling by targeting noggin3 during pharyngeal cartilage formation. Dev Cell. 2013;24:283–95. [DOI] [PubMed] [Google Scholar]
- [33].Mort JS, Geng Y, Fisher WD, et al. Aggrecan heterogeneity in articular cartilage from patients with osteoarthritis. BMC Musculoskelet Disord. 2016;17:1–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [34].Abo ElAtta AS, Ali YB, Bassyouni IH, et al. Upregulation of miR-221/222 expression in rheumatoid arthritis (RA) patients: correlation with disease activity. Clin Exp Med. 2019;19:47–53. [DOI] [PubMed] [Google Scholar]
- [35].Flamini V, Jiang WG, Cui Y. Therapeutic role of MiR-140-5p for the treatment of non-small cell lung cancer. Anticancer Res. 2017;37:4319–27. [DOI] [PubMed] [Google Scholar]
- [36].Zhuo E, Cai C, Liu W, et al. Downregulated microRNA-140-5p expression regulates apoptosis, migration and invasion of lung cancer cells by targeting zinc finger protein 800. Oncol Lett. 2020;20:1–1. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- [37].Li L, Chen XP, Li YJ. MicroRNA-146a and human disease. Scand J Immunol. 2010;71:227–31. [DOI] [PubMed] [Google Scholar]
- [38].Nahand JS, Karimzadeh MR, Nezamnia M, et al. The role of miR-146a in viral infection. IUBMB Life. 2020;72:343–60. [DOI] [PubMed] [Google Scholar]
- [39].Piletič K, Kunej T. MicroRNA epigenetic signatures in human disease. Arch Toxicol. 2016;90:2405–19. [DOI] [PubMed] [Google Scholar]
- [40].Ahmadi S, Sharifi M, Salehi R. Locked nucleic acid inhibits miR-92a-3p in human colorectal cancer, induces apoptosis and inhibits cell proliferation. Cancer Gene Ther. 2016;23:199–205. [DOI] [PubMed] [Google Scholar]
- [41].Guo Q, Wang Y, Xu D, et al. Rheumatoid arthritis: pathological mechanisms and modern pharmacologic therapies. Bone Res. 2018;6:15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [42].Pauley KM, Satoh M, Chan AL, et al. Upregulated miR-146a expression in peripheral blood mononuclear cells from rheumatoid arthritis patients. Arthritis Res Ther. 2008;10:R101–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [43].Ramasamy TS, Yee YM, Khan IM. Chondrocyte aging: the molecular determinants and therapeutic opportunities. Front Cell Dev Biol. 2021;9:625497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [44].Miyaki S, Asahara H. Macro view of microRNA function in osteoarthritis. Nat Rev Rheumatol. 2012;8:543–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [45].Menis J, Doussiere M, Touboul E, et al. Current characteristics of a population of psoriatic arthritis and gender disparities. J Clin Transl Res. 2023;9:84–92. [PMC free article] [PubMed] [Google Scholar]
- [46].Farrugia M, Baron B. The role of TNF-α in rheumatoid arthritis: a focus on regulatory T cells. J Clin Transl Res. 2016;2:84–90. [PMC free article] [PubMed] [Google Scholar]
- [47].Peng X, Wang Q, Li W, et al. Comprehensive overview of microRNA function in rheumatoid arthritis. Bone Res. 2023;11:8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [48].Pandis I, Ospelt C, Karagianni N, et al. Identification of microRNA-221/222 and microRNA-323-3p association with rheumatoid arthritis via predictions using the human tumour necrosis factor transgenic mouse model. Ann Rheum Dis. 2012;71:1716–23. [DOI] [PubMed] [Google Scholar]
- [49].Khan NM, Haqqi TM. Epigenetics in osteoarthritis: potential of HDAC inhibitors as therapeutics. Pharmacol Res. 2018;128:73–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [50].Chen W, Chen L, Zhang Z, et al. MicroRNA-455-3p modulates cartilage development and degeneration through modification of histone H3 acetylation. Biochim Biophys Acta. 2016;1863:2881–91. [DOI] [PubMed] [Google Scholar]
- [51].Kim HS, Shen Q, Nam SW. Histone deacetylases and their regulatory microRNAs in hepatocarcinogenesis. J Korean Med Sci. 2015;30:1375–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [52].Singhal R, Mitta SR, Das NK, et al. HIF-2α activation potentiates oxidative cell death in colorectal cancers by increasing cellular iron. J Clin Investig. 2021;131:e143691. [DOI] [PMC free article] [PubMed] [Google Scholar]