Abstract
Modern polymer science suffers from the curse of multidimensionality. The large chemical space imposed by including combinations of monomers into a statistical copolymer overwhelms polymer synthesis and characterization technology and limits the ability to systematically study structure−property relationships. To tackle this challenge in the context of 19F magnetic resonance imaging (MRI) agents, we pursued a computer-guided materials discovery approach that combines synergistic innovations in automated flow synthesis and machine learning (ML) method development. A software-controlled, continuous polymer synthesis platform was developed to enable iterative experimental−computational cycles that resulted in the synthesis of 397 unique copolymer compositions within a sixvariable compositional space. The nonintuitive design criteria identified by ML, which were accomplished by exploring <0.9% of the overall compositional space, lead to the identification of >10 copolymer compositions that outperformed state-of-the-art materials.
Graphical Abstract

INTRODUCTION
Next-generation challenges in soft materials will require the discovery of polymers that perform multiple functions simultaneously. Copolymerization, where two or more building blocks are included in a single material, is an effective strategy to achieve differentiated properties.1 The inclusion of multiple unique building blocks into a copolymer, however, often has interdependent effects on reactivity, structure, and properties, making the a priori prediction of material function for complex copolymers challenging.2 Additionally, traditional synthetic technology in polymer science is iterative, labor-intensive, capricious, and low-throughput, making rapid experimentation, purification, and analysis impractical.3 Polymer science therefore remains plagued by the “curse of multidimensionality”, where even simple combinations of monomeric building blocks lead to a high-dimensional chemical space that is too vast to explore without implementing limiting assumptions.
Computer-guided materials discovery has been shown to be an effective approach to detect meaningful patterns in data sets of high dimensionality, thus allowing the prediction of structure−function relationships while only requiring a small percentage of the chemical space to be experimentally explored.4 One such approach is the use of computer simulations to define molecular structure−property relationships and target specific polymer compositions.5 However, human intuition defines the inputs of these computational models, which restricts the diversity of the data set due to inherent biases or limitations in knowledge. Furthermore, computational and experimental cycles are often physically and temporally separated, which slows the speed of chemical structure optimization to achieve desired performance.
The use of artificial intelligence (AI) for computer-guided materials discovery is an alternative approach that holds promise for dramatically accelerating the optimization of polymer structure−property relationships, with the opportunity to close the loop between computational and experimental components of the materials discovery pipeline.6,7 Recent advances in both automated synthetic platforms and machine learning (ML) methods development have enabled experimental systems that provide high-quality training data to improve ML models and, at times, are driven by ML recommendations in the areas of small-molecule synthesis8–16 and nanomaterial synthesis.17–22 In a recent example, the Doyle group demonstrated a Bayesian optimization platform that allows chemists to iterate between experimentation and ML within their standard synthetic workflows, thus providing open-source tools to increase the efficiency of chemical synthesis.23
Despite impressive advances using ML for small-molecule and nanomaterial synthesis, the integration of synthesis, characterization, and ML in polymer science has lagged behind.24–28 A critical challenge is sourcing high-quality experimental data to train predictive ML models, which often requires a combination of high-throughput synthesis, purification, and characterization methods that remain underdeveloped in polymer chemistry.29,30 As an added challenge, the field of polymer science lacks a standardized data schema for reporting polymer structure and properties that contextualize the underlying measurement and its output.31–33 Applications of ML in polymer science have therefore mostly been isolated to a small subset of commonly reported properties34–36 or relied on legacy data collected within a single research group.37–39 In a seminal report, Pruksawan et al. demonstrated the utility of the synthesis and property evaluation of 42 epoxy adhesive samples and employed ML to generate a predictive model that accurately described the performance of 256 possible formulations.40 In complementary work, Reineke and coworkers made a multiparametric library of 43 copolymers to serve as gene-delivery vectors,30 and separately, Appel and coworkers synthesized a combinatorial library of 172 acrylamide hydrogels as antibiofouling coatings.29 Both groups used random forest classifiers to identify nonintuitive descriptors that led to high performance. In these examples, however, the reliance on labor-intensive batch synthesis or formulation, the need to probe a large percentage (>15%) of the compositional space to optimize an accurate model, and the lack of iterative experimental−computational cycles limit the translation of this approach to more complex problems in high-dimensional copolymer compositional space.
We identified the discovery of high-contrast 19F magnetic resonance imaging (MRI) agents as a challenge in need of a ML-driven discovery approach. 19F MRI is a high-contrast biomedical imaging modality with the potential to track cellular transport and quantitate oxygenation with spatiotemporal resolution.41–45 Synthetic polymers represent attractive 19F MRI agents due to their potential for multivalent displays of 19F atoms and their synthetic modularity. Despite decades of effort reporting hundreds of copolymer 19F MRI agents, challenges persist in the development of 19F MRI agents that are both water-soluble and contain enough fluorine nuclei to be visualized on clinical 3 T MRI scanners.46
Herein we developed automated tools to interface copolymer synthesis and characterization with ML, which enabled iterative feedback through numerous experimental−computational cycles. The nuanced structure−property trends uncovered through this ML-guided materials discovery approach motived us to reconsider the dogma that the 19F solution concentration is directly related to the signal intensity in the 19F MRI measurement and propose nonintuitive design elements that are critical to consider for next-generation 19F MRI agents. This combination of continuous-flow chemistry and ML represents a powerful approach to tackle high-dimensional challenges in polymer science, where the large number of interdependent variables makes structure−property relationships difficult to predict or model.
RESULTS AND DISCUSSION
Development of ML Approach.
Given the limited initial 19F MRI data set available to build a predictive ML model, we envisioned developing a platform that iterates between computational (i.e., software) and experimental (i.e., hardware) components to efficiently screen for high-performing 19F MRI agents (Figure 1). The choice of a computational approach proved to be challenging due to the conflicting performance criteria inherent to copolymers used as 19F MRI agents. The necessity for an imaging agent to possess a high density of hydrophobic fluorinated comonomers while also remaining water-soluble demands the simultaneous optimization of multiple objectives along a trade-off curve, otherwise known as a Pareto front.47
Figure 1.
Active-learning-guided discovery of copolymer 19F MRI agents relies on rapid feedback between computational and experimental nodes.
The capability to iterate between experiments and ML allowed us to leverage active learning (AL) for multiobjective optimization.48 AL is a semisupervised form of ML where the algorithm efficiently explores the chemical space by selecting maximally informative materials to evaluate through experimentation (i.e., exploration) or more narrowly identifies high-performing compositions (i.e., exploitation).48 Our attempt to implement AL, however, exposed a weakness in the ML pipeline: Almost all ML models are designed and tuned by hand, and there is no single ML model that works for all applications. Typically, a manuscript will report only the successful application of a particular method, but tuning these methods to a particular application inherently introduces model and sample selection biases. This leads to researchers selecting suboptimal models or investing a significant amount of time into model tuning for a particular application.
We hypothesized that an automated ML (AutoML) approach would streamline model development and allow a nonexpert to search for a high-quality ML model independently.49 A variety of approaches to AutoML have been developed recently that involve selecting an optimal ML algorithm, preprocessing input features, and selecting hyperparameters, including an extension of the scikit-learn library with meta-learning and ensemble construction50 and the use of genetic algorithms in the TPOT library.51 Our AutoML composite approach screens several validated ML learning methods including the XGBoost,53 and NGBoost,54 Gaussian Process, Random Forest, and Linear and Logistic Regression methods, as implemented in scikit-learn.52 Here we consider only the supervised learning setting. The models used are limited to fixed-length vectorial representations of the polymer composition and analytical characterization data. (See the SI for a technical description.)
The overall design of AutoML was expressed as a black-box optimization problem to optimize two objectives simultaneously. In such an AutoML workflow, the user provides data; then, the AutoML library autonomously samples the aforementioned methods, selects the optimal ML model parameters for the data set, and makes decisions about subsequent exploratory or exploitative experiments in real time. Overall, the automated ML cycle consists of four steps that can operate in a closed-loop fashion with the appropriate synthetic hardware: (i) train a proxy AutoML model to optimize for a given set of objectives on an initial data set; (ii) use the model constructed by AutoML to virtually screen the copolymer compositional space; (iii) select a subset of copolymer compositions that would increase the accuracy of the model; and (iv) perform the synthesis and experimental measurement of selected polymers and use these data to update the ML model(s).
Design and Implementation of Automated Continuous-Flow Copolymer Synthesis.
We identified continuous-flow chemistry as an ideal experimental platform for the iterative synthesis of novel copolymer materials due to its ease of automation, reproducible control of reaction conditions, potential for closed-looped optimization between synthesis and analysis, and simple translation to manufacturing scales.55–62 The majority of the previously reported high-throughput copolymer synthesis systems polymerize one sample at a time and thus require extremely short reaction times to achieve a high sample throughput. For example, Hedrick and coworkers developed a flow reactor capable of synthesizing 100 unique block copolymers in 8 min, but the technology relied on ringopening polymerizations with reaction times of <1 s.63 Unfortunately, the controlled radical polymerization techniques traditionally employed to make copolymer 19F MRI agents suffer from reaction kinetics that are orders of magnitude slower than this example,45,64 which required the design of a novel high-throughput flow reactor.
To combat the challenge of slow copolymerization kinetics, we designed a more general flow platform capable of polymerizing multiple samples simultaneously. We identified droplet flow as an enabling approach to achieving a high sample throughput regardless of the polymerization kinetics. Droplet-based flow systems manipulate discrete volumes of reaction mixtures that are separated by an immiscible inert fluid.56,65–67 As our lab previously demonstrated, polymerization in droplets reduces the residence time distribution and improves the control of polymer composition, molar mass, and dispersity.61 For this application, nitrogen gas was used as the immiscible fluid between large reaction droplets, or slugs. A custom liquid handler was fabricated that allowed precise formulations to be loaded into a sample loop before being injected into the heated reactor. Through experimentation, we discovered that a wash slug of dimethylformamide (DMF) was required between sequential reaction slugs to prevent crosscontamination. A simplified schematic of the flow reactor is shown in Figure 2, and a more detailed version is described in the (Supporting Information Figure S5).
Figure 2.
Automated continuous-flow reactor development. (A) Simplified reactor schematic. (B) Droplet-flow reactor. (C) Rapid prototyping enabled by 3-D printed hardware and a modular electronics platform. (D) Demonstration of droplet technology using colored dyes. Videos are provided as supplementary files.
To create a modular platform that could access a broad compositional space in a user-friendly fashion, we fully automated the reagent selection, comonomer formulation, slug injection sequence, and sample collection using custom hardware and software. (See the Supplementary Videos for visualization.) An Arduino microcontroller was chosen as the electronics platform to control the flow system, and integration of all individual components with LabVIEW software allowed full automation of complex reaction sequences. The use of readily accessible electronics and 3-D printed parts allowed for the rapid design and prototyping of hardware components optimized for high-throughput copolymer synthesis. The custom liquid handler enabled the efficient and precise formulation of reaction slugs containing radical initiators, up to six different comonomers, and a compatible RAFT chain-transfer agent. To achieve droplet flow, each 300 μL reaction slug was confined on both sides by two nitrogen slugs and loaded into the sample loop of a two-position, six-port switching valve. At a predetermined point during the automated reaction sequence, these slugs were injected into the flow stream. The heated reactor consisted of tubing embedded in a machined aluminum block, with a heating element and thermocouple to provide accurate temperature regulation. Upon exiting the reactor, samples were collected in a 30-slot sample collection carousal. Rotation of the carousal was triggered by a refractive index (RI) detector immediately upstream, which tracked the number of eluted slugs by monitoring the change in RI between reaction slugs and nitrogen slugs.
The entire droplet flow system occupied a small footprint (43 cm × 46 cm × 96 cm) and was fully touch-screen-enabled to allow use by nonexperts. LabVIEW software controlling the flow reactor was capable of extracting relevant reaction parameters from comma separated value (CSV) files generated by the user or the AL algorithm. The combination of these efforts afforded an easy-to-use system capable of synthesizing a new copolymer composition every 2 min, allowing the synthesis of 30 unique copolymers in 2 h using only 12 mL of reaction solution. The typical workflow for synthesis, purification, and analysis was optimized to evaluate batches of 30 unique 19F MRI contrast agents. This workflow consisted of (i) the preparation of reagent stock solutions, (ii) automated synthesis of copolymers in flow, (iii) transfer of samples to gravity-fed size exclusion chromatography (SEC) columns, (iv) drying of polymer containing fractions, and (v) 19F NMR analysis and data workup. Not including drying times, this method allowed the evaluation of 30 samples in a single 8 h workday. To evaluate the reproducibility of this workflow, 20 representative compositions were run in triplicate, and the tabulated results can be found in Table S4. The accumulative errors across all steps of the workflow resulted in a modest average standard deviation of four signal-to-noise ratio (SNR) units across the studied copolymer samples. Additionally, the automated and modular flow platform described herein will enable simple expansion to accommodate new chemistries and reaction sequences.
Synthesis and Characterization of Copolymer 19F MRI Agents.
The inherent tension between having a high density of hydrophobic fluorine atoms and maintaining water solubility for 19F MRI agents has been solved in previous literature through the statistical copolymerization of partially fluorinated monomers such as trifluoroethyl acrylate (TFEA) with hydrophilic monomers such as poly(ethylene-glycol) acrylate (PEGA) to afford 19F MRI agents with moderate sensitivity.64,68–77 These copolymers provided adequate materials for preclinical studies on high-resolution spectrometers but did not demonstrate the required sensitivity to be used on 3 T clinical-strength MRI instruments at realistic concentrations. Previous work has identified that a number of different hydrophilic and partially fluorinated comonomers can improve the 19F MRI sensitivity in isolated examples, but an understanding of how the polymer composition relates to the material performance is lacking.64,69,73–77 When attempting to data-mine literature examples of 19F MRI agents to apply ML methods, we encountered challenges in standardizing the SNR for 19F MRI agents across studies due to differences in the magnetic field strength, pulse sequence, reagent concentration, and reporting procedure.
To significantly advance the state of the art, we hypothesized that a systematic evaluation of the most promising fluorinated and solubilizing comonomers would provide a more comprehensive understanding of the structure−property relationships that dictate the performance of 19F MRI agents (Figure 3A). The partially fluorinated acrylic comonomers chosen include TFEA and the more densely fluorinated hexafluorooxy-ethyl acrylate and nonafluorooxy-ethyl acrylate (HexaFOEA and NonaFOEA, respectively). The water-solubilizing acrylic comonomers include PEGA and 2-(methylsulfiyl)ethyl acrylate and hydroxyethyl acrylate (MSEA and HEA, respectively).
Figure 3.
Synthesis of multicomponent copolymers as 19F MRI agents. (A) Six comonomers were chosen to synthesize statistical copolymers while balancing 19F content and water solubility. (B) 19F NMR spectra of 30 representative copolymers demonstrating the diversity of resonances arising from different copolymer compositions.
Copolymerization using thermally initiated reversible addition−fragmentation chain transfer (RAFT) was selected due to its tolerance of diverse functionality and its ability to provide control over the copolymer molar mass (Mn) and dispersity (Đ). A similar degree of polymerization (DP) was targeted for each copolymer 19F MRI agent to decrease the potential for chain-length effects to influence the material performance. A common challenge for high-throughput radical polymerization is the need to rigorously remove oxygen from each sample and thus limit the batch-to-batch variability.78–105 We took inspiration from the “polymerizing through” approach106–110 to oxygen-tolerant RAFT polymerization, where a large flux of radicals is introduced at the start of the reaction to consume dissolved oxygen, and a smaller and consistent radical flux subsequently provides controlled polymerization. A high radical flux was achieved through the addition of a low concentration of V-70, an azo radical initiator with a short half life at the reaction temperatures, in addition to the more typical radical initiator azobisisobutyronitrile (AIBN). (See the SI for the detailed copolymerization methodology.) The monomer conversation was between 60 and 80% for the conditions studied, with the range attributed to differences in the comonomer composition.
Following copolymer synthesis, we recognized that copolymer purification presented a potential bottleneck to the exploration of large compositional space. Precipitation was not broadly applicable because copolymers of different compositions possessed different solubilities, and dialysis was impractical in a high-throughput fashion. A purification procedure using aqueous SEC gravity desalting columns proved to be ideal.111 The workflow included taking polymer samples directly from the reaction and eluting them with a known amount of deionized water through the SEC column. This approach allowed for multiple polymers to be purified in parallel and excluded water insoluble copolymers that precipitated within the resin. The final isolated yields of the materials were therefore dependent on both the monomer conversion and the solubility.
The figure-of-merit chosen to evaluate the performance of multicomponent polymers as 19F MRI agents was the SNR ratio taken from 1D 19F experiments on a 400 MHz NMR. These SNR values for 19F NMR correlate with the 19F MRI sensitivity, with small variations that result from differences in the pulse sequences and probe design.112 Copolymer samples were diluted with phosphate-buffered saline (PBS) solution/D2O (90:10 v/v) at a concentration of 20 mg/mL. Each copolymer composition exhibited unique 19F resonances resulting from a combination of factors that included the copolymer composition and the copolymer solution conformation (Figure 3B). The unique chemical environment resulting from the copolymer solution structure influenced the 19F chemical shift values (δ) and spin−lattice relaxation times (T1), with shorter 19F T1 increasing the signal intensity observed during T1-weighted MRI sequences. Furthermore, polymers with high fluorinated comonomer content demonstrated significant peak broadening as a result of the short spin−spin relaxation (T2). The interdependent properties that contribute to the SNR value of multicomponent copolymers therefore are difficult to predict a priori and require experimental validation.
Implementation of ML-Guided Discovery of Champion 19F MRI Agents.
Whereas the six comonomers in Figure 2A established the compositional space for exploration, we sought to establish a number of boundary conditions to define the specific copolymer structures for synthesis and evaluation. First, the individual comonomer compositions would change by increments of 5%. Smaller shifts in composition were at the limit of our liquid handling technology. Second, the comonomers and chain-transfer agent (CTA) chosen produced only linear polymers, thus removing the potential for polymer topology to influence performance. Third, all polymerizations were assumed to be statistical, with the initial comonomer stoichiometry being the assumed stoichiometry incorporated into the polymer. Given these boundary conditions, the experimental exploration of six unique monomers revealed 47 854 possible copolymer compositions to explore. In every batch, the AutoML algorithm selected the optimal features from a range of representations, which included a vector of monomer fractions of length 6 for each composition (one-hot encoding; the sum of all 6 fractions of polymer composition equals 1 for each composition), the fraction of fluorine, and various constitutional descriptors from RDKit. (See the SI for a technical description.)
Typical ML approaches require a large portion of the overall chemical space to be explored (>5%) before converging onto an accurate model, which, in our case, would have required the synthesis of an impractical number (>2200) of individual copolymers. We hypothesized that because the polymer composition is a continuous variable whose boundary conditions can be adjusted, a hierarchical sampling of compositional space would be a more efficient approach. Therefore, the initial screening focused on a coarse compositional space where individual comonomer compositions could change in only 10% intervals, shrinking the explorable compositional space from 47 854 to 2486 possible copolymer combinations. We hypothesized that this would allow model development with a smaller library of initial data points, and, as the model performance improved, a switch to the larger compositional space of 47 854 potential copolymers with a 5% change in comonomer composition (fine compositional space) would be feasible. Furthermore, we required an approach that not only predicted the SNR for 19F MRI, but also overlaid that model with one that predicted the water solubility. Therefore, both properties are used for multiobjective optimization.
Our AL experiments were initialized from data containing 157 copolymer compositions, which represents 6.3% of the coarse compositional space. This initial data set was gathered from materials made previously in our lab,45 which targeted high-performing imaging agents and samples made during instrument optimization (Figure 4A). To simultaneously optimize for the water solubility and the SNR, we used two separate ML models: The first was a classification model that predicted whether a sample would be water-soluble, and the second was a regression model that predicted 19F NMR SNR values. The experimental solubility was determined qualitatively. Materials was considered soluble if the sample was fully transparent at a concentration of 20 mg/mL in phosphate buffer. Therefore, the solubility was treated similarly as a binary classification in the ML model. To balance exploration and exploitation, we used spherical exclusion clustering113 to reduce the number of candidate compositions to an experimental batch size of 30 while ensuring reasonable composition diversity. After running the system for two AL cycles, the mean absolute error (MAE) decreased to <8 SNR units and stabilized. Given the increasingly accurate model performance in the coarse compositional space, we sought to exploit the model to select high-performing materials in the larger compositional space of 47 854 potential copolymers (Figure 4, batch 3). This initial effort, which included data from only 0.45% of the fine compositional space, led to an experimental batch of 30 copolymers that were all insoluble in water.
Figure 4.
(A) Data acquisition and ML model performance throughout the AL steps. The top panel shows the MAE error for 19F SNR ML models; confidence intervals are obtained through 10 shuffle splits. The bottom panel shows actual data points and corresponding box plots for the data distribution. The insoluble materials are depicted as gray points at the bottom, and the total number of molecules per batch is equal to 30 in batches 1−8. The experiments using the coarse compositional space (10% step) are highlighted in purple, and those using the fine compositional space (5% step) are highlighted in white. The batches run targeting exploitation are colored in pink. (B) Uniform manifold approximation and projection (UMAP) of representative batches. Colored circles represent experimentally validated water-soluble structures and gray circles represent insoluble samples. The background is color coded by the 19F SNR ML prediction of the given batch.
As evidenced by the poor performance of batch 3, moving from a coarse (10% interval) to a fine (5% interval) compositional space required more experimental data points. For batch 4, we synthesized 30 copolymers to target exploration of the fine compositional space. The SNR predictions remained quite accurate, but the multiobjective optimization that included the solubility required a significant number of experimental results to converge. As shown in Figure 4A, batches 3 and 4 resulted in the synthesis of many insoluble samples as the algorithm worked to define solubility parameters. Three additional rounds of exploration (batches 5−7) improved the predictive power and resulted in an ML model that could accurately predict the SNR values of soluble copolymers with a mean absolute error of <7 SNR units. Given that the experimental error of the automated synthesis system is 4 SNR units, the model reached a high value of accuracy. A selection of the samples identified by the algorithm in each batch and experimentally produced by the flow system is shown in Figure 4B (full representation in Figure S2).
To study the influence of the molecular weight on the copolymer 19F MRI performance, we analyzed a large subset of the data by SEC, and the polymer Mn and Đ were calculated. The Mn values were used as an input for the ML model (Figure S6), but no statistically significant effect on predicting the material performance was identified. Given that all polymerizations targeted the same DP, the modest differences in the Mn did not influence the polymer 19F MRI agent sensitivity. Therefore, the design criteria identified herein could be translated to polymers of higher or lower molecular weight to tune application-specific properties such as the biodistribution, metabolic fate, or pharmacokinetics without significantly influencing the 19F MRI performance.
Providing the increasingly accurate model predictions, we initiated an exploitation AL cycle by having the model greedily select 19F MRI agents with potentially high performance (batch 8). The batch of 30 samples included 15 copolymers with an SNR of >80 and 2 that exceeded the values of the highest performing copolymers reported in our previous study.45 The batch also included 11 samples that were insoluble, which represented a significant improvement over the attempt at exploitation prior to model development in the larger compositional space. Overall, this hierarchical AL model development workflow produced a robust model to predict the structure−performance relationships of 47 854 potential copolymer 19F MRI agents while experimentally exploring <0.9% of the compositional space (397 copolymers).
Analysis of Compositional Space and Structure−Property−Performance Relationships.
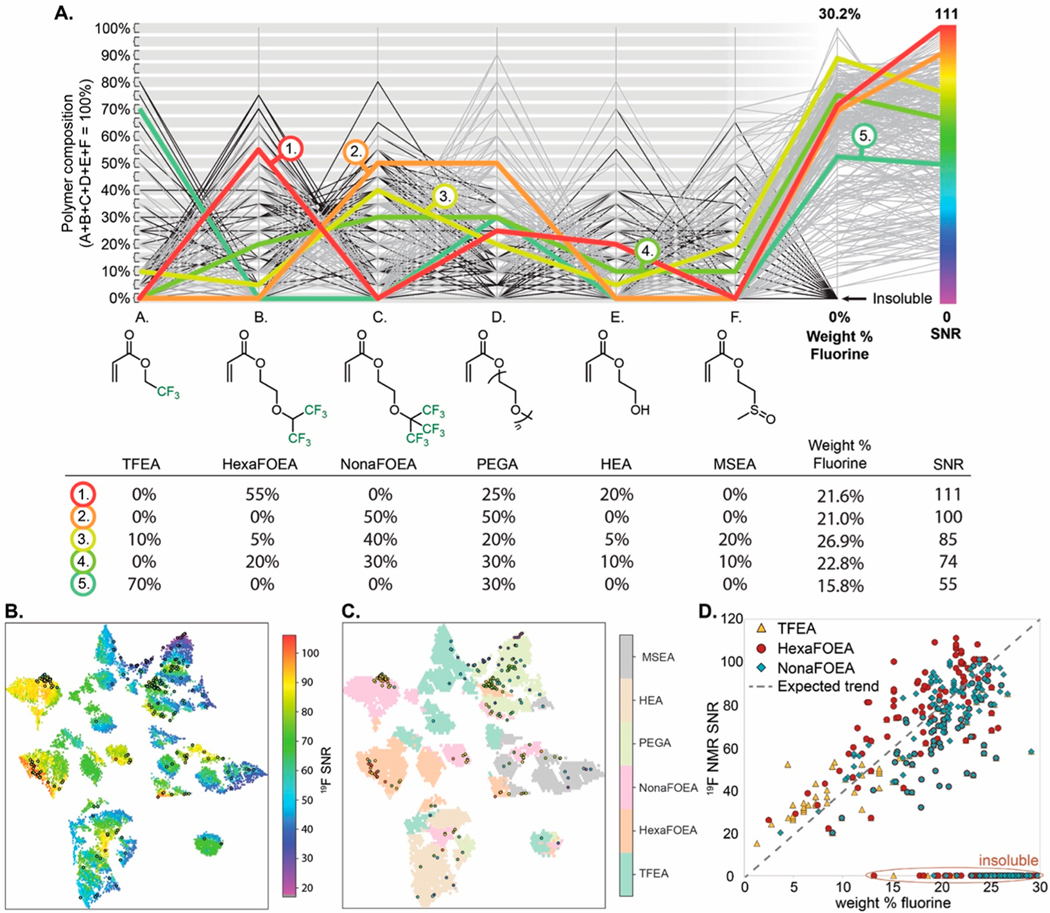
The central dogma in this field is that the copolymers with the higher 19F content have a higher SNR in 19F MRI experiments.45,64,74,76,77 Considering the three partially fluorinated monomers chosen for this study (Figure 3A) and our previous observations,45 we hypothesized that copolymers made with NonaFOEA would have the highest sensitivity given that NonaFOEA has the highest weight percent (wt %) 19F. The parallel coordinate diagram in Figure 5A collects data for the copolymer composition, wt % 19F, and SNR for each copolymer produced in this study. An initial evaluation of this data demonstrated an unexpected but clear discontinuity between the wt % 19F and the SNR. Comparing a few representative copolymers is instructive to describe these effects in more detail. Copolymer 1, which was identified by the ML model during the AL exploitation step (batch 8), represents the highest performing copolymer (SNR of 111). The sample, along with >80% of the samples that achieved an SNR of >100, had HexaFOEA as the fluorine-containing comonomer. Copolymer 1 contained only 21.6 wt % fluorine yet outperformed the dozens of copolymer samples that contained a higher fluorine density. Copolymer 1 also contained more than one solubilizing comonomer, which is a trend we observed for most high-performing copolymers and has not been demonstrated in previous studies.
Figure 5.
Visualization of experimental 19F compositional space. (A) Parallel coordinate diagram of the 397 samples that describes copolymer composition and performance, with six representative compositions colored and shown in table format. (B) UMAP projection of the copolymer compositional space with the 19F SNR ML prediction color coded. Circled samples represent experimentally validated water-soluble structures. (C) UMAP projection of the copolymer compositional space with the major comonomer component color coded. Circled samples represent experimentally validated water-soluble structures. (D) Plot demonstrating the relationship between the wt % 19F in soluble copolymers and the 19F NMR SNR.
Comparing 1 to other copolymers provides comparative structure−property information. Copolymer 2 was synthesized in batch 0 and contains approximately the same wt % fluorine at 1, but the use of NonaFOEA instead of HexaFOEA and the lack of solubilizing monomers beyond PEGA results in a lower SNR of 100. Copolymers 3 and 4 both have higher wt % fluorine than 1, but the higher fluorine density is the result of a combination of partially fluorinated monomers in the copolymers, which limits the SNR of any one 19F resonance. Lastly, copolymer 5 demonstrates the limitations of TFEA to achieve a high SNR despite its high mol % incorporation.
Figure 5B shows a 2-D UMAP of the complete compositional space of all possible copolymers.114 The UMAP estimates the topology of the high-dimensional data and uses this information to construct a low-dimensional representation that preserves the relationships present in the data. The resulting projections show strong local clustering. A consequence of the topology of this low dimensional data representation is that the distance between data points in the UMAP projection is difficult to interpret. In the context of polymer composition, UMAP finds the nearest neighbors for each polymer and creates a low-dimensional representation that groups closely related compositions together in space. The computationally derived SNR values are represented by the color gradient in the image, whereas the soluble copolymers samples that were produced experimentally are represented as circular icons (UMAP including soluble and insoluble samples in Figure S3). Most striking in this image are the many disconnected “islands” within the chemical space where high SNR copolymers are located. Considering this plot in tandem with the representation of the chemical composition (Figure 5C and Figure S4) demonstrates that high-performing copolymers predominately contain HexaFOEA and NonaFOEA, but identifying a pattern for the nonfluorinated comonomers that leads to the high SNR is nonintuitive.
A visualization of the unexpected structure−property trends can be seen in Figure 5D. The occurrence of a large population of copolymers containing HexaFOEA above the expected linear trend between the wt % 19F and the SNR demonstrates its privileged selection as a fluorinated comonomer. Additionally, the complexity of structure−solubility relationships is evident by the many insoluble copolymer compositions that have the same wt % 19F as high-performing materials. Regression analysis (Figure S11) of the soluble copolymers provides quantitative evidence of the privileged nature of HexaFOEA-containing copolymers. The R2 values are 0.67 and 0.39 for TFEA-containing copolymers and HexaFOEA-containing copolymers, respectively, revealing how the linear relationships between the wt % 19F and the SNR is less evident for HexaFOEA. These results underscore the importance of the high-throughput synthesis coupled to the ML-guided materials design, especially for identifying materials that display nonintuitive structure−property relationships.
Contextualizing the ML model outputs by comparing both the SNR and solubility predictions reveals the complexity of identifying high-performing 19F MRI agents (Figure 6A). The “islands” where the high SNR is predicted overlay quite closely areas in which few copolymers are predicted to be soluble, which is expected due to the intrinsic relationships between the fluorine density and the hydrophobicity. A more detailed visualization in Figure 6B,C shows a zoom-in on the western region of the chemical space, with both the SNR and the water solubility prediction shown. The solubility prediction clearly shows the complex geography of the chemical space where the high SNR and the water solubility coincide. Whereas this UMAP representation communicates the ability of the ML algorithm to differentiate subtle structure−property relationships visually, the representation is only qualitative. The ML model itself, working in a 6-D space, provides specific predictions of the material performance and the solubility to identify otherwise difficult-to-predict materials. Without the aid of ML, discovering the ideal combination of comonomers to yield a soluble copolymer is this region is unlikely. We hypothesize that the nonintuitive relationship between the polymer composition and the solubility is due to the subtle influence that sequence and comonomer identity can have on the solution conformation of a flexible polymer chain and thus the functionality present on the exterior of the globule that must interact with water to maintain solubility.
Figure 6.
(A) UMAP representation of the copolymer compositional space with ML algorithm predictions for water solubility. Colored circles represent experimentally validated water-soluble copolymers and gray circles represent insoluble copolymers. (B,C) Zoomed-in portion of the copolymer compositional space where the highest performing copolymers resided. Comparison of the water solubility (B) and SNR (C) models within this area of interest. (D) 19F MRI analysis of eight representative copolymer samples, including SNR values and phantom magnetic resonance images.
Eight representative copolymers with a range of SNR values and compositions were selected for the analysis using application-specific techniques. These included the evaluation of 19F NMR T1 and T2 relaxation times and MRI imaging using a T1-weighted fast low-angle shot (FLASH) pulse sequence115,116 Other imaging sequences can also be used for 19F MRI, and the power of our approach is that the ML algorithm can be easily modified to optimize for output variables that provide high performance for a particular imaging sequence. (Figure 6D and Tables S2 and S3). These MRI studies confirmed a number of observations that the ML algorithm identified. First, copolymers that contained three or more comonomers generally outperformed two-component copolymers, which we hypothesize is a result of the difficulty for fluorinated moieties to segregate into dense phases within a compositionally complex polymer globule. Second, although previous work117 set a detection limit of 126 mM 19F for visualization on a 3 T clinical MRI scanner, we demonstrate that the concentration of 19F alone is not an accurate predictor of the 19F MRI sensitivity. For example, the highest performing HexaFOEA and NonaFOEA multicomponent copolymers, containing a concentration of ~240 mM 19F and 230 mM 19F, respectively, both displayed nearly 1.4 times higher 19F MRI SNR than the highest sensitivity previously reported, which was assessed using the same imaging sequence on the same scanner and used NonaFOEA at a concentration of 220 mM 19F.45 Therefore, the increase in 19F MRI SNR cannot be solely attributed to an increase in the concentration of fluorine nuclei and further illustrates the interdependent nature of the variables responsible for 19F MRI sensitivity. Lastly, both the HexaFOEA and NonaFOEA copolymers reached a limit in the achievable 19F MRI SNR at 240 mM 19F, which could represent the threshold of fluorine concentration before the water insolubility and detrimental 19F T2 broadening impact the MRI sensitivity.
CONCLUSIONS
We demonstrated an ML-guided materials discovery approach that combines synergistic innovations in automated flow synthesis and ML method development. Iterative feedback between polymer synthesis, characterization, and ML, combined with a hierarchical exploration of compositional space, enabled the development of an ML algorithm that accurately predicts structure−property relationships while requiring only <0.9% of the compositional space (397 copolymers) to be experimentally explored. Our approach facilitated the discovery of a number of copolymer 19F MRI agents with imaging sensitivities higher than those of previously reported materials. Additionally, the trends uncovered herein have conclusively demonstrated that 19F concentration is not directly related to the signal intensity in the 19F MRI measurement. The nonintuitive material design elements for 19F MRI agents identified in our study, including the privileged function of HexaFOEA as a fluorinated comonomer, the benefits of using multiple solubilizing comonomers in a single imaging agent, and the observation that wt % fluorine is not directly related to the SNR, are critical to consider in the search for next-generation 19F MRI agents.
Materials discovery typically relies on human intuition, which suffers from inherent biases and limitations in knowledge. As this study demonstrates, the continued integration of software-enabled high-throughput polymer synthesis and ML represents a powerful approach to accelerate materials discovery, especially in areas of polymer science where the large number of interdependent variables makes structure−property relationships difficult to predict or model.
Supplementary Material
ACKNOWLEDGMENTS
F.A.L. acknowledges funding from the Beckman Foundation under the Beckman Young Investigator grant and award R35-GM142666 from the National Institute of General Medical Sciences. The UNC Department of Chemistry’s NMR Core Laboratory provided expertise and instrumentation that enabled this study with support from the National Science Foundation (CHE-1828183 and CHE-0922858). O.I. acknowledges support from NSF CHE-1802789 and CHE-2041108. This work was performed, in part, at the Center for Integrated Nanotechnologies, an Office of Science User Facility operated for the U.S. Department of Energy (DOE) Office of Science. We acknowledge Extreme Science and Engineering Discovery Environment (XSEDE) award DMR110088, which is supported by NSF grant number ACI-1053575. This research is part of the Frontera computing project at the Texas Advanced Computing Center. Frontera is made possible by the National Science Foundation award OAC-1818253.
Footnotes
ASSOCIATED CONTENT
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/jacs.1c08181.
Synthesis, experimental details, computational methods, characterization data, and supplementary figures (PDF)
Polymer composition (XLSX)
Animation describing how each component of the droplet-flow system is integrated (MP4)
Video of the liquid handling carousel and sample loop (MP4)
Video of reaction slugs moving through the heated reactor (MP4)
Video of the individual collection of each slug after reaction (MP4)
Complete contact information is available at: https://pubs.acs.org/10.1021/jacs.1c08181
The authors declare no competing financial interest.
Contributor Information
Marcus Reis, Department of Chemistry, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina 27599, United States.
Filipp Gusev, Department of Chemistry, Mellon College of Science and Computational Biology Department, School of Computer Science, Carnegie Mellon University, Pittsburgh, Pennsylvania 15213, United States.
Nicholas G. Taylor, Department of Chemistry, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina 27599, United States.
Sang Hun Chung, Department of Biomedical Engineering, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina 27599, United States.
Matthew D. Verber, Department of Chemistry, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina 27599, United States
Yueh Z. Lee, Department of Radiology, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina 27599, United States
Olexandr Isayev, Department of Chemistry, Mellon College of Science and Computational Biology Department, School of Computer Science, Carnegie Mellon University, Pittsburgh, Pennsylvania 15213, United States.
Frank A. Leibfarth, Department of Chemistry, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina 27599, United States
REFERENCES
- (1).Moad G; Solomon DH Copolymerization. In The Chemistry of Radical Polymerization 2005, 333–412.
- (2).Li Y; Abberton BC; Kröger M; Liu WK Challenges in Multiscale Modeling of Polymer Dynamics. Polymers 2013, 5, 751–832. [Google Scholar]
- (3).Oliver S; Zhao L; Gormley AJ; Chapman R; Boyer C. Living in the Fast Lane—High Throughput Controlled/Living Radical Polymerization. Macromolecules 2019, 52 (1), 3–23. [Google Scholar]
- (4).Himanen L; Geurts A; Foster AS; Rinke P. Data-Driven Materials Science: Status, Challenges, and Perspectives. Adv. Sci 2019, 6 (21), 1900808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (5).Gartner TE; Jayaraman A. Modeling and Simulations of Polymers: A Roadmap. Macromolecules 2019, 52 (3), 755–786. [Google Scholar]
- (6).Dimitrov T; Kreisbeck C; Becker JS; Aspuru-Guzik A; Saikin SK Autonomous Molecular Design: Then and Now. ACS Appl. Mater. Interfaces 2019, 11 (28), 24825–24836. [DOI] [PubMed] [Google Scholar]
- (7).Winicov H; Schainbaum J; Buckley J; Longino G; Hill J; Berkoff CE Chemical Process Optimization by Computer - a Self-Directed Chemical Synthesis System. Anal. Chim. Acta 1978, 103, 469–476. [Google Scholar]
- (8).Coley CW; Thomas DA; Lummiss JAM; Jaworski JN; Breen CP; Schultz V; Hart T; Fishman JS; Rogers L; Gao H. A Robotic Platform for Flow Synthesis of Organic Compounds Informed by AI Planning. Science 2019, 365 (6453), eaax1566. [DOI] [PubMed] [Google Scholar]
- (9).Granda JM; Donina L; Dragone V; Long DL; Cronin L. Controlling an Organic Synthesis Robot with Machine Learning to Search for New Reactivity. Nature 2018, 559 (7714), 377–381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (10).Ahneman DT; Estrada JG; Lin S; Dreher SD; Doyle AG Predicting Reaction Performance in C−N Cross-Coupling Using Machine Learning. Science 2018, 360 (6385), 186–190. [DOI] [PubMed] [Google Scholar]
- (11).Trobe M; Burke MD The Molecular Industrial Revolution: Automated Synthesis of Small Molecules. Angew. Chem., Int. Ed 2018, 57 (16), 4192–4214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (12).de Almeida AF; Moreira R; Rodrigues T. Synthetic Organic Chemistry Driven by Artificial Intelligence. Nat. Rev. Chem 2019, 3 (10), 589–604. [Google Scholar]
- (13).Strieth-Kalthoff F; Sandfort F; Segler MHS; Glorius F. Machine Learning the Ropes: Principles, Applications and Directions in Synthetic Chemistry. Chem. Soc. Rev 2020, 49 (17), 6154–6168. [DOI] [PubMed] [Google Scholar]
- (14).Maryasin B; Marquetand P; Maulide N. Machine Learning for Organic Synthesis: Are Robots Replacing Chemists? Angew. Chem., Int. Ed 2018, 57 (24), 6978–6980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (15).Ekins S; Puhl AC; Zorn KM; Lane TR; Russo DP; Klein JJ; Hickey AJ; Clark AM Exploiting Machine Learning for End-to-End Drug Discovery and Development. Nat. Mater 2019, 18 (5), 435–441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (16).Gromski PS; Henson AB; Granda JM; Cronin L. How to Explore Chemical Space Using Algorithms and Automation. Nat. Rev. Chem 2019, 3 (2), 119–128. [Google Scholar]
- (17).Kusne AG; Yu H; Wu C; Zhang H; Hattrick-Simpers J; DeCost B; Sarker S; Oses C; Toher C; Curtarolo S; et al. On-the-Fly Closed-Loop Materials Discovery via Bayesian Active Learning. Nat. Commun 2020, 11 (1), 5966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (18).Epps RW; Volk AA; Reyes KG; Abolhasani M. Accelerated AI Development for Autonomous Materials Synthesis in Flow. Chem. Sci 2021, 12 (17), 6025–6036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (19).Braham EJ; Davidson RD; Al-Hashimi M; Arróyave R; Banerjee S. Navigating the Design Space of Inorganic Materials Synthesis Using Statistical Methods and Machine Learning. Dalt. Trans 2020, 49 (33), 11480–11488. [DOI] [PubMed] [Google Scholar]
- (20).Tabor DP; Roch LM; Saikin SK; Kreisbeck C; Sheberla D; Montoya JH; Dwaraknath S; Aykol M; Ortiz C; Tribukait H; et al. Accelerating the Discovery of Materials for Clean Energy in the Era of Smart Automation. Nat. Rev. Mater 2018, 3 (5), 5–20. [Google Scholar]
- (21).Volk AA; Epps RW; Abolhasani M. Accelerated Development of Colloidal Nanomaterials Enabled by Modular Microfluidic Reactors: Toward Autonomous Robotic Experimentation. Adv. Mater 2021, 33 (4), 2004495. [DOI] [PubMed] [Google Scholar]
- (22).Pollice R; dos Passos Gomes G; Aldeghi M; Hickman RJ; Krenn M; Lavigne C; Lindner-D’Addario M; Nigam A; Ser CT; Yao Z; et al. Data-Driven Strategies for Accelerated Materials Design. Acc. Chem. Res 2021, 54 (4), 849–860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (23).Shields BJ; Stevens J; Li J; Parasram M; Damani F; Alvarado JIM; Janey JM; Adams RP; Doyle AG Bayesian Reaction Optimization as a Tool for Chemical Synthesis. Nature 2021, 590 (7844), 89–96. [DOI] [PubMed] [Google Scholar]
- (24).Sha W; Li Y; Tang S; Tian J; Zhao Y; Guo Y; Zhang W; Zhang X; Lu S; Cao Y; et al. Machine Learning in Polymer Informatics. InfoMat 2021, 3 (4), 353–361. [Google Scholar]
- (25).Kuenneth C; Rajan A; Tran H; Chen L; Kim C; Ramprasad R. Polymer Informatics with Multi-Task Learning. Patterns 2021, 2 (4), 100238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (26).Audus DJ; de Pablo JJ Polymer Informatics: Opportunities and Challenges. ACS Macro Lett. 2017, 6, 1078–1082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (27).Jackson NE; Webb MA; de Pablo JJ Recent Advances in Machine Learning towards Multiscale Soft Materials Design. Curr. Opin. Chem. Eng 2019, 23, 106–114. [Google Scholar]
- (28).Chen L; Pilania G; Batra R; Huan TD; Kim C; Kuenneth C; Ramprasad R. Polymer Informatics: Current Status and Critical next Steps. Mater. Sci. Eng., R 2021, 144, 100595. [Google Scholar]
- (29).Chan D; Chien J-C; Axpe E; Blankemeier L; Baker SW; Swaminathan S; Piunova VA; Zubarev DY; Soh HT; Appel EA Combinatorial Polyacrylamide Hydrogels for Preventing Biofouling on Implantable Biosensors. bioRxiv 2020, DOI: 10.1101/2020.05.25.115675. [DOI] [PMC free article] [PubMed]
- (30).Kumar R; Le N; Tan Z; Brown ME; Jiang S; Reineke TM Efficient Polymer-Mediated Delivery of Gene-Editing Ribonucleoprotein Payloads through Combinatorial Design, Parallelized Experimentation, and Machine Learning. ACS Nano 2020, 14 (12), 17626–17639. [DOI] [PubMed] [Google Scholar]
- (31).Doan Tran H; Kim C; Chen L; Chandrasekaran A; Batra R; Venkatram S; Kamal D; Lightstone JP; Gurnani R; Shetty P. Machine-Learning Predictions of Polymer Properties with Polymer Genome. J. Appl. Phys 2020, 128 (17), 171104. [Google Scholar]
- (32).Lin TS; Coley CW; Mochigase H; Beech HK; Wang W; Wang Z; Woods E; Craig SL; Johnson JA; Kalow JA; et al. BigSMILES: A Structurally-Based Line Notation for Describing Macromolecules. ACS Cent. Sci 2019, 5 (9), 1523–1531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (33).Lin TS; Rebello NJ; Beech HK; Wang Z; El-Zaatari B; Lundberg DJ; Johnson JA; Kalow JA; Craig SL; Olsen BD PolyDAT: A Generic Data Schema for Polymer Characterization. J. Chem. Inf. Model 2021, 61, 1150–1163. [DOI] [PubMed] [Google Scholar]
- (34).Kim C; Chandrasekaran A; Jha A; Ramprasad R. Active-Learning and Materials Design: The Example of High Glass Transition Temperature Polymers. MRS Commun. 2019, 9 (3), 860–866. [Google Scholar]
- (35).Zhang Y; Xu X. Machine Learning Glass Transition Temperature of Polyacrylamides Using Quantum Chemical Descriptors. Polym. Chem 2021, 12 (6), 843–851. [Google Scholar]
- (36).Pilania G; Iverson CN; Lookman T; Marrone BL Machine-Learning-Based Predictive Modeling of Glass Transition Temperatures: A Case of Polyhydroxyalkanoate Homopolymers and Copolymers. J. Chem. Inf. Model 2019, 59 (12), 5013–5025. [DOI] [PubMed] [Google Scholar]
- (37).Park NH; Zubarev DY; Hedrick JL; Kiyek V; Corbet C; Lottier S. A Recommender System for Inverse Design of Polycarbonates and Polyesters. Macromolecules 2020, 53 (24), 10847–10854. [Google Scholar]
- (38).Wu S; Kondo Y; Kakimoto M. aki; Yang B; Yamada H; Kuwajima I; Lambard G; Hongo K; Xu Y; Shiomi J. MachineLearning-Assisted Discovery of Polymers with High Thermal Conductivity Using a Molecular Design Algorithm. NPJ Comput. Mater 2019, 5 (1), 66. [Google Scholar]
- (39).Kholodovych V; Gubskaya AV; Bohrer M; Harris N; Knight D; Kohn J; Welsh WJ Prediction of Biological Response for Large Combinatorial Libraries of Biodegradable Polymers: Polymethacrylates as a Test Case. Polymer 2008, 49 (10), 2435–2439. [Google Scholar]
- (40).Pruksawan S; Lambard G; Samitsu S; Sodeyama K; Naito M. Prediction and Optimization of Epoxy Adhesive Strength from a Small Dataset through Active Learning. Sci. Technol. Adv. Mater 2019, 20 (1), 1010–1021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (41).Srinivas M; Morel PA; Ernst LA; Laidlaw DH; Ahrens ET Fluorine-19 MRI for Visualization and Quantification of Cell Migration in a Diabetes Model. Magn. Reson. Med 2007, 58 (4), 725–734. [DOI] [PubMed] [Google Scholar]
- (42).Janjic JM; Ahrens ET Fluorine-Containing Nanoemulsions for MRI Cell Tracking. WIREs Nanomed Nanobiol. 2009, 1, 492–501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (43).Srinivas M; Heerschap A; Ahrens ET; Figdor CG; de Vries IJM 19F MRI for Quantitative in Vivo Cell Tracking. Trends Biotechnol. 2010, 28 (7), 363–370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (44).Chapelin F; Capitini CM; Ahrens ET Fluorine-19 MRI for Detection and Quantification of Immune Cell Therapy for Cancer. J. Immunother. Cancer 2018, 6 (1), 105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (45).Taylor NG; Chung SH; Kwansa AL; Johnson RR; Teator AJ; Milliken NJB; Koshlap KM; Yingling YG; Lee YZ; Leibfarth FA Partially Fluorinated Copolymers as Oxygen Sensitive 19F MRI Agents. Chem. - Eur. J 2020, 26 (44), 9982–9990. [DOI] [PubMed] [Google Scholar]
- (46).Tirotta I; Dichiarante V; Pigliacelli C; Cavallo G; Terraneo G; Bombelli FB; Metrangolo P; Resnati G. 19F Magnetic Resonance Imaging (MRI): From Design of Materials to Clinical Applications. Chem. Rev 2015, 115 (2), 1106–1129. [DOI] [PubMed] [Google Scholar]
- (47).Gopakumar AM; Balachandran PV; Xue D; Gubernatis JE; Lookman T. Multi-Objective Optimization for Materials Discovery via Adaptive Design. Sci. Rep 2018, 8 (1), 1–12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (48).Settles B. From Theories to Queries: Active Learning in Practice. In Learning Active and Experimental Design Workshop in Conjunction with AISTATS 2010; Guyon I; Cawley G; Dror G; Lemaire V; Statnikov A, Eds.; Proceedings of Machine Learning Research 16; PMLR: Sardinia, Italy, 2011. [Google Scholar]
- (49).Automated Machine Learning: Methods, Systems, and Challenges; Hutter F, Vanschoren J, t L, Eds.; Springer: Cham, Switzerland, 2019. [Google Scholar]
- (50).Feurer M; Eggensperger K; Falkner S; Lindauer M; Hutter F. Auto-Sklearn 2.0: The Next Generation. arXiv [cs.LG], July 8, 2020, 2007.04074. https://arxiv.org/abs/2007.04074 (accessed August 10, 2020).
- (51).Olson RS; Bartley N; Urbanowicz RJ; Moore JH Evaluation of a Tree-Based Pipeline Optimization Tool for Automating Data Science. In GECCO ‘16: Proceedings of the Genetic and Evolutionary Computation Conference 2016; Association for Computing Machinery: New York, 2016; pp 485–492. [Google Scholar]
- (52).Pedregosa F; Varoquaux G; Gramfort A; Michel V; Thirion B; Grisel O; Blondel M; Prettenhofer P; Weiss R; Dubourg V; et al. Scikit-Learn: Machine Learning in Python. J. Mach. Learn. Res 2011, 12, 2825–2830. [Google Scholar]
- (53).Chen T; Guestrin C. XGBoost: A Scalable Tree Boosting System. In KDD ‘16: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining; Association for Computing Machinery: New York, 2016; pp 785–794. [Google Scholar]
- (54).Duan T; Avati A; Ding DY; Basu S; Ng AY; Schuler A. NGBoost: Natural Gradient Boosting for Probabilistic Prediction. arXiv [cs.LG], October 8, 2019, 1910.03225. https://arxiv.org/abs/1910.03225 (accessed August 8, 2020).
- (55).Reis MH; Leibfarth FA; Pitet LM Polymerizations in Continuous Flow: Recent Advances in the Synthesis of Diverse Polymeric Materials. ACS Macro Lett. 2020, 9 (1), 123–133. [DOI] [PubMed] [Google Scholar]
- (56).Zhou Y; Gu Y; Jiang K; Chen M. Droplet-Flow Photopolymerization Aided by Computer: Overcoming the Challenges of Viscosity and Facilitating the Generation of Copolymer Libraries. Macromolecules 2019, 52 (15), 5611–5617. [Google Scholar]
- (57).Reis MH; Davidson CLG; Leibfarth FA Continuous-Flow Chemistry for the Determination of Comonomer Reactivity Ratios. Polym. Chem 2018, 9 (13), 1728–1734. [Google Scholar]
- (58).Tonhauser C; Natalello A; Löwe H; Frey H. Microflow Technology in Polymer Synthesis. Macromolecules 2012, 45 (24), 9551–9570. [Google Scholar]
- (59).Walsh DJ; Schinski DA; Schneider RA; Guironnet D. General Route to Design Polymer Molecular Weight Distributions through Flow Chemistry. Nat. Commun 2020, 11 (1), 3094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (60).Rubens M; Vrijsen JH; Laun J; Junkers T. Precise Polymer Synthesis by Autonomous Self-Optimizing Flow Reactors. Angew. Chem 2019, 131 (10), 3215–3219. [DOI] [PubMed] [Google Scholar]
- (61).Reis MH; Varner TP; Leibfarth FA The Influence of Residence Time Distribution on Continuous-Flow Polymerization. Macromolecules 2019, 52 (9), 3551–3557. [Google Scholar]
- (62).Lu S; Wang K. Kinetic Study of TBD Catalyzed δ-Valerolactone Polymerization Using a Gas-Driven Droplet Flow Reactor. React. Chem. Eng 2019, 4 (7), 1189–1194. [Google Scholar]
- (63).Lin B; Hedrick JL; Park NH; Waymouth RM Programmable High-Throughput Platform for the Rapid and Scalable Synthesis of Polyester and Polycarbonate Libraries. J. Am. Chem. Soc 2019, 141 (22), 8921–8927. [DOI] [PubMed] [Google Scholar]
- (64).Wang K; Peng H; Thurecht KJ; Puttick S; Whittaker AK Segmented Highly Branched Copolymers: Rationally Designed Macromolecules for Improved and Tunable 19F MRI. Biomacromolecules 2015, 16 (9), 2827–2839. [DOI] [PubMed] [Google Scholar]
- (65).Reizman BJ; Jensen KF Simultaneous Solvent Screening and Reaction Optimization in Microliter Slugs. Chem. Commun 2015, 51 (68), 13290–13293. [DOI] [PubMed] [Google Scholar]
- (66).Sohrabi S; Kassir N; Keshavarz Moraveji M. Droplet Microfluidics: Fundamentals and Its Advanced Applications. RSC Adv. 2020, 10 (46), 27560–27574. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- (67).Li M; Zhang Y; Zhang J; Peng M; Yan L; Tang Z; Wu Q. Continuous Gas−Liquid−Solid Slug Flow for Sustainable Heterogeneously Catalyzed PET-RAFT Polymerization. Ind. Eng. Chem. Res 2021, 60, 5451–5462. [Google Scholar]
- (68).Ardana A; Whittaker AK; Thurecht KJ PEG-Based Hyperbranched Polymer Theranostics: Optimizing Chemistries for Improved Bioconjugation. Macromolecules 2014, 47 (15), 5211–5219. [Google Scholar]
- (69).Zhao W; Ta HT; Zhang C; Whittaker AK Polymerization-Induced Self-Assembly (PISA) - Control over the Morphology of 19F-Containing Polymeric Nano-Objects for Cell Uptake and Tracking. Biomacromolecules 2017, 18 (4), 1145–1156. [DOI] [PubMed] [Google Scholar]
- (70).Wang K; Peng H; Thurecht KJ; Puttick S; Whittaker AK Multifunctional Hyperbranched Polymers for CT/ 19 F MRI Bimodal Molecular Imaging. Polym. Chem 2016, 7 (5), 1059–1069. [Google Scholar]
- (71).Zhang C; Moonshi SS; Peng H; Puttick S; Reid J; Bernardi S; Searles DJ; Whittaker AK Ion-Responsive 19 F MRI Contrast Agents for the Detection of Cancer Cells. ACS Sensors 2016, 1 (6), 757–765. [Google Scholar]
- (72).Wang K; Peng H; Thurecht KJ; Whittaker AK Fluorinated POSS-Star Polymers for 19 F MRI. Macromol. Chem. Phys 2016, 217 (20), 2262–2274. [Google Scholar]
- (73).Wang K; Peng H; Thurecht KJ; Puttick S; Whittaker AK PH-Responsive Star Polymer Nanoparticles: Potential 19F MRI Contrast Agents for Tumour-Selective Imaging. Polym. Chem 2013, 4 (16), 4480–4489. [Google Scholar]
- (74).Zhang C; Moonshi SS; Han Y; Puttick S; Peng H; Magoling BJA; Reid JC; Bernardi S; Searles DJ; Král P; et al. PFPE-Based Polymeric 19F MRI Agents: A New Class of Contrast Agents with Outstanding Sensitivity. Macromolecules 2017, 50 (15), 5953–5963. [Google Scholar]
- (75).Munkhbat O; Canakci M; Zheng S; Hu W; Osborne B; Bogdanov AA; Thayumanavan S. 19 F MRI of Polymer Nanogels Aided by Improved Segmental Mobility of Embedded Fluorine Moieties. Biomacromolecules 2019, 20 (2), 790–800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (76).Thurecht KJ; Blakey I; Peng H; Squires O; Hsu S; Alexander C; Whittaker AK Functional Hyperbranched Polymers: Toward Targeted in Vivo 19F Magnetic Resonance Imaging Using Designed Macromolecules. J. Am. Chem. Soc 2010, 132 (15), 5336–5337. [DOI] [PubMed] [Google Scholar]
- (77).Fu C; Zhang C; Peng H; Han F; Baker C; Wu Y; Ta H; Whittaker AK Enhanced Performance of Polymeric 19F MRI Contrast Agents through Incorporation of Highly Water-Soluble Monomer MSEA. Macromolecules 2018, 51 (15), 5875–5882. [Google Scholar]
- (78).Bagheri A; Bainbridge CWA; Engel KE; Qiao GG; Xu J; Boyer C; Jin J. Oxygen Tolerant PET-RAFT Facilitated 3D Printing of Polymeric Materials under Visible LEDs. ACS Appl. Polym. Mater 2020, 2 (2), 782–790. [Google Scholar]
- (79).Song Y; Kim Y; Noh Y; Singh VK; Behera SK; Abudulimu A; Chung K; Wannemacher R; Gierschner J; Lüer, L.; et al. Organic Photocatalyst for Ppm-Level Visible-Light-Driven Reversible Addition−Fragmentation Chain-Transfer (RAFT) Polymerization with Excellent Oxygen Tolerance. Macromolecules 2019, 52 (15), 5538–5545. [Google Scholar]
- (80).Szymański JK; Pérez-Mercader J. Direct Optical Observations of Vesicular Self-Assembly in Large-Scale Polymeric Structures during Photocontrolled Biphasic Polymerization. Polym. Chem 2016, 7 (47), 7211–7215. [Google Scholar]
- (81).Albertsen AN; Szymański JK; Pérez-Mercader J . Emergent Properties of Giant Vesicles Formed by a Polymerization-Induced Self-Assembly (PISA) Reaction. Sci. Rep 2017, 7 (1), 41534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (82).Ren K; Perez-Mercader J. Thermoresponsive Gels Directly Obtained via Visible Light-Mediated Polymerization-Induced Self-Assembly with Oxygen Tolerance. Polym. Chem 2017, 8 (23), 3548–3552. [Google Scholar]
- (83).Niu J; Page ZA; Dolinski ND; Anastasaki A; Hsueh AT; Soh HT; Hawker CJ Rapid Visible Light-Mediated Controlled Aqueous Polymerization with In Situ Monitoring. ACS Macro Lett. 2017, 6 (10), 1109–1113. [DOI] [PubMed] [Google Scholar]
- (84).Xu J; Jung K; Boyer C. Oxygen Tolerance Study of Photoinduced Electron Transfer−Reversible Addition−Fragmentation Chain Transfer (PET-RAFT) Polymerization Mediated by Ru(Bpy) 3 Cl 2. Macromolecules 2014, 47 (13), 4217–4229. [Google Scholar]
- (85).Shanmugam S; Xu J; Boyer C. Photoinduced Electron Transfer−Reversible Addition−Fragmentation Chain Transfer (PET-RAFT) Polymerization of Vinyl Acetate and N -Vinylpyrrolidinone: Kinetic and Oxygen Tolerance Study. Macromolecules 2014, 47 (15), 4930–4942. [Google Scholar]
- (86).Yang P; Pageni P; Kabir MP; Zhu T; Tang C. Metallocene-Containing Homopolymers and Heterobimetallic Block Copolymers via Photoinduced RAFT Polymerization. ACS Macro Lett. 2016, 5 (11), 1293–1300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (87).Xu J; Shanmugam S; Fu C; Aguey-Zinsou K-F; Boyer C. Selective Photoactivation: From a Single Unit Monomer Insertion Reaction to Controlled Polymer Architectures. J. Am. Chem. Soc 2016, 138 (9), 3094–3106. [DOI] [PubMed] [Google Scholar]
- (88).Shanmugam S; Xu J; Boyer C. Utilizing the Electron Transfer Mechanism of Chlorophyll a under Light for Controlled Radical Polymerization. Chem. Sci 2015, 6 (2), 1341–1349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (89).Xu J; Shanmugam S; Duong HT; Boyer C. Organo-Photocatalysts for Photoinduced Electron Transfer-Reversible Addition−Fragmentation Chain Transfer (PET-RAFT) Polymerization. Polym. Chem 2015, 6 (31), 5615–5624. [Google Scholar]
- (90).Lee I-H; Discekici EH; Anastasaki A; de Alaniz JR; Hawker CJ Controlled Radical Polymerization of Vinyl Ketones Using Visible Light. Polym. Chem 2017, 8 (21), 3351–3356. [Google Scholar]
- (91).Shanmugam S; Xu J; Boyer C. Exploiting Metalloporphyrins for Selective Living Radical Polymerization Tunable over Visible Wavelengths. J. Am. Chem. Soc 2015, 137 (28), 9174–9185. [DOI] [PubMed] [Google Scholar]
- (92).Shanmugam S; Xu J; Boyer C. Aqueous RAFT Photopolymerization with Oxygen Tolerance. Macromolecules 2016, 49 (24), 9345–9357. [Google Scholar]
- (93).Shen L; Lu Q; Zhu A; Lv X; An Z. Photocontrolled RAFT Polymerization Mediated by a Supramolecular Catalyst. ACS Macro Lett. 2017, 6 (6), 625–631. [DOI] [PubMed] [Google Scholar]
- (94).Xu S; Ng G; Xu J; Kuchel RP; Yeow J; Boyer C. 2-(Methylthio)Ethyl Methacrylate: A Versatile Monomer for Stimuli Responsiveness and Polymerization-Induced Self-Assembly in the Presence of Air. ACS Macro Lett. 2017, 6 (11), 1237–1244. [DOI] [PubMed] [Google Scholar]
- (95).Ng G; Yeow J; Xu J; Boyer C. Application of Oxygen Tolerant PET-RAFT to Polymerization-Induced Self-Assembly. Polym. Chem 2017, 8 (18), 2841–2851. [Google Scholar]
- (96).Yeow J; Shanmugam S; Corrigan N; Kuchel RP; Xu J; Boyer C. A Polymerization-Induced Self-Assembly Approach to Nanoparticles Loaded with Singlet Oxygen Generators. Macromolecules 2016, 49 (19), 7277–7285. [Google Scholar]
- (97).Corrigan N; Almasri A; Taillades W; Xu J; Boyer C. Controlling Molecular Weight Distributions through Photoinduced Flow Polymerization. Macromolecules 2017, 50 (21), 8438–8448. [Google Scholar]
- (98).Corrigan N; Rosli D; Jones JWJ; Xu J; Boyer C. Oxygen Tolerance in Living Radical Polymerization: Investigation of Mechanism and Implementation in Continuous Flow Polymerization. Macromolecules 2016, 49 (18), 6779–6789. [Google Scholar]
- (99).Fu Q; Xie K; McKenzie TG; Qiao GG Trithiocarbonates as Intrinsic Photoredox Catalysts and RAFT Agents for Oxygen Tolerant Controlled Radical Polymerization. Polym. Chem 2017, 8 (9), 1519–1526. [Google Scholar]
- (100).McKenzie TG; Fu Q; Wong EHH; Dunstan DE; Qiao GG Visible Light Mediated Controlled Radical Polymerization in the Absence of Exogenous Radical Sources or Catalysts. Macromolecules 2015, 48 (12), 3864–3872. [Google Scholar]
- (101).Quinn JF; Barner L; Barner-Kowollik C; Rizzardo E; Davis TP Reversible Addition−Fragmentation Chain Transfer Polymerization Initiated with Ultraviolet Radiation. Macromolecules 2002, 35 (20), 7620–7627. [Google Scholar]
- (102).Lu L; Zhang H; Yang N; Cai Y. Toward Rapid and Well-Controlled Ambient Temperature RAFT Polymerization under UV−Vis Radiation: Effect of Radiation Wave Range. Macromolecules 2006, 39 (11), 3770–3776. [Google Scholar]
- (103).Wang J; Rivero M; Muñoz Bonilla A; Sanchez-Marcos J; Xue W; Chen G; Zhang W; Zhu X. Natural RAFT Polymerization: Recyclable-Catalyst-Aided, Opened-to-Air, and Sunlight-Photolyzed RAFT Polymerizations. ACS Macro Lett. 2016, 5 (11), 1278. [DOI] [PubMed] [Google Scholar]
- (104).Bagheri A; Engel KE; Bainbridge CWA; Xu J; Boyer C; Jin J. 3D Printing of Polymeric Materials Based on Photo-RAFT Polymerization. Polym. Chem 2020, 11 (3), 641–647. [Google Scholar]
- (105).Lamb JR; Qin KP; Johnson JA Visible-Light-Mediated, Additive-Free, and Open-to-Air Controlled Radical Polymerization of Acrylates and Acrylamides. Polym. Chem 2019, 10 (13), 1585–1590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (106).Kuroki A; Martinez-Botella I; Hornung CH; Martin L; Williams EGL; Locock KES; Hartlieb M; Perrier S. Looped Flow RAFT Polymerization for Multiblock Copolymer Synthesis. Polym. Chem 2017, 8 (21), 3249–3254. [Google Scholar]
- (107).Gurnani P; Floyd T; Tanaka J; Stubbs C; Lester D; Sanchez-Cano C; Perrier S. PCR-RAFT: Rapid High Throughput Oxygen Tolerant RAFT Polymer Synthesis in a Biology Laboratory. Polym. Chem 2020, 11 (6), 1230–1236. [Google Scholar]
- (108).Tanaka J; Gurnani P; Cook AB; Häkkinen S; Zhang J; Yang J; Kerr A; Haddleton DM; Perrier S; Wilson P. Microscale Synthesis of Multiblock Copolymers Using Ultrafast RAFT Polymerisation. Polym. Chem 2019, 10 (10), 1186–1191. [Google Scholar]
- (109).Parkinson S; Knox ST; Bourne RA; Warren NJ Rapid Production of Block Copolymer Nano-Objects via Continuous-Flow Ultrafast RAFT Dispersion Polymerisation. Polym. Chem 2020, 11 (20), 3465–3474. [Google Scholar]
- (110).Gody G; Barbey R; Danial M; Perrier S. Ultrafast RAFT Polymerization: Multiblock Copolymers within Minutes. Polym. Chem 2015, 6 (9), 1502–1511. [Google Scholar]
- (111).Upadhya R; Kanagala MJ; Gormley AJ Purifying Low-Volume Combinatorial Polymer Libraries with Gel Filtration Columns. Macromol. Rapid Commun 2019, 40 (24), 1900528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (112).Webb A. Increasing the Sensitivity of Magnetic Resonance Spectroscopy and Imaging. Anal. Chem 2012, 84 (1), 9–16. [DOI] [PubMed] [Google Scholar]
- (113).Butina D. Unsupervised Data Base Clustering Based on Daylight’s Fingerprint and Tanimoto Similarity: A Fast and Automated Way To Cluster Small and Large Data Sets. J. Chem. Inf. Comput. Sci 1999, 39 (4), 747–750. [Google Scholar]
- (114).McInnes L; Healy J; Melville J. UMAP: Uniform Manifold Approximation and Projection for Dimension Reduction. arXiv [stat.ML]. February 9, 2018, 1802.03426v3. https://arxiv.org/abs/1802.03426 (accessed February 20, 2019).
- (115).Haase A; Frahm J; Matthaei D; Hanicke W; Merboldt K-D FLASH Imaging. Rapid NMR Imaging Using Low Flip-Angle Pulses. J. Magn. Reson 1986, 67 (2), 258–266. [DOI] [PubMed] [Google Scholar]
- (116).Zhang S; Block KT; Frahm J. Magnetic Resonance Imaging in Real Time: Advances Using Radial FLASH. J. Magn. Reson. Imaging 2010, 31 (1), 101–109. [DOI] [PubMed] [Google Scholar]
- (117).Jiang Z-X; Liu X; Jeong E-K; Yu YB Symmetry-Guided Design and Fluorous Synthesis of a Stable and Rapidly Excreted Imaging Tracer for 19F MRI. Angew. Chem., Int. Ed 2009, 48 (26), 4755–4758. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.