SUMMARY
The metazoan-specific Integrator complex catalyzes 3' end processing of small nuclear RNAs (snRNAs) and premature termination that attenuates transcription of many protein-coding genes. Integrator has RNA endonuclease and protein phosphatase activities, but it remains unclear if both are required for complex function. Here, we show IntS6 (Integrator subunit 6) over-expression blocks Integrator function at a subset of Drosophila protein-coding genes, while having no effect on snRNAs or attenuation of other loci. Over-expressed IntS6 titrates protein phosphatase 2A (PP2A) subunits, thereby only affecting gene loci where phosphatase activity is necessary for Integrator function. IntS6 functions analogous to a PP2A regulatory B subunit as over-expression of canonical B subunits, which do not bind Integrator, are also sufficient to inhibit Integrator activity. These results show that the phosphatase module is critical at only a subset of Integrator regulated genes and point to PP2A recruitment as a tunable step that modulates transcription termination efficiency.
Keywords: RNA polymerase II, 3' end processing, snRNA, protein phosphatase 2A (PP2A), Integrator subunit 6, Integrator complex, transcription, promoter-proximal termination, INTAC
Graphical Abstract

eTOC BLURB
Fujiwara et al. show that the phosphatase module of the Integrator complex is differentially required for transcription termination across Drosophila gene loci. When protein phosphatase 2A subunits are limiting, Integrator complex function is blocked at a subset of protein-coding genes, yet snRNA processing and transcription attenuation of other loci continue.
INTRODUCTION
Transcription of all eukaryotic protein-coding genes and most noncoding RNAs is catalyzed by RNA polymerase II (Pol II), which itself is regulated by many transcription factors and co-factors that act during the initiation, elongation, and termination steps (for review, see 1-3). These multiple macromolecular assemblies collectively help dictate the levels, processing, and functions of the transcripts produced. In recent years, the multi-subunit Integrator complex (sometimes referred to as INTAC) has emerged as a key transcriptional regulator across metazoans that controls the fates of many nascent RNAs (for review, see 4-8). Integrator is > 1.5 MDa and interacts with the C-terminal domain (CTD) of Pol II, with most of the more than 14 subunits in the Integrator complex lacking identifiable paralogs.9 The notable exceptions are Integrator subunits 11 (IntS11) and 9 (IntS9), which are homologous to the RNA endonuclease CPSF73 (cleavage and polyadenylation specificity factor 73, also known as CPSF3) and CPSF100 (also known as CPSF2) that cleave mRNA 3' ends prior to poly(A) tail addition.10 The catalytic and scaffolding subunits of protein phosphatase 2A (PP2A) are also stably associated with Integrator, and thus the complex can have dual RNA cleavage and phosphatase catalytic activities.11-18
It has long been recognized that Integrator catalyzes RNA cleavage at the 3' ends of nascent small nuclear RNA (snRNA) gene loci, enabling termination of transcription and release of the snRNA transcript that goes on to function in pre-mRNA splicing.9,19 Integrator has further been implicated in the processing of a variety of other Pol II transcripts, including enhancer RNAs,20,21 telomerase RNA,22 viral microRNAs,23 piwi-interacting RNAs,24,25 replication-dependent histone mRNAs,26 and many canonical protein-coding genes.27-32 It is thus perhaps not surprising that mis-regulation of Integrator is associated with developmental and disease phenotypes, including in humans.4,33-37
Analogous to how Integrator functions at snRNAs, the complex can endonucleolytically cleave nascent mRNAs to enable transcription termination at protein-coding genes coupled to nascent RNA release.27-30 These cleavage events predominantly do not occur at the 3' ends of genes where CPSF73 acts but instead close to the transcription start site (TSS), often within 100 nucleotides at sites of Pol II pausing. This is at least in part because the Integrator phosphatase module can antagonize kinases that promote release of paused Pol II.11-14,38 Instead of allowing Pol II to productively elongate, Integrator causes transcription to prematurely terminate, and the released short mRNAs are rapidly degraded by the RNA exosome.27-29 In some cases, Integrator cleavage has been proposed to activate transcription by enabling removal of stalled, inactive Pol II,32,38 but termination driven by Integrator can also attenuate gene expression by blocking full length mRNA production. Attenuation can be potent and readily observed in Drosophila cells, as exemplified by our prior genome-scale RNAi screen that revealed depletion of Integrator subunits resulted in the largest increase (among all genes tested) in the output of the Metallothionein A (MtnA) promoter.28 RNA-seq further revealed more than 400 mRNAs that were up-regulated in Drosophila cells upon depletion of IntS9, some by more than 100-fold, compared to only 49 mRNAs that were down-regulated. Integrator is thus a major attenuator of protein-coding gene outputs in Drosophila, yet how Integrator activity can be toggled on/off at a given gene locus depending on cellular transcriptional needs remains poorly understood.
There is emerging evidence that Integrator recruitment and/or activity is differentially regulated across gene loci (for review, see 4). For example, depletion of many non-catalytic Integrator subunits has only a minimal effect on snRNA 3' end processing, yet these subunits are critical (especially IntS1, 2, 5, 6, 7, 8, and 12) for the ability of Integrator to attenuate the outputs of protein-coding genes.28,39,40 This suggests these non-catalytic Integrator subunits may act to ensure the proper balance between full-length mRNA production and premature termination at protein-coding genes, but the underlying molecular mechanisms remain unclear. There is also evidence that Integrator subunits can function independently of the rest of the complex, e.g. as part of the DNA damage response41 or to activate enhancers.21
Here, to identify regulatory subunits that enable the Integrator complex to be distinctly regulated in a locus-specific manner, we tested the effect of individually over-expressing each Integrator subunit in Drosophila cells. This revealed that IntS6 over-expression uniquely blocks Integrator function at a subset of protein-coding and long non-coding RNA genes. IntS6 can thus function in a dominant-negative manner, but IntS6 over-expression is not sufficient to fully disable Integrator across the genome: nascent snRNAs continue to be processed and the expression of many protein-coding genes remains attenuated by Integrator. The only loci affected by IntS6 over-expression are those where the PP2A phosphatase module is required for Integrator function, which is only a subset of gene loci bound by Integrator. We found that over-expressed IntS6 acts like a molecular sponge that titrates PP2A subunits and that IntS6 functions in a manner analogous to canonical PP2A regulatory B subunits. Altogether, these results show that the PP2A phosphatase module is critical at some, but not all genes for Integrator function and point to recruitment of PP2A via IntS6 as a tunable step that can control Integrator activity in a locus-specific manner.
RESULTS
Over-expression of IntS6 uniquely inhibits Integrator function at a subset of protein-coding promoters
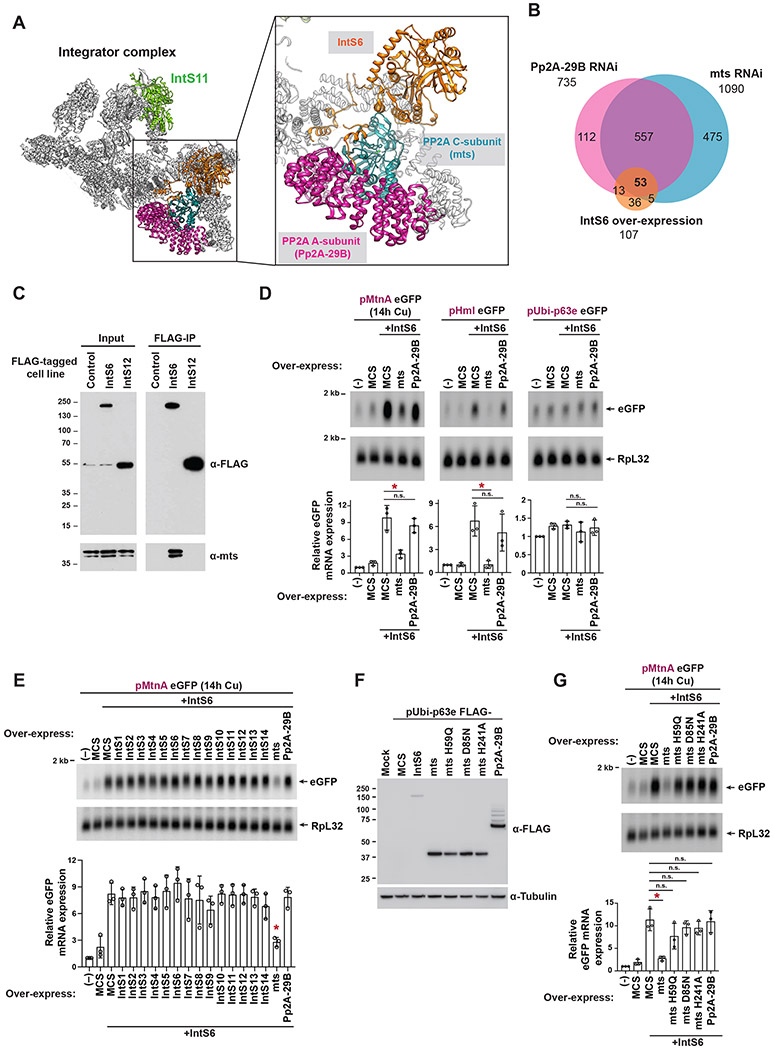
Cleavage of nascent snRNA transcripts by Integrator is critical for production of functional snRNAs,9 while cleavage of nascent mRNAs by Integrator triggers premature transcription termination coupled to nascent transcript degradation and attenuation of protein-coding gene expression.27,28 Given these opposite effects on the ultimate gene outputs, we reasoned that the Integrator complex must be regulated in a distinct manner at protein-coding genes compared to snRNAs. To reveal the critical regulatory differences, we took advantage of a set of eGFP reporters that are driven by Drosophila protein-coding promoters (Figure 1A, left) and compared their regulation patterns to that of snRNA readthrough reporters that produce eGFP when Integrator fails to process the 3' end of the encoded snRNA (Figure 1A, right). Each of the examined protein-coding gene promoters, which included MtnA, Hemolectin (Hml), CG8620, anachronism (ana), and the small subunit of ribonucleoside reductase (RnrS), are normally attenuated by Integrator in Drosophila DL1 cells as eGFP mRNA expression increased upon treatment with double-stranded RNA (dsRNA) targeting IntS4, a scaffolding component of the Integrator endonuclease cleavage module,42,43 relative to treatment with a control (β-galactosidase, β-gal) dsRNA (Figures 1B, left and S1A-C). As expected, the U4:39B and U5:34A snRNA readthrough reporters were likewise sensitive to depletion of IntS4 as demonstrated by a potent increase in eGFP levels due to Integrator not cleaving near the 3' box sequence (Figure 1B, left). In contrast, a reporter driven by the ubiquitin-63E (Ubi-p63e) promoter was not affected by IntS4 depletion (Figure 1B, left), consistent with prior RNA-seq results that showed endogenous Ubi-p63e mRNA levels are not regulated by Integrator.28
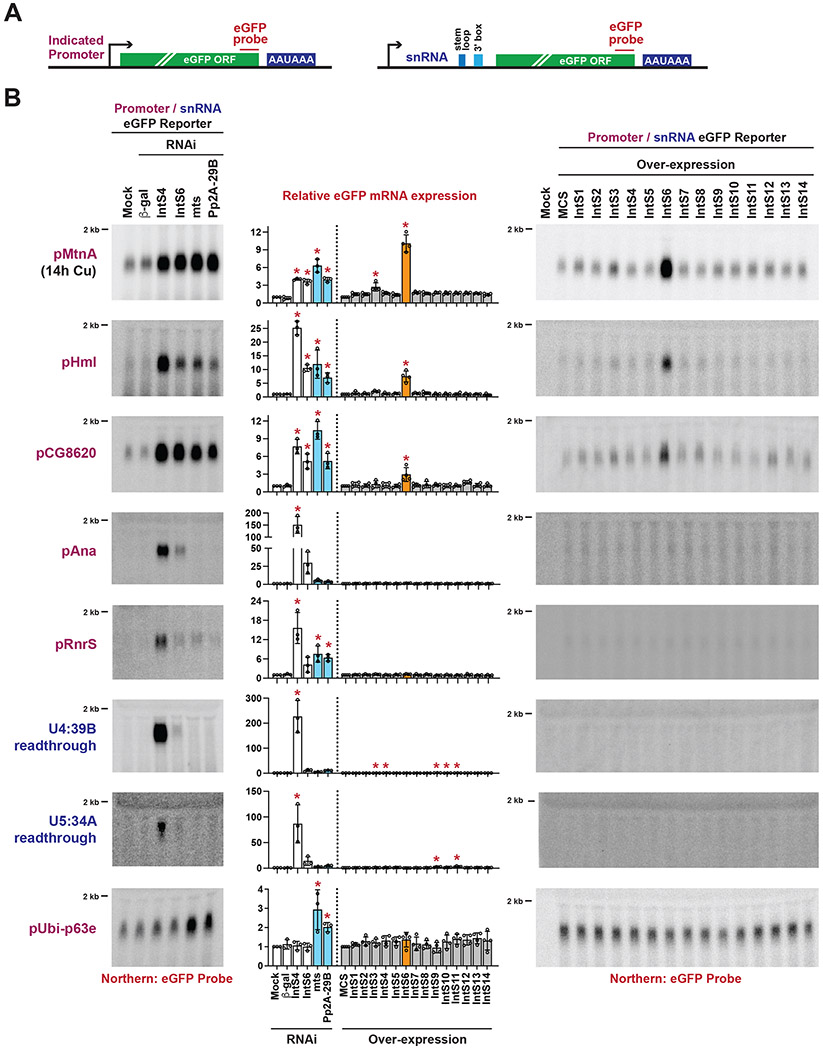
Figure 1. Integrator activity at a subset of reporter genes is lost upon over-expression of IntS6.
(A) eGFP-based reporters to examine Integrator activity at protein-coding genes (left) and snRNAs (right). The promoter and 5’ UTR of each indicated protein-coding gene was cloned upstream of eGFP (left). The snRNA promoter, coding sequence, and downstream region were cloned upstream of eGFP, thereby enabling eGFP production when Integrator fails to process the snRNA 3' end between the stem loop and 3' box sequences (right).
(B) Each individual reporter plasmid was transfected into DL1 cells that had been treated with the indicated dsRNAs (left) or was co-transfected with 100 ng of a plasmid that over-expresses a FLAG-tagged Integrator subunit from the Ubi-p63e promoter (right). A plasmid containing a multi-cloning site (MCS) driven by the Ubi-p63e promoter was used as a control for the over-expression experiments. CuSO4 was added for the last 14 h only when measuring eGFP production from the MtnA promoter. Total RNA was isolated and Northern blots (20 μg/lane) were used to measure expression of each eGFP reporter mRNA. Representative blots are shown and loading controls are provided in Figure S1C. RT-qPCR was used to quantify eGFP mRNA expression levels (middle). RNAi data were normalized to the mock samples and over-expression data were normalized to the MCS samples. Data are shown as mean ± SD, N ≥ 3. (*) P < 0.05.
See also Figures S1, S2, and Table S5.
We next took advantage of the eGFP reporters and examined the functional effects of individually over-expressing a FLAG-tagged version of each Integrator subunit (IntS1-IntS14) (Figure S1D). Compared to over-expressing a control transgene that encodes a multicloning site (MCS), individual over-expression of almost all Integrator subunits had no or minimal effect on the amounts of eGFP mRNA produced from the reporters (Figures 1B, right and S1C). The notable exception was IntS6 over-expression which caused significantly increased amounts of eGFP mRNA to be produced from the MtnA, Hml, and CG8620 promoters (Figure 1B, right). There was, however, no change in the outputs of the Ana, RnrS, or snRNA readthrough reporters upon IntS6 over-expression (Figure 1B, right). This suggested IntS6 over-expression may have a dominant-negative effect and be sufficient to disable Integrator function at a subset of protein-coding promoters, while having no effect on the ability of Integrator to attenuate other protein-coding promoters or process nascent snRNAs. To confirm the uniqueness of the IntS6 over-expression effects, increasing amounts of plasmids that over-express IntS6, IntS8 (which forms part of the Integrator shoulder module12), or IntS12 (which contains a PHD finger and can interact with the Integrator backbone module44) were transfected into DL1 cells. Effects on the eGFP reporters were then examined (Figures S2A and S2B). A progressive increase in eGFP levels from the MtnA promoter was observed with titration of the IntS6 plasmid, while IntS8 and IntS12 over-expression had no or minimal effect even at the highest tested levels.
The 1,284 amino acid Drosophila IntS6 protein contains a conserved Von Willebrand factor type A (VWA) domain at its N-terminus, a > 250 amino acid region in the middle of the protein (a.a. 950-1220) that is absent from IntS6 homologs in other species, and a conserved C-terminal domain that is observed in several additional proteins (Figure S2C). To define the key regions in Drosophila IntS6 responsible for the dominant-negative effect, FLAG-tagged IntS6 over-expression plasmids with deletions from the N- or C-terminus were generated (Figure S2C) and transfected into DL1 cells (Figure S2D). This revealed that the N-terminus (amino acids 1-600) and the C-terminus (amino acids 1197-1284) are each sufficient on their own to inhibit Integrator activity at the MtnA promoter (Figure S2E). Both functional regions of IntS6 contain conserved protein domains, and we further addressed whether the Drosophila-specific IntS6 region may play a role by testing the effect of over-expressing IntS6 homologs from human or zebrafish (Figure S2F). Human IntS6 and IntS6L (IntS6-like) were sufficient to inhibit Integrator activity at select promoters in Drosophila cells, arguing against a key role for the Drosophila-specific region of IntS6 (Figure S2G).
Depletion of IntS6 using RNAi interestingly revealed that this subunit was differentially required for Integrator function across the eGFP reporters (Figures 1B, left and S1B). When compared to IntS4 depletion, IntS6 depletion had only a minimal effect on the output of the snRNA readthrough reporters (e.g., 226 vs. 11-fold increase in eGFP expression for the U4:39B reporter) while IntS4 and IntS6 depletion resulted in near equal increases in eGFP mRNA expression from the MtnA promoter (4.0 and 3.6-fold, respectively) (Figure 1B, left). Hence, we concluded that IntS6 varies from dispensable (or near dispensable) to essential for Integrator function in Drosophila depending on which reporter is examined.
A subset of endogenous protein-coding genes normally attenuated by Integrator are up-regulated upon IntS6 over-expression
To extend the results from the eGFP reporters to endogenous genes, we generated DL1 cell lines that stably maintain IntS6 or IntS12 transgenes driven by the copper inducible MtnA promoter (Figure 2A). Parental DL1 cells and the inducible cell lines were grown in the presence of Cu2+ for 24 h, which enabled robust induction of IntS6 and IntS12 proteins (15 and 12-fold, respectively) in the stable cell lines (Figures 2B, left and S3A). As a control, we verified that expression of additional Integrator subunits (IntS8, IntS11, and mts) was unchanged upon Cu2+ addition in all cell lines (Figure 2B, left). Total RNA from 3 biological replicates was isolated from each cell line to generate ribosomal RNA depleted RNA-seq libraries (Figure 2A and Table S1) which were then analyzed to identify differentially expressed transcripts (Figure S3B).
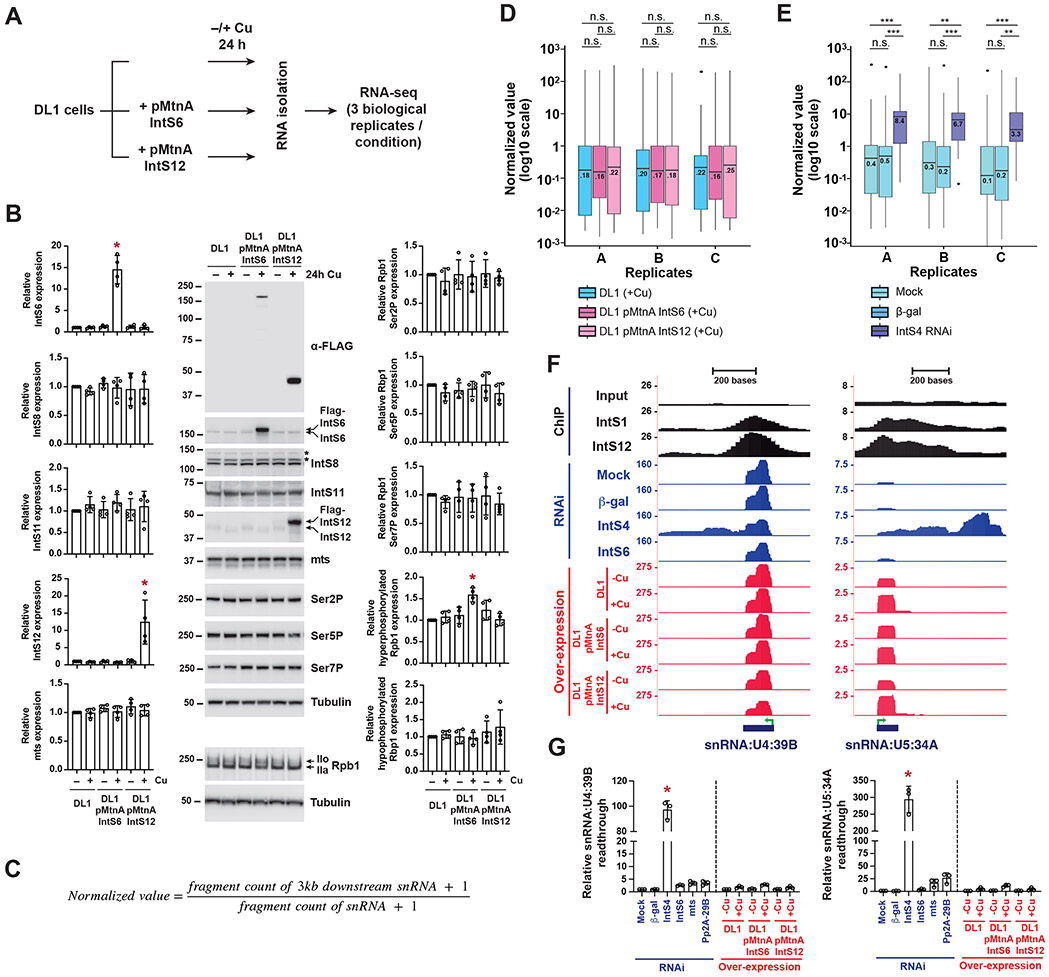
Figure 2. Over-expression of IntS6 does not affect Integrator activity at endogenous Drosophila snRNA loci.
(A) Parental DL1 cells or DL1 cells stably maintaining IntS6 or IntS12 transgenes driven by the copper inducible MtnA promoter were grown for 3 d. 500 μM CuSO4 was added for the last 24 h prior to total RNA isolation from three independent biological replicates. rRNA depleted RNA-seq libraries were then generated, sequenced, and analyzed.
(B) Cell lines were seeded in 12-well plates (5 x 105 cells per well) and grown for 3 d. As indicated, a final concentration of 500 μM CuSO4 was added to cells for the last 24 h prior to harvesting total protein. Western blot analysis was then performed using antibodies that recognize FLAG, IntS6, IntS8, IntS11, IntS12, mts, Rbp1 phosphorylated at Ser2, Ser5, or Ser7, and Rbp1 (C-terminal domain). * denotes non-specific band. IIo denotes hyperphosphorylated Rbp1, while IIa denotes hypophosphorylated Rbp1. α-tubulin was used as a loading control. Subunit expression data were normalized to the parental DL1 cells without CuSO4 treatment and are shown as mean ± SD, N = 3. (*) P < 0.05.
(C) To quantify readthrough transcription downstream of endogenous snRNAs, the levels of RNA-seq fragments that map to the 3 kb downstream of mature snRNA 3' ends were normalized to the levels of fragments that map to mature snRNA sequences.
(D-E) Normalized values of endogenous snRNA readthrough among (D) CuSO4 treated parental DL1 cells and DL1 cells stably maintaining IntS6 or IntS12 transgenes, and (E) DL1 cells subjected to mock, control (β-gal) dsRNA, or IntS4 dsRNA treatments. Center lines represent medians, boxes represent interquartile ranges (IQRs), and whiskers represent extreme data points within 1.5× IQRs. Black points were outliners exceeding 1.5× IQRs. P values were calculated by Wilcoxon signed-rank test. (**) P < 0.01; (***) P < 0.001; n.s., not significant.
(F) UCSC genome browser tracks depicting exemplar snRNA loci. IntS1 and IntS12 ChIP-seq profiles in DL1 cells (GSE114467) are shown in black. RNA-seq data generated from DL1 cells treated for 3 d with control (β-gal), IntS4, or IntS6 dsRNAs are shown in blue. RNA-seq data generated from parental DL1 cells or DL1 cells stably maintaining copper-inducible IntS6 or IntS12 transgenes are shown in red. 500 μM CuSO4 was added for 24 h as indicated. Green arrow, transcription start site (TSS).
(G) Readthrough downstream from snRNA transcripts was quantified using RT-qPCR. Data are shown as mean ± SD, N = 3. (*) P < 0.05.
To first examine effects on endogenous snRNA transcription termination, sequencing reads that mapped within 3 kb downstream of mature snRNA sequences were quantified and normalized to mature snRNA levels (Figure 2C). Over-expression of IntS6 or IntS12 had no significant effect on endogenous snRNA readthrough transcript levels (Figure 2D and Table S2), mirroring the results obtained with the eGFP reporters (Figure 1B, right). In contrast, depletion of IntS4 using RNAi resulted in the expected increase in endogenous snRNA readthrough (median increase >30-fold) due to disabled Integrator endonuclease activity (Figure 2E and Table S2). These expression effects were readily observed at the endogenous U4:39B and U5:34A loci (Figure 2F) and confirmed using RT-qPCR (Figure 2G).
We next examined changes in protein-coding gene expression and found that over-expression of IntS6 (Figures 3A and 3B) but not IntS12 (Figure 3C) resulted in up-regulation of more than 100 genes with almost no genes being down-regulated (fold change > 1.5 and adjusted P < 0.001). Depending on whether the IntS6 over-expression data were compared to parental DL1 cells (Figure 3A) or to the IntS12 over-expression cell line (Figure 3B), slightly different numbers of up-regulated genes were identified but the vast majority (107 genes) overlapped (Figure 3D and Table S3). These 107 genes are enriched in axon generation, protein folding, and heat response pathways (Figure S3C), are significantly longer in length than the average Drosophila gene (Figure S3D), and they have a high Pol II pausing index in parental Drosophila DL1 cells as determined by Precision Run-On sequencing (PRO-seq) (Figures S3E and S3F).27 Compared to genes that become up-regulated in DL1 cells upon RNAi depletion of IntS4 or IntS6, genes affected by IntS6 over-expression normally have even higher PRO-seq signal at the promoter regions coupled to low PRO-seq within the gene body (Figure S3F). Pol II thus typically initiates but fails to transition to productive elongation at these loci.
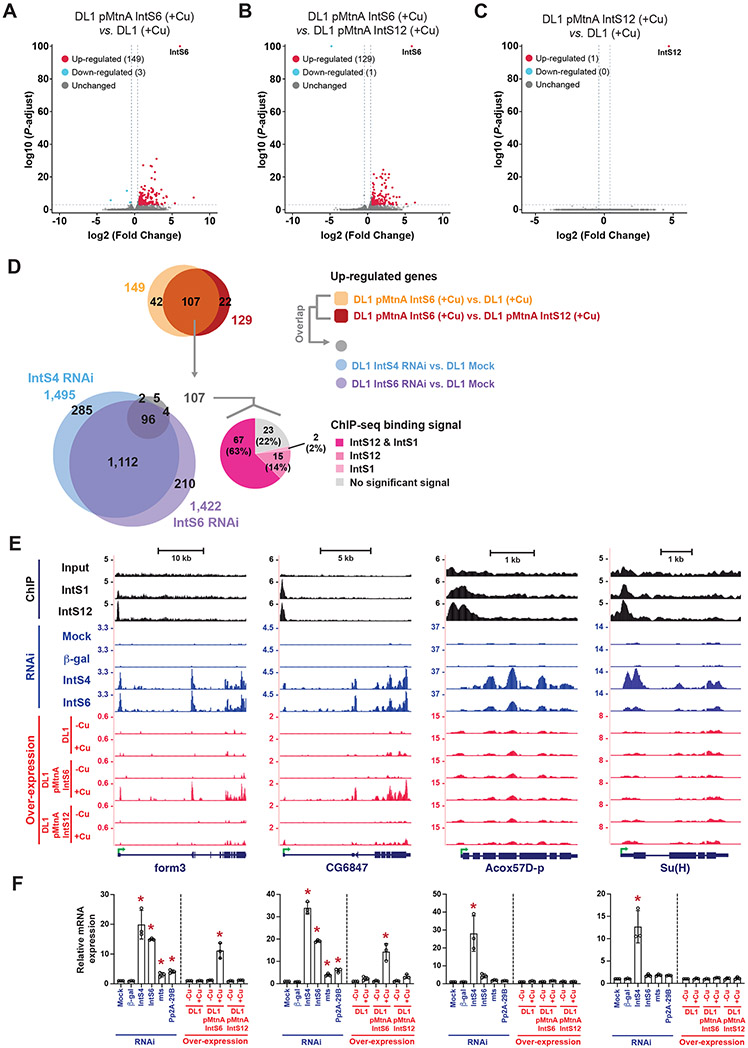
Figure 3. Over-expression of IntS6 blocks Integrator activity at a subset of endogenous Drosophila protein-coding genes.
(A-C) The magnitude of change in mRNA expression compared with statistical significance (adjusted P-value) is shown as volcano plots. Endogenous mRNA expression levels upon IntS6 over-expression were compared to that in parental DL1 cells (A) or upon IntS12 over-expression (B). mRNA expression levels upon IntS12 over-expression were compared to parental DL1 cells (C). Threshold used to define differentially expressed mRNAs was ∣log2(fold change)∣ > 0.585 and adjusted P < 0.001.
(D) The overlapping set of 107 protein-coding genes that were up-regulated upon IntS6 over-expression were compared to the sets of genes up-regulated upon RNAi depletion of IntS4 or IntS6 (left) and the sets of genes bound by IntS12 and/or IntS1 in DL1 cells in previously published (GSE114467) ChIP-seq experiments (right).
(E) UCSC genome browser tracks depicting example protein-coding loci that are (form3, CG6847) or are not (Acox57D-p, Su(H)) affected by IntS6 over-expression. IntS1 and IntS12 ChIP-seq profiles in DL1 cells (GSE114467) are shown in black. RNA-seq data generated from DL1 cells treated for 3 d with control (β-gal), IntS4, or IntS6 dsRNAs are shown in blue. RNA-seq data generated from parental DL1 cells or DL1 cells stably maintaining copper-inducible IntS6 or IntS12 transgenes are shown in red. 500 μM CuSO4 was added for 24 h as indicated. Green arrow, transcription start site (TSS).
(F) Expression of the indicated mRNAs (order as in E) was quantified using RT-qPCR. Data are shown as mean ± SD, N = 3. (*) P < 0.05.
Consistent with the PRO-seq observations, there is evidence that almost all of the 107 genes are directly attenuated by Integrator in DL1 cells under normal conditions (Figure 3D). IntS1 and/or IntS12 chromatin immunoprecipitation sequencing (ChIP-seq) peaks were detected at 84 of the 107 genes (78%) (Figures 3D and S3G-J), and 102 of these genes (95%) were up-regulated (fold change > 1.5 and adjusted P < 0.001) upon RNAi depletion of IntS4 and/or IntS6 in the parental DL1 cells (Figures 3D, S4A, S4B, and Table S4). There was, in fact, a strong positive correlation (~0.8) between the degree of up-regulation observed for these genes upon IntS6 over-expression with that observed upon depletion of IntS4 (Figure S4C) or IntS6 (Figure S4D). Upon examining additional RNA-seq datasets, we found that the vast majority of genes up-regulated upon IntS6 over-expression (89/107 genes, 83%) were also up-regulated upon IntS8 depletion (Figure S4E, left and Table S1).11 In addition, attenuation of many of the 107 genes is clearly dependent on the IntS11 endonuclease as their mRNA levels increased upon depletion of IntS11, and this could be rescued by expression of a wild-type IntS11 transgene but not by a catalytically dead (E203Q) IntS11 transgene (Figure S4E, right and Table S1).9,27
Using the form3 and CG6847 genes as examples, Integrator subunits are normally bound at their 5’ ends in DL1 cells (Figure 3E, top) and the encoded mRNAs were up-regulated when IntS4 or IntS6 was depleted by RNAi (Figure 3E, middle). These mRNAs were likewise up-regulated when IntS6, but not IntS12, was over-expressed (Figure 3E, bottom), and RT-qPCR confirmed the expression changes at both the mRNA (Figure 3F) and pre-mRNA levels (Figure S5A). The latter was done using qPCR primers that amplify across exon-intron boundaries. Similar effects on transcriptional outputs were confirmed at additional exemplar loci (CG8547, Kal1, wun2, and wdp) as shown in Figures S5B-E. To further confirm the unique ability of IntS6 over-expression to inhibit Integrator activity, we generated DL1 cell lines that stably maintain IntS7, IntS13, or IntS14 transgenes driven by the copper inducible MtnA promoter (Figure S5F). Each cell line was grown in the presence of Cu2+ for 24 h to induce robust transgene expression (Figure S5F), but RT-qPCR did not detect any effect of these other over-expressed Integrator subunits on endogenous gene outputs (Figure S5G).
IntS6 over-expression thus uniquely resulted in up-regulation of a set of protein-coding genes normally attenuated by Integrator, but the vast majority of Integrator regulated genes were unaffected by IntS6 over-expression. More than 1,100 genes became up-regulated (fold change > 1.5 and adjusted P < 0.001) when IntS4 or IntS6 was depleted using RNAi (Figures 3D, S4A, S4B, and Table S4) and many of these genes are normally bound by IntS1 and/or IntS12, suggesting they are direct Integrator targets (discussed further in Figure 6). For example, Integrator subunits are bound at the 5’ ends of the Acox57D-p and Su(H) genes (Figure 3E, top) and these mRNAs were up-regulated upon depletion of IntS4 (Figure 3E, middle), yet they were minimally affected by IntS6 over-expression at both the mRNA (Figures 3E, bottom and 3F) and pre-mRNA levels (Figure S6A). Similar expression patterns were observed at the endogenous Ana (Figure S6B) and RnrS loci (Figure S6C), mirroring the lack of effect that was observed when these promoters were tested in the eGFP reporter assay (Figure 1B). We thus conclude that IntS6 over-expression has a dominant-negative effect on Integrator function at only a subset of endogenous target loci.
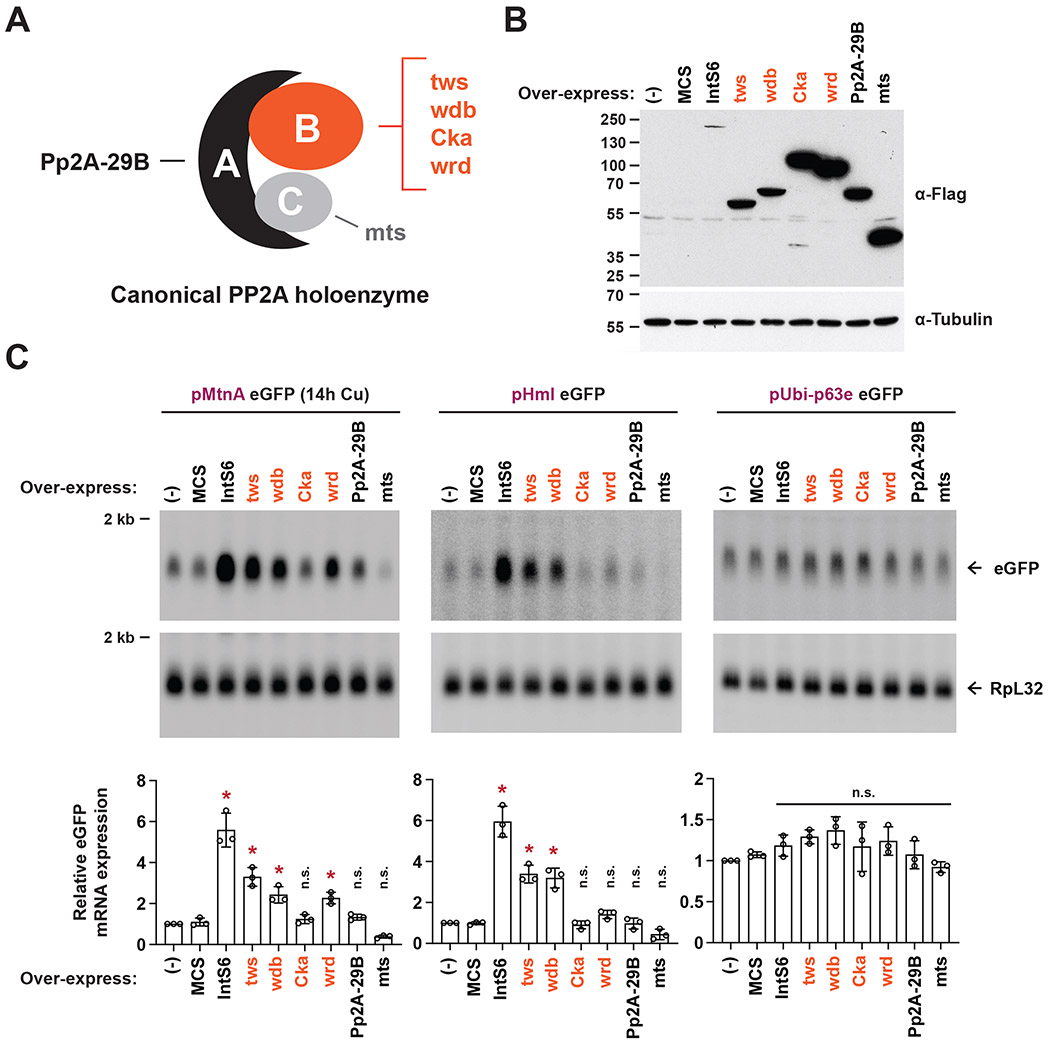
Figure 6. Over-expression of canonical PP2A B subunits can inhibit Integrator activity.
(A) Schematic of the canonical Drosophila PP2A holoenzyme that consists of a scaffolding A subunit (Pp2A-29B), a catalytic subunit (mts), and a variable regulatory B subunit (tws, wdb, Cka, or wrd).
(B) DL1 cells were transfected with 500 ng of the indicated FLAG-tagged expression plasmid and total protein was harvested after 48 h. A plasmid containing a multi-cloning site (MCS) was used as a control. Western blot analysis using an antibody that recognizes FLAG was used to confirm expression of each subunit. α-tubulin was used as a loading control.
(C) DL1 cells were co-transfected with 400 ng of eGFP reporter plasmid and 100 ng of the indicated PP2A subunit over-expression plasmid (driven by the Ubi-p63e promoter). CuSO4 was added for the last 14 h only when measuring eGFP production from the MtnA promoter. Northern blots (20 μg/lane) were used to quantify expression of each eGFP reporter mRNA. Representative blots are shown and RpL32 mRNA was used as a loading control. Data are shown as mean ± SD, N = 3. (*) P < 0.05; n.s., not significant.
IntS6 over-expression blocks Integrator function by titrating PP2A
We next aimed to understand why only a subset of Integrator regulated loci are affected by IntS6 over-expression. Recent cryo-electron microscopy (cryoEM) efforts characterizing the Integrator complex revealed that IntS6 is located far (> 75 angstroms) from the IntS11 endonuclease active site and instead contacts the protein phosphatase 2A (PP2A) subunits (Figure 4A).12,14,18 This led us to hypothesize that IntS6 over-expression may somehow affect the phosphatase module of the Integrator complex (in line with recent work13). PP2A typically functions as a trimer in which a regulatory B subunit (twins (tws), widerborst (wdb), Connector of kinase to AP-1 (Cka), or well-rounded (wrd) in Drosophila) enables the scaffolding (A) and catalytic (C) subunits (Pp2A-29B and microtubule star (mts), respectively, in Drosophila) to be targeted to a variety of substrates in cells.45-47 Both the A (Pp2A-29B) and C (mts) subunits are present in the Integrator complex, but no canonical B subunits have been detected in association with Integrator (Figure 4A).11-14 Eliminating the association of Integrator with PP2A subunits by depleting IntS8,11,12,38 knocking out IntS6,13 or mutating/truncating the N-terminus of IntS811,38 has been shown to increase phosphorylation levels of the C-terminal domain (CTD) of Rbp1, the largest subunit of Pol II. Consistent with these studies, we observed increased (~2 fold) hyperphosphorylated Rpb1 and increased Ser7 CTD phosphorylation when IntS6 or IntS8 was depleted (Figure S7A). We also noted that the levels of IntS6 and IntS8 proteins appear to be co-regulated, as IntS6 protein was depleted when a dsRNA targeting IntS8 was used and vice versa (Figure S7A).
Figure 4. IntS6 over-expression titrates the catalytic subunit of PP2A and causes it to be limiting for Integrator activity.
(A) Cryo-EM structure (PDB: 7PKS) of the human Integrator complex, 14 highlighting the positions of the RNA endonuclease IntS11 (green), IntS6 (orange), and PP2A subunits (teal and pink). There are direct contacts between IntS6 and the PP2A subunits.
(B) DL1 cells were treated for 3 d with control (β-gal), Pp2A-29B, or mts dsRNAs and RNA-seq data generated. The sets of endogenous genes up-regulated upon Pp2A-29B or mts depletion (fold change > 1.5 and adjusted P value < 0.001) were compared to the set of 107 genes that were up-regulated upon over-expression of IntS6.
(C) Parental DL1 cells (Control) and DL1 cells stably maintaining inducible FLAG-tagged IntS6 or IntS12 transgenes were treated with 500 μM CuSO4 for 24 h to induce transgene expression. Immunoprecipitation (IP) using anti-FLAG resin was then performed. Western blots of input nuclear extracts (left, 0.16% input for FLAG and 0.25% for mts) and IP (right, 0.2% eluate for FLAG and 25% for mts) are shown.
(D, E) DL1 cells were co-transfected with 300 ng of eGFP reporter plasmid, 100 ng of IntS6 over-expression plasmid (driven by the Ubi-p63e promoter), and 100 ng of the indicated PP2A subunit (D) or Integrator subunit (E) over-expression plasmid (driven by the Ubi-p63e promoter). Empty vector (pUb 3xFLAG MCS) was added as needed so that 500 ng DNA was transfected in all samples. CuSO4 was added for the last 14 h only when measuring eGFP production from the MtnA promoter. Northern blots (20 μg/lane) were used to quantify expression of each eGFP reporter mRNA. Representative blots are shown and RpL32 mRNA was used as a loading control. Data are shown as mean ± SD, N = 3. (*) P < 0.05; n.s., not significant.
(F) DL1 cells were transfected with 500 ng of the indicated FLAG-tagged expression plasmids and total protein was harvested after 48 h. A plasmid containing a multi-cloning site (MCS) was used as a control. Western blot analysis using an antibody that recognizes FLAG was used to confirm expression of individual Integrator/PP2A subunits. α-tubulin was used as a loading control.
(G) DL1 cells were co-transfected with 300 ng of eGFP reporter plasmid, 100 ng of IntS6 over-expression plasmid (driven by the Ubi-p63e promoter), and 100 ng of the indicated PP2A subunit over-expression plasmid (driven by the Ubi-p63e promoter). CuSO4 was added for the last 14 h and Northern blots (20 μg/lane) were used to quantify expression of the eGFP reporter mRNA. Representative blots are shown and RpL32 mRNA was used as a loading control. Data are shown as mean ± SD, N = 3. (*) P < 0.05; n.s., not significant.
Upon over-expression of IntS6 in DL1 cells, there was also a statistically significant increase (~1.5 fold) in hyperphosphorylated Rpb1 levels, although no change was observed for any of the individual CTD phosphorylation sites (Ser2, Ser5, or Ser7) examined (Figure 2B, right). To address whether Integrator phosphatase activity may indeed be disrupted, we asked whether IntS6 over-expression altered the expression of genes normally controlled by PP2A subunits. Indeed, significant overlap was observed when comparing the list of 107 up-regulated genes to those that were up-regulated when the A (Pp2A-29B) or C (mts) subunit was depleted using RNAi in DL1 cells (Figures 4B, Tables S3 and S4). We thus hypothesized that IntS6 over-expression may sponge or titrate PP2A subunits from the rest of the Integrator complex. Consistent with such a model, the PP2A C (mts) subunit was co-immunoprecipitated with over-expressed FLAG-tagged IntS6, but not with over-expressed IntS12 (Figure 4C). This result nicely mirrors our observation that only IntS6 over-expression (not IntS12) alters Integrator complex function (Figures 1, 2, and 3).
Prior work in S. cerevisiae has shown that the PP2A A subunit is expressed in excess over the C subunit,48 suggesting a possible role for stoichiometry in control of PP2A function. If PP2A subunits are indeed limiting in DL1 cells, we reasoned that over-expressing the C (mts) and/or A (Pp2A-29B) subunits should be sufficient to reverse the gene expression changes caused by IntS6 over-expression. Increasing the expression of the C (mts) subunit, but not the A (Pp2A-29B) subunit was sufficient to reduce the amount of eGFP mRNA produced from the MtnA (Figure 4D, left) and Hml promoter reporters (Figure 4D, middle) to near wild-type levels, indicating a restoration of Integrator activity at these promoters. As expected, there was no change in the output of the control Ubi-p63e promoter, which is not regulated by Integrator (Figure 4D, right). Over-expression of other individual Integrator subunits (Figures 4E and S7B) or catalytically dead PP2A C (mts) subunits that bear mutations that disrupt metal binding (H59Q, D85N, or H241A)49,50 all were unable to rescue the gene expression changes (Figures 4F, 4G, and S7C). Collectively, these data indicate that the phosphatase activity of the PP2A C (mts) subunit can be limiting for Integrator function and that over-expression of IntS6 blocks the ability of PP2A to function at specific gene loci.
The phosphatase module of Integrator is critical at only a subset of Integrator-regulated loci
Given that IntS6 over-expression only affects the outputs of a subset of Integrator regulated eGFP reporters (Figure 1) and endogenous genes (Figures 2 and 3), the titration model suggests that the PP2A phosphatase activity is differentially required across loci for regulation by the Integrator complex. To test this hypothesis, RNAi was used to deplete the PP2A A (Pp2A-29B) or C (mts) subunits (Figure S1A) and we then examined the effect on the outputs of the eGFP reporters (Figure 1B, left). eGFP expression from the MtnA, Hml, and CG8620 promoters all increased upon depletion of PP2A subunits, consistent with a critical role for phosphatase activity in enabling Integrator function at these promoters. In stark contrast, there was no or minimal change in the outputs of the Ana, RnrS, and snRNA readthrough reporters upon depletion of PP2A subunits (Figure 1B, left). These results perfectly mirror the effects observed with IntS6 over-expression: only at promoters where PP2A subunits are required (MtnA, Hml, and CG8620) does IntS6 over-expression result in increased eGFP reporter expression (Figure 1B, right). Promoters unaffected by depletion of PP2A subunits were further only minimally affected by depletion of IntS6 (Figure 1B, left), with modest, but reproducible depressive effects observed that may be due to co-depletion of other Integrator subunits, including IntS8 (Figure S7A) and perhaps IntS5.38
We next examined individual endogenous snRNA (Figure 2G) and protein-coding genes (Figures 3F, S5B-E, S6B, and S6C) using RT-qPCR and again observed that only genes that require PP2A subunits for Integrator function were significantly affected by IntS6 over-expression. For example, the Integrator phosphatase module is not required at the endogenous Acox57D-p and Su(H) genes as there was only a minimal increase in expression of these mRNAs upon depletion of the PP2A A (Pp2A-29B) or C (mts) subunits, and these loci were unaffected when IntS6 was over-expressed (Figure 3F). In contrast, PP2A subunits were required for Integrator function at the form3 and CG6847 loci, and the expression of these genes was increased with IntS6 over-expression (Figure 3F). Mirroring the results with the eGFP reporter genes (Figure 1B, left), the expression of PP2A-independent endogenous loci was only modestly increased upon IntS6 depletion (Figure 3F and S6B).
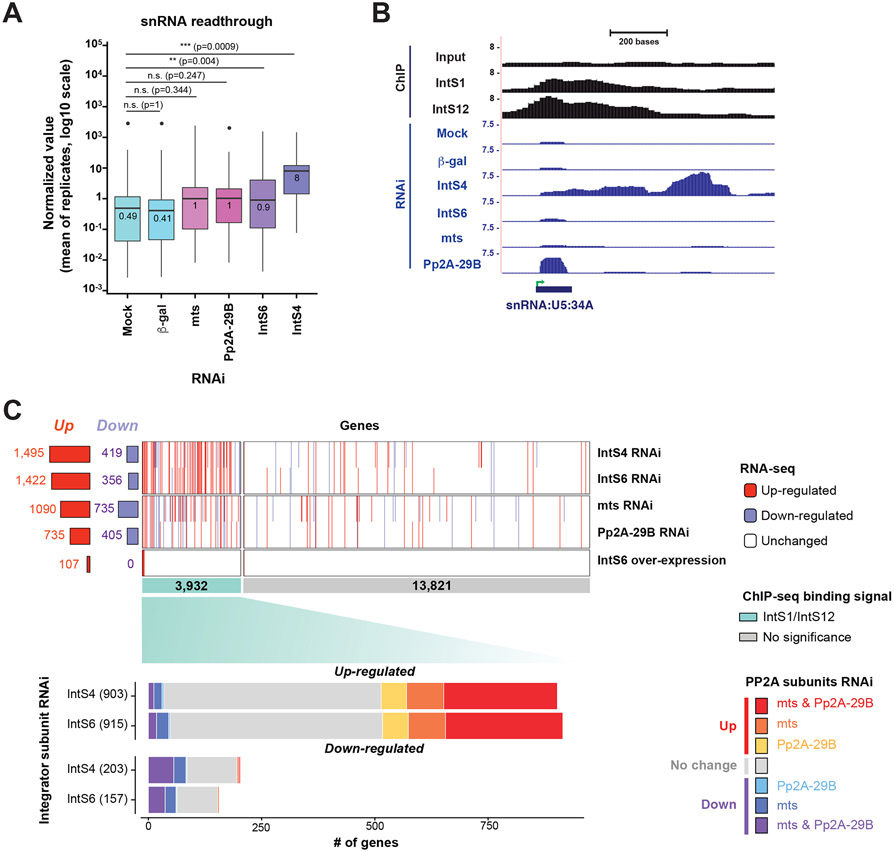
To determine how broadly the phosphatase activity is required for Integrator function across the genome, we first examined readthrough transcription downstream of all endogenous snRNA transcripts. Depletion of PP2A C (mts) or A (Pp2A-29B) resulted in modest effects (compared to IntS4 depletion) that were not statistically significant (Figures 5A and 5B). Note that the levels of mature U5:34A snRNA increased upon depletion of Pp2A-29B (Figure 5B), but similar changes were not observed at the vast majority of snRNA loci, suggesting an idiosyncratic effect particular to this snRNA (Table S2). We then used ChIP-seq data from DL1 cells27 to identify 3,932 protein-coding genes with peaks of IntS1 and/or IntS12 binding located ±1 kb of gene bodies (Figures 5C, middle and S3G-J). Most (>70%) of these peaks were close to TSSs (Figures S3H and S3I) and we reasoned these 3,932 genes represent loci where Integrator may have direct effects. We thus determined if their expression changed upon depletion of IntS4, IntS6, PP2A C (mts), or PP2A A (Pp2A-29B) (Figure 5C, top and Table S4). Consistent with Integrator catalyzing premature transcription termination, most of the protein-coding genes that changed were up-regulated (fold change > 1.5 and adjusted P < 0.001) upon depletion of IntS4 or IntS6 (Figure 5C, top). Of the 3,932 genes bound by Integrator, 903 and 915 were up-regulated upon IntS4 or IntS6 depletion, respectively, and ~40% of these genes were also up-regulated upon depletion of PP2A C (mts) and/or A (Pp2A-29B) (Figure 5C, bottom). This suggests that the phosphatase activity of Integrator may be required at ~40% of protein-coding genes attenuated by Integrator, while it is dispensable at many other genes that are nonetheless still controlled by the complex. This result helps explain why IntS6 over-expression had no effect on most Integrator regulated genes (Figure 3D). The 107 protein-coding genes that were up-regulated by IntS6 over-expression may represent those genes that are most sensitive to tuning by PP2A and indeed these loci had higher Pol II pausing indexes (Figure S3E and S3F).
Figure 5. The phosphatase module is differentially required for Integrator activity across the genome.
(A) Means of normalized values of endogenous snRNA readthrough. Readthrough values (see Figure 2C) were calculated from RNA-seq data of DL1 cells subjected to a mock treatment or treatment with β-gal, mts, Pp2A-29B, IntS6, or IntS4 dsRNAs (3 independent biological replicates). Center lines represent medians, boxes represent interquartile ranges (IQRs), and whiskers represent extreme data points within 1.5× IQRs. Black points were outliners exceeding 1.5× IQRs. P values were calculated by Wilcoxon signed-rank test. (**) P < 0.01; (***) P < 0.001; n.s., not significant.
(B) ChIP-seq and RNA-seq tracks at the U5:34A snRNA locus. IntS1 and IntS12 ChIP-seq profiles in DL1 cells (GSE114467) are shown in black. RNA-seq data generated from DL1 cells treated for 3 d with control (β-gal), IntS4, IntS6, mts, or Pp2A-29B dsRNAs are shown in blue. Green arrow, transcription start site (TSS).
(C) RNA-seq was used to define genes that were up- or down-regulated (∣log2(fold change)∣ > 0.585 and adjusted P < 0.001) upon IntS4, IntS6, mts, or Pp2A-29B depletion using RNAi or upon IntS6 over-expression (top). These gene lists were then stratified by ChIP-seq data that identified 3,932 protein-coding genes with peaks of IntS1 and/or IntS12 binding located ±1 kb of gene bodies in DL1 cells (green, middle). For genes bound by Integrator subunits and differentially regulated upon IntS4 or IntS6 depletion, the effect of depleting PP2A subunits on their expression is graphed (bottom).
IntS6 behaves like a canonical PP2A B subunit
The PP2A holoenzyme typically functions as a heterotrimer consisting of an A, B, and C subunit (Figure 6A), but no canonical B subunits have been detected in the Integrator complex.11-14 This has made it somewhat unclear how the Integrator phosphatase activity is controlled and regulated. Because IntS6 binds the PP2A A and C subunit in Integrator (Figure 4A) and IntS6 over-expression is sufficient to titrate PP2A activity (Figure 4D), we reasoned that IntS6 may be a previously unappreciated regulatory B subunit. If true, this model predicts that over-expression of canonical B subunits (tws, wdb, Cka, or wrd in Drosophila) may likewise be sufficient to titrate the PP2A C subunit and inhibit the attenuation activity of Integrator. We thus individually over-expressed FLAG-tagged versions of each PP2A B subunit (Figure 6B) and examined the functional effects on the eGFP reporters (Figure 6C). Over-expression of the canonical B subunits tws and wdb did indeed lead to increased eGFP expression from the MtnA and Hml promoters, but not from the control Ubi-p63e promoter that is not regulated by Integrator (Figure 6C). It should be noted the effects were not as large as that observed with IntS6 over-expression, despite the FLAG-tagged versions of tws and wdb being expressed at higher levels than FLAG-tagged IntS6 (Figure 6B). In total, these results indicate that IntS6 behaves, at least in some ways, similar to canonical B subunits and that alterations in the levels of B subunits not normally associated with Integrator can still affect the efficiency of select premature transcription termination events.
DISCUSSION
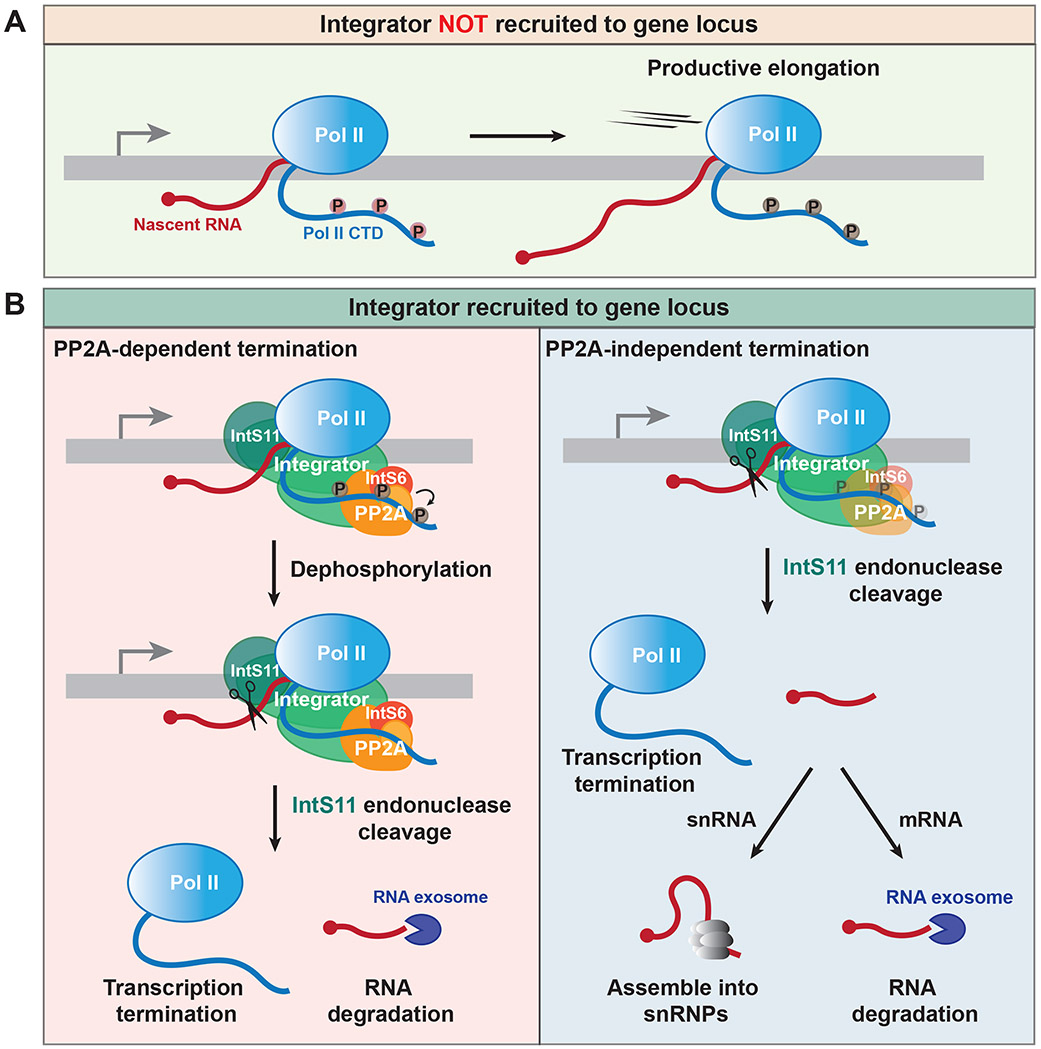
The Integrator complex has RNA endonuclease and protein phosphatase activities, but it has remained unclear if the coordinated action of both activities is required for complex function or if they can be independently harnessed in a locus-specific manner. Here, using reporter genes as well as transcriptomics, we revealed that (i) the phosphatase module is functionally required only at a subset of Drosophila genes that are regulated by Integrator and (ii) that the ability of the PP2A catalytic subunit to act as part of Integrator can be tuned by the levels of IntS6 as well as canonical PP2A regulatory B subunits. Integrator regulates transcription elongation (Figure 7A) and, on one hand, we identified a set of protein-coding genes that became potently de-attenuated when IntS4 (a component of the endonuclease module), IntS6, or PP2A subunit levels were modulated, indicating that the endonuclease and phosphatase modules are both critical for Integrator function at these genes (Figure 7B, left). Meanwhile, cleavage of nascent snRNAs as well as attenuation of many other protein-coding genes required the Integrator endonuclease module but was largely unaffected by modulation of IntS6 or PP2A subunit levels. This suggests the phosphatase module plays little or perhaps even no functional role at these loci (Figure 7B, right). Prior work suggested the phosphatase module is broadly necessary for Integrator function across Drosophila protein-coding genes and snRNAs,11 which contrasts with the locus specificity we observed. Most of the prior contrasting conclusions were based on either (i) depleting IntS8 using RNAi, which we found to also deplete IntS6 protein (Figure S7A) and possibly other closely associated Integrator subunits (e.g. IntS5, as was shown recently in human cells38), or (ii) mutating a 4 amino acid region in IntS8 that binds the Pp2A-29B subunit. Direct roles for PP2A subunits themselves were only previously examined at 4 Drosophila protein-coding genes after RNAi depletion of PP2A subunits or treatment with chemical PP2A inhibitors.11 When the catalytic subunit of PP2A, IntS6, or IntS8 were individually depleted in human cell lines, no or minimal effect on transcription termination of exemplar snRNA genes was observed,12,13 which mirrors the results described here.
Figure 7. Integrator can catalyze transcription termination via PP2A-dependent and - independent mechanisms.
(A) In the absence of Integrator, Pol II is able to productively elongate.
(B) Integrator recruitment facilitates transcription termination. (Left) At some protein-coding genes, the Integrator phosphatase module must act prior to or, at minimum, simultaneously with the IntS11 endonuclease to enable cleavage of the nascent RNA, which is subsequently degraded by the RNA exosome. (Right) In contrast, the phosphatase module is dispensable for Integrator function at snRNA and many other protein-coding gene loci. Cleavage by IntS11 enables stable snRNAs to be produced and prematurely terminated mRNAs to be rapidly degraded. The exact CTD phosphorylation status at these genes when Integrator is acting remains to be determined.
Structural studies using cryoEM have made it increasingly clear that Integrator contains discrete endonuclease, phosphatase, and auxiliary modules that each bind around a core set of Integrator subunits.12,14,18,42,51-53 Overall complex integrity is disrupted upon depletion of a core subunit (e.g., IntS2), while depletion of module specific subunits, including IntS6, does not appreciably affect the binding of other modules.11,13 Our functional data support this concept of Integrator complex modularity as we found that altering IntS6 or PP2A subunit levels had no effect on endonuclease activity at many gene loci (e.g., snRNAs continued to be efficiently processed at their 3' ends). Why then is the Integrator phosphatase module required at a subset of loci? It has been recently shown that PP2A can dephosphorylate components of the Pol II complex, e.g. Spt5 or the C-terminal domain of Pol II itself, to antagonize kinases that stimulate Pol II pause release and productive elongation.11-13 Based on our data, it appears that the order by which the Integrator RNA endonuclease and protein phosphatase activities act can be critical. In particular, at a subset of genes, the Integrator phosphatase module must act prior to or, at minimum, simultaneously with the IntS11 endonuclease to slow/pause Pol II and thereby enable cleavage of the nascent RNA (Figure 7B, left). In the absence of phosphatase activity, the temporal window of opportunity for IntS11 to cleave may be too short and/or IntS11 may fail to achieve an active conformation that accommodates RNA.12,14 The Pol II juggernaut2 is hence unable to be stopped and instead elongates to produce full length mRNAs. Given that RNA cleavage by IntS11 is presumably a non-reversible step, modulating PP2A activity and thus the IntS11 temporal window of opportunity represents an important way that the efficiency of premature termination events can be tuned. Here, we found that changes in protein levels can modulate the activity of the Integrator phosphatase module, and we propose that post-translational modifications (e.g., on IntS6 or the PP2A subunits) or the action of additional binding partners likely also tunes Integrator phosphatase activity.
At genes regulated by Integrator in a manner independent of the phosphatase module (Figure 7B, right), there are presumably alternative mechanisms that enable a sufficiently long temporal window of opportunity for IntS11 to cleave. The exact phosphorylation status of the Pol II CTD when Integrator is acting at these loci remains unclear, however. Ser7 phosphorylation has been implicated as being critical for Integrator recruitment to snRNAs,54-56 and a conserved but relatively degenerate 3' box sequence located 9-19 nucleotides downstream from the 3' ends of mature snRNA transcripts is required for Integrator cleavage.57 Details of the underlying molecular mechanism unfortunately remain poorly understood and sequences resembling a 3' box are not present at most protein-coding genes attenuated by Integrator. Instead, nucleosome occupancy32 or redundant phosphatases may act to stall or pause Pol II near protein-coding promoters, e.g. protein phosphatase 1 (PP1) which is known to help slow Pol II near polyadenylation signals.58-61 Differences in Pol II speed/dynamics caused by histone modifications or RNA sequence/structure may also be at play.
Due to its high abundance, PP2A dephosphorylates many important substrates in eukaryotes and participates in many signaling cascades (for review, see 45,46,47). We nonetheless found that expression of the PP2A catalytic subunit (mts) can be limiting for Integrator function. Increasing the levels of IntS6 or canonical regulatory B subunits was sufficient to titrate PP2A subunits and block Integrator, pointing to a critical role for stoichiometry in controlling PP2A function and Integrator activity. Consistent with this idea, PP2A regulatory B subunits are differentially expressed across tissues and developmental time, and IntS6 is known to be lost or down-regulated in several types of human cancer.62,63 We thus propose that modulating the expression, localization, or modification status of B subunits (including IntS6) may be used by cells to couple rapid dephosphorylation of protein substrates to longer term transcriptional changes via modulation of Integrator activity. It is also noteworthy that IntS6 underwent a gene duplication in many mammals, including humans (but not Drosophila), and it will be interesting in the future to determine whether IntS6 and IntS6-like (IntS6L) modulate PP2A activity in distinct ways and/or at distinct sets of Integrator regulated genes.
In summary, the Integrator complex controls the fates of many nascent RNAs in metazoans and our work has revealed key insights into how its functional modules, especially the IntS6/PP2A phosphatase module, can be differentially employed across gene loci. Our results make clear that the Integrator RNA endonuclease and protein phosphatase activities do not always have to be coordinated with one another, but instead can act independently (consistent with recent work38). Modulation of phosphatase activity by IntS6 is sufficient to disable Integrator at some loci, and we anticipate there will be additional mechanisms that tune Integrator in a gene-specific manner to meet a cell’s ever-changing transcriptional needs.
Limitations of the study
It remains unclear if all the Integrator complex modules are normally recruited as a group in vivo or if they can be selectively assembled at a gene locus. In addition, the exact characteristics that cause a gene locus to require the phosphatase module for Integrator activity need to be defined. Future work will be required to clarify if IntS6 over-expression affects PP2A activity beyond the Integrator complex as well as to define how the IntS6-PP2A interaction is naturally modulated, including in species beyond Drosophila, to tune Integrator activity at specific loci in vivo.
STAR Methods
RESOURCE AVAILABILITY
Lead Contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Jeremy E. Wilusz (jeremy.wilusz@bcm.edu).
Materials Availability
All unique materials generated in this study are available upon request. Plasmids generated in this study have been deposited to Addgene.
Data and Code Availability
RNA-seq data have been deposited at GEO and original Northern and western blot images have been deposited at Mendeley. All data are publicly available as of the date of publication. Accession numbers and DOIs are listed in the key resources table.
This paper does not report original code.
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Drosophila IntS6 | This paper | UP2581 Bleed 3 |
| Drosophila IntS8 | This paper | UP2577 Bleed 3 |
| Drosophila IntS11 | This paper | BCM444 Bleed 2 |
| Drosophila IntS12 | This paper | BCM446 Bleed 2 |
| FLAG | Sigma #F1804-1MG | AB_262044 |
| α-tubulin | Sigma #T6074 | AB_477582 |
| PP2A C (mts) | Cell Signaling #2038 | AB_2169495 |
| RNA Pol II CTD (Clone 8WG16) | Sigma #05-952-I-100UG | AB_492629 |
| RNA Pol II CTD phospho Ser2 (Clone 3E10) | Active Motif #61984 | AB_2687450 |
| RNA Pol II CTD phospho Ser5 (Clone 3E8) | Active Motif #61986 | AB_2687451 |
| RNA Pol II CTD phospho Ser7 (Clone 4E12) | Active Motif #61987 | AB_2687452 |
| Anti-FLAG beads | Sigma #A2220 | AB_10063035 |
| Amersham ECL Rabbit IgG, HRP-linked whole Ab | Cytiva #NA934 | AB_772206 |
| Amersham ECL Mouse IgG, HRP-linked whole Ab | Cytiva #NA931 | AB_772210 |
| Anti-rat IgG, HRP-linked Antibody | Cell Signaling #7077S | AB_10694715 |
| Bacterial and virus strains | ||
| Biological samples | ||
| Chemicals, peptides, and recombinant proteins | ||
| Effectene | Qiagen | Cat #301427 |
| Copper sulfate | Fisher BioReagents | Cat #BP346-500 |
| Trizol | ThermoFisher Scientific | Cat #15596018 |
| Protease inhibitors | Roche | Cat #11836170001 |
| Critical commercial assays | ||
| MEGAscript Kit | ThermoFisher Scientific | Cat #AMB13345 |
| NorthernMax Formaldehyde Load Dye | ThermoFisher Scientific | Cat #AM8552 |
| NorthernMax 10X Running Buffer | ThermoFisher Scientific | Cat #AM8671 |
| NorthernMax 10X Denaturing Gel Buffer | ThermoFisher Scientific | Cat #AM8676 |
| ULTRAhyb-Oligo | ThermoFisher Scientific | Cat #AM8663 |
| NuPAGE 4-12% Bis-Tris gels | ThermoFisher Scientific | Cat #NP0323 |
| PVDF membrane | Bio-Rad | Cat #1620177 |
| NuPAGE 3-8% Tris-Acetate gels | ThermoFisher Scientific | Cat #EA03755 |
| SuperSignal West Pico PLUS Chemiluminescent Substrate | ThermoFisher Scientific | Cat #PI34080 |
| TURBO DNase | ThermoFisher Scientific | Cat #AM2238 |
| TaqMan Reverse Transcription Reagents | ThermoFisher Scientific | Cat #N8080234 |
| 2x Power SYBR Green PCR Master Mix | ThermoFisher Scientific | Cat #4368708 |
| Deposited data | ||
| RNA-seq from DL1 cells, DL1 cells over-expressing IntS6, and DL1 cells over-expressing IntS12 | This paper | GEO: GSE223973 |
| RNA-seq from DL1 cells depleted of IntS4, IntS6, mts, or PP2A-29B | This paper | GEO: GSE223974 |
| PRO-seq from DL1 cells treated with β-gal dsRNA | Elrod et al., 201927 | GEO: GSE114467 |
| RNA-seq from DL1 cells depleted of IntS11 and rescued with WT IntS11 or E203Q mutant IntS11 | Elrod et al., 201927 | GEO: GSE114467 |
| RNA-seq from DL1 cells depleted of IntS8 and rescued with WT IntS8 or WFEF/A mutant IntS8 | Huang et al., 202011 | GEO: GSE150844 |
| IntS1 and IntS12 ChIP-seq from DL1 cells | Elrod et al., 201927 | GEO: GSE114467 |
| Raw image files | Mendeley | 10.17632/r46b2sgkkb.1 |
| Experimental models: Cell lines | ||
| DL1 | Sara Cherry, Univ of Pennsylvania | CVCL_Z231 |
| DL1 pMtnA IntS6 | This paper | N/A |
| DL1 pMtnA IntS12 | This paper | N/A |
| DL1 pMtnA IntS7 | This paper | N/A |
| DL1 pMtnA IntS13 | This paper | N/A |
| DL1 pMtnA IntS14 | This paper | N/A |
| Experimental models: Organisms/strains | ||
| Oligonucleotides | ||
| Table S5 | This paper | N/A |
| Recombinant DNA | ||
| Hy_pMT eGFP SV40 | Kramer et al., 201566 | Addgene #69911 |
| Hy_pHml eGFP SV40 | Tatomer et al., 201928 | Addgene #132645 |
| Hy_pCG8620 eGFP SV40 | Tatomer et al., 201928 | Addgene #132646 |
| Hy_pUbi-p63e eGFP SV40 | Tatomer et al., 201928 | Addgene #132650 |
| Hy_pRnrS eGFP SV40 | This paper | Addgene #195062 |
| Hy_pAna eGFP SV40 | This paper | Addgene #195063 |
| pUC U4:39B eGFP | Chen et al., 201344 | N/A |
| Hy_U5:34A eGFP SV40 | This paper | Addgene #195064 |
| pUb 3xFLAG MCS | Chen et al., 201267 | N/A |
| pUb FLAG-Mts | This paper | Addgene #195065 |
| pUb FLAG-Mts H59Q | This paper | Addgene #208402 |
| pUb FLAG-Mts D85N | This paper | Addgene #208403 |
| pUb FLAG-Mts H241A | This paper | Addgene #208404 |
| pUb FLAG-Pp2A-29B | This paper | Addgene #195066 |
| pUb FLAG-Cka | This paper | Addgene #195067 |
| pUb FLAG-tws | This paper | Addgene #195068 |
| pUb FLAG-wdb | This paper | Addgene #195069 |
| pUb FLAG-wrd | This paper | Addgene #195070 |
| pUb FLAG-IntS6 AA 1-400 | This paper | Addgene #195071 |
| pUb FLAG-IntS6 AA 1-600 | This paper | Addgene #195072 |
| pUb FLAG-IntS6 AA 101-1284 | This paper | Addgene #195073 |
| pUb FLAG-IntS6 AA 1197-1284 | This paper | Addgene #195074 |
| pUb FLAG-IntS6 AA 101-1200 | This paper | Addgene #195075 |
| pUb FLAG-Human IntS6 | This paper | Addgene #198408 |
| pUb FLAG-Human IntS6L | This paper | Addgene #198409 |
| pUb FLAG-Zebrafish IntS6 | This paper | Addgene #198410 |
| pUb FLAG-Zebrafish IntS6L | This paper | Addgene #198411 |
| pUb FLAG-IntS1 | Tatomer et al., 201928 | N/A |
| pUb FLAG-IntS2 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS3 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS4 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS5 | Tatomer et al., 201928 | N/A |
| pUb FLAG-IntS6 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS7 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS8 | Huang et al., 202011 | N/A |
| pUb FLAG-IntS9 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS10 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS11 | Tatomer et al., 201928 | N/A |
| pUb FLAG-IntS12 | Chen et al., 201344 | N/A |
| pUb FLAG-IntS13 | Mascibroda et al., 202234 | N/A |
| pUb FLAG-IntS14 | Eric Wagner, Univ of Rochester | N/A |
| pMT FLAG MCS puro | Elrod et al., 201927 | N/A |
| pMtnA FLAG-IntS6 puro | This paper | Addgene #195076 |
| pMtnA FLAG-IntS7 puro | This paper | Addgene #208405 |
| pMtnA FLAG-IntS12 puro | This paper | Addgene #195077 |
| pMtnA FLAG-IntS13 puro | This paper | Addgene #208406 |
| pMtnA FLAG-IntS14 puro | This paper | Addgene #208407 |
| pRSFDuet-1 | Novagen | Cat #71341 |
| pRSFDuet-1 IntS6 AA 1035-1284 | This paper | Addgene #196904 |
| pRSFDuet-1 IntS8 AA 1-308 | This paper | Addgene #196905 |
| pRSFDuet-1 IntS11 AA 300-597 | This paper | Addgene #199329 |
| pGEX-6P-1 | Amersham | Cat #27-4597-01 |
| pGEX-6P-1 IntS12 | This paper | Addgene #210521 |
| Software and algorithms | ||
| Trimmomatic | Bolger et al., 201468 | N/A |
| ImageQuant | Cytiva Life Sciences | N/A |
| HISAT2 | Kim et al., 201969 | N/A |
| featureCounts | Liao et al., 201470 | N/A |
| DESeq2 | Love et al., 201465 | N/A |
| R 3.6.3 | https://www.r-project.org | N/A |
| clusterProfiler | Yu et al., 201264 | N/A |
| bowtie2 | Langmead and Salzberg, 201271 | N/A |
| sambamba | Tarasov et al., 201572 | N/A |
| MACS2 | Zhang et al., 200873 | N/A |
| ChIPseeker | Yu et al., 201574 | N/A |
| GenomicFeatures | Lawrence et al., 201375 | N/A |
| Prism v10.0.0 | GraphPad | N/A |
| Bowtie | Langmead et al., 200976 | N/A |
| Other | ||
EXPERIMENTAL MODEL AND STUDY PARTICIPANT DETAILS
Cell lines
Drosophila DL1 cells were cultured at 25°C in Schneider’s Drosophila medium (Thermo Fisher Scientific 21720024), supplemented with 10% (v/v) fetal bovine serum (HyClone SH30396.03), 1% (v/v) penicillin-streptomycin (Thermo Fisher Scientific 15140122), and 1% (v/v) L-glutamine (Thermo Fisher Scientific 35050061).
METHOD DETAILS
Expression plasmids
Drosophila reporter plasmids expressing eGFP under the control of the inducible Metallothionein A promoter (Hy_pMT eGFP SV40; Addgene #69911), Hml promoter (Hy_pHml eGFP SV40; Addgene #132645), CG8620 promoter (Hy_pCG8620 eGFP SV40; Addgene #132646), or Ubi-p63e promoter (Hy_pUbi-p63e eGFP SV40; Addgene #132650) were described previously.28,66 Reporter plasmids expressing eGFP under the control of the RnrS promoter (Hy_pRnrS eGFP SV40, Addgene #195062) or Ana promoter (Hy_pAna eGFP SV40, Addgene #195063) were generated from Hy_pPepck1 eGFP SV40 (Addgene #132644) as detailed in Methods S1. Plasmids expressing individual Integrator or PP2A subunits under the control of the Ubi-p63e promoter were generated using the previously described pUb 3xFLAG MCS plasmid67 as detailed in Methods S1. Full-length IntS1-14 expression plasmids were obtained from Eric J. Wagner, University of Rochester. Plasmids expressing IntS6 (pMtnA FLAG-IntS6 puro, Addgene #195076), IntS7 (pMtnA FLAG-IntS7 puro, Addgene #208405), IntS12 (pMtnA FLAG-IntS12 puro, Addgene #195077), IntS13 (pMtnA FLAG-IntS13 puro, Addgene #208406), or IntS14 (pMtnA FLAG-IntS14 puro, Addgene #208407) under the control of the Metallothionein A promoter were generated using the previously described pMT FLAG MCS puro plasmid.27 The reporter plasmid expressing eGFP downstream of the U4:39B snRNA gene was previously described.44 An analogous reporter expressing eGFP downstream of U5:34A snRNA (Hy_U5:34A eGFP SV40, Addgene #195064) was generated from Hy_pPepck1 eGFP SV40. Details of all cloning, including full plasmid sequences, are described in Methods S1.
Double-stranded RNAs
Double-stranded RNAs (dsRNAs) from the DRSC (Drosophila RNAi Screening Center) were generated by in vitro transcription (MEGAscript kit, Thermo Fisher Scientific AMB13345) of PCR templates containing the T7 promoter sequence on both ends as previously described in detail.77 Primer sequences are provided in Table S5.
Generation of stable cell lines
To generate DL1 cells stably maintaining the inducible IntS6, IntS7, IntS12, IntS13, or IntS14 transgenes, 2 x 106 cells were first plated in complete media in 6-well dishes. After 24 h, 2 μg of pMtnA FLAG-IntS6 puro (Addgene #195076), pMtnA FLAG-IntS7 puro (Addgene #208405), pMtnA FLAG-IntS12 puro (Addgene #195077), pMtnA FLAG-IntS13 puro (Addgene #208406), or pMtnA FLAG-IntS14 puro (Addgene #208407) plasmid was transfected using Effectene (Qiagen 301427; 16 μL Enhancer and 30 μL Effectene Reagent). On the following day, 5 μg/mL puromycin was added to the media to select and maintain the cell population.
Transfections, RNAi, and RNA isolation
To determine the effect of Integrator subunit over-expression on the output of the eGFP reporters, DL1 cells were seeded in 12-well plates (5 x 105 cells per well) in complete Schneider’s Drosophila medium and cultured overnight. On the following day, 500 ng of plasmid DNA was transfected into each well using Effectene (4 μL of enhancer and 5 μL of Effectene reagent; Qiagen 301427). Unless otherwise noted, 400 ng of eGFP plasmid was transfected along with 100 ng of the Integrator/PP2A subunit expression plasmid. Total RNA was isolated ~48 h later using TRIzol (Thermo Fisher Scientific 15596018) according to the manufacturer’s instructions.
To determine the effect of Integrator subunit down-regulation on the output of the eGFP reporters, 1 x 106 DL1 cells were first bathed in 2 μg of dsRNAs in 12-well plates in 0.5 mL Schneider’s Drosophila medium without FBS, penicillin/streptomycin, or L-glutamine. After incubating cells at 25°C for 45 min, 1 mL of complete Drosophila medium was added and cells were grown at 25°C. After 24 h, Effectene (4 μL of enhancer and 5 μL of Effectene reagent; Qiagen 301427) was used to transfect 500 ng of the indicated eGFP plasmid. Total RNA was isolated ~48 h later using TRIzol (Thermo Fisher Scientific 15596018) according to the manufacturer’s instructions. When examining reporters driven by the Metallothionein A promoter, a final concentration of 500 μM copper sulfate (Fisher BioReagents BP346-500) was added to cells for the last 14 h prior to RNA isolation.
Northern blotting
Northern blots using 1.2% denaturing formaldehyde agarose gels, NorthernMax reagents (Thermo Fisher Scientific), and oligonucleotide probes were performed as previously described in detail.78 Oligonucleotide probe sequences are provided in Table S5. Blots were hybridized overnight at 42°C with oligonucleotide probes in ULTRAhyb-Oligo (Thermo Fisher Scientific AM8663), washed two times with 2x SSC, 0.5% SDS, and viewed with the Amersham Typhoon scanner (Cytiva) followed by quantification using ImageQuant (Cytiva).
Antibody production
The C-terminal region of Drosophila IntS6 (amino acids 1035-1284) and the N-terminal region of Drosophila IntS8 (amino acids 1-308) were individually cloned into pRSFDuet-1 using the BamHI and HindIII restriction enzyme sites to generate pRSFDuet-1 IntS6 AA 1035-1284 (Addgene #196904) and pRSFDuet-1 IntS8 AA 1-308 (Addgene #196905), respectively. The C-terminal region of Drosophila IntS11 (amino acids 300-597) was cloned into pRSFDuet-1 using the SalI and HindIII restriction enzyme sites to generate pRSFDuet-1 IntS11 AA 300-597 (Addgene #199329). Full length Drosophila IntS12 was cloned into pGEX-6P-1 using EcoRI and NotI to generate pGEX-6P-1 IntS12 (Addgene #210521). Details of cloning, including full plasmid sequences, are described in Methods S1. Plasmids were transformed into BL21 Star (DE3) E. coli and grown in terrific broth media supplemented with 50 μg/mL kanamycin for IntS6, IntS8, and IntS11, and with 100 μg/mL ampicillin for IntS12. Expression of the His-tagged proteins was induced at OD600 ~0.8 by addition of 0.3 mM IPTG. IntS6, IntS8, IntS11, and IntS12 cultures were incubated at 16°C for 20 h, 25°C for 7 h, 16°C for 20 h, and 16°C for 16 h, respectively, before cells were harvested. All purification steps were carried out at 4°C unless otherwise noted.
For purification of His-tagged IntS6, the cell pellet was lysed by sonication in lysis buffer (50 mM Tris pH 8.0, 500 mM NaCl, 0.5 mM DTT, 25 mM imidazole, 1 mM PMSF, 100 μM leupeptin, 10 μM pepstatin A, and 1 mM benzamidine). Cell debris was removed by centrifugation at 20,000 x g for 15 min. The supernatant was filtered and loaded onto a Ni-column (Cytiva 17524801). The column was then washed with wash buffer (30 mM Tris pH 8.0, 300 mM NaCl, 50 mM imidazole) and eluted by gradient of imidazole to 400 mM. Fractions containing His-tagged IntS6 were pooled, concentrated, and loaded onto a Superdex 200 column in PBS. Fractions containing His-tagged IntS6 were pooled, concentrated, flash frozen, and stored at −80°C.
His-tagged IntS8 and IntS11 were purified under denaturing conditions. Cells were resuspended and incubated in lysis buffer (30 mM Tris pH 7.5, 300 mM NaCl, 1 mM DTT, 1 mM EDTA, 100 μg/mL lysozyme, 1 mM PMSF, 100 μM leupeptin, 10 μM pepstatin A, and 1 mM benzamidine) on ice for 30 min, and then sonicated for a total of 5 min. Sample was pelleted by centrifugation at 20,000 x g for 15 min and then resuspended in denaturing buffer (30 mM Tris pH 7.5, 300 mM NaCl, and 8 M urea) at 4°C for 30 min while stirring. Cell debris was removed by centrifugation at 20,000 x g for 15 min. The solubilized IntS8 protein was diluted with buffer to lower the urea concentration to 1 M, loaded onto a Ni-column (Cytiva 29051021), and eluted by adding elution buffer (30 mM Tris pH 7.5, 300 mM NaCl, and 500 mM imidazole). Fractions containing His-tagged IntS8 were pooled, dialyzed against PBS, concentrated, flash frozen, and stored at −80°C. The solubilized IntS11 protein in denaturing buffer was flash frozen and stored at −80°C.
For purification of IntS12, the cell pellet was lysed by sonication in lysis buffer (50 mM Tris pH 7.5, 300 mM NaCl, 5 mM DTT, 0.1% Triton X-100, 1 mM PMSF, 100 μM leupeptin, 10 μM pepstatin A, and 1 mM benzamidine). Cell debris was removed by centrifugation at 12,000 x g for 1 h. The supernatant was incubated with pre-equilibrated Glutathione Sepharose 4B resin for 40 min while rotating. The protein bound resin was washed with lysis buffer, then with buffer A (50 mM Tris pH 7.5, 1M NaCl, 3 mM DTT, 1 mM PMSF, 100 μM leupeptin, 10 μM pepstatin A, and 1 mM benzamidine), and then with buffer B (50 mM Tris pH 7.5, 100 mM NaCl, 3 mM DTT, and 5% glycerol). IntS12 was released from the resin by incubating the resin with PreScission protease for 17 h at 4°C. IntS12 was eluted with buffer B and fractions containing IntS12 were passed through a heparin column to remove contamination. Flow through was collected, dialyzed in PBS, concentrated, flash frozen, and stored at −80°C.
Purified His-tagged IntS6, IntS8, and IntS12 proteins in PBS were shipped to a commercial vendor (Cocalico Biologicals) and used to inoculate rabbits. His-tagged IntS11 was purified from SDS-PAGE gels and the cut bands were likewise used to inoculate rabbits. The reactivity and specificity of antisera was confirmed with Western blots using whole cell extracts from DL1 cells treated for 3 d with dsRNAs targeting either IntS6, IntS8, IntS11, or IntS12.
Western blotting
In experiments where Pol II levels were analyzed using western blots, cells were resuspended in RIPA buffer (150 mM NaCl, 1% Triton X-100, 50 mM Tris pH 7.5, 0.1% SDS, 0.5% sodium deoxycholate, 1% NP-40) supplemented with protease/phosphatase inhibitor (Cell Signaling 5872S) and benzonase nuclease (Sigma E1014-25KU) and then incubated on ice for 1 h. For western blot experiments where Pol II levels were not measured, cells were resuspended in RIPA buffer supplemented with protease inhibitors (Roche 11836170001) and incubated on ice for 20 min. Cell lysates were cleared at 21,000 x g for 20 min at 4°C and protein concentrations were measured using a standard Bradford assay (Bio-Rad 5000006). Lysates containing 20 μg protein were then resolved on NuPAGE 3-8% Tris-Acetate gels (Thermo Fisher Scientific EA03755) when the Rbp1 CTD antibody was used to resolve hyper- and hypophosphorylated isoforms. For Westerns using all other antibodies, lysates containing 20 μg protein were resolved on NuPAGE 4-12% Bis-Tris gels (Thermo Fisher Scientific NP0323). All gels were transferred to PVDF membranes (Bio-Rad 1620177). Membranes were blocked with 10% nonfat milk in TBST for 1 h before incubation in primary antibody (diluted in 1x TBST) overnight at 4°C. Membranes were then washed with 1x TBST (4 x 10 min) followed by incubation in secondary antibody (diluted in 1x TBST) at room temperature for 1 h. Antibody incubation conditions are summarized in Table S6. Membranes were washed with 1x TBST (4 x 10 min) and processed using SuperSignal West Pico PLUS Chemiluminescent Substrate (Thermo Fisher Scientific PI34080). The intensity of protein bands were quantified using ImageQuantTL (Cytiva).
RT-qPCR
5 μg of total RNA (quantified by Nanodrop) was digested with TURBO DNase (Thermo Fisher Scientific AM2238) in a 20 μL reaction following the manufacturer’s protocol. Samples were then incubated at 75°C for 10 min in the presence of 15 mM EDTA. 1 μg of the digested RNA was reverse transcribed to cDNA in a 20 μL reaction using TaqMan Reverse Transcription Reagents (Thermo Fisher Scientific N8080234) with random hexamers following the manufacturer’s protocol except that 4 μL of 25 mM MgCl2 instead of 1.4 μL was used. RT-qPCR reactions were performed in 15 μL reactions that contained 1.5 μL of cDNA (diluted up to 10-fold in H2O), 7.5 μL 2x Power SYBR Green PCR Master Mix (Thermo Fisher Scientific 4368708), and 6 μL 1.5 μM gene-specific primer pairs. Primer sequences are provided in Table S5.
Using the QuantStudio 3 Real-Time PCR System (Thermo Fisher Scientific A28566) and clear plates (Thermo Fisher Scientific 4346907), the following cycling conditions were used: 95°C for 10 min, 40 amplification cycles of 95°C for 15 s followed by 60°C for 1 min, and a final melting cycle of 95°C for 10 s, 65°C for 1 min, and 97°C for 1 s. Subsequently, a melt curve was performed to verify that amplified products were a single discrete species. Threshold cycle (CT) values were calculated by the QuantStudio 3 system and relative transcript levels (compared to RpL32) were calculated using the 2−ΔΔCT method. RT-qPCR reactions were performed using at least three independent biological replicates, with each replicate having two technical replicates.
Immunoprecipitation
Parental DL1 cells and DL1 cells stably maintaining inducible FLAG-tagged IntS6 or IntS12 transgenes (DL1 pMtnA IntS6 and DL1 pMtnA IntS12, respectively) were grown in T75 flasks (4 flasks/each) for 3 d and 500 μM copper sulfate (Fisher BioReagents BP346-500) was added for the last 24 h. Cells were harvested by centrifugation at 340 x g for 5 min and then washed with ice-cold PBS. Cells were resuspended in hypotonic buffer (50 mM Tris pH 7.5, 10 mM KCl, 1 mM DTT, and protease inhibitor mix containing 1 mM PMSF, 100 μM leupeptin, 10 μM pepstatin A, and 1 mM benzamidine) and incubated for 20 min on ice. Cell suspension was then homogenized using a glass Dounce homogenizer. Nuclei were pelleted at 1000 x g for 10 min at 4°C and washed once with hypotonic buffer. Nuclei were incubated in lysis buffer (40 mM Tris pH 7.5, 300 mM NaCl, 1 mM DTT, 10% glycerol, 0.75% Triton X-100 and protease inhibitors) for 20 min on ice. Insoluble proteins and cell debris were removed by centrifugation at 21,000 x g for 30 min and the supernatant was passed through a 0.45 μM filter (Thermo Fisher Scientific F2513-14). The supernatant was diluted with dilution buffer (20 mM Tris pH 7.5, 10 mM NaCl, 1 mM DTT, 10% glycerol, and protease inhibitors) such that the final NaCl and Triton X-100 concentrations became 200 mM and 0.5%, respectively. The diluted supernatant was then incubated with pre-equilibrated anti-FLAG beads (Sigma A2220) while rotating for 2 h at 4°C. The beads were washed five times with wash buffer (20 mM Tris pH 7.5, 150 mM NaCl, 1 mM DTT, 10% glycerol, 0.25% Triton X-100, and protease inhibitors). Bound proteins were eluted with 0.1 M glycine pH 3.5, TCA precipitated, and resuspended in 1x loading dye containing 5 mM DTT.
RNA-seq library generation
To determine the effect of Integrator subunit over-expression on the endogenous transcriptome, DL1, DL1 pMtnA IntS6, and DL1 pMtnA IntS12 cells were seeded in 6-well plates (2 x 106 cells per well) in 2 mL complete Schneider’s Drosophila medium (with FBS, penicillin/streptomycin, and L-glutamine) and grown for 3 d. As indicated, a final concentration of 500 μM copper sulfate (Fisher BioReagents BP346-500) was added to cells for the last 24 h prior to RNA isolation using TRIzol (Thermo Fisher Scientific 15596018). Total RNA was isolated from three biological replicates, treated with DNase I, depleted of ribosomal RNAs, and strand-specific RNA-seq libraries were generated by Genewiz/Azenta Life Sciences. Libraries were sequenced using Illumina HiSeq, 2 x 150 bp configuration.
To determine the effect of Integrator/PP2A subunit down-regulation on the endogenous transcriptome, 5 x 105 DL1 cells were bathed in 2 μg of dsRNAs in 12-well plates in 0.5 mL Schneider’s Drosophila medium without FBS, penicillin/streptomycin, or L-glutamine. After incubating cells at 25°C for 45 min, 1 mL of complete Drosophila medium was added and cells were grown at 25°C for 3 d. Total RNA was isolated using TRIzol (Thermo Fisher Scientific 15596018) according to the manufacturer’s instructions. RNAs were treated with DNase I, depleted of ribosomal RNAs, and strand-specific RNA-seq libraries were generated by Genewiz/Azenta Life Sciences. Libraries were sequenced using Illumina HiSeq, 2 x 150 bp configuration.
The GEO accession numbers for the RNA-seq datasets generated in this study are GSE223973 and GSE223974.
QUANTIFICATION AND STATISTICAL ANALYSIS
RNA-seq analysis
For paired-end RNA-seq samples generated in this study, Trimmomatic68 (version 0.39; parameters: PE -threads 3 -phred33 TruSeq3PE-2.fa:2:30:8:true LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:30) was used for quality control of RNA-seq datasets, including removal of adaptor sequences and low-quality bases at both ends of reads. Reads were next aligned to the D. melanogaster dm6/BDGP6.22 reference genome using HISAT269 (version 2.1.0, parameters: --no-softclip --score-min L,−16,0 --mp 7,7 --rfg 0,7 --rdg 0,7 --dta -k 1 --max-seeds 20). Mapping summaries of RNA-seq datasets are described in Table S1. Fragment counts were calculated per gene using featureCounts70 (version 2.0.1, parameters: -s 2 -p --fraction -O -T 16 -t exon -g gene_id) in a strand-specific manner based on the D. melanogaster Ensembl gene annotation (BDGP6.22.96). Differentially expressed genes (DEGs) were then identified using DESeq265 (version: 1.26.0) under R 3.6.3 with a threshold of an adjusted P value < 0.001 and ∣log2(fold change, FC)∣ > 0.585. Pearson correlation coefficient (PCC) of FC of 96 up-regulated genes between IntS6 over-expression, IntS4 depletion, and IntS6 depletion were calculated under R 3.6.3. GO enrichment analysis (biological process) for the 107 genes up-regulated with IntS6 over-expression was performed and visualized with clusterProfiler64 (version 3.14.3, parameters: pvalueCutoff = 0.05, qvalueCutoff = 0.2).
Fragment counts mapped to 3 kb downstream of snRNA gene bodies were calculated using featureCounts70 (version 2.0.1, parameters: -s 2 -p --fraction -O -T 16 -t exon -g gene_id) in a strand-specific manner based on the D. melanogaster Ensembl gene annotation (BDGP6.22.96). DESeq265 (version: 1.26.0) was then used to compare differences for each snRNA between treatments under R 3.6.3. Next, to compare the overall endogenous snRNA readthrough levels between treatments, readthrough transcription from a given snRNA was calculated by dividing RNA-seq fragment counts aligned to 3 kb downstream of the snRNA gene body with the fragment counts aligned to the mature snRNA. Statistical difference between treatments was assessed using Wilcoxon signed-rank test under R 3.6.3.
For single-end RNA-seq samples downloaded from GEO (GSE114467),27 Trimmomatic68 (version 0.39; parameters: SE -threads 5 -phred33 TruSeq3PE-2.fa:2:30:8:true LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:30) was used for quality control, including removal of adaptor sequences and low-quality bases. Reads were then aligned to D. melanogaster dm6/BDGP6.22 reference genome using HISAT269 (version 2.1.0, parameters: -k 1 --max-seeds 2). The mapping summaries of RNA-seq datasets are described in Table S1. Fragment counts were calculated per gene using featureCounts70 (version 2.0.1, parameters: -s 1 -p --fraction -O -T 16 -t exon -g gene_id) and then normalized to FPKM (fragments per kilobase of gene per million fragments mapped).79
Lengths of all genes were extracted from the D. melanogaster Ensembl gene annotation (BDGP6.22.96).
ChIP-seq analysis
Published ChIP-seq data for IntS1 and IntS12 in DL1 cells (3 replicates/each) were downloaded from GEO (GSE114467).27 Raw sequences were filtered and trimmed using Trimmomatic68 (version 0.39; parameters: PE -threads 3 -phred33 TruSeq3PE-2.fa:2:30:8:true LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:30). Reads were then aligned to the D. melanogaster dm6/BDGP6.22 reference genome using bowtie271 (version 2.3.5) with default parameters. Multi-mapped reads were removed using sambamba72 (version 0.8.0, parameter: view -F "[XS] == null and not unmapped"). Mapping summaries of ChIP-seq datasets are described in Table S1. IntS1 and IntS12 ChIP-seq peaks were next called using MACS273 (version 2.2.7.1, parameters: -g 142573017 -q 0.001). ChIP-seq peaks were annotated by ChIPseeker74 (version 1.22.1), with the TxDb object of D. melanogaster Ensembl gene annotation (BDGP6.22.96) generated using the makeTxDbFromGFF function in GenomicFeatures package (version 1.38.0).75 Genes closest to peaks and with peaks located within ±1 kb of gene bodies in all 3 replicates were defined as IntS1 or IntS12 binding genes. Finally, IntS1 binding genes and IntS12 binding genes were merged to define a set of 3,932 Integrator bound genes.
PRO-seq analysis
Published paired-end PRO-seq data for DL1 cells (3 replicates of control (β-gal dsRNA) samples, spiked with mouse embryonic stem cells) were downloaded from GEO (GSE114467).27 Cutadapter (version v1.18, parameters: -a TGGAATTCTCGGGTGCCAAGG -A NNNNNNGATCGTCGGACTGTAGAACTCTGAAC -q 10 -m 20) was used to remove adapter sequences, low quality bases at 3' ends, and reads that are <20 nt after trimming. Remaining fragments were first aligned to the mouse mm10 reference genome. Unmapped fragments were then aligned to the D. melanogaster dm6/BDGP6.22 reference genome. Bowtie76 was used in above two alignment steps with same parameters (version 1.12, parameters: -m1 -v2 --un). Mapping summaries of PRO-seq datasets are described in Table S1.
Fragments mapped to the promoter and gene body for each gene were calculated using featurecounts59 (version 2.0.1, parameters: -s 2 -F SAF -Q 20 -p --fraction -O -a). Transcription start sites (TSSs) were extracted from D. melanogaster Ensembl gene annotation (BDGP6.22.96). TSSs ± 250 nt were selected to represent the promoter region and +250 to +1250 nt from TSSs were selected to represent gene body region for each gene. Fragment counts were then normalized to FPKM (fragments per kilobase of gene per million fragments mapped) 79 and FPKM values for 3 replicates were averaged to denote the PRO-seq signal.
Statistical tests
For Northern blots, Western blots, and RT-qPCRs, statistical significance for comparison of means was assessed by one-way ANOVA using GraphPad Prism. Statistical analyses and error bars are explained in the corresponding figure legends, when applicable.
Supplementary Material
Table S1. Quality control and mapping statistics of RNA-seq, ChIP-seq, and PRO-seq experiments. Related to Figures 2, 3, and 5.
Table S2. Quantification of endogenous snRNA readthrough transcription and mature snRNA levels upon over-expression or depletion of Integrator subunits. Related to Figures 2 and 5.
Comparisons of normalized RNA-seq fragment counts aligned to 3 kb downstream of each endogenous snRNA gene body (top) or to the mature snRNA sequence (bottom) in DL1 cells between treatments (depletion of IntS4, IntS6, mts, or Pp2A-29B using RNAi, or over-expression of IntS6 or IntS12). Statistical difference for each snRNA locus was assessed by DESeq2.65
Table S3. Endogenous protein-coding and long non-coding RNA genes up-regulated upon IntS6 over-expression. Related to Figures 3 and 5.
List of protein-coding and long non-coding genes up-regulated in DL1 cells upon IntS6 over-expression when compared to parental DL1 cells or DL1 cells over-expressing IntS12. All cell lines were grown in the presence of Cu2+ for 24 h prior to total RNA isolation and RNA-seq library generation. Threshold used to define differentially expressed genes was ∣log2(fold change)∣ > 0.585 and adjusted P < 0.001. An overlapping set of 107 genes was up-regulated upon IntS6 over-expression.
Table S4. Endogenous protein-coding genes differentially expressed upon depletion of IntS4, IntS6, mts, or Pp2A-29B. Related to Figures 2, 3, and 5.
List of genes that were up- or down-regulated in Drosophila DL1 cells after RNAi depletion of the indicated factor as determined by RNA-seq. Threshold used to define differentially expressed genes was ∣log2(fold change)∣ > 0.585 and adjusted P < 0.001.
Table S5. Oligonucleotide sequences. Related to Figures 1, 2, 3, 4, and 6.
The oligonucleotide sequences used for Northern blots, RT-qPCR, and dsRNA synthesis are provided.
Table S6. Antibodies for Western blots. Related to Figures 2, 4, and 6.
For each antibody used in this study, the source and experimental conditions are provided.
HIGHLIGHTS.
The Integrator phosphatase module is critical at only a subset of regulated genes
IntS6 over-expression blocks Integrator function by titrating PP2A subunits
IntS6 functions analogous to a PP2A regulatory B subunit
PP2A recruitment can be tuned to modulate transcription termination efficiency
ACKNOWLEDGMENTS
We thank Veerle Janssens and members of the Wilusz and Yang labs for helpful discussions. This paper is dedicated to the memory of our colleague Deirdre Tatomer, who passed away while this work was ongoing. Supported by National Institutes of Health grant R35-GM119735 (to J.E.W.), Cancer Prevention & Research Institute of Texas grant RR210031 (to J.E.W.), and National Natural Science Foundation of China grant 31925011 (to L.Y.). J.E.W. is a CPRIT Scholar in Cancer Research.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
DECLARATION OF INTERESTS
J.E.W. serves as a consultant for Laronde.
REFERENCES
- 1.Schier AC, and Taatjes DJ (2020). Structure and mechanism of the RNA polymerase II transcription machinery. Genes Dev 34, 465–488. 10.1101/gad.335679.119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Proudfoot NJ (2016). Transcriptional termination in mammals: Stopping the RNA polymerase II juggernaut. Science 352, aad9926. 10.1126/science.aad9926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Cramer P. (2019). Organization and regulation of gene transcription. Nature 573, 45–54. 10.1038/S41586-019-1517-4. [DOI] [PubMed] [Google Scholar]
- 4.Mendoza-Figueroa MS, Tatomer DC, and Wilusz JE (2020). The Integrator Complex in Transcription and Development. Trends Biochem Sci 45, 923–934. 10.1016/j.tibs.2020.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Welsh SA, and Gardini A (2022). Genomic regulation of transcription and RNA processing by the multitasking Integrator complex. Nat Rev Mol Cell Biol 24, 204–220. 10.1038/S41580-022-00534-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sabath K, and Jonas S (2022). Take a break: Transcription regulation and RNA processing by the Integrator complex. Curr Opin Struct Biol 77, 102443. 10.1016/j.sbi.2022.102443. [DOI] [PubMed] [Google Scholar]
- 7.Kirstein N, Gomes Dos Santos H, Blumenthal E, and Shiekhattar R (2021). The Integrator complex at the crossroad of coding and noncoding RNA. Curr Opin Cell Biol 70, 37–43. 10.1016/j.ceb.2020.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Pfleiderer MM, and Galej WP (2021). Emerging insights into the function and structure of the Integrator complex. Transcription 12, 251–265. 10.1080/21541264.2022.2047583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Baillat D, Hakimi MA, Naar AM, Shilatifard A, Cooch N, and Shiekhattar R (2005). Integrator, a multiprotein mediator of small nuclear RNA processing, associates with the C-terminal repeat of RNA polymerase II. Cell 123, 265–276. 10.1016/j.cell.2005.08.019. [DOI] [PubMed] [Google Scholar]
- 10.Shi Y, and Manley JL (2015). The end of the message: multiple protein-RNA interactions define the mRNA polyadenylation site. Genes Dev 29, 889–897. 10.1101/gad.261974.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Huang KL, Jee D, Stein CB, Elrod ND, Henriques T, Mascibroda LG, Baillat D, Russell WK, Adelman K, and Wagner EJ (2020). Integrator Recruits Protein Phosphatase 2A to Prevent Pause Release and Facilitate Transcription Termination. Mol Cell 80, 345–358 e349. 10.1016/j.molcel.2020.08.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zheng H, Qi Y, Hu S, Cao X, Xu C, Yin Z, Chen X, Li Y, Liu W, Li J, et al. (2020). Identification of Integrator-PP2A complex (INTAC), an RNA polymerase II phosphatase. Science 370, abb5872. 10.1126/science.abb5872. [DOI] [PubMed] [Google Scholar]
- 13.Vervoort SJ, Welsh SA, Devlin JR, Barbieri E, Knight DA, Offley S, Bjelosevic S, Costacurta M, Todorovski I, Kearney CJ, et al. (2021). The PP2A-Integrator-CDK9 axis fine-tunes transcription and can be targeted therapeutically in cancer. Cell 184, 3143–3162 e3132. 10.1016/j.cell.2021.04.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Fianu I, Chen Y, Dienemann C, Dybkov O, Linden A, Urlaub H, and Cramer P (2021). Structural basis of Integrator-mediated transcription regulation. Science 374, 883–887. 10.1126/science.abk0154. [DOI] [PubMed] [Google Scholar]
- 15.Malovannaya A, Li Y, Bulynko Y, Jung SY, Wang Y, Lanz RB, O'Malley BW, and Qin J (2010). Streamlined analysis schema for high-throughput identification of endogenous protein complexes. Proc Natl Acad Sci U S A 107, 2431–2436. 10.1073/pnas.0912599106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Yadav L, Tamene F, Goos H, van Drogen A, Katainen R, Aebersold R, Gstaiger M, and Varjosalo M (2017). Systematic Analysis of Human Protein Phosphatase Interactions and Dynamics. Cell Syst 4, 430–444 e435. 10.1016/j.cels.2017.02.011. [DOI] [PubMed] [Google Scholar]
- 17.Herzog F, Kahraman A, Boehringer D, Mak R, Bracher A, Walzthoeni T, Leitner A, Beck M, Hartl FU, Ban N, et al. (2012). Structural probing of a protein phosphatase 2A network by chemical cross-linking and mass spectrometry. Science 337, 1348–1352. 10.1126/science.1221483. [DOI] [PubMed] [Google Scholar]
- 18.Zheng H, Jin Q, Wang X, Qi Y, Liu W, Ren Y, Zhao D, Xavier Chen F, Cheng J, Chen X, and Xu Y (2023). Structural basis of INTAC-regulated transcription. Protein Cell 14, 698–702. 10.1093/procel/pwad010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Matera AG, and Wang Z (2014). A day in the life of the spliceosome. Nat Rev Mol Cell Biol 15, 108–121. 10.1038/nrm3742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lai F, Gardini A, Zhang A, and Shiekhattar R (2015). Integrator mediates the biogenesis of enhancer RNAs. Nature 525, 399–403. 10.1038/nature14906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Barbieri E, Trizzino M, Welsh SA, Owens TA, Calabretta B, Carroll M, Sarma K, and Gardini A (2018). Targeted Enhancer Activation by a Subunit of the Integrator Complex. Mol Cell 71, 103–116 e107. 10.1016/j.molcel.2018.05.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Rubtsova MP, Vasilkova DP, Moshareva MA, Malyavko AN, Meerson MB, Zatsepin TS, Naraykina YV, Beletsky AV, Ravin NV, and Dontsova OA (2019). Integrator is a key component of human telomerase RNA biogenesis. Sci Rep 9, 1701. 10.1038/S41598-018-38297-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Xie M, Zhang W, Shu MD, Xu A, Lenis DA, DiMaio D, and Steitz JA (2015). The host Integrator complex acts in transcription-independent maturation of herpesvirus microRNA 3' ends. Genes Dev 29, 1552–1564. 10.1101/gad.266973.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Berkyurek AC, Furlan G, Lampersberger L, Beltran T, Weick EM, Nischwitz E, Cunha Navarro I, Braukmann F, Akay A, Price J, et al. (2021). The RNA polymerase II subunit RPB-9 recruits the integrator complex to terminate Caenorhabditis elegans piRNA transcription. EMBO J 40, e105565. 10.15252/embj.2020105565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Beltran T, Pahita E, Ghosh S, Lenhard B, and Sarkies P (2021). Integrator is recruited to promoter-proximally paused RNA Pol II to generate Caenorhabditis elegans piRNA precursors. EMBO J 40, e105564. 10.15252/embj.2020105564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Skaar JR, Ferris AL, Wu X, Saraf A, Khanna KK, Florens L, Washburn MP, Hughes SH, and Pagano M (2015). The Integrator complex controls the termination of transcription at diverse classes of gene targets. Cell Res 25, 288–305. 10.1038/cr.2015.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Elrod ND, Henriques T, Huang KL, Tatomer DC, Wilusz JE, Wagner EJ, and Adelman K (2019). The Integrator Complex Attenuates Promoter-Proximal Transcription at Protein-Coding Genes. Mol Cell 76, 738–752 e737. 10.1016/j.molcel.2019.10.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Tatomer DC, Elrod ND, Liang D, Xiao MS, Jiang JZ, Jonathan M, Huang KL, Wagner EJ, Cherry S, and Wilusz JE (2019). The Integrator complex cleaves nascent mRNAs to attenuate transcription. Genes Dev 33, 1525–1538. 10.1101/gad.330167.119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lykke-Andersen S, Zumer K, Molska ES, Rouviere JO, Wu G, Demel C, Schwalb B, Schmid M, Cramer P, and Jensen TH (2021). Integrator is a genome-wide attenuator of non-productive transcription. Mol Cell 81, 514–529 e516. 10.1016/j.molcel.2020.12.014. [DOI] [PubMed] [Google Scholar]
- 30.Stein CB, Field AR, Mimoso CA, Zhao C, Huang KL, Wagner EJ, and Adelman K (2022). Integrator endonuclease drives promoter-proximal termination at all RNA polymerase II-transcribed loci. Mol Cell 82, 4232–4245 e4211. 10.1016/j.molcel.2022.10.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Rosa-Mercado NA, Zimmer JT, Apostolidi M, Rinehart J, Simon MD, and Steitz JA (2021). Hyperosmotic stress alters the RNA polymerase II interactome and induces readthrough transcription despite widespread transcriptional repression. Mol Cell 81, 502–513.e504. 10.1016/j.molcel.2020.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Beckedorff F, Blumenthal E, daSilva LF, Aoi Y, Cingaram PR, Yue J, Zhang A, Dokaneheifard S, Valencia MG, Gaidosh G, et al. (2020). The Human Integrator Complex Facilitates Transcriptional Elongation by Endonucleolytic Cleavage of Nascent Transcripts. Cell Rep 32, 107917. 10.1016/j.celrep.2020.107917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Oegema R, Baillat D, Schot R, van Unen LM, Brooks A, Kia SK, Hoogeboom AJM, Xia Z, Li W, Cesaroni M, et al. (2017). Human mutations in integrator complex subunits link transcriptome integrity to brain development. PLoS Genet 13, e1006809. 10.1371/journal.pgen.1006809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Mascibroda LG, Shboul M, Elrod ND, Colleaux L, Hamamy H, Huang KL, Peart N, Singh MK, Lee H, Merriman B, et al. (2022). INTS13 variants causing a recessive developmental ciliopathy disrupt assembly of the Integrator complex. Nat Commun 13, 6054. 10.1038/s41467-022-33547-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Federico A, Rienzo M, Abbondanza C, Costa V, Ciccodicola A, and Casamassimi A (2017). Pan-Cancer Mutational and Transcriptional Analysis of the Integrator Complex. Int J Mol Sci 18, 936. 10.3390/ijms18050936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yoshimi A, Lin KT, Wiseman DH, Rahman MA, Pastore A, Wang B, Lee SC, Micol JB, Zhang XJ, de Botton S, et al. (2019). Coordinated alterations in RNA splicing and epigenetic regulation drive leukaemogenesis. Nature 574, 273–277. 10.1038/S41586-019-1618-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tepe B, Macke EL, Niceta M, Weisz Hubshman M, Kanca O, Schultz-Rogers L, Zarate YA, Schaefer GB, Granadillo De Luque JL, Wegner DJ, et al. (2023). Bi-allelic variants in INTS11 are associated with a complex neurological disorder. Am J Hum Genet 110, 774–789. 10.1016/j.ajhg.2023.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hu S, Peng L, Song A, Ji YX, Cheng J, Wang M, and Chen FX (2023). INTAC endonuclease and phosphatase modules differentially regulate transcription by RNA polymerase II. Mol Cell 83, 1588–1604 e1585. 10.1016/j.molcel.2023.03.022. [DOI] [PubMed] [Google Scholar]
- 39.Ezzeddine N, Chen J, Waltenspiel B, Burch B, Albrecht T, Zhuo M, Warren WD, Marzluff WF, and Wagner EJ (2011). A subset of Drosophila integrator proteins is essential for efficient U7 snRNA and spliceosomal snRNA 3'-end formation. Mol Cell Biol 31, 328–341. 10.1128/MCB.00943-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Replogle JM, Saunders RA, Pogson AN, Hussmann JA, Lenail A, Guna A, Mascibroda L, Wagner EJ, Adelman K, Lithwick-Yanai G, et al. (2022). Mapping information-rich genotype-phenotype landscapes with genome-scale Perturb-seq. Cell 185, 2559–2575 e2528. 10.1016/j.cell.2022.05.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zhang F, Ma T, and Yu X (2013). A core hSSB1-INTS complex participates in the DNA damage response. J Cell Sci 126, 4850–4855. 10.1242/jcs.132514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Pfleiderer MM, and Galej WP (2021). Structure of the catalytic core of the Integrator complex. Mol Cell 81, 1246–1259 e1248. 10.1016/j.molcel.2021.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Albrecht TR, Shevtsov SP, Wu Y, Mascibroda LG, Peart NJ, Huang KL, Sawyer IA, Tong L, Dundr M, and Wagner EJ (2018). Integrator subunit 4 is a 'Symplekin-like' scaffold that associates with INTS9/11 to form the Integrator cleavage module. Nucleic Acids Res 46, 4241–4255. 10.1093/nar/gky100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Chen J, Waltenspiel B, Warren WD, and Wagner EJ (2013). Functional analysis of the integrator subunit 12 identifies a microdomain that mediates activation of the Drosophila integrator complex. J Biol Chem 288, 4867–4877. 10.1074/jbc.M112.425892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Seshacharyulu P, Pandey P, Datta K, and Batra SK (2013). Phosphatase: PP2A structural importance, regulation and its aberrant expression in cancer. Cancer Lett 335, 9–18. 10.1016/j.canlet.2013.02.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Xu Y, Xing Y, Chen Y, Chao Y, Lin Z, Fan E, Yu JW, Strack S, Jeffrey PD, and Shi Y (2006). Structure of the protein phosphatase 2A holoenzyme. Cell 127, 1239–1251. 10.1016/j.cell.2006.11.033. [DOI] [PubMed] [Google Scholar]
- 47.Janssens V, and Goris J (2001). Protein phosphatase 2A: a highly regulated family of serine/threonine phosphatases implicated in cell growth and signalling. Biochem J 353, 417–439. 10.1042/0264-6021:3530417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Ghaemmaghami S, Huh WK, Bower K, Howson RW, Belle A, Dephoure N, O'Shea EK, and Weissman JS (2003). Global analysis of protein expression in yeast. Nature 425, 737–741. 10.1038/nature02046. [DOI] [PubMed] [Google Scholar]
- 49.Ogris E, Du X, Nelson KC, Mak EK, Yu XX, Lane WS, and Pallas DC (1999). A protein phosphatase methylesterase (PME-1) is one of several novel proteins stably associating with two inactive mutants of protein phosphatase 2A. J Biol Chem 274, 14382–14391. 10.1074/jbc.274.20.14382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Ogris E, Mudrak I, Mak E, Gibson D, and Pallas DC (1999). Catalytically inactive protein phosphatase 2A can bind to polyomavirus middle tumor antigen and support complex formation with pp60(c-src). J Virol 73, 7390–7398. 10.1128/JVI.73.9.7390-7398.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Sabath K, Staubli ML, Marti S, Leitner A, Moes M, and Jonas S (2020). INTS10-INTS13-INTS14 form a functional module of Integrator that binds nucleic acids and the cleavage module. Nat Commun 11, 3422. 10.1038/s41467-020-17232-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Lin MH, Jensen MK, Elrod ND, Huang KL, Welle KA, Wagner EJ, and Tong L (2022). Inositol hexakisphosphate is required for Integrator function. Nat Commun 13, 5742. 10.1038/S41467-022-33506-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Xu C, Li C, Chen J, Xiong Y, Qiao Z, Fan P, Li C, Ma S, Liu J, Song A, et al. (2023). R-loop-dependent promoter-proximal termination ensures genome stability. Nature 621, 610–619. 10.1038/s41586-023-06515-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Egloff S, O'Reilly D, Chapman RD, Taylor A, Tanzhaus K, Pitts L, Eick D, and Murphy S (2007). Serine-7 of the RNA polymerase II CTD is specifically required for snRNA gene expression. Science 318, 1777–1779. 10.1126/science.1145989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Egloff S, Szczepaniak SA, Dienstbier M, Taylor A, Knight S, and Murphy S (2010). The integrator complex recognizes a new double mark on the RNA polymerase II carboxyl-terminal domain. J Biol Chem 285, 20564–20569. 10.1074/jbc.M110.132530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Egloff S, Zaborowska J, Laitem C, Kiss T, and Murphy S (2012). Ser7 phosphorylation of the CTD recruits the RPAP2 Ser5 phosphatase to snRNA genes. Mol Cell 45, 111–122. 10.1016/j.molcel.2011.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Hernandez N. (1985). Formation of the 3' end of U1 snRNA is directed by a conserved sequence located downstream of the coding region. EMBO J 4, 1827–1837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Cortazar MA, Sheridan RM, Erickson B, Fong N, Glover-Cutter K, Brannan K, and Bentley DL (2019). Control of RNA Pol II Speed by PNUTS-PP1 and Spt5 Dephosphorylation Facilitates Termination by a "Sitting Duck Torpedo" Mechanism. Mol Cell 76, 896–908.e894. 10.1016/j.molcel.2019.09.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Kecman T, Kus K, Heo DH, Duckett K, Birot A, Liberatori S, Mohammed S, Geis-Asteggiante L, Robinson CV, and Vasiljeva L (2018). Elongation/Termination Factor Exchange Mediated by PP1 Phosphatase Orchestrates Transcription Termination. Cell Rep 25, 259–269 e255. 10.1016/j.celrep.2018.09.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Eaton JD, Francis L, Davidson L, and West S (2020). A unified allosteric/torpedo mechanism for transcriptional termination on human protein-coding genes. Genes Dev 34, 132–145. 10.1101/gad.332833.119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Parua PK, Booth GT, Sanso M, Benjamin B, Tanny JC, Lis JT, and Fisher RP (2018). A Cdk9-PP1 switch regulates the elongation-termination transition of RNA polymerase II. Nature 558, 460–464. 10.1038/s41586-018-0214-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Filleur S, Hirsch J, Wille A, Schon M, Sell C, Shearer MH, Nelius T, and Wieland I (2009). INTS6/DICE1 inhibits growth of human androgen-independent prostate cancer cells by altering the cell cycle profile and Wnt signaling. Cancer Cell Int 9, 28. 10.1186/1475-2867-9-28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Ropke A, Buhtz P, Bohm M, Seger J, Wieland I, Allhoff EP, and Wieacker PF (2005). Promoter CpG hypermethylation and downregulation of DICE1 expression in prostate cancer. Oncogene 24, 6667–6675. 10.1038/sj.onc.1208824. [DOI] [PubMed] [Google Scholar]
- 64.Yu G, Wang LG, Han Y, and He QY (2012). clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287. 10.1089/omi.2011.0118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Love MI, Huber W, and Anders S (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15, 550. 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Kramer MC, Liang D, Tatomer DC, Gold B, March ZM, Cherry S, and Wilusz JE (2015). Combinatorial control of Drosophila circular RNA expression by intronic repeats, hnRNPs, and SR proteins. Genes Dev 29, 2168–2182. 10.1101/gad.270421.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Chen J, Ezzeddine N, Waltenspiel B, Albrecht TR, Warren WD, Marzluff WF, and Wagner EJ (2012). An RNAi screen identifies additional members of the Drosophila Integrator complex and a requirement for cyclin C/Cdk8 in snRNA 3'-end formation. RNA 18, 2148–2156. 10.1261/rna.035725.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Bolger AM, Lohse M, and Usadel B (2014). Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. 10.1093/bioinformatics/btu170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Kim D, Paggi JM, Park C, Bennett C, and Salzberg SL (2019). Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol 37, 907–915. 10.1038/s41587-019-0201-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Liao Y, Smyth GK, and Shi W (2014). featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930. 10.1093/bioinformatics/btt656. [DOI] [PubMed] [Google Scholar]
- 71.Langmead B, and Salzberg SL (2012). Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359. 10.1038/nmeth.1923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Tarasov A, Vilella AJ, Cuppen E, Nijman IJ, and Prins P (2015). Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034. 10.1093/bioinformatics/btv098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nusbaum C, Myers RM, Brown M, Li W, and Liu XS (2008). Model-based analysis of ChIP-Seq (MACS). Genome Biol 9, R137. 10.1186/gb-2008-9-9-r137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Yu G, Wang LG, and He QY (2015). ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383. 10.1093/bioinformatics/btv145. [DOI] [PubMed] [Google Scholar]
- 75.Lawrence M, Huber W, Pages H, Aboyoun P, Carlson M, Gentleman R, Morgan MT, and Carey VJ (2013). Software for computing and annotating genomic ranges. PLoS Comput Biol 9, e1003118. 10.1371/journal.pcbi.1003118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Langmead B, Trapnell C, Pop M, and Salzberg SL (2009). Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10, R25. 10.1186/gb-2009-10-3-r25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Tatomer DC, Liang D, and Wilusz JE (2021). RNAi Screening to Identify Factors That Control Circular RNA Localization. Methods Mol Biol 2209, 321–332. 10.1007/978-1-0716-0935-4_20. [DOI] [PubMed] [Google Scholar]
- 78.Tatomer DC, Liang D, and Wilusz JE (2017). Inducible Expression of Eukaryotic Circular RNAs from Plasmids. Methods Mol Biol 1648, 143–154. 10.1007/978-1-4939-7204-3_11. [DOI] [PubMed] [Google Scholar]
- 79.Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, and Pachter L (2010). Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28, 511–515. 10.1038/nbt.1621. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1. Quality control and mapping statistics of RNA-seq, ChIP-seq, and PRO-seq experiments. Related to Figures 2, 3, and 5.
Table S2. Quantification of endogenous snRNA readthrough transcription and mature snRNA levels upon over-expression or depletion of Integrator subunits. Related to Figures 2 and 5.
Comparisons of normalized RNA-seq fragment counts aligned to 3 kb downstream of each endogenous snRNA gene body (top) or to the mature snRNA sequence (bottom) in DL1 cells between treatments (depletion of IntS4, IntS6, mts, or Pp2A-29B using RNAi, or over-expression of IntS6 or IntS12). Statistical difference for each snRNA locus was assessed by DESeq2.65
Table S3. Endogenous protein-coding and long non-coding RNA genes up-regulated upon IntS6 over-expression. Related to Figures 3 and 5.
List of protein-coding and long non-coding genes up-regulated in DL1 cells upon IntS6 over-expression when compared to parental DL1 cells or DL1 cells over-expressing IntS12. All cell lines were grown in the presence of Cu2+ for 24 h prior to total RNA isolation and RNA-seq library generation. Threshold used to define differentially expressed genes was ∣log2(fold change)∣ > 0.585 and adjusted P < 0.001. An overlapping set of 107 genes was up-regulated upon IntS6 over-expression.
Table S4. Endogenous protein-coding genes differentially expressed upon depletion of IntS4, IntS6, mts, or Pp2A-29B. Related to Figures 2, 3, and 5.
List of genes that were up- or down-regulated in Drosophila DL1 cells after RNAi depletion of the indicated factor as determined by RNA-seq. Threshold used to define differentially expressed genes was ∣log2(fold change)∣ > 0.585 and adjusted P < 0.001.
Table S5. Oligonucleotide sequences. Related to Figures 1, 2, 3, 4, and 6.
The oligonucleotide sequences used for Northern blots, RT-qPCR, and dsRNA synthesis are provided.
Table S6. Antibodies for Western blots. Related to Figures 2, 4, and 6.
For each antibody used in this study, the source and experimental conditions are provided.
Data Availability Statement
RNA-seq data have been deposited at GEO and original Northern and western blot images have been deposited at Mendeley. All data are publicly available as of the date of publication. Accession numbers and DOIs are listed in the key resources table.
This paper does not report original code.
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Drosophila IntS6 | This paper | UP2581 Bleed 3 |
| Drosophila IntS8 | This paper | UP2577 Bleed 3 |
| Drosophila IntS11 | This paper | BCM444 Bleed 2 |
| Drosophila IntS12 | This paper | BCM446 Bleed 2 |
| FLAG | Sigma #F1804-1MG | AB_262044 |
| α-tubulin | Sigma #T6074 | AB_477582 |
| PP2A C (mts) | Cell Signaling #2038 | AB_2169495 |
| RNA Pol II CTD (Clone 8WG16) | Sigma #05-952-I-100UG | AB_492629 |
| RNA Pol II CTD phospho Ser2 (Clone 3E10) | Active Motif #61984 | AB_2687450 |
| RNA Pol II CTD phospho Ser5 (Clone 3E8) | Active Motif #61986 | AB_2687451 |
| RNA Pol II CTD phospho Ser7 (Clone 4E12) | Active Motif #61987 | AB_2687452 |
| Anti-FLAG beads | Sigma #A2220 | AB_10063035 |
| Amersham ECL Rabbit IgG, HRP-linked whole Ab | Cytiva #NA934 | AB_772206 |
| Amersham ECL Mouse IgG, HRP-linked whole Ab | Cytiva #NA931 | AB_772210 |
| Anti-rat IgG, HRP-linked Antibody | Cell Signaling #7077S | AB_10694715 |
| Bacterial and virus strains | ||
| Biological samples | ||
| Chemicals, peptides, and recombinant proteins | ||
| Effectene | Qiagen | Cat #301427 |
| Copper sulfate | Fisher BioReagents | Cat #BP346-500 |
| Trizol | ThermoFisher Scientific | Cat #15596018 |
| Protease inhibitors | Roche | Cat #11836170001 |
| Critical commercial assays | ||
| MEGAscript Kit | ThermoFisher Scientific | Cat #AMB13345 |
| NorthernMax Formaldehyde Load Dye | ThermoFisher Scientific | Cat #AM8552 |
| NorthernMax 10X Running Buffer | ThermoFisher Scientific | Cat #AM8671 |
| NorthernMax 10X Denaturing Gel Buffer | ThermoFisher Scientific | Cat #AM8676 |
| ULTRAhyb-Oligo | ThermoFisher Scientific | Cat #AM8663 |
| NuPAGE 4-12% Bis-Tris gels | ThermoFisher Scientific | Cat #NP0323 |
| PVDF membrane | Bio-Rad | Cat #1620177 |
| NuPAGE 3-8% Tris-Acetate gels | ThermoFisher Scientific | Cat #EA03755 |
| SuperSignal West Pico PLUS Chemiluminescent Substrate | ThermoFisher Scientific | Cat #PI34080 |
| TURBO DNase | ThermoFisher Scientific | Cat #AM2238 |
| TaqMan Reverse Transcription Reagents | ThermoFisher Scientific | Cat #N8080234 |
| 2x Power SYBR Green PCR Master Mix | ThermoFisher Scientific | Cat #4368708 |
| Deposited data | ||
| RNA-seq from DL1 cells, DL1 cells over-expressing IntS6, and DL1 cells over-expressing IntS12 | This paper | GEO: GSE223973 |
| RNA-seq from DL1 cells depleted of IntS4, IntS6, mts, or PP2A-29B | This paper | GEO: GSE223974 |
| PRO-seq from DL1 cells treated with β-gal dsRNA | Elrod et al., 201927 | GEO: GSE114467 |
| RNA-seq from DL1 cells depleted of IntS11 and rescued with WT IntS11 or E203Q mutant IntS11 | Elrod et al., 201927 | GEO: GSE114467 |
| RNA-seq from DL1 cells depleted of IntS8 and rescued with WT IntS8 or WFEF/A mutant IntS8 | Huang et al., 202011 | GEO: GSE150844 |
| IntS1 and IntS12 ChIP-seq from DL1 cells | Elrod et al., 201927 | GEO: GSE114467 |
| Raw image files | Mendeley | 10.17632/r46b2sgkkb.1 |
| Experimental models: Cell lines | ||
| DL1 | Sara Cherry, Univ of Pennsylvania | CVCL_Z231 |
| DL1 pMtnA IntS6 | This paper | N/A |
| DL1 pMtnA IntS12 | This paper | N/A |
| DL1 pMtnA IntS7 | This paper | N/A |
| DL1 pMtnA IntS13 | This paper | N/A |
| DL1 pMtnA IntS14 | This paper | N/A |
| Experimental models: Organisms/strains | ||
| Oligonucleotides | ||
| Table S5 | This paper | N/A |
| Recombinant DNA | ||
| Hy_pMT eGFP SV40 | Kramer et al., 201566 | Addgene #69911 |
| Hy_pHml eGFP SV40 | Tatomer et al., 201928 | Addgene #132645 |
| Hy_pCG8620 eGFP SV40 | Tatomer et al., 201928 | Addgene #132646 |
| Hy_pUbi-p63e eGFP SV40 | Tatomer et al., 201928 | Addgene #132650 |
| Hy_pRnrS eGFP SV40 | This paper | Addgene #195062 |
| Hy_pAna eGFP SV40 | This paper | Addgene #195063 |
| pUC U4:39B eGFP | Chen et al., 201344 | N/A |
| Hy_U5:34A eGFP SV40 | This paper | Addgene #195064 |
| pUb 3xFLAG MCS | Chen et al., 201267 | N/A |
| pUb FLAG-Mts | This paper | Addgene #195065 |
| pUb FLAG-Mts H59Q | This paper | Addgene #208402 |
| pUb FLAG-Mts D85N | This paper | Addgene #208403 |
| pUb FLAG-Mts H241A | This paper | Addgene #208404 |
| pUb FLAG-Pp2A-29B | This paper | Addgene #195066 |
| pUb FLAG-Cka | This paper | Addgene #195067 |
| pUb FLAG-tws | This paper | Addgene #195068 |
| pUb FLAG-wdb | This paper | Addgene #195069 |
| pUb FLAG-wrd | This paper | Addgene #195070 |
| pUb FLAG-IntS6 AA 1-400 | This paper | Addgene #195071 |
| pUb FLAG-IntS6 AA 1-600 | This paper | Addgene #195072 |
| pUb FLAG-IntS6 AA 101-1284 | This paper | Addgene #195073 |
| pUb FLAG-IntS6 AA 1197-1284 | This paper | Addgene #195074 |
| pUb FLAG-IntS6 AA 101-1200 | This paper | Addgene #195075 |
| pUb FLAG-Human IntS6 | This paper | Addgene #198408 |
| pUb FLAG-Human IntS6L | This paper | Addgene #198409 |
| pUb FLAG-Zebrafish IntS6 | This paper | Addgene #198410 |
| pUb FLAG-Zebrafish IntS6L | This paper | Addgene #198411 |
| pUb FLAG-IntS1 | Tatomer et al., 201928 | N/A |
| pUb FLAG-IntS2 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS3 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS4 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS5 | Tatomer et al., 201928 | N/A |
| pUb FLAG-IntS6 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS7 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS8 | Huang et al., 202011 | N/A |
| pUb FLAG-IntS9 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS10 | Eric Wagner, Univ of Rochester | N/A |
| pUb FLAG-IntS11 | Tatomer et al., 201928 | N/A |
| pUb FLAG-IntS12 | Chen et al., 201344 | N/A |
| pUb FLAG-IntS13 | Mascibroda et al., 202234 | N/A |
| pUb FLAG-IntS14 | Eric Wagner, Univ of Rochester | N/A |
| pMT FLAG MCS puro | Elrod et al., 201927 | N/A |
| pMtnA FLAG-IntS6 puro | This paper | Addgene #195076 |
| pMtnA FLAG-IntS7 puro | This paper | Addgene #208405 |
| pMtnA FLAG-IntS12 puro | This paper | Addgene #195077 |
| pMtnA FLAG-IntS13 puro | This paper | Addgene #208406 |
| pMtnA FLAG-IntS14 puro | This paper | Addgene #208407 |
| pRSFDuet-1 | Novagen | Cat #71341 |
| pRSFDuet-1 IntS6 AA 1035-1284 | This paper | Addgene #196904 |
| pRSFDuet-1 IntS8 AA 1-308 | This paper | Addgene #196905 |
| pRSFDuet-1 IntS11 AA 300-597 | This paper | Addgene #199329 |
| pGEX-6P-1 | Amersham | Cat #27-4597-01 |
| pGEX-6P-1 IntS12 | This paper | Addgene #210521 |
| Software and algorithms | ||
| Trimmomatic | Bolger et al., 201468 | N/A |
| ImageQuant | Cytiva Life Sciences | N/A |
| HISAT2 | Kim et al., 201969 | N/A |
| featureCounts | Liao et al., 201470 | N/A |
| DESeq2 | Love et al., 201465 | N/A |
| R 3.6.3 | https://www.r-project.org | N/A |
| clusterProfiler | Yu et al., 201264 | N/A |
| bowtie2 | Langmead and Salzberg, 201271 | N/A |
| sambamba | Tarasov et al., 201572 | N/A |
| MACS2 | Zhang et al., 200873 | N/A |
| ChIPseeker | Yu et al., 201574 | N/A |
| GenomicFeatures | Lawrence et al., 201375 | N/A |
| Prism v10.0.0 | GraphPad | N/A |
| Bowtie | Langmead et al., 200976 | N/A |
| Other | ||