Abstract
More than two hundred papers have reported genome-wide data from ancient humans. While the raw data for the vast majority are fully publicly available testifying to the commitment of the paleogenomics community to open data, formats for both raw data and meta-data differ. There is thus a need for uniform curation and a centralized, version-controlled compendium that researchers can download, analyze, and reference. Since 2019, we have been maintaining the Allen Ancient DNA Resource (AADR), which aims to provide an up-to-date, curated version of the world’s published ancient human DNA data, represented at more than a million single nucleotide polymorphisms (SNPs) at which almost all ancient individuals have been assayed. The AADR has gone through six public releases at the time of writing and review of this manuscript, and crossed the threshold of >10,000 individuals with published genome-wide ancient DNA data at the end of 2022. This note is intended as a citable descriptor of the AADR.
Subject terms: Genetic variation, Genetic variation
Background & Summary
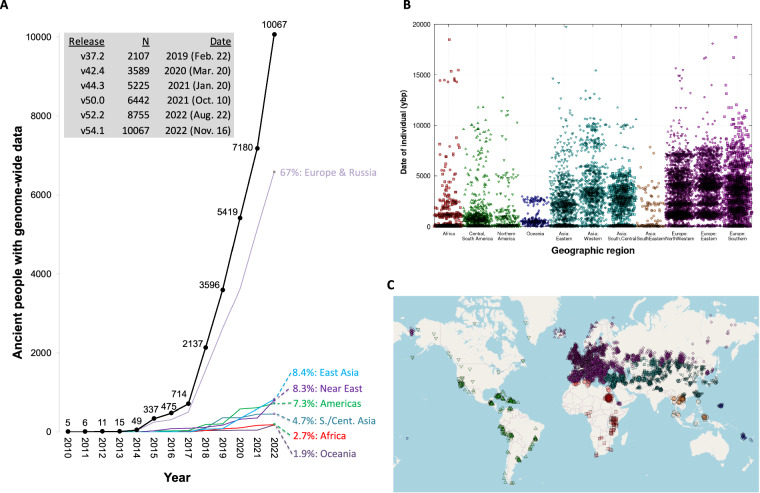
The first genome-wide ancient DNA data were published in 20101–3. However, it was only in 2015 with the advent of large-scale studies of Holocene genomes, in-solution enrichment of ancient DNA libraries for targeted single nucleotide polymorphisms (SNPs)4–6, and the introduction of automated protocols and liquid handling robots for processing of ancient DNA libraries7,8, that the number of individuals with genome-wide data began to increase rapidly. Between 2010 and 2014, data from an average of about 10 individuals with genome-wide data were published each year. Between 2015 and 2017, the numbers increased to about 200 annually. Since 2018, data from thousands of individuals have been published every year (Fig. 1). About 67% of the data are from Europe and Russia, a proportion that has held relatively steady since the beginning of the field of ancient DNA. The proportion of data from East Asia has grown from about 1% of all data in 2015 to 8% today. The proportion of data from Africa has grown from none in 2014 to 3% today (Fig. 1).
Fig. 1.
Growth in world’s published human genome-wide ancient DNA data. (A) By year of publication (broken down by geography). (B) By date (color and symbol both indicate geographic location). (C) By geography (using same color and symbol scheme as in previous panel).
A challenge in analyzing ancient DNA data is that it has been reported over hundreds of independent studies. Thus, while raw sequence data for more than 99% of individuals9 are fully available in public repositories such as ENA10 and SRA11, the uploaded data exist in diverse formats, as do the meta-data such as archaeological, chronological, and geographic information. Some resources exist which consolidate subsets of publicly available ancient DNA data, including a Y-chromosome database with assembled information from nearly two thousand ancient Eurasian individuals12, a mitochondrial DNA database with more than two thousand individuals13, and the Online Ancient Genome Repository14 which copies publicly available data and encapsulates each dataset into an archived tar file. However, none of these provide a regularly curated dataset that attempts to include all published data in an easily co-analyzable format, such as a single genotype file with complete annotations in a single tab-delimited form.
Sources of data
To bring data generated outside our own laboratory into the AADR, we usually start with available sequences from a public repository, most often the European Nucleotide Archive (https://www.ebi.ac.uk/ena), following accession numbers given in the published papers. In some cases we start with alternatively formatted versions that we request directly from the authors. All source articles are cited in the reference list of this paper. For data generated in our laboratory, we start from our own raw sequence files, which are the basis for data uploaded to established public repositories.
Methods
The raw data generated outside our laboratory come in diverse formats, usually fastq files (for raw sequence data) or bams (for either unaligned reads or reads aligned to a reference genome)15. A challenge is that there can be considerable variation in fastq and bam files, reflecting the formatting, filtering and processing choices made by researchers in generating data. This includes:
Base calls and associated quality scores in raw sequences are often modified by the researchers who generated the data. One common modification is to recalibrate base quality scores16. Another modification is to ignore information from the ends of sequences, either by masking terminal bases in the sequences that are uploaded and marking them as “N”, or clipping (removing) them altogether17. This reduces error rates associated with cytosine deamination typical of ancient DNA data. However, it also means that users cannot make choices about whether to use the valuable data that have been masked and clipped (such as sites unaffected by deamination). In addition, this procedure has the effect of making it difficult to identify damaged molecules which are a strong indicator that those molecules indeed are ancient and not derived from some potential contaminating modern human source.
Sequences may be aligned to different human reference genomes, typically hg19, hs37d5, or hg20, each with their own unique coordinate systems. To build a homogeneous dataset, we therefore have to map to a unified coordinate system, currently based on hg1918–20. A further challenge is that chromosomes may have inconsistent naming conventions (for example ‘chr1’ v. ‘1’, or ‘chrMT’ v. ‘MT’ v. ‘chrM’), or the sorting order of chromosomes can differ. This results in practical difficulties in merging datasets.
Data may be deposited either (i) by library, or (ii) by-individual with multiple libraries in a single file. If data are deposited by library, then it may be necessary to identify and perform a merging step. There are pitfalls that arise in such merging, as in some cases “readgroup” names (a tag which groups reads together) are the same across individuals, and so joint processing of many individuals can inadvertently lead to in-silico contamination.
Filtering of data
To add data to the AADR, we manually process the dataset from each individual, tailoring the processing procedure according to the characteristics of the data. We create a bam file aligned to the hg19 genome reference sequence. The bam files used to generate AADR constitute tens of terabytes in size altogether. We process these bams to produce genotypes at a set of about 1.23 million SNPs that have been assayed for nearly all published individuals with ancient DNA data. For the great majority of ancient DNA datasets, the genotypes are “pseudohaploid”, meaning that we represent the individual by a randomly sampled sequence at each analyzed position. For the small fraction of individuals for whom coverage is sufficient to allow full genotyping, we also release diploid genotypes21,22.
Combining datasets
To increase the usefulness of the AADR, we have added into the AADR data from diverse modern humans, including shotgun sequencing data from sets of individuals included within the 1000 Genomes Project23, the Simons Genome Diversity Project24, and the Human Genome Diversity Project25 To integrate these data, we had to address challenges of different reference genomes (for example transforming from hg20 to hg19 coordinates). There are 6399 modern individuals with shotgun data in the v54.1 AADR release.
We also integrated a dataset of 4114 modern individuals genotyped on the Affymetrix Human Origins array at approximately ~600,000 SNPs26. This is a sufficiently valuable dataset that the AADR provides two releases: one on all 1.23 million targets (excluding the Human Origins data), and one on just the Human Origins targets.
Since the v52.2 release, we have also maintained a mitochondrial repository, which now includes mitochondrial genomes for 4122 ancient individuals in the AADR.
Technical Validation
Prior to each release, several steps are performed to verify that new and updated data components have been added correctly and are co-analyzable with the full datasets.
An initial assessment considers coverage of each individual and evidence for contamination, updating annotations as needed. In addition, we manually curate the genotypes to check that the data from each individual has sensible population genetic properties, by looking for potential outliers based on ADMIXTURE27 and principal components analysis28.
Curation of metainformation and integration of ongoing community feedback
Because we are trying to keep AADR current, we err on the side of inclusivity, and thus try to bring data into the dataset even when meta-information and metrics are incomplete. Each AADR release updates meta-information and identifiers as appropriate. We rely on ongoing curation of the dataset as well as feedback from the user community which we invite through communication with the corresponding authors, to identify individuals with erroneous meta-information or corrupted genetic data, which we then seek to correct in subsequent releases.
Data Record
The AADR dataset is available at Harvard Dataverse29 (https://dataverse.harvard.edu/dataverse/reich_lab). The latest release at the time of writing and peer review is 8.0.
Each data release consists of three standard files in EIGENSTRAT format (.ind,.snp, and.geno). We also include an annotation file that is rich in meta-information for the dataset (.anno). The.anno file includes meta-data manually extracted from the papers reporting the data, in some cases supplemented by information that appeared later or that reflect clarifications from authors or the user community. For archaeological information, we attempt to provide:
Skeletal codes and grave numbers and sometimes other identifiers, always also including the code used for genetic analysis.
Latitude and longitude.
Location information, with a separate column for “Political entity” such as country, and locality information.
Chronological information in a standard format. When a radiocarbon date is available, we include the laboratory number and calibrated 95.4% confidence interval obtained in OxCal v4.4.2 using either the IntCal20 or SHCal20 calibration curve (if we make an alternative choice, it is explicitly explained in a “Methods for Determining Date” column). We also report the posterior mean and standard deviation of the calibrated radiocarbon date. When no radiocarbon date is available, we present a date uncertainty range based on archaeological context, usually rounded to the nearest 50 or 100 years, and quote the mean and standard deviation assuming a uniform distribution over its range (the standard deviation of a uniform distribution is the range of that distribution divided by the square root of 12).
We include an estimate of the age of the individual at their death based on physical anthropology when we are able to obtain it.
We include a group name for the individual, using a naming convention that aims to be systematic30.
Data from individuals generated using shotgun sequencing methods have a suffix “.SG” (for pseudohaploid representations) or “.DG” (for diploid representations).
We include many metrics computed on the genetic data, including not just amount of data (such as average coverage assayed at the subset of 1.15 million autosomal sites targeted in the 1.23 million SNP enrichment assays), but also molecular sex determination, cytosine-to-thymine rate in the final nucleotide31, fraction of the genome in multi-megabase runs of homozygosity32, identification of close relatives in the dataset (in a dedicated “family information” column), and estimates of contamination33,34. We have added additional metrics in each release to further improve the usefulness of the dataset.
When data from an individual have been published in multiple studies using the same methodology such as in-solution enrichment, the AADR typically includes only the best quality version which is usually the latest one (for such individuals, the “publication” columns in the.anno file notes the date of the publication that first reported data from the individual, as well as the publication that report the version that is actually included within the AADR). For some individuals, we include multiple representations of data, for example from shotgun sequencing, in-solution enrichment, restricted to UDG-treated libraries, or restricted to sequences showing characteristic ancient DNA damage to reduce the possible impact of contaminating sequences (“_d” suffix). The different versions have unique “Version IDs” but the same “Master ID” (which seeks to uniquely identifies an individual). These IDs may change from data release to data release; if data from two different Master IDs are found to come from the same individual, they are merged into a single Master ID.
The group name may include a suffix that mark individuals such as potentially contaminated (“_contam”), or as a population genetic outlier (“_o”), or as having relatively little data (low coverage – “lc”).
Citation guidance
Researchers who use the AADR as the starting point for analyses are encouraged to give two citations for the individual datasets: (1) this paper, and (2) the AADR Dataverse citation29 specifying the version of the AADR downloaded. Citing the AADR paper is not a substitute for citing the original publications that produced data, which should be specifically referenced in each publication. Supplementary Data Table 1 provides the full list of references in the component papers for the v54.1 release, and will be updated at Dataverse for each new release going forward29.
All source articles are additionally cited in the reference list of this paper1–6,8,25,26,35–260.
Supplementary information
Acknowledgements
We are grateful to Rebecca Bernardos, Aisling Kearns, Nadin Rohland, Arie Shaus, Katie Mika and many in the user community for help in curating and improving the AADR. Portions of this research were conducted on the O2 High Performance Compute Cluster, supported by the Research Computing Group, at Harvard Medical School. Construction and maintenance of the AADR was supported by NIH grant HG012287, by the Allen Discovery Center program, a Paul G. Allen Frontiers Group advised program of the Paul G. Allen Family Foundation, by John Templeton Foundation grant 61220, and by the Howard Hughes Medical Institute (HHMI). This article is subject to HHMI’s Open Access to Publications policy. HHMI lab heads have previously granted a nonexclusive CC BY 4.0 license to the public and a sublicensable license to HHMI in their research articles. Pursuant to those licenses, the author-accepted manuscript of this article can be made freely available under a CC BY 4.0 license immediately upon publication.
Author contributions
S.M. built the original pipeline and data structures which were reimplemented with performance improvements and upgrades by M.M. A.M. managed data production, merging of datasets, and quality control; he also improved the pipeline. H.R., I.L. and I.O. determined relatedness patterns, population genetic affinities, runs of homozygosity, and contributed to data quality control. N.P. wrote components of the pipeline used for generating pseudo-haploid genotypes. D.R. managed and curated genetic and archaeological annotations. S.M. and D.R. wrote the manuscript.
Code availability
The pipeline used for processing raw data generated within the Reich lab is available in the ‘Workflow Description Language’ (WDL) here: https://github.com/DReichLab/adna-workflow, and includes individual python scripts for components of the pipeline.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Swapan Mallick, Email: shop@genetics.med.harvard.edu.
David Reich, Email: reich@genetics.med.harvard.edu.
Supplementary information
The online version contains supplementary material available at 10.1038/s41597-024-03031-7.
References
- 1.Green RE, et al. A draft sequence of the Neandertal genome. Science. 2010;328(5979):710–722. doi: 10.1126/science.1188021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rasmussen M, et al. Ancient human genome sequence of an extinct Palaeo-Eskimo. Nature. 2010;463(7282):757–762. doi: 10.1038/nature08835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Reich D, et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature. 2010;468(7327):1053–1060. doi: 10.1038/nature09710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Fu Q, et al. DNA analysis of an early modern human from Tianyuan Cave, China. Proc Natl Acad Sci USA. 2013;110(6):2223–2227. doi: 10.1073/pnas.1221359110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Haak W, et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature. 2015;522(7555):207–211. doi: 10.1038/nature14317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Mathieson I, et al. Genome-wide patterns of selection in 230 ancient Eurasians. Nature. 2015;528(7583):499–503. doi: 10.1038/nature16152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Rohland N, et al. Partial uracil-DNA-glycosylase treatment for screening of ancient DNA. Philos Trans R Soc Lond B Biol Sci. 2015;370(1660):p20130624. doi: 10.1098/rstb.2013.0624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Slon V, et al. Neandertal and Denisovan DNA from Pleistocene sediments. Science. 2017;356(6338):605–608. doi: 10.1126/science.aam9695. [DOI] [PubMed] [Google Scholar]
- 9.Anagnostou P, et al. When data sharing gets close to 100%: what human paleogenetics can teach the open science movement. PLoS One. 2015;10(3):pe0121409. doi: 10.1371/journal.pone.0121409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Yuan, D. et al. The European Nucleotide Archive in 2023. Nucleic Acids Res, gkad1067 (2023) [DOI] [PMC free article] [PubMed]
- 11.Leinonen R, et al. The sequence read archive. Nucleic Acids Res. 2011;39(Database issue):D19–D21. doi: 10.1093/nar/gkq1019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Freeman L, Brimacombe CS, Elhaik E. aYChr-DB: a database of ancient human Y haplogroups. NAR Genom Bioinform. 2020;2(4):lqaa081. doi: 10.1093/nargab/lqaa081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ehler E, et al. AmtDB: a database of ancient human mitochondrial genomes. Nucleic Acids Res. 2019;47(D1):D29–D32. doi: 10.1093/nar/gky843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.www.oagr.org.
- 15.Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(14):1754–1760. doi: 10.1093/bioinformatics/btp324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ginolhac A, et al. mapDamage: testing for damage patterns in ancient DNA sequences. Bioinformatics. 2011;27(15):2153–2155. doi: 10.1093/bioinformatics/btr347. [DOI] [PubMed] [Google Scholar]
- 17.Briggs AW, et al. Patterns of damage in genomic DNA sequences from a Neandertal. Proc Natl Acad Sci USA. 2007;104(37):14616–14621. doi: 10.1073/pnas.0704665104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lander ES, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409(6822):860–921. doi: 10.1038/35057062. [DOI] [PubMed] [Google Scholar]
- 19.Schneider VA, et al. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res. 2017;27(5):849–864. doi: 10.1101/gr.213611.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Fairley S, et al. The International Genome Sample Resource (IGSR) collection of open human genomic variation resources. Nucleic Acids Res. 2020;48(D1):D941–D947. doi: 10.1093/nar/gkz836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.McKenna A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20(9):1297–1303. doi: 10.1101/gr.107524.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Prufer K. snpAD: an ancient DNA genotype caller. Bioinformatics. 2018;34(24):4165–4171. doi: 10.1093/bioinformatics/bty507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Genomes Project, C. et al. A global reference for human genetic variation. Nature. 2015;526(7571):68–74. doi: 10.1038/nature15393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Mallick S, et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature. 2016;538(7624):201–6. doi: 10.1038/nature18964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Bergstrom A, et al. Insights into human genetic variation and population history from 929 diverse genomes. Science. 2020;367(6484):eaay5012. doi: 10.1126/science.aay5012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Patterson N, et al. Ancient admixture in human history. Genetics. 2012;192(3):1065–1093. doi: 10.1534/genetics.112.145037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Alexander DH, Novembre J, Lange K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 2009;19(9):1655–1664. doi: 10.1101/gr.094052.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Patterson N, Price AL, Reich D. Population structure and eigenanalysis. PLoS genetics. 2006;2(12):e190. doi: 10.1371/journal.pgen.0020190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mallick S, Reich D. 2023. The Allen Ancient DNA Resource (AADR): A curated compendium of ancient human genomes. Harvard Dataverse. [DOI] [PubMed]
- 30.Eisenmann S, et al. Reconciling material cultures in archaeology with genetic data: The nomenclature of clusters emerging from archaeogenomic analysis. Sci Rep. 2018;8(1):13003. doi: 10.1038/s41598-018-31123-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Skoglund P, et al. Separating endogenous ancient DNA from modern day contamination in a Siberian Neandertal. Proc Natl Acad Sci USA. 2014;111(6):2229–2234. doi: 10.1073/pnas.1318934111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ringbauer H, Novembre J, Steinrucken M. Parental relatedness through time revealed by runs of homozygosity in ancient DNA. Nat Commun. 2021;12(1):5425. doi: 10.1038/s41467-021-25289-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Korneliussen TS, Albrechtsen A, Nielsen R. ANGSD: Analysis of Next Generation Sequencing Data. BMC Bioinformatics. 2014;15(1):356. doi: 10.1186/s12859-014-0356-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Fu Q, et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr Biol. 2013;23(7):553–559. doi: 10.1016/j.cub.2013.02.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Jakobsson M, et al. Genotype, haplotype and copy-number variation in worldwide human populations. Nature. 2008;451(7181):998–1003. doi: 10.1038/nature06742. [DOI] [PubMed] [Google Scholar]
- 36.Keller A, et al. New insights into the Tyrolean Iceman’s origin and phenotype as inferred by whole-genome sequencing. Nat Commun. 2012;3(1):698. doi: 10.1038/ncomms1701. [DOI] [PubMed] [Google Scholar]
- 37.Meyer M, et al. A high-coverage genome sequence from an archaic Denisovan individual. Science. 2012;338(6104):222–226. doi: 10.1126/science.1224344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Fu Q, et al. Genome sequence of a 45,000-year-old modern human from western Siberia. Nature. 2014;514(7523):445–449. doi: 10.1038/nature13810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Gamba C, et al. Genome flux and stasis in a five millennium transect of European prehistory. Nat Commun. 2014;5(1):5257. doi: 10.1038/ncomms6257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Lazaridis I, et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature. 2014;513(7518):409–413. doi: 10.1038/nature13673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Malaspinas AS, et al. Two ancient human genomes reveal Polynesian ancestry among the indigenous Botocudos of Brazil. Curr Biol. 2014;24(21):R1035–R1037. doi: 10.1016/j.cub.2014.09.078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Olalde I, et al. Derived immune and ancestral pigmentation alleles in a 7,000-year-old Mesolithic European. Nature. 2014;507(7491):225–228. doi: 10.1038/nature12960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Prüfer K, et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature. 2014;505(7481):43–49. doi: 10.1038/nature12886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Raghavan M, et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature. 2014;505(7481):87–91. doi: 10.1038/nature12736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Raghavan M, et al. The genetic prehistory of the New World Arctic. Science. 2014;345(6200):1255832. doi: 10.1126/science.1255832. [DOI] [PubMed] [Google Scholar]
- 46.Rasmussen M, et al. The genome of a Late Pleistocene human from a Clovis burial site in western Montana. Nature. 2014;506(7487):225–229. doi: 10.1038/nature13025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Seguin-Orlando A, et al. Genomic structure in Europeans dating back at least 36,200 years. Science. 2014;346(6213):1113–1118. doi: 10.1126/science.aaa0114. [DOI] [PubMed] [Google Scholar]
- 48.Skoglund P, et al. Genomic diversity and admixture differs for Stone-Age Scandinavian foragers and farmers. Science. 2014;344(6185):747–750. doi: 10.1126/science.1253448. [DOI] [PubMed] [Google Scholar]
- 49.Allentoft ME, et al. Population genomics of Bronze Age Eurasia. Nature. 2015;522(7555):167–172. doi: 10.1038/nature14507. [DOI] [PubMed] [Google Scholar]
- 50.Fu Q, et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature. 2015;524(7564):216–219. doi: 10.1038/nature14558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Gallego Llorente M, et al. Ancient Ethiopian genome reveals extensive Eurasian admixture throughout the African continent. Science. 2015;350(6262):820–822. doi: 10.1126/science.aad2879. [DOI] [PubMed] [Google Scholar]
- 52.Günther T, et al. Ancient genomes link early farmers from Atapuerca in Spain to modern-day Basques. Proc Natl Acad Sci USA. 2015;112(38):11917–11922. doi: 10.1073/pnas.1509851112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Jones ER, et al. Upper Palaeolithic genomes reveal deep roots of modern Eurasians. Nat Commun. 2015;6(1):8912. doi: 10.1038/ncomms9912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Olalde I, et al. A Common Genetic Origin for Early Farmers from Mediterranean Cardial and Central European LBK Cultures. Mol Biol Evol. 2015;32(12):3132–3142. doi: 10.1093/molbev/msv181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Qin P, et al. Denisovan Ancestry in East Eurasian and Native American Populations. Mol Biol Evol. 2015;32(10):2665–2674. doi: 10.1093/molbev/msv141. [DOI] [PubMed] [Google Scholar]
- 56.Raghavan M, et al. Genomic evidence for the Pleistocene and recent population history of Native Americans. Science. 2015;349(6250):aab3884. doi: 10.1126/science.aab3884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Rasmussen M, et al. The ancestry and affiliations of Kennewick Man. Nature. 2015;523(7561):455–458. doi: 10.1038/nature14625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Skoglund P, et al. Genetic evidence for two founding populations of the Americas. Nature. 2015;525(7567):104–108. doi: 10.1038/nature14895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Broushaki F, et al. Early Neolithic genomes from the eastern Fertile Crescent. Science. 2016;353(6298):499–503. doi: 10.1126/science.aaf7943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Cassidy LM, et al. Neolithic and Bronze Age migration to Ireland and establishment of the insular Atlantic genome. Proc Natl Acad Sci USA. 2016;113(2):368–373. doi: 10.1073/pnas.1518445113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Fu Q, et al. The genetic history of Ice Age Europe. Nature. 2016;534(7606):200–205. doi: 10.1038/nature17993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Hofmanová Z, et al. Early farmers from across Europe directly descended from Neolithic Aegeans. Proc Natl Acad Sci USA. 2016;113(25):6886–6891. doi: 10.1073/pnas.1523951113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Jeong C, et al. Long-term genetic stability and a high-altitude East Asian origin for the peoples of the high valleys of the Himalayan arc. Proc Natl Acad Sci USA. 2016;113(27):7485–7490. doi: 10.1073/pnas.1520844113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Kılınç GM, et al. The Demographic Development of the First Farmers in Anatolia. Curr Biol. 2016;26(19):2659–2666. doi: 10.1016/j.cub.2016.07.057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Lazaridis I, et al. Genomic insights into the origin of farming in the ancient Near East. Nature. 2016;536(7617):419–424. doi: 10.1038/nature19310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Martiniano R, et al. Genomic signals of migration and continuity in Britain before the Anglo-Saxons. Nat Commun. 2016;7(1):10326. doi: 10.1038/ncomms10326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Mondal M, et al. Genomic analysis of Andamanese provides insights into ancient human migration into Asia and adaptation. Nat Genet. 2016;48(9):1066–1070. doi: 10.1038/ng.3621. [DOI] [PubMed] [Google Scholar]
- 68.Omrak A, et al. Genomic Evidence Establishes Anatolia as the Source of the European Neolithic Gene Pool. Curr Biol. 2016;26(2):270–275. doi: 10.1016/j.cub.2015.12.019. [DOI] [PubMed] [Google Scholar]
- 69.Schiffels S, et al. Iron Age and Anglo-Saxon genomes from East England reveal British migration history. Nat Commun. 2016;7(1):10408. doi: 10.1038/ncomms10408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Skoglund P, et al. Genomic insights into the peopling of the Southwest Pacific. Nature. 2016;538(7626):510–513. doi: 10.1038/nature19844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.González-Fortes G, et al. Paleogenomic Evidence for Multi-generational Mixing between Neolithic Farmers and Mesolithic Hunter-Gatherers in the Lower Danube Basin. Curr Biol. 2017;27(12):1801–1810. doi: 10.1016/j.cub.2017.05.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Haber M, et al. Continuity and Admixture in the Last Five Millennia of Levantine History from Ancient Canaanite and Present-Day Lebanese Genome Sequences. Am J Hum Genet. 2017;101(2):274–282. doi: 10.1016/j.ajhg.2017.06.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Japan, et al. A partial nuclear genome of the Jomons who lived 3000 years ago in Fukushima, Japan. J Hum Genet. 2017;62(2):213–221. doi: 10.1038/jhg.2016.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Jones ER, et al. The Neolithic Transition in the Baltic Was Not Driven by Admixture with Early European Farmers. Curr Biol. 2017;27(4):576–582. doi: 10.1016/j.cub.2016.12.060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Kennett DJ, et al. Archaeogenomic evidence reveals prehistoric matrilineal dynasty. Nat Commun. 2017;8(1):14115. doi: 10.1038/ncomms14115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Lazaridis I, et al. Genetic origins of the Minoans and Mycenaeans. Nature. 2017;548(7666):214–218. doi: 10.1038/nature23310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Lindo J, et al. Ancient individuals from the North American Northwest Coast reveal 10,000 years of regional genetic continuity. Proc Natl Acad Sci USA. 2017;114(16):4093–4098. doi: 10.1073/pnas.1620410114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Lipson M, et al. Parallel palaeogenomic transects reveal complex genetic history of early European farmers. Nature. 2017;551(7680):368–372. doi: 10.1038/nature24476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Martiniano R, et al. The population genomics of archaeological transition in west Iberia: Investigation of ancient substructure using imputation and haplotype-based methods. PLoS Genet. 2017;13(7):e1006852. doi: 10.1371/journal.pgen.1006852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Nakatsuka N, et al. The promise of discovering population-specific disease-associated genes in South Asia. Nat Genet. 2017;49(9):1403–1407. doi: 10.1038/ng.3917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Prüfer K, et al. A high-coverage Neandertal genome from Vindija Cave in Croatia. Science. 2017;358(6363):655–658. doi: 10.1126/science.aao1887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Rodríguez-Varela R, et al. Genomic Analyses of Pre-European Conquest Human Remains from the Canary Islands Reveal Close Affinity to Modern North Africans. Curr Biol. 2017;27(21):3396–3402. doi: 10.1016/j.cub.2017.09.059. [DOI] [PubMed] [Google Scholar]
- 83.Saag L, et al. Extensive Farming in Estonia Started through a Sex-Biased Migration from the Steppe. Curr Biol. 2017;27(14):2185–2193. doi: 10.1016/j.cub.2017.06.022. [DOI] [PubMed] [Google Scholar]
- 84.Schlebusch CM, et al. Southern African ancient genomes estimate modern human divergence to 350,000 to 260,000 years ago. Science. 2017;358(6363):652–655. doi: 10.1126/science.aao6266. [DOI] [PubMed] [Google Scholar]
- 85.Schuenemann VJ, et al. Ancient Egyptian mummy genomes suggest an increase of Sub-Saharan African ancestry in post-Roman periods. Nat Commun. 2017;8(1):15694. doi: 10.1038/ncomms15694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Sikora M, et al. Ancient genomes show social and reproductive behavior of early Upper Paleolithic foragers. Science. 2017;358(6363):659–662. doi: 10.1126/science.aao1807. [DOI] [PubMed] [Google Scholar]
- 87.Siska V, et al. Genome-wide data from two early Neolithic East Asian individuals dating to 7700 years ago. Sci Adv. 2017;3(2):e1601877. doi: 10.1126/sciadv.1601877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Skoglund P, et al. Reconstructing Prehistoric African Population Structure. Cell. 2017;171(1):59–71. doi: 10.1016/j.cell.2017.08.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Unterländer M, et al. Ancestry and demography and descendants of Iron Age nomads of the Eurasian Steppe. Nat Commun. 2017;8(1):14615. doi: 10.1038/ncomms14615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.van, den, Brink ECM, et al. A Late Bronze Age II clay coffin from Tel Shaddudin the Central Jezreel Valley, Israel: context andhistorical implications. Levant. 2017;49(2):105–135. doi: 10.1080/00758914.2017.1368204. [DOI] [Google Scholar]
- 91.Yang MA, et al. 40,000-Year-Old Individual from Asia Provides Insight into Early Population Structure in Eurasia. Curr Biol. 2017;27(20):3202–3208. doi: 10.1016/j.cub.2017.09.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Amorim CEG, et al. Understanding 6th-century barbarian social organization and migration through paleogenomics. Nat Commun. 2018;9(1):3547. doi: 10.1038/s41467-018-06024-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Damgaard PB, et al. 137 ancient human genomes from across the Eurasian steppes. Nature. 2018;557(7705):369–374. doi: 10.1038/s41586-018-0094-2. [DOI] [PubMed] [Google Scholar]
- 94.de, et al. The first horse herders and the impact of early Bronze Age steppe expansions into Asia. Science. 2018;360(6396):eaar7711. doi: 10.1126/science.aar7711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.De, et al. Genomic insights into the origin and diversification of late maritime hunter-gatherers from the Chilean Patagonia. Proc Natl Acad Sci USA. 2018;115(17):E4006–E4012. doi: 10.1073/pnas.1715688115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Ebenesersdóttir SS, et al. Ancient genomes from Iceland reveal the making of a human population. Science. 2018;360(6392):1028–1032. doi: 10.1126/science.aar2625. [DOI] [PubMed] [Google Scholar]
- 97.Fernandes DM, et al. A genomic Neolithic time transect of hunter-farmer admixture in central Poland. Sci Rep. 2018;8(1):14879. doi: 10.1038/s41598-018-33067-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Fregel R, et al. Ancient genomes from North Africa evidence prehistoric migrations to the Maghreb from both the Levant and Europe. Proc Natl Acad Sci USA. 2018;115(26):6774–6779. doi: 10.1073/pnas.1800851115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Günther T, et al. Population genomics of Mesolithic Scandinavia: Investigating early postglacial migration routes and high-latitude adaptation. PLoS Biol. 2018;16(1):e2003703. doi: 10.1371/journal.pbio.2003703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Hajdinjak M, et al. Reconstructing the genetic history of late Neanderthals. Nature. 2018;555(7698):652–656. doi: 10.1038/nature26151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Harney É, et al. Ancient DNA from Chalcolithic Israel reveals the role of population mixture in cultural transformation. Nat Commun. 2018;9(1):3336. doi: 10.1038/s41467-018-05649-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Jeong C, et al. Bronze Age population dynamics and the rise of dairy pastoralism on the eastern Eurasian steppe. Proc Natl Acad Sci USA. 2018;115(48):E11248–55. doi: 10.1073/pnas.1813608115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Krzewińska M, et al. Genomic and Strontium Isotope Variation Reveal Immigration Patterns in a Viking Age Town. Curr Biol. 2018;28(17):2730–2738. doi: 10.1016/j.cub.2018.06.053. [DOI] [PubMed] [Google Scholar]
- 104.Krzewińska M, et al. Ancient genomes suggest the eastern Pontic-Caspian steppe as the source of western Iron Age nomads. Sci Adv. 2018;4(10):eaat4457. doi: 10.1126/sciadv.aat4457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Lamnidis TC, et al. Ancient Fennoscandian genomes reveal origin and spread of Siberian ancestry in Europe. Nat Commun. 2018;9(1):5018. doi: 10.1038/s41467-018-07483-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Lindo J, et al. The genetic prehistory of the Andean highlands 7000 years BP though European contact. Sci Adv. 2018;4(11):eaau4921. doi: 10.1126/sciadv.aau4921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Lipson M, et al. Ancient genomes document multiple waves of migration in Southeast Asian prehistory. Science. 2018;361(6397):92–95. doi: 10.1126/science.aat3188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Lipson M, et al. Population Turnover in Remote Oceania Shortly after Initial Settlement. Curr Biol. 2018;28(7):1157–1165. doi: 10.1016/j.cub.2018.02.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Mathieson I, et al. The genomic history of southeastern Europe. Nature. 2018;555(7695):197–203. doi: 10.1038/nature25778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.McColl H, et al. The prehistoric peopling of Southeast Asia. Science. 2018;361(6397):88–92. doi: 10.1126/science.aat3628. [DOI] [PubMed] [Google Scholar]
- 111.Mittnik A, et al. The genetic prehistory of the Baltic Sea region. Nat Commun. 2018;9(1):442. doi: 10.1038/s41467-018-02825-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Moreno-Mayar JV, et al. Terminal Pleistocene Alaskan genome reveals first founding population of Native Americans. Nature. 2018;553(7687):203–207. doi: 10.1038/nature25173. [DOI] [PubMed] [Google Scholar]
- 113.Moreno-Mayar JV, et al. Early human dispersals within the Americas. Science. 2018;362(6419):eaav2621. doi: 10.1126/science.aav2621. [DOI] [PubMed] [Google Scholar]
- 114.O’Sullivan N, et al. Ancient genome-wide analyses infer kinship structure in an Early Medieval Alemannic graveyard. Sci Adv. 2018;4(9):eaao1262. doi: 10.1126/sciadv.aao1262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Olalde I, et al. The Beaker phenomenon and the genomic transformation of northwest Europe. Nature. 2018;555(7695):190–196. doi: 10.1038/nature25738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Posth C, et al. Language continuity despite population replacement in Remote Oceania. Nat Ecol Evol. 2018;2(4):731–740. doi: 10.1038/s41559-018-0498-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Posth C, et al. Reconstructing the Deep Population History of Central and South America. Cell. 2018;175(5):1185–1197. doi: 10.1016/j.cell.2018.10.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Scheib CL, et al. Ancient human parallel lineages within North America contributed to a coastal expansion. Science. 2018;360(6392):1024–1027. doi: 10.1126/science.aar6851. [DOI] [PubMed] [Google Scholar]
- 119.Schroeder H, et al. Origins and genetic legacies of the Caribbean Taino. Proc Natl Acad Sci USA. 2018;115(10):2341–2346. doi: 10.1073/pnas.1716839115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Slon V, et al. The genome of the offspring of a Neanderthal mother and a Denisovan father. Nature. 2018;561(7721):113–116. doi: 10.1038/s41586-018-0455-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Valdiosera C, et al. Four millennia of Iberian biomolecular prehistory illustrate the impact of prehistoric migrations at the far end of Eurasia. Proc Natl Acad Sci USA. 2018;115(13):3428–3433. doi: 10.1073/pnas.1717762115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.van, de, Loosdrecht M, et al. Pleistocene North African genomes link Near Eastern and sub-Saharan African human populations. Science. 2018;360(6388):548–552. doi: 10.1126/science.aar8380. [DOI] [PubMed] [Google Scholar]
- 123.Veeramah KR, et al. Population genomic analysis of elongated skulls reveals extensive female-biased immigration in Early Medieval Bavaria. Proc Natl Acad Sci USA. 2018;115(13):3494–3499. doi: 10.1073/pnas.1719880115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Zalloua P, et al. Ancient DNA of Phoenician remains indicates discontinuity in the settlement history of Ibiza. Sci Rep. 2018;8(1):17567. doi: 10.1038/s41598-018-35667-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Amazonia, et al. The Current Genomic Landscape of Western South America: Andes, Amazonia, and Pacific Coast. Mol Biol Evol. 2019;36(12):2698–2713. doi: 10.1093/molbev/msz174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Antonio ML, et al. Ancient Rome: A genetic crossroads of Europe and the Mediterranean. Science. 2019;366(6466):708–714. doi: 10.1126/science.aay6826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Biagini SA, et al. People from Ibiza: an unexpected isolate in the Western Mediterranean. Eur J Hum Genet. 2019;27(6):941–951. doi: 10.1038/s41431-019-0361-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Brace S, et al. Ancient genomes indicate population replacement in Early Neolithic Britain. Nat Ecol Evol. 2019;3(5):765–771. doi: 10.1038/s41559-019-0871-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Fan S, et al. African evolutionary history inferred from whole genome sequence data of 44 indigenous African populations. Genome Biol. 2019;20(1):82. doi: 10.1186/s13059-019-1679-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Feldman M, et al. Ancient DNA sheds light on the genetic origins of early Iron Age Philistines. Sci Adv. 2019;5(7):eaax0061. doi: 10.1126/sciadv.aax0061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Feldman M, et al. Late Pleistocene human genome suggests a local origin for the first farmers of central Anatolia. Nat Commun. 2019;10(1):1218. doi: 10.1038/s41467-019-09209-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Flegontov P, et al. Palaeo-Eskimo genetic ancestry and the peopling of Chukotka and North America. Nature. 2019;570(7760):236–240. doi: 10.1038/s41586-019-1251-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.González-Fortes G, et al. A western route of prehistoric human migration from Africa into the Iberian Peninsula. Proc Biol Sci. 2019;286(1895):20182288. doi: 10.1098/rspb.2018.2288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Haber M, et al. A Transient Pulse of Genetic Admixture from the Crusaders in the Near East Identified from Ancient Genome Sequences. Am J Hum Genet. 2019;104(5):977–984. doi: 10.1016/j.ajhg.2019.03.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Harney É, et al. Ancient DNA from the skeletons of Roopkund Lake reveals Mediterranean migrants in India. Nat Commun. 2019;10(1):3670. doi: 10.1038/s41467-019-11357-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Järve M, et al. Shifts in the Genetic Landscape of the Western Eurasian Steppe Associated with the Beginning and End of the Scythian Dominance. Curr Biol. 2019;29(14):2430–2441. doi: 10.1016/j.cub.2019.06.019. [DOI] [PubMed] [Google Scholar]
- 137.Jensen TZT, et al. A 5700 year-old human genome and oral microbiome from chewed birch pitch. Nat Commun. 2019;10(1):5520. doi: 10.1038/s41467-019-13549-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Jeong C, et al. The genetic history of admixture across inner Eurasia. Nat Ecol Evol. 2019;3(6):966–976. doi: 10.1038/s41559-019-0878-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Mallick S, et al. The genomic history of the Iberian Peninsula over the past 8000 years. Science. 2019;363(6432):1230–1234. doi: 10.1126/science.aav4040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Malmström H, et al. The genomic ancestry of the Scandinavian Battle Axe Culture people and their relation to the broader Corded Ware horizon. Proc Biol Sci. 2019;286(1912):20191528. doi: 10.1098/rspb.2019.1528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Mittnik A, et al. Kinship-based social inequality in Bronze Age Europe. Science. 2019;366(6466):731–734. doi: 10.1126/science.aax6219. [DOI] [PubMed] [Google Scholar]
- 142.Narasimhan VM, et al. The formation of human populations in South and Central Asia. Science. 2019;365(6457):eaat7487. doi: 10.1126/science.aat7487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Nikitin AG, et al. Interactions between earliest Linearbandkeramik farmers and central European hunter gatherers at the dawn of European Neolithization. Sci Rep. 2019;9(1):19544. doi: 10.1038/s41598-019-56029-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Ning C, et al. Ancient Genomes Reveal Yamnaya-Related Ancestry and a Potential Source of Indo-European Speakers in Iron Age Tianshan. Curr Biol. 2019;29(15):2526–2532. doi: 10.1016/j.cub.2019.06.044. [DOI] [PubMed] [Google Scholar]
- 145.Prendergast ME, et al. Ancient DNA reveals a multistep spread of the first herders into sub-Saharan Africa. Science. 2019;365(6448):eaaw6275. doi: 10.1126/science.aaw6275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Reynolds AW, et al. Comparing signals of natural selection between three Indigenous North American populations. Proc Natl Acad Sci USA. 2019;116(19):9312–9317. doi: 10.1073/pnas.1819467116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Saag L, et al. The Arrival of Siberian Ancestry Connecting the Eastern Baltic to Uralic Speakers further East. Curr Biol. 2019;29(10):1701–1711. doi: 10.1016/j.cub.2019.04.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Sánchez-Quinto F, et al. Megalithic tombs in western and northern Neolithic Europe were linked to a kindred society. Proc Natl Acad Sci USA. 2019;116(19):9469–9474. doi: 10.1073/pnas.1818037116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Scheib CL, et al. East Anglian early Neolithic monument burial linked to contemporary Megaliths. Ann Hum Biol. 2019;46(2):145–149. doi: 10.1080/03014460.2019.1623912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Schroeder H, et al. Unraveling ancestry, kinship, and violence in a Late Neolithic mass grave. Proc Natl Acad Sci USA. 2019;116(22):10705–10710. doi: 10.1073/pnas.1820210116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Shinde V, et al. An Ancient Harappan Genome Lacks Ancestry from Steppe Pastoralists or Iranian Farmers. Cell. 2019;179(3):729–735. doi: 10.1016/j.cell.2019.08.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Sikora M, et al. The population history of northeastern Siberia since the Pleistocene. Nature. 2019;570(7760):182–188. doi: 10.1038/s41586-019-1279-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Villalba-Mouco V, et al. Survival of Late Pleistocene Hunter-Gatherer Ancestry in the Iberian Peninsula. Curr Biol. 2019;29(7):1169–1177. doi: 10.1016/j.cub.2019.02.006. [DOI] [PubMed] [Google Scholar]
- 154.Wang CC, et al. Ancient human genome-wide data from a 3000-year interval in the Caucasus corresponds with eco-geographic regions. Nat Commun. 2019;10(1):590. doi: 10.1038/s41467-018-08220-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Agranat-Tamir L, et al. The Genomic History of the Bronze Age Southern Levant. Cell. 2020;181(5):1146–1157. doi: 10.1016/j.cell.2020.04.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Bongers JL, et al. Integration of ancient DNA with transdisciplinary dataset finds strong support for Inca resettlement in the south Peruvian coast. Proc Natl Acad Sci USA. 2020;117(31):18359–18368. doi: 10.1073/pnas.2005965117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Brunel S, et al. Ancient genomes from present-day France unveil 7,000 years of its demographic history. Proc Natl Acad Sci USA. 2020;117(23):12791–12798. doi: 10.1073/pnas.1918034117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Burger J, et al. Low Prevalence of Lactase Persistence in Bronze Age Europe Indicates Ongoing Strong Selection over the Last 3,000 Years. Curr Biol. 2020;30(21):4307–4315. doi: 10.1016/j.cub.2020.08.033. [DOI] [PubMed] [Google Scholar]
- 159.Cassidy LM, et al. A dynastic elite in monumental Neolithic society. Nature. 2020;582(7812):384–388. doi: 10.1038/s41586-020-2378-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Coutinho A, et al. The Neolithic Pitted Ware culture foragers were culturally but not genetically influenced by the Battle Axe culture herders. Am J Phys Anthropol. 2020;172(4):638–649. doi: 10.1002/ajpa.24079. [DOI] [PubMed] [Google Scholar]
- 161.Csáky V, et al. Early medieval genetic data from Ural region evaluated in the light of archaeological evidence of ancient Hungarians. Sci Rep. 2020;10(1):19137. doi: 10.1038/s41598-020-75910-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Fernandes DM, et al. The spread of steppe and Iranian-related ancestry in the islands of the western Mediterranean. Nat Ecol Evol. 2020;4(3):334–345. doi: 10.1038/s41559-020-1102-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Furtwängler A, et al. Comparison of target enrichment strategies for ancient pathogen DNA. Biotechniques. 2020;69(6):455–459. doi: 10.2144/btn-2020-0100. [DOI] [PubMed] [Google Scholar]
- 164.Gokhman D, et al. Differential DNA methylation of vocal and facial anatomy genes in modern humans. Nat Commun. 2020;11(1):1189. doi: 10.1038/s41467-020-15020-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165.Haber M, et al. A Genetic History of the Near East from an aDNA Time Course Sampling Eight Points in the Past 4,000 Years. Am J Hum Genet. 2020;107(1):149–157. doi: 10.1016/j.ajhg.2020.05.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Jeong C, et al. A Dynamic 6,000-Year Genetic History of Eurasia’s Eastern Steppe. Cell. 2020;183(4):890–904. doi: 10.1016/j.cell.2020.10.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Linderholm A, et al. Corded Ware cultural complexity uncovered using genomic and isotopic analysis from south-eastern Poland. Sci Rep. 2020;10(1):6885. doi: 10.1038/s41598-020-63138-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Lipson M, et al. Ancient West African foragers in the context of African population history. Nature. 2020;577(7792):665–670. doi: 10.1038/s41586-020-1929-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Lipson M, et al. Three Phases of Ancient Migration Shaped the Ancestry of Human Populations in Vanuatu. Curr Biol. 2020;30(24):4846–4856. doi: 10.1016/j.cub.2020.09.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Liu D, et al. Extensive Ethnolinguistic Diversity in Vietnam Reflects Multiple Sources of Genetic Diversity. Mol Biol Evol. 2020;37(9):2503–2519. doi: 10.1093/molbev/msaa099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Mafessoni F, et al. A high-coverage Neandertal genome from Chagyrskaya Cave. Proc Natl Acad Sci USA. 2020;117(26):15132–15136. doi: 10.1073/pnas.2004944117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Marcus JH, et al. Genetic history from the Middle Neolithic to present on the Mediterranean island of Sardinia. Nat Commun. 2020;11(1):939. doi: 10.1038/s41467-020-14523-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Margaryan A, et al. Population genomics of the Viking world. Nature. 2020;585(7825):390–396. doi: 10.1038/s41586-020-2688-8. [DOI] [PubMed] [Google Scholar]
- 174.Massilani D, et al. Denisovan ancestry and population history of early East Asians. Science. 2020;370(6516):579–583. doi: 10.1126/science.abc1166. [DOI] [PubMed] [Google Scholar]
- 175.Nägele K, et al. Genomic insights into the early peopling of the Caribbean. Science. 2020;369(6502):456–460. doi: 10.1126/science.aba8697. [DOI] [PubMed] [Google Scholar]
- 176.Nakatsuka N, et al. A Paleogenomic Reconstruction of the Deep Population History of the Andes. Cell. 2020;181(5):1131–1145. doi: 10.1016/j.cell.2020.04.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Nakatsuka N, et al. Ancient genomes in South Patagonia reveal population movements associated with technological shifts and geography. Nat Commun. 2020;11(1):3868. doi: 10.1038/s41467-020-17656-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 178.Nieves-Colón MA, et al. Ancient DNA Reconstructs the Genetic Legacies of Precontact Puerto Rico Communities. Mol Biol Evol. 2020;37(3):611–626. doi: 10.1093/molbev/msz267. [DOI] [PubMed] [Google Scholar]
- 179.Ning C, et al. Ancient genomes from northern China suggest links between subsistence changes and human migration. Nat Commun. 2020;11(1):2700. doi: 10.1038/s41467-020-16557-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Rivollat M, et al. Ancient genome-wide DNA from France highlights the complexity of interactions between Mesolithic hunter-gatherers and Neolithic farmers. Sci Adv. 2020;6(22):eaaz5344. doi: 10.1126/sciadv.aaz5344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Skourtanioti E, et al. Genomic History of Neolithic to Bronze Age Anatolia, Northern Levant, and Southern Caucasus. Cell. 2020;181(5):1158–1175. doi: 10.1016/j.cell.2020.04.044. [DOI] [PubMed] [Google Scholar]
- 182.Teschler-Nicola M, et al. Ancient DNA reveals monozygotic newborn twins from the Upper Palaeolithic. Commun Biol. 2020;3(1):650. doi: 10.1038/s42003-020-01372-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183.Wang, C. C. et al. The Genomic Formation of Human Populations in East Asia. bioRxiv, 2020. 2020.03.25.004606: 10.1101/2020.03.25.004606.
- 184.Wang K, et al. Ancient genomes reveal complex patterns of population movement, interaction, and replacement in sub-Saharan Africa. Sci Adv. 2020;6(24):eaaz0183. doi: 10.1126/sciadv.aaz0183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Yang MA, et al. Ancient DNA indicates human population shifts and admixture in northern and southern China. Science. 2020;369(6501):282–288. doi: 10.1126/science.aba0909. [DOI] [PubMed] [Google Scholar]
- 186.Yu H, et al. Paleolithic to Bronze Age Siberians Reveal Connections with First Americans and across Eurasia. Cell. 2020;181(6):1232–1245. doi: 10.1016/j.cell.2020.04.037. [DOI] [PubMed] [Google Scholar]
- 187.Bortolini E, et al. Early Alpine occupation backdates westward human migration in Late Glacial Europe. Curr Biol. 2021;31(11):2484–2493. doi: 10.1016/j.cub.2021.03.078. [DOI] [PubMed] [Google Scholar]
- 188.Capodiferro MR, et al. Archaeogenomic distinctiveness of the Isthmo-Colombian area. Cell. 2021;184(7):1706–1723. doi: 10.1016/j.cell.2021.02.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Carlhoff S, et al. Genome of a middle Holocene hunter-gatherer from Wallacea. Nature. 2021;596(7873):543–547. doi: 10.1038/s41586-021-03823-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Clemente F, et al. The genomic history of the Aegean palatial civilizations. Cell. 2021;184(10):2565–2586. doi: 10.1016/j.cell.2021.03.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Cooke NP, et al. Ancient genomics reveals tripartite origins of Japanese populations. Sci Adv. 2021;7(38):eabh2419. doi: 10.1126/sciadv.abh2419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192.Egfjord AF, et al. Genomic Steppe ancestry in skeletons from the Neolithic Single Grave Culture in Denmark. PLoS One. 2021;16(1):e0244872. doi: 10.1371/journal.pone.0244872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.Fernandes DM, et al. A genetic history of the pre-contact Caribbean. Nature. 2021;590(7844):103–110. doi: 10.1038/s41586-020-03053-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Freilich S, et al. Reconstructing genetic histories and social organisation in Neolithic and Bronze Age Croatia. Sci Rep. 2021;11(1):16729. doi: 10.1038/s41598-021-94932-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195.Gnecchi-Ruscone GA, et al. Ancient genomic time transect from the Central Asian Steppe unravels the history of the Scythians. Sci Adv. 2021;7(13):eabe4414. doi: 10.1126/sciadv.abe4414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Hajdinjak M, et al. Initial Upper Palaeolithic humans in Europe had recent Neanderthal ancestry. Nature. 2021;592(7853):253–257. doi: 10.1038/s41586-021-03335-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197.Harney É, et al. A minimally destructive protocol for DNA extraction from ancient teeth. Genome Res. 2021;31(3):472–483. doi: 10.1101/gr.267534.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198.Immel A, et al. Genome-wide study of a Neolithic Wartberg grave community reveals distinct HLA variation and hunter-gatherer ancestry. Commun Biol. 2021;113(1):113. doi: 10.1038/s42003-020-01627-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 199.Ingman T, et al. Turkey during the 2nd millennium BC: Integration of isotopic and genomic evidence. PLoS One. 2021;16(6):e0241883. doi: 10.1371/journal.pone.0241883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Kılınç GM, et al. Human population dynamics and Yersinia pestis in ancient northeast Asia. Sci Adv. 2021;7(2):eabc4587. doi: 10.1126/sciadv.abc4587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201.Larena M, et al. Multiple migrations to the Philippines during the last 50,000 years. Proc Natl Acad Sci USA. 2021;118(13):e2026132118. doi: 10.1073/pnas.2026132118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.López S, et al. Evidence of the interplay of genetics and culture in Ethiopia. Nat Commun. 2021;12(1):3581. doi: 10.1038/s41467-021-23712-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 203.Mao X, et al. The deep population history of northern East Asia from the Late Pleistocene to the Holocene. Cell. 2021;184(12):3256–3266. doi: 10.1016/j.cell.2021.04.040. [DOI] [PubMed] [Google Scholar]
- 204.Novak M, et al. Genome-wide analysis of nearly all the victims of a 6200 year old massacre. PLoS One. 2021;16(3):e0247332. doi: 10.1371/journal.pone.0247332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 205.Papac L, et al. Dynamic changes in genomic and social structures in third millennium BCE central Europe. Sci Adv. 2021;7(35):eabi6941. doi: 10.1126/sciadv.abi6941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.Popović D, et al. Ancient genomes reveal long-range influence of the pre-Columbian culture and site of Tiwanaku. Sci Adv. 2021;7(39):eabg7261. doi: 10.1126/sciadv.abg7261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207.Posth C, et al. The origin and legacy of the Etruscans through a 2000-year archeogenomic time transect. Sci Adv. 2021;7(39):eabi7673. doi: 10.1126/sciadv.abi7673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 208.Prüfer K, et al. A genome sequence from a modern human skull over 45,000 years old from Zlatý kůň in Czechia. Nat Ecol Evol. 2021;5(6):820–825. doi: 10.1038/s41559-021-01443-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 209.Pugach I, et al. Ancient DNA from Guam and the peopling of the Pacific. Proc Natl Acad Sci USA. 2021;118(1):e2022112118. doi: 10.1073/pnas.2022112118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 210.Robbeets M, et al. Triangulation supports agricultural spread of the Transeurasian languages. Nature. 2021;599(7886):616–621. doi: 10.1038/s41586-021-04108-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Saag L, et al. Genetic ancestry changes in Stone to Bronze Age transition in the East European plain. Sci Adv. 2021;7(4):eabd6535. doi: 10.1126/sciadv.abd6535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 212.Saupe T, et al. Ancient genomes reveal structural shifts after the arrival of Steppe-related ancestry in the Italian Peninsula. Curr Biol. 2021;31(12):2576–2591. doi: 10.1016/j.cub.2021.04.022. [DOI] [PubMed] [Google Scholar]
- 213.Seguin-Orlando A, et al. Heterogeneous Hunter-Gatherer and Steppe-Related Ancestries in Late Neolithic and Bell Beaker Genomes from Present-Day France. Curr Biol. 2021;31(5):1072–1083. doi: 10.1016/j.cub.2020.12.015. [DOI] [PubMed] [Google Scholar]
- 214.Silva M, et al. Biomolecular insights into North African-related ancestry, mobility and diet in eleventh-century Al-Andalus. Sci Rep. 2021;11(1):18121. doi: 10.1038/s41598-021-95996-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215.Sirak KA, et al. Social stratification without genetic differentiation at the site of Kulubnarti in Christian Period Nubia. Nat Commun. 2021;12(1):7283. doi: 10.1038/s41467-021-27356-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 216.Svensson E, et al. Genome of Peştera Muierii skull shows high diversity and low mutational load in pre-glacial Europe. Curr Biol. 2021;31(14):2973–2983. doi: 10.1016/j.cub.2021.04.045. [DOI] [PubMed] [Google Scholar]
- 217.Villalba-Mouco V, et al. Genomic transformation and social organization during the Copper Age-Bronze Age transition in southern Iberia. Sci Adv. 2021;7(47):eabi7038. doi: 10.1126/sciadv.abi7038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 218.Wang CC, et al. Genomic insights into the formation of human populations in East Asia. Nature. 2021;591(7850):413–419. doi: 10.1038/s41586-021-03336-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 219.Wang T, et al. Human population history at the crossroads of East and Southeast Asia since 11,000 years ago. Cell. 2021;184(14):3829–3841. doi: 10.1016/j.cell.2021.05.018. [DOI] [PubMed] [Google Scholar]
- 220.Yaka R, et al. Variable kinship patterns in Neolithic Anatolia revealed by ancient genomes. Curr Biol. 2021;31(11):2455–2468. doi: 10.1016/j.cub.2021.03.050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 221.Žegarac A, et al. Ancient genomes provide insights into family structure and the heredity of social status in the early Bronze Age of southeastern Europe. Sci Rep. 2021;11(1):10072. doi: 10.1038/s41598-021-89090-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 222.Zhang F, et al. The genomic origins of the Bronze Age Tarim Basin mummies. Nature. 2021;599(7884):256–261. doi: 10.1038/s41586-021-04052-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 223.Alpaslan-Roodenberg S, et al. The genetic history of the Southern Arc: A bridge between West Asia and Europe. Science. 2022;377(6609):eabm4247. doi: 10.1126/science.abm4247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 224.Aneli S, et al. The Genetic Origin of Daunians and the Pan-Mediterranean Southern Italian Iron Age Context. Mol Biol Evol. 2022;39(2):msac014. doi: 10.1093/molbev/msac014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225.Changmai P, et al. Ancient DNA from Protohistoric Period Cambodia indicates that South Asians admixed with local populations as early as 1st-3rd centuries CE. Sci Rep. 2022;12(1):22507. doi: 10.1038/s41598-022-26799-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 226.Changmai P, et al. Indian genetic heritage in Southeast Asian populations. PLoS Genet. 2022;18(2):e1010036. doi: 10.1371/journal.pgen.1010036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 227.Childebayeva A, et al. Population Genetics and Signatures of Selection in Early Neolithic European Farmers. Mol Biol Evol. 2022;39(6):msac108. doi: 10.1093/molbev/msac108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 228.De, et al. First Glimpse into the Genomic Characterization of People from the Imperial Roman Community of Casal Bertone (Rome, First–Third Centuries AD) Genes. 2022;13(1):136. doi: 10.3390/genes13010136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 229.Dulias K, et al. Ancient DNA at the edge of the world: Continental immigration and the persistence of Neolithic male lineages in Bronze Age Orkney. Proc Natl Acad Sci USA. 2022;119(8):e2108001119. doi: 10.1073/pnas.2108001119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 230.Fowler C, et al. A high-resolution picture of kinship practices in an Early Neolithic tomb. Nature. 2022;601(7894):584–587. doi: 10.1038/s41586-021-04241-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 231.Gelabert P, et al. Northeastern Asian and Jomon-related genetic structure in the Three Kingdoms period of Gimhae, Korea. Curr Biol. 2022;32(15):3232–3244. doi: 10.1016/j.cub.2022.06.004. [DOI] [PubMed] [Google Scholar]
- 232.Gelabert P, et al. Genomes from Verteba cave suggest diversity within the Trypillians in Ukraine. Sci Rep. 2022;12(1):7242. doi: 10.1038/s41598-022-11117-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 233.Gnecchi-Ruscone GA, et al. Ancient genomes reveal origin and rapid trans-Eurasian migration of 7th century Avar elites. Cell. 2022;185(8):1402–1413. doi: 10.1016/j.cell.2022.03.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 234.Gretzinger J, et al. The Anglo-Saxon migration and the formation of the early English gene pool. Nature. 2022;610(7930):112–119. doi: 10.1038/s41586-022-05247-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 235.Hannah, M. et al. A Genetic History of Continuity and Mobility in the Iron Age Central Mediterranean. bioRxiv, 2022. 2022.03.13.483276: 10.1101/2022.03.13.483276.
- 236.Kennett DJ, et al. South-to-north migration preceded the advent of intensive farming in the Maya region. Nat Commun. 2022;13(1):1530. doi: 10.1038/s41467-022-29158-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 237.Kumar V, et al. Bronze and Iron Age population movements underlie Xinjiang population history. Science. 2022;376(6588):62–69. doi: 10.1126/science.abk1534. [DOI] [PubMed] [Google Scholar]
- 238.Lindo J, et al. The genomic prehistory of the Indigenous peoples of Uruguay. PNAS Nexus. 2022;1(2):pgac047. doi: 10.1093/pnasnexus/pgac047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 239.Lipson M, et al. Ancient DNA and deep population structure in sub-Saharan African foragers. Nature. 2022;603(7900):290–296. doi: 10.1038/s41586-022-04430-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 240.Liu CC, et al. Ancient genomes from the Himalayas illuminate the genetic history of Tibetans and their Tibeto-Burman speaking neighbors. Nat Commun. 2022;13(1):1203. doi: 10.1038/s41467-022-28827-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241.Liu YC, et al. Ancient DNA reveals five streams of migration into Micronesia and matrilocality in early Pacific seafarers. Science. 2022;377(6601):72–79. doi: 10.1126/science.abm6536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 242.Margaret, L. et al. Stable population structure in Europe since the Iron Age, despite high mobility. bioRxiv, 2022. 2022.05.15.491973: 10.1101/2022.05.15.491973.
- 243.Maróti Z, et al. The genetic origin of Huns, Avars, and conquering Hungarians. Curr Biol. 2022;32(13):2858–2870. doi: 10.1016/j.cub.2022.04.093. [DOI] [PubMed] [Google Scholar]
- 244.Maryland. et al. Technical Report on Ancient DNA analysis of 27 African Americans from Catoctin Furnace, Maryland. bioRxiv, 2022. 2022.06.12.495320.: 10.1101/2022.06.12.495320.
- 245.Oliveira S, et al. Ancient genomes from the last three millennia support multiple human dispersals into Wallacea. Nat Ecol Evol. 2022;6(7):1024–1034. doi: 10.1038/s41559-022-01775-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 246.Patterson N, et al. Large-scale migration into Britain during the Middle to Late Bronze Age. Nature. 2022;601(7894):588–594. doi: 10.1038/s41586-021-04287-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 247.Reitsema LJ, et al. The diverse genetic origins of a Classical period Greek army. Proc Natl Acad Sci USA. 2022;119(41):e2205272119. doi: 10.1073/pnas.2205272119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 248.Rivollat M, et al. Ancient DNA gives new insights into a Norman Neolithic monumental cemetery dedicated to male elites. Proc Natl Acad Sci USA. 2022;119(18):e2120786119. doi: 10.1073/pnas.2120786119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 249.Scorrano G, et al. Bioarchaeological and palaeogenomic portrait of two Pompeians that died during the eruption of Vesuvius in 79 AD. Sci Rep. 2022;12(1):6468. doi: 10.1038/s41598-022-10899-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 250.Spyrou MA, et al. The source of the Black Death in fourteenth-century central Eurasia. Nature. 2022;606(7915):718–724. doi: 10.1038/s41586-022-04800-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 251.Srigyan M, et al. Bioarchaeological evidence of one of the earliest Islamic burials in the Levant. Commun Biol. 2022;5(1):554. doi: 10.1038/s42003-022-03508-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 252.Tiesler, et al. Life and death in early colonial Campeche: New insights from ancient DNA. Antiquity. 2022;96(388):937–954. doi: 10.15184/aqy.2022.79. [DOI] [Google Scholar]
- 253.Ullinger J, et al. A Bioarchaeological Investigation of Fraternal Stillborn Twins from Tell el-Hesi. Near Eastern Archaelogy. 2022;85(3):228–237. doi: 10.1086/720748. [DOI] [Google Scholar]
- 254.Waldman S, et al. Genome-wide data from medieval German Jews show that the Ashkenazi founder event pre-dated the 14th century. Cell. 2022;185(25):4703–4716. doi: 10.1016/j.cell.2022.11.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 255.Winkelbach L, et al. The genomic origins of the world’s first farmers world. Cell. 2022;185(11):1842–1859. doi: 10.1016/j.cell.2022.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 256.Wohns AW, et al. A unified genealogy of modern and ancient genomes. Science. 2022;375(6583):eabi8264. doi: 10.1126/science.abi8264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 257.Yu H, et al. Genomic and dietary discontinuities during the Mesolithic and Neolithic in Sicily. iScience. 2022;25(5):104244. doi: 10.1016/j.isci.2022.104244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 258.Armit I, et al. Kinship practices in Early Iron Age South-east Europe: Genetic and isotopic analysis of burials from the Dolge njive barrow cemetery, Dolenjska, Slovenia. Antiquity. 2023;97(392):403–418. doi: 10.15184/aqy.2023.2. [DOI] [Google Scholar]
- 259.Brielle ES, et al. Entwined African and Asian genetic roots of medieval peoples of the Swahili coast. Nature. 2023;615(7954):866–873. doi: 10.1038/s41586-023-05754-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 260.Koptekin D, et al. Spatial and temporal heterogeneity in human mobility patterns in Holocene Southwest Asia and the East Mediterranean. Curr Biol. 2023;33(1):41–57. doi: 10.1016/j.cub.2022.11.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Citations
Supplementary Materials
Data Availability Statement
The pipeline used for processing raw data generated within the Reich lab is available in the ‘Workflow Description Language’ (WDL) here: https://github.com/DReichLab/adna-workflow, and includes individual python scripts for components of the pipeline.