Abstract
In this study, two novel bacterial strains were isolated from coastal sediment of Weihai, China. The two strains were Gram-stain-negative and facultatively aerobic, designated 3-1745T and A346T. Based on phenotypic, genetic and phylogenetic properties, strains 3-1745T and A346T represent two novel species of the genus Marinobacterium. The results of genome analysis revealed many central carbohydrate metabolism pathways such as gluconeogenesis, pyruvate oxidation, tricyclic acid cycle, pentose phosphate pathway and PRPP biosynthesis in the genus Marinobacterium. The ability of strains 3-1745T and A346T to utilize volatile fatty acids was experimentally confirmed. Polyhydroxyalkanoate synthases (PhaA, PhaB and PhaC) for the synthesis of polyhydroxyalkanoates were prevalent in the genus Marinobacterium. Multiple BGCs (biosynthetic gene clusters) including betalactone, ectoine, ranthipeptide, redox-cofactor, RiPPs (ribosomally synthesized post-translationally modified peptides) and T3PKS (polyketide synthases) in the genome of the genus Marinobacterium were found. Additional genome analyses suggested that the genus Marinobacterium contained diverse potential mechanisms of salt tolerance and mainly utilized oligosaccharides. This is the first report on broad genomic analyses of the genus Marinobacterium with the description of two novel species and potential ecological and biotechnological implications.
Keywords: comparative genomic analysis, Marinobacterium, polyhydroxyalkanoates, polyphasic taxonomy
Abbreviations
AAI, average amino acid identity; ANI, average nucleotide identity; BGC, biosynthetic gene cluster; CAZyme, carbohydrate-active enzyme; DDH, DNA–DNA hybridization; hserlactone, homoserine lactone; LAP, linear azol(in)e-containing peptides; ML, maximum-likelihood; PHA, polyhydroxyalkanoate; PRPP, phosphoribosyl diphosphate; RiPPs, ribosomally synthesized post-translationally modified peptides; SOX, sulphur-oxidizing; T3PKS, polyketide synthases.
Impact Statement
The genus Marinobacterium belongs to the family Oceanospirillaceae of the class Gammaproteobacteria in the phylum Pseudomonadota. Most members of the genus Marinobacterium have been isolated from different marine environments, such as seawater, sediment, coral, tidal flat and mangrove roots. In this work, we studied two strains isolated from marine sediment. Based on phenotypic, genetic and phylogenetic properties, strains A346T and 3-1745T represent two novel species of the genus Marinobacterium. Comparative genome analysis was conducted to study their properties and functions. Strains A346T and 3-1745T have potential to produce polyhydroxyalkanoates and harbour multiple secondary metabolites (biosynthetic gene clusters). They can oxidize thiosulphate to produce sulphate and may play roles in sulphur cycling. In this study, the isolation and identification of strains A346T and 3-1745T further enriched the marine microbial resources and gene library. The genus Marinobacterium has complete nitrogen fixation, dissimilatory nitrate reduction and denitrification pathways, which might foster future projects to develop and utilize the strains industrially. Our understanding of the genus Marinobacterium with regard to metabolic pathways and multiple potential mechanisms of salt tolerance was also deepened.
Data Summary
This study generated sequencing data for strains A346T and 3-1745T and all sequence data have been uploaded to the National Center for Biotechnology Information. The GenBank accession number for the 16S rRNA gene sequence and draft genome of strain A346T is MZ434947 and JAHQZT000000000. The GenBank accession number for the 16S rRNA gene sequence and draft genome of strain 3-1745T is MW391814 and JACEMT000000000. Detailed information can be found in Table S1 (available in the online version of this article).
Introduction
The genus Marinobacterium was first described in 1997, and was isolated from marine pulp mill effluent enrichment cultures [1]. The genus Marinobacterium widely exists in a variety of habitats; 18 species of the genus have been isolated from different environments, such as seawater [2], sediment [3], coral [4], tidal flat [5], mangrove roots [6] and high-salinity soil [7]. In addition, the genus Marinobacterium has also been detected in other samples including marine sediments taken from the Yellow Sea of South Korea [8], oil seep marine sediments from the southern Gulf of Mexico [9], rhizosphere soil from India [10], and different oil fields from Brazil, Malaysia and China [11–13]. The genome analysis results in this study suggest that the genus Marinobacterium contains diverse potential mechanisms of salt tolerance, and thus halotolerance is a general characteristic of members of this genus.
Some members of the genus Marinobacterium are capable of nitrogen-fixation [6], benzene-degradation [14], producing antimicrobial and cytotoxic ortho-dialkylbenzene-class metabolites [15], and the decomposition of marine aromatic compounds [16]. It has been reported that members of this genus can produce polyhydroxyalkanoates (PHAs) from various carbohydrates and volatile fatty acids [17]. PHAs have a variety of medical applications: biocontrol agents, drug carriers, biodegradable implants, anti-osteoporosis agents, medical devices, antibacterials, tissue engineering, memory enhancers and anticancer agents [18].
In order to further enrich marine microbial resources and understand their survival mechanism and application potentialities, our laboratory has carried out research on the mining of microbial resources. In this study, we collected coastal intertidal sediments from Xiaoshi Island, Weihai, China, which is located in a National Special Marine Reserve with a good degree of protection and little human exploitation. Measured salinity and pH values of the coastal sediment samples were 25‰ and 7.5, respectively. Two new strains of the genus Marinobacterium were isolated from marine sediments, the taxonomic positions of strains A346T and 3-1745T were determined by exploring phenotypic, genetic and phylogenetic properties, and comparative genome analysis was conducted to study their properties and potential functions. The two newly isolated strains had a short growth cycle, obvious colony formation within 1 day of inoculation, and were able to survive in a highly saline and alkaline environment, and through genomic analysis of the genus Marinobacterium, members of this group have the potential to produce PHAs. Reducing sulphides can be utilized by a variety of bacteria, the most commonly known being sulphur-oxidizing bacteria of the genera Thiobacillus [19], Thiomicrospira [20] and Thiothrix [21]. To date, thiosulphate oxidation in the genus Marinobacterium has not been reported. The current study showed that the genus Marinobacterium has the potential to oxidize thiosulphate and may participate in nitrogen fixation, dissimilatory nitrate reduction and denitrification, indicating that members of the genus may play an important role in geochemical cycles.
Methods
Bacterial isolation and identification
Strains A346T and 3-1745T were isolated from coastal sediment taken from Weihai, China (37° 31′ 3″ N 122° 1′ 6″ E). Sample pore water was extracted using a Rhizonsphere solution sampler (https://www.rhizosphere.com/rhizons) to determine sample salinity. Temperature and pH were measured by using a digital display rapid thermometer (TM902C; Qiankuang) and portable pH meter (PHB-5; Qiwei). Measured salinity, temperature and pH of the coastal sediment samples were 25 ‰, 20 °C and 7.5, respectively. The sediment sample was continuously diluted and resuspended in sterile seawater. Marine agar 2216 (MA; BD) spread with 10-fold diluted samples was incubated at 30 °C for 7 days. Strains A346T and 3-1745T, which formed yellowish-white, circular, shiny colonies, were picked and purified. They were stored at −80 °C in sterile 1 % (w/v) saline supplemented with 15 % (v/v) glycerol. The genomes of strains A346T and 3-1745T were extracted by using a DNA extraction kit (Takara). Details of the PCR conditions for amplification of the 16S rRNA gene were as given in Weisburg et al. [22, 23].
Almost-complete 16S rRNA gene sequences were obtained by cloning with pMD18-T vector (Takara) and sequencing primers M13F and M13R as previously described [24]. Sequencing was conducted by BGI (Qingdao, PR China). The nearly complete 16S rRNA gene sequences of strains A346T (1541 bp, with GenBank accession number MZ434947.1) and 3-1745T (1503 bp, with GenBank accession number MW391814.1) were obtained. The level of 16S rRNA gene sequence similarity between strains A346T and 3-1745T was calculated using the EzBioCloud (https://www.ezbiocloud.net/) [25] and NCBI databases (https://www.ncbi.nlm.nih.gov/). The nucleotide collections (nr) from the Standard database (NCBI databases) was used in BLASTn similarity searches. To determine their exact taxonomic status, we performed further phylogenetic and taxonomic studies. Phylogenetic trees were established by the maximum-likelihood (ML) [26] algorithm in the software package mega v11.0 [27], and all bootstrap values were based on 1000 replications. The ML tree was reconstructed using the best-fit substitution model GTR+G+I. The phylogenetic relationship based on a group of 120 conserved genes was analysed via GTDB-Tk [28], and the phylogenetic trees were reconstructed using IQ-TREE [29] with the LG+F+I+G4 model and 1000 bootstrap replicates, and against the GTDB database [30] to confirm the species novelty.
Marinobacterium stanieri DSM 7027T (obtained from Deutsche Sammlung von Mikroorganismen und Zellkulturen, DSMZ), Marinobacterium georgiense JCM 21667T (obtained from Japan Collection of Microorganisms, JCM) and Marinobacterium maritimum JCM 15134T (obtained from JCM) were used as reference strains. M. georgiense is the type species of the genus Marinobacterium. M. stanieri DSM 7027T and M. maritimum JCM 15134T have a close phylogenetic relationship with strains 3-1745T and A346T, respectively.
Genomic analysis
The genomes of strains A346T and 3-1745T were sequenced on the Hiseq X Ten platform (Illumina) at Beijing Novogene Bioinformatics Technology, using the pair-end 350 bp sequencing protocol. The raw data were filtered by fastp software [31] and all cleaned data were assembled by SOAPdenovo software v2.04 [32]. The quality of the assembled genomes was evaluated by using CheckM (v1.1.6) [33]. The genomes of other related strains were downloaded from the NCBI Prokaryotic reference genomes database. The draft genome content was annotated using the NCBI Prokaryotic Genome Annotation Pipeline (PGAP). To further confirm the taxonomic status of strains A346T and 3-1745T, the average nucleotide identity (ANI) was calculated using online tools of the EzBioCloud database (https://www.ezbiocloud.net/tools/ani) [34], DNA–DNA hybridization (DDH) values wee calculated by using the Genome-to-Genome Distance Calculator v2.1 [35], and the average amino acid identity (AAI) was calculated by CompareM (https://github.com/dparks1134/CompareM). To investigate the metabolic pathways and potential functions of bacteria, genomes were annotated using the KEGG database (v3.0) [36]. Bacterial resistance genes were annotated through RAST databases (v2.0), Proksee (https://proksee.ca/) [37] and the Comprehensive Antibiotic Resistance Database (CARD) (v3.2.4) [38, 39]. Clusters of orthologous groups (COGs) were annotated in eggNOG-mapper (v2.1.12) [40]. Prediction of secondary metabolites from draft genomes were conducted by using the antiSMASH 6.0 database [41]. Carbohydrate-active enzymes were annotated by the dbCAN2 meta server [42]. Pan-genomic analysis of bacteria was performed by BPGA v1.3 [43], determining core, accessory and unique gene numbers. The percentage of COG and KEGG categories of genes were calculated based on protein blast against reference COG and KEGG databases.
Phenotypic, physiological and biochemical characteristics
To examine phenotypic and physiological characteristics, the strains were cultured at 35 °C on MA plates. Both strains A346T and 3-1745T were able to grow at 35℃ and grew rapidly at this temperature. After incubation for 3 days, Gram-staining was examined using the method described previously described [44]. Determination of catalase activity was conducted by observing bubble production in 3 % (v/v) H2O2. Oxidase was determined by an oxidase reagent kit (bioMérieux) according to the manufacturer’s instructions. Cell morphology and size were assessed by using a light microscope (E600; Nikon), a scanning electron microscope (model Nova NanoSEM450; FEI) and a transmission electron microscope (JEM-1200; JEOL). Optimum temperatures for growth were tested at various temperatures (0, 4, 10, 15, 20, 25, 28, 30, 33, 35, 37, 40, 43, 44, 45 and 46 °C) on MA plates. Growth at different NaCl concentrations (0, 0.5, 1.0, 1.5, 2.0, 2.5, 3.0, 4.0, 5.0, 7.0, 8.0, 10.0, 12.0, 14.0, 16.0, 18.0, 19.0 and 20.0 %, w/v) was observed using a medium (0.5 % peptone, 0.1 % yeast extract and 2 % agar), prepared with artificial seawater (MgCl2 0.23 %, MgSO4 0.32 %, CaCl2 0.12 %, KCl 0.07 % and NaHCO3 0.02 %; all w/v) [45]. Growth at various pH (from pH 5.5 to 9.5, in increments of 0.5 pH units) was determined on Marine Broth 2216 (MB; BD). pH values were adjusted with buffers containing MES (pH 5.5–6.0), PIPES (pH 6.5–7.0), HEPES (pH 7.5–8.0), Tricine (pH 8.5) and CAPSO (pH 9.0–14.0) at 35 °C. Cell motility was determined on modified MA medium with 0.3 % agar. The abilities to hydrolyse DNA, starch, agar, casein, alginate, CM-cellulose and Tweens (20, 40, 60 and 80) were tested according to classical methods in bacterial taxonomy [44]. For testing anaerobic growth, strains were grown in anaerobic (10 % H2, 10 % CO2 and 80 % N2) and aerobic (5 % O2, 10 % CO2 and 85 % N2) environments on MA medium with or without 0.1 % (w/v) KNO3 for 2 weeks at 35 °C in an anaerobic jar. Antibiotic susceptibility tests of strains A346T and 3-1745T were performed on MA plates at 35 °C for up to 5 days using the disc-diffusion method [46]. Physiological and biochemical characteristics were described by using the results from API 20E kit (bioMérieux), API 50CHB kit (bioMérieux), API ZYM kits (bioMérieux) and Biolog GEN III MicroPlates according to the manufacturers’ instructions. Expression of the sulphur-oxidizing (SOX) system was tested by using a medium (0.5 % peptone, 0.1 % yeast extract and 0.03 % sodium pyruvate), prepared with artificial seawater (MgCl2 0.23 %, MgSO4 0.32 %, CaCl2 0.12 %, KCl 0.07 % and NaS2O4 0.5 %, all w/v). S2O3 2− and SO4 2− were detected by FeCl3 and BaCl2 solutions at 0 and 7 days [47].
Chemotaxonomic analyses
To determine chemotaxonomic characteristics, cells of strains A346T and 3-1745T, M. georgiense JCM 21667T, M. stanieri DSM 7027T and M. maritimum JCM 15134T were cultured in MB for 2 days at 30 °C, and then cells were collected and made into lyophilized powder. Isoprenoid quinones were determined using HPLC [48]. The extraction of polar lipids was extracted using a solution consisting of chloroform, methanol, and water (2.5:5:2, v/v/v) and the spilt off in two different directions on TLC plates according to a previously published method [49]. Fatty acids were obtained and detected using HPLC after adding 50 mg freeze-dried powder and a series of reagents into a 10 ml glass tube, followed by saponification, methylation, extraction and alkali washing [50].
Results
Bacterial isolation and identification
Strains A346T and 3-1745T, showing yellowish-white, circular and shiny colonies, were isolated from coastal sediment taken from Weihai, China (Fig. 1). The purified culture was stored at −80 °C in sterile 1 % (w/v) saline supplemented with 15 % (v/v) glycerol.
Fig. 1.
Map showing the sample collection site. The blue dot represents the sampling site, and the upper right corner shows an enlarged image of the sample collection site.
Nearly full-length 16S rRNA gene sequences of strains A346T (1541 bp) and 3-1745T (1503 bp) were obtained. Based on comparison against the EzBioCloud database, both strains A346T and 3-1745T had the highest 16S rRNA gene sequence similarity to M. stanieri DSM 7027T (97.0 and 98.4 %, respectively). The 16S rRNA gene sequence similarity between strain A346T (3-1745T) and other species of the genus Marinobacterium were 92.3–97.0 % (92.4–98.4 %). The 16S rRNA gene sequence similarity between the sediment-derived strains and other members of the genus Marinobacterium are detailed in Table S1. According to the results of 16S rRNA gene sequence analysis, the threshold between different bacterial species is 98.7 % [51]. The phylogenetic tree reconstructed with the ML algorithm indicated that strains A346T and 3-1745T were affiliated to the genus Marinobacterium (Fig. 2). The 16S rRNA gene sequence similarity between strains A346T and 3-1745T was 96.5 % and they were thus considered to represent two different novel species. Strain A346T was more closely related to M. maritimum JCM 15134T, and strain 3-1745T was more closely related to M. stanieri DSM 7027T in the 16S rRNA gene phylogenetic tree. The genomic phylogenetic tree also showed that both strains A346T and 3-1745T were affiliated to the genus Marinobacterium (Fig. 3). DDH, ANI and AAI values are listed in Fig. 4. It was clear that the DDH values were all <41 % and the ANI values were all <91%, which were lower than the species delineation thresholds (70 % for DDH and 95–96 % for ANI) [52, 53], indicating that strains A346T and 3-1745T represent two novel species of the genus Marinobacterium. The AAI values between all genomes were >65.7 %, which were higher than the genus delineation thresholds (65 % for AAI) [54], indicating that they were affiliated to the same genus. However, due to the large number of species in the genus Marinobacterium, the range of 16S rRNA gene sequence similarity, DDH, AAI and ANI values between strains A346T and 3-1745T with other members of this genus is relatively large, and 16S rRNA gene sequence similarity values even exceed the threshold range [55] for the same genus, which may also explain the presence of Neptuniibacter and Neptunomonas strains within the Marinobacterium cluster in the phylogenetic tree.
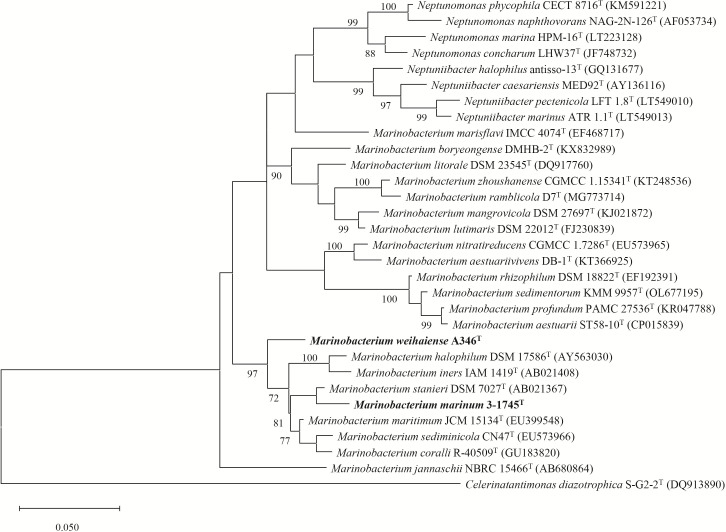
Fig. 2.
Maximum-likelihood phylogenetic tree based on 16S rRNA gene sequences showing the position of strains A346T and 3-1745T. Bootstrap values >70 % are shown at branch nodes. Celerinatantimonas diazotrophica S-G2-2T (DQ913890.1) was used as an outgroup. The 16S rRNA gene sequences of strains A346T and 3-1745T were extracted from the genome. Bar, 0.050 substitutions per nucleotide position.
Fig. 3.
IQ-TREE based on a group of 120 conserved genes showing the relationships between strains A346T, 3-1745T and related taxa. Bootstrap values >70 % are shown at branch nodes. Celerinatantimonas diazotrophica S-G2-2T (NZ_SMGD01000011.1) was used as an outgroup. Bar, 0.20 substitutions per nucleotide position.
Fig. 4.
Comparisons of the average nucleotide identity (ANI), DNA–DNA hybridization (DDH) and average amino acid identity (AAI) values between strains A346T, 3-1745T and related Marinobacterium type strains. (a) ANI and DDH values; (b) AAI values. Strains: 1, M. weihaiense A346T; 2, M. marinum 3-1745T; 3, M. stanieri DSM 7027T; 4, M. georgiense JCM 21667T; 5, M. halophilum DSM 17586T; 6, M. aestuarii ST58-10T; 7, M. alkalitolerans AK62T; 8, M. arenosum CAU 1594T; 9, M. jannaschii DSM 6295T; 10, M. litorale DSM 23545T; 11, M. lutimaris DSM 22012T; 12, M. mangrovicola DSM 27697T; 13, M. nitratireducens CGMCC 1.7286T; 14, M. profundum PAMC 27536T; 15, M. ramblicola D7T; 16, M. rhizophilum DSM 18822T; 17, M. zhoushanense CGMCC 1.15341T; 18, M. sedimentorum KMM 9957T.
Genomic characterization and quality assessment of the genus Marinobacterium
The draft genome of strain A346T (accession number GCA_019061305.1) was annotated with the NCBI PGAP. It contained 3351 genes (in total), including 3228 protein-coding genes and 77 RNAs genes (a complete 5S rRNA gene, a complete 16S rRNA gene, a complete 23S rRNA gene, 70 tRNA genes and four non-coding RNAs). One 16S rRNA gene sequence (1 540 bp) was detected from the genome of strain A346T and shared 99.9 % similarity with the one obtained by PCR. By contrast, a total of 3086 genes were annotated in the genome of strain 3-1745T (accession number GCA_013868415.1), including 2971 protein-coding genes and 79 RNA genes (three complete 5S rRNA genes, a complete 16S rRNA gene, three complete 23S rRNA genes, 68 tRNA genes and four non-coding RNAs). Only one 16S rRNA gene sequence (1 576 bp) was detected from the genome of strain 3-1745T and found to have 100 % similarity with the 16S rRNA gene sequence obtained by amplification. All genomic data (excluding strains A346T and 3-1745T) were obtained from the NCBI Genome Database (download before 20 May 2023). Detailed genomic data are shown in Table S2. The genomes of members of the genus Marinobacterium comprised 3.2–5.6 Mp with DNA G+C contents of 54.9–62.1 %. The DNA G+C contents of strains A346T and 3-1745T were respectively 58.9 and 56.4 %, occupying a medium level in the genus Marinobacterium. The genomic completeness and contamination were respectively above 98 % and below 3 %. Except for two strains (Marinobacterium arenosum CAU 1594T and Marinobacterium profundum PAMC 27536T genomes), the contig count of other bacteria was <100, and the value of N50 was >100 000. All this suggested that these genomic data were highly reliable. These two bacteria were not very different from other members of the genus Marinobacterium, which supported them belonging to this genus.
Comparative genomic analysis of the genus Marinobacterium was performed to determine the consistency and diversity of its members. Members of the genus have a total of 1085 core genes, which make up 20–40 % of the total (Fig. S1). The core genome is mainly distributed in amino acid metabolism, carbohydrate metabolism, energy metabolism, metabolism of cofactors and vitamins, nucleotide metabolism, overview and translation metabolism (Table S3). The accessory genome comprises to a large proportion lipid metabolism (fatty acid biosynthesis, fatty acid degradation, glycerolipid metabolism, etc), membrane transport (ABC transporters), signal transduction (two-component system), xenobiotic biodegradation and metabolism (benzoate degradation, aminobenzoate degradation, chloroalkane and chloroalkene degradation, etc), indicating that there are significant differences among members of this genus in substrate utilization, cell signal transduction and membrane transporter composition.
Gene functional classification of strains A346T and 3-1745T was predicted by the eggNOG database. The function classifications of strains A346T and 3-1745T mainly included RNA processing and modification (1, 1) (gene count of the two strains), chromatin structure and conversion (1, 1), energy production and conversion (217, 203), cell division/cycle control and chromosome partitioning (46, 45), amino acid transport and metabolism (223, 231), nucleotide transport and metabolism (72, 66), carbohydrate transport and metabolism (99, 126), coenzyme transport and metabolism (137, 137), lipid transport and metabolism (142, 103), translation, ribosomal structure and biogenesis (198, 201), transcription (189, 200), replication, recombination and repair (228, 200), cell wall/membrane/envelope biogenesis (174, 155), cell motility (121, 99), post-translational modification, protein turnover, chaperones (126, 113), inorganic ion transport and metabolism (169, 176), secondary metabolite biosynthesis, transport and catabolism (92, 73), function unknown (539, 497), signal transduction mechanisms (231, 166), intracellular trafficking, secretion, vesicular transport (106, 103), and defence mechanisms (49, 36) (Fig. S2). For example, the C4-dicarboxylate transporter stimulates the uptake of fumarate by bacteria [56]. Multiple ATP-binding cassette (ABC) transporters are involved in the absorption of nutrients and micronutrients [57]. In addition, pyruvate kinase, 3-phosphate dehydrogenase, fructose-1,6-bisphosphatase I and phosphoenolpyruvate carboxykinase in strains play an important role in central metabolism. The large proportion of substrate transport and metabolism genes and cell motility genes suggested that they might play a role in the growth and survival of bacteria.
Metabolic pathway analysis of the genus Marinobacterium
The genomes of 18 species of the genus Marinobacterium were analysed. Based on functional analyses, all Marinobacterium strains are primarily heterotrophic, with many complete central carbohydrate metabolism, energy metabolism, lipid metabolism, nucleotide metabolism and amino acid metabolism pathways (Fig. 5). The description of metabolism pathways is given in Table S4. Nine species, including those represented by strains A346T and 3-1745T, had the incomplete Embden–Meyerhoff pathway (M00001) and no glucokinase encoding gene was found. Central carbohydrate metabolism pathways such as gluconeogenesis, pyruvate oxidation, tricarboxylic acid (TCA) cycle, pentose phosphate pathway and PRPP (Phosphoribosyl diphosphate) biosynthesis were annotated in all bacterial genomes. However, strains A346T and 3-1745T were deficient in the Entner–Doudoroff pathway (M00008), which distinguishes the two novel strains from other species of the genus Marinobacterium. In terms of energy metabolism pathways, the nitrogen cycle is an important part of the global biogeochemical cycle, and plays an important role in biodiversity, climate change and human life [58]. Four strains had a complete nitrogen fixation pathway, indicating that they were important in providing a nitrogen source and improve oligotrophy in the marine environment. A complete pathway for dissimilatory nitrate reduction in five strains and denitrification in two strains were detected in their genomes; both consumption and utilization of nitrate make the novel strains of important potential value in improving water quality. Also, eight species including that of strain 3-1745T possessed a complete assimilatory sulphate reduction pathway (M00176), and six species including those of strains A346T and 3-1745T had a complete thiosulphate oxidation pathway by SOX complex (M00595), indicating that the genus Marinobacterium may play roles in sulphur and nitrogen cycling. Experimental results for the SOX system showed that strains A346T, 3-1745T, M. stanieri DSM 7027T and M. maritimum JCM 15134T could oxidize thiosulphate to produce sulphate. In contrast, M. georgiense JCM 21667T could not oxidize thiosulphate to produce sulphate, and the genome was annotated with an incomplete thiosulphate oxidation pathway. In terms of amino acid metabolism, strains A346T and 3-1745T had almost the same metabolic pathways, including the complete pathways for biosynthesis of leucine, threonine, cysteine and isoleucine, as well as incomplete betaine biosynthesis, ectoine degradation, polyamine biosynthesis, histidine degradation and GABA shunt. Histidine biosynthesis was annotated in the genomes of M. lutimaris DSM 22012T, M. nitratireducens CGMCC 1.7286T and M. marinum 3-1745T, which were different from the other strains. Significantly, almost all members of the genus Marinobacterium possessed partly complete fatty acid biosynthesis and beta-oxidation pathways.
Fig. 5.
Complete and incomplete metabolism pathways of the KEGG database in the genus Marinobacterium: 0, incomplete metabolism pathways; 1, complete metabolism pathways. Different colour codes in the y-axis represent different module types.
PHA production potential of the genus Marinobacterium
PHAs are microbial intracellular metabolites with plastic-like material properties, biodegradable properties and broad application prospects. PHAs are always produced under imbalanced metabolic conditions [59]. PHAs in bacteria serve as carbon and energy reserves that help strains to resist adverse stress conditions [60, 61]. As far as we are aware, three species of the genus Marinobacterium (M. nitratireducens, M. sediminicola and M. zhoushanense) have the ability to produce PHAs [17].
PhaA, PhaB and PhaC for the biosynthesis of PHAs were prevalent in the genus Marinobacterium. All genomes contained multiple PHA synthases (Fig. 6). Strains A346T and 3-1745T both possessed three PHA synthase (phaC) genes and one acetoacetyl-CoA reductase (phaB) gene. In particular, strain A346T encoded up to seven acetyl-CoA C-acetyltransferase (phaA) genes, compared to up to four phaA genes for strain 3-1745T. PHA production was accomplished through three steps sequentially mediated by acetyl-CoA C-acetyltransferase (EC 2.3.1.9; ACAT), acetoacetyl-CoA re-ductase (EC 1.1.1.36; phaB) and polyhydroxyalkanoate synthase (EC 2.3.1.304; phaC) or poly[(R)-3-hydroxyalkanoate] polymerase subunit PhaE (phaE). Micro-organisms are able to use different carbon sources such as sugars and fatty acids to form acetyl-CoA through multiple metabolic pathways, by acetyl-CoA C-acetyltransferase (phaA) to produce acetoacetyl-CoA. Acetoacetyl-CoA was reduced to (R)-3-hydroxybutanoyl-CoA by acetoacetyl-CoA reductase (phaB), and then (R)-3-hydroxybutanoyl-CoA was reduced to poly-β-hydroxybutyrate by PHA synthase (phaC). Specific complete metabolic pathways are shown in Fig. 7. All strains possessed the complete beta-oxidation pathway (M00086), producing acetyl-CoA. These results indicated that these 15 species, including those represented by the two novel strains A346T and 3-1745T, of the genus Marinobacterium might synthesize PHAs.
Fig. 6.
PHA synthase genes in Marinobacterium genomes. Numbers represent the gene count. K00626: ACAT, atoB; acetyl-CoA C-acetyltransferase; K00023: phbB; acetoacetyl-CoA reductase; K03821: phaC, phbC; poly[(R)−3-hydroxyalkanoate] polymerase subunit PhaC; K22881: phaE; poly[(R)-3-hydroxyalkanoate] polymerase subunit PhaE; K03737: por, nifJ; pyruvate-ferredoxin/flavodoxin oxidoreductase.
Fig. 7.
PHA synthetic pathways in the genus Marinobacterium. The enzymes shown in yellow are present in strains A346T and 3-1745T.
Prediction of biosynthetic gene clusters
In the genomes of the 18 type strains of the genus Marinobacterium, potential production of secondary metabolites was annotated by antiSMASH. All results are showed in Fig. 8. The results of genome analysis revealed the presence of multiple biosynthetic gene clusters (BGCs), including arylpolyene, betalactone, ectoine, hserlactone (homoserine lactone), LAP (Linear azol(in)e-containing peptides), NAGGN (nacetyl glutaminyl glutamine amide), NRPS (non-ribosomal peptide synthetases), ranthipeptide, redox-cofactor, RiPPs (ribosomally synthesised and post-translationally modified peptide product), RRE-containing (containing -RiPP recognition element-containing cluster), siderophore, T1PKS (polyketide synthases), T3PKS, terpene and tropodithietic acid. All genomes contained some common BGCs (betalactone, ectoine and T3PKS). The gene cluster for ectoine biosynthesis probably underlies the salinity tolerance strategy. Strain 3-1745T had gene clusters for the synthesis of five products, including ectoine, redox-cofactor, ranthipeptide, betalactone and T3PKS gene cluster. Strain A346T contained six gene clusters for the synthesis of ectoine, redox-cofactor, ranthipeptide, beta-lactone, RiPP-like and T3PKS. As all species of the genus Marinobacterium live in saline environments, the synthesis of multiple BGCS may be an adaptation to such environments [62]. M. jannaschii DSM 6295T and M. nitratireducens CGMCC 1.7286T had the largest number of gene clusters (15) among all strains. Members of the genus Marinobacterium isolated from scleractinian corals can produce marinoquinolones and marinobactoic acid that have antimicrobial activity and cytotoxicity [15]. These results could be utilized to support future microbial prospecting of the genus Marinobacterium.
Fig. 8.
Prediction of biosynthetic gene clusters for Marinobacterium strains annotated by the antiSMASH database. Bubble sizes represent the number of gene clusters.
In summary, strains A346T and 3-1745T carried some genes encoding compatible solute production (proA, proB, proC, proQ) and different genes encoding compatible solute transport as well as the presence of multiple BGCs including ectoine, and can survive in highly saline environments.
Resistome analyses
The results obtained in the antimicrobial susceptibility tests showed that strains 3-1745T and A346T were resistant to lincomycin (2 µg), vancomycin (30 µg), kanamycin (30 µg), streptomycin (10 µg), neomycin (30 µg) and tobramycin (10 µg), but susceptible to ofloxacin (5 µg), ampicillin (10 µg), penicillin (10 µg), polymyxin B (300 µg), ceftriaxone (30 µg), chloramphenicol (30 µg) and cefotaxime sodium (30 µg). There were some differences between the two strains. Strain A346T was resistant to carbenicillin (100 µg), norfloxacin (30 µg), erythromycin (15 µg), clarithromycin (10 µg), rifampin (5 µg), tetracycline (30 µg) and gentamycin (10 µg), but strain 3-1745T was susceptible to these compounds. The genomes of strains 3-1745T and A346T had the genes AAC (3)-IIa, CrcB, mexY, npmA and rmtB that are resistant to aminoglycoside antibiotics, which may explain their resistance to tobramycin and streptomycin. This may also be the cause of the resistance of strain A346T to gentamicin. Mobile genetic elements with replication/recombination and integration/excision/repair functions have been annotated around these genes, indicating that they have horizontal transfer risk [63–65]. In both strains 3-1745T and A346T, there are vancomycin resistance (van) gene clusters, tet (O), adeb and HelR, which may explain their resistance to vancomycin, tetracycline and rifampicin. Mobile genetic elements with replication/recombination and integration/excision/repair functions are also annotated around these genes, suggesting that they may have horizontal transfer risk [63–65]. Genome analyses of strains 3-1745T and A346T showed that they contained the resistance-nodulation-cell division (RND) antibiotic efflux pump gene family, multiple antimicrobial extrusion protein (Na+/drug antiporter) and MATE family of MDR efflux pumps. This suggested that resistance in these bacteria might be associated with resistance genes. Strain 3-1745T was resistant to multiple classes of antibiotics (lincosamides, aminoglycosides and glycopeptides), and strain A346T was resistant to lincosamides, ansamycins, macrolides, quinolones, aminoglycosides and glycopeptides (among others). Due to the multi-resistance characteristics of these two strains [66], this may also provide a new way to add antibiotics in industrial production to prevent contamination by other bacteria.
Microbial substrate preference
To investigate if the genus Marinobacterium could utilize oligosaccharides and polysaccharides, the composition of carbohydrate-active enzymes (CAZymes) was compared via the Carbohydrate-Active Enzymes Database. The results of genome analysis revealed that all strains had analogous CAZymes compositions, and glycosyltransferase (GTs) and glycoside hydrolase (GHs) were their dominant CAZymes (Figs 9 and 10). The description of CAZymes families is shown in Table S5. Polysaccharide lyases were absent in almost all strains. Strains A346T and 3-1745T contained 110 and 103 CAZymes, respectively. Glycoside hydrolase families GH23, GH24, GH25 and GH108 potentially involved in lysozyme production were detected, which could reflect a certain bacteriostatic ability of these strains, with possible application in food preservation and preservation, medicine and biological engineering. The other CAZyme families related to degradation of xylan (GH38, GH43_12 and GH51), cellobiose (GH5_12, GT94) and other multiple oligosaccharides (AA3, GT2 and GT4) were annotated in some strains. These results suggest that the genus Marinobacterium mainly utilizes oligosaccharides, consistent with carbon utilization results for strains A346T, 3-1745T, M. stanieri DSM 7027T, M. maritimum JCM 15134T and M. georgiense JCM 21667T (Tables 1 and S6).
Fig. 9.
Comparison of carbohydrate-active enzymes (CAZymes) between various Marinobacterium strains. AA: auxiliary activities; CBM: carbohydrate-binding module; CE: carbohydrate esterases; GH: glycoside hydrolases; GT: glycosyltransferases; PL: polysaccharide lyases.
Fig. 10.
Predicted number of carbohydrate-active enzymes detected in the genus Marinobacterium. The shade of the colour represents the number of genes.
Table 1.
Differential phenotypic characteristics between strains A346T, 3-1745T and related Marinobacterium type strains
Strains: 1, A346T; 2, 3-1745T; 3, M. georgiense JCM 21667T; 4, M.stanieri DSM 7027T; 5, M. maritimum JCM 15134T. +, Growth; w, weak growth; −, no growth. All data were obtained in this study unless indicated otherwise.
|
Characteristic |
1 |
2 |
3 |
4 |
5 |
|---|---|---|---|---|---|
|
Temperature for growth (°C) |
|||||
|
Range |
4–44 |
4–40 |
4–41* |
10–37† |
7–37‡ |
|
Optimum |
30–37 |
37 |
37* |
25–30† |
25–28‡ |
|
Cell size (μm) |
0.6–0.7×1.2–2.3 |
0.4–0.5×1.1–1.7 |
0.5–0.7×1.6–2.3* |
0.6–0.8×1.3–2.2† |
0.5–0.6×0.8–0.9‡ |
|
Salinity growth range (%) |
0.5–18 |
0–12 |
0–12* |
0.5–8† |
0.5–7‡ |
|
Optimum NaCl concentration (%, w/v) |
1 |
0.5 |
0.5–2* |
2† |
1–2‡ |
|
pH growth range |
6.0–10.0 |
6.0–9.5 |
5.0–10.0* |
5.0–12.0† |
5.5–9.0‡ |
|
Optimum pH growth |
7.0 |
7.0 |
7.0–8.0* |
7.0† |
7.5–8.0‡ |
|
Motility |
+ |
+ |
+* |
+† |
+‡ |
|
Nitrate reduction |
– |
– |
–* |
–† |
–‡ |
|
Hydrolysis of: |
|||||
|
Tween 20 |
– |
+ |
– |
– |
– |
|
Tween 40 |
– |
+ |
– |
+ |
– |
|
Tween 60 |
– |
– |
– |
– |
– |
|
Tween 80 |
– |
– |
+∗ |
– |
–‡ |
|
Citrate utilization |
– |
– |
w |
+ |
+ |
|
Enzymatic activity |
|||||
|
Oxidase |
+ |
+ |
+* |
+† |
+‡ |
|
Catalase |
+ |
– |
+* |
+ |
+‡ |
|
Acid phosphatase |
+ |
– |
+ |
+ |
+ |
|
Acid production from: |
|||||
|
l-Arabinose |
+ |
– |
– |
– |
– |
|
Ribose |
– |
+ |
+ |
+ |
+ |
|
l-Xylose |
+ |
– |
– |
– |
– |
|
Oxidation of: |
|||||
|
Saccharic acid |
+ |
– |
– |
– |
– |
|
Malic acid |
– |
w |
w |
– |
+ |
|
DNA G+C content (%) |
58.9 |
56.4 |
54.9* |
55.6† |
57.9‡ |
In addition, to explore the ability of this genus to utilize other substrates, annotated analysis of the genome revealed the existence of a variety of esterases (esterase, acetyl esterase and pimeloyl-[acyl-carrier protein] methyl ester esterase) and enzymes (acetate-CoA synthase, acetate kinase and phosphate acetyltransferase) related to acetic acid metabolism. It has also been confirmed experimentally that strains A346T, 3-1745T, M. stanieri DSM 7027T, M. maritimum JCM 15134T and M. georgiense JCM 21667T were positive for activity of alkaline phosphatase, esterase (C4), esterase lipase (C8) and lipase (C14) and positive for the utilization of l-lactic acid, α-hydroxy-butyric acid, β-hydroxy-d, l-butyric acid, propionic acid, acetic acid, sodium bromate, sodium butyrate, etc. In addition, strain 3-1745T was positive for hydrolysis of Tweens 20 and 40. It is reported that several species from the genus Marinobacterium including M. nitratireducens, M. sediminicola, and M. zhoushanense can producePHAs using sugars and volatile fatty acids as the carbon source [24].
Flagellar motor capacity
Flagellar movement plays an active role in many biological functions of bacteria, such as the formation of bacteria–host symbiosis, pathogenicity and antibiotic resistance [67, 68]. Through scanning electron microscopy and transmission electron microscopy, cell morphology showed that cells of strains A346T and 3-1745T were both rod-shaped with a single polar flagellum and approximately 0.6–0.7 µm wide and 1.2–2.3 µm long for strain A346T and approximately 0.4–0.5 µm wide and 1.1–1.7 µm long for strain 3-1745T (Table 1 and Fig. S3). Colony sizes for strains A346T and 3-1745T were 0.2–1.2 and 0.1–2.0 mm, respectively. Genome analysis revealed that each bacterium in the genus Marinobacterium contained multiple genes that encode flagella, which were used to synthesize its components (filament, filament cap, H ring, hook-filament junction, MS/C ring, type III secretion system, P/L ring and T ring Stator, etc.) [69, 70] (Fig. 11). Bacteria may make directional responses to environmental factors of different gradients through flagellar movement, so as to favour stimuli and avoid harmful stimuli [71]. Cells of members of the genus Marinobacterium were motile by a single or two polar flagella [72]. In the same family, the discovery of thick, polar flagellar filaments in Oleibacter marinus has expanded the known diversity of flagellar architecture [73]. The results suggested that flagella are a common feature of the genus, which could help bacteria actively gravitate toward nutrients or avoid harmful chemicals.
Fig. 11.
Flagella synthase genes in Marinobacterium sp. genomes. Numbers represent the gene count.
Physiological and chemotaxonomic analyses
Strain A346T could grow at 4–44 °C (optimum, 30–37 °C), with 0.5–18 % (w/v) NaCl (optimum, 1%) and at pH 6.0–10.0 (optimum, pH 7.0). Strain 3-1745T could grow at 4–40 °C (optimum, 37 °C), with 0–12 % (w/v) NaCl (optimum, 0.5 %) and at pH 6.0–9.5 (optimum, pH 7.0) (Table 1). By analysing the genome of the strains, A346T and 3-1745T contained multiple salt-tolerance genes (trkA, trkH, proA, proB, proC and proQ), which may be related to a wide salt tolerance shown by both strains during rearing in the lab.
The dominant fatty acids (>5 %) of strain A346T were summed feature 3 (C16 : 1ω7c/C16 : 1ω6c) (29.2 %), C16 : 0 (24.8 %), summed feature 8 (C18 : 1ω7c/C18 : 1ω6c) (22.0 %), C12 : 0 (6.4 %) and C10 : 0 3-OH (5.9 %). The dominant fatty acids (>5 %) of strain 3-1745T were summed feature 8 (C18 : 1ω7c/C18 : 1ω6c) (31.0 %), summed feature 3 (C16 : 1ω7c/C16 : 1ω6c) (27.9 %) and C16 : 0 (27.1 %). The cellular fatty acid compositions of A346T and 3-1745T and related type strains are shown in Table 2. Some dominant fatty acids of strains A346T and 3-1745T were similar with those of the other related type strains, such as C16 : 0 and C10 : 0 3-OH, but there were differences in the proportions of some fatty acids (C17 : 0 cyclo, C14 : 0). In strain A346T, the contents of C12 : 0, C14 : 0 and C10 : 0 3-OH were greater than those of the other four strains. The sole respiratory quinone of both strains 3-1745T and A346T was ubiquinone-8. The major polar lipids of strain A346T were phosphatidylglycerol (PG), phosphatidyl-ethanolamine (PE), an unidentified amino lipid (AL) and three unidentified lipids (L). The major polar lipids of strain 3-1745T were phosphatidylglycerol (PG), phosphatidyl-ethanolamine (PE), diphosphatidylglycerol (DPG), an unidentified aminolipid (AL), an unidentified phospholipid and five unidentifified lipids (L) (Fig. S4). The polar lipids of strains A346T, 3-1745T and related type strains had some in common (PE, PG), and strains A346T and 3-1745T had an unidentified aminolipid (AL) which was different from the other strains. In addition, strain 3-1745T contained one unidentified phospholipid (PL) and five unidentifified lipids, which were not found in the other strains.
Table 2.
Cellular fatty acid composition (%) of strains A346T, 3-1745T and related Marinobacterium type strains
Strains: 1, A346T; 2, 3-1745T; 3, M. georgiense JCM 21667T; 4, M. stanieri DSM 7027T; 5, M. maritimum JCM 15134T. All data were obtained in this study. Fatty acids present at >5 % are marked in bold. −, Not detected; tr, trace (<1 %).
|
Fatty acid |
1 |
2 |
3 |
4 |
5 |
|---|---|---|---|---|---|
|
Straight-chain fatty acids |
|||||
|
C10 : 0 |
4.2 |
2.4 |
3.8 |
tr |
2.7 |
|
C12 : 0 |
6.4 |
2.6 |
1.9 |
4.3 |
2.9 |
|
C14 : 0 |
1.1 |
tr |
tr |
tr |
tr |
|
C16 : 0 |
24.8 |
27.1 |
30.6 |
22.9 |
24.4 |
|
C18 : 0 |
tr |
1.3 |
1.2 |
2.0 |
tr |
|
Cyclopropane acids |
|||||
|
C17 : 0 cyclo |
– |
– |
– |
– |
3.4 |
|
Hydroxy fatty acids |
|||||
|
C10 : 0 3-OH |
6.0 |
4.3 |
4.3 |
4.4 |
4.5 |
|
Summed features* |
|||||
|
3 |
29.17 |
27.93 |
19.05 |
21.13 |
25.26 |
|
8 |
22.0 |
31.0 |
36.5 |
40.5 |
32.7 |
*Summed features represent groups of two or three fatty acids that could not be separated by GLC with the MIDI system. Summed feature 3 consisted of C16 : 1ω7c/C16 : 1ω6c and summed feature 8 consisted of C18 : 1ω7c/C18 : 1ω6c.
Description of Marinobacterium weihaiense sp. nov.
Marinobacterium weihaiense (wei.hai.en’se. N.L. neut. n. weihaiense originating from Weihai, China).
Cells are Gram-stain-negative, facultatively anaerobic, rod-shaped, motile and almost 0.6–0.7 µm width and 1.2–2.3 µm length. Colony size is 0.2–1.2 mm. Colonies are circular at 35 °C on MA after 2 days. Grows at 4–44 °C (optimum, 30–37 °C), with 0.5–18 % (w/v) NaCl (optimum, 1%) and at pH 6.0–10.0 (optimum, pH 7.0). Positive for Voges–Proskauer reaction. The activity of oxidase and catalase, as well as gelatinase, alkaline phosphatase, esterase (C4), leucine arylamidase, acid phosphatase and naphthol-AS-BI-phosphohydrolase are positive. The activity of esterase lipase (C8), lipase (C14), cystine arylamidase, N-acetyl-β-glucosaminidase and valine arylamidase are weakly positive. Negative for hydrolysis of agar, alginate, CM-cellulose, starch, Tweens 20, 40, 60 and 80, DNA and casein. Positive for utilization of tagatose, potassium 2-ketogluconate, potassium 5-ketogluconate, aesculin ferric citrate, amygdalin, methyl-β-d-xylopyranoside, l-xylose, l-arabinose, cellobiose, mannitol, galactose, l-rhamnose, fusidic acid, serine, l-alanine, glucuronamide, saccharic acid, α-ketoglutaric acid and acetic acid. The major fatty acids are summed feature 3 (C16 : 1ω7c/C16 : 1ω6c), C16 : 0, summed feature 8 (C18 : 1ω7c/C18 : 1ω6c) and C12 : 0. The sole respiratory quinone is ubiquinone-8. The major polar lipids are phosphatidylglycerol, phosphatidylethanolamine, an unidentified amino lipid and three unidentifified lipids. The genomic DNA G+C content of the type strain is 58.9 %.
The type strain, A346T (=KCTC 92007T=MCCC 1H00492T=SDUM032111T), was isolated from marine sediment from the coast of Weihai, China (36° 58′ 37″ N 122° 2 ′37″ E). The GenBank accession number for the 16S rRNA gene sequence of strain A346T is MZ434947.1 and the draft genome has been deposited in GenBank under accession number JAHQZT000000000.1.
Description of Marinobacterium marinum sp. nov.
Marinobacterium marinum (ma.ri’num. L. neut. adj. marinum of or belonging to the sea, marine).
Cells are rod-shaped, Gram-stain-negative, facultatively anaerobic, approximately 0.4–0.5 µm width and 1.1–1.7 µm length, and motile by means of a single polar flagellum. Colony size is 0.1–2.0 mm. Grows at 4–40 °C (optimum, 37 °C), with 0–12 % (w/v) NaCl (optimum, 0.5 %) and at pH 6.0–9.5 (optimum, pH 7.0). Positive results in tests for oxidase activity, Voges–Proskauer reaction and hydrolysis of Tweens 20 and 40, but negative for catalase activity, nitrate reduction, hydrolysis of agar, alginate, CM-cellulose, starch, Tweens 60 and 80, DNA and casein. Positive for activity of gelatinase, alkaline phosphatase, esterase (C4), leucine arylamidase and naphthol-AS-BI-phosphohydrolase and positive for utilization of glycerol, d-ribose, l-rhamnose, aesculin ferric citrate, cellobiose, raffinose, tagatose, potassium 2-ketogluconate, potassium 5-ketogluconate, fucose, fusidic acid, serine, l-alanine, l-glutamic acid, l-pyroglutamic acid, galacturonic acid, glucuronic acid, glucuronamide, l-lactic acid, α-ketoglutaric acid, propionic acid and acetic acid. The activity of esterase lipase (C8), lipase (C14) and valine arylamidase are weakly positive. The major fatty acids are summed feature 8 (C18 : 1ω7c/C18 : 1ω6c), summed feature 3 (C16 : 1ω7c/C16 : 1ω6c) and C16 : 0. The major polar lipids are phosphatidylglycerol, phosphatidylethanolamine, diphosphatidylglycerol, an unidentified amino lipid, an unidentified phospholipid and five unidentified lipids. The sole respiratory quinone is ubiquinone-8. The genomic DNA G+C content of the type strain is 56.4 %.
The type strain, 3-1745T (=KCTC 72925T=MCCC 1H00422T=SDUM 032109T), was isolated from marine sediment from the coast of Weihai, China (36° 58′ 37″ N 122° 2′ 37″ E). The GenBank accession number for 16S rRNA gene sequence of strain 3-1745T is MW391814.1 and the draft genome has been deposited in GenBank with accession number JACEMT000000000.1.
Conclusions
In this study, strains A346T and 3-1745T were isolated from coastal sediment taken from China. They were affiliated to the genus Marinobacterium, and represented the novel species Marinobacterium weihaiense and Marinobacterium marinum, respectively. Strains A346T and 3-1745T carried multiple genes encoding compatible solute production and compatible solute transport to help them resist the high-salinity environment. Strain 3-1745T had five BGCs, including ectoine, redox-cofactor, ranthipeptide, betalactone and T3PKS. Strain A346T contained six BGCs for synthesis of ectoine, redox-cofactor, ranthipeptide, beta-lactone, RiPP-like and T3PKS.
Genomic analyses of the two new strains indicated they have the capacity to utilize lipids and volatile fatty acids, and can grow rapidly in a high-salinity environment and have a strong ability to adapt to the environment. Through genomic analysis of the genus Marinobacterium, members of this group have a complete PHA synthesis pathway. This indicates that some members of the genus Marinobacterium have the ability to oxidize thiosulphate to sulphate. Some members had a complete nitrogen fixation, dissimilatory nitrate reduction and denitrification pathway, indicating that they may be important in providing a nitrogen source, improve oligotrophic marine environments and improve water quality. In this study, the isolation and identification of strains A346T and 3-1745T has further enriched the marine microbial resources and gene pool particularly known for this bacterial genus which has been poorly exploited from a biotechnological standpoint. There are multiple metabolic pathways in strains A346T and 3-1745T, including carbohydrate metabolism, energy metabolism, lipid metabolism, nucleotide metabolism and amino acid metabolism, which play an important role in the global carbon, nitrogen and sulphur cycle and are of great significance for maintaining global ecological balance. They have the potential ability to synthesize PHAs and multiple secondary metabolites, and have broad application prospects in the materials, agriculture, food and biomedical fields.
Supplementary Data
Funding information
This work was funded by Guangdong Basic and Applied Basic Research Foundation (2022A1515110773), National Natural Science Foundation of China (32200003) and Natural Science Foundation of Shandong Province (ZR2022QC106).
Acknowledgements
Scanning electron microscopy was supported by the Physical-Chemical Materials Analytical and Testing Center of Shandong University at Weihai.
Author contributions
X.-J.L. and Z.-T.H. isolated strains A346T and 3- 1745T, X.-J.L., K.-L.Z., Y.-Q.Y., Z.-T.H. and X.-Y.T. performed material preparation, experimental operation, data collection and analysis. K.-L.Z. helped process cell samples and photograph cell morphology. X.-J.L. finished the experiment and manuscript. Z.-J.D. and M.-Q.Y. offered experimental guidance and critical revision of manuscript. All authors contributed to the article and approved the submitted version.
Conflicts of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
- 1.González JM, Mayer F, Moran MA, Hodson RE, Whitman WB. Microbulbifer hydrolyticus gen. nov., sp. nov., and Marinobacterium georgiense gen. nov., sp. nov., two marine bacteria from a lignin-rich pulp mill waste enrichment community. Int J Syst Bacteriol. 1997;47:369–376. doi: 10.1099/00207713-47-2-369. [DOI] [PubMed] [Google Scholar]
- 2.Kang JY, Kim MJ, Chun J, Son KP, Jahng KY. Marinobacterium boryeongense sp. nov., isolated from seawater. Int J Syst Evol Microbiol. 2019;69:493–497. doi: 10.1099/ijsem.0.003184. [DOI] [PubMed] [Google Scholar]
- 3.Huo Y-Y, Xu X-W, Cao Y, Wang C-S, Zhu X-F, et al. Marinobacterium nitratireducens sp. nov. and Marinobacterium sediminicola sp. nov., isolated from marine sediment. Int J Syst Evol Microbiol. 2009;59:1173–1178. doi: 10.1099/ijs.0.005751-0. [DOI] [PubMed] [Google Scholar]
- 4.Chimetto LA, Cleenwerck I, Brocchi M, Willems A, De Vos P, et al. Marinobacterium coralli sp. nov., isolated from mucus of coral (Mussismilia hispida) Int J Syst Evol Microbiol. 2011;61:60–64. doi: 10.1099/ijs.0.021105-0. [DOI] [PubMed] [Google Scholar]
- 5.Park S, Jung YT, Kim S, Yoon JH. Marinobacterium aestuariivivens sp. nov., isolated from a tidal flat. Int J Syst Evol Microbiol. 2016;66:1718–1723. doi: 10.1099/ijsem.0.000927. [DOI] [PubMed] [Google Scholar]
- 6.Alfaro-Espinoza G, Ullrich MS. Marinobacterium mangrovicola sp. nov., a marine nitrogen-fixing bacterium isolated from mangrove roots of Rhizophora mangle. Int J Syst Evol Microbiol. 2014;64:3988–3993. doi: 10.1099/ijs.0.067462-0. [DOI] [PubMed] [Google Scholar]
- 7.Durán-Viseras A, Castro DJ, Reina JC, Béjar V, Martínez-Checa F. Taxogenomic and metabolic insights into Marinobacterium ramblicola sp. nov., a new slightly halophilic bacterium isolated from Rambla Salada, Murcia. Microorganisms. 2021;9:1654. doi: 10.3390/microorganisms9081654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Velmurugan N, Kalpana D, Cho J-Y, Lee G-H, Park S-H, et al. Phylogenetic analysis of culturable marine bacteria in sediments from the South Korean Yellow Sea. Mikrobiologiia. 2011;80:275–286. [PubMed] [Google Scholar]
- 9.Rosano-Hernández MC, Fernández-Linares LC. Bacterial diversity of marine seeps in the southeastern Gulf of Mexico. Pak J Biol Sci. 2009;12:683–689. doi: 10.3923/pjbs.2009.683.689. [DOI] [PubMed] [Google Scholar]
- 10.Sorty AM, Meena KK, Choudhary K, Bitla UM, Minhas PS, et al. Effect of plant growth promoting bacteria associated with halophytic weed (Psoralea corylifolia L) on germination and seedling growth of wheat under saline conditions. Appl Biochem Biotechnol. 2016;180:872–882. doi: 10.1007/s12010-016-2139-z. [DOI] [PubMed] [Google Scholar]
- 11.Sierra-Garcia IN, Dellagnezze BM, Santos VP, Chaves B MR, Capilla R, et al. Microbial diversity in degraded and non-degraded petroleum samples and comparison across oil reservoirs at local and global scales. Extremophiles. 2017;21:211–229. doi: 10.1007/s00792-016-0897-8. [DOI] [PubMed] [Google Scholar]
- 12.Li D, Midgley DJ, Ross JP, Oytam Y, Abell GCJ, et al. Microbial biodiversity in a Malaysian oil field and a systematic comparison with oil reservoirs worldwide. Arch Microbiol. 2012;194:513–523. doi: 10.1007/s00203-012-0788-z. [DOI] [PubMed] [Google Scholar]
- 13.Gao P, Tian H, Li G, Sun H, Ma T. Microbial diversity and abundance in the Xinjiang Luliang long-term water-flooding petroleum reservoir. Microbiologyopen. 2015;4:332–342. doi: 10.1002/mbo3.241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bae SS, Jung J, Chung D, Baek K. Marinobacterium aestuarii sp. nov., a benzene-degrading marine bacterium isolated from estuary sediment. Int J Syst Evol Microbiol. 2018;68:651–656. doi: 10.1099/ijsem.0.002561. [DOI] [PubMed] [Google Scholar]
- 15.Ul Karim MdR, Fukaya K, In Y, Sharma AR, Harunari E, et al. Marinoquinolones and marinobactoic acid: antimicrobial and cytotoxic ortho -dialkylbenzene-class metabolites produced by a marine obligate gammaproteobacterium of the genus Marinobacterium . J Nat Prod. 2022;85:1763–1770. doi: 10.1021/acs.jnatprod.2c00281. [DOI] [PubMed] [Google Scholar]
- 16.Wang M, Xi W, Li Z. Analysis of the genome sequencing data of the Marinobacterium genus. Chin J Biotechnol. 2020;36:2695–2706. doi: 10.13345/j.cjb.200366. [DOI] [PubMed] [Google Scholar]
- 17.Wang MR, Li HF, Yi JJ, Tao SY, Li ZJ. Production of polyhydroxyalkanoates by three novel species of Marinobacterium . Int J Biol Macromol. 2022;195:255–263. doi: 10.1016/j.ijbiomac.2021.12.019. [DOI] [PubMed] [Google Scholar]
- 18.Ray S, Kalia VC. Biomedical applications of polyhydroxyalkanoates. Indian J Microbiol. 2017;57:261–269. doi: 10.1007/s12088-017-0651-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Robertson LA, Kuenen JG. In: The Prokaryotes: Volume 5: Proteobacteria: Alpha and Beta Subclasses. Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E, editors. New York, NY: Springer New York; 2006. The genus Thiobacillus ; pp. 812–827. [Google Scholar]
- 20.Kuenen JG, Veldkamp H. Thiomicrospira pelophila, gen. n., sp. n., a new obligately chemolithotrophic colourless sulfur bacterium. Antonie van Leeuwenhoek. 1972;38:241–256. doi: 10.1007/BF02328096. [DOI] [PubMed] [Google Scholar]
- 21.Brock TD. In: Bergey’s Manual of Determinative Bacteriology. 8th. Buchanan RE, Gibbons NE, editors. Baltimore, MD, USA: The Williams & Wilkins Co; 1974. Genus Thiothrix ; p. 119. edn. p. [Google Scholar]
- 22.Weisburg WG, Barns SM, Pelletier DA, Lane DJ. 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol. 1991;173:697–703. doi: 10.1128/jb.173.2.697-703.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lane DJ. In: Nucleic Acid Techniques in Bacterial Systematics. Stackebrandt E, Goodfellow M, editors. New York,NY: Wiley; 1991. 16S/23S rRNA sequencing; pp. 115–175. [Google Scholar]
- 24.Wang XM, Wang XT, Wang XQ, Mu DS, Du ZJ. Algoriphagus lacus sp. nov., isolated from a freshwater lake. Int J Syst Evol Microbiol. 2020;70:193–198. doi: 10.1099/ijsem.0.003734. [DOI] [PubMed] [Google Scholar]
- 25.Yoon S-H, Ha S-M, Kwon S, Lim J, Kim Y, et al. Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol. 2017;67:1613–1617. doi: 10.1099/ijsem.0.001755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Fitch WM. Toward defining the course of evolution: minimum change for a specific tree topology. Syst Biol. 1971;20:406–416. doi: 10.1093/sysbio/20.4.406. [DOI] [Google Scholar]
- 27.Tamura K, Stecher G, Kumar S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol Evol. 2021;38:3022–3027. doi: 10.1093/molbev/msab120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chaumeil P-A, Mussig AJ, Hugenholtz P, Parks DH. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics. 2019;36:1925–1927. doi: 10.1093/bioinformatics/btz848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Trifinopoulos J, Nguyen L-T, von Haeseler A, Minh BQ. W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016;44:W232–5. doi: 10.1093/nar/gkw256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Parks DH, Chuvochina M, Rinke C, Mussig AJ, Chaumeil P-A, et al. GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 2022;50:D785–D794. doi: 10.1093/nar/gkab776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i890. doi: 10.1093/bioinformatics/bty560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Li R, Yu C, Li Y, Lam T-W, Yiu S-M, et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics. 2009;25:1966–1967. doi: 10.1093/bioinformatics/btp336. [DOI] [PubMed] [Google Scholar]
- 33.Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015;25:1043–1055. doi: 10.1101/gr.186072.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yoon SH, Ha SM, Lim J, Kwon S, Chun J. A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie van Leeuwenhoek. 2017;110:1281–1286. doi: 10.1007/s10482-017-0844-4. [DOI] [PubMed] [Google Scholar]
- 35.Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics. 2013;14:60. doi: 10.1186/1471-2105-14-60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016;44:D457–D462. doi: 10.1093/nar/gkv1070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Grant JR, Enns E, Marinier E, Mandal A, Herman EK, et al. Proksee: in-depth characterization and visualization of bacterial genomes. Nucleic Acids Res. 2023;51:W484–W492. doi: 10.1093/nar/gkad326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics. 2008;9:75. doi: 10.1186/1471-2164-9-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Alcock BP, Raphenya AR, Lau TTY, Tsang KK, Bouchard M, et al. CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 2020;48:D517–D525. doi: 10.1093/nar/gkz935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Huerta-Cepas J, Forslund K, Coelho LP, Szklarczyk D, Jensen LJ, et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. Mol Biol Evol. 2017;34:2115–2122. doi: 10.1093/molbev/msx148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Blin K, Shaw S, Kloosterman AM, Charlop-Powers Z, van Wezel GP, et al. antiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res. 2021;49:W29–W35. doi: 10.1093/nar/gkab335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Zhang H, Yohe T, Huang L, Entwistle S, Wu P, et al. dbCAN2: a meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 2018;46:W95–W101. doi: 10.1093/nar/gky418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Chaudhari NM, Gupta VK, Dutta C. BPGA- an ultra-fast pan-genome analysis pipeline. Sci Rep. 2016;6:24373. doi: 10.1038/srep24373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Smibert RM, Krieg NR. In: Methods for General and Molecular. Gerhardt P, Murray RGE, Wood WA, Krieg NR, editors. Bacteriology, DC: American Society for Microbiology; 1994. Phenotypic characterization; pp. 607–654. [Google Scholar]
- 45.Sarwar G, Oda H, Sakata T, Kakimoto D. Potentiality of artificial sea water salts for the production of carrageenase by a marine Cytophaga sp. Microbiol Immunol. 1985;29:405–411. doi: 10.1111/j.1348-0421.1985.tb00841.x. [DOI] [PubMed] [Google Scholar]
- 46.Du ZJ, Wang Y, Dunlap C, Rooney AP, Chen GJ. Draconibacterium orientale gen. nov., sp. nov., isolated from two distinct marine environments, and proposal of Draconibacteriaceae fam. nov. Int J Syst Evol Microbiol. 2014;64:1690–1696. doi: 10.1099/ijs.0.056812-0. [DOI] [PubMed] [Google Scholar]
- 47.Zhu KL, Wang XQ, Zhang TS, Shang DD, Du ZJ. Salibaculum halophilum gen. nov., sp. nov. and Salibaculum griseiflavum sp. nov., in the family Rhodobacteraceae . Int J Syst Evol Microbiol. 2021;71 doi: 10.1099/ijsem.0.004808. [DOI] [PubMed] [Google Scholar]
- 48.Kroppenstedt RM. Separation of bacterial menaquinones by HPLC using reverse Phase (RP18) and a silver loaded ion exchanger as stationary phases. J Liq Chromatogr. 1982;5:2359–2367. doi: 10.1080/01483918208067640. [DOI] [Google Scholar]
- 49.Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, et al. An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods. 1984;2:233–241. doi: 10.1016/0167-7012(84)90018-6. [DOI] [Google Scholar]
- 50.Athalye M, Noble WC, Minnikin DE. Analysis of cellular fatty acids by gas chromatography as a tool in the identification of medically important coryneform bacteria. J Appl Bacteriol. 1985;58:507–512. doi: 10.1111/j.1365-2672.1985.tb01491.x. [DOI] [PubMed] [Google Scholar]
- 51.Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, et al. Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol. 2014;12:635–645. doi: 10.1038/nrmicro3330. [DOI] [PubMed] [Google Scholar]
- 52.Richter M, Rosselló-Móra R. Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci U S A. 2009;106:19126–19131. doi: 10.1073/pnas.0906412106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, et al. DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol. 2007;57:81–91. doi: 10.1099/ijs.0.64483-0. [DOI] [PubMed] [Google Scholar]
- 54.Konstantinidis KT, Rosselló-Móra R, Amann R. Uncultivated microbes in need of their own taxonomy. ISME J. 2017;11:2399–2406. doi: 10.1038/ismej.2017.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, et al. Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol. 2014;12:635–645. doi: 10.1038/nrmicro3330. [DOI] [PubMed] [Google Scholar]
- 56.Rhie MN, Yoon HE, Oh HY, Zedler S, Unden G, et al. A Na+-coupled C4-dicarboxylate transporter (Asuc_0304) and aerobic growth of Actinobacillus succinogenes on C4-dicarboxylates. Microbiology. 2014;160:1533–1544. doi: 10.1099/mic.0.076786-0. [DOI] [PubMed] [Google Scholar]
- 57.Locher KP. Mechanistic diversity in ATP-binding cassette (ABC) transporters. Nat Struct Mol Biol. 2016;23:487–493. doi: 10.1038/nsmb.3216. [DOI] [PubMed] [Google Scholar]
- 58.Galloway JN. The global nitrogen cycle: past, present and future. Sci China C Life Sci. 2005;48 Suppl 2:669–678. doi: 10.1007/BF03187108. [DOI] [PubMed] [Google Scholar]
- 59.Chen G-Q, Jiang X-R. Next generation industrial biotechnology based on extremophilic bacteria. Curr Opin Biotechnol. 2018;50:94–100. doi: 10.1016/j.copbio.2017.11.016. [DOI] [PubMed] [Google Scholar]
- 60.Anderson AJ, Dawes EA. Occurrence, metabolism, metabolic role, and industrial uses of bacterial polyhydroxyalkanoates. Microbiol Rev. 1990;54:450–472. doi: 10.1128/mr.54.4.450-472.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Dawes EA, Senior PJ. The role and regulation of energy reserve polymers in micro-organisms. Adv Microb Physiol. 1973;10:135–266. doi: 10.1016/s0065-2911(08)60088-0. [DOI] [PubMed] [Google Scholar]
- 62.Khan MA, Sahile AA, Jan R, Asaf S, Hamayun M, et al. Halotolerant bacteria mitigate the effects of salinity stress on soybean growth by regulating secondary metabolites and molecular responses. BMC Plant Biol. 2021;21:176. doi: 10.1186/s12870-021-02937-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Brown CL, Mullet J, Hindi F, Stoll JE, Gupta S, et al. mobileOG-db: a manually curated database of protein families mediating the life cycle of bacterial mobile genetic elements. Appl Environ Microbiol. 2022;88:e0099122. doi: 10.1128/aem.00991-22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Rankin DJ, Rocha EPC, Brown SP. What traits are carried on mobile genetic elements, and why? Heredity. 2011;106:1–10. doi: 10.1038/hdy.2010.24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Davies J. Origins and evolution of antibiotic resistance. Microbiologia. 1996;12:9–16. [PubMed] [Google Scholar]
- 66.Magiorakos A-P, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012;18:268–281. doi: 10.1111/j.1469-0691.2011.03570.x. [DOI] [PubMed] [Google Scholar]
- 67.Wadhwa N, Berg HC. Bacterial motility: machinery and mechanisms. Nat Rev Microbiol. 2022;20:161–173. doi: 10.1038/s41579-021-00626-4. [DOI] [PubMed] [Google Scholar]
- 68.Raina J-B, Fernandez V, Lambert B, Stocker R, Seymour JR. The role of microbial motility and chemotaxis in symbiosis. Nat Rev Microbiol. 2019;17:284–294. doi: 10.1038/s41579-019-0182-9. [DOI] [PubMed] [Google Scholar]
- 69.Morimoto YV, Minamino T. Structure and function of the bi-directional bacterial flagellar motor. Biomolecules. 2014;4:217–234. doi: 10.3390/biom4010217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Aldridge P, Hughes KT. Regulation of flagellar assembly. Curr Opin Microbiol. 2002;5:160–165. doi: 10.1016/s1369-5274(02)00302-8. [DOI] [PubMed] [Google Scholar]
- 71.Keegstra JM, Carrara F, Stocker R. The ecological roles of bacterial chemotaxis. Nat Rev Microbiol. 2022;20:491–504. doi: 10.1038/s41579-022-00709-w. [DOI] [PubMed] [Google Scholar]
- 72.Whitman WB. Bergey’s Manual of Systematics of Archaea and Bacteria. 2021. pp. 1–20. [DOI] [Google Scholar]
- 73.Thomson NM, Ferreira JL, Matthews-Palmer TR, Beeby M, Pallen MJ, et al. Giant flagellins form thick flagellar filaments in two species of marine γ-proteobacteria. PLoS One. 2018;13:e0206544. doi: 10.1371/journal.pone.0206544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Baumann P, Bowditch RD, Baumann L, Beaman B. Taxonomy of marine Pseudomonas species: P. stanieri sp. nov.; P. perfectomarina sp. nov., nom. rev.; P. nautica: and P. doudoroffii . Int J Syst Bacteriol. 1983;33:857–865. doi: 10.1099/00207713-33-4-857. [DOI] [Google Scholar]
- 75.Kim S-J, Park S-J, Yoon D-N, Park B-J, Choi B-R, et al. Marinobacterium maritimum sp. nov., a marine bacterium isolated from Arctic sediment. Int J Syst Evol Microbiol. 2009;59:3030–3034. doi: 10.1099/ijs.0.009134-0. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.