Summary
Tissue-resident macrophages (TRMs) are abundant immune cells within pre-metastatic sites, yet their functional contributions to metastasis remain incompletely understood. Here, we show that alveolar macrophages (AMs), the main TRMs of the lung, are susceptible to downregulation of the immune stimulatory transcription factor IRF8, impairing anti-metastatic activity in models of metastatic breast cancer. G-CSF is a key tumor-associated factor (TAF) that acts upon AMs to reduce IRF8 levels and facilitate metastasis. Translational relevance of IRF8 downregulation was observed among macrophage precursors in breast cancer and a CD68hiIRF8loG-CSFhi gene signature suggests poorer prognosis in triple-negative breast cancer (TNBC), a G-CSF-expressing subtype. Our data highlight the underappreciated, pro-metastatic roles of AMs in response to G-CSF and identify the contribution of IRF8-deficient AMs to metastatic burden. AMs are an attractive target of local neoadjuvant G-CSF blockade to recover anti-metastatic activity.
Subject areas: Microenvironment, Immunology, Cancer
Graphical abstract

Highlights
-
•
IRF8 declines in distal macrophages and monocytes during early mammary tumor growth
-
•
Loss of IRF8 in AMs serves as a pre-conditioning mechanism for lung metastasis
-
•
G-CSF reduces AM IRF8 levels and is a therapeutic target for anti-metastatic activity
-
•
A CD68hiIRF8loG-CSFhi gene signature in TNBC patients portends poorer prognosis
Microenvironment; Immunology; Cancer
Introduction
Metastasis remains a major challenge in the clinic and unfortunately, is the main cause of cancer-related deaths due to a lack of effective and durable treatments that prevent or inhibit tumor progression.1,2 Distal, potential metastatic microenvironments are ‘conditioned’ during primary tumor growth to enable tumor colonization and proliferation.3 Resident or recruited stromal populations within distal sites are often exploited by tumor-derived cues to sustain metastatic outgrowth.4,5 Multiple types of myeloid cells, including granulocytes, myeloid-derived suppressor cells (MDSCs), macrophages, and monocytes comprise the stroma, enhancing metastasis and immunosuppression.6 Understanding the molecular basis underlying the functional transition of these stromal elements toward pro-metastatic roles is necessary to uncover newer therapeutic targets to prevent or eliminate metastasis.
Macrophages are prominent stromal components comprised of tissue-resident and recruited populations that significantly contribute to cancer outcome.7,8,9 The macrophage response in cancer is complex as macrophages arise from different sources, including embryonic tissue and bone marrow, and may exert a continuum of functional responses, ranging from anti-metastatic “defenders” to pro-metastatic “remodelers”.10 The functional roles of macrophages recruited from bone marrow have been extensively investigated in cancer biology,11,12,13,14,15,16,17 overshadowing the roles of tissue-resident macrophages (TRMs) that inhabit pre-cancerous tissue. Few reports have shed light on TRM contributions in supporting primary tumor growth18,19,20,21,22 and metastasis23,24 or conversely, eliminating metastatic tumor spread.25,26 It is thought that tumor-associated factors (TAFs), which may be tumor- or stroma-derived, alter macrophage biology to dampen anti-metastatic activity and aid metastatic outgrowth. The identity and breadth of TAFs that initiate molecular cues to dysregulate TRM function remain to be fully understood.
Granulocyte-colony stimulating factor (G-CSF) is a prominent TAF of interest due to its tumor-intrinsic growth effects27,28 and pro-metastatic roles,29,30,31 which are largely mediated by granulocytes or polymorphonuclear (PMN)-MDSCs. The effects of G-CSF on other myeloid cells are overlooked, especially TRMs that may be affected by G-CSF prior to granulocyte or PMN-MDSC arrival. Few reports have assessed the functional consequences of G-CSF on macrophage biology and suggest enhanced immunosuppressive properties.32,33
Downstream of TAFs, the tightly controlled coordination of activated transcriptional networks direct the diversity and plasticity of macrophage effector functions.34,35 Transcriptional regulators that promote gene expression for macrophage-mediated tumoricidal activities and induction of adaptive immunity are instrumental for robust anti-metastatic responses.36 Interferon regulatory factor-8 (IRF8) is one such positive determinant that upregulates genes crucial for pathogen recognition, anti-microbial host defense, antigen processing, and immune activation during microbial pathogenesis.37,38,39 Our laboratory recently identified the importance of macrophage IRF8 expression for improved clinical outcomes in kidney cancer40 and reduced lung metastasis in preclinical models of mammary cancer,41 tumor types characterized by heavy macrophage infiltration. However, several important questions remained unanswered, namely: 1) the dynamics of IRF8 downregulation in macrophages at distal sites during tumor progression; 2) the relevant macrophage population(s) reliant on IRF8 for anti-metastatic activity; and 3) the signals downregulating macrophage IRF8 expression. As TRMs inhabit the front line of tissue defense,8,42 we tested the hypothesis that anti-metastatic activity by TRMs is IRF8-dependent. We further hypothesized that G-CSF, a negative regulator of IRF8 expression in other myeloid systems,43,44 impairs anti-metastatic activity of TRMs by downregulating IRF8.
Lung tissue is a common and lethal metastatic site of solid tumors, including triple-negative breast cancer (TNBC) and other breast cancer subtypes.45 Therefore, we utilized mammary tumor models of spontaneous and experimental lung metastasis to focus on alveolar macrophages (AMs), the main TRM population of the lung.42,46 We show that AMs are susceptible to IRF8 downregulation during early mammary tumor growth and prior to the formation of detectable micro-metastasis. Utilizing a conditional model of IRF8 deletion in AMs, we demonstrate that IRF8-deficient AMs enhance the frequency and size of metastatic nodules. We observed that G-CSF is a TAF that acts within the lung to reduce IRF8 levels within AMs and facilitate metastasis. Importantly, local G-CSF blockade recovers AM-mediated anti-metastatic activity in an IRF8-dependent manner. Lastly, a CD68hiIRF8loG-CSFhi gene signature in TNBC patients infers prognostic significance, demonstrating translational relevance of IRF8 downregulation in G-CSF-expressing cancer types.
Results
Decline of IRF8 levels in macrophages and monocytes throughout the periphery precedes micro-metastasis in the lung
To determine the dynamics of IRF8 expression in TRMs and recruited macrophages during early primary tumor growth in relation to endpoint micro-metastasis within lung tissue, we used orthotopic models of metastatic breast cancer, including syngeneic implantable, autochthonous and mouse-human xenograft (Figures S1A and S1B). First, we utilized the 4T1 model which resembles TNBC and spontaneously metastasizes to the lung.47 Three groups of BALB/c female wild-type (WT) mice bearing orthotopic 4T1 mammary tumors with distinct surface volumes [1) <50 mm3; 2) 50-100 mm3; and 3) 100-300mm3] were assessed in comparison to healthy tissue from non-tumor-bearing (NTB) mice. Lung tissue was dissociated, cultured in vitro for 1–3 weeks and stained for metastatic tumor colony formation (CF). Within one week of in vitro growth, numerous 4T1 colonies were detected from lungs of mice bearing >100mm3-sized 4T1 tumors, which aligned with enlarged tumor and spleen weights as splenomegaly is a well-known pathology of 4T1 tumor progression (Figures 1A and 1B). In contrast, mice bearing <100mm3-sized 4T1 tumors had no detectable colonies after 3 weeks of monitoring. Micro-metastasis was tissue-specific to the lung at these observed early time points of primary tumor growth and not detected within the brain, a later metastatic site.
Figure 1.
IRF8 expression declines among alveolar macrophages (AMs), recruited macrophages and monocytes prior to 4T1 micro-metastasis in lung tissue
(A) Mammary tumor, lung and spleen weights of BALB/c orthotopic 4T1-bearing WT mice compared to non-tumor-bearing (NTB) control tissue.
(B) 4T1 micro-metastasis detection in lung tissue by staining of colony formation (CF). Scale bar: 370 μm.
(C) Myeloid cell percentages within lung tissue or blood during 4T1 growth.
(D) Myeloid intracellular IRF8 levels within lung tissue of NTB or 4T1-bearing mice.
(E) Flow-sorted AMs have reduced Irf8 and target gene expression during 4T1 growth.
(F) IRF8 levels as in (D) within blood. Data are represented as mean ± SEM. Significance was determined by Wilcoxon rank-sum tests with Holm-Bonferroni correction for pre-planned comparisons (A and B), Dunnett’s test for correction of comparisons to NTB control (C; D and F, left), Spearman correlation (D and F, right; line indicates simple linear regression), or Mann-Whitney (E; n ≥ 5 mice/group). ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001; ∗∗∗∗p < 0.0001. See also Figures S1–S3.
Lung tissue and blood were collected to analyze intracellular IRF8 levels of macrophages, monocytes or granulocytes by flow cytometry (Figure S2). Specificity of IRF8 staining was demonstrated with the Irf8−/− model and IRF8 expression was calculated as log2 transformation48 based on normalization of the anti-IRF8 antibody median fluorescence intensity (MFI) to the isotype control MFI for each tissue per mouse. AMs (CD11chiSiglecFhiCD11bint-loLy6G−F4/80+) are the largest TRM population that reside among alveoli, outnumbering interstitial macrophages (IMs; CD11bhiMHCIIhiCD11cloSiglecFloLy6CloLy6G−F4/80+), a minor TRM population that resides within the interstitial space, by 10-fold (Figure 1C). CD11bhi macrophages (CD11bhiCD11cloSiglecFloLy6CloLy6G−F4/80+) and monocytes (CD11bhiLy6ChiLy6G−F4/80-) recruited to lung tissue from the bone marrow are also 10-fold less abundant than AMs. As primary tumors enlarged beyond 50 mm3, IRF8 was significantly reduced in AMs, CD11bhi macrophages and monocytes (Figure 1D). An inverse correlation of IRF8 levels was also observed with increased tumor weight. In contrast, IMs maintained high IRF8 levels throughout early 4T1 growth similar to the NTB controls. AMs were flow-sorted from lung tissue of NTB or 4T1-bearing mice (average < 150 mm3) to further assess the impact of IRF8 reduction on downstream targets. Decreased expression of Irf8, target genes Il-12β and Nos237 and the homeostatic lung factor Tgf-β1 were observed, while the AM-associated factor Car442 remained unchanged (Figure 1E). Select chemokine and growth factor expression including Cxcl9 and Vefg-α trended upward, highlighting differential gene expression in AMs during early 4T1 growth.
At the earliest size of 4T1 tumor growth (<50 mm3), IRF8 levels were significantly reduced in monocytes within the blood (Figure 1F), recognizing that the decline in IRF8 expression was not exclusive to macrophages within lung tissue and precedes detection of micro-metastasis. While granulocytes (CD11bhiLy6CloLy6G+F4/80-) were readily detected in lung tissue and blood (Figure 1C), IRF8 expression remained low as expected within granulocyte biology44 and unchanged throughout early 4T1 progression (Figures 1D and 1F). Similarly, basal IRF8 levels were also low within a CD11bhiCD11cloSiglecFhiLy6CloLy6G−F4/80+ population of lung tissue, remained unchanged during tumor growth and not a further focus of this study. Overall, our results demonstrate that select distal TRMs (AMs; 4T1>50 mm3), recruited macrophages (4T1>50 mm3) and monocytes (4T1<50 mm3) within the periphery are susceptible to IRF8 loss prior to detectable 4T1 lung micro-metastasis (4T1>100 mm3), suggesting that IRF8 loss in these myeloid populations carried broad biologic relevance.
Next, we assessed endpoint micro-metastasis and the dynamics of IRF8 expression in AMs, recruited macrophages and monocytes within additional spontaneous metastatic mammary tumor models. C57BL/6 female transgenic mice expressing the polyomavirus middle T antigen under the control of the mouse mammary tumor virus (MMTV-PyMT; abbreviated as MTAG) develop multi-focal tumors with lung metastasis evident within 6 months of age.49 We observed early tumor growth between two groups of female MTAG mice within 4 months of age and bearing ≤3 palpable primary tumors [1) each individual tumor volume <150 mm3; 2) at least one primary tumor volume >150mm3]. Lung micro-metastasis was undetectable within either group as total tumor surface volume and weight enlarged (Figures 2A and 2B). While AMs outnumbered the other lung macrophage populations by 10 to 20-fold, IRF8 levels declined among AMs, IMs and CD11bhi macrophages (Figures 2C and 2D). Monocytes collected from the lung or blood also had reduced IRF8 expression, emphasizing systemic effects. Myeloid IRF8 downregulation thus also precedes micro-metastasis in the autochthonous MTAG model. In contrast, in female WT mice bearing syngeneic implantable EMT6 (BALB/c)50 or E0771.ML-1 (C57BL/6)51 orthotopic tumors (average < 150 mm3), lung micro-metastasis occurred while IRF8 expression remained intact in macrophages or monocytes of lung tissue and blood (Figure S3). Our results indicate that IRF8 downregulation in macrophages and monocytes occurs in some, but not all preclinical mammary tumor models.
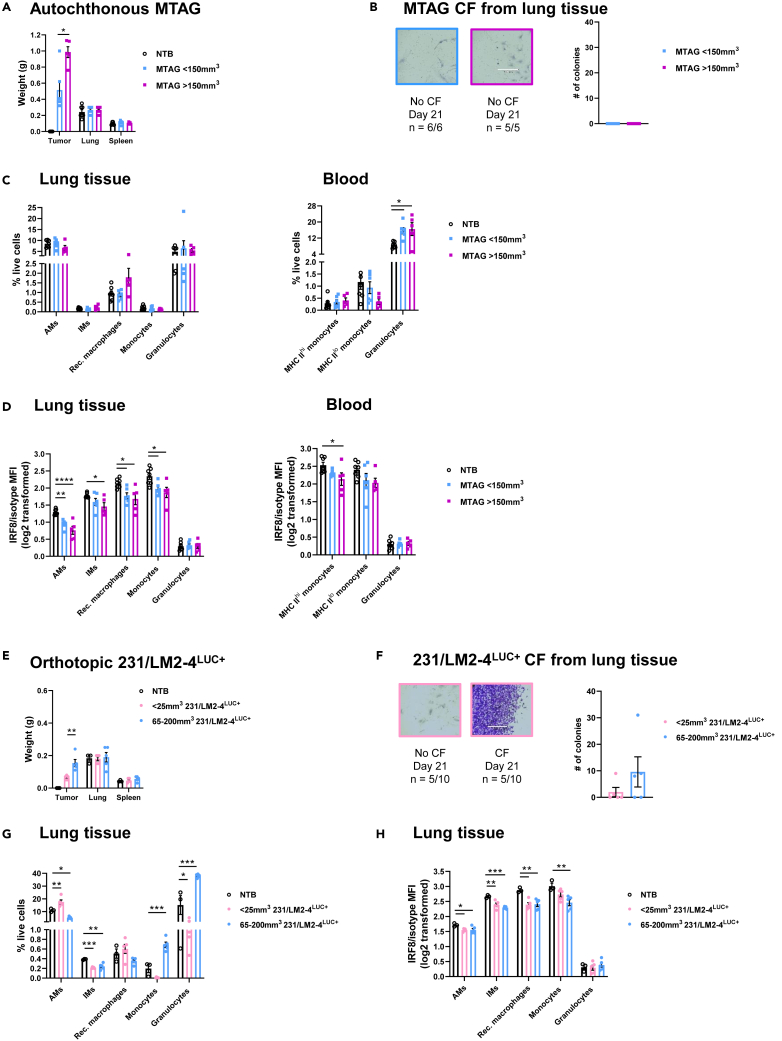
Figure 2.
Macrophage IRF8 downregulation also occurs in the autochthonous MTAG model and within a mouse-human xenograft model
(A) Multifocal mouse mammary tumor virus-polyomavirus middle T antigen (MTAG) tumors, lung and spleen weights compared to NTB tissue of C57BL/6 WT mice.
(B) Lung tissue of MTAG mice lack micro-metastasis. Scale bar: 370 μm.
(C and D) Myeloid cell percentages (C) and IRF8 levels (D) within lung tissue or blood during MTAG growth.
(E) Mammary tumor, lung and spleen weights of BALB/c orthotopic 231/LM2-4LUC+-bearing SCID mice compared to NTB tissue.
(F) Detection of 231/LM2-4LUC+ micro-metastasis in lung tissue. Scale bar: 370 μm.
(G and H) Myeloid cell percentages (G) and IRF8 levels (H) within lung tissue during 231/LM2-4LUC+ tumor growth. Data are represented as mean ± SEM. Significance was determined by Wilcoxon rank-sum tests with Holm-Bonferroni correction for pre-planned comparisons (A and E), Mann-Whitney (B and F) or Dunnett’s test for correction of comparisons to NTB control (C, D, G, and H; n ≥ 3 mice/group). ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001; ∗∗∗∗p < 0.0001.
IRF8 dynamics were then evaluated within a mouse-human xenograft tumor setting, enabling an assessment of human tumors on alteration of myeloid IRF8 expression in the absence of adaptive immunity. A lung metastatic variant of MDA-MB-231 (termed 231/LM2-4LUC+)52 was orthotopically implanted into BALB/c female SCID mice. Within one week of tumor growth (average < 25 mm3), spontaneous lung micro-metastasis occurred as primary tumors enlarged in surface volume and weight (Figures 2E and 2F). AMs initially expanded in cell percentage and remained 10 to 20-fold more abundant than other lung macrophage populations (Figure 2G). IRF8 quickly declined within AMs, IMs, CD11bhi macrophages, and monocytes in lung tissue (Figure 2H). Overall, our data suggest that myeloid-intrinsic IRF8 downregulation serves as a pre-conditioning mechanism to facilitate micro-metastasis within lung tissue in several but not all tumor settings.
AM-mediated metastatic tumor growth is enhanced by IRF8-deficiency
Since we observed a significant reduction in IRF8 levels of AMs, the largest macrophage population observed prior to micro-metastasis, we evaluated the role of IRF8 for anti-metastatic activity using a conditional knockout (cKO) model. This genetic approach enabled functional assessment of IRF8-deficiency, bypassing TAF-mediated mechanisms of IRF8 downregulation. The IRF8 cKO model driven by the lysozyme M-Cre recombinase system (Lyz2CreIRF8fl/fl; Figure S1C) does not alter myeloid development41 nor abundance of TRMs within multiple tissues, including the lung (Figure S4A). IRF8 cKO AMs have significantly reduced IRF8 expression in both BALB/c and C57BL/6 strains (Figure S4B). As described elsewhere,53,54 we confirmed that AMs highly express lysozyme compared to other macrophages and monocytes among lung tissue or peripheral sites (Figure S4C). Granulocytes highly express lysozyme but express low levels of IRF8 and thus were not likely targeted within this genetic model. Our results thus provide an explanation of nearly complete deletion of IRF8 within AMs. To assess the impact of IRF8-deficient AMs on spontaneous endpoint micro-metastasis, 4T1 tumors were orthotopically implanted into BALB/c female IRF8fl/fl or IRF8 cKO mice. Significantly more 4T1 colonies were visible from the lungs of 4T1-bearing IRF8 cKO mice compared to IRF8fl/fl mice with similar-sized primary tumors (average < 150 mm3; Figures 3A, and S4D). Interestingly, by bypassing potential upstream TAFs with direct genetic deletion of IRF8, spontaneous lung micro-metastasis occurred more proficiently in BALB/c EMT6-bearing IRF8 cKO mice (average < 200 mm3; Figures S4E and S4F).
Figure 3.
IRF8 expression in AMs promotes anti-metastatic activity
(A) Greater spontaneous 4T1 metastatic tumor CF occurs in BALB/c IRF8 cKO (Lyz2CreIRF8fl/fl) lung tissue than IRF8fl/fl lung tissue.
(B) IRF8 cKO hosts have greater 4T1 experimental lung metastasis and larger nodule sizes than IRF8fl/fl controls.
(C) Schematic of control or clodronate-encapsulated liposome treatment before and during 4T1 experimental metastasis.
(D and E) AM depletion via intranasal (i.n.) clodronate treatment reduces 4T1 experimental metastatic burden in IRF8 cKO hosts (D). Depletion of systemic macrophages, but not AMs, by intraperitoneal (i.p.) clodronate treatment increases 4T1 experimental metastasis in IRF8 cKO hosts (E). Representative H&E staining and images of lung metastasis displayed on the left. Scale bar: 370 μm. All data are represented as mean ± SEM. Significance was determined by Mann-Whitney (A and B) or Wilcoxon rank-sum tests with Holm-Bonferroni correction for pre-planned comparisons (D and E; n ≥ 5 mice/group). ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001. See also Figures S1–S6.
To focus our studies on post-extravasation mechanisms within the lung, a prominent end-stage metastatic site, we utilized models of experimental metastasis (Figure S1D) including 4T1 (BALB/c) and parental E0771 (C57BL/6).16 This approach enabled an assessment of the AM-tumor interaction without the confounding influence of the primary tumor.55,56 Mice with IRF8-deficient AMs have significantly more 4T1 or E0771 metastatic foci within 3 weeks compared to IFR8fl/fl controls (Figures 3B and S4G–S4I). Large metastatic nodule sizes, including >3 mm in the 4T1 model, were more prominent in IRF8 cKO hosts. IRF8-deficiency thus contributes to greater lung metastatic nodule abundance and growth regardless of TAF expression.
To demonstrate that AMs are a key TRM population that influence metastatic outcome, we utilized two routes of pharmacological macrophage depletion: 1) intranasal; and 2) intraperitoneal. The effects of continuous macrophage depletion on post-extravasation metastatic mechanisms were assessed by experimental metastasis as a less toxic approach than settings of spontaneous metastasis. First, compared to control liposome treatment, intranasal administration of clodronate-encapsulated liposomes reduced AMs 8-fold and other minor lung macrophages 2-fold, but not splenic macrophages, demonstrating specificity of local drug administration (Figure S5A). Pre-depletion prior to 4T1 injection and twice weekly clodronate administration (Figure 3C) was utilized for rigorous depletion throughout 4T1 experimental metastasis in comparison to control liposome treatment.57 Prolonged intranasal clodronate treatment induced lung enlargement (Figure S5B) and sterile pneumonia characteristic of inflammation,42 debris and granulocyte infiltration within alveoli, while control liposome treatment only induced lung debris (Figure 3D, left). Despite such lung pathology, intranasal clodronate treatment similarly reduced overall metastatic formation within IRF8-proficient and -deficient hosts (Figure 3D, right). IRF8 cKO mice also had less large-sized nodules in response to local targeting of AMs for depletion. These data demonstrate that although AMs serve important roles for maintaining lung tissue homeostasis and preventing debris accumulation, AMs also exert pro-metastatic activities. To dissect a role of AMs on metastatic seeding, a single treatment of intranasal clodronate or control liposomes were administered prior to 4T1 experimental metastasis (Figure S5C). Single, pre-depletion of AMs did not alter lung pathology or overall metastatic burden (Figures S5D and S4E). Pre-depletion of AMs in IRF8 cKO hosts may hinder colonization proficiency, as observed by a significant decrease in small-sized nodules. However, an abundance of large-sized nodules within IRF8-deficient hosts, regardless of pre-depletion, supports pro-metastatic activity of AMs beyond colonization.
Continuous, intraperitoneal administration of clodronate was then selected to target IMs and CD11bhi recruited macrophages for depletion (Figure S5F) since such macrophages are replenished by bone marrow precursors.58 Murine AMs do not heavily rely on bone marrow precursors for self-renewal59 and remain intact during intraperitoneal clodronate treatment. In contrast to intranasal administration, intraperitoneal clodronate treatment was toxic to a proportion of mice arising from systemic depletion of multiple macrophage populations (Figure S5F). Although lung weight was unaltered (Figure S5G) and alveoli structures were not as obstructed by intraperitoneal control or clodronate liposome treatment (Figure 3E, left), repeated intraperitoneal clodronate administration induced numerous lung metastatic nodules in mice evaluable at endpoint. IRF8 cKO hosts had significantly more metastasis and enlarged nodules (Figure 3E, right). Granulocyte infiltration within lung tissue resulted from intranasal or intraperitoneal clodronate administration (Figures S5A and S4F). The opposing experimental metastatic outcomes in response to differing clodronate administration routes are therefore most likely due to changes in macrophage populations rather than lung neutrophilia. We tested this by depleting granulocytes with intraperitoneal administration of anti-Ly6G in comparison to isotype controls. Experimental metastasis increased in IRF8-proficient hosts while metastatic burden remained higher in IRF8-deficient hosts regardless of granulocyte depletion (Figures S6A–S6D).
To validate these findings that the intact AM population enhances experimental metastatic outcome, we utilized intraperitoneal administration of a CCR2 inhibitor (CCR2i) as an additional approach by titrating the CCR2i dose to deplete CD11bhi recruited macrophages in lung tissue and reduce IMs (Figure S6E). Other peripheral macrophages and monocytes remained intact, reflective of less toxic treatment. Granulocyte infiltration within lungs was not as large as induced by intraperitoneal clodronate, enabling an assessment of AMs on metastatic burden. Numerous metastatic nodules grew in response to continuous CCR2i treatment compared to vehicle control, with larger nodules observed in IRF8 cKO hosts (Figures S6F–S6H). These results are consistent with reports of increased tumor load within lung tissue during CCR2-deficiency.20,55 Collectively, our results demonstrate that IRF8-deficient AMs promote metastatic growth post-extravasation in the absence of recruited macrophages.
An in vitro source of AMs augment tumor growth in absence of IRF8
Our results thus far indicate that IRF8-deficient AMs expand metastatic outgrowth. To further study the AM response mechanistically in vitro, we developed a neonate liver-derived AM (NLDAM) culture system, reflective of the embryonic origin of AMs.60 NLDAMs display a surface phenotype paralleling in vivo AMs, highly express lysozyme and characteristic AM gene expression including Cd206 and Car4 (Figures S7A–S7C). This expression profile is unique to NLDAMs as traditional bone marrow-derived macrophages (BMDMs) express less lysozyme and differential macrophage effector genes. The BALB/c IRF8 cKO model was applied to investigate the effects of IRF8 cKO NLDAMs, which have significantly reduced IRF8 mRNA and protein levels (Figures 4A and S7D). IRF8 targets Il-12β and Nos2, which elicit anti-metastatic properties, were significantly downregulated in IRF8 cKO NLDAMs in basal or inducible settings. IRF8-deficient NLDAMs upregulated Bmp2, Osm and Tgf-β1 in response to IRF8-inducible factors IFN-γ and LPS,37,38 while Vefg-α was upregulated regardless of IRF8-deficiency (Figure S7E). Such genes have well-known roles in tissue remodeling or immunosuppression.10
Figure 4.
IRF8-deficient neonate liver-derived alveolar macrophages (NLDAMs) directly facilitate tumor growth
(A) IRF8 cKO NLDAMs have reduced Irf8 and target gene expression (top) while increased inducible pro-tumor factor gene expression (bottom) compared to IRF8fl/fl NLDAMs.
(B) Schematic of 4T1 indirect co-culture with NLDAMs or BMDMs.
(C and D) IRF8 cKO NLDAMs stimulate the greatest CF compared to IRF8fl/fl controls or BMDMs (C). IRF8-transfected WT NLDAMs stimulate lower CF than IRF8-transfected IRF8 cKO NLDAMs (D). Scale bar: 370 μm.
(E) Schematic of i.n. adoptive cell transfers of NLDAMs during 4T1 experimental metastasis.
(F and G) IRF8 cKO NLDAM transfers augment 4T1 experimental metastasis in IRF8 cKO hosts (F). Reduced 4T1 experimental metastatic burden in IRF8 cKO hosts following transfers of IRF8-transfected WT NLDAMs than IRF8-transfected IRF8 cKO NLDAMs (G). Representative images are displayed. All data are represented as mean ± SEM. Significance was determined by Wilcoxon rank-sum tests with Holm-Bonferroni correction for pre-planned comparisons ($ indicates significantly different from all groups; n ≥ 6 mice/group). ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001; ∗∗∗∗p < 0.0001. See also Figures S7 and S8.
The impact of IRF8 cKO NLDAMs on in vitro 4T1 or parental E0771 mammary tumor cell growth was examined by two approaches: 1) conditioned media; and 2) indirect co-culture via transwell assay. NLDAM-conditioned media increased 4T1 tumor growth compared to control media alone, demonstrating a pro-tumor role of AM secretory products (Figure S8A). Significantly more and larger 4T1 colonies grew in response to IRF8 cKO NLDAM-conditioned media than IRF8fl/fl NLDAM-conditioned media. IRF8 cKO NLDAM-conditioned media also increased E0771 CF (Figure S8B). Next, indirect co-culture of NLDAMs expanded 4T1 or E0771 colony number and size in a cell-contact-independent manner, reinforcing the impact of AM-derived products on tumor growth (Figures 4B, 4C, and S8C). Importantly, greater CF occurred in response to indirect contact with IRF8 cKO NLDAMs than IRF8fl/fl controls. Results were compared to BMDM-conditioned media or indirect co-culture (Figures 4C and S8A). While IRF8 cKO BMDMs have reduced IRF8 mRNA and protein levels (Figures S7F and S7G), 4T1 colonies were not as plentiful nor dense as observed in response to indirect co-culture with IRF8 cKO NLDAMs. NLDAMs thus exert a rapid ability to potentiate tumor growth in vitro, especially upon IRF8-deficiency.
A complementary gain-of-function approach was performed by transfecting WT or IRF8 cKO NLDAMs with an IRF8-expression plasmid. For proof-of-concept, we compared IRF8-transfected cKO NLDAMs as a control versus IRF8-transfected WT NLDAMs. While higher IRF8 levels were detected in control- or IRF8-transfected WT NLDAMs as anticipated by augmenting basal IRF8 expression, IRF8-transfected IRF8 cKO NLDAMs displayed IRF8 levels greater than the non-transfected WT controls (Figure S7H). High transfection efficiency and stability was separately confirmed with an EGFP-expressing plasmid (Figure S7I). Low 4T1 CF was then observed upon indirect co-culture with control- or IRF8-transfected NLDAMs (Figure 4D). Importantly, fewer colonies grew in response to IRF8-transfected WT NLDAMs than IRF8-transfected IRF8 cKO NLDAMs, which directly paralleled their relationship with the enforced IRF8 levels (Figures 4D and S7H).
NLDAM expansion in vitro provides a unique source of AMs for in vivo adoptive cell transfer. We tested whether transferring NLDAMs with differential IRF8 expression by the intranasal route alters post-extravasation metastatic growth of 4T1. NLDAM transfers were administered in non-depleted, IRF8-deficient hosts to eliminate confounding factors involving the proficient background of IRF8fl/fl hosts and depletion-induced inflammation or granulocyte infiltration.61 Metastatic nodules were detectable and enhanced in IRF8 cKO hosts that received two transfers of IRF8fl/fl NLDAMs (Figures 4E, 4F, and S8D), supporting an AM pro-metastatic role. Intranasal transfers of IRF8-deficient NLDAMs doubled the number of metastatic nodules, covered lobes entirely and strengthened evidence that IRF8-deficient AMs augment metastatic outcome. Separately, we tested a gain-of-function approach by transferring IRF8-transfected NLDAMs. Intranasal transfers of IRF8-transfected NLDAMs led to fewer metastatic nodules compared to transfers of IRF8-transfected IRF8 cKO NLDAMs (Figures 4G and S8E). These results reinforce the importance of high AM-intrinsic IRF8 expression for reducing metastasis.
G-CSF is a TAF that reduces IRF8 expression in AMs and a therapeutic target for recovering AM-mediated anti-metastatic activity
To examine TAFs that may negatively regulate IRF8, we focused on G-CSF, an abundant TAF of mammary cancer including TNBC.28,32,43,44 G-CSF is highly expressed by 4T1, followed by 231/LM2-4LUC+ and MTAG models as measured within tumor-conditioned media in vitro and lysate from orthotopic primary tumors in vivo (Figure 5A). In contrast, EMT6 and E0771.ML-1 (a metastatic variant of E0771) models express low G-CSF levels and did not induce myeloid IRF8 downregulation, consistent with the broader notion that TAF expression and concentration varies among tumor subtypes. Translational relevance of G-CSF to human breast cancer was assessed among surgical tumor resections from pathological stage- and race-matched non-TNBC and TNBC patients (Table S1 and Figure S9A). Lysate from tumor cell suspensions of TNBC patients had significantly higher G-CSF (Figure 5B), supporting specificity of G-CSF as a TAF secreted by certain breast cancer types.
Figure 5.
Mammary tumor expression of G-CSF reduces IRF8 expression in AMs and promotes lung metastasis
(A) G-CSF levels within in vitro tumor-conditioned media (left) or in vivo orthotopic mammary tumors (right).
(B) Triple-negative breast cancer (TNBC) cells express more G-CSF than non-TNBC cells.
(C) Lung and spleen weights of NTB WT mice following local recombinant G-CSF (rG-CSF) exposure.
(D and E) Myeloid cell percentages (D) and IRF8 levels (E) within lung tissue of control or rG-CSF-treated mice.
(F) Schematic of local G-CSF blockade during orthotopic 4T1 growth. Tumor, lung and spleen weights from treated 4T1-bearing WT mice compared to isotype controls and baseline NTB tissue.
(G) I.n. anti-G-CSF reduces lung micro-metastasis compared to isotype control treatment of 4T1-bearing mice.
(H) Myeloid cell percentages within lung tissue following anti-G-CSF treatment.
(I) Schematic of local G-CSF blockade during orthotopic 231/LM2-4LUC+ growth and weights as in (F).
(J) I.n. blockade of human or murine G-CSF reduces lung micro-metastasis of 231/LM2-4LUC+-bearing SCID mice compared to isotype controls.
(K) Myeloid cell percentages within lung tissue following human or murine G-CSF blockade. All data are represented as mean ± SEM. Significance was determined by Wilcoxon rank-sum tests with Holm-Bonferroni correction for pre-planned comparisons (A, F, I, and J), unpaired t test with Welch’s correction (B and E), Mann-Whitney (C, D and G), or Dunnett’s test for correction of comparisons to NTB control (H and K; n ≥ 5 mice/group). ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001; ∗∗∗∗p < 0.0001. See also Figure S9 and Table S1.
To assess how G-CSF alters basal IRF8 expression of AMs in absence of granulocyte infiltration, BALB/c WT NTB mice were locally treated twice with recombinant G-CSF (rG-CSF) by intranasal route (Figures 5C and 5D). Short-acting rG-CSF significantly decreased IRF8 in AMs and was maintained 48 h post-exposure (Figure 5E). Other lung macrophage IRF8 levels were not altered, suggesting AM-specific effects in response to local G-CSF exposure.
To assess the impact of local G-CSF on detectable spontaneous micro-metastasis, BALB/c female WT mice bearing orthotopic 4T1 tumor volumes >100 mm3 were randomized and treated twice with anti-G-CSF or an isotype control by intranasal route (Figure 5F). Short-term local G-CSF blockade significantly decreased 4T1 lung micro-metastasis compared to the isotype controls (Figure 5G). The selected dose of local G-CSF blockade did not reduce granulocyte percentages within lung tissue of 4T1-bearing mice (Figure 5H), suggesting anti-metastatic activity independent of the granulocyte response. To gain insights into the source of G-CSF among lung tissue, we employed the orthotopic 231/LM2-4LUC+ model in SCID mice bearing tumor volumes >20 mm3 (Figure 5I). As human G-CSF is well-known to cross-react in mice,62 mice were randomized for treatment with species-specific anti-G-CSF to target either murine (stroma-derived) or human (tumor-derived) G-CSF. Local blockade of human or murine G-CSF significantly reduced micro-metastasis formation in comparison to the isotype controls without altering percentages of lung granulocytes (Figures 5J and 5K). Furthermore, anti-metastatic activity observed within the SCID model underscores the myeloid role for therapeutic efficacy of local G-CSF blockade in the absence of adaptive immunity.
To evaluate local G-CSF blockade as a pre-clinical therapeutic agent, we utilized a neoadjuvant approach. Anti-G-CSF or isotype control was administered twice by intranasal route into BALB/c female WT mice bearing orthotopic 4T1 tumor volumes >100 mm3 (Figure 6A). Primary tumors were surgically removed and the impact of local neoadjuvant treatment on post-surgical survival and metastatic outcome was assessed. Local neoadjuvant anti-G-CSF treatment significantly improved survival and reduced metastatic burden compared to neoadjuvant isotype control treatment (Figures 6B and 6C). To determine whether therapeutic efficacy of local neoadjuvant G-CSF blockade depended on IRF8 expression in AMs, the same experiment was performed in BALB/c IRF8 cKO mice. Local neoadjuvant anti-G-CSF treatment did not extend survival of IRF8 cKO mice post-surgery (Figure 6B), demonstrating an IRF8-dependent therapeutic effect. Greater metastatic burden was observed in IRF8 cKO mice post-surgery compared to local neoadjuvant anti-G-CSF treatment in WT mice (Figure 6C). However, neoadjuvant anti-G-CSF treatment reduced the number of metastatic nodules <1 mm in size within IRF8 cKO hosts, suggesting IRF8-independent effects of G-CSF blockade. Finally, to further evaluate AMs in the local neoadjuvant treatment response, AMs were depleted with intranasal clodronate 24 h after G-CSF blockade. Surgery was then performed 24 h post-AM depletion and compared to WT mice treated with control liposomes following G-CSF blockade (Figure 6D). AM depletion did not alter the survival advantage of local neoadjuvant G-CSF blockade in WT mice (Figure 6E). However, AM depletion augmented metastatic burden and enlarged nodule sizes (Figure 6F). Collectively, efficacy of local neoadjuvant G-CSF blockade is dependent on IRF8-expressing AMs. Our results provide evidence for utilizing G-CSF blockade in a neoadjuvant setting and as a potential macrophage ‘reprogramming’ agent to promote anti-metastatic activity.10
Figure 6.
Local G-CSF targeting recovers anti-metastatic activity via AM- and IRF8-dependent mechanisms
(A) Schematic of local neoadjuvant G-CSF blockade during orthotopic 4T1 growth compared to neoadjuvant isotype treatment. Tumor weights measured upon surgical removal. Post-surgery endpoint weights of lung or spleen.
(B and C) Local neoadjuvant G-CSF blockade extends survival (B) and reduces overall metastasis (C) of WT but not IRF8 cKO mice.
(D) Schematic of local neoadjuvant G-CSF blockade followed by single i.n. control or clodronate treatment during orthotopic 4T1 growth. Weights as in (A).
(E and F) AM depletion via i.n. clodronate treatment does not alter post-surgery survival (E) of WT mice but negates anti-metastatic effects of local neoadjuvant G-CSF blockade (F). Representative images at post-surgery endpoint. All data are represented as mean ± SEM. Significance was determined by Wilcoxon rank-sum tests with Holm-Bonferroni correction for pre-planned comparisons (A and C), Log rank (B and E) or Mann-Whitney (D and F; n ≥ 8 mice/group). ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001; ∗∗∗∗p < 0.0001.
Translational relevance of IRF8 levels in macrophage precursors of breast cancer patients
Evidence indicate that circulating monocytes sustain AM populations throughout human adulthood63 and commonly differentiate into macrophages at metastatic sites.8 We hypothesized that peripheral blood mononuclear cell (PBMC) analyses could serve as a non-invasive approach to quantify systemic IRF8 levels of macrophage precursors among various cancer stages and subtypes. Monocytes (CD33brHLA-DRbrCD15−CD11b+CD14+) collected from PBMCs of treatment-naïve non-TNBC or TNBC patients were examined for translational significance of IRF8 loss in comparison to healthy donor controls (Table S2, Figures S9B and S9C). Patient pathological primary tumor, regional lymph node and metastasis (TNM) composite scores were calculated64 and significantly increased as pathological stage progressed (Figure S9D). Patients predisposed to distant metastasis, evident by tumor cell presence in lymph node (N), blood (M) or bone marrow (M) with N or M composite integer values ≥ 1, were selected for IRF8 analysis. Utilizing flow cytometry, the specificity of IRF8 staining was confirmed in healthy donors (Figure S9C) and IRF8 expression was calculated as log2 transformation48 based on corresponding isotype control MFI values per patient sample. Monocyte IRF8 levels were determined as high or low based on the Youden index (see statistical methods).
TNM composite scores trended upward in patients with IRF8lo monocytes (Figure 7A, left). For patients with Stage II or III disease, significant correlations of TNM score were revealed in relation to IRF8 quantification. IRF8 levels and TNM scores proportionally increased in patients with IRF8hi monocytes (Figure S9E), indicating that some patients maintain higher IRF8 levels in advanced, untreated disease. In contrast, patients with IRF8lo monocytes had reduced IRF8 levels as TNM score increased (Figure 7A, right), underscoring the impact of IRF8 downregulation on breast cancer metastasis. While not all breast cancer patients are susceptible to IRF8 downregulation, such differential monocyte IRF8 expression parallels our murine studies.
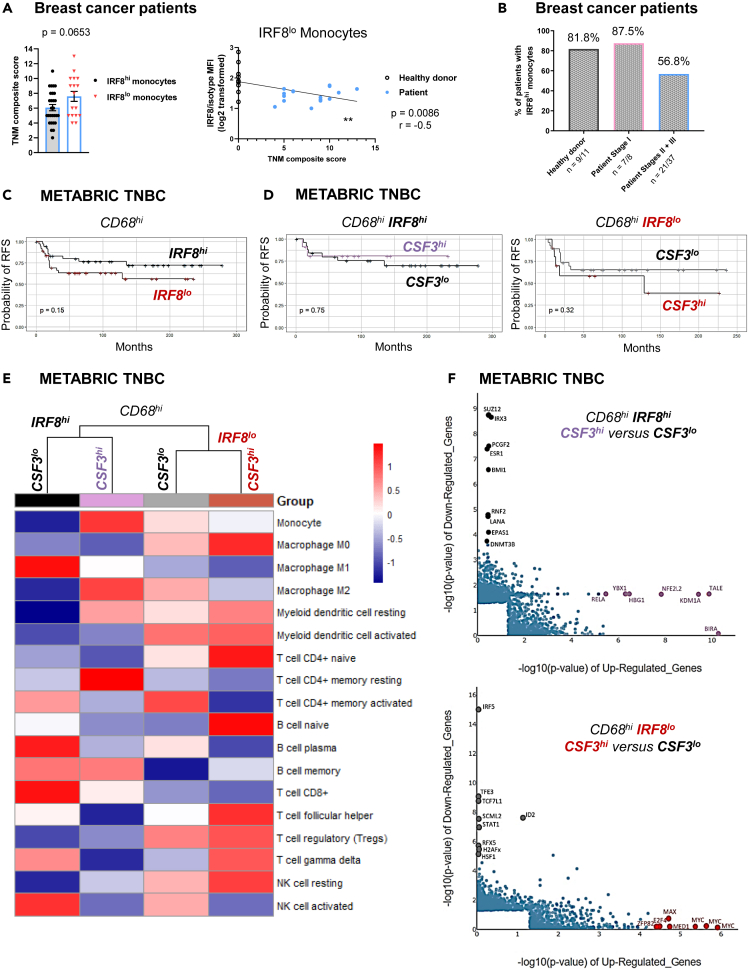
Figure 7.
CD68hiIRF8hiG-CSFlo signatures within TNBC predict immune effector gene enrichment
(A and B) Monocyte IRF8 levels of healthy donor or breast cancer patients were quantified by flow cytometry. Primary tumor, regional lymph node and metastasis (TNM) composite scores trend higher in patients with IRF8lo monocytes (A, left) and inversely correlate with IRF8 levels (A, right). Downward trend in the percentage of patients with IRF8hi monocytes as breast cancer stage advances (B).
(C) Relapse-free survival (RFS) of METABRIC TNBC patients (n = 223) with a CD68hiIRF8hi signature trends toward more favorable outcome than CD68hiIRF8lo.
(D) High G-CSF (encoded by CSF3) predicts a trend in poorer outcome of the TNBC CD68hiIRF8lo signature (right), but not in the CD68hiIRF8hi signature (left).
(E and F) Immune enrichment (E) and MR analyses (F) based on differential IRF8 and G-CSF expression in TNBC CD68hi signatures. Data are represented as mean ± SEM. Significance was determined by unpaired t test with Welch’s correction (A, left), Spearman correlation (A, right; line indicates simple linear regression) or Log rank test (C and D). ∗∗p < 0.01. Trend analyses were determined by one-sided Cochrane-Armitage test (B; p = 0.03402; n ≥ 8 patients/group). See also Figures S9, S10 and Table S2.
We then quantified the number of patients with IRF8hi monocytes based on the total number of patients per stage and performed a trend analysis. Patients with stages II and III were combined to measure greater metastatic propensity compared to stage I. A large percentage of patients with IRF8hi monocytes were observed in stage I disease, similar to the percentage of healthy donor controls. An overall decreasing trend in the percentage of patients with IRF8hi monocytes was observed in higher breast cancer stages (Figure 7B). Overall, the percentages of live monocytes did not differ among patients and healthy donors (Figure S9F), reinforcing quantitative effects of IRF8 levels on associations with TNM score or stage.
As an additional translational approach, we examined the transcriptomic relevance of CD68, a pan-macrophage marker, IRF8 and G-CSF in TNBC patients. We focused on TNBC as a G-CSF-expressing cancer and analyzed the METABRIC cohort as one of the largest available TNBC patient dataset. High expression of IRF8 alone significantly associated with improved outcome (Figure S10A), while a CD68hiIRF8hi signature trended toward better relapse-free survival (RFS; Figure 7C). G-CSF, encoded by CSF3, did not alter RFS of the CD68hiIRF8hi signature (Figure 7D). However, differences in immune gene enrichment were observed as the CD68hiIRF8hiG-CSFlo signature predicted enrichment of “M1” macrophages, plasma B cells, CD8+ T cells, and activated natural killer (NK) cells (Figure 7E). Within the CD68hiIRF8lo signature, in contrast, high G-CSF expression predicted a trend of worse RFS and enrichment of “M0” macrophages, activated dendritic cells (DCs), naive lymphocytes, regulatory T cells, and resting NK cells. High G-CSF expression results in downstream transcriptional signatures that enrich for three different motifs of MYC proliferative factors associated with up-regulated genes, while STAT1 and IRF5 associated with down-regulated genes (Figure 7F). STAT1 is an inducer of IRF8 while IRF5 is a critical binding partner of IRF8 for macrophage host defense properties.34 The resultant combination of predicted gene enrichment involving pro-tumor factors, naive or resting immune cells and the lack of effector immune gene expression may underlie poorer prognosis of TNBC patients displaying a CD68hiIRF8loG-CSFhi signature.
The beneficial CD68hiIRF8hi signature extended to improved RFS of all METABRIC breast cancer subtypes while the single IRF8hi signature did not (Figures S10B and S10C). The differential effects of G-CSF on the RFS or gene enrichment of the CD68hiIRF8lo signature did not extend to all breast cancer subtypes (Figures S10D and S10E). Furthermore, an inverse trend of G-CSF and IRF8 was only observed in TNBC CD68hi signatures (Figure S10F). Our results support context-specificity of a CD68hiIRF8loG-CSFhi axis in TNBC, reinforcing G-CSF as an important and influential TAF on the macrophage response in TNBC.
Discussion
While macrophages within the primary tumor significantly impact clinical outcome,7,65 much less is known about the contributions of TRMs to prognosis and therapy. Understanding the molecular cues that dysregulate TRM “defender” activities among pre-metastatic sites hold significant therapeutic implications for preventing metastasis. Here, we show that AMs mediate IRF8-dependent anti-metastatic activity in multiple models of mammary cancer. G-CSF renders AMs susceptible to IRF8 loss prior to detectable lung micro-metastasis, signifying an unrecognized molecular cue in impairment of anti-metastatic activity. Neoadjuvant G-CSF blockade is a therapeutic intervention to recover IRF8-dependent anti-metastatic activity of AMs. Altogether, we identified molecular signals in AMs that impact metastasis and advance the pro-metastatic role of G-CSF to TRMs, which are underappreciated responders of G-CSF compared to granulocytes.
To focus on the AM response, we analyzed early tumor growth proceeding the large recruitment of granulocytes, monocytes, or macrophages. Prior studies have established the multiple pro-metastatic roles of recruited granulocytes29,30 or macrophages.12,16,17 Macrophages recruited to lung tissue by CCL2 facilitate extravasation, colonization and growth of metastatic tumors.66 Evidence during lung infection also demonstrate that recruited macrophages may replace the original, embryonic-derived AM population.61,67 The potential impact of such bone-marrow derived AMs on metastasis remains less defined.68 Through selective routes of macrophage depletion or adoptive transfers of embryonic-derived AMs, we identified a distinct pro-metastatic role of IRF8-deficient AMs in the absence of recruited macrophages and regardless of lung neutrophilia. Thus, at early time points of mammary tumor growth, we observed that AM populations were mainly intact and preceded the large recruitment of macrophages into lung tissue. Additionally, we observed an AM phenotype with high Siglec-F expression, suggesting the retention of an embryonic-derived AM population at those time points. Whether embryonic-derived AMs are replaced at later time points during mammary tumor progression and whether bone marrow-derived AMs or transitional lung macrophages play potential pro-metastatic roles remain to be determined.
Multifactorial immune-dependent or -independent components may underlie the functional mechanisms by which IRF8 downregulation renders AMs pro-metastatic. IRF8-regulated host defense properties, such as phagocytosis, secretion of tumoricidal effector molecules or antigen presentation, may be reduced with IRF8-deficiency.37,39 Such impaired innate immune mechanisms may increase immune suppression, limiting anti-metastatic adaptive immunity. Interactions of IRF8-deficient AMs with T cells in vivo remain to be investigated and may uncover immunosuppressive modes for therapeutic targeting.23,69 Independent of the immune response, IRF8-deficient AMs may enhance early metastatic outgrowth through tissue-remodeling activities. We demonstrated that IRF8-deficient AMs upregulate inducible Bmp2, Osm and Tgf-β1 which are proliferative factors that may directly augment metastatic tumor growth or remodel the stroma.4,70,71 Thus, while we used an NLDAM culture system for these studies to reflect the embryonic origin of AMs,72 the secretion of such proliferative factors and subsequent tissue-remodeling functions of IRF8-deficient AMs remain to be evaluated in vivo, which will further strengthen the biologic merit of the NLDAM model.
Along with IRF binding partners, IRF8 is vital for amplifying STAT1-induced gene transcription for host defense and immune activation,34 including immunization against COVID-19.73 Interestingly, a recent study reported that macrophage IRF8 expression contributes to CD8+ T cell exhaustion and enhanced primary tumor growth in a murine multifocal mammary tumor model.74 IRF8-proficient macrophages expressed a gene signature involving antigen presentation and T cell chemoattractants consistent with type-1 DCs. While the reasons for this dichotomous biology remain unclear, immune exhaustion may be consequential of persistent antigen exposure and presentation or exacerbated by the multifocal, immunogenic transgenic tumor model. In our earlier41 and current studies, primary tumor growth was unaltered, yet an integral impact of macrophage IRF8 expression on metastasis was identified. Thus, context-specificity regarding the tumor model, immune interactions within the primary versus distal sites and evaluation of macrophage ontogeny may influence the role of IRF8 on the macrophage response in cancer.
We provided evidence for systemic IRF8 downregulation among multiple macrophage and monocyte populations. G-CSF most likely downregulates IRF8 expression through STAT3 activation, based on our previous studies in myeloid biology.44 Other TAFs, such as TGF-β75 or IL-1,76 may act in synergy with G-CSF to downregulate IRF8, suggesting potential TAF blockade combinations. However, micro-metastasis may occur independently of G-CSF3,5 or IRF8 downregulation, such as in EMT6 and E0771.ML-1 models. Tumor heterogeneity and the resultant cues of TAFs may contribute to differential IRF8 expression in TRMs and monocytes across different cancer types.11 Knowledge of TAF composition may thus define context-specificity in which macrophage IRF8 signatures hold prognostic merit. Our results provide the rationale for monitoring TRM IRF8 levels in G-CSF-expressing cancers. The translational relevance of IRF8 downregulation to patient outcome was assessed in blood monocytes and in our previous work that evaluated macrophages at metastatic sites.40 As abundant macrophage precursors in aging adults and cancer patients,8 blood monocytes may serve as an non-invasive surrogate for IRF8 quantification of inaccessible TRMs.
To locally target AMs, short-term intranasal administration of G-CSF blockade served as an efficacious intervention to reduce lung metastasis and extend survival in neoadjuvant settings. Our pre-clinical data has translational implications for G-CSF blockade as a potential therapeutic regimen to ‘reprogram’ macrophage function for host defense against metastasis and emphasizes local ‘reprogramming’ prior to standard-of-care treatment to enhance efficacy.10 This neoadjuvant approach may benefit patients with high risk of metastasis by delaying or preventing disease progression.77 Local AM targeting may extend to other G-CSF-expressing tumors that metastasize to the lung78 and adjuvant treatment may improve durability. Alternatively, systemic G-CSF blockade may offer ‘reprogramming’ effects across TRMs of potential metastatic sites, including bone.79 Systemic G-CSF blockade confers pre-clinical survival advantages in combination with other cancer therapies43 and blockade of the G-CSF receptor is being assessed in a Phase I interventional clinical trial for skin inflammatory diseases (NCT03972280). While no clinical trials of intranasal G-CSF blockade are reported, inhalation therapy is of current interest for lung diseases, notably inhalation of GM-CSF in COVID-19 (NCT04326920)80 or metastasis (NCT00066365).
The source of G-CSF may provide insight for the ideal TAF blockade route and our data demonstrate that local blockade of tumor- or stroma-derived G-CSF reduces micro-metastasis without inducing neutropenia. While our studies of local G-CSF targeting in a mouse-human xenograft model unveiled anti-metastatic activity independent of adaptive immunity, we cannot exclude potential anti-metastatic responses involving IRF8-dependent type-1 DCs and cytotoxic T cells that may be reintegrated by G-CSF blockade.43 Furthermore, IRF8-independent effects of G-CSF blockade may reduce metastatic colonization such as changes in vasculature.31
Overall, our results underscore the importance of IRF8 in AMs for reducing metastatic outcome, which is compromised upon exposure to certain TAFs, namely G-CSF. Our work recognizes TRMs as an attractive therapeutic target for strengthening “defender” anti-metastatic activity. We propose that IRF8lo signatures among distal TRMs may carry prognostic value as a potential biomarker of metastatic propensity in the context of G-CSF-expressing cancers.
Limitations of the study
While our study identified IRF8 downregulation in AMs as a molecular cue impairing anti-metastatic activity, the downstream events that enhance metastasis through interactions of AMs with immune-dependent or -independent elements remain to be further defined. The value of IRF8 expression in AMs for lung metastasis was supported by proof-of-concept IRF8 loss- and gain-of-function approaches. For IRF8 overexpression, we made use of a transient expression plasmid-based strategy. Our translational approaches for evaluating IRF8 expression were limited to macrophage precursors in blood or genomic assessments of primary tumors. Patient AM assessment may strengthen the prognostic utility of IRF8 downregulation on the metastatic propensity of G-CSF-expressing cancers. Finally, the potential ‘reprogramming’ mechanisms driven by local neoadjuvant G-CSF blockade that augment anti-metastatic activity of AMs and subsequent adaptive immune responses remain to be identified.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Rat anti-mouse G-CSF (clone 67604) | Leinco Technologies | Cat#G669; RRID:AB_2830259 |
| Rat IgG1 isotype control | Leinco Technologies | Cat#I-1195; RRID:AB_2830523 |
| Mouse anti-human G-CSF (clone 3316) | Leinco Technologies | Cat#G664; RRID:AB_2830254 |
| Mouse IgG1 isotype control | Leinco Technologies | Cat#I-536; RRID:AB_2737545 |
| Rat anti-mouse Ly6G (clone 1A8) | Leinco Technologies | Cat#L280; RRID:AB_2737551 |
| Rat IgG2a isotype control | Leinco Technologies | Cat#I-1177; RRID:AB_2737530 |
| Anti-mouse CD16/32 (clone 93) | BioLegend | Cat#101302; RRID:AB_312801 |
| Human IgG | Millipore Sigma | Cat#I2511; MDL:MFCD00163923; RRID:AB_1163604 |
| FITC anti-human CD11b (clone Bear1) | Beckman Coutler | Cat#IM0530U; RRID:AB_130987 |
| PE-Cy7 anti-human CD33 (clone D3HL60.251) | Beckman Coutler | Cat#A54824; RRID:AB_131163 |
| APC-H7 anti-human CD14 (clone MφP9) | BD Biosciences | Cat#560180; RRID:AB_1645464 |
| V450 anti-human CD15 (clone HI98) | BD Biosciences | Cat#561584; RRID:AB_10897841 |
| BUV395 anti-mouse CD45.2 (clone 104) | BD Horizon | Cat#564616; RRID:AB_2738867 |
| BV421 anti-mouse Siglec F (clone E50-2440) | BD Horizon | Cat#562681; RRID:AB_2722581 |
| PE-CF594 anti-mouse Siglec F (clone E50-2440) | BD Horizon | Cat#562757; RRID:AB_2687994 |
| R780/60 fixable viability dye | BD Horizon | Cat#565388 |
| BV510 anti-human/mouse B220 (clone RA3-6B2) | BioLegend | Cat#103248; RRID:AB_2650679 |
| BV510 anti-mouse CCR2 (clone SA203G11) | BioLegend | Cat#150617; RRID:AB_2721535 |
| BV711 anti-human/mouse CD11b (clone M1/70) | BioLegend | Cat#101242; RRID:AB_2563310 |
| BV785 anti-mouse CD11c (clone N418) | BioLegend | Cat#117336; RRID:AB_2565268 |
| APC anti-mouse CD115 (clone AFS98) | BioLegend | Cat#135509; RRID:AB_2085222 |
| APC anti-mouse F4/80 (clone BM8) | BioLegend | Cat#123116; RRID:AB_893481 |
| Ax700 anti-mouse Ly6C (clone HK1.4) | BioLegend | Cat#128024; RRID:AB_10643270 |
| PE-Cy7 anti-mouse Ly6G (clone 1A8) | BioLegend | Cat#127617; RRID:AB_1877262 |
| BV510 anti-mouse I-A/I-E (clone M5/114.15.2) | BioLegend | Cat#107635; RRID:AB_2561397 |
| BV605 anti-mouse I-A/I-E (clone M5/114.15.2) | BioLegend | Cat#107639; RRID:AB_2565894 |
| APC anti-human HLA-DR (clone TU36) | Invitrogen | Cat#MHLDR05; RRID:AB_10374598 |
| Aqua fixable viability dye | Invitrogen | Cat#L34966 |
| DAPI viability dye | Invitrogen | Cat#D1306 |
| PE anti-human/mouse IRF8 (clone REA516) | Miltenyi Biotec | Cat#130-122-927; RRID:AB_2857568 |
| PE recombinant antibody control isotype | Miltenyi Biotec | Cat#130-113-462; RRID:AB_2751113 |
| Bacterial and virus strains | ||
| LPS derived from E coli 0111:B4 | Millipore Sigma | Cat#L2630; MDL:MFCD00164401 |
| Biological samples | ||
| Surgical primary breast tumor resections, see Table S1 | Roswell Park Comprehensive Cancer Center | N/A |
| Peripheral blood mononuclear cells from healthy donors or breast cancer patients, see Table S2 | Roswell Park DataBank and Biorepository | N/A |
| Chemicals, peptides, and recombinant proteins | ||
| Matrigel matrix | Corning | Cat#356234 |
| Human recombinant G-CSF (Filgrastim-sndz) | Zarxio/Sandoz | Cat#46228891; NDC:61314-318-05 |
| Clodronate-encapsulated liposomes (Clophosome, neutral) | FormuMax Scientific | Cat#F70101C-N; CAS:88416-50-6 |
| Control liposomes (neutral) | FormuMax Scientific | Cat#F70101-N |
| CCR2 inhibitor (PF-4136309) | MedChemExpress | Cat#HY-13245; CAS:1341224-83-6 |
| Vehicle control (corn oil) | MedChemExpress | Cat#HY-Y1888; CAS:8001-30-7 |
| Collagenase/hyaluronidase solution | Stem Cell Technologies | Cat#07912; CAS:9001-12-1 |
| DNase I | Millipore Sigma | Cat#10104159001; CAS:9003-98-9 |
| RIPA buffer | Cell Signaling Technologies | Cat#9806S |
| PMSF | Cell Signaling Technologies | Cat#8553S; CAS:329-98-6 |
| Murine recombinant M-CSF | Peprotech | Cat#315-02 |
| Murine recombinant GM-CSF | Peprotech | Cat#315-03 |
| Murine recombinant IFN-γ | Peprotech | Cat#315-05 |
| Cellstripper non-enzymatic cell dissociation solution | Corning | Cat#25-056-CI |
| Hucker ammonium oxalate crystal violet | Carolina Biological Supply Company | Cat#867485 |
| SYBR Select mastermix | Applied Biosystems/ThermoFisher Scientific | Cat#4472908 |
| Zinc formalin fixative | Millipore Sigma | Cat#Z2902 |
| Formalin 10% neutral buffered with 0.03% eosin | Millipore Sigma | Cat#F5304 |
| Critical commercial assays | ||
| DirectPCR lysis reagent | Viagen Biotech | Cat#102-T |
| jetOPTIMUS transfection reagent | Polyplus/VWR | Cat#101000051 |
| Transcription factor staining buffer set | Miltenyi Biotec | Cat#130-122-981 |
| Murine G-CSF DuoSet ELISA | R&D Systems | Cat#DY414 |
| Human G-CSF DuoSet ELISA | R&D Systems | Cat#DY214 |
| DuoSet ELISA ancillary reagent kit | R&D Systems | Cat#DY008B |
| RNeasy miniprep kit | Qiagen | Cat#74106 |
| iScript cDNA synthesis kit | Bio-Rad | Cat#1708891 |
| High-capacity cDNA reverse transcription kit with MultiScribe reverse transcriptase | Applied Biosystems/ThermoFisher Scientific | Cat#4368814 |
| Experimental models: Cell lines | ||
| Mouse: 4T1 mammary tumor cell line | ATCC | Cat#CRL-2539 |
| Mouse: EMT6 mammary tumor cell line | ATCC | Cat#CRL-2755 |
| Mouse: E0771 mammary tumor cell line | Dr. Rosandra Kaplan, NIH | N/A |
| Mouse: E0771.ML-1 metastatic variant | Dr. Vivek Mittal, Weill Medical College at Cornell University | N/A |
| Human: 231/LM2-4LUC+ metastatic variant | Dr. John Ebos, Roswell Park | N/A |
| Experimental models: Organisms/strains | ||
| Mouse: BALB/c IRF8fl/fl: BALB/cJ(Cg)-Irf8tm1.1Hm | Speed congenic methodologies by Roswell Park Gene Targeting and Transgenic Shared Resource | N/A |
| Mouse: BALB/c Lyz2Cre: BALB/cJ(Cg)-Lyz2tm1(cre)Ifo | Speed congenic methodologies by Roswell Park Gene Targeting and Transgenic Shared Resource; Vannella et al., 201480 | N/A |
| Mouse: BALB/c IRF8 cKO (Lyz2CreIRF8fl/fl) | This paper | N/A |
| Mouse: C57BL/6 IRF8fl/fl: B6(Cg)-Irf8tm1.1Hm/J | The Jackson Laboratory | Cat#014175 |
| Mouse: C57BL/6 Lyz2Cre: B6.129P2-Lyz2tm1(cre)Ifo/J | The Jackson Laboratory | Cat#004781 |
| Mouse: C57BL/6 IRF8 cKO (Lyz2CreIRF8fl/fl) | Twum et al., 201941 | N/A |
| Mouse: C57BL/6 IRF8 gKO (Irf8−/−): B6(Cg)-Irf8tm1.2Hm/J | Dr. Keiko Ozato, NIH; Holtschke et al., 199678; The Jackson Laboratory | Cat#018298 |
| Mouse: BALB/c IRF8 gKO (Irf8−/−): BALB/cJ(Cg)-Irf8tm1.2Hm | Speed congenic methodologies by Roswell Park Gene Targeting and Transgenic Shared Resource; Colligan et al., 202279 | N/A |
| Mouse: C57BL/6 MTAG | Dr. Sandra Gendler, Mayo Clinic; Basu et al., 200481 | N/A |
| Mouse: BALB/c SCID: C.B-Igh-1b/lcrTac-Prkdcscid | Roswell Park Comparative Oncology Shared Resource | N/A |
| Mouse: BALB/c WT: BALB/c/Crl | Charles River Laboratories/NCI | Cat#555 |
| Mouse: C57BL/6 WT: C57BL/6/Crl | Charles River Laboratories/NCI | Cat#556 |
| Mouse: C57BL/6 ROSA-EYFP: B6.129X1-Gt(ROSA)26Sortm1(EYFP)Cos/J | The Jackson Laboratory | Cat#006148 |
| Mouse: C57BL/6 Lyz2CreROSA-EYFP | Kramer et al., 202351 | N/A |
| Oligonucleotides | ||
| Primers for PCR assessment of murine genotyping, see Table S3 | The Jackson Laboratory; Holtschke et al., 199678; Basu et al., 200481 | N/A |
| Primers for RT-qPCR analyses, see Table S4 | OriGene Technologies; Waight et al., 201344; Kramer et al., 202351; Colligan et al., 202279; Tamura et al., 200582; Bardina et al., 201583; Lee et al., 201884 | N/A |
| Recombinant DNA | ||
| pcDNA3.1+ control plasmid | GenoQuest; Banik et al., 201585 | N/A |
| pcDNA3.1+ IRF8 plasmid (full-length murine IRF8) | GenoQuest; Banik et al., 201585 | N/A |
| pEGFP-C1 plasmid (mammalian expression of EGFP) | Clonetech; Husain et al., 200286 | N/A |
| Software and algorithms | ||
| FCS Express (Version 7) | De Novo Software | https://denovosoftware.com/full-access/download-landing/ |
| Gen5 (Version 3) | Agilent | https://www.agilent.com/en/product/microplate-instrumentation/microplate-instrumentation-control-analysis-software/imager-reader-control-analysis-software/biotek-gen5-software-for-detection-1623227 |
| CFX Maestro (Version 1) | Bio-Rad | https://www.bio-rad.com/en-us/product/cfx-maestro-software-for-cfx-real-time-pcr-instruments?ID=OKZP7E15 |
| QuantStudio 3 Design & Analysis (Version 1) | Applied Biosystems/ThermoFisher Scientific | https://www.thermofisher.com/us/en/home/global/forms/life-science/quantstudio-3-5-software.html |
| Prism (Version 9) | GraphPad | https://www.graphpad.com/features |
| RStudio (Version 3.6.1 or 4.1.2) | posit | https://posit.co/products/open-source/rstudio/ |
| cBioPortal | Gao et al., 201387 | https://www.cbioportal.org |
| Molecular Signatures Database (MSigDB) | Liberzon et al., 201588 | https://www.gsea-msigdb.org/gsea/msigdb |
| Tumor Immune Estimation Resource (TIMER) | Li et al., 202089 | http://cistrome.org/TIMER/ |
| Lisa | Qin et al., 202090 | http://lisa.cistrome.org/ |
| Other | ||
| gentleMACS system and dissociator tubes | Miltenyi Biotec | Cat#130-096-334 |
| Microvette capillary blood tubes | Sarstedt | Cat#15-1671-100 |
| Untreated culture dishes | Corning | Cat#430591 |
| 1 μm pore size cell culture insert | Falcon | Cat#353102 |
| ZOE Fluorescent Cell Imager | Bio-Rad | Cat#1450031 |
| Echo Revolve 4 microscope | Avantor/VWR | Cat#76490-302 |
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact, Dr. Scott Abrams (scott.abrams@roswellpark.org).
Materials availability
This study did not generate new unique reagents.
Data and code availability
Data reported in this paper will be shared by the lead contact upon request. This paper does not report original code. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Experimental model and study participant details
Patient samples
Surgical primary tumor resections were collected from 10 female breast cancer patients (protocol BDR 030312; Table S1). In independent studies, peripheral blood mononuclear cells (PBMCs) were collected from a total of 11 female healthy donors and 45 female breast cancer patients (protocol NHR 008510; Table S2) prior to treatment according to standard collection procedures and stored in liquid nitrogen until analysis. All patients provided written informed consent. All research was performed according to guidelines and regulations of Roswell Park Comprehensive Cancer Center Institutional Review Board, with sample collection according to the Declaration of Helsinki.
Animals
Sources of all BALB/c or C57BL/6 murine strains are identified (key resources table). Global IRF8 knockout (a kind gift from Dr. Keiko Ozato, NIH)81,91 and IRF8fl/fl mice on a C57BL/6 background were backcrossed to >99% purity onto a BALB/c background via marker-assisted backcrossing (speed congenics) using polymorphic microsatellites in collaboration with Dr. Aimee Stablewski (Roswell Park Gene Targeting and Transgenic Shared Resource) and Dr. Fernando Benavides (MD Anderson Cancer Center). Lyz2Cre mice (kindly provided by Dr. Thomas Winn, NIH)85 on a BALB/c background were further backcrossed to >99% purity onto a BALB/c background. IRF8 cKO (BALB/c or C57BL/6), IRF8 gKO (BALB/c), Lyz2CreROSA-EYFP (C57BL/6), MTAG (C57BL/6), and SCID (BALB/c) mice were bred at Roswell Park. All research was approved and performed according to Roswell Park Institutional Animal Care and Use Committee (IACUC) protocols 1108M, 1117M and 1407M, and the National Institutes of Health (NIH) Guide for the Care and Use of Laboratory Animals. Mice were housed in specific pathogen-free conditions within an American Association for the Accreditation of Laboratory Animals Care (AAALAC)-designated facility at Roswell Park. For all mammary tumor experiments, female mice aged 8–12 weeks were utilized. For MTAG experiments, female mice were monitored beginning at 3 months and evaluated by 4 months. Male mice aged 8–12 weeks were utilized for non-tumor experiments, including in vitro BMDM analyses and in vivo rG-CSF treatment. Littermates of the same sex were randomly assigned to control or experimental treatment groups. Tail clips provided a source of DNA for routine genotyping using the DirectPCR lysis reagent according to the manufacturer’s protocol. Primer sequences for genotyping (Table S3) were obtained from The Jackson Laboratory or published reports.81,86
Tumor mouse models
For orthotopic tumor implantation, 5 × 104 4T1 cells or 3 × 105 EMT6 cells were implanted into the 4th mammary gland of female BALB/c mice. 1 × 105 E0771.ML-1 cells were implanted with Matrigel into the 4th mammary gland of female C57BL/6 mice. Tumor volumes were calculated with the equation (width2 × length)/2. Mice were euthanized at indicated primary tumor volumes, spanning 3–4 weeks of tumor growth. 1 × 106 231/LM2-4LUC+ cells were implanted into the 4th mammary gland of female BALB/c SCID mice and mice were euthanized within 1.5 weeks of tumor growth. For MTAG studies, female C57BL/6 mice were monitored for spontaneous mammary tumor formation beginning at 3 months.49 Mice were euthanized by 4 months, with three or less palpable primary mammary tumors. For experimental metastasis, 5 × 104 4T1 cells (BALB/c) or 2 × 105 parental E0771 cells (C57BL/6) were injected intravenously by tail vein. For macrophage depletion studies, 1 × 105 4T1 cells were injected intravenously by tail vein. Mice were euthanized within 3 weeks post-intravenous injection of tumor cells. 4T1 and EMT6 cells were purchased from ATCC and maintained in complete RPMI (Figure S1B) at 37°C. E0771 cells were a kind gift from Dr. Rosandra Kaplan (NIH) while the metastatic variant E0771.ML-1 was from Dr. Vivek Mittal (Weill Medical College at Cornell University) and the metastatic variant 231/LM2-4LUC+ was from Dr. John Ebos (Roswell Park). E0771, E0771.ML-1 and 231/LM2-4LUC+ cells were maintained in supplemented DMEM (Figure S1B) at 37°C. All cell lines were authenticated at ATCC by STR profiling and tested regularly for mycoplasma to confirm no evidence of infection. Tumor-conditioned media was obtained 24 h after seeding 5 × 105 tumor cells/mL and banked at −80°C.
Method details
In vivo treatments
For 4T1 orthotopic tumors, intranasal (i.n.) treatment of 10 μg rat anti-mouse G-CSF or rat IgG1 isotype control was initiated at 3 weeks of tumor growth (tumor volumes >100 mm3) under anesthesia. A total of two treatments were administered, 4 days apart as indicated in experimental schemes. At non-surgical endpoint, 48 h following anti-G-CSF or isotype treatment, lungs were digested to detect micro-metastasis. In neoadjuvant anti-G-CSF experiments combined with AM depletion, 60 μL clodronate-encapsulated or control liposomes were administered by i.n. route, under anesthesia, 24 h after anti-G-CSF treatment. For 231/LM2-4LUC+ orthotopic tumors, one i.n. treatment of 10 μg mouse anti-human G-CSF, rat anti-mouse G-CSF or mouse IgG1 isotype control was initiated at 1 week of tumor growth (tumor volumes >20 mm3) under anesthesia. Lung tissue was digested 96 h following anti-G-CSF or isotype treatment to assess micro-metastasis. To observe the local effects of G-CSF in absence of primary tumor growth, i.n. treatment of 40 μg human recombinant G-CSF (rG-CSF also known as Filgrastim) was administered under anesthesia twice, 4 days apart compared to untreated controls. At endpoint, 48 h following rG-CSF treatment, lungs were digested for flow cytometry analysis.
To induce macrophage depletion prior to and during the course of 4T1 experimental metastasis, clodronate-encapsulated or control liposomes were administered as described by our laboratory previously.57 Briefly, 60 μL clodronate was administered by i.n. route under anesthesia, while 100 μL clodronate was administered by intraperitoneal (i.p.) route compared to control liposome treatment. As an independent approach, 1 mg of a CCR2 inhibitor (PF-4136309) was administered by i.p. route compared to vehicle control (corn oil). At endpoint, lungs were collected and fixed to quantify metastatic tumor burden. For pathologist evaluation and histological assessment of lung tissue in response to clodronate-induced macrophage depletion, formalin-fixed paraffin-embedded blocks were prepared for hematoxylin and eosin (H&E) staining. Images of H&E staining were taken with the Echo Revolve 4 microscope (Avantor/VWR). To deplete granulocytes prior to and throughout 4T1 experimental metastasis, 50 μg rat anti-mouse Ly6G was administered by i.p. route compared to rat IgG2a isotype. Lung tissue was fixed to quantify metastatic tumor burden at endpoint.
Surgery
For 4T1 orthotopic tumors, surgery was performed at ∼4 weeks of tumor growth, following neoadjuvant anti-G-CSF or isotype control treatment. Adjacent skin and muscle tissue was microscopically evaluated for residual tumor tissue to achieve a complete resection of the primary tumor. The skin was repaired with 6-0 vicryl or black silk sutures and Vet-bond was applied to the incision site. No primary tumor regrowth occurred. Overall survival (OS) was monitored until approved ethical endpoints. Lungs were collected and fixed to quantify metastatic tumor burden.
Murine tissue harvests
Lung or mammary tissue was sliced, digested with a collagenase/hyaluronidase solution supplemented with 30 μg/mL DNase I and dissociated using the gentleMACS system (Miltenyi Biotec). Tissue was incubated at 37°C for 1 h and manually swirled. Tissue samples were filtered through 100 μm strainers and red blood cells (RBCs) in lung samples were briefly lysed on ice for 2 min. Single-cell suspensions were stained for flow cytometry analyses and plated to detect micro-metastasis in lung tissue. Single-cell suspensions obtained from MTAG tumors were seeded 5 × 105cells/mL to collect 24-h tumor-conditioned media. To obtain lysate from primary mammary tumors, protein was extracted from sliced tissue using 1 mL 1X RIPA buffer supplemented with 5 μL 1 mM PMSF per 0.1g of tissue. Tissue was incubated on ice for 5 min and pelleted at 14000xg for 10 min. Lysate was banked at −80°C until ELISA analysis.
Peripheral blood was collected into Microvette capillary blood tubes via retro-orbital bleed under anesthesia. RBCs were lysed for 10 min on ice. Blood cell suspensions were stained and analyzed by flow cytometry. Spleen was manually mashed, filtered through 100 μm strainers, and RBCs were lysed for 10 min on ice. Single-cell suspensions were stained and analyzed by flow cytometry.
Processing of human tumors
Small pieces of fresh surgical resections were sliced, dissociated using the gentleMACS system (Miltenyi Biotec) and digested with a collagenase/hyaluronidase solution at 37°C for 1 h. Tumor cells were filtered and single-cell suspensions were stored in liquid nitrogen until analysis. Single-cell suspensions were thawed in RPMI containing 0.1 mg/mL DNase I and pelleted at 1200 rpm for 5 min. To obtain lysate from 1 × 106 tumor cell suspensions, protein was extracted using 40 μL 1X RIPA buffer supplemented with 0.2 μL 1 mM PMSF, incubated on ice for 5 min and pelleted at 14000xg for 10 min. Lysate was banked at −80°C until ELISA analysis.
In vitro macrophage differentiation
Traditional bone marrow-derived macrophages (BMDMs) were differentiated in vitro from bone marrow progenitors as described previously by our laboratory.41 Briefly, 2 × 106 bone marrow cells (2 × 105cells/mL) were plated in untreated culture dishes in complete RPMI (Figure S1B) with 30 ng/mL M-CSF for 5 days. To reflect the embryonic origin of TRMs, neonate liver provided a source of progenitors for the in vitro differentiation of neonate liver-derived alveolar macrophages (NLDAMs), based on an adapted protocol.60 Briefly, 2 × 106 neonate liver cells (2 × 105cells/mL) were plated in untreated culture dishes in complete RPMI (Figure S1B) with 30 ng/mL GM-CSF for 7–14 days, with weekly replenishment of 30 ng/mL GM-CSF. GM-CSF, the main growth factor of AMs, was selected as the differentiation cytokine. This differentiation assay may be applied to other types of TRMs when M-CSF is utilized as the differentiation cytokine and are referred to as neonate liver-derived TRMs (NLTRMs).
Macrophage-conditioned media was obtained from confluent NLDAMs or BMDMs, utilized in a 1:1 ratio with complete media for in vitro assays and banked at −80°C. For RT-qPCR studies, NLDAMs or BMDMs were treated with 100 U/mL IFN-γ and 1 μg/mL LPS for 24 h as a positive inducer of Irf8 expression. NLDAMs or BMDMs were harvested from untreated culture dishes using Cellstripper as described92 for downstream analyses including flow cytometry, in vitro indirect co-culture or in vivo adoptive transfer studies. For in vitro indirect co-culture, 1 × 105 NLDAMs were placed in a sterile, 1 μm pore size cell culture insert within a 6-well plate to assess 48-h growth of 1 × 103 mammary tumor cells. NLDAMs were indirectly co-cultured in excess to mimic the abundance of AMs within lung tissue. For in vivo adoptive transfer studies, 2.5 × 105 NLDAMs were administered by i.n. route once a week under anesthesia during 4T1 experimental metastasis. Unlike BMDMs, NLDAMs grow in vitro for multiple passages and may be frozen in heat-inactivated FBS supplemented with 10% DMSO for future cultures. For all assays, at least three biological replicates of macrophages/genotype were utilized.
NLDAM transfection
WT or IRF8 cKO NLDAMs were transfected with a full-length murine IRF8 expression plasmid82 driven by the CMV promoter in comparison to a control pcDNA3.1+ plasmid using the jetOPTIMUS transfection reagent and the manufacturer’s protocol (Polyplus). Briefly, the transfection mix consisted of 1 μg DNA and 1 μL jetOPTIMUS reagent in 500 μL jetOPTIMUS buffer. After confluent NLDAMs were incubated with the transfection mix for 1.5 h, the transfection mix was removed and NLDAMs were cultured in fresh complete RPMI with GM-CSF supplementation. Transfection efficiency and stability was evaluated with an EGFP expression plasmid.83 EGFP-transfected NLDAMs were observed with the ZOE Fluorescent Cell Imager (Bio-Rad) for proliferation and maintained EGFP expression for at least a month post-transfection. Control- or IRF8-transfected NLDAMs were utilized 4 days post-transfection for flow cytometry, indirect co-culture or in vivo adoptive transfer studies.
In vitro micro-metastasis detection
To assess metastatic tumor colony formation (CF) within lung tissue of mice bearing orthotopic, primary mammary tumors, 3 × 105 lung cells were plated in 75 cm2 flasks and incubated at 37°C. Within 24 h, media was discarded to remove non-adherent cells. Fresh supplemented media (Figure S1B) was added and flasks were monitored 1–3 weeks for micro-metastasis. For in vitro macrophage studies, 1 × 103 mammary tumor cells were plated in 6-well plates for 48-h incubation at 37°C. Three distinct passages of tumor cells were observed in triplicate wells/macrophage genotype. At endpoint, flask or well media was removed and rinsed prior to colony fixation with 100% methanol for 30 min. Fixed CF were stained with 3% crystal violet solution for 30 min. CF was observed under a light microscope to assess tumor morphology and counted manually for total number of colonies. Images were taken with the Echo Revolve 4 microscope (Avantor/VWR).
Flow cytometry
1 × 106 murine single-cell suspensions underwent Fc receptor (FcR) blocking with 0.5 μL anti-mouse CD16/32 in 50 μL flow buffer (1X PBS supplemented with 2 mM EDTA and 0.5% BSA) for 10 min at 4°C. Single-cell suspensions were surface stained with murine antibody (key resources table) panel mastermixes in 50 μL flow buffer for 30 min at 4°C and kept dark. For samples undergoing intracellular staining, viability dye was added during surface staining and samples were thoroughly washed to remove excess viability dye prior to fixation. For intracellular staining of IRF8, two tubes of surface stained single-cell suspensions were prepared for each sample to compare IRF8 staining with isotype control. Single-cell suspensions were fixed and permeabilized based on the manufacturer’s protocol (Miltenyi Biotec) for 30 min at 4°C and kept dark. 0.5 μL anti-mouse CD16/32 and 5 μL human IgG in 100 μL 1X perm buffer was added as an additional blocking step for 10 min at 4°C. Murine single-cell suspensions were intracellularly stained with 100ug/mL anti-IRF8 or isotype control overnight at 4°C and kept dark until flow analyses.
1 × 106 human PBMCs underwent FcR blocking with 5 μL human IgG in 50 μL flow buffer for 10 min at 4°C. PBMCs were surface stained with a human antibody (key resources table) panel mastermix in 50 μL flow buffer for 30 min at 4°C and kept dark. Two tubes of surface stained single-cell suspensions were prepared for each sample to compare IRF8 staining with isotype control, similarly to murine studies. Following fixation and permeabilization, 5 μL human IgG in 100 μL 1X perm buffer was added as an additional blocking step for 10 min at 4°C. Human PBMCs were intracellularly stained with 50ug/mL anti-IRF8 or isotype control for 30 min at 4°C and kept dark until flow analyses.
Samples were acquired on a Fortessa flow cytometer (BD Biosciences) and analyzed using the FCS Express software with depicted gating strategies (Figures S2 and S9). Published reports were consulted to fine-tune gating strategies of murine lung59 and mammary84 tissues. IRF8 expression was calculated using the log2 transformation equation LOG (Sample MFI/isotype MFI, 2) based on normalization of the anti-IRF8 MFI to the isotype control MFI.48
AM flow-sort
Murine lung tissue was prepared as a single-cell suspension and 1x106 cells were surface stained as described above. Samples were sorted for SiglecFhiLy6G−F4/80+ AMs with the Aria II sorter (BD Biosciences).
ELISA
Tumor-conditioned media and murine primary mammary tumor lysate were utilized undiluted except for 4T1-conditioned media and 4T1 tumor lysate diluted 1:30. Lysate from patient tumor cell suspensions were utilized as a 1:30 dilution. G-CSF within tumor-conditioned media or lysate was assessed in technical duplicates and detected based on the manufacturer’s protocol (R&D Systems). Absorbance at 450 nm was quantified using the Gen5 program on the Synergy H1 Hybrid multi-mode microplate reader (BioTek). Background correction was performed by subtracting background absorbance observed at 540 nm. G-CSF expression was calculated based on a four parameter logistic curve.
RT-qPCR
RNA was isolated from sorted AMs, in vitro NLDAMs or BMDMs based on the manufacturer’s protocol (Qiagen). iScript reverse transcriptase was utilized to synthesize 0.1 μg cDNA. cDNA was diluted 1:5 in at least technical duplicates for real-time quantitative PCR (RT-qPCR) assessment in a 96-well plate, SYBR Select mastermix was quantified by the CFX96 Real Time system (Bio-Rad) and expression was analyzed using the Bio-Rad CFX Maestro. For Figure 1E, cDNA was synthesized with MultiScribe reverse transcriptase, RT-qPCR assessed by the QuantStudio 3 Real-Time PCR Instrument (Applied Biosystems/ThermoFisher Scientific) and expression analyzed with the QuantStudio 3 Design & Analysis Software. 25 μM primer pairs (Table S4) were used and assessed for a single melting curve per sample. Primer sequences were obtained from OriGene Technologies or published reports44,51,87,91,93,94 and validated by NCBI primer BLAST. Relative expression was normalized to Ppia expression and calculated using the equation POWER(2, Ppia mean – experimental value).
Metastatic nodule quantification
Lungs were collected and fixed in zinc formalin (for experimental metastasis endpoints) or formalin (for post-surgery endpoints) for 24 h prior to storage in 70% ethanol at room temperature. Lobes of the lung were separated to individually visualize and count macroscopic metastatic nodules with a stereoscope. Macroscopic metastatic nodules ≥1 mm were measured with a ruler to quantify nodule size.
Quantification and statistical analysis
Statistical methods
Data was analyzed with Prism software and multiple comparisons were performed with RStudio software (Version 4.1.2). For murine flow cytometry analyses, unpaired Welch’s t test (unequal variances) was utilized to assess two groups while multiple comparisons were assessed by Dunnett’s test for correction of comparisons to NTB control or Holm-Bonferroni correction for pre-planned comparisons. IRF8 flow cytometry data was also assessed by Spearman correlation in tumor settings. For other murine data analyses, Mann-Whitney tests were utilized to assess two groups while multiple comparisons were assessed by Wilcoxon rank-sum tests with Holm-Bonferroni correction for pre-planned comparisons. All data are depicted as mean ± SEM. Log rank tests were utilized to determine differences in OS as depicted by Kaplan-Meier curves. Asterisks are noted in figures and p values are detailed within legends. P-values <0.05 were considered statistically significant. Three independent experiments with a minimum of three biological replicates were performed for in vitro analyses. Two independent experiments with large biological replicates were performed for in vivo analyses. Randomization of mice occurred prior to administration of in vivo treatments.
For patient data analysis, pathological TNM composite scores were calculated based on the sum of composite integer T (1–5), N (0–7) and M (0–2) values guided by the American Joint Committee on Cancer (AJCC) designations of advancing breast cancer. Patients with N or M composite integer values ≥ 1 were selected and analyzed by Spearman correlation or unpaired Welch’s t test (unequal variances) in Prism software. Subsequent analyses utilized RStudio software (Version 4.1.2). Two independent experiments with large biological replicates were performed. A regression analysis on batch and stage was performed to estimate and compensate for the batch effect. An optimal threshold of IRF8 expression was selected by maximizing the Youden index for discriminating healthy/stage I versus stage II/III subjects. IRF8 values greater than the selected cutoff of 1.76 were characterized as IRF8hi. One-sided Cochrane Armitage test was utilized to assess the overall decreasing trend of patients with IRF8hi monocytes. All data are depicted as mean ± SEM. Asterisks are noted in figures and p values are detailed within legends. P-values <0.05 were considered statistically significant.
METABRIC patient stratification and analysis
Statistical analyses were performed in RStudio (Version 3.6.1) and related dependencies. Publically available transcriptome data from the Molecular Taxonomy of Breast Cancer International Consortium (METABRIC; n = 1904) was obtained from cBioPortal95 as normalized counts. In addition to counts, clinical data, which included all PAM50 molecular subtypes,88 OS and relapse-free survival (RFS) was obtained. TNBC patients were selected based on negative hormonal receptor status of ER, PR and HER2 (n = 223). Designation of highly expressing a gene was determined using tertiles, distinguishing high (33%) from low (67%) gene expression for CD68, IRF8 and CSF3. A combination signature of all three was then applied to delineate clinical populations for comparison. Kaplan-Meier curves for RFS were constructed based on the TNBC and total METABRIC datasets using the ‘SurvMiner’ package. Log rank p values <0.05 were considered statistically significant. Transcriptomic counts were normalized prior to download. Therefore, data was scaled and differential expression was performed on normalized counts using ‘Limma’.89 Differential expression rank order was used for subsequent gene set enrichment analysis (GSEA),90 performed using the ‘clusterprofile’ package in R. Significant pathways were indicated by an adjusted p value q < 0.05. Gene sets queried included the Hallmark, Canonical pathways and GO Biological Processes Oncology collections available in the Molecular Signatures Database (MSigDB).96 Immune cell type deconvolution was performed using the Tumor Immune Estimation Resource (TIMER)97 and all heatmaps resulted from utilization of the ‘pheatmap’ package. Master regulator (MR) analysis was conducted on the top statistically significant (adjusted p < 0.05, ǀlogFCǀ>0.58, unless otherwise indicated), using LISA.98 Correlation of expression was conducted utilizing the ‘cor.stat’ function in R, with a Spearman rank correlation coefficient test.
Acknowledgments
We gratefully thank our patients and their families. We thank Dr. Jennifer Guerriero (Brigham and Women’s Hospital, Dana-Farber/Harvard Cancer Center) for intriguing macrophage discussions, Dr. Jessica Young for clinical insights, Mary Lynn Hensen for mouse colony management, and Dr. Pragya Srivastava, Dr. Andrea Monell and Cameron MacDonald for technical assistance. For flow cytometry expertise, we thank Dr. Hans Minderman, Dr. Orla Maguire, Kieran O′ Loughlin, and Dr. Joe Tario. We thank Dr. Rosandra Kaplan (NIH) for the E0771 cell line and Dr. Vivek Mittal (Weill Medical College at Cornell University) for the E0771.ML-1 variant. Graphical abstract and summaries of experimental methods were created with BioRender.com. This work was funded by NIH R01CA172105 (S.I.A.), R01CA250412 (S.S.E. & S.I.A.), R01CA251545 (G.M.D.), R21CA256321 (C.M.B. and S.I.A.), F31CA243304 (S.L.T.), F30CA254327 (E.D.K.), and the Roswell Park Alliance Foundation (M.J.N. and S.I.A.). Previous funding includes NIH T32CA085183 (S.L.T. & E.D.K.). The following Roswell Park Shared Resources are supported by NIH P30CA016056: Comparative Oncology, Data Bank and Biorepository, Flow and Immune Analysis, Gene Modulation Services, Gene Targeting and Transgenic, and Pathology Network.
Author contributions
S.L.T., M.O., M.J.N., S.S.E., and S.I.A. designed experiments. S.L.T., E.D.K., H.M., M.K., M.R.D., C.C.O., and B.G.M. performed experiments. S.L.T., E.D.K., M.K., S.R.R., P.N.B., A.B.S., F.J.B., C.M.B., M.O., M.J.N., S.S.E., and S.I.A. performed analyses or data interpretations. S.R.R. and H.Y. performed statistical analyses. A.B.S., F.J.B, C.M.B., J.M.L.E., G.M.D., and S.S.E. provided essential reagents or models. S.L.T. and S.I.A. developed the conceptualization, wrote the manuscript and supervised the study. All authors provided manuscript editorial review.
Declaration of interests
M.O. has received research support from Alphageneron, AIM Therapeutics, Eli Lilly, and Pfizer. The other authors declare no competing interests.
Published: February 10, 2024
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2024.109187.
Supplemental information
References
- 1.Anderson R.L., Balasas T., Callaghan J., Coombes R.C., Evans J., Hall J.A., Kinrade S., Jones D., Jones P.S., Jones R., et al. A framework for the development of effective anti-metastatic agents. Nat. Rev. Clin. Oncol. 2019;16:185–204. doi: 10.1038/s41571-018-0134-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ganesh K., Massagué J. Targeting metastatic cancer. Nat. Med. 2021;27:34–44. doi: 10.1038/s41591-020-01195-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Liu Y., Cao X. Characteristics and Significance of the Pre-metastatic Niche. Cancer Cell. 2016;30:668–681. doi: 10.1016/j.ccell.2016.09.011. [DOI] [PubMed] [Google Scholar]
- 4.Correia A.L. Locally sourced: site-specific immune barriers to metastasis. Nat. Rev. Immunol. 2023;23:522–538. doi: 10.1038/s41577-023-00836-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Patras L., Shaashua L., Matei I., Lyden D. Immune determinants of the pre-metastatic niche. Cancer Cell. 2023;41:546–572. doi: 10.1016/j.ccell.2023.02.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Engblom C., Pfirschke C., Pittet M.J. The role of myeloid cells in cancer therapies. Nat. Rev. Cancer. 2016;16:447–462. doi: 10.1038/nrc.2016.54. [DOI] [PubMed] [Google Scholar]
- 7.Cassetta L., Pollard J.W. Targeting macrophages: therapeutic approaches in cancer. Nat. Rev. Drug Discov. 2018;17:887–904. doi: 10.1038/nrd.2018.169. [DOI] [PubMed] [Google Scholar]
- 8.Guilliams M., Thierry G.R., Bonnardel J., Bajenoff M. Establishment and Maintenance of the Macrophage Niche. Immunity. 2020;52:434–451. doi: 10.1016/j.immuni.2020.02.015. [DOI] [PubMed] [Google Scholar]
- 9.Ruffell B., Coussens L.M. Macrophages and therapeutic resistance in cancer. Cancer Cell. 2015;27:462–472. doi: 10.1016/j.ccell.2015.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tzetzo S.L., Abrams S.I. Redirecting macrophage function to sustain their "defender" antitumor activity. Cancer Cell. 2021;39:734–737. doi: 10.1016/j.ccell.2021.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cassetta L., Fragkogianni S., Sims A.H., Swierczak A., Forrester L.M., Zhang H., Soong D.Y.H., Cotechini T., Anur P., Lin E.Y., et al. Human Tumor-Associated Macrophage and Monocyte Transcriptional Landscapes Reveal Cancer-Specific Reprogramming, Biomarkers, and Therapeutic Targets. Cancer Cell. 2019;35:588–602.e10. doi: 10.1016/j.ccell.2019.02.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Consonni F.M., Bleve A., Totaro M.G., Storto M., Kunderfranco P., Termanini A., Pasqualini F., Alì C., Pandolfo C., Sgambelluri F., et al. Heme catabolism by tumor-associated macrophages controls metastasis formation. Nat. Immunol. 2021;22:595–606. doi: 10.1038/s41590-021-00921-5. [DOI] [PubMed] [Google Scholar]
- 13.Franklin R.A., Liao W., Sarkar A., Kim M.V., Bivona M.R., Liu K., Pamer E.G., Li M.O. The cellular and molecular origin of tumor-associated macrophages. Science. 2014;344:921–925. doi: 10.1126/science.1252510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gordon S.R., Maute R.L., Dulken B.W., Hutter G., George B.M., McCracken M.N., Gupta R., Tsai J.M., Sinha R., Corey D., et al. PD-1 expression by tumour-associated macrophages inhibits phagocytosis and tumour immunity. Nature. 2017;545:495–499. doi: 10.1038/nature22396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Guerriero J.L., Sotayo A., Ponichtera H.E., Castrillon J.A., Pourzia A.L., Schad S., Johnson S.F., Carrasco R.D., Lazo S., Bronson R.T., et al. Class IIa HDAC inhibition reduces breast tumours and metastases through anti-tumour macrophages. Nature. 2017;543:428–432. doi: 10.1038/nature21409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kitamura T., Qian B.Z., Soong D., Cassetta L., Noy R., Sugano G., Kato Y., Li J., Pollard J.W. CCL2-induced chemokine cascade promotes breast cancer metastasis by enhancing retention of metastasis-associated macrophages. J. Exp. Med. 2015;212:1043–1059. doi: 10.1084/jem.20141836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ruffell B., Chang-Strachan D., Chan V., Rosenbusch A., Ho C.M.T., Pryer N., Daniel D., Hwang E.S., Rugo H.S., Coussens L.M. Macrophage IL-10 blocks CD8+ T cell-dependent responses to chemotherapy by suppressing IL-12 expression in intratumoral dendritic cells. Cancer Cell. 2014;26:623–637. doi: 10.1016/j.ccell.2014.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Baer J.M., Zuo C., Kang L.I., de la Lastra A.A., Borcherding N.C., Knolhoff B.L., Bogner S.J., Zhu Y., Yang L., Laurent J., et al. Fibrosis induced by resident macrophages has divergent roles in pancreas inflammatory injury and PDAC. Nat. Immunol. 2023;24:1443–1457. doi: 10.1038/s41590-023-01579-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Casanova-Acebes M., Dalla E., Leader A.M., LeBerichel J., Nikolic J., Morales B.M., Brown M., Chang C., Troncoso L., Chen S.T., et al. Tissue-resident macrophages provide a pro-tumorigenic niche to early NSCLC cells. Nature. 2021;595:578–584. doi: 10.1038/s41586-021-03651-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Loyher P.L., Hamon P., Laviron M., Meghraoui-Kheddar A., Goncalves E., Deng Z., Torstensson S., Bercovici N., Baudesson de Chanville C., Combadière B., et al. Macrophages of distinct origins contribute to tumor development in the lung. J. Exp. Med. 2018;215:2536–2553. doi: 10.1084/jem.20180534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhu Y., Herndon J.M., Sojka D.K., Kim K.W., Knolhoff B.L., Zuo C., Cullinan D.R., Luo J., Bearden A.R., Lavine K.J., et al. Tissue-Resident Macrophages in Pancreatic Ductal Adenocarcinoma Originate from Embryonic Hematopoiesis and Promote Tumor Progression. Immunity. 2017;47:323–338.e6. doi: 10.1016/j.immuni.2017.07.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Zhang J., Song J., Tang S., Zhao Y., Wang L., Luo Y., Tang J., Ji Y., Wang X., Li T., et al. Multi-omics analysis reveals the chemoresistance mechanism of proliferating tissue-resident macrophages in PDAC via metabolic adaptation. Cell Rep. 2023;42 doi: 10.1016/j.celrep.2023.112620. [DOI] [PubMed] [Google Scholar]
- 23.Chow A., Schad S., Green M.D., Hellmann M.D., Allaj V., Ceglia N., Zago G., Shah N.S., Sharma S.K., Mattar M., et al. Tim-4(+) cavity-resident macrophages impair anti-tumor CD8(+) T cell immunity. Cancer Cell. 2021;39:973–988.e9. doi: 10.1016/j.ccell.2021.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Klemm F., Maas R.R., Bowman R.L., Kornete M., Soukup K., Nassiri S., Brouland J.P., Iacobuzio-Donahue C.A., Brennan C., Tabar V., et al. Interrogation of the Microenvironmental Landscape in Brain Tumors Reveals Disease-Specific Alterations of Immune Cells. Cell. 2020;181:1643–1660.e17. doi: 10.1016/j.cell.2020.05.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Tacconi C., Commerford C.D., Dieterich L.C., Schwager S., He Y., Ikenberg K., Friebel E., Becher B., Tugues S., Detmar M. CD169(+) lymph node macrophages have protective functions in mouse breast cancer metastasis. Cell Rep. 2021;35 doi: 10.1016/j.celrep.2021.108993. [DOI] [PubMed] [Google Scholar]
- 26.Timmers M., Vekemans K., Vermijlen D., Asosingh K., Kuppen P., Bouwens L., Wisse E., Braet F. Interactions between rat colon carcinoma cells and Kupffer cells during the onset of hepatic metastasis. Int. J. Cancer. 2004;112:793–802. doi: 10.1002/ijc.20481. [DOI] [PubMed] [Google Scholar]
- 27.Li W., Tanikawa T., Kryczek I., Xia H., Li G., Wu K., Wei S., Zhao L., Vatan L., Wen B., et al. Aerobic Glycolysis Controls Myeloid-Derived Suppressor Cells and Tumor Immunity via a Specific CEBPB Isoform in Triple-Negative Breast Cancer. Cell Metab. 2018;28:87–103.e6. doi: 10.1016/j.cmet.2018.04.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Waight J.D., Hu Q., Miller A., Liu S., Abrams S.I. Tumor-derived G-CSF facilitates neoplastic growth through a granulocytic myeloid-derived suppressor cell-dependent mechanism. PLoS One. 2011;6 doi: 10.1371/journal.pone.0027690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kowanetz M., Wu X., Lee J., Tan M., Hagenbeek T., Qu X., Yu L., Ross J., Korsisaari N., Cao T., et al. Granulocyte-colony stimulating factor promotes lung metastasis through mobilization of Ly6G+Ly6C+ granulocytes. Proc. Natl. Acad. Sci. USA. 2010;107:21248–21255. doi: 10.1073/pnas.1015855107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Swierczak A., Cook A.D., Lenzo J.C., Restall C.M., Doherty J.P., Anderson R.L., Hamilton J.A. The promotion of breast cancer metastasis caused by inhibition of CSF-1R/CSF-1 signaling is blocked by targeting the G-CSF receptor. Cancer Immunol. Res. 2014;2:765–776. doi: 10.1158/2326-6066.Cir-13-0190. [DOI] [PubMed] [Google Scholar]
- 31.Yip R.K.H., Rimes J.S., Capaldo B.D., Vaillant F., Mouchemore K.A., Pal B., Chen Y., Surgenor E., Murphy A.J., Anderson R.L., et al. Mammary tumour cells remodel the bone marrow vascular microenvironment to support metastasis. Nat. Commun. 2021;12:6920. doi: 10.1038/s41467-021-26556-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hollmén M., Karaman S., Schwager S., Lisibach A., Christiansen A.J., Maksimow M., Varga Z., Jalkanen S., Detmar M. G-CSF regulates macrophage phenotype and associates with poor overall survival in human triple-negative breast cancer. OncoImmunology. 2016;5 doi: 10.1080/2162402x.2015.1115177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Karagiannidis I., de Santana Van Vilet E., Said Abu Egal E., Phinney B., Jacenik D., Prossnitz E.R., Beswick E.J. G-CSF and G-CSFR Induce a Pro-Tumorigenic Macrophage Phenotype to Promote Colon and Pancreas Tumor Growth. Cancers. 2020;12 doi: 10.3390/cancers12102868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lawrence T., Natoli G. Transcriptional regulation of macrophage polarization: enabling diversity with identity. Nat. Rev. Immunol. 2011;11:750–761. doi: 10.1038/nri3088. [DOI] [PubMed] [Google Scholar]
- 35.Natoli G., Ostuni R. Adaptation and memory in immune responses. Nat. Immunol. 2019;20:783–792. doi: 10.1038/s41590-019-0399-9. [DOI] [PubMed] [Google Scholar]
- 36.Guerriero J.L. Macrophages: The Road Less Traveled, Changing Anticancer Therapy. Trends Mol. Med. 2018;24:472–489. doi: 10.1016/j.molmed.2018.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Langlais D., Barreiro L.B., Gros P. The macrophage IRF8/IRF1 regulome is required for protection against infections and is associated with chronic inflammation. J. Exp. Med. 2016;213:585–603. doi: 10.1084/jem.20151764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Mancino A., Termanini A., Barozzi I., Ghisletti S., Ostuni R., Prosperini E., Ozato K., Natoli G. A dual cis-regulatory code links IRF8 to constitutive and inducible gene expression in macrophages. Genes Dev. 2015;29:394–408. doi: 10.1101/gad.257592.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Marquis J.F., Kapoustina O., Langlais D., Ruddy R., Dufour C.R., Kim B.H., MacMicking J.D., Giguère V., Gros P. Interferon regulatory factor 8 regulates pathways for antigen presentation in myeloid cells and during tuberculosis. PLoS Genet. 2011;7 doi: 10.1371/journal.pgen.1002097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Muhitch J.B., Hoffend N.C., Azabdaftari G., Miller A., Bshara W., Morrison C.D., Schwaab T., Abrams S.I. Tumor-associated macrophage expression of interferon regulatory Factor-8 (IRF8) is a predictor of progression and patient survival in renal cell carcinoma. J. Immunother. Cancer. 2019;7:155. doi: 10.1186/s40425-019-0630-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Twum D.Y., Colligan S.H., Hoffend N.C., Katsuta E., Cortes Gomez E., Hensen M.L., Seshadri M., Nemeth M.J., Abrams S.I. IFN regulatory factor-8 expression in macrophages governs an antimetastatic program. JCI Insight. 2019;4 doi: 10.1172/jci.insight.124267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Aegerter H., Lambrecht B.N., Jakubzick C.V. Biology of lung macrophages in health and disease. Immunity. 2022;55:1564–1580. doi: 10.1016/j.immuni.2022.08.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Meyer M.A., Baer J.M., Knolhoff B.L., Nywening T.M., Panni R.Z., Su X., Weilbaecher K.N., Hawkins W.G., Ma C., Fields R.C., et al. Breast and pancreatic cancer interrupt IRF8-dependent dendritic cell development to overcome immune surveillance. Nat. Commun. 2018;9:1250. doi: 10.1038/s41467-018-03600-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Waight J.D., Netherby C., Hensen M.L., Miller A., Hu Q., Liu S., Bogner P.N., Farren M.R., Lee K.P., Liu K., Abrams S.I. Myeloid-derived suppressor cell development is regulated by a STAT/IRF-8 axis. J. Clin. Invest. 2013;123:4464–4478. doi: 10.1172/jci68189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kennecke H., Yerushalmi R., Woods R., Cheang M.C.U., Voduc D., Speers C.H., Nielsen T.O., Gelmon K. Metastatic behavior of breast cancer subtypes. J. Clin. Oncol. 2010;28:3271–3277. doi: 10.1200/jco.2009.25.9820. [DOI] [PubMed] [Google Scholar]
- 46.Harker J.A., Lloyd C.M. Overlapping and distinct features of viral and allergen immunity in the human lung. Immunity. 2021;54:617–631. doi: 10.1016/j.immuni.2021.03.010. [DOI] [PubMed] [Google Scholar]
- 47.Pulaski B.A., Ostrand-Rosenberg S. Mouse 4T1 breast tumor model. Curr Protoc Immunol Chapter. Curr. Protoc. Immunol. 2001;Chapter.20 doi: 10.1002/0471142735.im2002s39. Unit.20.2, Unit 20.22. [DOI] [PubMed] [Google Scholar]
- 48.Srivastava P., Tzetzo S.L., Gomez E.C., Eng K.H., Jani Sait S.N., Kuechle J.B., Singh P.K., De Jong K., Wiatrowski K.R., Peresie J., et al. Inhibition of LSD1 in MDS progenitors restores differentiation of CD141(Hi) conventional dendritic cells. Leukemia. 2020;34:2460–2472. doi: 10.1038/s41375-020-0765-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Stewart T.J., Lutsiak M.E.C., Abrams S.I. Immune consequences of protracted host-tumor interactions in a transgenic mouse model of mammary carcinoma. Cancer Invest. 2008;26:237–249. doi: 10.1080/07357900701708419. [DOI] [PubMed] [Google Scholar]
- 50.Piranlioglu R., Lee E., Ouzounova M., Bollag R.J., Vinyard A.H., Arbab A.S., Marasco D., Guzel M., Cowell J.K., Thangaraju M., et al. Primary tumor-induced immunity eradicates disseminated tumor cells in syngeneic mouse model. Nat. Commun. 2019;10:1430. doi: 10.1038/s41467-019-09015-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kramer E.D., Tzetzo S.L., Colligan S.H., Hensen M.L., Brackett C.M., Clausen B.E., Taketo M.M., Abrams S.I. β-Catenin signaling in alveolar macrophages enhances lung metastasis through a TNF-dependent mechanism. JCI Insight. 2023;8 doi: 10.1172/jci.insight.160978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ebos J.M.L., Lee C.R., Bogdanovic E., Alami J., Van Slyke P., Francia G., Xu P., Mutsaers A.J., Dumont D.J., Kerbel R.S. Vascular endothelial growth factor-mediated decrease in plasma soluble vascular endothelial growth factor receptor-2 levels as a surrogate biomarker for tumor growth. Cancer Res. 2008;68:521–529. doi: 10.1158/0008-5472.Can-07-3217. [DOI] [PubMed] [Google Scholar]
- 53.Abram C.L., Roberge G.L., Hu Y., Lowell C.A. Comparative analysis of the efficiency and specificity of myeloid-Cre deleting strains using ROSA-EYFP reporter mice. J. Immunol. Methods. 2014;408:89–100. doi: 10.1016/j.jim.2014.05.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Jakubzick C., Bogunovic M., Bonito A.J., Kuan E.L., Merad M., Randolph G.J. Lymph-migrating, tissue-derived dendritic cells are minor constituents within steady-state lymph nodes. J. Exp. Med. 2008;205:2839–2850. doi: 10.1084/jem.20081430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Tapmeier T.T., Howell J.H., Zhao L., Papiez B.W., Schnabel J.A., Muschel R.J., Gal A. Evolving polarisation of infiltrating and alveolar macrophages in the lung during metastatic progression of melanoma suggests CCR1 as a therapeutic target. Oncogene. 2022;41:5032–5045. doi: 10.1038/s41388-022-02488-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Wang T., Zhang J., Wang Y., Li Y., Wang L., Yu Y., Yao Y. Influenza-trained mucosal-resident alveolar macrophages confer long-term antitumor immunity in the lungs. Nat. Immunol. 2023;24:423–438. doi: 10.1038/s41590-023-01428-x. [DOI] [PubMed] [Google Scholar]
- 57.Burkard-Mandel L., O'Neill R., Colligan S., Seshadri M., Abrams S.I. Tumor-derived thymic stromal lymphopoietin enhances lung metastasis through an alveolar macrophage-dependent mechanism. OncoImmunology. 2018;7 doi: 10.1080/2162402x.2017.1419115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Gibbings S.L., Thomas S.M., Atif S.M., McCubbrey A.L., Desch A.N., Danhorn T., Leach S.M., Bratton D.L., Henson P.M., Janssen W.J., Jakubzick C.V. Three Unique Interstitial Macrophages in the Murine Lung at Steady State. Am. J. Respir. Cell Mol. Biol. 2017;57:66–76. doi: 10.1165/rcmb.2016-0361OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Kopf M., Schneider C., Nobs S.P. The development and function of lung-resident macrophages and dendritic cells. Nat. Immunol. 2015;16:36–44. doi: 10.1038/ni.3052. [DOI] [PubMed] [Google Scholar]
- 60.Fejer G., Wegner M.D., Györy I., Cohen I., Engelhard P., Voronov E., Manke T., Ruzsics Z., Dölken L., Prazeres da Costa O., et al. Nontransformed, GM-CSF-dependent macrophage lines are a unique model to study tissue macrophage functions. Proc. Natl. Acad. Sci. USA. 2013;110:E2191–E2198. doi: 10.1073/pnas.1302877110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Aegerter H., Kulikauskaite J., Crotta S., Patel H., Kelly G., Hessel E.M., Mack M., Beinke S., Wack A. Influenza-induced monocyte-derived alveolar macrophages confer prolonged antibacterial protection. Nat. Immunol. 2020;21:145–157. doi: 10.1038/s41590-019-0568-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Nguyen-Jackson H.T., Li H.S., Zhang H., Ohashi E., Watowich S.S. G-CSF-activated STAT3 enhances production of the chemokine MIP-2 in bone marrow neutrophils. J. Leukoc. Biol. 2012;92:1215–1225. doi: 10.1189/jlb.0312126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Byrne A.J., Powell J.E., O'Sullivan B.J., Ogger P.P., Hoffland A., Cook J., Bonner K.L., Hewitt R.J., Wolf S., Ghai P., et al. Dynamics of human monocytes and airway macrophages during healthy aging and after transplant. J. Exp. Med. 2020;217 doi: 10.1084/jem.20191236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Jones G.W., Browman G., Goodyear M., Marcellus D., Hodson D.I. Comparison of the addition of T and N integer scores with TNM stage groups in head and neck cancer. Head Neck. 1993;15:497–503. doi: 10.1002/hed.2880150604. [DOI] [PubMed] [Google Scholar]
- 65.Pittet M.J., Michielin O., Migliorini D. Clinical relevance of tumour-associated macrophages. Nat. Rev. Clin. Oncol. 2022;19:402–421. doi: 10.1038/s41571-022-00620-6. [DOI] [PubMed] [Google Scholar]
- 66.Qian B.Z., Li J., Zhang H., Kitamura T., Zhang J., Campion L.R., Kaiser E.A., Snyder L.A., Pollard J.W. CCL2 recruits inflammatory monocytes to facilitate breast-tumour metastasis. Nature. 2011;475:222–225. doi: 10.1038/nature10138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Li F., Piattini F., Pohlmeier L., Feng Q., Rehrauer H., Kopf M. Monocyte-derived alveolar macrophages autonomously determine severe outcome of respiratory viral infection. Sci. Immunol. 2022;7:eabj5761. doi: 10.1126/sciimmunol.abj5761. [DOI] [PubMed] [Google Scholar]
- 68.Iliakis C.S., Kulikauskaite J., Aegerter H., Li F., Piattini F., Jakubzick C.V., Guilliams M., Kopf M., Wack A. The role of recruitment versus training in influenza-induced lasting changes to alveolar macrophage function. Nat. Immunol. 2023;24:1639–1641. doi: 10.1038/s41590-023-01602-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Sun L., Kees T., Almeida A.S., Liu B., He X.Y., Ng D., Han X., Spector D.L., McNeish I.A., Gimotty P., et al. Activating a collaborative innate-adaptive immune response to control metastasis. Cancer Cell. 2021;39:1361–1374.e9. doi: 10.1016/j.ccell.2021.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Araujo A.M., Abaurrea A., Azcoaga P., López-Velazco J.I., Manzano S., Rodriguez J., Rezola R., Egia-Mendikute L., Valdés-Mora F., Flores J.M., et al. Stromal oncostatin M cytokine promotes breast cancer progression by reprogramming the tumor microenvironment. J. Clin. Invest. 2022;132 doi: 10.1172/jci148667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Wu C.K., Wei M.T., Wu H.C., Wu C.L., Wu C.J., Liaw H., Su W.P. BMP2 promotes lung adenocarcinoma metastasis through BMP receptor 2-mediated SMAD1/5 activation. Sci. Rep. 2022;12 doi: 10.1038/s41598-022-20788-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Li F., Okreglicka K.M., Piattini F., Pohlmeier L.M., Schneider C., Kopf M. Gene therapy of Csf2ra deficiency in mouse fetal monocyte precursors restores alveolar macrophage development and function. JCI Insight. 2022;7 doi: 10.1172/jci.insight.152271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Li C., Lee A., Grigoryan L., Arunachalam P.S., Scott M.K.D., Trisal M., Wimmers F., Sanyal M., Weidenbacher P.A., Feng Y., et al. Mechanisms of innate and adaptive immunity to the Pfizer-BioNTech BNT162b2 vaccine. Nat. Immunol. 2022;23:543–555. doi: 10.1038/s41590-022-01163-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Nixon B.G., Kuo F., Ji L., Liu M., Capistrano K., Do M., Franklin R.A., Wu X., Kansler E.R., Srivastava R.M., et al. Tumor-associated macrophages expressing the transcription factor IRF8 promote T cell exhaustion in cancer. Immunity. 2022;55:2044–2058.e5. doi: 10.1016/j.immuni.2022.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Papaspyridonos M., Matei I., Huang Y., do Rosario Andre M., Brazier-Mitouart H., Waite J.C., Chan A.S., Kalter J., Ramos I., Wu Q., et al. Id1 suppresses anti-tumour immune responses and promotes tumour progression by impairing myeloid cell maturation. Nat. Commun. 2015;6:6840. doi: 10.1038/ncomms7840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Allen B.M., Hiam K.J., Burnett C.E., Venida A., DeBarge R., Tenvooren I., Marquez D.M., Cho N.W., Carmi Y., Spitzer M.H. Systemic dysfunction and plasticity of the immune macroenvironment in cancer models. Nat. Med. 2020;26:1125–1134. doi: 10.1038/s41591-020-0892-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Wolf D.M., Yau C., Wulfkuhle J., Brown-Swigart L., Gallagher R.I., Lee P.R.E., Zhu Z., Magbanua M.J., Sayaman R., O'Grady N., et al. Redefining breast cancer subtypes to guide treatment prioritization and maximize response: Predictive biomarkers across 10 cancer therapies. Cancer Cell. 2022;40:609–623.e6. doi: 10.1016/j.ccell.2022.05.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Karagiannidis I., Salataj E., Said Abu Egal E., Beswick E.J. G-CSF in tumors: Aggressiveness, tumor microenvironment and immune cell regulation. Cytokine. 2021;142 doi: 10.1016/j.cyto.2021.155479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Hirbe A.C., Uluçkan O., Morgan E.A., Eagleton M.C., Prior J.L., Piwnica-Worms D., Trinkaus K., Apicelli A., Weilbaecher K. Granulocyte colony-stimulating factor enhances bone tumor growth in mice in an osteoclast-dependent manner. Blood. 2007;109:3424–3431. doi: 10.1182/blood-2006-09-048686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Bosteels C., Van Damme K.F.A., De Leeuw E., Declercq J., Maes B., Bosteels V., Hoste L., Naesens L., Debeuf N., Deckers J., et al. Loss of GM-CSF-dependent instruction of alveolar macrophages in COVID-19 provides a rationale for inhaled GM-CSF treatment. Cell Rep. Med. 2022;3 doi: 10.1016/j.xcrm.2022.100833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Holtschke T., Löhler J., Kanno Y., Fehr T., Giese N., Rosenbauer F., Lou J., Knobeloch K.P., Gabriele L., Waring J.F., et al. Immunodeficiency and chronic myelogenous leukemia-like syndrome in mice with a targeted mutation of the ICSBP gene. Cell. 1996;87:307–317. doi: 10.1016/s0092-8674(00)81348-3. [DOI] [PubMed] [Google Scholar]
- 82.Banik D., Netherby C.S., Bogner P.N., Abrams S.I. MMP3-mediated tumor progression is controlled transcriptionally by a novel IRF8-MMP3 interaction. Oncotarget. 2015;6:15164–15179. doi: 10.18632/oncotarget.3897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Husain M., Gusella G.L., Klotman M.E., Gelman I.H., Ross M.D., Schwartz E.J., Cara A., Klotman P.E. HIV-1 Nef induces proliferation and anchorage-independent growth in podocytes. J. Am. Soc. Nephrol. 2002;13:1806–1815. doi: 10.1097/01.asn.0000019642.55998.69. [DOI] [PubMed] [Google Scholar]
- 84.Dawson C.A., Pal B., Vaillant F., Gandolfo L.C., Liu Z., Bleriot C., Ginhoux F., Smyth G.K., Lindeman G.J., Mueller S.N., et al. Tissue-resident ductal macrophages survey the mammary epithelium and facilitate tissue remodelling. Nat. Cell Biol. 2020;22:546–558. doi: 10.1038/s41556-020-0505-0. [DOI] [PubMed] [Google Scholar]
- 85.Vannella K.M., Barron L., Borthwick L.A., Kindrachuk K.N., Narasimhan P.B., Hart K.M., Thompson R.W., White S., Cheever A.W., Ramalingam T.R., Wynn T.A. Incomplete deletion of IL-4Rα by LysM(Cre) reveals distinct subsets of M2 macrophages controlling inflammation and fibrosis in chronic schistosomiasis. PLoS Pathog. 2014;10 doi: 10.1371/journal.ppat.1004372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Basu G.D., Pathangey L.B., Tinder T.L., Lagioia M., Gendler S.J., Mukherjee P. Cyclooxygenase-2 inhibitor induces apoptosis in breast cancer cells in an in vivo model of spontaneous metastatic breast cancer. Mol. Cancer Res. 2004;2:632–642. [PubMed] [Google Scholar]
- 87.Tamura T., Thotakura P., Tanaka T.S., Ko M.S.H., Ozato K. Identification of target genes and a unique cis element regulated by IRF-8 in developing macrophages. Blood. 2005;106:1938–1947. doi: 10.1182/blood-2005-01-0080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Parker J.S., Mullins M., Cheang M.C.U., Leung S., Voduc D., Vickery T., Davies S., Fauron C., He X., Hu Z., et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J. Clin. Oncol. 2009;27:1160–1167. doi: 10.1200/jco.2008.18.1370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Ritchie M.E., Phipson B., Wu D., Hu Y., Law C.W., Shi W., Smyth G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47. doi: 10.1093/nar/gkv007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Subramanian A., Tamayo P., Mootha V.K., Mukherjee S., Ebert B.L., Gillette M.A., Paulovich A., Pomeroy S.L., Golub T.R., Lander E.S., Mesirov J.P. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA. 2005;102:15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Colligan S.H., Amitrano A.M., Zollo R.A., Peresie J., Kramer E.D., Morreale B., Barbi J., Singh P.K., Yu H., Wang J., et al. Inhibiting the biogenesis of myeloid-derived suppressor cells enhances immunotherapy efficacy against mammary tumor progression. J. Clin. Invest. 2022;132 doi: 10.1172/jci158661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Zhang X., Gonçalves R., Mosser D.M. The isolation and characterization of murine macrophages. Curr. Protoc. Immunol. 2008;14:14.1.1–14.1.14. doi: 10.1002/0471142735.im1401s83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Bardina S.V., Michlmayr D., Hoffman K.W., Obara C.J., Sum J., Charo I.F., Lu W., Pletnev A.G., Lim J.K. Differential Roles of Chemokines CCL2 and CCL7 in Monocytosis and Leukocyte Migration during West Nile Virus Infection. J. Immunol. 2015;195:4306–4318. doi: 10.4049/jimmunol.1500352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Lee C.C., Lin J.C., Hwang W.L., Kuo Y.J., Chen H.K., Tai S.K., Lin C.C., Yang M.H. Macrophage-secreted interleukin-35 regulates cancer cell plasticity to facilitate metastatic colonization. Nat. Commun. 2018;9:3763. doi: 10.1038/s41467-018-06268-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Gao J., Aksoy B.A., Dogrusoz U., Dresdner G., Gross B., Sumer S.O., Sun Y., Jacobsen A., Sinha R., Larsson E., et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013;6:pl1. doi: 10.1126/scisignal.2004088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Liberzon A., Birger C., Thorvaldsdóttir H., Ghandi M., Mesirov J.P., Tamayo P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1:417–425. doi: 10.1016/j.cels.2015.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Li B., Li T., Liu J.S., Liu X.S. Computational Deconvolution of Tumor-Infiltrating Immune Components with Bulk Tumor Gene Expression Data. Methods Mol. Biol. 2020;2120:249–262. doi: 10.1007/978-1-0716-0327-7_18. [DOI] [PubMed] [Google Scholar]
- 98.Qin Q., Fan J., Zheng R., Wan C., Mei S., Wu Q., Sun H., Brown M., Zhang J., Meyer C.A., Liu X.S. Lisa: inferring transcriptional regulators through integrative modeling of public chromatin accessibility and ChIP-seq data. Genome Biol. 2020;21:32. doi: 10.1186/s13059-020-1934-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data reported in this paper will be shared by the lead contact upon request. This paper does not report original code. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.