Highlights
-
•
Some biological responses could be restricted to some interferon subtypes.
-
•
Soluble IFNAR levels may be useful to predict disease severity or mortality risk.
-
•
IFNAR2 variants may be linked to disease severity due to its high biding affinity.
Keywords: COVID-19, Type I interferon, Single nucleotide polymorphisms
Abstract
Coronavirus Disease 2019 (COVID-19) is an infectious disease caused by SARS-CoV-2. According to the World Health Organization (WHO), there have been over 760 million reported cases and over 6 million deaths caused by this disease worldwide. The severity of COVID-19 is based on symptoms presented by the patient and is divided as asymptomatic, mild, moderate, severe, and critical. The manifestations are interconnected with genetic variations. The innate immunity is the quickest response mechanism of an organism against viruses. Type I interferon pathway plays a key role in antiviral responses due to viral replication inhibition in infected cells and adaptive immunity stimulation induced by interferon molecules. Thus, variants in type I interferon pathway's genes are being studied in different COVID-19 manifestations. This review summarizes the role of variants in type I interferon pathway's genes on prognosis and severity progression of COVID-19.
1. Introduction
Coronavirus Disease 2019 (COVID-19) is an infectious disease that originated in Wuhan, China, in December 2019 and rapidly spread to all continents, prompting the World Health Organization (WHO) to declare it a pandemic situation on March 11, 2020 (Azkur et al., 2020; Dhama et al., 2020; World Health Organization, 2023). Four years since its emergence, over 770 million cases and over 6 million deaths have been notified worldwide (World Health Organization, 2023). The causative agent of COVID-19 is identified as the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). While it shares a lineage with SARS-CoV, SARS-CoV-2 leads to a broader clinical spectrum of infection, encompassing asymptomatic infection; mild illness; moderate illness; severe illness; and critical illness (Dhama et al., 2020; National Institutes of Health, 2021).
Once the virus invades the organism, membrane-bound or intracellular pattern recognition receptors (PRRs) present in host cells are able to recognize it, triggering activation of interferon regulatory factors (IRFs) and nuclear factor kappa-beta (NF-κβ); culminating in the production of different signaling molecules known as interferon (IFN) (Matsuyama et al., 2020; Shah et al., 2020). There are different types of interferon, but the main molecules produced by virus-infected cells are the type I interferons (IFN-I) as neighboring cells can capture this alert signal through type I interferon receptor (IFNAR) and initiate an antiviral state to fight the infection (Solimani et al., 2021). Although this pathway has been implicated to generate protective responses against SARS-CoV-2, such as the release of interleukin 10 (IL-10) to limit excessive virus-dependent inflammatory lung lesions and the regulation of angiotensin converting enzyme 2 (ACE2) expression, the receptor responsible for SARS-CoV-2 cellular entry, high levels of IFN-I in critical COVID-19 patients has been associated with mortality, suggesting that a dysregulation of this pathway can favor pathogenesis (Matsuyama et al., 2020; Smith et al., 2022; Solimani et al., 2021; Sun et al., 2023).
As susceptibility or resistance to other infectious diseases have already been associated with host genetics and COVID-19 is a disease that manifests differently in each patient, the role of host genetics has been explored to better understand the variability of its possible clinical manifestations (Beck and Aksentijevich, 2020; Elhabyan et al., 2020; Murray et al., 2020; Ovsyannikova et al., 2020; Öztürk et al., 2020). Genome-wide studies can detect populational genetic variability, which are often a result of the modification of one base pair within a gene sequence, known as single nucleotide polymorphisms (SNPs) (Elhabyan et al., 2020; Murray et al., 2020; Öztürk et al., 2020). These studies are crucial for a better understanding of the pathophysiology of SARS-CoV-2 since it can identify regions of genetic variations that regulate complex traits or affect disease risk (Dehghan, 2018; Ovsyannikova et al., 2020).
In this study we reviewed IFN-I gene variants, as they have been suggested to play a role in the protective response against SARS-CoV-2 based on findings from small-scale studies (LoPresti et al., 2020, Van Der Made et al., 2020). We also identified the most studied gene candidates on this pathway. These genes should be further investigated when exploring the role of innate immune response against SARS-CoV-2.
2. COVID-19
2.1. Epidemiology
In the past two decades there were epidemics caused by other viral diseases that originated in the East, including the severe acute respiratory syndrome (SARS) in China in 2002, and the Middle East respiratory syndrome (MERS), in Saudi Arabia in 2012 (Boban, 2021; Dhama et al., 2020; Ovsyannikova et al., 2020; Sariol and Perlman, 2020; Shah et al., 2020). SARS-CoV-2, due to its higher transmissibility, asymptomatic transmission, and long infectious window before symptom onset, managed to have a broader scope of infection compared to MERS and SARS epidemics, but had a higher case fatality rate (CFR) of approximately 35 % and ∼10 %, respectively, compared to COVID-19 (∼2 %) (Azkur et al., 2020; Boban, 2021; Dhama et al., 2020; Sariol and Perlman, 2020).
Several factors have been identified as associated with a higher risk of severe COVID-19, including male gender, blood type A, old age and comorbidities (Boban, 2021; Dhama et al., 2020; Funk et al., 2020; National Institutes of Health, 2021; Ovsyannikova et al., 2020). Hypertension, obesity, diabetes, lipid metabolism disorders, and coronary artery disease are among the most common comorbidities reported, while other comorbidities such as chronic obstructive pulmonary disease and cancer have also been observed in severely ill patients (Boban, 2021; Kompaniyets et al., 2021; Ovsyannikova et al., 2020). The absence of comorbidities may lead to a CFR as low as 0.9 %, while the presence of some comorbidities, such as cardiovascular diseases, diabetes, chronic lung disease, hypertension, cancer, anxiety and fear-related disorders, chronic kidney disease, and neurocognitive disorders may be linked to critical illness, leading to a CFR as high as 49 % (Kompaniyets et al., 2021; Öztürk et al., 2020). Furthermore, the development of acute respiratory distress syndrome (ARDS) has been associated with a higher risk of death in patients with COVID-19 pneumonia. This association is influenced by factor such as old age, presence of comorbidities, high fever, neutrophilia, lymphopenia, elevated high-sensitivity C-reactive protein, and coagulation function-related indicators, such as prothrombin time and d-dimer (Wu et al., 2020).
2.2. Clinical manifestations
Fever, cough, nasal congestion, headaches, myalgia, tiredness, and dyspnea are common symptoms related to COVID-19. However, the clinical manifestations of this disease depends on the infected host's ability to have an immune response against the virus (Boban, 2021; Carfì et al., 2020; Dhama et al., 2020; Funk et al., 2020; Kaur and Gupta, 2020; Öztürk et al., 2020; Shah et al., 2020). Consequently, less common symptoms such as anosmia, dysgeusia, loss of appetite, intestinal discomfort, diarrhea, articular pain, dizziness or malaise, and chills have also been reported in these patients (Boban, 2021; Funk et al., 2020; Öztürk et al., 2020; Tay et al., 2020).
The United States National Institute of Health (NIH) has categorized SARS-CoV-2 clinical spectrum as asymptomatic infection, mild illness, moderate illness, severe illness, and critical illness. Table 1 shows the definitions for each category within the clinical spectrum of this disease.
Table 1.
Clinical spectrum of COVID-19 according to NIH treatment guidelines.
| Severity of illness | Symptoms | Observations |
|---|---|---|
| Asymptomatic or Pre symptomatic | No symptoms consistent with COVID-19. | Positive for virologic tests (NAAT or antigen test). |
| Mild Illness | Fever, cough, sore throat, malaise, headache, muscle pain, nausea, vomiting, diarrhea, loss of taste and smell. | No imaging or specific laboratory evaluations are routinely indicated in otherwise healthy patients. Patients with underlying comorbidities are at higher risk and should be closely monitored until recovery. |
| Moderate Illness | Decreased oxygen saturation (SpO2 ≥ 94 %) on room air at sea level. Shortness of breath or dyspnea. | Clinical assessment showing imaging evidence of lower respiratory disease. |
| Severe Illness | SpO2 〈 94 % on room air at sea level, PaO2/FiO2 < 300 mmHg, respiratory rate 〉 30 breaths/min. or lung infiltrates > 50 %. | Patients may experience rapid clinical deterioration. Immediate administration of oxygen therapy using a nasal cannula or a high-flow oxygen device. |
| Critical Illness | Respiratory failure, septic shock, and/or multiple organ dysfunction. | In addition to pulmonary disease, these patients may experience cardiac, hepatic, renal, central nervous system or thrombotic disease. |
NAAT: Nucleic Acid Amplification Test; PaO2/FiO2: Ratio of arterial partial pressure of oxygen to fraction of inspired oxygen.
As COVID-19 patients recovered, a subset has experienced persistent symptoms lasting for weeks past primary viral infection, giving rise to a post-infectious syndrome known as Long COVID or post-acute sequelae of COVID-19 (Pagaduan and Altawallbeh, 2020; Phetsouphanh et al., 2022; Rio et al., 2020). This multisystemic condition often includes severe symptoms persisting for at least two months, following a history of probable or confirmed SARS-CoV-2 infection without an alternative diagnosis (Davis et al., 2023; Li et al., 2023; Nalbandian et al., 2023). While estimates suggests that at least 65 million individuals worldwide may be affected by Long COVID, the actual number is difficult to ascertain, and it is likely higher due to undocumented cases (Davis et al., 2023; Nalbandian et al., 2023). Some common manifestations are similar to the acute disease, such as fatigue, shortness of breath, and a dysfunctional sense of smell, while others are different, such as neurocognitive impairment, autonomic dysfunction, and psychiatric ailments such as anxiety and depression (Davis et al., 2023; Li et al., 2023; Nalbandian et al., 2023; Shaffer, 2022). Obese and older patients have a higher risk of developing Long COVID and this may be due to their association with acute COVID-19 severity, since severely ill patients are more likely to exhibit persistent symptoms. However, this syndrome is not restricted to this population alone, as young and healthy individuals have also been subject to the long-lasting effects of COVID-19 (Davis et al., 2023; Nalbandian et al., 2023).
SARS-CoV-2 variants may play a role in the progression of long COVID, with data from studies suggesting that Omicron infection is associated with fewer clinical manifestations of this post-infectious syndrome. However, the increased transmissibility and relatively low morbidity of this variant have the potential to expose more people to acute COVID-19 and the risk of progressing to persistent symptoms (Fernández-de-las-Peñas et al., 2022; Li et al., 2023; Torjesen, 2021). Conversely, vaccination may prevent long COVID, since they can prevent acute COVID-19 or severe illness progression. A study in Italy showed a correlation between the number of vaccine doses and the prevalence of long COVID, demonstrating that unvaccinated individuals were the most prevalent, while those with three vaccine doses were the least prevalent (Azzolini et al., 2022; Li et al., 2023). Although vaccination may reduce the risk of long COVID, it can also cause adverse effects since excessive immune response during vaccination may lead to a higher antibody titer, which has been associated with worse sequelae (Li et al., 2023; Tsuchida et al., 2022).
3. Type I interferon pathway
The human organism's initial defense line, comprising epithelial cells and innate immune cells, expresses PRRs to identify damage-associated molecular patterns (DAMPs) or pathogen-associated molecular patterns (PAMPs), recognizing viruses upon entry into host cells. PRRs fall into three classes responsible for intracellular activation of the innate immune system: Toll-like receptors (TLR), RIG-1-like receptors (RLR), and NOD-like receptors (NLR). Although the genome of SARS-CoV-2 is composed of unsegmented single-stranded RNA, receptors that recognize double-stranded RNA (dsRNA) can also participate in viral recognition due to intermediate dsRNA formation during replication (Dhama et al., 2020; Kaur and Gupta, 2020; Lu et al., 2020; Schreiber, 2020; Yin et al., 2021). Toll-like receptors (TLR) 3 and 7 contribute significantly to the detection of SARS-CoV-2. However, Melanoma Differentiation-Associated protein 5 (MDA5), a member of the RIG-I-like receptor family, has been identified as a mediator of the protective immune response against the virus. MDA5 facilitates the induction of Type I interferons (IFN-I) through its interaction with the mitochondrial antiviral-signaling protein (MAVS) adaptor. Notably, this process occurs without stimulating the production of pro-inflammatory cytokines, which, when elevated, can exacerbate disease severity (Ramasamy and Subbian, 2021; Sampaio et al., 2021; Yin et al., 2021). When these receptors recognize the virus, dimers of interferon regulating factors (IRF3/IRF7) are formed to activate the production of IFN-I, signaling the presence of a viral infection to neighboring cells (Balkhi, 2021; Carter-Timofte et al., 2020; Mazewski et al., 2020; Ramasamy and Subbian, 2021; Schreiber, 2020).
The IFNAR, located in the membrane of neighboring cells, detects the IFN-I signal, and their interaction results in the activation of the intracellularly associated proteins of the Janus kinase family (JAK2 and TYK2) through cross-phosphorylation, which also phosphorylates proteins from the signal transducer and activator of transcription family (STAT) associated with the receptor (Bagheri et al., 2021; Carter-Timofte et al., 2020; Mazewski et al., 2020; Schreiber, 2020). Consequently, STAT proteins dissociate from the receptor and form homo- and heterodimers that are transported to the nucleus to function as transcription factors for the expression of interferon-stimulated genes (ISGs). These ISGs possess antiviral activities to protect these cells against the infection (Fig. 1) (Bagheri et al., 2021; Matsuyama et al., 2020; Mazewski et al., 2020; Schreiber, 2020).
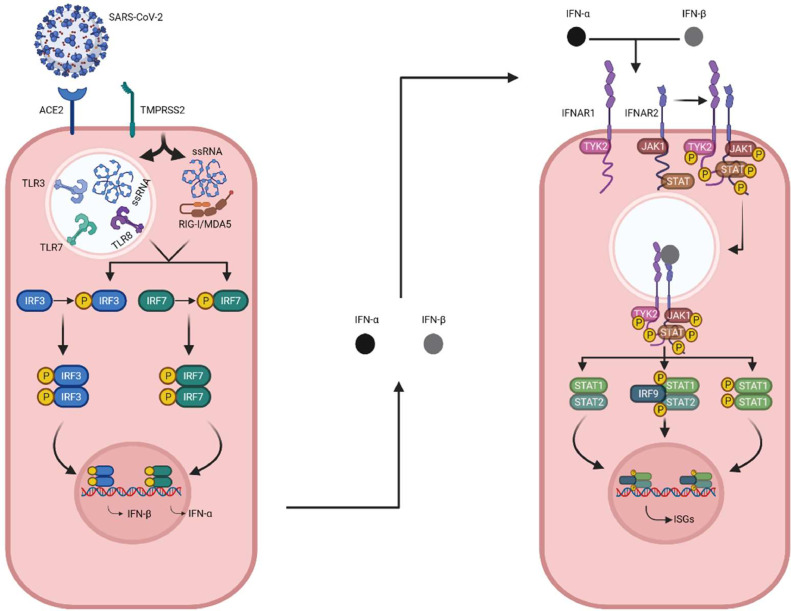
Fig. 1.
Type I interferon pathway activation by SARS-CoV-2 infection. SARS-CoV-2 enters the host epithelial alveolar cells through the ACE2 protein located at the cell surface, with the assistance of TMPRSS2 protein. The cell membrane fuses with the viral envelope and the single-stranded virus RNA enters the cell in an endosome. TLR3, TLR7, and TLR8 are endomosomaly expressed and can detect viral RNA. RIG-I and MDA5 are located at the cytoplasm but can also detect viral RNA if it reaches this location without detection. The expression of interferon regulatory factors (IRF3 and IRF7) is stimulated after detection by PRRs. IRF3 and IRF7 are phosphorylated and homodimerized to serve as transcription factors to produce IFN-α and IFN-β. These type I interferon (IFN-I) molecules are then detected by type I interferon receptor subunits (IFNAR1 and IFNAR2), which have JAK and STAT proteins that bind to their intracellular domain (ICD). The JAK (TYK2 and JAK1) and STAT proteins are responsible for signal transduction. TYK2 binds to IFNAR1 ICD and JAK1 and STAT1 bind to IFNAR2 ICD. When the IFNAR subunits are bind to IFN-I, these proteins cross-phosphorylate and internalize the receptor through endosome formation. STAT proteins form homo- and heterodimers such as the IRF9/STAT1/STAT2 complex. The resulting complexes serve as transcription factors for interferon stimulated genes (ISGs), which have several antiviral functions (Figure created with Biorender, adapted from Schreiber 2020; Carter-Timofte et al., 2020).
IFN-I play a crucial role in activating an antiviral state in neighboring cells to limit virus spread. However, these molecules are highly active and cannot be limited to that, being also responsible for the modulation of innate immune system response, such as natural killer (NK) cell functions and antigen presentation. Simultaneously, IFN-I restrains pro-inflammatory pathways and activates adaptive immune system, generating high-affinity antigen-specific B and T cell responses (Schreiber, 2020). To better understand the different functions of the IFN-I family, it is essential to know that it is composed of multi-genes encoding 13 highly homologous IFN-α (IFN-α1, -α2, -α4, -α5, -α6, -α7, -α8, -α10, -α13, -α14, -α16, -α17, and -α21) and genes encoding lower homology single IFN-I subtypes (IFN-ω, -κ, -ϵ, -β) (da Silva et al., 2021; Mazewski et al., 2020; Schreiber, 2020, 2017). The differences among IFN-I subtypes affect their receptor binding affinity, which is suggested to impact the biological responses that these molecules stimulate, with greater affinity related to a more intense signal (Schreiber, 2020, 2017). Therefore, complex biological responses, such as antiproliferative (proapoptotic and antitumor) activity and immune-modulation (modulation of antigen presentation and natural killer cell functions), can be induced by IFN-I, but seem to be restricted for some types with stronger affinity to the receptor, like IFN-β, for instance. Common IFN-I responses, including antiviral cellular response, can be induced by any type of IFN-I, even weak binding ones, like IFN-ϵ (Fox et al., 2020; Mazewski et al., 2020; Schreiber, 2020, 2017).
Although greater receptor affinity contributes to the activation of complex responses, other factors also influence their trigger, thus not limiting them to the strongest affinity IFN-I, as high concentrations of not so strong-affinity IFN-I can also activate these responses (Fox et al., 2020; Schreiber, 2020, 2017). Additionally, cell types play a role in biological response activation, since all cells have access to common IFN-I responses, but only some cells can activate antiproliferative and immune-modulation responses (Schreiber, 2020, 2017). Differences in interferon subtypes affinity is a vast subject, leaving room for much more discoveries about the responses they can generate; therefore, future studies should widely explore this field (Mazewski et al., 2020).
Type I interferon receptor (IFNAR) comprises two transmembrane subunits, IFNAR1 and IFNAR2, each containing two domains: an extracellular domain (ECD), which binds to IFN-I, and an intracellular domain (ICD), which binds to the JAK and STAT proteins (Akter et al., 2021; Schreiber, 2020; Thomas et al., 2012). While JAK1, STAT1, STAT2, and possibly other STAT proteins bind to the ICD of IFNAR2, only TYK2 is bound to the ICD of IFNAR1 (Akter et al., 2021; Schreiber, 2020; Thomas et al., 2012). These subunits have distinct structures, with IFNAR1 optimized for functional plasticity, causing conformational changes that result in energetic loss and consequently lower binding affinity, while IFNAR2 is optimized for high binding affinity to IFN-I (Schreiber, 2020; Thomas et al., 2012).
3.1. IFN-I impairment mechanisms
As host immunity restrains viral persistence, the virus is pressured to adapt, leading to mechanisms to evade host immunity (Zhang et al., 2022). In COVID-19, early IFN-I production is key for viral suppression, making it a crucial target for these viral evasion mechanisms (Smith et al., 2022). Consequently, SARS-CoV-2 may inhibit IFN-I production through evasion of PRR detection or interference in IRF dimer formation, or hinder IFN-I signaling through disruption of STAT phosphorylation or obstruction of the transport of antiviral transcription factors to cell nucleus (Rubio-Casillas et al., 2022; Zhang et al., 2022). For instance, MDA5-mediated sensing of coronavirus RNA has been implicated as a significant inductor of IFN-I responses. However, these viruses have developed genomic methylation mechanisms to mimic host mRNA and evade recognition (Zhang et al., 2022; Züst et al., 2011). Moreover, dysregulation of the IFN-I has been suggested as pathogenic during SARS-CoV-2 infection since highly impaired IFN-I response have been associated with exacerbated inflammatory response in severe COVID-19 patients, while increased IFN-I levels have been associated with mortality, which emphasizes the importance of the timing of pathway activation (Hadjadj et al., 2020; Smith et al., 2022; Sun et al., 2023).
IFN-I responses can also be impaired by autoimmunity. Although not consistently associated with susceptibility to viral diseases, the presence of IFN-I autoantibodies has been observed in critical COVID-19 patients with pneumonia (Bastard et al., 2020; Casanova and Anderson, 2023; Manry et al., 2022). Bastard et al. (2020) suggest IFN-I autoimmunity to be a cause of severe COVID-19 rather than a consequence of SARS-CoV-2 infection, even though it may intensify its effects. This can be acquired through autoimmune polyglandular syndrome type I (APS-I), systemic lupus erythematosus, and previous IFN-I treatment (Casanova and Anderson, 2023). Manry et al. (2022) associated the presence of these autoantibodies, especially anti-IFN-α2 and anti-IFN-ω, with high COVID-19 mortality risk. They observed that anti-IFN-I antibodies have a higher impact in the mortality risk of younger patients compared to older patients, even though IFN-I autoimmunity occurrence increases with age. This emphasizes the potential of IFN-I autoantibody detection for COVID-19 severity progression in all age groups, suggesting that a risk prediction screening test could be elaborated through the association of these antibodies with patient age (Casanova and Anderson, 2023; Manry et al., 2022). Casanova and Anderson (2023) suggest that IFN-I autoantibody detection could also assist in patient treatment correction, raising the hypothesis of enhancement of early antiviral responses through IFN-β administration in these patients, as autoimmunity to this subtype is rare.
3.2. Soluble IFNARs
Although certain mechanisms can impair IFN-I responses during viral infections a reaction can be set by the host through the modulation of the activity of this cytokine. Soluble isoforms of cytokine receptors are fundamental for modulating their cytokine activity, and they can be produced by protein modification processes, which can occur before translation, through alternative splicing, or after translation, through proteolytic cleavage (Han et al., 2001; Hurtado-Guerrero et al., 2020). Therefore, a potential role for soluble IFNAR (sIFNAR) isoforms has been proposed to protect this cytokine from degradation or excretion, even though it was initially proposed to antagonize IFN-I (Han et al., 2001; Hurtado-Guerrero et al., 2020). Fricke-Galindo et al. (2022) observed that COVID-19 survivors exhibited higher sIFNAR2 levels compared to non-survivors while investigating an association between plasmatic levels of sIFNAR2 and clinical outcome in severe COVID-19 patients in Mexico. The results indicate that increased sIFNAR2 production during severe disease could assist in recovery through IFN-I stability improvement, confirming its role in cytokine protection. Additionally, Yaugel-Novoa et al. (2023) also noted increased sIFNAR2 concentrations in severe patients while comparing serum levels of IFNAR1 and IFNAR2 in different COVID-19 patients in France, suggesting that a decreased IFN-I sensitivity, which may be a result of viral mechanisms for host immune evasion or acquired IFN-I autoimmunity found in severe patients can increase sIFNAR2 production. Collectively, these studies suggest that sIFNAR2 may counteract mechanisms that inhibit type I interferon responses, thereby enhancing antiviral defenses. Although there is an implication that sIFNAR2 levels could serve as indicators of COVID-19 severity or mortality likelihood, further research is necessary to substantiate its predictive value for disease progression.
4. Host genetics in COVID-19
4.1. Single nucleotide polymorphisms
In the context of the broad clinical spectrum observed in COVID-19, researchers think that genetic differences may be linked to inborn variations in critical cell signaling pathway genes, which may interfere with antiviral responses, thereby impacting disease progression, as previous studies already evidenced the influence of genetic variation in other infectious diseases (Casanova and Abel, 2018; Elhabyan et al., 2020; Ovsyannikova et al., 2020; Öztürk et al., 2020). One notable example is the CCR5∆32, a loss-of-function mutation in a chemokine receptor-encoding gene crucial for HIV entry in the host cell that results in revoked protein expression and provides resistance to homozygous individuals, while heterozygous individuals experience delayed disease progression (Gómez et al., 2020; Ovsyannikova et al., 2020; Öztürk et al., 2020). Single nucleotide polymorphisms represent the most common genetic variation in humans and consist of a single change in a nucleotide within a genetic sequence that can modify protein formation through altered amino acid codification, influence the gene expression, and translation efficiency processes, and affect messenger RNA stability (Shastry, 2009).
4.2. Type I interferon pathway genes and their variants
As the IFN-I pathway plays an important role in antiviral protection, alteration in genes acting in this pathway can modify the efficiency of the host's immune response to the virus, making it more or less efficient (Beck and Aksentijevich, 2020). Several studies have focused on the receptor-ligand (IFNAR-IFN) interaction, exploring how altered receptor affinity or expression may generate different timing of antiviral responses. Therefore, numerous IFNAR subunit-encoding gene variants have been identified, with a particular focus on their potential associations with COVID-19 severity (Akter et al., 2021; Dieter et al., 2023; Fricke-Galindo et al., 2022; Ma et al., 2021; Pairo-Castineira et al., 2020; Schmiedel et al., 2021).
Akter et al. (2021) carried out in silico comparative analyses of different conformational protein structures for the IFNAR2, analyzing the difference in interferon binding affinity. P136R and P136S protein structures, identified as rs768348126 and rs746695388 respectively, showed lower interferon binding affinity. Their in silico analysis suggested that P136R expression is highly pathogenic, making it a good target for further investigating, especially in severely ill patients. The same study performed in silico analysis for IFNAR2 intronic variants, which can modify the regulation of their target genes either by affecting their splicing mechanism or changing its expression. Three variants (rs2250226, rs2300371, and rs6517156) reduced the expression of IFNAR2 and five variants (rs2252650, rs2284550, rs2834158, rs2834164, and rs2834165) affected the splicing mechanism of this gene, resulting in a higher risk of severe COVID-19. Pairo-Castineira et al. (2020) explored different genetic variants to understand their potential role in critical COVID-19, highlighting the presence of the A allele in an IFNAR2 variant, rs2236757, as a risk factor. Supposedly, the presence of this allele can impact gene expression and reduce the presence of this receptor in host cells rendering the patient susceptible to disease progression, as increased expression of IFNAR2 has shown reduced odds of severe COVID-19.
In the Brazilian population, Dieter et al. (2023) analyzed the same IFNAR2 variant, rs2236757, comparing genotypes among critical unvaccinated COVID-19 patients and non-critical unvaccinated COVID-19 patients. After stratification by ethnicity and statistical adjustment for sex and age, the A/A genotype exhibited association not only to Intensive Care Unit (ICU) admission risk, as observed previously by Pairo-Castineira et al. (2020), but also to mortality risk, since non-survivors showed higher frequency of the A allele and A/A genotype compared to survivors. This emphasizes the importance of the rs2236757 A allele in COVID-19 severity and outcome. In Mexico, Fricke-Galindo et al. (2022) obtained similar results regarding rs2236757, associating the A/A genotype to mortality risk in severe COVID-19 patients. They analyzed different IFNAR2 variants in hospitalized patients, and observed that mortality risk was also associated with further IFNAR2 variants, including a missense variant (rs1051393) that may affect IFNAR2 membrane mobility through an amino acid change (phenylalanine to valine). In Palestine, Abdelhafez et al. (2023) analyzed household members and found that patients with the rs2236757 A allele had a higher probability of developing dyspnea while also associating the presence this allele with critical COVID-19.
Schmiedel et al. (2021) identified an IFNAR2 variant (rs12482556) capable of influencing the expression of a neighboring gene, IL10RB, in natural killer cells and T-cell subsets through the interaction with its promoter. IL10RB expression could affect IL-10 cell responsiveness, given that this gene is responsible for IL-10 receptor beta production, contributing to IL-10 immunomodulation. Therefore, the authors suggest that an increased expression of this gene might play a role in COVID-19 severity by enhancing IL-10 cell responsiveness. Similarly, Ma et al. (2021) proposed a role for IL10RB gene expression modification in SARS-CoV-2 infection by also identifying an IFNAR2 variant (rs9976829) that regulates IL10RB alternative splicing, consequently being a factor for its expression in different tissue types, particularly in the lungs.
Although IFNAR1 alterations may interfere in IFN-I pathway function and its genetic variants have been linked to other viral diseases, such as rs2257167 in susceptibility to hepatitis B, they have not been implicated in the occurrence of severe COVID-19 as IFNAR2 (de Weerd et al., 2020; Zhang et al., 2020). Abolhassani et al. (2022) reported a case of critical COVID-19 pneumonia in a child with autosomal recessive IFNAR1 deficiency, suggesting a possible association of receptor subunit deficiency with critical disease. On the other hand, genetic variation of signal transduction protein encoding genes, such as TYK2, may interfere in pathway signal uptake, making them potential targets as COVID-19 severity risk factors. A variant of this gene, rs2304256, could not be associated to disease outcome alone, but a Brazilian study found that the interaction among TYK2, TMPRSS2, and IFIH1 polymorphisms was associated with mortality risk when four or more minor alleles were present (Dieter et al., 2023).
The responses generated by the IFN-I pathway, including the antiviral response, occur through the expression of interferon stimulated genes (ISGs), which are defined as any gene induced during an interferon response of any kind. The impact of their variation is being studied in the context of COVID-19 severity progression (Alghamdi et al., 2020; Kim and Jeong, 2021; Schoggins, 2019). For instance, IFITM3, an ISG crucial in antiviral responses through the inhibition of viral entry on host cells and its rs12252 and rs6598045 variants were related to severe illness progression in Arabic and Chinese population (Alghamdi et al., 2020; Kim and Jeong, 2021). The OAS gene cluster, which encodes proteins responsible for activation of latent RNAse L that results in viral RNA degradation and consequent inhibition of viral replication, has also been suggested to be associated with critical COVID-19 (Pairo-Castineira et al., 2020; Schmiedel et al., 2021; Zhou et al., 2021). Schmiedel et al. (2021) associated the presence of the OAS1 variant (rs4767032) and the OAS3 variant (rs1298962) in severe COVID-19 patients with reduced protein expression in non-classical monocytes and T-cell subsets, respectively, suggesting a potential role for reduced expression of this protein in the impairment of viral RNA degradation in COVID-19 severity progression. Similarly, Zhou et al. (2021) noted that the presence of the rs10774671 OAS1 variant G allele has a role in the alternative splicing and production of a protein isoform with enhanced enzymatic activity against viruses, potentially contributing to COVID-19 protection, as increased OAS1 levels during non-infectious state is strongly associated with reduced risk of hospitalization and disease severity. Although increased OAS3 expression has been associated with critical disease and worse outcomes in SARS-CoV-2 infection, Abdelhafez et al. (2023) could not associate the presence of rs10735079, a OAS3 variant, with disease severity, illustrating the demand for further knowledge about the complex part that OAS plays in this disease (Pairo-Castineira et al., 2020; Zhou et al., 2021).
Ziegler et al. (2020) demonstrated that ACE2 can be classified as an ISG and suggested that SARS-CoV-2 might exploit tissue-protection responses mediated by ACE2 to increase virus cellular entry (Ziegler et al., 2020). Recent studies indicate that interferons (IFNs) might prompt the production of a variant of the angiotensin-converting enzyme 2, termed deltaACE2 (dACE2), which seems to be deficient in its ability to adhere to SARS-CoV-2′s spike protein. (Oliveto et al., 2022; Onabajo et al., 2020). This suggests deltaACE2 may not promote increased viral entry into host cells. While ACE2 serves as the entry receptor for SARS-CoV-2, it also plays a critical role in modulating blood pressure and fluid-electrolyte balance by interacting with angiotensin II. The binding of SARS-CoV-2 to ACE2 can diminish its availability for angiotensin II regulation, potentially leading to elevated angiotensin II levels. (Onabajo et al., 2020). High plasmatic levels of angiotensin II have been associated with ARDS, lung damage and high mortality in coronavirus infection in mouse models. (Imai et al., 2005; Kuba et al., 2005; Onabajo et al., 2020). This association has prompted investigations into ACE2 deficiency as a potential contributing factor to the morbidity observed in COVID-19 cases. Onabajo et al. (2020) proposed that dACE2 isoform could be beneficial for COVID-19 patients through the preservation of ACE2 protein levels, making it still accessible for the renin-angiotensin-aldosterone system function. The type I interferon pathway participating genes are detailed in Table 2.
Table 2.
Type I interferon pathway genes studied to understand the link between this pathway and COVID-19 severity, susceptibility, or resistance.
| Gene | Protein | Function | Action in COVID-19 | Variants | Refs. |
|---|---|---|---|---|---|
| IFNAR2 | Type I interferon receptor subunit 2. | High-affinity binding to type I interferon. | High-affinity binding to type I interferon. | rs9976829; rs1332014803; rs751841284; rs1568885326; rs1190725243; rs758825586; rs1286287301; rs147496374; rs773793948; rs772583115; rs1265189745; rs768348126; rs746695388; rs2250226; rs2300371; rs6517156; rs2252650; rs2284550; rs2834158; rs2834164; rs2834165; rs2236757; rs12482556; rs3153; rs2834158; rs1051393 | Ma et al. (2021), Akter et al. (2021), Pairo-Castineira et al. (2020), Schmiedel et al. (2021), Fricke-Galindo et al. (2022), Dieter et al. (2023) |
| IFIH1 | Melanoma Differentiation-Associated protein 5 (MDA5), a RIG-1-like receptor. | Sensing RNA length and secondary structure leading to a proinflammatory response that includes interferons. | Senses intracellular viral RNA triggers innate immune response and assists in cell-mediated and humoral immunity. | rs1990760 | Maiti (2020), Dieter et al. (2023) |
| IFITM3 | Interferon-induced transmembrane protein 3 | Restricts cellular entry by diverse viral pathogens. | Block the fusion pore formation between the virus and host membrane. | rs12252 rs6598045 | Alghamdi et al. (2020), Kim and Jeong (2021) |
| OAS1 | 2′−5′-oligoadenylate synthetase 1 | Degrades viral RNA by activating latent RNase L. | Inhibits viral replication. | rs10774671 rs4767032 | Zhou et al. (2021), Schmiedel et al. (2021) |
| OAS3 | 2′−5′-oligoadenylate synthetase 3 | rs1298962 rs10735079 | Schmiedel et al. (2021), Abdelhafez et al. (2023) | ||
| TYK2 | Tyrosine Kinase 2 | This protein associates with the ICD of IFNAR1 and transduces signal by phosphorylating receptor subunits. | Signal transduction. | rs74956615 rs2304256 | Pairo-Castineira et al. (2020), Solimani et al. (2021), Dieter et al. (2023) |
IFIH1: Interferon Induced With Helicase C Domain 1.
5. Future perspectives
There is still a lot to learn about the reasons underlying the different clinical manifestations of COVID-19. Populational studies are crucial to clarify which genes are essential for disease severity. These genes, once identified, could function as severity biomarkers. Therefore, studies aiming at identifying severity biomarkers for COVID-19 must include different races and global regions to investigate genetic variations of target genes. A study performed only in an European population may not identify the same biomarkers as a study performed only in an African population, for instance. Once these severity biomarkers are identified for several populations worldwide, it might be possible to differ the high genetic risk patients, resulting in proper care, and the reduction of deaths. Notably, insights into the origin of genetic variants in innate immunity genes may provide its populational distribution and assist in the identification of protected or risk populations. For instance, some OAS gene cluster variants originated in archaic hominins like Neanderthals and Denisovans, have a higher probability of being inherited by specific populations and although they could be advantageous in infection immune response at that time, they could be disadvantageous in present infections (Deschamps et al., 2016; Zeberg and Pääbo, 2021; Zhou et al., 2021).
The type I interferon pathway has a protective role against SARS-CoV-2 and variants in this pathway and disruption of its function can be related to disease severity. Few studies indicate that a delayed activation of this pathway may exacerbate inflammation, contributing to progression of disease severity and increased mortality (Lee and Shin, 2020; Schreiber, 2020). Comprehensive studies are necessary to understand the participation of this pathway in disease progression and virus response.
CRediT authorship contribution statement
A.F.L. Montenegro: Conceptualization, Investigation, Visualization, Writing – original draft. M.A.F. Clementino: Supervision, Writing – review & editing. J.N.U. Yaochite: Supervision, Project administration, Funding acquisition, Writing – review & editing.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgment
This study was funded by JBS S.A. through the “Fazer o Bem Faz Bem” Program and by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES).
Data availability
No data was used for the research described in the article.
References
- Abdelhafez M., Nasereddin A., Shamma O.A., Abed R., Sinnokrot R., Marof O., Heif T., Erekat Z., Al-Jawabreh A., Ereqat S. Association of IFNAR2 rs2236757 and OAS3 rs10735079 polymorphisms with susceptibility to COVID-19 infection and severity in palestine. Interdiscip. Perspect. Infect. Dis. 2023;11 doi: 10.1155/2023/9551163. 2023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abolhassani H., Landegren N., Bastard P., Materna M., Modaresi M., Du L., Aranda-Guillén M., Sardh F., Zuo F., Zhang P., Marcotte H., Marr N., Khan T., Ata M., Al-Ali F., Pescarmona R., Belot A., Béziat V., Zhang Q., Casanova J.L., Kämpe O., Zhang S.Y., Hammarström L., Pan-Hammarström Q. Inherited IFNAR1 deficiency in a child with both critical COVID-19 pneumonia and multisystem inflammatory syndrome. J. Clin. Immunol. 2022;42:471–483. doi: 10.1007/s10875-022-01215-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akter S., Roy A.S., Tonmoy M.I.Q., Islam M.S. Deleterious single nucleotide polymorphisms (SNPs) of human IFNAR2 gene facilitate COVID-19 severity in patients: a comprehensive in silico approach. J. Biomol. Struct. Dyn. 2021;0:1–17. doi: 10.1080/07391102.2021.1957714. [DOI] [PubMed] [Google Scholar]
- Alghamdi J., Alaamery M., Barhoumi T., Rashid M., Alajmi H. Interferon-induced transmembrane protein-3 genetic variant rs12252 is associated with COVID-19 mortality. Genomics. 2020;113:1733–1741. doi: 10.1016/j.ygeno.2021.04.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Azkur A.K., Akdis M., Azkur D., Sokolowska M., van de Veen W., Brüggen M.C., O'Mahony L., Gao Y., Nadeau K., Akdis C.A. Immune response to SARS-CoV-2 and mechanisms of immunopathological changes in COVID-19. Allergy Eur. J. Allergy Clin. Immunol. 2020;75:1564–1581. doi: 10.1111/all.14364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Azzolini E., Levi R., Sarti R., Pozzi C., Mollura M., Mantovani A., Rescigno M. Association between BNT162b2 vaccination and long COVID after infections not requiring hospitalization in health care workers. JAMA. 2022;328:676–678. doi: 10.1001/jama.2022.11691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bagheri A., Mohammad S., Moezzi I., Mosaddeghi P. International immunopharmacology interferon-inducer antivirals: potential candidates to combat COVID-19. Int. Immunopharmacol. 2021;91:1–17. doi: 10.1016/j.intimp.2020.107245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balkhi M.Y. Mechanistic understanding of innate and adaptive immune responses in SARS-CoV-2 infection. Mol. Immunol. 2021;135:268–275. doi: 10.1016/j.molimm.2021.04.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bastard P., Rosen L.B., Zhang Q., Michailidis E., Hoffmann H.H., Zhang Y., Dorgham K. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science. 2020;(80):370. doi: 10.1126/science.abd4585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beck D.B., Aksentijevich I. Susceptibility to severe COVID-19. Science. 2020;370(80):404–405. doi: 10.1126/science.abe7591. [DOI] [PubMed] [Google Scholar]
- Boban M. Novel coronavirus disease (COVID-19) update on epidemiology, pathogenicity, clinical course and treatments. Int. J. Clin. Pract. 2021;75:1–7. doi: 10.1111/ijcp.13868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carfì A., Bernabei R., Landi F. Persistent symptoms in patients after acute COVID-19. JAMA. 2020;369:603–605. doi: 10.1136/bmj.m1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carter-Timofte M.E., Jørgensen S.E., Freytag M.R., Thomsen M.M., Brinck Andersen N.S., Al-Mousawi A., Hait A.S., Mogensen T.H. Deciphering the role of host genetics in susceptibility to severe COVID-19. Front. Immunol. 2020;11:1–14. doi: 10.3389/fimmu.2020.01606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Casanova J., Abel L. Human genetics of infectious diseases: unique insights into immunological redundancy. Semin. Immunol. 2018;36:1–12. doi: 10.1016/j.smim.2017.12.008.Human. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Casanova J.L., Anderson M.S. Unlocking life-threatening COVID-19 through two types of inborn errors of type I IFNs. J. Clin. Invest. 2023;133:1–11. doi: 10.1172/JCI166283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- da Silva R.P., Gonçalves J.I.B., Zanin R.F., Schuch F.B., de Souza A.P.D. Circulating type I interferon levels and COVID-19 severity: a systematic review and meta-analysis. Front. Immunol. 2021;12:1–11. doi: 10.3389/fimmu.2021.657363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis H.E., McCorkell L., Vogel J.M., Topol E.J. Long COVID: major findings, mechanisms and recommendations. Nat. Rev. Microbiol. 2023;21:133–146. doi: 10.1038/s41579-022-00846-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Weerd N.A., Vivian J.P., Lim S.S., Huang S.U.S., Hertzog P.J. Structural integrity with functional plasticity: what type I IFN receptor polymorphisms reveal. J. Leukoc. Biol. 2020;108:909–924. doi: 10.1002/JLB.2MR0420-152R. [DOI] [PubMed] [Google Scholar]
- Dehghan A. Genetic Epidemiology: Methods and Protocols. Evangelos Evangelou; 2018. Genome-wide association studies; pp. 37–48. [Google Scholar]
- Deschamps M., Laval G., Fagny M., Itan Y., Abel L., Casanova J.L., Patin E., Quintana-Murci L. Genomic signatures of selective pressures and introgression from archaic hominins at human innate immunity genes. Am. J. Hum. Genet. 2016;98:5–21. doi: 10.1016/j.ajhg.2015.11.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dhama K., Khan S., Tiwari R., Sircar S., Bhat S., Malik Y.S., Singh K.P., Chaicumpa W., Bonilla-Aldana D.K., Rodriguez-Morales A.J. Coronavirus disease 2019–COVID-19. Clin. Microbiol. Rev. 2020;33:1–48. doi: 10.1128/CMR.00028-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dieter C., Brondani L.D.A., Schaeffer A.F., Zanotto C., Ramos D.T., Girardi E., Pellenz F.M., Camargo J.L., Moresco K.S., Lima L., Aubin M.R., Oliveira M.S.De, Rech T.H., Gerchman F., Bauermann C., Crispim D. Polymorphisms in ACE1, TMPRSS2, IFIH1, IFNAR2, and TYK2 genes are associated with worse clinical outcomes in COVID-19. Genes (Basel) 2023;14:29. doi: 10.3390/genes14010029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elhabyan A., Elyaacoub S., Sanad E., Abukhadra A., Elhabyan A., Dinu V. The role of host genetics in susceptibility to severe viral infections in humans and insights into host genetics of severe COVID-19 : a systematic review. Virus Res. 2020;289:1–12. doi: 10.1016/j.virusres.2020.198163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fernández-de-las-Peñas C., Notarte K.I., Peligro P.J., Velasco J.V., Ocampo M.J., Henry B.M., Arendt-Nielsen L., Torres-Macho J., Plaza-Manzano G. Long-COVID symptoms in individuals infected with different SARS-CoV-2 variants of concern: a systematic review of the literature. Viruses. 2022;14 doi: 10.3390/v14122629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox L.E., Locke M.C., Lenschow D.J. Context Is key: delineating the unique functions of IFNα and IFNβ in disease. Front. Immunol. 2020;11:1–28. doi: 10.3389/fimmu.2020.606874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fricke-Galindo I., Martínez-Morales A., Chávez-Galán L., Ocaña-Guzmán R., Buendía-Roldán I., Pérez-Rubio G., Hernández-Zenteno R., de J., Verónica-Aguilar A., Alarcón-Dionet A., Aguilar-Duran H., Gutiérrez-Pérez I.A., Zaragoza-García O., Alanis-Ponce J., Camarena A., Bautista-Becerril B., Nava-Quiroz K.J., Mejía M., Guzmán-Guzmán I.P., Falfán-Valencia R. IFNAR2 relevance in the clinical outcome of individuals with severe COVID-19. Front. Immunol. 2022;13:1–9. doi: 10.3389/fimmu.2022.949413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Funk C.D., Laferrière C., Ardakani A. A snapshot of the global race for vaccines targeting SARS-CoV-2 and the COVID-19 pandemic. Front. Pharmacol. 2020;11:1–17. doi: 10.3389/fphar.2020.00937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gómez J., Albaiceta G.M., García-clemente M., López-larrea C., Amado-Rodríguez L., Lopez-Alonso I., Hermida T., Enriquez A.I., Herrero P., Melón S., Alvarez-Argüelles M.E., Boga J.A., Rojo-Alba S., Cuesta-Llavona E., Alvarez V., Lorca R., Coto E. Angiotensin-converting enzymes (ACE, ACE2) gene variants and COVID-19 outcome. Gene. 2020;762:1–5. doi: 10.1016/j.gene.2020.145102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hadjadj J., Yatim N., Barnabei L., Corneau A., Boussier J. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science. 2020;724(80):718–724. doi: 10.1126/science.abc6027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han C.S., Chen Y., Ezashi T., Roberts R.M. Antiviral activities of the soluble extracellular domains of type I interferon receptors. Proc. Natl. Acad. Sci. U. S. A. 2001;98:6138–6143. doi: 10.1073/pnas.111139598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hurtado-Guerrero I., Hernáez B., Pinto-Medel M.J., Calonge E., Rodriguez-Bada J.L., Urbaneja P., Alonso A., Mena-Vázquez N., Aliaga P., Issazadeh-Navikas S., Pavia J., Leyva L., Alcamí J., Alcamí A., Fernández Ó., Oliver-Martos B. Antiviral, immunomodulatory and antiproliferative activities of recombinant soluble IFNAR2 without IFN-ß mediation. J. Clin. Med. 2020;9 doi: 10.3390/jcm9040959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imai Y., Kuba K., Rao S., Huan Y., Guo F., Guan B., Yang P., Sarao R., Wada T., Leong-Poi H., Crackower M.A., Fukamizu A., Hui C.C., Hein L., Uhlig S., Slutsky A.S., Jiang C., Penninger J.M. Angiotensin-converting enzyme 2 protects from severe acute lung failure. Nature. 2005;436:112–116. doi: 10.1038/nature03712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaur S.P., Gupta V. COVID-19 vaccine : a comprehensive status report. Virus Res. 2020;288:1–12. doi: 10.1016/j.virusres.2020.198114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim Y.C., Jeong B.H. Strong correlation between the case fatality rate of covid-19 and the rs6598045 single nucleotide polymorphism (Snp) of the interferon-induced transmembrane protein 3 (ifitm3) gene at the population-level. Genes (Basel) 2021;12:1–9. doi: 10.3390/genes12010042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kompaniyets L., Pennington A.F., Goodman A.B., Rosenblum H.G., Belay B., Ko J.Y., Chevinsky J.R., Schieber L.Z., Summers A.D., Lavery A.M., Preston L.E., Danielson M.L., Cui Z., Namulanda G., Yusuf H., Gundlapalli A.V. Underlying medical conditions and severe illness among 540,667 adults hospitalized underlying medical conditions and severe with COVID-19, March 2020–March 2021. Prev. Chronic Dis. 2021;18:1–13. doi: 10.5888/pcd18.210123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuba K., Imai Y., Rao S., Gao H., Guo F., Guan B., Huan Y., Yang P., Zhang Y., Deng W., Bao L., Zhang B., Liu G., Wang Z., Chappell M., Liu Y., Zheng D., Leibbrandt A., Wada T., Slutsky A.S., Liu D., Qin C., Jiang C., Penninger J.M. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus-induced lung injury. Nat. Med. 2005;11:875–879. doi: 10.1038/nm1267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee J.S., Shin E.C. The type I interferon response in COVID-19: implications for treatment. Nat. Rev. Immunol. 2020;20:585–586. doi: 10.1038/s41577-020-00429-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J., Zhou Y., Ma J., Zhang Q., Shao J., Liang S., Yu Y., Li W., Wang C. The long-term health outcomes, pathophysiological mechanisms and multidisciplinary management of long COVID. Signal Transduct. Target. Ther. 2023;8 doi: 10.1038/s41392-023-01640-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LoPresti M., Beck D.B., Duggal P., Cummings D.A.T., Solomon B.D. The role of host genetic factors in coronavirus susceptibility: review of animal and systematic review of human literature. Am. J. Hum. Genet. 2020;107:381–402. doi: 10.1016/j.ajhg.2020.08.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu R., Zhao X., Li J., Niu P., Yang B., Wu H., Wang W., Song H., Huang B., Zhu N., Bi Y., Ma X., Zhan F., Wang L., Hu T., Zhou H., Hu Z., Zhou W., Zhao L., Chen J., Meng Y., Wang J., Lin Y., Yuan J., Xie Z., Ma J., Liu W.J., Wang D., Xu W., Holmes E.C., Gao G.F., Wu G., Chen W., Shi W., Tan W. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395:565–574. doi: 10.1016/S0140-6736(20)30251-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma Y., Huang Y., Zhao S., Yao Y., Zhang Y., Qu J., Wu N., Su J. Integrative genomics analysis reveals a 21q22.11 locus contributing risk to COVID-19. Hum. Mol. Genet. 2021;30:1247–1258. doi: 10.1093/hmg/ddab125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maiti A.K. The African-American population with a low allele frequency of SNP rs1990760 (T allele) in IFIH1 predicts less IFN-beta expression and potential vulnerability to COVID-19 infection. Immunogenetics. 2020;72:387–391. doi: 10.1007/s00251-020-01174-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manry J., Bastard P., Gervais A., Le Voyer T., Rosain J., Philippot Q., Michailidis E., Hoffmann H.H., Eto S., Garcia-Prat M., Bizien L., Parra-Martínez A., Yang R., Haljasmägi L., Migaud M., Särekannu K., Maslovskaja J., de Prost N., Tandjaoui-Lambiotte Y., Luyt C.E., Amador-Borrero B., Gaudet A., Poissy J., Morel P., Richard P., Cognasse F., Troya J., Trouillet-Assant S., Belot A., Saker K., Garçpn P., Rivière J.G., Lagier J.C., Gentile S., Rosen L.B., Shaw E., Morio T., Tanaka J., Dalmau D., Tharaux P.L., Sene D., Stepanian A., Mégarbane B., Triantafyllia V., Fekkar A., Heath J.R., Franco J.L., Anaya J.M., Solé-Violán J., Imberti L., Biondi A., Bonfanti P., Castagnoli R., Delmonte O.M., Zhang Y., Snow A.L., Holland S.M., Biggs C.M., Moncada-Vélez M., Arias A.A., Lorenzo L., Boucherit S., Anglicheau D., Planas A.M., Haerynck F., Duvlis S., Ozcelik T., Keles S., Bousfiha A.A., El Bakkouri J., Ramirez-Santana C., Paul S., Pan-Hammarström Q., Hammarström L., Dupont A., Kurolap A., Metz C.N., Aiuti A., Casari G., Lampasona V., Ciceri F., Barreiros L.A., Dominguez-Garrido E., Vidigal M., Zatz M., van de Beek D., Sahanic S., Tancevski I., Stepanovskyy Y., Boyarchuk O., Nukui Y., Tsumura M., Vidaur L., Tangye S.G., Burrel S., Duffy D., Quintana-Murci L., Klocperk A., Kann N.Y., Shcherbina A., Lau Y.L., Leung D., Coulongeat M., Marlet J., Koning R., Reyes L.F., Chauvineau-Grenier A., Venet F., Monneret G., Nussenzweig M.C., Arrestier R., Boudhabhay I., Baris-Feldman H., Hagin D., Wauters J., Meyts I., Dyer A.H., Kennelly S.P., Bourke N.M., Halwani R., Sharif-Askari F.S., Dorgham K., Sallette J., Sedkaoui S.M., AlKhater S., Rigo-Bonnin R., Morandeira F., Roussel L., Vinh D.C., Erikstrup C., Condino-Neto A., Prando C., Bondarenko A., Spaan A.N., Gilardin L., Fellay J., Lyonnet S., Bilguvar K., Lifton R.P., Mane S., Anderson M.S., Boisson B., Béziat V., Zhang S.Y., Andreakos E., Hermine O., Pujol A., Peterson P., Mogensen T.H., Rowen L., Mond J., Debette S., de Lamballerie X., Burdet C., Bouadma L., Zins M., Soler-Palacin P., Colobran R., Gorochov G., Solanich X., Susen S., Martinez-Picado J., Raoult D., Vasse M., Gregersen P.K., Piemonti L., Rodríguez-Gallego C., Notarangelo L.D., Su H.C., Kisand K., Okada S., Puel A., Jouanguy E., Rice C.M., Tiberghien P., Zhang Q., Casanova J.L., Abel L., Cobat A. The risk of COVID-19 death is much greater and age dependent with type I IFN autoantibodies. Proc. Natl. Acad. Sci. U. S. A. 2022;119:1–10. doi: 10.1073/pnas.2200413119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuyama T., Kubli S.P., Yoshinaga S.K., Pfeffer K., Mak T.W. An aberrant STAT pathway is central to COVID-19. Cell Death Differ. 2020;27:3209–3225. doi: 10.1038/s41418-020-00633-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mazewski C., Perez R.E., Fish E.N., Platanias L.C. Type I Interferon (IFN)-regulated activation of canonical and non-canonical signaling pathways. Front. Immunol. 2020;11:1–13. doi: 10.3389/fimmu.2020.606456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murray M.F., Kenny E.E., Ritchie M.D., Rader D.J., Bale A.E., Giovanni M.A., Abul-Husn N.S. COVID-19 outcomes and the human genome. Genet. Med. 2020;22:1175–1177. doi: 10.1038/s41436-020-0832-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nalbandian, A., Desai, A.D., Wan, E.Y., 2023. Post-COVID-19 condition. [DOI] [PubMed]
- National Institutes of Health, 2021. Treatment guidelines panel. Coronavirus Disease 2019 (COVID-19). [PubMed]
- Oliveto G., Scagnolari C., Frasca F., Sorrentino L., Matera L., Nenna R., Viscido A., Scordio M., Petrarca L., Maria Zicari A., Gentilini E., D'Ettorre G., Ceccarelli G., Midulla F., Antonelli G., Pierangeli A. The non-functional ACE2 isoform, but not the SARS-CoV-2 receptor, is induced as an interferon-stimulated gene, in SARS-CoV-2 infected adults. Cytokine. 2022;158 doi: 10.1016/j.cyto.2022.155997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Onabajo O.O., Banday A.R., Stanifer M.L., Yan W., Obajemu A., Santer D.M., Florez-Vargas O., Piontkivska H., Vargas J.M., Ring T.J., Kee C., Doldan P., Tyrrell D.L., Mendoza J.L., Boulant S., Prokunina-Olsson L. Interferons and viruses induce a novel truncated ACE2 isoform and not the full-length SARS-CoV-2 receptor. Nat. Genet. 2020;52:1283–1293. doi: 10.1038/s41588-020-00731-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ovsyannikova I.G., Haralambieva I.H., Crooke S.N., Poland G.A., Kennedy R.B. The role of host genetics in the immune response to SARS-CoV-2 and COVID-19 susceptibility and severity. Immunol. Rev. 2020;296:205–219. doi: 10.1111/imr.12897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Öztürk R., Taşova Y., Ayaz A. Covid-19: pathogenesis, genetic polymorphism, clinical features and laboratory findings. Turkish J. Med. Sci. 2020;50:638–657. doi: 10.3906/SAG-2005-287. [DOI] [PubMed] [Google Scholar]
- Pagaduan J.V., Altawallbeh G. Early clues regarding the pathogenesis of long-COVID. Adv. TB Test. 2020:02–24. [Google Scholar]
- Pairo-Castineira E., Clohisey S., Klaric L., Bretherick A.D., Rawlik K., Pasko D., Walker S., Parkinson N., Fourman M.H., Russell C.D., Furniss J., Richmond A., Gountouna E., Wrobel N., Harrison D., Wang B., Wu Y., Meynert A., Griffiths F., Oosthuyzen W., Kousathanas A., Moutsianas L., Yang Z., Zhai R., Zheng C., Grimes G., Beale R., Millar J., Shih B., Keating S., Zechner M., Haley C., Porteous D.J., Hayward C., Yang J., Knight J., Summers C., Shankar-Hari M., Klenerman P., Turtle L., Ho A., Moore S.C., Hinds C., Horby P., Nichol A., Maslove D., Ling L., McAuley D., Montgomery H., Walsh T., Pereira A.C., Renieri A., Shen X., Ponting C.P., Fawkes A., Tenesa A., Caulfield M., Scott R., Rowan K., Murphy L., Openshaw P.J.M., Semple M.G., Law A., Vitart V., Wilson J.F., Baillie J.K. Genetic mechanisms of critical illness in COVID-19. Nature. 2020;591:92–97. doi: 10.1038/s41586-020-03065-y. [DOI] [PubMed] [Google Scholar]
- Phetsouphanh C., Darley D.R., Wilson D.B., Howe A., Munier C.M.L., Patel S.K., Juno J.A., Burrell L.M., Kent S.J., Dore G.J., Kelleher A.D., Matthews G.V. Immunological dysfunction persists for 8 months following initial mild-to-moderate SARS-CoV-2 infection. Nat. Immunol. 2022;23:210–216. doi: 10.1038/s41590-021-01113-x. [DOI] [PubMed] [Google Scholar]
- Ramasamy S., Subbian S. Critical determinants of cytokine storm and type i interferon response in COVID-19 pathogenesis. Clin. Microbiol. Rev. 2021;34:1–24. doi: 10.1128/CMR.00299-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rio C.del, Collins L.F., Malani P. Management of post-acute covid-19 in primary care. JAMA. 2020;324:1723–1724. doi: 10.1136/bmj.m3026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubio-Casillas A., Redwan E.M., Uversky V.N. SARS-CoV-2: a master of immune evasion. Biomedicines. 2022;10:1–24. doi: 10.3390/biomedicines10061339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sampaio N.G., Chauveau L., Hertzog J., Bridgeman A., Fowler G., Moonen J.P., Dupont M., Russell R.A., Noerenberg M., Rehwinkel J. The RNA sensor MDA5 detects SARS-CoV-2 infection. Sci. Rep. 2021;11:1–10. doi: 10.1038/s41598-021-92940-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sariol A., Perlman S. CellPress; 2020. Lessons For COVID-19 Immunity from Other Coronavirus Infections; pp. 248–263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmiedel B.J., Rocha J., Gonzalez-Colin C., Bhattacharyya S., Madrigal A., Ottensmeier C.H., Ay F., Chandra V., Vijayanand P. COVID-19 genetic risk variants are associated with expression of multiple genes in diverse immune cell types. Nat. Commun. 2021;12:1–12. doi: 10.1038/s41467-021-26888-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schoggins J.W. Interferon-stimulated genes: what do they all do? Annu. Rev. Virol. 2019;6:567–584. doi: 10.1146/annurev-virology-092818-015756. [DOI] [PubMed] [Google Scholar]
- Schreiber G. The role of type I interferons in the pathogenesis and treatment of COVID-19. Front. Immunol. 2020;11:8–10. doi: 10.3389/fimmu.2020.595739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schreiber G. The molecular basis for differential type i interferon signaling. J. Biol. Chem. 2017;292:7285–7294. doi: 10.1074/jbc.R116.774562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaffer L. Lots of long COVID treatment leads, but few are proven. Proc. Natl. Acad. Sci. U. S. A. 2022;119:1–5. doi: 10.1073/pnas.2213524119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shah V.K., Firmal P., Alam A., Ganguly D., Chattopadhyay S. Overview of immune response during SARS-CoV-2 infection: lessons from the past. Front. Immunol. 2020;11:1–17. doi: 10.3389/fimmu.2020.01949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shastry, B.S., 2009. Single nucleotide polymorphisms: impact on gene function & phenotype, in: Single Nucleotide Polymorphisms: Methods & Protocols. p. 1–22. 10.1007/978-1-60327-411-1.
- Smith N., Possémé C., Bondet V., Sugrue J., Townsend L., Charbit B., Rouilly V., Saint-André V., Dott T., Pozo A.R., Yatim N., Schwartz O., Cervantes-Gonzalez M., Ghosn J., Bastard P., Casanova J.L., Szwebel T.A., Terrier B., Conlon N., O'Farrelly C., Cheallaigh C.N., Bourke N.M., Duffy D. Defective activation and regulation of type I interferon immunity is associated with increasing COVID-19 severity. Nat. Commun. 2022;13:1–14. doi: 10.1038/s41467-022-34895-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Solimani F., Meier K., Ghoreschi K. Janus kinase signaling as risk factor and therapeutic target for severe SARS-CoV-2 infection. Eur. J. Immunol. 2021;51:1071–1075. doi: 10.1002/eji.202149173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun Y., Wang G., Wang R., Ren L., Yuan Z., Liu Y., Wu Y., Chen R., Chen Y., Diao B. Serum levels of type I interferon (IFN-I) is associated with the severity of COVID-19. J. Med. Microbiol. 2023;72:1–9. doi: 10.1099/jmm.0.001694. [DOI] [PubMed] [Google Scholar]
- Tay M.Z., Poh C.M., Rénia L., MacAry P.A., Ng L.F.P. The trinity of COVID-19: immunity, inflammation and intervention. Nat. Rev. Immunol. 2020;20:363–374. doi: 10.1038/s41577-020-0311-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas C., Moraga I., Levin D., Krutzik P.O., Podoplelova Y., Trejo A., Lee C., Yarden G., Vleck S.E., Glenn J.S., Nolan P., Piehler J., Schreiber G., Garcia K.C. Structural linkage between ligand discrimination and receptor activation by type I interferons. Cell. 2012;146:621–632. doi: 10.1016/j.cell.2011.06.048.Structural. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torjesen I. Covid-19: omicron may be more transmissible than other variants and partly resistant to existing vaccines, scientists fear. BMJ. 2021;375:2. doi: 10.1136/bmj.n2943. [DOI] [PubMed] [Google Scholar]
- Tsuchida T., Hirose M., Inoue Y., Kunishima H., Otsubo T., Matsuda T. Relationship between changes in symptoms and antibody titers after a single vaccination in patients with Long COVID. J. Med. Virol. 2022;94:3416–3420. doi: 10.1002/jmv.27689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Der Made C.I., Simons A., Schuurs-Hoeijmakers J., Van Den Heuvel G., Mantere T., Kersten S., Van Deuren R.C., Steehouwer M., Van Reijmersdal S.V., Jaeger M., Hofste T., Astuti G., Corominas Galbany J., Van Der Schoot V., Van Der Hoeven H., Hagmolen Of Ten Have W., Klijn E., Van Den Meer C., Fiddelaers J., De Mast Q., Bleeker-Rovers C.P., Joosten L.A.B., Yntema H.G., Gilissen C., Nelen M., Van Der Meer J.W.M., Brunner H.G., Netea M.G., Van De Veerdonk F.L., Hoischen A. Presence of genetic variants among young men with severe COVID-19. JAMA - J. Am. Med. Assoc. 2020;324:663–673. doi: 10.1001/jama.2020.13719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- World Health Organization, 2023. WHO Coronavirus (COVID-19) Dashboard [WWW Document]. https://covid19.who.int/ (acessado 6.1.23).
- Wu, C., Chen, X., Cai, Y., Xia, J., Zhou, Xing, Xu, S., Huang, H., Zhang, L., Zhou, Xia, Du, C., Zhang, Y., Song, J., Wang, S., Chao, Y., Yang, Z., Xu, J., Zhou, Xin, Chen, D., Xiong, W., Xu, L., Zhou, F., Jiang, J., Bai, C., Zheng, J., Song, Y., 2020. Risk factors associated with acute respiratory distress syndrome and death in patients with coronavirus disease 2019 Pneumonia in Wuhan, China. JAMA Intern. Med. 180, 934–943. 10.1001/jamainternmed.2020.0994. [DOI] [PMC free article] [PubMed]
- Yaugel-Novoa M., Bourlet T., Longet S., Botelho-Nevers E., Paul S. Association of IFNAR1 and IFNAR2 with COVID-19 severity. Lancet Microbe. 2023;4:1. doi: 10.1016/S2666-5247(23)00095-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yin X., Riva L., Pu Y., Martin-Sancho L., Kanamune J., Yamamoto Y., Sakai K., Gotoh S., Miorin L., De Jesus P., Yang C.C., Herbert K., Yoh S., Hultquist J., Garcia-Sastre A., Chanda S.K. MDA5 governs the innate immune response to SARS-CoV-2 in lung epithelial cells. Cell Rep. 2021;34 doi: 10.2139/ssrn.3682826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeberg H., Pääbo S. A genomic region associated with protection against severe COVID-19 is inherited from Neandertals. Proc. Natl. Acad. Sci. U. S. A. 2021;118:3–7. doi: 10.1073/pnas.2026309118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang, Q., Liu, Z., Moncada-Velez, M., Chen, J., Ogishi, M., Bigio, B., Yang, R., Arias, A.A., Zhou, Q., Han, J.E., Ugurbil, A.C., Zhang, P., Rapaport, F., Li, J., Spaan, A.N., Boisson, B., Boisson-Dupuis, S., Bustamante, J., Puel, A., Ciancanelli, M.J., Zhang, S.Y., Béziat, V., Jouanguy, E., Abel, L., Cobat, A., Casanova, J.L., Bastard, P., Korol, C., Rosain, J., Philippot, Q., Chbihi, M., Lorenzo, L., Bizien, L., Neehus, A.L., Kerner, G., Seeleuthner, Y., Manry, J., Le Voyer, T., Boisson, B., Boisson-Dupuis, S., Bustamante, J., Puel, A., Zhang, S.Y., Béziat, V., Jouanguy, E., Abel, L., Cobat, A., Bastard, P., Rosain, J., Philippot, Q., Chbihi, M., Lorenzo, L., Bizien, L., Neehus, A.L., Kerner, G., Seeleuthner, Y., Manry, J., Le Voyer, T., Boisson, B., Boisson-Dupuis, S., Bustamante, J., Puel, A., Zhang, S.Y., Béziat, V., Jouanguy, E., Abel, L., Cobat, A., Le Pen, J., Schneider, W.M., Razooky, B.S., Hoffmann, H.H., Michailidis, E., Rice, C.M., Sabli, I.K.D., Hodeib, S., Sancho-Shimizu, V., Bilguvar, K., Ye, J., Maniatis, T., Bolze, A., Arias, A.A., Arias, A.A., Zhang, Y., Notarangelo, L.D., Su, H.C., Zhang, Y., Notarangelo, L.D., Su, H.C., Onodi, F., Korniotis, S., Karpf, L., Soumelis, V., Bonnet-Madin, L., Amara, A., Dorgham, K., Gorochov, G., Smith, N., Duffy, D., Moens, L., Meyts, I., Meade, P., García-Sastre, A., Krammer, F., Corneau, A., Masson, Cecile, Schmitt, Yohann, Schlüter, A., Pujol, A., Khan, T., Marr, N., Fellay, J., Fellay, J., Fellay, J., Roussel, L., Vinh, D.C., Shahrooei, Mohammad, Shahrooei, Mohammad, Alosaimi, M.F., Alsohime, F., Hasanato, R., Mansouri, Davood, Al-Saud, H., Almourfi, F., Al-Mulla, F., Al-Muhsen, S.Z., Al Turki, S., Al Turki, S., van de Beek, D., Biondi, A., Bettini, L.R., D'Angio, M., Bonfanti, P., Imberti, L., Sottini, A., Paghera, S., Quiros-Roldan, E., Rossi, C., Oler, A.J., Tompkins, M.F., Alba, C., Dalgard, C.L., Vandernoot, I., Smits, G., Goffard, J.C., Migeotte, I., Haerynck, F., Soler-Palacin, P., Martin-Nalda, A., Colobran, R., Morange, P.E., Keles, S., Çölkesen, F., Ozcelik, T., Yasar, K.K., Senoglu, S., Karabela, Ş.N., Rodríguez-Gallego, C., Rodríguez-Gallego, C., Novelli, G., Hraiech, S., Tandjaoui-Lambiotte, Y., Tandjaoui-Lambiotte, Y., Duval, X., Laouénan, C., Duval, X., Laouénan, C., Laouénan, C., Snow, A.L., Dalgard, C.L., Milner, J.D., Mogensen, T.H., Spaan, A.N., Bustamante, J., Ciancanelli, M.J., Maniatis, T., Soumelis, V., Nussenzweig, M., García-Sastre, A., García-Sastre, A., García-Sastre, A., Lifton, R.P., Lifton, R.P., Lifton, R.P., Foti, G., Bellani, G., Citerio, G., Contro, E., Pesci, A., Valsecchi, M.G., Cazzaniga, M., Abad, J., Blanco, I., Rodrigo, C., Aguilera-Albesa, S., Akcan, O.M., Darazam, I.A., Aldave, J.C., Ramos, M.A., Nadji, S.A., Alkan, G., Allardet-Servent, J., Allende, L.M., Alsina, L., Alyanakian, M.A., Amador-Borrero, B., Mouly, S., Sene, D., Amoura, Z., Mathian, A., Antolí, A., Blanch, G.R., Riera, J.S., Moreno, X.S., Arslan, S., Assant, S., Auguet, T., Azot, A., Bajolle, F., Bustamante, J., Lévy, R., Oualha, M., Baldolli, A., Ballester, M., Feldman, H.B., Barrou, B., Beurton, A., Bilbao, A., Blanchard-Rohner, G., Blandinières, A., Rivet, N., Blazquez-Gamero, D., Bloomfield, M., Bolivar-Prados, M., Clavé, P., Borie, R., Bosteels, C., Lambrecht, B.N., van Braeckel, E., Bousfiha, A.A., Bouvattier, C., Vincent, A., Boyarchuk, O., Bueno, M.R.P., Castro, M. V., Matos, L.R.B., Zatz, M., Agra, J.J.C., Calimli, S., Capra, R., Carrabba, M., Fabio, G., Casasnovas, C., Vélez-Santamaria, V., Caseris, M., Falck, A., Poncelet, G., Castelle, M., Castelli, F., de Vera, M.C., Catherinot, E., Chalumeau, M., Toubiana, J., Charbit, B., Li, Z., Pellegrini, S., Cheng, M.P., Clotet, B., Codina, A., Colkesen, F., Çölkesen, F., Colobran, R., Comarmond, C., Dalmau, D., Dalmau, D., Darley, D.R., Dauby, N., Dauger, S., Le Bourgeois, F., Levy, M., de Pontual, L., Dehban, A., Delplancq, G., Demoule, A., Diehl, J.L., Dobbelaere, S., Durand, S., Mircher, C., Rebillat, A.S., Vilaire, M.E., Eldars, W., Elgamal, M., Elnagdy, M.H., Emiroglu, M., Erdeniz, E.H., Aytekin, S.E., Euvrard, R., Evcen, R., Faivre, L., Fartoukh, M., Philippot, Q., Faure, M., Arquero, M.F., Flores, C., Flores, C., Flores, C., Flores, C., Francois, B., Fumadó, V., Fumadó, V., Fumadó, V., Fusco, F., Ursini, M.V., Solis, B.G., de Diego, Rebeca Pérez, van Den Rym, A.M., Gaussem, P., Gil-Herrera, J., Gilardin, L., Alarcon, M.G., Girona-Alarcón, M., Goffard, J.C., Gok, F., Yosunkaya, A., González-Montelongo, R., Íñigo-Campos, A., Lorenzo-Salazar, J.M., Muñoz-Barrera, A., Guerder, A., Gul, Y., Guner, S.N., Gut, M., Hadjadj, J., Haerynck, F., Halwani, R., Hammarström, L., Hatipoglu, N., Hernandez-Brito, E., Heijmans, C., Holanda-Peña, M.S., Horcajada, J.P., Hoste, L., Hoste, E., Hraiech, S., Humbert, L., Mordacq, C., Thumerelle, C., Vuotto, F., Iglesias, A.D., Jamme, M., Arranz, M.J., Jordan, I., Jorens, P., Kanat, F., Kapakli, H., Kara, I., Karbuz, A., Yasar, K.K., Senoglu, S., Keles, S., Demirkol, Y.K., Klocperk, A., Król, Z.J., Kuentz, P., Kwan, Y.W.M., Lagier, J.C., Lau, Y.L., Leung, D., Leo, Y.S., Young, B.E., Lopez, R.L., Levin, M., Linglart, A., Loeys, B., Louapre, C., Lubetzki, C., Luyt, C.E., Lye, D.C., Marjani, M., Pereira, J.M., Martin, A., Soler-Palacín, P., Pueyo, D.M., Martinez-Picado, J., Marzana, I., Matthews, G. V., Mayaux, J., Parizot, C., Quentric, P., Mège, J.L., Raoult, D., Melki, I., Meritet, J.F., Metin, O., Mezidi, M., Migeotte, I., Taccone, F., Millereux, M., Mirault, T., Mirsaeidi, M., Melián, A.M., Martinez, A.M., Morange, P., Morelle, G., Naesens, L., Nafati, C., Neves, J.F., Ng, L.F.P., Medina, Y.N., Cuadros, E.N., Gonzalo Ocejo-Vinyals, J., Orbak, Z., Özçelik, T., Pan-Hammarström, Q., Pascreau, T., Paz-Artal, E., Philippe, A., Planas-Serra, L., Schluter, A., Ploin, D., Viel, S., Poissy, J., Pouletty, M., Reisli, I., Ricart, P., Richard, J.C., Rivière, J.G., Rodriguez-Gallego, C., Rodriguez-Gallego, C., Rodríguez-Palmero, A., Romero, C.S., Rothenbuhler, A., Rozenberg, F., del Prado, M.Y.R., Sanchez, O., Sánchez-Ramón, S., Schmidt, M., Schweitzer, C.E., Scolari, F., Sediva, A., Seijo, L.M., Seppänen, M.R.J., Ilovich, A.S., Slabbynck, H., Smadja, D.M., Sobh, A., Solé-Violán, J., Soler, C., Stepanovskiy, Y., Stoclin, A., Tandjaoui-Lambiotte, Y., Taupin, J.L., Tavernier, S.J., Terrier, B., Tomasoni, G., Alvarez, J.T., Trouillet-Assant, S., Troya, J., Tucci, A., Uzunhan, Y., Vabres, P., Valencia-Ramos, J., van de Velde, S., van Praet, J., Vandernoot, I., Vatansev, H., Vilain, C., Voiriot, G., Yucel, F., Zannad, F., Belot, A., Bole-Feysot, C., Lyonnet, S., Masson, Cécile, Nitschke, P., Pouliet, A., Schmitt, Yoann, Tores, F., Zarhrate, M., Shahrooei, Mohammad, Abel, L., Andrejak, C., Angoulvant, F., Bachelet, D., Bhavsar, K., Bouadma, L., Chair, A., Couffignal, C., Silveira, C. Da, Debray, M.P., Duval, X., Eloy, P., Esposito-Farese, M., Ettalhaoui, N., Gault, N., Ghosn, J., Gorenne, I., Hoffmann, I., Kafif, O., Kali, S., Khalil, A., Laouénan, C., Laribi, S., Le, M., Le Hingrat, Q., Lescure, F.X., Lucet, J.C., Mentré, F., Mullaert, J., Peiffer-Smadja, N., Peytavin, G., Roy, C., Schneider, M., Mohammed, N.S., Tagherset, L., Tardivon, C., Tellier, M.C., Timsit, J.F., Trioux, T., Tubiana, S., Basmaci, R., Behillil, S., Beluze, M., Benkerrou, D., Dorival, C., Meziane, A., Téoulé, F., Bompart, F., Bouscambert, M., Gaymard, A., Lina, B., Rosa-Calatrava, M., Terrier, O., Caralp, M., Cervantes-Gonzalez, M., D'Ortenzio, E., Puéchal, O., Semaille, C., Coelho, A., Diouf, A., Hoctin, A., Mambert, M., Couffin-Cadiergues, S., Deplanque, D., Descamps, D., Visseaux, B., Desvallées, M., Khan, C., Diallo, A., Mercier, N., Paul, C., Petrov-Sanchez, V., Dubos, F., Enouf, V.V.E., Mouquet, H., Esperou, H., Jaafoura, S., Papadopoulos, A., Etienne, M., Gigante, T., Rossignol, B., Guedj, J., Le Nagard, H., Lingas, G., Neant, N., Kaguelidou, F., Lévy, Y., Wiedemann, A., Lévy, Y., Wiedemann, A., Levy-Marchal, C., Malvy, D., Noret, M., Pages, J., Picone, O., Rossignol, P., Tual, C., Veislinger, A., van der Werf, S., Vanel, N., Yazdanpanah, Y., Alavoine, L., Costa, Y., Duval, X., Ecobichon, J.L., Frezouls, W., Ilic-Habensus, E., Leclercq, A., Lehacaut, J., Letrou, S., Mandic, M., Nouroudine, M., Quintin, C., Rexach, J., Tubiana, S., Vignali, V., Amat, K.K.A., Behillil, S., Enouf, V., van der Werf, S., Bielicki, J., Bruijning, P., Burdet, C., Burdet, C., Caumes, E., Charpentier, C., Damond, F., Descamps, D., Le Hingrat, Q., Visseaux, B., Coignard, B., Couffin-Cadiergues, S., Delmas, C., Espérou, H., Roufai, L., Dechanet, A., Houhou, N., Kafif, O., Kikoine, J., Manchon, P., Piquard, V., Postolache, A., Terzian, Z., Lebeaux, D., Lina, B., Lucet, J.C., Malvy, D., Meghadecha, M., Motiejunaite, J., Thy, M., van Agtmael, M., Bomers, M., Chouchane, O., Geerlings, S., Goorhuis, B., Grobusch, M.P., Harris, V., Hermans, S.M., Hovius, J.W., Nellen, J., Peters, E., van der Poll, T., Prins, J.M., Reijnders, T., Schinkel, M., Sigaloff, K., Stijnis, C.S., van der Valk, M., van Vugt, M., Joost Wiersinga, W., Algera, A.G., van Baarle, F., Bos, L., Botta, M., de Bruin, S., Bulle, E., Elbers, P., Fleuren, L., Girbes, A., Hagens, L., Heunks, L., Horn, J., van Mourik, N., Paulus, F., Raasveld, J., Schultz, M.J., Smit, M., Stilma, W., Thoral, P., Tsonas, A., de Vries, H., Bax, D., Cloherty, A., Beudel, M., Brouwer, M.C., Koning, R., van de Beek, D., Bogaard, H.J., de Brabander, J., de Bree, G., Bugiani, M., Geerts, B., Hollmann, M.W., Preckel, B., Veelo, D., Geijtenbeek, T., Hafkamp, F., Hamann, J., Hemke, R., de Jong, M.D., Schuurman, A., Teunissen, C., Vlaar, A.P.J., Wouters, D., Zwinderman, A.H., Abel, L., Aiuti, A., Muhsen, S. Al, Al-Mulla, F., Anderson, M.S., Arias, A.A., Feldman, H.B., Bogunovic, D., Itan, Y., Bolze, A., Cirulli, E., Barrett, K.S., Washington, N., Bondarenko, A., Bousfiha, A.A., Brodin, P., Bryceson, Y., Bustamante, C.D., Butte, M., Casari, G., Chakravorty, S., Christodoulou, J., Le Mestre, S., Condino-Neto, A., Cooper, M.A., Dalgard, C.L., David, A., DeRisi, J.L., DeRisi, J.L., Desai, M., Drolet, B.A., Espinosa, S., Fellay, J., Flores, C., Franco, J.L., Gregersen, P.K., Haerynck, F., Hagin, D., Heath, J., Henrickson, S.E., Hsieh, E., Imai, K., Karamitros, T., Kisand, K., Ku, C.L., Lau, Y.L., Ling, Y., Lucas, C.L., Maniatis, T., Mansouri, Davoud, Marodi, L., Milner, J., Mironska, K., Morio, T., Notarangelo, L.D., Su, H.C., Novelli, A., O'Farrelly, C., Okada, S., Ozcelik, T., de Diego, Rebeca Perez, Planas, A.M., Prando, C., Pujol, A., Quintana-Murci, L., Renia, L., Renieri, A., Rodríguez-Gallego, C., Sancho-Shimizu, V., Sankaran, V., Shahrooei, Mohammed, Snow, A., Soler-Palacín, P., Spaan, A.N., Tangye, S., Turvey, S., Uddin, F., Uddin, M.J., Uddin, M.J., van de Beek, D., Vazquez, S.E., Vinh, D.C., von Bernuth, H., Zawadzki, P., Jing, H., Tung, W., Meguro, K., Shaw, E., Jing, H., Shafer, S., Zheng, L., Zhang, Z., Kubo, S., Chauvin, S.D., Meguro, K., Shaw, E., Lenardo, M., Luthers, C.R., Bauman, B.M., Shafer, S., Zheng, L., Zhang, Z., Kubo, S., Chauvin, S.D., Lenardo, M., Lack, J., Karlins, E., Hupalo, D.M., Rosenberger, J., Sukumar, G., Wilkerson, M.D., Zhang, X., 2020. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science (80). 370, 718–724. 10.1126/science.abd4570. [DOI] [PMC free article] [PubMed]
- Zhang S., Wang L., Cheng G. The battle between host and SARS-CoV-2: innate immunity and viral evasion strategies. Mol. Ther. 2022;30:1869–1884. doi: 10.1016/j.ymthe.2022.02.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou S., Butler-Laporte G., Nakanishi T., Morrison D.R., Afilalo J., Afilalo M., Laurent L., Pietzner M., Kerrison N., Zhao K., Brunet-Ratnasingham E., Henry D., Kimchi N., Afrasiabi Z., Rezk N., Bouab M., Petitjean L., Guzman C., Xue X., Tselios C., Vulesevic B., Adeleye O., Abdullah T., Almamlouk N., Chen Y., Chassé M., Durand M., Paterson C., Normark J., Frithiof R., Lipcsey M., Hultström M., Greenwood C.M.T., Zeberg H., Langenberg C., Thysell E., Pollak M., Mooser V., Forgetta V., Kaufmann D.E., Richards J.B. A Neanderthal OAS1 isoform protects individuals of European ancestry against COVID-19 susceptibility and severity. Nat. Med. 2021;27:659–667. doi: 10.1038/s41591-021-01281-1. [DOI] [PubMed] [Google Scholar]
- Ziegler C.G.K., Allon S.J., Nyquist S.K., Mbano I.M., Miao V.N., Tzouanas C.N., Cao Y., Yousif A.S., Bals J., Hauser B.M., Feldman J., Muus C., Wadsworth M.H., Kazer S.W., Hughes T.K., Doran B., Gatter G.J., Vukovic M., Taliaferro F., Mead B.E., Guo Z., Wang J.P., Gras D., Plaisant M., Ansari M., Angelidis I., Adler H., Sucre J.M.S., Taylor C.J., Lin B., Waghray A., Mitsialis V., Dwyer D.F., Buchheit K.M., Boyce J.A., Barrett N.A., Laidlaw T.M., Carroll S.L., Colonna L., Tkachev V., Peterson C.W., Yu A., Zheng H.B., Gideon H.P., Winchell C.G., Lin P.L., Bingle C.D., Snapper S.B., Kropski J.A., Theis F.J., Schiller H.B., Zaragosi L.E., Barbry P., Leslie A., Kiem H.P., Flynn J.A.L., Fortune S.M., Berger B., Finberg R.W., Kean L.S., Garber M., Schmidt A.G., Lingwood D., Shalek A.K., Ordovas-Montanes J., Banovich N., Brazma A., Desai T., Duong T.E., Eickelberg O., Falk C., Farzan M., Glass I., Haniffa M., Horvath P., Hung D., Kaminski N., Krasnow M., Kuhnemund M., Lafyatis R., Lee H., Leroy S., Linnarson S., Lundeberg J., Meyer K., Misharin A., Nawijn M., Nikolic M.Z., Pe'er D., Powell J., Quake S., Rajagopal J., Tata P.R., Rawlins E.L., Regev A., Reyfman P.A., Rojas M., Rosen O., Saeb-Parsy K., Samakovlis C., Schiller H., Schultze J.L., Seibold M.A., Shepherd D., Spence J., Spira A., Sun X., Teichmann S., Tsankov A., van den Berge M., von Papen M., Whitsett J., Xavier R., Xu Y., Zhang K. SARS-CoV-2 receptor ACE2 is an interferon-stimulated gene in human airway epithelial cells and is detected in specific cell subsets across tissues. Cell. 2020;181:1016–1035. doi: 10.1016/j.cell.2020.04.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Züst R., Cervantes-Barragan L., Habjan M., Maier R., Neuman B.W., Ziebuhr J., Szretter K.J., Baker S.C., Barchet W., Diamond M.S., Siddell S.G., Ludewig B., Thiel V. Ribose 2’-O-methylation provides a molecular signature for the distinction of self and non-self mRNA dependent on the RNA sensor Mda5. Nat. Immunol. 2011;12:137–143. doi: 10.1038/ni.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
No data was used for the research described in the article.