Abstract
Background:
Colon adenocarcinoma (COAD) is one of the most common cancer happened in gastrointestinal tract, with the overall incidence rate of 4%–5% among human beings. Like most malignancies, we uncovered the exact mechanisms of the pathogenesis of colorectal cancer yet. Therefore, there is an urgent need to explore the molecules that can be used as diagnostic maker at early stage. In addition, we also need to define the essential factors that related to the prognosis and treatment of the colon carcinoma.
Methods:
The study was conducted at the Third Affiliated Hospital of Qiqihar Medical University, Qiqihar, China in September 2020. The R language was used to identify the differentially expressed genes. We performed receiver operating characteristic curve analysis to determine the diagnostic markers for COAD. The machine learning strategy was used to assess the effectiveness of genes in the diagnosis of COAD. The molecular mechanism and prognostic value of genes were explored by bioinformatics analysis and molecular experiments.
Results:
The expression level of heat shock factor 4 (HSF4) was significantly elevated in COAD patients (P=1.89×10−29), according to The Cancer Genome Atlas (TCGA) database. Additionally, survival analysis showed the higher expression level of the HSF4 was correlated with the poor prognosis in COAD.
Conclusion:
The HSF4 was the target gene of zinc finger protein 692(ZNF692). HSF4 might promote the progression of COAD through the apoptosis pathway. It was diagnostic and prognosis maker of COAD. Furthermore, the upstream gene of HSF4, ZNF692, promotes the progression of colorectal cancer by regulating HSF4 expression.
Keywords: Colon adenocarcinoma, Biomarker, Heat shock factor 4, Zinc finger protein 692
Introduction
Colon adenocarcinoma (COAD) is common in the digestive system with high morbidity and mortality (1). The incidence of COAD is growing year by year in China, and the involved population are getting younger because of lifestyle and dietary changes in the past decades (2). The pathophysiology of COAD, like that of other malignant tumors, remains largely under investigated. COAD, as we know, is caused by a combination of genetic and environmental elements. Additionally, COAD is affected by a variety of factors, such as the environment, alcohol, cigarette addiction, intestinal homeostasis, diet, and physical activity (3). However, the treatment of COAD is still dominated by surgery, supplemented by radiotherapy and chemotherapy. Moreover, there are considerable individual variances in treatment effect among individuals with COAD. Although there are multiple choices for the COAD treatment, but the flaws of the foregoing strategies are clear in patients with advanced COAD, resulting in a dismal prognosis. With this in mind, the distant lymph node metastasis, peritoneal metastasis, and hematological metastasis are the most common types of postoperative metastasis for COAD, and they are accompanied with local recurrence typically (4, 5). As the fact that the most prevalent cause of therapeutic failure in COAD is hematogenous metastasis. The survival rates of 5 years among the patients, who are suffering with colon carcinoma, with distant, local, or no metastasis are ranging from 10% to 90%, in other words, are negatively corelated. The goal of this study is to find early detection indicators and to investigate the essential factors involved in COAD development and metastasis.
Heat shock factor 4 (HSF4) is a histone H3 variation of centromeric nucleosomes, which is necessary for normal chromosomal segregation assembly (6, 7). Artificial overexpressed HSF4 may lead to its cellular mislocalization, which will cause the dysfunction of the kinetochores, resulting in abnormal chromosomal separation and genomic instability (8–10). Furthermore, the abnormal expression of HSF4 and abnormalities of the function of HSF4 caused compromised genomic integrity led to improper cell division to occur, which may induce the development of cancers (11–13). Moreover, HSF4 nonphysiologically inclusion has been linked to human cancer in the past (14–16). On the other hand, HSF4, has only been documented in a few cases of COAD, and may be utilized as a diagnostic and therapeutic target for the disease.
The current study evaluated the involvement of the HSF4 in COAD progress, by the data from in vitro studies and clinical specimens from patients.
Methods
Data acquisition
Through the collecting of cancer-related omics data, the Cancer Genome Atlas (TCGA) offers a huge, free reference library for cancer research, which is publically accessible through the TCGA Data Portal (https://cancergenome.nih.gov). In September 2020, we retrieved the expression matrix of COAD patients as well as pertinent clinical data from the TCGA database.
Reagents and cell culture
COAD cell lines (LS513, CW2), normal human colon tissue cells (CCD-18CO cells) were all purchased from American Type Culture Collection (ATCC). In a 37 °C incubator with 5% CO2, RPMI-1640 medium (Solarbio, China) were used to grown the COAD and BEAS-2B cells. Additionally, 10% (v/v) fetal bovine serum (FBS, Noverse), 100 U/mL of penicillin, and 100 μg/mL of streptomycin (Solarbio) were added to the. Within the past three years, short tandem repeat profiling has been used to validate all human cell lines. All of the investigations were done on mycoplasma-free cells.
RNA extraction and qRT-PCR
TRIzol (Invitrogen, Carlsbad, CA, USA) was used to extract the total RNAs, followed the instruction by the manufacturer. 1 ug extracted RNA was reverse transcribed to cDNA, using the Moloney Leukemia Virus Reverse Transcriptase Kit (Madison, Wisconsin, Promega, USA). Then the qRT-PCR was carried out to amplify the samples to analyze mRNA expression level using Green Mix SYBR (Promega) kit. The housekeeping gene, GAPDH was referred to normalize the expression of the samples. Each experiment was repeated three times to determine the FC in the relative expression level, and the data were analyzed using the 2-ΔΔCT method. The primer sequences utilized in this work are included in Table S1. The primers were created by the NCBI website’s “pick primer” online tool.
Western blot
COAD cells were lysed in the RIPA buffer (Thermo Fisher Scientific) (Sigma-Aldrich) supplemented with protease inhibitor cocktail. Pierce Protein Assay Kit (Pierce, USA) was utilized to measure the protein concentration in the cell lysates. Proteins were separated by SDS-PAGE gel, then the proteins were transferred to polyvinylidene difluoride (PVDF) membranes. All the primary antibodies used in this study are listed here: MMP2 (Abcam, UK, ab37150), MMP9 (Abcam, ab73734), PLEKHG5 (Sigma-Aldrich), N-cadherin (Abcam, ab76011), Vimentin (Abcam, ab8069), mDia (Santa Cruz, USA, sc-373895), GAPDH (Cell Signaling Technology, USA, 5174). Chemiluminescence (Bio-Rad, USA) was used to quantify proteins in line with the methodology of the manufacturer.
Cell proliferation assay
The viability of the cells was measured by the Cell Counting Kit-8 (CCK-8) assay as indicated by the protocol from the manufacturer. Briefly, the cells were seeded in 96-well plates with appropriate amount of tumor cells. Then the varied amounts of TMZ or a dimethyl sulfoxide control was incubated with the cells. After that, the CCK-8 solution was supplemented into the wells. Cells were found in a microplate reader at 450 nm of absorbance.
Apoptosis Assay
When cells were treated with various doses of NG52, the apoptosis test was done using the Annexin-Vfluorescein isothiocyanate (FITC)/propidium iodide (PI) staining technique (KeyGen Biotech, Nanjing, China). The tests were conducted in line with the manufacturer’s recommendations.
Biological function and pathway enrichment analysis
There are certain genes that are functionally related because of the similar cellular expression distribution. Genes positively correlated with HSF4 in the selected modules above (shown by Cytoscape 3.6.1) were found by calculating the corresponding topological overlaps. After that, it underwent pathway analysis using the Kyoto Encyclopedia of Genes and Genomes (KEGG) and the Gene Ontology (GO). Then the enriched bioprocesses and KEGG pathways were obtained using the “clusterprofiler” package of the R/Bioconductor software for further analysis. Furthermore, HSF4 expression (median value) was utilized to divide TCGA samples into two groups, and gene set enrichment analysis (GSEA) software (http://software.broadinstitute.org/gsea/index.jsp) was utilized to confirm the results of GO and KEGG analysis in the two groups. Notably, GSEA had nominal P-value and false discovery rate (FDR) cut-offs of 0.05 and 0.25, respectively.
Statistical analysis
For statistical analysis, we utilized SPSS 23.0 (IBM Corp., Armonk, NY, USA) or GraphPad Prism 7 (GraphPad Prism, Inc., La Jolla, CA, USA). The final data were presented as the average SD of three separate studies. To compare two or three groups, the Student’s t-test or ANOVA were utilized, accordingly.
Results
COAD patients and COAD cells had high expression levels of HSF4
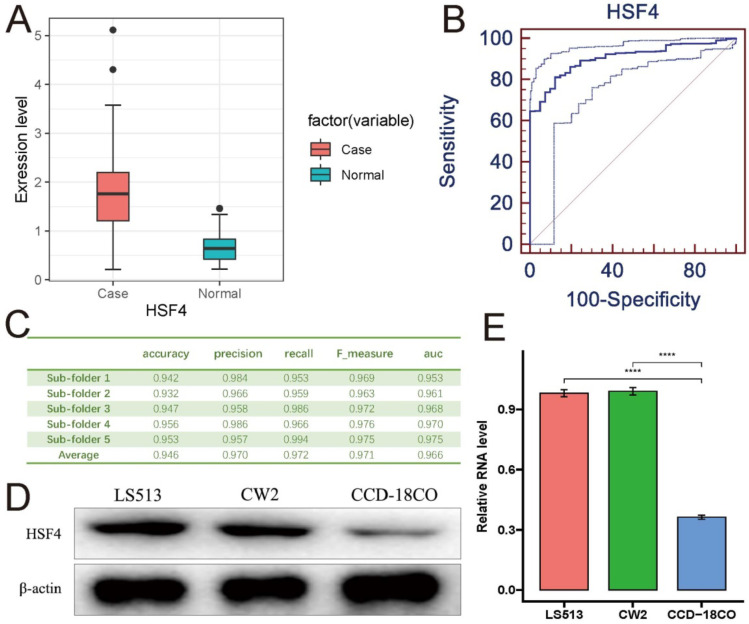
To explore the function of HSF4 in COAD, we determined the HSF4 expression levels in the TCGA database. We discovered the expression level of HSF4 was considerably higher in COAD patients (T=1.988, P=1.89×10−29 Fig.1A). Moreover, the sensitivity and specificity of HSF4 for the diagnosis of COAD in the TCGA database were examined using ROC curve analysis. For COAD, HSF4 demonstrated a strong diagnostic significance (AUC=0.903 [0.873, 0.927], P<0.0001) (Fig. 1B). Furthermore, the efficiency of HSF4 in diagnosing COAD was then evaluated using logistic regression. After constructing the confusion matrix, the mean of AUC (0.903) on the ROC curve of HSF4 was calculated (Fig. 1C). The averages for accuracy, precision, recall, and F1-score were 0.900, 0.924, 0.970, and 0.946 (Fig. 1C). RT-qPCR and western blot results revealed that the COAD cell lines had greater amounts of HSF4 mRNA and protein levels than that of normal human colon tissue cells (CCD-18CO cells) (Fig. 1D,E). Taken together, our findings indicated that the HSF4 performed well in distinguishing COAD samples from normal controls.
Fig. 1:
HSF4 is elevated in COAD patients and COAD cells. (A)TCGA database showed that HSF4 expression was considerably elevated in COAD patients. (B) ROC curve analysis showed that HSF4 had high diagnostic significance for COAD. (C). Evaluation metrics of each fold. (D). Western blot was used to detect the protein levels of HSF4. (E). RT-qPCR was used to detect the mRNA levels of HSF4. ****P < 0.0001
Survival analysis of HSF4 in COAD
In order to explore the relationship between HSF4 and prognosis of COAD, survival analysis determined that the increased level of HSF4 was negatively correlated with prognosis or positively corelated with the cancer-related mortality of COAD patients (P=0.026 HR=1.56;95%CI=1.05–2.32, Fig. 2A,B). Likewise, in 2-, 3-, and 4-year survival, the area under the curve of time dependent ROC curve was 0.635, 0.656, and 0.642 (Fig. 2C).
Fig. 2:
Survival analysis of HSF4 in COAD. (A) HSF4 expression and the survival time scatter gram of COAD patients; (B) The survival curves for groups with different expression levels of HSF4; (C) Time-dependent receiver operating characteristic (ROC) curve of HSF4 expression in predicting OS of COAD
Knockdown of HSF4 promoted cell apoptosis and inhibited cell proliferation in vitro
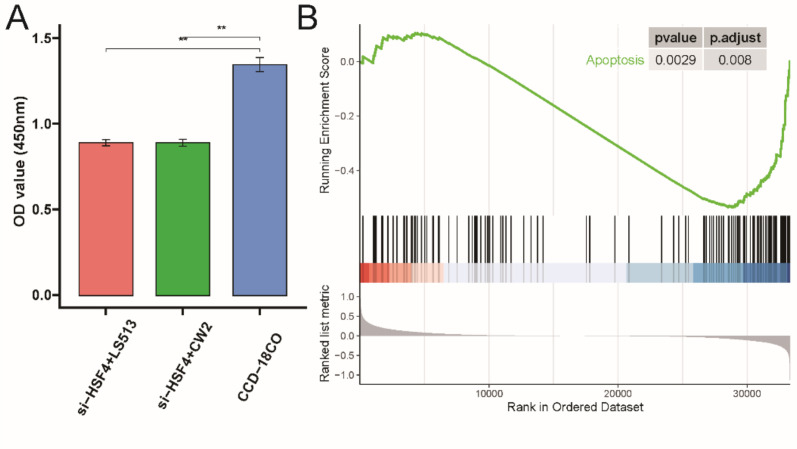
The COAD cell lines LS513, CW2 were transfected with si-HSF4 and the negative control siRNA (siRNA NC). Then the cell proliferation ability was detected by CCK-8 assay, and it was found that the level of cell proliferation was dramatically reduced after HSF4 knockdown (Fig. 3A). In addition, the GSEA software was utilized to examine the relationship between HSF4 expression and KEGG pathway gene sets. In fact, the cell apoptosis was shown to be considerably increased (NES= −1.603, P= 0.0029) (Fig. 3B).
Fig. 3:
Knockdown of HSF4 promoted apoptosis and inhibited cell proliferation in vitro. (A). The cell proliferation was measured by CCK-8 assay after HSF4 knockdown. (B). GSEA analysis showed that apoptosis were significantly enriched. **P< 0.01
HSF4 was the target gene of ZNF692
We discovered that HSF4 might be the target gene of ZNF692, analyzed by the R package “Dorothea”. The expression level of ZNF692 in TCGA database was shown to be considerably higher in COAD patients. (T=1.997, P=2.606×10−31 Fig. 4A). Furthermore, RT-qPCR and western blot results revealed that the COAD cell lines had higher level of ZNF692 mRNA and protein level than that of normal human colon tissue cells (CCD-18CO cells) (Fig. 4B, C). Additionally, we created a ZNF692 overexpression construct and transfected it into LS513 cells. Overexpression of ZNF692 induced HSF4 expression (Fig. 4D). In contrast, knocking down ZNF692 with shRNA produced the opposite effect (Fig. 4E), demonstrating that ZNF692 controls the expression of HSF4.
Fig. 4:
HSF4 was the target gene of ZNF692. (A) TCGA database showed that ZNF692 expression was elevated considerably in COAD patients. (B) RT-qPCR indicated that the mRNA level of ZNF692 was raised in the COAD cell lines compared to those in normal human colon tissue cells (CCD-18CO cells). (C) Western blot revealed that the protein level of ZNF692 was raised compared to those in normal human colon tissue cells (CCD-18CO cells). (D) We found that overexpression of ZNF692 did induce the expression of HSF4. (E) shRNA-mediated knockdown of ZNF692 did reduce the expression of HSF4. **P < 0.01, ****P < 0.0001
ZNF692 stimulates COAD cell proliferation in a HSF4 dependent manner
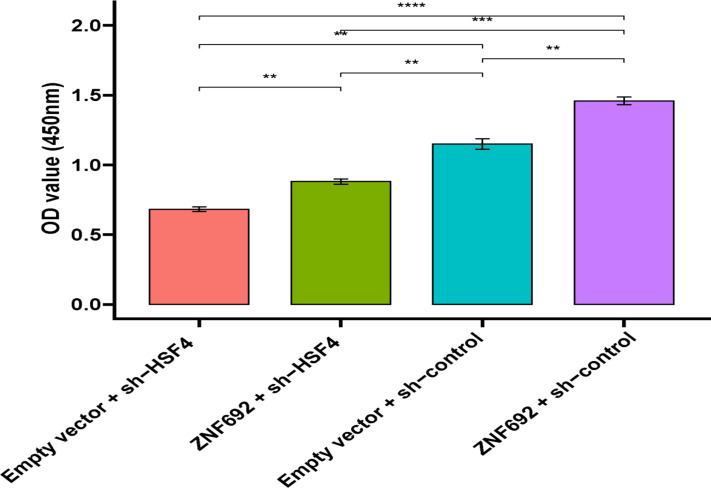
HSF4 was knocked down in ZNF692-overexpressing LS513 cells to test this theory. The CCK8 experiment revealed that ZNF692 + sh-Control LS513 cells proliferated faster than Empty vector + sh-Control LS513 cells, but it was reversed by HSF4 knockdown (ZNF692 + sh-HSF4) (Fig. 5), suggesting that ZNF692 increases COAD cell proliferation through HSF4.
Fig. 5:
ZNF692 stimulates COAD cell proliferation in a HSF4 dependent manner. CCK-8 was utilized to assess the proliferation of LS513 cells that were stable in their expression of ZNF692 + sh-HSF4. **P < 0.01, ***P < .001,****P< 0.0001
Discussion
Actually, COAD is the second-leading cause of carcinoma-related death all over the word (17). The inner epithelial cells of colorectal tissue are generated from endoderm cells through a multi-step process, which are started from normal epithelial cells to adenomatous polyps, then invasive colorectal cancer, also together with the inactivation of tumor suppressor genes and the activation of oncogenes (18). Patients with advanced COAD had a worse treatment response and survival rate than those with early COAD. Specifically, the rates of 5-year survival declining from 50% to 10% in the advanced instances. The most frequent radical treatment for locally advanced COAD is surgical tumor removal; however, for the metastatic carcinoma, which could not be able to cure by the traditional surgical, radiation, or chemotherapy, there is no optional choice to cure it (19). The COAD quantitative investigations show that conversion between stem cells and malignant carcinoma cells are happened within 10 years, and those carcinoma cells gain the potential to spread nearby tissues in the following 5 years (20). There are about 30%–40% of COAD patients experience recurrence following therapy, so the early and accurate identification is critical for disease management and prevention. Given the clinical hurdles of improving the prognosis of COAD patients with therapeutic techniques, a considerable number of patients getting standard therapy still suffer recurrence. To develop effective treatments, it is critical to uncover novel approaches, particularly molecular markers, for conducting quick colorectal cancer diagnosis and assessments, discovering novel therapy targets, and greater understanding of the fundamental COAD molecular mechanisms.
ZNF692 is the member of Krüppel C2H2 type zinc finger protein family; it contains five repeats of the C2H2-type zinc finger. Functionally, ANF692 may be able to regulate the gene transcription (21–23). In the past decades, there are growing data suggests that ZNF692 plays an critical role in tumor initiation and progression (24,25). Additionally, based on the publicly available data from the UALCAN database, ZNF692 was overexpressed in fourteen different types of cancer, such as head and neck squamous cell carcinoma, renal clear cell carcinoma, COAD, lung squamous cell carcinoma, bladder urothelial carcinoma, lung adenocarcinoma, and prostate adenocarcinoma (26, 27). Furthermore, in four of these cancers, ZNF692 was linked to a poor prognosis, such as COAD, adrenocortical carcinoma hepatocellular carcinoma, and renal clear cell carcinoma (28). ZFX expression was shown to be strongly connected with advanced stage and poor survival, and to play a significant role in COAD tumor growth (29, 30).
Furthermore, potential oncogenes with diagnostic and prognostic relevance were the focus of this investigation. HSF4 and ZNF692 were discovered to be elevated in COAD samples from the TCGA dataset. The mRNA and protein levels of HSF4 and ZNF692 were both raised when compared to normal human colon tissue cells, according to RT-qPCR and western blot. High HSF4 and ZNF692 expression levels were strongly linked with a poor prognosis in COAD. In addition, HSF4 gene expression level has a certain prognostic value in the long-term survival of COAD patients, according to a time-dependent ROC curve.
Our results from CCK8 assay suggested that knockdown of ZNF692 significantly decreased COAD cell proliferation. Moreover, we discovered that overexpression of ZNF692 induced higher expression of HSF4. In contrast, knocking down ZNF692 with shRNA produced the opposite effect, demonstrating that ZNF692 regulates the expression of HSF4. In this case, through HSF4 signaling, we observed that ZNF692 increased COAD cell proliferation.
Conclusion
The expression of ZNF692 is considerably elevated in COAD tissues and cell lines. Furthermore, ZNF692 increases colorectal cancer growth by modulating HSF4 expression. As a result, the current findings clearly imply that ZNF692 is a carcinogen in COAD and might be a new prognostic marker for the illness.
Journalism Ethics considerations
Ethical issues (Including plagiarism, informed consent, misconduct, data fabrication and/or falsification, double publication and/or submission, redundancy, etc.) have been completely observed by the authors.
Acknowledgements
This work was supported by the Basic research funding for higher education institutions in Heilongjiang Province in 2021(2021-KYYWF-0360).
Footnotes
Conflict of Interest
The authors declare that there is no conflict of interest.
References
- 1.Wong KE, Ngai SC, Chan KG, Lee LH, Goh BH, Chuah LH. (2019). Curcumin nanoformulations for colorectal cancer: A review. Front Pharmacol, 10:152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Edwards BK, Ward E, Kohler BA, et al. (2010). Annual report to the nation on the status of cancer, 1975–2006, featuring colorectal cancer trends and impact of interventions (risk factors, screening, and treatment) to reduce future rates. Cancer, 116:544–573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hao Y, Wang Y, Qi M, He X, Zhu Y, Hong J. (2020). Risk factors for recurrent colorectal polyps. Gut Liver,14:399–411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Pohl C, Hombach A, Kruis W. (2000). Chronic inflammatory bowel disease and cancer. Hepatogastroenterology, 47:57–70. [PubMed] [Google Scholar]
- 5.Mehta A, Patel BM. (2019). Therapeutic opportunities in COAD: Focus on phosphodiesterase inhibitions. Life Sci,230:150–161. [DOI] [PubMed] [Google Scholar]
- 6.Valdivia MM, Hamdouch K, Ortiz M, Astola A. (2009). CENPA a genomic marker for centromere activity and human diseases. Curr Genomics, 10(5):326–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Howman EV, Fowler KJ, Newson AJ, et al. (2000). Early disruption of centromeric chromatin organization in centromere protein A (Cenpa) null mice. Proc Natl Acad Sci U S A, 97(3):1148–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Heun P, Erhardt S, Blower MD, Weiss S, Skora AD, Karpen GH. (2006). Mislocalization of the Drosophila centromere-specific histone CID promotes formation of functional ectopic kinetochores. Dev Cell, 10(3):303–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Shrestha RL, Ahn GS, Staples MI, et al. (2017). Mislocalization of centromeric histone H3 variant CENP-A contributes to chromosomal instability (CIN) in human cells. Oncotarget, 8(29):46781–46800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zhang W, Mao JH, Zhu W, et al. (2016). Centromere and kinetochore gene misexpression predicts cancer patient survival and response to radiotherapy and chemotherapy. Nat Commun, 7:12619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zhang W, Karpen GH, Zhang Q. (2017). Exploring the role of CENP-A Ser18 phosphorylation in CIN and tumorigenesis. Cell Cycle, 16(24):2323–2325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Sharma AB, Dimitrov S, Hamiche A, Van Dyck E. (2019). Centromeric and ectopic assembly of CENP-A chromatin in health and cancer: old marks and new tracks. Nucleic Acids Res, 47(3):1051–1069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Sullivan LL, Boivin CD, Mravinac B, Song IY, Sullivan BA. (2011). Genomic size of CENPA domain is proportional to total alpha satellite array size at human centromeres and expands in cancer cells. Chromosome Res, 19(4):457–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Athwal RK, Walkiewicz MP, Baek S, et al. (2015). CENP-A nucleosomes localize to transcription factor hotspots and subtelomeric sites in human cancer cells. Epigenetics Chromatin, 8:2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.McGovern SL, Qi Y, Pusztai L, Symmans WF, Buchholz TA. (2012). Centromere protein-A, an essential centromere protein, is a prognostic marker for relapse in estrogen receptor-positive breast cancer. Breast Cancer Res, 14(3):R72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. (2018). Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin, 68(6):394–424. [DOI] [PubMed] [Google Scholar]
- 17.Arnold CN, Goel A, Blum HE, Boland CR. (2005). Molecular pathogenesis of colorectal cancer: implications for molecular diagnosis. Cancer, 104(10):2035–2047. [DOI] [PubMed] [Google Scholar]
- 18.Vogel JD, Eskicioglu C, Weiser MR, Feingold DL, Steele SR. (2017). The American Society of Colon and Rectal Surgeons clinical practice guidelines for the treatment of COAD. Dis Colon Rectum, 60(10):999–1017. [DOI] [PubMed] [Google Scholar]
- 19.Jones S, Chen WD, Parmigiani G, et al. (2008). Comparative lesion sequencing provides insights into tumor evolution. Proc Natl Acad Sci U S A, 105(11):4283–4288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Jeffery M, Hickey BE, Hider PN. (2019). Follow-up strategies for patients treated for nonmetastatic colorectal cancer. Cochrane Database Syst Rev, 9:CD002200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Xing Y, Zhao Z, Zhu Y, Zhao L, Zhu A, Piao D. (2018). Comprehensive analysis of differential expression profiles of mRNAs and lncRNAs and identification of a 14-lncRNA prognostic signature for patients with colon adenocarcinoma. Oncol Rep, 39:2365–2375. [DOI] [PubMed] [Google Scholar]
- 22.Dahmane N, Lee J, Robins P, Heller P, Ruiz i Altaba A. (1997). Activation of the transcription factor Gli1 and the Sonic hedgehog signalling pathway in skin tumours. Nature, 389:876–881. [DOI] [PubMed] [Google Scholar]
- 23.Nonet GH, Stampfer MR, Chin K, Gray JW, Collins CC, Yaswen P. (2001). The ZNF217 gene amplified in breast cancers promotes immortalization of human mammary epithelial cells. Cancer Res, 61:1250–1254. [PubMed] [Google Scholar]
- 24.Imoto I, Yuki Y, Sonoda I, et al. (2003). Identification of ZASC1 encoding a Krüppel-like zinc finger protein as a novel target for 3q26 amplification in esophageal squamous cell carcinomas. Cancer Res, 63:5691–5696. [PubMed] [Google Scholar]
- 25.Jiang R, Wang JC, Sun M, Zhang XY, Wu H. (2012). Zinc finger X-chromosomal protein (ZFX) promotes solid agar colony growth of osteosarcoma cells. Oncol Res, 20:565–570. [DOI] [PubMed] [Google Scholar]
- 26.Lai KP, Chen J, He M, et al. (2014). Overexpression of ZFX confers self-renewal and chemo-resistance properties in hepatocellular carcinoma. Int J Cancer, 135:1790–1799. [DOI] [PubMed] [Google Scholar]
- 27.Fang X, Huang Z, Zhou W, et al. (2014). The zinc finger transcription factor ZFX is required for maintaining the tumorigenic potential of glioblastoma stem cells. Stem Cells, 32:2033–2047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yan X, Shan Z, Yan L, et al. (2016). High expression of Zinc-finger protein X-linked promotes tumor growth and predicts a poor outcome for stage II/III colorectal cancer patients. Oncotarget, 7:19680–19692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Robinson MD, McCarthy DJ, Smyth GK. (2010). edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics, 26:139–140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Edge SB, Compton CC. (2010). The American Joint Committee on Cancer: the 7th Edition of the AJCC Cancer Staging Manual and the Future of TNM. Ann Surg Oncol, 17:1471–1474. [DOI] [PubMed] [Google Scholar]