Abstract.
Purpose
Diagnostic performance of prostate MRI depends on high-quality imaging. Prostate MRI quality is inversely proportional to the amount of rectal gas and distention. Early detection of poor-quality MRI may enable intervention to remove gas or exam rescheduling, saving time. We developed a machine learning based quality prediction of yet-to-be acquired MRI images solely based on MRI rapid localizer sequence, which can be acquired in a few seconds.
Approach
The dataset consists of 213 (147 for training and 64 for testing) prostate sagittal T2-weighted (T2W) MRI localizer images and rectal content, manually labeled by an expert radiologist. Each MRI localizer contains seven two-dimensional (2D) slices of the patient, accompanied by manual segmentations of rectum for each slice. Cascaded and end-to-end deep learning models were used to predict the quality of yet-to-be T2W, DWI, and apparent diffusion coefficient (ADC) MRI images. Predictions were compared to quality scores determined by the experts using area under the receiver operator characteristic curve and intra-class correlation coefficient.
Results
In the test set of 64 patients, optimal versus suboptimal exams occurred in 95.3% (61/64) versus 4.7% (3/64) for T2W, 90.6% (58/64) versus 9.4% (6/64) for DWI, and 89.1% (57/64) versus 10.9% (7/64) for ADC. The best performing segmentation model was 2D U-Net with ResNet-34 encoder and ImageNet weights. The best performing classifier was the radiomics based classifier.
Conclusions
A radiomics based classifier applied to localizer images achieves accurate diagnosis of subsequent image quality for T2W, DWI, and ADC prostate MRI sequences.
Keywords: prostate, magnetic resonance imaging MRI, quality, gas, artifact
1. Introduction
Prostate magnetic resonance imaging (MRI) diagnostic test accuracy for tumor localization and local staging is dependent on a variety of patient, radiologist, and proceduralist factors that may be of influence.1 Chief among patient factors is the amount of rectal distention and rectal gas present at time of MRI. Rectal distention, in particular distention with gas, correlates inversely with prostate MRI quality.1,2 Diffusion weighted imaging (DWI) is a critical sequence for evaluation of the prostate and DWI quality is highly influenced by rectal gas, which creates magnetic susceptibility artifacts that are magnified with echo-planar imaging techniques. As practices move toward bi-parametric MRI protocols that eliminate dynamic contrast enhanced (DCE)-MRI,3,4 poor quality DWI may render a study non-diagnostic requiring costly repeat exams, which reduce access to MRI, delay patient care, and create patient morbidity. A variety of patient preparation techniques have been explored prior to prostate MRI scanning, including cleansing enema and dietary modification.5,6 These techniques are effective in many patients, but are dependent on patient compliance and may not be necessary in all patients. The use of a rectal tube, inserted at time of MRI, to withdraw unwanted rectal gas is an option that has been proposed for patients with too much rectal gas.7 The use of a rectal catheter was shown to improve image quality in one study.8 A microenema can also be administered at time of MRI, in patients discovered with too much rectal gas at time of MRI.9
A critical aspect of prostate MRI quality is immediate recognition of rectal distention or rectal gas, which may negatively impact the MRI exam. At this step, the intervention by having the patient evacuate rectal contents in an adjacent restroom or by manual withdrawal of rectal gas by catheter insertion could salvage an otherwise limited or non-diagnostic exam.10 MRI technologists can be trained to recognize a distended rectum with gas; however, this is operator dependent and not all patients with rectal distention or rectal gas have the same poor diagnostic quality MRI. Prior to any prostate MRI scans, a rapid localizer scan is acquired to locate the prostate. We propose to utilize this MRI rapid localizer to predict the rectal content and in turn, the quality of the yet to be acquired MRI images using cascaded and end-to-end approaches. The use of an automated technique that can automatically detect and segment the rectum and quantify rectal volume and content using rapid MRI localizer sequences may predict quality of subsequent prostate MRI exam. If successful, this system could alert a technologist to intervene and seek rectal evacuation prior to continuing the exam. To date, application of machine learning techniques for prostate MRI quality have focused on quantifying the quality of the diagnostic prostate MRI images.11–13 While this is of importance at time of reporting, predicting the quality of the diagnostic images as optimal and suboptimal before they have been acquired could be more impactful enabling potential intervention to salvage the exam or termination and rescheduling to save time. The purpose of this study was therefore to develop a fully automated method for rectal detection and segmentation, rectal volume estimation, rectal content determination, and ultimately prediction of image quality in T2-weighted (T2W), DWI, and apparent diffusion coefficient (ADC) MRI images using rapid three-plane MRI localizer. To our knowledge, this is the first study to utilize the rapid MRI localizer for prediction of image quality in subsequence high-resolution MRI images.
2. Materials and Methods
2.1. Patients and Image Acquisition
This study was approved by the local institutional review board with waiver of consent. 213 consecutive men (mean age of years) referred for prostate MRI at a single academic referral center between January, 2014, and March, 2016, were included. Examination indications included: pre-biopsy, prior negative biopsy, and active surveillance. Patients post-prostatectomy or pelvic radiotherapy and with hip prosthesis or other metallic implant were excluded. The data were available to the (BLINDED) through a data-sharing agreement.
All patients underwent multi-parametric MRI at 3 T using the same clinical system (Discovery 750W, GE Healthcare) at a tertiary care referral center for prostate. During the study dates, patients either underwent no preparation or a Fleet™ enema self-administered 3 h prior to MRI. A summary of the MRI protocol used at our institution is provided in Table 1.
Table 1.
Multi-parametric MRI technique [Integrated pelvic surface coils (32 channels) with activated spine coils (12 channels). Clinical 3 T system: discovery 750 W (General Electric, Milwaukee, Wisconsin] performed during the study period.
| T2 fast spin echoa | DWIb | T1 GREc dynamic contrast | |||
|---|---|---|---|---|---|
| Imaging plane | Coronal | Sagittal | Axial | Axial | Axial |
| Field of view (mm) | 220 × 220 | — | — | 220 × 220 | 220 × 220 |
| Matrix size | 320 × 256 | — | — | 128 × 80 | 128 × 128 |
| Slice thickness/gap (mm) | 4.0/0 | 3.0/0 | 3.0/0 | 4.0/0 | 4.0/0 |
| TR/TE (ms) | 5250/125 | — | — | 4200/90 | 4.3/1.3 |
| Echo train length | 35 | — | — | 1 | N/A |
| Flip angle | 111 | — | — | 90 | 12 |
| Acceleration factor | N/A | — | — | 2 | 2 |
| Receiver bandwidth (Hz/voxel) | 122 | — | — | 1950 | 488 |
| Acquisition time (min) | 4 | 4 | 4 | 5 | 2 |
| Number of signals averaged | 2 | — | — | 10-Apr | 1 |
Fast spin echo.
DWI, diffusion weighted imaging performed with spectral fat suppression echo planar imaging with tri-directional motion probing gradients and values of 0, 500, with automatic ADC map generation derived. DWI acquired or calculated separately.
Dynamic fast spoiled 3D gradient recalled echo performed with a temporal resolution of 9 s after injection of 0.1 mmol/kg of gadobutrol (Gadovist, Bayer Inc., Toronto, Ontario) at a rate of 3 mL/s.
For the purposes of this study, the three-plane localizer sequence, which is performed at the start of every MRI examination was used for rectal segmentation. This is a large field of view (FOV = 40 cm, matrix ) T2W (TE = 72 ms, TR 640 ms, flip angle 90) single shot fast spin echo (ssFSE) free-breathing sequence with 10 mm thick slices, which acquires 21 images (seven each in the axial, coronal, and sagittal planes) in . Localizer sequences are performed first in all prostate MRI examinations across institutions and additional prostate MRI sequences are typically planned from the localizer images, which are otherwise not generally used for diagnosis.
2.2. Manual Rectal Segmentation and Rectal Content Determination
Rectum was segmented by a medical imaging student working with a fellowship trained genitourinary radiologist with 10 years of post-fellowship experience in prostate MRI (BLINDED). The student and radiologist manually segmented the rectum from the anal verge to the sacral promontory (approximate level of the rectosigmoid junction) on all sagittal images in which the rectum was visible, Fig. 1, using ITKSnap 3.8.0. Manual segmentation was performed on 213 patients. Rectal content was judged subjectively, by the radiologist, as follows: 0-collapsed rectum, no significant rectal content; 1-mostly solid stool, 2-mostly liquid stool, 3-mostly gas.
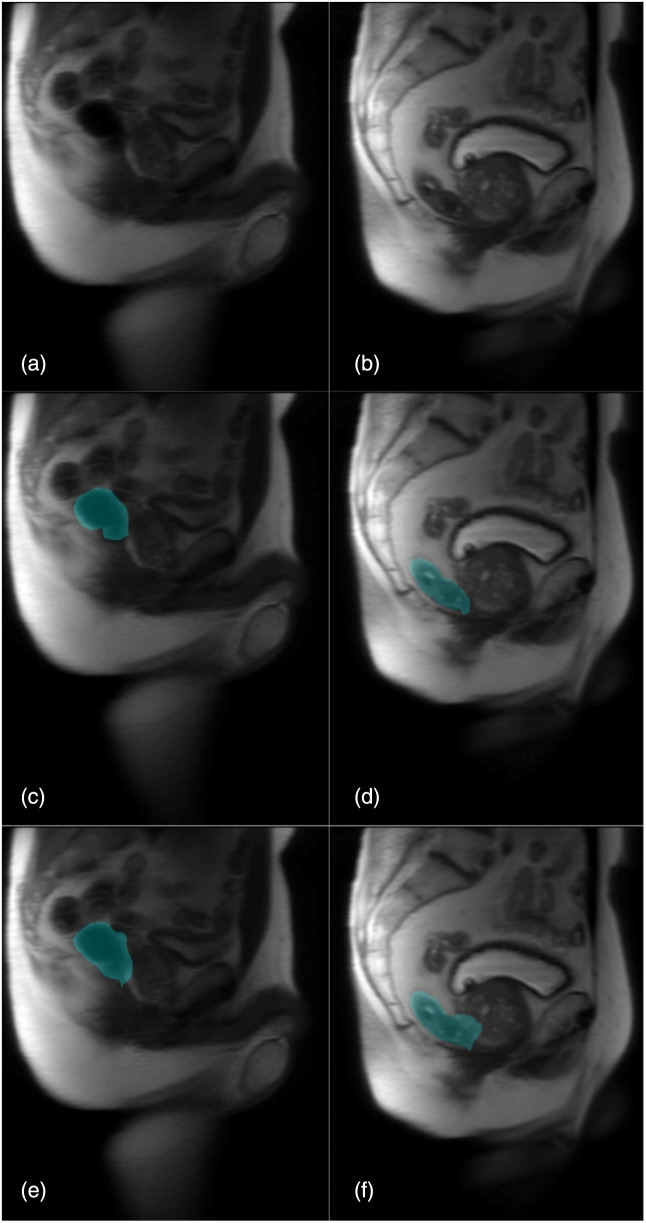
Fig. 1.
Sagittal T2W large field of view ssFSE MRI-localizer images in two different patients (a) and (b). Segmentation in (c) and (d) corresponds to manual segmentation. In (c), the rectum is decompressed and empty, image quality of subsequent diagnostic T2W, DWI, and ADC map images were rated as excellent. In (d), the rectum is distended with gas, image quality of the subsequent diagnostic T2W, DWI, and ADC map images were rated as suboptimal or non-diagnostic. In (e) and (f), the predicted segmentations for patients in (a) and (b), respectively, are shown.
Overall image quality scores for T2W, DWI, and ADC were subjectively based on the radiologist’s experience reporting prostate MRI and were rated using a five-point Likert type scale where: 1 = non-diagnostic, 2 = below average image quality but remains diagnostic, 3 = average image quality, 4 = above average image quality, and 5 = excellent image quality as described previously.14 In this study, a rating of 1 (non-diagnostic) and 2 (below average) was considered suboptimal compared to a rating of , which was considered acceptable or optimal.
2.3. Overview of MRI Image Quality Prediction Pipeline
The proposed MRI image quality prediction pipeline is shown in Fig. 2. The pipeline utilizes a cascaded approach, where an input image is input into a segmentation model, and the generated mask is input into a classification to generate a prediction. The segmentation stage is applied on the input images, where the rectum is located and a corresponding segmentation mask is generated. The generated segmentation masks will be used as one of the inputs for classification to identify the rectum’s content based on the mask and then derive a quality prediction. The data were first pre-processed using image intensity normalization and down-sampling. Data augmentation was then applied to synthetically increase the number of samples by introducing images of different orientations and sizes, and to also imitate clinical settings where the images acquired are not perfectly aligned in all axes. Three deep learning models were experimented with for rectal segmentation. The segmentation models tested were two-dimensional (2D) U-Net, attention U-Net, and U-Net++, with separate experiments using ImageNet pre-trained weights and ResNet-34 and VGG-16 as an encoder. 2D U-Net contains skip connections between the encoder and decoder for spatial content and context preservation.14 Attention U-Net integrates attention gates into the U-Net architecture, which allows the model to focus on specific regions of the image by weighting the feature maps. This mechanism helps in emphasizing certain structures or regions in the image, making it particularly useful for medical imaging where certain structures or anomalies are of primary interest. U-Net++ extends the notion of skip connections in 2D-U-Net by utilizing nested and densely connected skip connections for better spatial information preservation between the encoder and decoder.15 ImageNet pre-trained weights, and ResNet-34, and VGG-16 as encoders were used for the segmentation models to utilize transfer learning. VGG-16 uses small convolutional filters, whereas ResNet-34 uses residual networks for model training.16 Both networks have shown impressive performance in segmentation tasks, especially when pre-trained on the ImageNet dataset.16
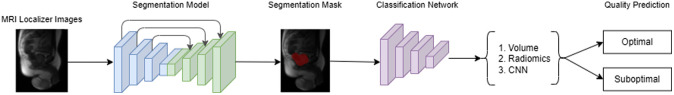
Fig. 2.
Automated quality prediction pipeline. An image is an input into the segmentation model, where a deep learning model generates rectum segmentation masks. The image along with the predicted segmentation mask is passed into the classification network to generate a quality output as either optimal or suboptimal. The classification can be based on rectal content, deep learning models, or using radiologist-assigned quality scores as features. The output is a final prediction for the yet to be acquired T2W, DWI, and ADC images.
Given the nature of the dataset, where data sparsity is a prevalent issue, transfer learning with pre-trained weights from ImageNet was experimented with. ImageNet is a dataset that contains millions of images and has been used as the de facto dataset for several computer vision tasks. Deep learning models that are trained on the ImageNet dataset have shown better performance compared to networks that are trained from scratch.17 We experimented using different backbones to the base 2D U-Net model and U-Net++ using ResNet-34 and VGG-16 as the encoder, and ImageNet pre-trained weights to utilize transfer learning.
Classification of images to generate a quality score as optimal or suboptimal was performed through three approaches: utilizing radiomic-based classifier deep learning-based classifier and using algorithm-generated rectal volume computed from the U-Net models as a feature. The best performing classifier in terms of evaluation metrics was then used to predict quality. Quality prediction was performed in two ways: (1) using the radiologist assigned quality labels as labels for the classifiers mentioned before, therefore, each classifier would be predicting a quality score, and (2) using rectal content as a label for the classification models, therefore, the predicted rectal content would be translated to a quality score. Rectal content is considered as an intermediate step, where a quality score will be generated based on the predicted outcome of rectal content. We used two classification labels to experiment on whether using rectal content as an intermediate label improves quality prediction compared to direct radiologist quality labels.
Radiomic features within the rectum were first extracted using a Python package named Pyradiomics18 and were used in classification via XGBoost. CNN based classifiers experimented with were ResNet-34, VGG-16, Inception V3, and DenseNet169. Similar to radiomic features, rectal volume was computed and used as a feature in classification via XGBoost.
The patient data were split into 70% train and 30% test. Image pixel intensities were normalized between 0 and 1, and the images were downsampled to . The image size selected was based on resource availability to train the deep learning models. Image augmentation techniques used were rotation of 100 deg, flipping along the vertical axis, and elastic affine transforms of (, ). The choice of augmentations was to synthetically increase the number of data samples and to provide more scenarios for the trained network to be exposed to. The features were normalized to a zero mean and unit variance for them to be treated equally. Since the MRI rapid localizer images are noisy and relatively low quality, image denoising using random Gaussian blur with a probability of 0.5 was performed. Gaussian blurring smoothes the image, which eases the detection of rectal boundaries. Finally, contrast enhancement was performed to increase the contrast of the rectum within the image using ITKSnap. The minimum and maximum intensity values were determined via visual inspection. Adaptive histogram equalization was utilized over regular histogram equalization because of the varying levels of contrast across the image.19 It was noticed that the lower extremities of the image were deemed to be of lower contrast than the content in the middle and higher locations.
Given the heavy class imbalance of data present, synthetic minority over-sampling technique (SMOTE) was utilized.20 A severe class imbalance can lead to both deep learning based and machine learning-based models having a bias to the dominant class, thus generating inaccurate classifications.21,22 SMOTE synthetically over samples minority classes, in this case empty, fluid, fluid + gas classes, to generate an even data split.
2.4. Deep Learning Based Image Quality Prediction
The CNN-based classification models used for image quality prediction were ResNet-34, VGG-16, Inception V3, and DenseNet169. ResNet-34 is a 34 convolutional layer network that utilizes residual blocks with skip connections.23 VGG-16 addresses the potential issues that come with deep networks, thus, VGG-16 reduces number of parameters and training time.23 Inception V3 is composed of several kernel sizes within a convolutional layer and concatenates spatial information as the output map.24 The size of kernels increases with the increase of convolutional layers numbers instead of the network becoming deeper.24 This variability in kernel sizes aims to extract as much information from a network as possible, with the hopes of not neglecting any spatial regions that may be critical to the segmentation mask.25 Finally, DenseNet was used to address the potential problem of vanishing gradients.26 Each layer in the network receives two inputs: the feature map from the direct previous layer and a concatenated output from all the previous layers as an input to the current layer.26,27 This connectivity pattern aims to preserve spatial information and also is more parameter efficient.26,27
The settings for the CNN classifiers were set to a kernel size of , stride of 2, and zero padding of . Cross entropy loss was used as the loss function, and Adam optimizer with a learning rate of was used. The model was trained for 200 epochs and 10-fold cross validation was applied.
2.6. End-to-End Quality Prediction Pipeline
In addition to the two-stage approach, an end-to-end convolutional neural network (CNN) model was developed for joint segmentation and classification. The model architecture consists of a ResNet-34 encoder backbone, followed by separate decoder and classifier heads. The encoder extracts 64-dimensional feature representations from each localizer image. The decoder module upsamples these features using transpose convolutions and skip connections to produce a pixel-wise rectum segmentation mask. The classifier head passes the encoder features through two fully connected layers of 128 and 64 units, followed by a sigmoid output layer to predict image quality. The model is trained end-to-end using a combined weighted loss function:
where and are the soft dice loss and binary cross-entropy losses for segmentation and classification, respectively. and are the weights to balance the losses. The combined loss enables joint optimization of both tasks. During training, online data augmentation including rotations, scaling, and brightness changes is used to improve generalization. The model is trained for 100 epochs using Adam optimization with a learning rate of and a batch size of 16 images. During inference, the end-to-end model takes a localizer image as input and directly outputs both the rectum segmentation mask and predicted image quality score in a single forward pass.
2.7. Radiomics Features and Volume Based Image Quality Prediction
Radiomics is a quantitative analysis approach that includes size, shape, and textural features extraction of the region of interest.18,28 Radiomics and its features are based on the assumption that biomedical images contain information of disease specific processes that are hard to visualize through the human eye, thus not accessible via traditional visual inspection techniques of images. Recently, several studies started utilizing radiomics to improve diagnostic and predictive performance of models in prostate cancer related problems.29–31 Radiomics enable medical images to be converted to higher dimension, mineable, and quantitative features for further analysis by machine learning models.32,33
Radiomic features were extracted using Pyradiomics library. The main textural features extracted when analyzing the radiomics of a region of interest are the first order statistics, 2D and three-dimensional (3D) shaped based features, and gray level cooccurrence/run length/size zone/gray tone difference/dependence matrix.34 This set of feature classes generates features per image, which consist of 16 shape descriptors and features extracted from original and derived images.34,35 To mitigate overfitting risk, we applied Boruta for feature selection. Other dimensionality reduction techniques, such as principal component analysis (PCA), were considered. Boruta was ultimately chosen because it selects features based on predictive importance for the specific prediction task. By comparison, PCA selects features that explain variance in the data but not necessarily predictive variance. For example, PCA would retain highly correlated features that explain significant variance but provide redundant information for prediction. In contrast, Boruta explicitly penalizes correlated features and removes uninformative features that simply fit noise in the data. By selecting features according to importance for predicting prostate MRI quality, Boruta provides superior dimensionality reduction compared to variance-based techniques, such as PCA, that do not consider the prediction task. This enables retaining only the most useful features for optimizing model performance. Boruta removes features that are highly correlated or have low correlation with the target variable. This is important because highly correlated features provide redundant information and inflate model variance. Features with low correlation to the target provide little useful signal and can allow fitting to noise. Boruta ranks features by importance calculated using a random forest classifier. At each iteration, it compares real features to randomly permuted copies to assess significance. Features that are significantly more important than random are selected. This process removed the vast majority of the initial 1500 features, reducing the set to only the most predictive and independent features. The final Boruta selected feature set contained 10 features. We chose 10 features based on observing the sharp decline in importance scores after the top 10 features in Boruta ranking. The 11th and later features did not improve cross-validation accuracy, indicating the top 10 captured the majority of useful signal in the data. Selecting 10 features avoided retaining irrelevant features while focusing on a meaningful but compact set of predictive radiomic signatures. The radiomic features used in our method were all original features and percentile deviations offered by the library were discarded.18,28 Rectal volume was computed by computing the number of voxels within the segmented mask and then multiplying it by the voxel size. The voxel size is the product of the , , and coordinates, which equates to the in-plane resolution multiplied by the slice thickness in the coordinate.
Several classification algorithms were tested using the two types of features mentioned earlier. The first classification algorithm tested was random forests. Random forests comprised of decision trees ensembles that utilize feature randomness and bagging when constructing each individual tree.36 The randomness in constructing trees avoid overfitting that may be caused by an individual tree.36 Following random forests, tree boosting systems were considered as they tend to outperform base tree-based algorithms.37 Tree boosting is beneficial as it capitalizes on weak learners within a tree to improve on its predictions.37 AdaBoost and extreme gradient boosting (XGBoost) were both investigated. XGBoost was selected because it performs well for sparse data,37 which is the case for this dataset. XGBoost has been shown to outperform conventional machine learning and deep learning-based algorithms in terms of accuracy and inference speed.38 10-fold cross validation during training was implemented.
2.8. Statistical Analysis and Evaluation Metrics
Radiologist assigned quality scores were grouped into suboptimal (scores of 1 and 2) and average or optimal (scores 3, 4, and 5), the latter representing a diagnostic image quality. Similarly, when using the predicted rectal content classifications and mapping them to a quality score, a rectal content of either empty or fluid would be classed as optimal, whereas a rectal content of either fluid + gas or gas would be classed as suboptimal.
To evaluate the performance of the segmentation models, Dice score coefficient (DSC) confidence intervals were used. To assess the performance of the classification models, area under the receiver operator characteristic curve (AUC) confidence intervals across the cross-validation folds were used. Quality assessment agreement between the radiologist and the machine learning algorithm was assessed using intra-class correlation coefficient (ICC). Agreement scores between the two evaluation methods indicate poor reliability if ICC is below 0.50, moderate reliability for values between 0.50 and 0.75, good reliability for values between 0.75 and 0.90, and excellent reliability for values of 0.90 and above.39,40 All statistical analyses were performed using Scikit-learn on Python 3.7.
3. Results
The test set consisted of 64 unseen patient data. The manually determined quality scores determined by the radiologist were classified as average or optimal (quality score ) versus suboptimal (quality score ) on the test set as follows: 95.3% (61/64) versus 4.7% (3/64) for T2W, 90.6% (58/64) versus 9.4% (6/64) for DWI, and 89.1% (57/64) versus 10.9% (7/64) for ADC (Table 2). The patients’ rectal content as rated by the radiologist was: 23.4% (15/64) empty, 10.9% (7/64) fluid, 10.9% (7/64) fluid + gas, and 54.7% (35/64) gas (Table 3).
Table 2.
Radiologist assigned image quality scores for the train and test set.
| Quality scores | Sequence | ||
|---|---|---|---|
| T2Wa | ADCb | DWIc | |
|
Train set
| |||
| Quality score 1 (non-diagnostic) | 1 | 3 | 5 |
| Quality score 2 (below average) | 5 | 7 | 10 |
| Quality score 3 (average) | 18 | 22 | 22 |
| Quality score 4 (above average) | 54 | 61 | 37 |
| Quality score 5 (excellent) | 69 | 54 | 73 |
| Overall optimal exam (quality score ≥3) | 95.9% (141/147) | 93.2% (137/147) | 89.8% (132/147) |
|
Overall suboptimal exam (quality score ≤2)
|
4.1% (6/147) |
6.8% (10/147) |
10.2% (15/147) |
|
Test set
| |||
| Quality score 1 (non-diagnostic) | 1 | 2 | 2 |
| Quality score 2 (below average) | 2 | 4 | 5 |
| Quality score 3 (average) | 30 | 27 | 18 |
| Quality score 4 (above average) | 13 | 16 | 15 |
| Quality score 5 (excellent) | 18 | 15 | 24 |
| Optimal exams | 95.3% (61/64) | 90.6% (58/64) | 89.1% (57/64) |
| Suboptimal exams | 4.7% (3/64) | 9.4% (6/64) | 10.9% (7/64) |
T2-weighted.
Apparent diffusion coefficient.
Diffusion weighted imaging.
Table 3.
Ground truth radiologist assigned rectal content for the train and test set.
| Rectal content | Empty | Fluid | Fluid + gas | Gas |
|---|---|---|---|---|
|
Train set
| ||||
| Proportion of patients |
23.8% (35/147) |
14.3% (21/147) |
11.6% (17/147) |
50.3% (74/147) |
|
Test set
| ||||
| Proportion of patients | 23.4% (15/64) | 10.9% (7/64) | 10.9% (7/64) | 54.7% (35/64) |
In terms of segmentation, 2D U-Net, using ResNet-34 encoder and ImageNet weights for transfer learning, resulted with a DSC of . 2D U-Net with VGG-16 as an encoder and ImageNet weights resulted with a DSC of . Attention U-Net resulted with a DSC of . Finally, U-Net++ with ResNet-34 encoder and ImageNet weights resulted with a DSC of (Table 4).
Table 4.
Results for segmentation of various models compared to ground truth segmentation of rectum by radiologist.
| Model | DSCa (mean ± 95% CIb) |
|---|---|
| 2D U-Net | 0.79 ± 0.07 |
| 2D U-Net (ResNet-34 encoder) | 0.83 ± 0.04 |
| 2D U-Net (ResNet-34 encoder + ImageNet weights) | 0.85 ± 0.07 |
| 2D U-Net (VGG-16 encoder + ImageNet weights) | 0.84 ± 0.06 |
| U-Net ++ | 0.80 ± 0.05 |
| Attention U-Net | 0.82 ± 0.05 |
| U-Net ++ (ResNet-34 encoder + ImageNet weights) | 84.84 ± 0.05 |
Dice score coefficient.
95% confidence interval.
Using radiologist assigned image quality scores as labels for our classification models, a radiomics based classifier predicted the following classification for image quality (average or optimal versus suboptimal): 87.5% (56/64) versus 12.5% (8/64) for T2W, 82.8% (53/64) versus 17.2% (11/64) for DWI, and 81.2% (52/64) versus 18.8% (12/64) for ADC. The CNN based classifier resulted in the following predictions (average or optimal versus suboptimal): 79.7% (51/64) versus 20.3% (13/64) for T2W, 76.5% (49/64) versus 23.4% (15/64) for DWI, and 76.5% (49/64) versus 23.4% (15/64) for ADC. The end-to-end based classifier resulted in the following predictions (average or optimal versus suboptimal): 75% (48/64) versus 25% (16/64) for T2W, 76.5% (49/64) versus 23.4% (15/64) for DWI, and 79.7% (51/64) versus 20.3% (13/64) for ADC. Finally, the volume based classifier resulted in the following predictions (average or optimal versus suboptimal): 70.3% (45/64) versus 29.7% (19/64) for T2W, 67.2% (43/64) versus 32.8% (21/64) for DWI, and 71.9% (46/64) versus 28.1% (18/64) for ADC. An ensemble classification model using radiomic, CNN, and volume as features resulted in the following predictions (average or optimal versus suboptimal): 85.9% (55/64) versus 14.1% (9/64) for T2W, 82.8% (53/64) versus 17.2% (11/64) for DWI, and 78.1% (50/64) versus 21.9% (14/64) for ADC. Results are summarized in Table 5.
Table 5.
Predicted quality scores using radiologist assigned quality scores as labels, and using rectal content as an intermediate step to determine quality as labels. Results represent predicted scores on the test set (64 images).
| Label | Model features | Sequence | |||
|---|---|---|---|---|---|
| T2W | ADC | DWI | |||
| Radiologist assigned quality scores | Volume (XGBoost) | Optimal exams | 70.3% (45/64) | 71.9% (46/64) | 67.2% (43/64) |
| Suboptimal exams | 29.7% (19/64) | 28.1% (18/64) | 32.8% (21/64) | ||
| CNN (ResNet-34) | Optimal exams | 79.7% (51/64) | 76.5% (49/64) | 76.5% (49/64) | |
| Suboptimal exams | 20.3% (13/64) | 23.4% (15/64) | 23.4% (15/64) | ||
| End-to-end | Optimal exams | 75% (48/64) | 76.5% (49/64) | 79.7% (51/64) | |
| Suboptimal exams | 25% (16/64) | 23.4% (15/64) | 20.3% (13/64) | ||
| Radiomics (XGBoost) | Optimal exams | 87.5% (56/64) | 81.2% (52/64) | 82.8% (53/64) | |
| Suboptimal exams | 12.5% (8/64) | 18.8% (12/64) | 17.2% (11/64) | ||
| Quality scores derived from rectal content | Radiomics (XGBoost) | Optimal exams | 82.8% (53/64) | 79.7% (51/64) | 78.1% (50/64) |
| Suboptimal exams | 17.2% (11/64) | 20.3% (13/64) | 21.9% (14/64) | ||
In terms of the classifiers’ performance, the radiomics based classifier, using quality scores as labels resulted in an AUC of for T2W, for ADC, and for DWI. The CNN based classifier, using radiologist assigned quality scores resulted with an AUC of for T2W, for ADC, for DWI. The end-to-end based quality predictor, using radiologist assigned quality scores resulted with an AUC of for T2W, for ADC, and for DWI. The rectal volume based classifier, using radiologist assigned quality scores as labels resulted with an AUC of for T2W, for ADC, and for DWI. Results are summarized in Table 6.
Table 6.
Optimal performing classification models for diagnosis of suboptimal image quality from localizer sequence after 10-fold cross validation using quality scores assigned by radiologist as labels.
| Model features | Sequence AUC ± 95% CI | ||
|---|---|---|---|
| T2W | ADC | DWI | |
| Volume (XGBoost) | 0.71 ± 0.04 | 0.70 ± 0.04 | 0.70 ± 0.04 |
| CNN (ResNet-34) | 0.84 ± 0.04 | 0.83 ± 0.02 | 0.83 ± 0.06 |
| End-to-end | 0.83 ± 0.05 | 0.83 ± 0.04 | 0.84 ± 0.04 |
| Radiomics (XGBoost) | 0.87 ± 0.03 | 0.84 ± 0.05 | 0.84 ± 0.04 |
When comparing quality evaluation of the radiologist and the best performing quality prediction algorithm, radiomics based classifier using assigned radiologist quality scores as labels, an ICC value of 0.77 was obtained, which shows a substantial agreement between the two quality scorers. When comparing the reliability of the worst performing quality scoring algorithm, rectal diameter using quality scores derived from rectal content as labels, an ICC of 0.59 was achieved, which indicates that there is a moderate agreement between the two scorers.
4. Discussion
Our study describes a fully automated pipeline to predict the yet to be acquired T2W, ADC, and DWI prostate MRI image quality from MRI-localizer images. The fully automated pipeline utilizes variants of 2D U-Net to generate segmentation masks, and several types of features to generate quality score predictions for T2W, DWI, and ADC sequences compared to radiologist assigned quality scores. Our results indicate that our fully automated quality estimation method achieved moderate-to-substantial agreement compared to radiologist determination of image quality of T2W, DWI, and ADC map images. Furthermore, we were able to decrease exams’ rejection rate from up to 20% down to a best of 1.5%. These results indicate that a fully automated method of estimating diagnostic image quality from localizer images is possible and could flag suboptimal or non-diagnostic exams enabling early intervention to improve image quality (e.g., catheter removal of gas or microenema) or exam rescheduling, saving MRI time, resources, and patient call-backs.
The CNN architectures implemented in this study to segment the rectum were able to delineate the contours of the rectum well. Prior work in rectum segmentation involved a focus on rectal tumor segmentation, with emphasis on rectum localization. Using T2W images, a mean DSC and a standard deviation of 0.70 () was achieved when the rectum is localized and the tumors are segmented.41 Knuth et al. achieved a median DSC score of 0.77 when performing rectal segmentation on T2W,42 and a median DSC of 0.76 when performing segmentations on DWI.42 As it can be observed, the DSC score is lower than what we achieved, and this can be attributed to the simple architectures used in the CNN design. When using plain 2D U-Net to segment the rectum, similar DSC has been achieved, but the metric improves with added complexities to the architectures design by using pre-trained weights and transfer learning. It is worth noting that our top performing segmentation algorithms performed similarly to each other, which infers that the choice of model architecture is not a critical aspect of the design. However, it should be noted that vanilla 2D U-Net and simpler variants did perform relatively poorer than more sophisticated models that utilized transfer learning and weights from bigger datasets. The difference between the best performing architectures, though minor, was evident in the regions of capture, where each model showed some bias toward different regions of the rectum. The reason as to why such bias exists is not fully understood, but a possibility to explain this phenomenon is the poor quality of localizer images, which may omit certain regions in a batch of images, and have them clearly shown in a different batch. A common theme among the best performing segmentation models was over segmentation, and given the application we are using it for, is appropriate. Under segmentation has the potential of discarding critical information from the rectum, thus deeming the predictions unrepresentative of the final image quality.
Experimenting and end-to-end pipeline to generate predictions was in consideration regarding the potential disadvantages of a two-stage training approach. In a two-stage approach, errors from the initial segmentation models can propagate to downstream image quality classification tasks, potentially impacting performance. Our results corroborate this concern, especially when observing the volume-based classification, which yielded inferior performance compared to other feature types. In contrast, the end-to-end model, which was developed to jointly handle segmentation and classification, showed a performance that was intermediate between the radiomics-based classifier and the volume-based classifier and similar to that of the CNN. This suggests that the end-to-end approach, by jointly optimizing both tasks, can mitigate some of the errors that might arise in a two-stage approach. The choice between the two approaches might also depend on the specific application and the importance of interpretability. While the two-stage approach allows for a clear separation between segmentation and classification, making it easier to pinpoint and address errors in each stage, the end-to-end approach offers a more streamlined process, potentially reducing computational time during inference. Another aspect to consider is the robustness of the model. While the end-to-end model might be more resilient to certain errors, it is crucial to ensure that it does not become a black box, making it harder to understand and interpret its predictions. Future work might explore hybrid approaches that combine the strengths of both methods or delve deeper into understanding the feature importance in the end-to-end model to ensure both high performance and interpretability.
Predicting the quality of the yet to be acquired T2W, ADC, and DWI based on localizer sequences through radiomics, CNNs, and rectal volume using radiologist assigned quality scores as labels, to the authors’ knowledge, has not been previously researched. In our classification models, radiomic features showed to be superior to those of CNN based features, which was surprising given that the number of feature parameters in CNNs is greater than that of radiomics. However, this could be explained by the heavy emphasis on gray level features generated in radiomics, which translate to more meaningful predictions. Volume features were also underperforming. This could be explained by the sparse number of features for the model to train on, compared to the hundreds in case of radiomics and thousands in case of CNNs. Another consideration is that rectal volume alone may not predict the final image quality. For example, a collapsed rectum containing a small amount of anterior gas could degrade DWI and, conversely, a rectum distended with fluid or solid stool may not degrade image quality.
The integration of a quality control pipeline to current MRI image acquisition pipelines could have a positive impact and salvaging potentially uninterpretable scans, saving time, and resources. Ideally, this automated pipeline would be implemented to current MRI technology as a feature of alerting technologists of a potentially uninterpretable scan from an MRI localizer image. At this stage, a technologist and radiologist could decide whether insertion of a rectal catheter to withdraw gas, patient attempt at a bowel movement (with or without the use of a micro-enema) or exam rescheduling might be considered rather than blindly acquiring the full prostate MRI acquisition (in cases of protocols with DCE MRI having unnecessary administration of Gadolinium based contrast agent).
Our study has limitations. There is a small sample size for training, with a total number of 213 patients. The localizer images also did not contain a large number of slices, with each patient having seven slices in each plane (axial, coronal, and sagittal). Image quality was evaluated using a previously validated five-point Likert type scale7,43,44 for T2W and DWI/ADC. A limitation of this work could be that we did not evaluate the MRI using PI-QUAL, a proposed standardized prostate quality assessment tool.45,46 We chose not to use PI-QUAL in this study because we aimed to determine quality of each individual sequence (rather than the study overall) and because DCE-MRI, which is included in PI-QUAL, was not assessed in this study since our institution presently applies bi-parametric MRI. We used augmentation techniques to synthetically increase the number of training images and to also imitate potential scenarios the image would be displayed as. Furthermore, SMOTE was applied to counter the data imbalance. Ideally, the data used for this study would have an even split for each of the classes in terms of rectal content and quality to avoid synthetic pre-processing steps. The images were also of low resolution, which posed several difficulties when training CNNs for both segmentation and classification and extracting relevant radiomic features. Some trial and error steps in terms of architectural design were performed to identify the ideal depth and number of convolutional layers of the network.
Acknowledgments
We acknowledge A.I. Vali Inc., Compute Canada, Compute Ontario, and SHARCNET for providing expertise and computing resources that made this research possible. Grant support was provided by the Mitacs Accelerate Scholarship and the Queen Elizabeth II Graduate Scholarship in Science and Technology (QEII-GSST).
Biographies
Eranga Ukwatta received his master’s and PhD degrees in electrical and computer engineering and biomedical engineering from Western University, Canada, in 2009 and 2013, respectively. From 2013 to 2015, he was a multicenter postdoctoral fellow with Johns Hopkins University and University of Toronto. He is currently an associate professor with the School of Engineering, University of Guelph, Canada, and an adjunct professor in Systems and Computer Engineering with Carleton University, Canada. He has been an author/coauthor of more than 100 journal articles and conference proceedings. His research interests include medical image segmentation and registration, deep learning for computer-aided diagnosis, and computational modeling. He is also an IEEE senior member and a professional engineer in Canada.
Biographies of the other authors are not available.
Contributor Information
Abdullah Al-Hayali, Email: aalhayal@uoguelph.ca.
Amin Komeili, Email: akomeili@ualberta.ca.
Azar Azad, Email: azar@aivali.org.
Paul Sathiadoss, Email: psathiadoss@toh.ca.
Nicola Schieda, Email: nschieda@toh.on.ca.
Eranga Ukwatta, Email: eukwatta@uoguelph.ca.
Disclosure
The authors have no financial conflicts of interest to disclose.
Code and Data Availability
The data are not publicly available due to the local institutional review board’s waiver of consent that indicates the data being used for this specific research project, and the data not being used farther than the confines of this research project as described in Sec. 2.1.
The full codebase for this project is not publicly available; however, core code scripts are available on the main author’s GitHub profile, and it can be used replicate the experiments conducted in this paper for different types of MRI data. The code can be found on GitHub:
References
- 1.Caglic I., et al. , “Evaluating the effect of rectal distension on prostate multiparametric MRI image quality,” Eur. J. Radiol. 90, 174–180 (2017). 10.1016/j.ejrad.2017.02.029 [DOI] [PubMed] [Google Scholar]
- 2.Giganti F., et al. , “Prostate imaging quality (PI-QUAL): a new quality control scoring system for multiparametric magnetic resonance imaging of the prostate from the precision trial,” Eur. Urol. Oncol. 3(5), 615–619 (2020). 10.1016/j.euo.2020.06.007 [DOI] [PubMed] [Google Scholar]
- 3.Choi M. H., et al. , “Prebiopsy biparametric MRI for clinically significant prostate cancer detection with PI-RADS version 2: a multicenter study,” Am. J. Roentgenol. 212(4), 839–846 (2019). 10.2214/AJR.18.20498 [DOI] [PubMed] [Google Scholar]
- 4.Barth B. K., et al. , “Detection of clinically significant prostate cancer: short dual-pulse sequence versus standard multiparametric MR imaging: a multireader study,” Radiology 284(3), 725–736 (2017). 10.1148/radiol.2017162020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Vincent M., Mellnick M. D., “Preparation strategies for prostate MRI,” Can. Assoc. Radiol. J. 73(2), 346–354 (2022). 10.1177/08465371211039215 [DOI] [PubMed] [Google Scholar]
- 6.Purysko A. S., et al. , “Influence of enema and dietary restrictions on prostate MR image quality: a multireader study,” Acad. Radiol. 29(1), 4–14 (2022). 10.1016/j.acra.2020.10.019 [DOI] [PubMed] [Google Scholar]
- 7.Lim C., et al. , “Does a cleansing enema improve image quality of 3T surface coil multiparametric prostate MRI?” J. Magn. Reson. Imaging 42(3), 689–697 (2015). 10.1002/jmri.24833 [DOI] [PubMed] [Google Scholar]
- 8.Huang Y. H., et al. , “Impact of 18-French rectal tube placement on image quality of multiparametric prostate MRI,” Am. J. Roentgenol. 217(4), 919–920 (2021). 10.2214/AJR.21.25732 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Plodeck V., et al. , “Rectal gas-induced susceptibility artefacts on prostate diffusion-weighted MRI with EPI read-out at 3.0 T: does a preparatory micro-enema improve image quality?” Abdom. Radiol. 45(12), 4244–4251 (2020). 10.1007/s00261-020-02600-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Weinreb J. C., et al. , “PI-RADS prostate imaging – reporting and data system: 2015, version 2,” Eur. Urol. 69(1), 16–40 (2016). 10.1016/j.eururo.2015.08.052 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cipollari S., et al. , “Convolutional neural networks for automated classification of prostate multiparametric magnetic resonance imaging based on image quality,” J. Magn. Reson. Imaging 55(2), 480–490 (2022). 10.1002/jmri.27879 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bosse S., et al. , “Deep neural networks for no-reference and full-reference image quality assessment,” IEEE Trans. Image Process. 27(1), 206–219 (2018). 10.1109/TIP.2017.2760518 [DOI] [PubMed] [Google Scholar]
- 13.Amirshahi S. A., Pedersen M., Yu S. X., “Image quality assessment by comparing CNN features between images,” J. Imaging Sci. Technol. 60(6), 060410 (2016). 10.2352/J.ImagingSci.Technol.2016.60.6.060410 [DOI] [Google Scholar]
- 14.Weng W., Zhu X., “I-Net: convolutional networks for biomedical image segmentation,” IEEE Access 9, 16591–16603 (2021). 10.1109/ACCESS.2021.3053408 [DOI] [Google Scholar]
- 15.Zhou Z., et al. , “UNet++: a nested U-Net architecture for medical image segmentation.” [DOI] [PMC free article] [PubMed]
- 16.He K., et al. , “Deep residual learning for image recognition,” http://image-net.org/challenges/LSVRC/2015/ (accessed June 8, 2022).
- 17.van Griethuysen J. J. M., et al. , “Computational radiomics system to decode the radiographic phenotype,” Cancer Res. 77(21), e104–e107 (2017). 10.1158/0008-5472.CAN-17-0339 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Pizer S. M., et al. , “Adaptive histogram equalization and its variations,” Comput. Vision Graphics Image Process. 39(3), 355–368 (1987). 10.1016/S0734-189X(87)80186-X [DOI] [Google Scholar]
- 19.Chawla N. V., et al. , “SMOTE: synthetic minority over-sampling technique,” J. Artif. Intell. Res. 16, 321–357 (2002). 10.1613/jair.953 [DOI] [Google Scholar]
- 20.Johnson J. M., Khoshgoftaar T. M., “Survey on deep learning with class imbalance,” J. Big Data 6(1), 27 (2019). 10.1186/S40537-019-0192-5/TABLES/18 [DOI] [Google Scholar]
- 21.“Learning imbalanced datasets with label-distribution-aware margin loss,” https://papers.nips.cc/paper/2019/hash/621461af90cadfdaf0e8d4cc25129f91-Abstract.html (accessed June 8, 2022).
- 22.Trebeschi S., et al. , “Deep learning for fully-automated localization and segmentation of rectal cancer on multiparametric MR,” Sci. Rep. 7(1), 5301 (2017). 10.1038/s41598-017-05728-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Soomro M. H., et al. , “Automated segmentation of colorectal tumor in 3D MRI using 3D multiscale densely connected convolutional neural network,” J. Healthc. Eng. 2019, 1075434 (2019). 10.1155/2019/1075434 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Huh M., Agrawal P., Efros A. A., “What makes ImageNet good for transfer learning?”
- 25.Simonyan K., Zisserman A., “Very deep convolutional networks for large-scale image recognition,” 2015, http://www.robots.ox.ac.uk/ (accessed June 8, 2022).
- 26.Szegedy C., et al. , “Rethinking the inception architecture for computer vision,” in Proc. IEEE Comput. Soc. Conf. Comput. Vision and Pattern Recognit., pp. 2818–2826 (2016). [Google Scholar]
- 27.Sharma N., et al. , “Automated medical image segmentation techniques,” J. Med. Phys. 35(1), 3–14 (2010). 10.4103/0971-6203.58777 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Steen M., et al. , “DenseNet: densely connected convolutional networks,” arXiv:1608.06993v5 28(4), 362–371 (2018). [Google Scholar]
- 29.Wang J., et al. , “Radiomics features on radiotherapy treatment planning CT can predict patient survival in locally advanced rectal cancer patients,” Sci. Rep. 9(1), 15346 (2019). 10.1038/s41598-019-51629-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wibmer A., et al. , “Haralick texture analysis of prostate MRI: utility for differentiating non-cancerous prostate from prostate cancer and differentiating prostate cancers with different Gleason scores,” Eur. Radiol. 25(10), 2840–2850 (2015). 10.1007/s00330-015-3701-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Nketiah G., et al. , “T2-weighted MRI-derived textural features reflect prostate cancer aggressiveness: preliminary results,” Eur. Radiol. 27(7), 3050–3059 (2017). 10.1007/s00330-016-4663-1 [DOI] [PubMed] [Google Scholar]
- 32.Ma S., et al. , “MRI-based radiomics signature for the preoperative prediction of extracapsular extension of prostate cancer,” J. Magn. Reson. Imaging 50(6), 1914–1925 (2019). 10.1002/jmri.26777 [DOI] [PubMed] [Google Scholar]
- 33.Aerts H. J. W. L., et al. , “Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach,” Nat. Commun. 5, 4006 (2014). 10.1038/ncomms5006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Huang Y. Q., et al. , “Development and validation of a radiomics nomogram for preoperative prediction of lymph node metastasis in colorectal cancer,” J. Clin. Oncol. 34(18), 2157–2164 (2016). 10.1200/JCO.2015.65.9128 [DOI] [PubMed] [Google Scholar]
- 35.Huang Y., et al. , “Radiomics signature: a potential biomarker for the prediction of disease-free survival in early-stage (I or II) non-small cell lung cancer,” Radiology 281(3), 947–957 (2016). 10.1148/radiol.2016152234 [DOI] [PubMed] [Google Scholar]
- 36.Sun Y., et al. , “Radiomic features of pretreatment MRI could identify T stage in patients with rectal cancer: preliminary findings,” J. Magn. Reson. Imaging 48(3), 615–621 (2018). 10.1002/jmri.25969 [DOI] [PubMed] [Google Scholar]
- 37.Dong Y., et al. , “Preoperative prediction of sentinel lymph node metastasis in breast cancer based on radiomics of T2-weighted fat-suppression and diffusion-weighted MRI,” Eur. Radiol. 28(2), 582–591 (2018). 10.1007/s00330-017-5005-7 [DOI] [PubMed] [Google Scholar]
- 38.Dashevsky B. Z., et al. , “MRI features predictive of negative surgical margins in patients with HER2 overexpressing breast cancer undergoing breast conservation,” Sci. Rep. 8(1), 315 (2018). 10.1038/s41598-017-18758-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Gnep K., et al. , “Haralick textural features on T2-weighted MRI are associated with biochemical recurrence following radiotherapy for peripheral zone prostate cancer,” J. Magn. Reson. Imaging 45(1), 103–117 (2017). 10.1002/jmri.25335 [DOI] [PubMed] [Google Scholar]
- 40.Bobak C. A., Barr P. J., O’Malley A. J., “Estimation of an inter-rater intra-class correlation coefficient that overcomes common assumption violations in the assessment of health measurement scales,” BMC Med. Res. Methodol. 18(1), 93 (2018). 10.1186/s12874-018-0550-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Vignati A., et al. , “Texture features on T2-weighted magnetic resonance imaging: new potential biomarkers for prostate cancer aggressiveness,” Phys. Med. Biol. 60(7), 2685–2701 (2015). 10.1088/0031-9155/60/7/2685 [DOI] [PubMed] [Google Scholar]
- 42.Knuth F., et al. , “MRI-based automatic segmentation of rectal cancer using 2D U-Net on two independent cohorts,” Acta Oncol. 61(2), 255–263 (2022). 10.1080/0284186X.2021.2013530 [DOI] [PubMed] [Google Scholar]
- 43.Tomaszewski M. R., Gillies R. J., “The biological meaning of radiomic features,” Radiology 298(3), 505 (2021). 10.1148/radiol.2021202553 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sathiadoss P., et al. , “Comparison of 5 rectal preparation strategies for prostate MRI and impact on image quality,” Can. Assoc. Radiol. J. 73(2), 346–354 (2022). 10.1177/08465371211033753 [DOI] [PubMed] [Google Scholar]
- 45.Lakshmanaprabu S. K., et al. , “Random forest for big data classification in the internet of things using optimal features,” Int. J. Mach. Learn. Cybern. 10(10), 2609–2618 (2019). 10.1007/s13042-018-00916-z [DOI] [Google Scholar]
- 46.Giganti F., et al. , “Understanding PI-QUAL for prostate MRI quality: a practical primer for radiologists,” Insights Imaging 12(1), 59 (2021). 10.1186/s13244-021-00996-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The data are not publicly available due to the local institutional review board’s waiver of consent that indicates the data being used for this specific research project, and the data not being used farther than the confines of this research project as described in Sec. 2.1.
The full codebase for this project is not publicly available; however, core code scripts are available on the main author’s GitHub profile, and it can be used replicate the experiments conducted in this paper for different types of MRI data. The code can be found on GitHub: