Abstract
Background
PTGS2 encodes cyclooxygenase‐2 (COX‐2), which catalyses the committed step in prostaglandin synthesis. Various in vivo and in vitro data suggest that COX‐2 mediates the VEGF signalling pathway. In silico analysis performed in TCGA, PanCancer Atlas for head and neck cancers, demonstrated significant expression and co‐expression of PTGS2 and genes that regulate VEGF signalling. This study was designed to elucidate the expression pattern of PTGS2 and genes regulating VEGF signalling in patients with locally advanced oral squamous cell carcinoma (OSCC).
Methodology
Tumour and normal tissue samples were collected from patients with locally advanced OSCC. RNA was isolated from tissue samples, followed by cDNA synthesis. The cDNA was used for gene expression analysis (RT‐PCR) using target‐specific primers. The results obtained were compared with the in silico gene expression of the target genes in the TCGA datasets. Co‐expression analysis was performed to establish an association between PTGS2 and VEGF signalling genes.
Results
Tumour and normal tissue samples were collected from 24 OSCC patients. Significant upregulation of PTGS2 expression was observed. Furthermore, VEGFA, KDR, CXCR1 and CXCR2 were significantly upregulated in tumour samples compared with paired normal samples, except for VEGFB, whose expression was not statistically significant. A similar expression pattern was observed in silico, except for CXCR2 which was highly expressed in the normal samples. Co‐expression analysis showed a significant positive correlation between PTGS2 and VEGF signalling genes, except for VEGFB which showed a negative correlation.
Conclusion
PTGS2 and VEGF signalling genes are upregulated in OSCC, which has a profound impact on clinical outcomes.
Keywords: angiogenesis, gene expression, metastasis, Oral cancer, PTGS2, VEGF
This study was designed to elucidate the expression pattern of PTGS2 and genes regulating VEGF signalling in patients with locally advanced oral squamous cell carcinoma. Significant upregulation of PTGS2, VEGFA, KDR, CXCR1, and CXCR2 expression were observed in tumour samples compared with paired normal samples. This study underscores the clinical relevance of these genes, pointing to potential links between disease aggressiveness and adverse prognostic factors.

1. INTRODUCTION
Oral squamous cell carcinoma (OSCC) is a global health concern, with global projections reporting 202,000 cases annually. 1 According to recent GLOBOCAN data, 354,864 new lip and oral cavity cancer cases have been diagnosed, with South and South‐Central Asia reporting the highest incidence rates. 2 Locally advanced OSCC is treated using a multimodal approach, with standard surgery followed by adjuvant chemoradiotherapy or radiotherapy, depending on the various risk factors. 3 , 4 Poor disease prognosis, along with logistical issues such as a delay in access to tertiary care, often exacerbates disease progression following diagnosis and early recurrence in OSCC. 5 Therefore, it is essential to understand the biological pathways involved in disease progression, metastasis and resistance to treatment. This will help in developing regimens that prevent metastasis and early recurrence while improving survival rates, and also help identify certain disease‐specific molecular signatures that enable us to predict treatment response.
Vascular endothelial growth factor (VEGF), a well‐known angiogenic driver molecule in both normal and pathological conditions, is widely recognised. 6 VEGF signalling has a profound effect on the pathophysiology of OSCC as a crucial regulator of angiogenesis and a factor that helps differentiate new blood vessels from pre‐existing ones. Tumour cells secrete VEGF protein, which promotes angiogenesis and results in a continuous supply of oxygen and nutrients for tumour growth and proliferation. VEGF has several isoforms, including VEGF‐A, VEGF‐B, VEGF‐C and VEGF‐D, of which the most extensively investigated is VEGF‐A, which is largely involved in angiogenesis and vascular permeability. 7 , 8 Various studies have reported that VEGFA and VEGFB expression is higher in malignant tissues than in normal oral mucosa, and they play a significant role in enhancing angiogenesis and tumour growth in OSCC. 9 VEGFB to a limited extent has demonstrated the ability to stimulate the development of new blood vessels. 10 , 11 , 12 VEGF binds to specific endothelial cell receptors, VEGFR‐1 (Flt‐1) and VEGFR‐2 (KDR/Flk‐1) out of which VEGFR‐2 is thought to be the primary mediator of VEGF‐induced angiogenesis in OSCC. 13 , 14 By promoting the development of new blood vessels within the tumour, VEGF signalling increases the proliferation of cancer cells and facilitates tumour invasion into surrounding healthy tissues. 15 VEGF can also stimulate lymphangiogenesis or the development of new lymphatic vessels in addition to stimulating angiogenesis. Lymphatic vessels aid in tumour metastasis by allowing cancer cells to migrate to localised lymph nodes and distant areas. 13
Cyclooxygenase‐2 (COX‐2), also known as prostaglandin‐endoperoxide synthase 2 (PTGS2), has been investigated as a pro‐angiogenic because the prostaglandins generated by COX‐2 can increase the expression of angiogenic factors such as VEGF. 15 In OSCC, COX‐2 overexpression can modulate angiogenesis, which stabilises the tumour microenvironment and promotes tumour progression. 16 Poor prognosis in OSCC, including large tumours, lymph node metastases and low overall survival rates, has been linked to high COX‐2 expression. 17 Thus, it has the potential to serve as a biomarker for determining the disease progression in patients. It also affects the tumour microenvironment by increasing vascular permeability and promoting cancer cell migration and invasion. In particular, chemokine receptors with the C‐X‐C motif (CXCR), particularly CXCR1 and CXCR2, which were previously shown to be involved in inflammation and tumourigenesis, are crucial mediators of the VEGF signalling cascade. 18 Immune, endothelial and cancer cells are a few examples of many cell types that contain these receptors. 19 Owing to their role in promoting angiogenesis, CXCR1 and CXCR2 have been correlated with the development, invasion and metastasis of OSCC tumours. 20 This promotes the establishment of new blood vessels, which are essential for the development and spread of tumours.
PTGS2 and VEGF signalling pathway genes, including VEGFA, VEGFB, KDR, CXCR1 and CXCR2, were found to be highly expressed in tumour samples compared to their paired normal tissues in the Cancer Genome Atlas for Head and Neck Squamous Cell Carcinoma (TCGA‐HNSCC) datasets, with CXCR2 being the only exception, with higher expression in normal tissues than in tumour tissues. 21 Hence, in the present study, the differential expression in tumour tissues from patients with locally advanced OSCC was examined, and the expression pattern was compared with TCGA‐HNSCC datasets computationally. It is important to note that our understanding of VEGF signalling in OSCC is still evolving, and our work aims to provide more insight into how the VEGF signalling pathway is correlated with the expression of PTGS2. In this study, a significant increase in PTGS2 with VEGFA, KDR, CXCR1 and CXCR2 expression was observed in tumour samples compared to their paired normal samples, with the exception of VEGFB, whose expression was not statistically significant. Further co‐expression analysis revealed a strong positive correlation between PTGS2 and VEGF signalling genes. The clinical characteristics of the cohort further assisted in a more accurate assessment of gene expression based on various risk factors.
2. MATERIALS AND METHODS
2.1. Patient enrolment and sample collection
Patients diagnosed with histopathologically proven locally advanced OSCC (pT3 and above, pN1 and above) who underwent treatment at Kasturba Hospital, Manipal from November 2021 to October 2022 with adequate renal, liver, cardiac and thyroid function, falling between the age groups of 18–80 years were included in the study. Patients with early OSCC (cT1, pT2 and cN0), patients with recurrent/second primary oral malignancy, pregnant women and patients unwilling to participate in the study were excluded. Tumour and adjacent normal tissues were collected during surgery. The tissue samples were snap‐frozen and stored at −80°C.
The demographic profile, histopathological characteristics (stage of the disease, tumour grade, nodal metastasis status, extranodal extension (ENE) status, lymphovascular invasion status, perineural invasion status etc) and data pertaining to treatment received by the patients were collected.
2.2. RNA isolation and cDNA synthesis
Total RNA was extracted from 50 to 100 mg of tissue using TRI Reagent (Sigma Aldrich), according to the manufacturer's protocol. The RNA purity was then assessed calorimetrically for the final preparation, considering an A260 to A280 ratio of ≥1.7, to avoid contamination. cDNA was then synthesised using the PrimeScript RT Reagent Kit (TaKaRa) and the reverse transcriptase reactions contained 500 ng of RNA samples, 2 μL of 5X PrimeScript® Buffer (for real time), 0.5 μL of PrimeScript® RT Enzyme Mix I, 4.5 μL of RNase‐free water and 1 μL of primers (0.5 μL of oligo dT primer and 0.5 μL of random 6 mers). The 10 μL reactions were incubated for 15 min at 37°C and for 5 s at 85°C and maintained at 4°C. The synthesised cDNA product was stored at −20°C for later use.
2.3. Differential gene expression analysis in OSCC
Real‐time polymerase chain reaction (RT‐PCR) was carried out differentially (tumour compared to paired normal) with SYBR® Premix Ex Taq II (TaKaRa) and target gene‐specific primers (VEGFA, VEGFB, KDR, CXCR1, CXCR2 and PTGS2) (Table 1) using QuantStudio 5 Real‐Time PCR (Thermo Fisher). Glyceraldehyde‐3‐phosphate dehydrogenase (GAPDH) was used as the internal control. Samples were run in triplicates for 40 cycles at 95°C for 30 s and 60°C for 30 s. 2^ΔΔCT method was employed to calculate the fold quantification among paired samples. All experiments were repeated three times. Negative controls with Milli‐Q water instead of template DNA were included.
TABLE 1.
Primers suitable for genes PTGS2, VEGFA, VEGFB, KDR, CXCR1, CXCR2 and GAPDH.
| S. no. | Primer name | Primer sequence 5′ → 3′ | No of bases | Product size (BP) |
|---|---|---|---|---|
| 1 | PTGS2‐F | CCTGTGCCTGATGATTGC | 18 | 165 |
| 2 | PTGS2‐R | CTGATGCGTGAAGTGCTG | 18 | |
| 3 | VEGFA‐F | TCTTCAAGCCATCCTGTGTG | 20 | 102 |
| 4 | VEGFA‐R | TCTGCATGGTGATGTTGGAC | 20 | |
| 5 | VEGFB‐F | CCTCATGATCCGGTACCCGA | 20 | 110 |
| 6 | VEGFB‐R | CCTTGGCAACGGAGGAAGCT | 20 | |
| 7 | KDR‐F | GGAACCTCACTATCCGCAGAGT | 22 | 132 |
| 8 | KDR‐R | CCAAGTTCGTCTTTTCCTGGGC | 22 | |
| 9. | CXCR1‐F | TCCTTTTCCGCCAGGCTTACCA | 22 | 127 |
| 10. | CXCR1‐R | GGCACGATGAAGCCAAAGGTGT | 22 | |
| 11. | CXCR2‐F | TCCGTCACTGATGTCTACCTGC | 22 | 140 |
| 12. | CXCR2‐R | TCCTTCAGGAGTGAGACCACCT | 22 | |
| 13. | GAPDH‐F | CGACCACTTTGTCAAGCTCA | 20 | 150 |
| 14. | GAPDH‐R | GAGGGTCTCTCTCTTCCTCT | 20 |
2.4. Computational gene expression and Kaplan–Meier analysis
Gene expression of PTGS2, VEGFA, VEGFB, KDR, CXCR1 and CXCR2, was analysed based on patient gender, nodal metastasis status and individual cancer stages in the Cancer Genome Atlas for Head and Neck Squamous Cell Carcinoma (TCGA‐HNSCC) datasets using the University of Alabama at Birmingham Cancer Data Analysis Portal (UALCAN) (http://ualcan.path.uab.edu). 22 , 23 mRNA expression was expressed as transcription per million (TPM) for all selected genes. To elucidate the clinical significance of PTGS2 and VEGF signalling pathway gene expression, Kaplan–Meier plots were generated computationally with log‐rank test p‐values using the GEPIA2 database (http://gepia2.cancer‐pku.cn). 24 Stratification of patient groups was performed based on the median TPM gene expression values extracted from TCGA‐HNSCC datasets, dividing them into high and low gene expression groups.
2.5. Co‐expression analysis
Co‐expression analysis was conducted to ascertain the correlation between the expression patterns of PTGS2 and genes involved in regulating VEGF signalling. Pearson's correlation analysis was used to assess the co‐expression relationships among the target genes.
2.6. Statistical analysis
All continuous variables are reported as median, interquartile range (IQR) and standard deviation (SD) after assessing the normality of the data. Wilcoxon–Mann–Whitney test was used to evaluate the expression between the tumour and normal tissue. One‐way ANOVA was performed to determine significant differences between the groups (expression based on patient sex, tobacco exposure, tumour stage and grade, ENE status, recurrence and nodal metastasis status). Pearson's correlation analysis was performed to evaluate co‐expression between target genes. Kaplan–Meier analysis was used to assess survival rates using time‐to‐event data. Patient groups with high and low expression levels were compared over a 5‐year period, and statistical significance was determined using log‐rank tests.
3. RESULTS
3.1. Patient characteristics and demographic details
Twenty‐four patients with locally advanced OSCC were enrolled in the study after obtaining informed consent. Among the enrolled patients, 16 were male and 8 were female. Two patients were reported to have a smoking habit, and 16 patients reported a habit of tobacco chewing. Of the 24 patients, 17 patients had buccal mucosa as the sub‐site of the tumour's origin, while 2 patients had gingivobuccal sulcus, 2 had retromolar trigone and 1 patient each with hard palate, tongue and upper alveolus as primary sub‐site. Eleven patients were reported to have pT4 tumours, followed by six patients with pT3 tumours and seven with pT2 tumours. Nodal metastasis was positive in 17 patients, of which 6 were pN1 followed by pN2 and pN3 in 6 and 5 patients, respectively, whereas nodal metastasis was negative (pN0) in 7 patients. Six patients were positive for ENE, seven patients were positive for lymphovascular invasion (LVI), and 15 patients were positive for perineural invasion (PNI) (details enclosed in Table 2). All recruited patients underwent surgery followed by adjuvant radiotherapy or chemoradiotherapy according to risk factors, such as advanced T‐stage(T3/T4), N‐stage (N2/N3), margin positive status, depth of invasion and PNI.
TABLE 2.
Patient characteristics and pathological staging of the tumour (n = 24).
| Characteristic | Value (n = 24) |
|---|---|
| Demographics | |
| Male | 16 |
| Female | 8 |
| Smoking | |
| Yes | 2 |
| No | 22 |
| Tobacco chewing | |
| Yes | 16 |
| No | 8 |
| Diet | |
| Vegetarian | 4 |
| Mixed | 20 |
| Alcohol | |
| Yes | 2 |
| No | 22 |
| Histopathology pT stage | |
| pT1 | 0 |
| pT2 | 7 |
| pT3 | 6 |
| pT4 | 11 |
| Histopathology pN stage | |
| pN0 | 7 |
| pN1 | 6 |
| pN2 | 6 |
| pN3 | 5 |
| Extranodal extension | |
| Positive | 6 |
| Negative | 18 |
| Perineural invasion | |
| Positive | 15 |
| Negative | 9 |
| Lymphovascular invasion | |
| Positive | 7 |
| Negative | 17 |
| Worst pathology pattern | |
| 2 | 3 |
| 3 | 9 |
| 4 | 7 |
| 5 | 5 |
| Tumour stage and grade (AJCC) | |
| Stage 3 | 7 |
| Stage 4 | 17 |
| Grade 1 | 12 |
| Grade 2 | 12 |
| Adjuvant treatment | |
| Chemoradiotherapy | 13 |
| Radiotherapy | 9 |
| Defaulted | 2 |
| Margin status | |
| Positive | 1 |
| Negative | 23 |
| Recurrence status | |
| Locoregional recurrence | 1 |
| Distant recurrence | 5 |
| No recurrence | 18 |
3.2. Differential gene expression in locally advanced OSCC
Differential expression analysis using RT‐PCR was performed for all 24 patients. Based on the results obtained, PTGS2, VEGFA, KDR, CXCR1 and CXCR2 were found to be significantly overexpressed in tumour samples compared to paired normal samples. Similar trend was observed with VEGFB gene expression but the results were not statistically significant (Figure 1A). Gene expression analysis based on the patient's gender revealed that CXCR1, CXCR2, KDR, VEGFB and VEGFA were overexpressed in males whereas only PTGS2 expression was observed to be expressed more in females (Figure 1B). When the cohort was stratified based on smokeless tobacco exposure, we observed that all genes, including PTGS2, were upregulated in patients with a smokeless tobacco chewing habit (Figure 2A). Patients with recurrence (distant and locoregional) had higher expression of PTGS2, CXCR1, KDR and VEGFB (Figure 2B).
FIGURE 1.

(A) The differential gene expression analysis of I. PTGS2, II. VEGFA, III. VEGFB, IV. KDR, V. CXCR1 and VI. CXCR2 genes in OSCC tumour samples as compared to normal. (*p < 0.05 and ns p >0.05) (n = 24). (B) The differential gene expression analysis of PTGS2, VEGFA, VEGFB, KDR, CXCR1 and CXCR2 genes in OSCC tumour samples (n = 24) based on gender stratification, that is, male (n = 16) and female (n = 8), where ***p = 0.001 and **p = 0.01.
FIGURE 2.

(A) The gene expression analysis of PTGS2, VEGFA, VEGFB, KDR, CXCR1 and CXCR2 genes in OSCC tumour samples (n = 24) based on smokeless tobacco consumption status, that is, tobacco consumers (n = 16) and non‐consumers (n = 8), where ***p = 0.001, **p = 0.01 and *p < 0.05. (B) The gene expression analysis of PTGS2, VEGFA, VEGFB, KDR, CXCR1 and CXCR2 genes in OSCC tumour samples (n = 24) based on recurrence status, that is, recurrence positive (n = 6) and recurrence negative (n = 18), where ***p = 0.001, **p = 0.01 and *p < 0.05.
Overexpression of all genes was found in stage 3 cancer patients compared to that in normal tissues, except CXCR1 which was upregulated in patients with stage 4 disease (Figure 3A). All five genes were overexpressed in grade 2 tumours compared with those in grade 1 tumours (Figure 3B). When expression was evaluated based on the nodal metastasis status of the patients, all genes were observed to be more highly expressed in samples with positive nodal metastasis status compared to the samples with negative nodes. Furthermore, PTGS2, CXCR1, CXCR2 and VEGFA were observed to be more highly expressed in patients with pN2 status, with significantly higher median expression compared to samples with pN0 status. VEGFB and KDR were observed to have a significantly higher median expression when compared to samples with pN0 in tumour samples with pN3 status (Figure 4A). When further investigating the expression based on ENE status, all genes were significantly upregulated in tumour samples with an ENE‐positive status (Figure 4B).
FIGURE 3.

(A) The gene expression analysis of PTGS2, VEGFA, VEGFB, KDR, CXCR1 and CXCR2 genes in OSCC tumour samples (n = 24) based on cancer stage (AJCC), that is, stage 3 (n = 7) and stage 4 (n = 17), where ***p = 0.001. (B) The gene expression analysis of PTGS2, VEGFA, VEGFB, KDR, CXCR1 and CXCR2 genes in OSCC tumour samples (n = 24) based on cancer grade (AJCC), that is, grade 1 (n = 12) and grade 2 (n = 12), where ***p = 0.001, **p = 0.01 and *p < 0.05.
FIGURE 4.

(A) The gene expression analysis of PTGS2, VEGFA, VEGFB, KDR, CXCR1 and CXCR2 genes in OSCC tumour samples (n = 24) based on nodal metastasis status, that is, pN0 (n = 7), pN1 (n = 6), pN2 (n = 6) and pN3 (n = 5), where ***p = 0.001, **p = 0.01 and *p < 0.05. (B) The gene expression analysis of PTGS2, VEGFA, VEGFB, KDR, CXCR1 and CXCR2 genes in OSCC tumour samples (n = 24) based on extranodal extension (ENE) status, that is, ENE‐positive (n = 6) and ENE negative (n = 18), where ***p = 0.001, **p = 0.01 and *p < 0.05.
3.3. Computational gene expression and Kaplan–Meier analysis in TCGA‐HNSCC data
Computational gene expression analysis based on patient gender revealed that PTGS2, KDR, CXCR1 and CXCR2 genes had significantly higher expression in female patients, whereas VEGFA and VEGFB genes had significantly higher expression in male patients (Figure 5). Expression based on individual cancer stages showed that VEGFA and VEGFB were highly expressed in patients with stage 4 cancer. The expression of PTGS2 and KDR was significantly higher in stage 3 and stage 4 cancer, respectively. CXCR2 genes were more highly expressed in normal samples compared to tumour samples (Figure 6A). Expression based on nodal metastasis status showed that VEGFA and KDR were significantly upregulated in samples with N2 metastasis. CXCR1 and PTGS2 genes had significantly higher expression in patients with N1 metastasis, whereas VEGFB expression was significantly higher in tumour samples with N3 metastasis. CXCR2 expression was higher in the normal samples (Figure 6B). The computational Kaplan–Meier analysis did not achieve statistical significance. However, the analysis revealed that individuals with low expression levels of PTGS2, VEGFA, VEGFB and CXCR1 demonstrated better 5‐year survival rates than those with high expression levels. Conversely, patients expressing elevated levels of KDR and CXCR2 displayed better 5‐year survival rates compared to those expressing lower levels (see Figure S1).
FIGURE 5.

The gene expression analysis of genes, that is, I. PTGS2, II. VEGFB, III. VEGFA, IV. KDR, V. CXCR1 and VI. CXCR2 in head and neck squamous cell carcinoma based on patient gender as compared to normal samples using the UALCAN web server (***p = 0.001).
FIGURE 6.

(A) The gene expression analysis of genes, that is, I. PTGS2, II. VEGFB, III. VEGFA, IV. KDR, V. CXCR1 and VI. CXCR2 in head and neck squamous cell carcinoma based on individual cancer stage using the UALCAN web server (***p = 0.001). (B) The gene expression analysis of genes, that is, I. PTGS2, II. VEGFB, III. VEGFA, IV. KDR, V. CXCR1 and VI. CXCR2 in head and neck squamous cell carcinoma is based on nodal metastasis status using the UALCAN web server (***p = 0.001).
3.4. Co‐expression analysis
Co‐expression analysis was performed using Pearson's correlation to assess the significance of the linear association between the expression of the two genes. Co‐expression analysis was performed between PTGS2 with VEGFA, VEGFB, KDR, CXCR1 and CXCR2. A statistically significant positive correlation was observed between the expression of PTGS2 with VEGFA, KDR and CXCR1 genes. In contrast, CXCR2 and VEGFB showed positive and negative correlations, respectively, but these were not statistically significant (Figure 7).
FIGURE 7.
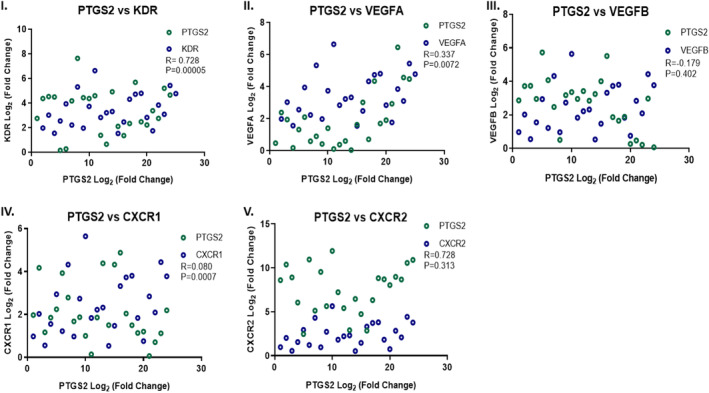
Co‐expression analysis in OSCC samples (n = 24) between I. KDR versus PTGS2, II. VEGFA versus PTGS2, III. VEGFB versus PTGS2, IV. CXCR1 versus PTGS2 and V. CXCR2 versus PTGS2 (R = Pearson correlation coefficient).
4. DISCUSSION
Like many other cancers, oral malignancies are significantly influenced by the VEGF signalling pathway. Angiogenesis is necessary for oral malignancies to establish a blood supply like other solid tumours. In a previous in silico study on TCGA‐HNSCC datasets (PanCancer Atlas), encompassing 523 samples, 328 displayed altered gene expression. 21 Tumour tissues exhibited higher median expression levels, except for CXCR2, which showed a higher median expression in normal tissues. Analysis based on tumour grade using the UALCAN portal revealed specific expression patterns such as increased VEGFA expression in grade 2 and grade 3 tumours, elevated VEGFB expression in grade 3 tumours and high KDR expression in grade 4 tumours. This study further identified unique gene expression patterns associated with HPV and TP53 mutation status. HPV‐negative patients displayed significant upregulation of PTGS2, KDR, CXCR1 and CXCR2, whereas VEGFA and VEGFB were more highly expressed in HPV‐positive patients. TP53‐mutated tumours exhibit elevated expression of VEGFA, VEGFB, CXCR1 and PTGS2. Co‐expression analysis using cBioPortal indicated a positive correlation between PTGS2 and VEGF pathway genes and a negative correlation with VEGFB. Gene set enrichment analysis revealed associations between various biological pathways linked to angiogenesis. 21 However, this study has several limitations, as the identified expression patterns were confined to the populations represented in the databases, thus challenging the precision of extrapolation to broader ethnic groups. Notably, the absence of experimental validation, particularly gene expression studies on patient samples, undermines the confirmation of the results. Moreover, the study's clinical relevance was compromised by limited correlations with clinical data, underscoring the importance of a more thorough analysis in this regard. These constraints contributed significantly to further investigating the role played by PTGS2 and VEGF signalling in OSCC, further offering insights into their clinical relevance, particularly in the context of locally advanced and aggressive disease.
In our study, we found that PTGS2 and VEGF signalling pathway genes (VEGFA, VEGFB, KDR, CXCR1 and CXCR2) were upregulated in OSCC tumour tissues compared to their paired normal tissues. While further investigating the expression by stratifying the patient data based on sex, pathological staging, tumour grade, ENE status and nodal metastasis status, an intriguing expression pattern was observed in all genes. All the targeted genes were more highly expressed in male patients, except for PTGS2, which was more highly expressed in female patients, which was also observed in TCGA‐HNSCC datasets. This can be associated with oestrogen and progesterone, which influence PTGS2 expression, as documented in a few studies. 25 , 26 In the HNSCC‐TCGA data, the KDR and CXCR1 genes were highly expressed in female patients. However, the available literature suggests that this might also be the case because of the female sex hormones, but it is not well established. PTGS2 expression is upregulated in tumours following smokeless tobacco exposure, and there is ample evidence that increased PTGS2 expression in oral tissues is associated with tobacco chewing habits. 27 The inflammatory reaction triggered by exposure to harmful compounds in smokeless tobacco may be an underlying cause of the elevated expression of PTGS2. 28 VEGFA, VEGFB and CXCR1 expression were higher in patients with smokeless tobacco exposure, which might be in response to cellular damage caused by exposure to smokeless tobacco. 29 CXCR1, which is associated with the inflammatory response brought on by exposure to carcinogens and toxins in smokeless tobacco, contributes to the greater expression of CXCR1. 30 Ongoing inflammation increases the risk of cancer, whereas smokeless tobacco can cause cellular damage and localised inflammation in the oral mucosa. Increased CXCR1 expression might be a factor in this inflammatory reaction which in the context of cancer can affect tumour growth, invasion and progression. 31
In a stage‐wise analysis based on the American Joint Committee on Cancer (AJCC) classification (stage 3 and stage 4), all genes were upregulated in stage 3 cancers, except CXCR1, which showed higher expression in stage 4 tumours. Increased CXCR1 expression in stage 4 entails disease aggressiveness, as CXCR1 plays a crucial role in tumour cell migration and invasion. 32 VEGFA and VEGFB genes were more highly expressed in stage 4 disease in HNSCC‐TCGA data, likely due to the highly vascularised nature of the head and neck (HN) region. With respect to the nodal metastasis status, all genes were significantly expressed in patients with positive nodal metastasis. In oral squamous cell carcinoma (OSCC), PTGS2‐regulated VEGF signalling promotes the formation of new blood and lymphatic vessels, facilitating the spread of cancer to the regional lymph nodes. Elevated PTGS2 and VEGF signalling pathway expression in OSCC correlates with a higher likelihood of lymph node metastasis. 33 VEGFB induces lymphangiogenesis, and KDR aids in maintaining lymphatic vessel structure and function. 34 , 35 Higher expression of all genes was observed in tumours with ENE‐positive status and grade 2 tumours, indicating a more aggressive disease. 36 , 37 ENE and grade 2 tumours are poor prognostic indicators associated with treatment resistance and an increased likelihood of spread or recurrence, emphasising the potential for targeted treatment strategies involving PTGS2 and VEGF signalling pathway genes for optimal oncological outcomes. 38
It is well established that PTGS2 expression plays a significant role in inducing the VEGF signalling pathway, as observed in co‐expression analysis results. 39 The increased expression of PTGS2, KDR, VEGFB and CXCR1 in samples from patients with positive recurrence can be explained by their association with treatment resistance. Inflammation caused by PTGS2 promotes the survival of cancer cells and their resistance to therapy because inflammatory cells can create inflammatory mediators that support cell survival and confer resistance to therapies that cause cell death. 40 Overexpression of PTGS2 can prevent apoptosis (programmed cell death), contributing to treatment resistance. 41 Inflammatory cytokines that activate CXCR1 can activate downstream signalling pathways that improve cancer cell survival and increase resistance to certain treatments. 42 CXCR1‐associated cancer cells express invasive attributes and metastasise faster, making the cells resistant to treatment. VEGFB and KDR are often associated with angiogenesis‐mediated resistance because they stabilise the tumour microenvironment by ensuring an adequate supply of oxygen and other nutrients required for tumour growth, further adjusting the tumour microenvironment for optimum disease progression. 43 Drugs, such as celecoxib and methotrexate (a combination used in low‐dose oral metronomic chemotherapy) and bevacizumab, a monoclonal antibody against VEGF, have demonstrated encouraging results in clinical trials to reduce angiogenesis and improve oncological outcomes in specific malignancies. 44 , 45
Our findings on the intriguing expression pattern and clinical implications of PTGS2 and VEGF signalling genes provide a foundation for further investigation with comprehensive multi‐omics approach to gain deeper insights into the molecular complexity of the disease. However, the study's limitations, including the small sample size, requires validation with a larger patient cohort. Considering the potential differences in gene expression among cells, tissues and various oral cavity sub‐sites, a more comprehensive study is needed to accurately capture the overall gene expression pattern. Although useful, in silico predictions involve complex data processing. The unique demographic focus of this study may limit its generalisability to other ethnic groups, underscoring the need for a more extensive study with a larger and diverse patient cohort and employing a disciplinary experimental methodology for validation. As the genes analysed in this study are limited, transcriptomic and proteomic investigations (like immunohistochemistry) of genes correlated with the PTGS2 and VEGF signalling pathway will further enhance our understanding of expression patterns at the translational level, providing insights into cancer biology.
5. CONCLUSION
This study highlights the significance of PTGS2 and VEGF signalling genes in locally advanced OSCC by identifying their significant overexpression in tumour tissues. Overexpression of PTGS2, VEGFA, VEGFB, KDR, CXCR1 and CXCR2 was observed, with a notable association with various risk factors for OSCC. Despite the small sample size, this study emphasises the clinical relevance of these genes, suggesting potential connections to disease aggressiveness and adverse prognostic factors. Acknowledging these limitations, the findings advocate for further exploration using a multi‐omics approach to unravel molecular intricacies of OSCC. This study establishes a foundational understanding, paving the way for future research with larger and more diverse cohorts, advanced methodologies and translational insights into disease biology.
AUTHOR CONTRIBUTIONS
Mehta Vedant Kamal: Conceptualization (equal); formal analysis (lead); investigation (lead); methodology (lead); validation (equal); visualization (equal); writing – original draft (lead). Rama Rao Damerla: Conceptualization (equal); data curation (equal); funding acquisition (supporting); investigation (supporting); methodology (supporting); project administration (supporting); resources (lead); software (lead); supervision (lead); writing – review and editing (supporting). Preetiparna Parida: Data curation (equal); formal analysis (supporting); investigation (equal); methodology (supporting); resources (supporting); software (supporting); validation (equal); visualization (equal); writing – original draft (supporting). Mahadev Rao: Conceptualization (supporting); funding acquisition (supporting); project administration (equal); resources (equal); supervision (supporting); validation (supporting); writing – review and editing (equal). Vijetha Shenoy Belle: Formal analysis (supporting); investigation (supporting); methodology (supporting); resources (equal); software (supporting); validation (equal); visualization (supporting); writing – review and editing (supporting). Punit Singh Dikhit: Data curation (equal); resources (lead); software (equal). Akhil Palod: Data curation (equal); resources (equal); software (equal); writing – review and editing (equal). Rinsha Gireesh: Data curation (equal); resources (equal); software (equal); writing – review and editing (supporting). Naveena AN Kumar: Conceptualization (equal); funding acquisition (lead); resources (equal); supervision (lead); writing – original draft (equal); writing – review and editing (equal).
FUNDING INFORMATION
This work was supported by the Manipal Academy of Higher Education under faculty seed money funding [MAHE FSM No. 00000676].
CONFLICT OF INTEREST STATEMENT
The authors declare no conflict of interest.
ETHICS STATEMENT
The above project was reviewed by the institutional ethics committee of Kasturba Medical College and Kasturba Hospital, and approval was obtained (IEC 595/2020). Informed consent was obtained from all patients whose samples were utilised in the study.
Supporting information
Figure S1.
ACKNOWLEDGMENTS
All the authors would like to acknowledge the support from the Global Cancer Consortium (https://glocacon.org/).
Kamal MV, Damerla RR, Parida P, et al. Expression of PTGS2 along with genes regulating VEGF signalling pathway and association with high‐risk factors in locally advanced oral squamous cell carcinoma. Cancer Med. 2024;13:e6986. doi: 10.1002/cam4.6986
DATA AVAILABILITY STATEMENT
The computational data used in this article are freely available in The Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov), UALCAN (http://ualcan.path.uab.edu) and GEPIA2 (http://gepia2.cancer‐pku.cn) database. The patient details and RT‐PCR expression raw data will be provided by the corresponding author upon request.
REFERENCES
- 1. Mummudi N, Agarwal JP, Chatterjee S, Mallick I, Ghosh‐Laskar S. Oral cavity cancer in the Indian subcontinent – challenges and opportunities. Clin Oncol. 2019;31(8):520‐528. [DOI] [PubMed] [Google Scholar]
- 2. Dixit S, Kumar A, Srinivasan K. Current review of machine learning and deep learning models in oral cancer diagnosis: recent technologies, open challenges, and future research directions. Diagnostics. 2023;13:1353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Anil Kumar S, Indu S, Gautami D, Varruchi S. Oral squamous cell carcinoma (OSCC) in humans: etiological factors, diagnostic and therapeutic relevance. Res J Biotechnol. 2020;15:141‐151. https://www.researchgate.net/publication/344374174 [Google Scholar]
- 4. Patra S, Patil S, Das S, Bhutia SK. Epigenetic dysregulation in autophagy signaling as a driver of viral manifested oral carcinogenesis. Biochim Biophys Acta Mol basis Dis. 2022;1868(11):166517. [DOI] [PubMed] [Google Scholar]
- 5. Al‐Ostoot FH, Salah S, Khamees HA, Khanum SA. Tumor angiogenesis: current challenges and therapeutic opportunities. Cancer Treat Res Commun. 2021;28:100422. [DOI] [PubMed] [Google Scholar]
- 6. Meng W, Hao Y, He C, Li L, Zhu G. Exosome‐orchestrated hypoxic tumor microenvironment. Mol Cancer. 2019;18:57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Hanrahan V, Currie MJ, Gunningham SP, et al. The angiogenic switch for vascular endothelial growth factor (VEGF)‐A, VEGF‐B, VEGF‐C, and VEGF‐D in the adenoma‐carcinoma sequence during colorectal cancer progression. J Pathol. 2003;200(2):183‐194. [DOI] [PubMed] [Google Scholar]
- 8. Sia D, Alsinet C, Newell P, Villanueva A. VEGF signaling in cancer treatment. Curr Pharm Des. 2014;20:2834‐2842. [DOI] [PubMed] [Google Scholar]
- 9. Costache MI, Ioana M, Iordache S, Ene D, lexandra CCA, Săftoiu A. VEGF expression in pancreatic cancer and other malignancies: a review of the literature. Rom J Intern Med. 2015;53:199‐208. [DOI] [PubMed] [Google Scholar]
- 10. Guo S, Colbert LS, Mcglothen TZ, Gonzalez‐Perez RR. 2 Regulation of Angiogenesis in Human Cancer Via Vascular Endothelial Growth Factor Receptor‐2 (VEGFR‐2). Accessed August 29, 2023. www.intechopen.com
- 11. Rmali KA, Puntis MCA, Jiang WG. Tumour‐associated angiogenesis in human colorectal cancer. Color Dis. 2007;9:3‐14. [DOI] [PubMed] [Google Scholar]
- 12. Anderson NM, Simon MC. The tumor microenvironment. Curr Biol. 2020;30:R921‐R925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Jones D. Parallels of resistance between angiogenesis and lymphangiogenesis inhibition in cancer therapy. Cells. 2020;9:762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Rathinavelu A, Alhazzani K, Dhandayuthapani S, Kanagasabai T. Anti‐cancer effects of F16: a novel vascular endothelial growth factor receptor–specific inhibitor. Tumour Biol. 2017;39(11):101042831772684. [DOI] [PubMed] [Google Scholar]
- 15. Santoro A, Bufo P, Russo G, et al. Expression and clinical implication of cyclooxygenase‐2 and E‐cadherin in oral squamous cell carcinomas. Cancer Biol Ther. 2020;21(8):667‐674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Frejborg E, Salo T, Salem A. Role of cyclooxygenase‐2 in head and neck tumorigenesis. Int J Mol Sci. 2020;21:1‐17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Saxena S, Singh RK. Chemokines orchestrate tumor cells and the microenvironment to achieve metastatic heterogeneity. Cancer Metastasis Rev. 2023;40:447‐476. doi: 10.1007/s10555-021-09970-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Liu Q, Li A, Tian Y, et al. The CXCL8‐CXCR1/2 pathways in cancer. Cytokine Growth Factor Rev. 2016;31:61‐71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Ahmadi M, Rezaie J. Tumor cells derived‐exosomes as angiogenenic agents: possible therapeutic implications. J Transl Med. 2020;18:249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Herrera‐Vargas AK, García‐Rodríguez E, Olea‐Flores M, Mendoza‐Catalán MA, Flores‐Alfaro E, Navarro‐Tito N. Pro‐angiogenic activity and vasculogenic mimicry in the tumor microenvironment by leptin in cancer. Cytokine Growth Factor Rev. 2021;62:23‐41. [DOI] [PubMed] [Google Scholar]
- 21. Kamal MV, Damerla RR, Dikhit PS, Kumar NA. Prostaglandin‐endoperoxide synthase 2 (PTGS2) gene expression and its association with genes regulating the VEGF signaling pathway in head and neck squamous cell carcinoma. J Oral Biol Craniofac Res. 2023;13(5):567‐574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Chandrashekar DS, Bashel B, Balasubramanya SAH, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia (United States). 2017;19(8):649‐658. doi: 10.1016/j.neo.2017.05.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Chandrashekar DS, Karthikeyan SK, Korla PK, et al. UALCAN: an update to the integrated cancer data analysis platform. Neoplasia. 2022;25:18‐27. doi: 10.1016/j.neo.2022.01.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Tang Z, Kang B, Li C, Chen T, Zhang Z. GEPIA2: an enhanced web server for large‐scale expression profiling and interactive analysis. Nucleic Acids Res. 2019;47(W1):W556‐W560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Jitprasertwong P, Charadram N, Kumphune S, Pongcharoen S, Sirisinha S. Female sex hormones modulate Porphyromonas gingivalis lipopolysaccharide‐induced Toll‐like receptor signaling in primary human monocytes. J Periodontal Res. 2016;51(3):395‐406. [DOI] [PubMed] [Google Scholar]
- 26. Santulli P, Borghese B, Noël JC, et al. Hormonal therapy deregulates prostaglandin‐endoperoxidase synthase 2 (PTGS2) expression in endometriotic tissues. J Clin Endocrinol Metab. 2014;99(3):881‐890. [DOI] [PubMed] [Google Scholar]
- 27. Huang R‐Y, Chen GG. Cigarette smoking, cyclooxygenase‐2 pathway and cancer. Biochim Biophys Acta Rev Cancer. 2011;1815(2):158‐169. doi: 10.1016/j.bbcan.2010.11.005 [DOI] [PubMed] [Google Scholar]
- 28. Sawhney M, Rohatgi N, Kaur J, et al. Expression of NF‐κB parallels COX‐2 expression in oral precancer and cancer: association with smokeless tobacco. Int J Cancer. 2007;120(12):2545‐2556. [DOI] [PubMed] [Google Scholar]
- 29. Grover V, Kapoor A, Sehgal K, Kaur G. Chronic inflammation and carcinogenesis – emerging role of chronic inflammatory periodontal disease. Cancer Res Front. 2016;2(2):200‐225. http://cancer‐research‐frontiers.org/2016‐2‐200/ [Google Scholar]
- 30. Oudega M. Molecular and cellular mechanisms underlying the role of blood vessels in spinal cord injury and repair. Cell Tissue Res. 2012;349:269‐288. [DOI] [PubMed] [Google Scholar]
- 31. Lee YS, Choi I, Ning Y, et al. Interleukin‐8 and its receptor CXCR2 in the tumour microenvironment promote colon cancer growth, progression and metastasis. Br J Cancer. 2012;106(11):1833‐1841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Jaffer T, Ma D. The emerging role of chemokine receptor CXCR2 in cancer progression. Transl Cancer Res. 2016;5:S616‐S628. [Google Scholar]
- 33. Shibuya M, Claesson‐Welsh L. Signal transduction by VEGF receptors in regulation of angiogenesis and lymphangiogenesis. Exp Cell Res. 2006;312:549‐560. [DOI] [PubMed] [Google Scholar]
- 34. Cueni LN, Hegyi I, Shin JW, et al. Tumor lymphangiogenesis and metastasis to lymph nodes induced by cancer cell expression of podoplanin. Am J Pathol. 2010;177(2):1004‐1016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Behl T, Kotwani A. Exploring the various aspects of the pathological role of vascular endothelial growth factor (VEGF) in diabetic retinopathy. Pharmacol Res. 2015;99:137‐148. [DOI] [PubMed] [Google Scholar]
- 36. Huang SH, Chernock R, O'Sullivan B, Fakhry C. Assessment criteria and clinical implications of extranodal extension in head and neck cancer. Am Soc Clin Oncol Educ Book. 2021;41:265‐278. [DOI] [PubMed] [Google Scholar]
- 37. Wong HM. Oral complications and management strategies for patients undergoing cancer therapy. Sci World J. 2014;2014:1‐14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Yaacoub K, Pedeux R, Tarte K, Guillaudeux T. Role of the tumor microenvironment in regulating apoptosis and cancer progression. Cancer Lett. 2016;378:150‐159. [DOI] [PubMed] [Google Scholar]
- 39. Gallo O, Masini E, Bianchi B, Bruschini L, Paglierani M, Franchi A. Prognostic significance of cyclooxygenase‐2 pathway and angiogenesis in head and neck squamous cell carcinoma. Hum Pathol. 2002;33(7):708‐714. [DOI] [PubMed] [Google Scholar]
- 40. Campagna R, Belloni A, Pozzi V, et al. Role played by paraoxonase‐2 enzyme in cell viability, proliferation and sensitivity to chemotherapy of oral squamous cell carcinoma cell lines. Int J Mol Sci. 2023;24(1):338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Bhat AA, Nisar S, Maacha S, et al. Cytokine‐chemokine network driven metastasis in esophageal cancer; promising avenue for targeted therapy. Mol Cancer. 2021;20:2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Gacche RN, Meshram RJ. Targeting tumor micro‐environment for design and development ofnovel anti‐angiogenic agents arresting tumor growth. Prog Biophys Mol Biol. 2013;113:333‐354. [DOI] [PubMed] [Google Scholar]
- 43. Larionova I, Kazakova E, Gerashchenko T, Kzhyshkowska J. New angiogenic regulators produced by tams: perspective for targeting tumor angiogenesis. Cancers (Basel). 2021;13(13):3253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Kamal MV, Rao M, Damerla RR, et al. A mechanistic review of methotrexate and celecoxib as a potential metronomic chemotherapy for oral squamous cell carcinoma. Cancer Investig. 2023;41(2):144‐154. doi: 10.1080/07357907.2022.2139840 [DOI] [PubMed] [Google Scholar]
- 45. Kumar NA, Dikhit PS, Jose A, et al. Oral metronomic chemotherapy in advanced and metastatic oral squamous cell carcinoma: a need of the hour. J Maxillofac Oral Surg. 2023:1‐8. doi: 10.1007/s12663-023-01963-y [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1.
Data Availability Statement
The computational data used in this article are freely available in The Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov), UALCAN (http://ualcan.path.uab.edu) and GEPIA2 (http://gepia2.cancer‐pku.cn) database. The patient details and RT‐PCR expression raw data will be provided by the corresponding author upon request.


