Abstract
Background and Aims
RNA-based, antibody-based, and genome editing-based therapies are currently under investigation to determine if the inhibition of angiopoietin-like protein-3 (ANGPTL3) could reduce lipoprotein-lipid levels and atherosclerotic cardiovascular disease (ASCVD) risk. Mendelian randomisation (MR) was used to determine whether genetic variations influencing ANGPTL3 liver gene expression, blood levels, and protein structure could causally influence triglyceride and apolipoprotein B (apoB) levels as well as coronary artery disease (CAD), ischaemic stroke (IS), and other cardiometabolic diseases.
Methods
RNA sequencing of 246 explanted liver samples and genome-wide genotyping was performed to identify single-nucleotide polymorphisms (SNPs) associated with liver expression of ANGPTL3. Genome-wide summary statistics of plasma protein levels of ANGPTL3 from the deCODE study (n = 35 359) were used. A total of 647 carriers of ANGPTL3 protein-truncating variants (PTVs) associated with lower plasma triglyceride levels were identified in the UK Biobank. Two-sample MR using SNPs that influence ANGPTL3 liver expression or ANGPTL3 plasma protein levels as exposure and cardiometabolic diseases as outcomes was performed (CAD, IS, heart failure, non-alcoholic fatty liver disease, acute pancreatitis, and type 2 diabetes). The impact of rare PTVs influencing plasma triglyceride levels on apoB levels and CAD was also investigated in the UK Biobank.
Results
In two-sample MR studies, common genetic variants influencing ANGPTL3 hepatic or blood expression levels of ANGPTL3 had a very strong effect on plasma triglyceride levels, a more modest effect on low-density lipoprotein cholesterol, a weaker effect on apoB levels, and no effect on CAD or other cardiometabolic diseases. In the UK Biobank, the carriers of rare ANGPTL3 PTVs providing lifelong reductions in median plasma triglyceride levels [−0.37 (interquartile range 0.41) mmol/L] had slightly lower apoB levels (−0.06 ± 0.32 g/L) and similar CAD event rates compared with non-carriers (10.2% vs. 10.9% in carriers vs. non-carriers, P = .60).
Conclusions
PTVs influencing ANGPTL3 protein structure as well as common genetic variants influencing ANGPTL3 hepatic expression and/or blood protein levels exhibit a strong effect on circulating plasma triglyceride levels, a weak effect on circulating apoB levels, and no effect on ASCVD. Near-complete inhibition of ANGPTL3 function in patients with very elevated apoB levels may be required to reduce ASCVD risk.
Keywords: Angiopoietin-like protein-3, Atherosclerotic cardiovascular diseases, Cardiometabolic diseases, Lipoprotein-lipid levels, RNA-based therapies, Antibody-based therapies, Gene-editing-based therapies, Mendelian randomisation, Protein-truncating variants
Structured Graphical Abstract
Structured Graphical Abstract.
ANGPTL3, angiopoietin-like protein-3; LDL, low-density lipoprotein; RNA, ribonucleic acid; SNP, single-nucleotide polymorphism.
See the editorial comment for this article ‘Inhibiting angiopoietin-like protein 3: clear skies or clouds on the horizon?’, by R.A. Hegele, https://doi.org/10.1093/eurheartj/ehae068.
Translational perspective.
Mendelian randomisation offers the possibility of investigating the potential of preventive therapies such as angiopoietin-like protein-3 (ANGPTL3) inhibitors in ‘natural’ randomised clinical trials. The results of this study showing a lack of effect of variants at the ANGPTL3 locus on cardiometabolic diseases suggest that the long-term cardiometabolic benefits of this therapeutic strategy in patients at high residual risk of atherosclerotic cardiovascular disease (ASCVD) with elevated triglyceride levels may be limited. Near-complete inhibition of ANGPTL3 function in patients with very elevated apolipoprotein B levels may be required to reduce ASCVD risk with this strategy.
Introduction
Elevated plasma lipoprotein and lipid levels including low-density lipoprotein (LDL) cholesterol and triglycerides are important risk factors for atherosclerotic cardiovascular disease (ASCVD) and some metabolic diseases.1,2 Every lipoprotein that is positively associated with long-term ASCVD risk contains one apolipoprotein B (apoB) molecule. A strong and dose-dependent effect of apoB on ASCVD risk has been reported in multiple observational studies.3 The results of several recent Mendelian randomisation (MR) studies and randomised cardiovascular outcome trials provided evidence that lipid-lowering therapies may be effective in reducing ASCVD risk if they provide significant reductions in plasma apoB levels regardless of their impact on other lipoprotein parameters such as triglycerides or high-density lipoprotein (HDL) cholesterol levels.4–6 Many triglyceride-lowering agents are currently under investigation for their potential cardiovascular benefits. While some triglyceride-lowering agents such as icosapent ethyl may provide cardiovascular benefits, other triglyceride-lowering agents such as fibrates or other formulations of omega-3 fatty acids may not provide cardiovascular benefits.7–9
The plasma levels of angiopoietin-like protein-3 (ANGPTL3) play a critical role in triglyceride-rich lipoprotein metabolism.10,11 ANGPTL3 is mostly expressed in the liver. Rare and common genetic variations at the ANGPTL3 locus are associated with lower plasma levels of LDL cholesterol and triglyceride levels.12,13 At least three ANGPTL3 inhibition strategies are currently under investigation for their impact on lipoprotein-lipid levels and potential cardiovascular benefits. The inhibition of ANGPTL3 synthesis by a liver-targeted antisense oligonucleotide has been shown to substantially reduce plasma triglyceride levels in patients with elevated triglyceride levels.14,15 The neutralisation of the circulating ANGPTL3 protein by a monoclonal antibody has also been shown to reduce triglyceride levels and LDL cholesterol levels, respectively, in patients with elevated triglyceride levels and homozygous familial hypercholesterolaemia (FH).16,17 A CRISPR-based therapy testing a one-time gene-editing strategy intended to potently and durably reduce plasma triglyceride and LDL cholesterol levels by inactivating hepatic ANGPTL3 via the introduction of a premature stop codon in the ANGPTL3 DNA sequence is also under development.18,19 Whether targeting ANGPTL3 via one of these strategies could ultimately prevent ASCVD events in high-risk patients remains to be demonstrated.
By establishing the impact of genetic variants predicting human traits on health outcomes (such as cardiometabolic traits and diseases), MR offers the possibility of investigating the association between genetically predicted drug targets and cardiometabolic outcomes in ‘natural’ randomised clinical trial (as genetic variants are randomly acquired at meiosis and immune to reverse causation and confounding). One of the most replicated MR studies presented the impact of sequence variants at the PCSK9 gene linked with low LDL cholesterol levels and protection against coronary artery disease (CAD).20 Randomised clinical trials subsequently confirmed that the PCSK9 neutralisation with monoclonal antibodies prevents atherosclerosis progression and protects against short-term ASCVD events.21–23 Previous genetic/MR studies testing the impact of rare and common variants at ANGPTL3 have provided conflicting results. While rare variants at the ANGPTL3 locus were associated with protection against CAD,24,25 more common variants linked with lifelong reductions in plasma triglyceride, LDL cholesterol, and apoB levels were not associated with CAD.26 In this study, only one independent genetic variant with an unknown function at the ANGPTL3 locus was associated with triglyceride levels. Combining genomics with other OMIC strategies such as transcriptomics and proteomics offers the opportunity to better understand how the genetic regulation of ANGPTL3 gene or protein expression might be related to cardiometabolic traits and diseases. These new developments could enable the investigation of the cardiometabolic impact of different ANGPTL3 inhibition strategies.
The first objective of this study was to identify genetic variants at the ANGPTL3 locus influencing (i) liver ANGPTL3 gene expression, (ii) circulating ANGPTL3 protein levels, and (iii) ANGPTL3 protein structure to respectively mimic ANGPTL3 RNA-based therapies, antibodies, and gene-editing strategies. We sought to determine whether the genetic variations at the ANGPTL3 locus influencing ANGPTL3 gene expression, circulating concentrations of ANGPTL3 levels, or rare ANGPTL3 sequence variants influence plasma lipoprotein-lipid levels as well as cardiometabolic diseases using a comprehensive MR approach. We also aimed at identifying and replicating genetic determinants of circulating ANGPTL3 levels. The schematic representation of the study design is presented in Figure 1.
Figure 1.
Schematic representation of the Mendelian randomisation study design. ANGPTL3 is a key regulator of blood lipid levels. ANGPTL3 inhibitors are currently under investigation for their impact on metabolic and cardiovascular health. ANGPTL3, angiopoietin-like protein-3; ApoB, apolipoprotein B; ASO, antisense oligonucleotides; eQTL, expression quantitative trait loci; HDL-C, high-density lipoprotein cholesterol; IDL-C, intermediate-density lipoprotein cholesterol; LDL-C, low-density lipoprotein cholesterol; mAb, monoclonal antibody; mRNA, messenger ribonucleic acid; pQTL, protein quantitative trait loci; PTVs, protein-truncating variants; siRNA, small interfering RNA; TG, triglycerides; VLDL-C, very-low-density lipoprotein cholesterol
Methods
Identification of genetic variants influencing liver ANGPTL3 expression
Patients undergoing bariatric surgery at the Quebec Heart and Lung Institute [Institut universitaire de cardiologie et de pneumologie de Québec (IUCPQ)] provided informed consent to participate to the institutional biobank (IUCPQ Obesity Biobank). Clinical information at the time of surgery (sex, age, anthropometry, medication use, medical history, and comorbidities and glycaemic, lipoprotein, and liver enzyme profile) was available for all patients. A total of 246 participants passed the genotyping and RNA sequencing quality controls. Baseline characteristics of the 246 participants of the cohort are presented in Supplementary data online, Table S1. Liver samples were obtained by incisional biopsy of the left lobe and were not cauterised. The grading and staging of histological lesions have been carried out according to the protocol of Brunt et al.27 by pathologists who were blind to the objectives of the study. The liver sample procedure and position of the liver sample are standardised for among all surgeons. A detailed description of DNA and RNA extraction, genome-wide genotyping, RNA sequencing, and mapping of expression quantitative trait loci (eQTL) is available in Supplementary data online, Methods.
Genome-wide association studies of circulating ANGPTL3 levels
The instruments for genetically predicted circulating ANGPTL3 protein levels were obtained from a genetic association study in the deCODE genetics cohort.28 In this study, 4719 plasma protein levels, including ANGPTL3, were measured in 35 559 Icelanders with 4907 aptamers from SomaScan version 4. A detailed description of the proteomic measurements, the genotyping, and genome-wide association analyses have been described previously.28 For the replication, we used the Fenland population-based cohort, where Pietzner et al.29 performed a genome-wide association study (GWAS) for 4775 plasma protein levels, including ANGPTL3, in 10 708 European-descent participants using the SomaScan v4 assay.
Identification of protein-truncating variants in ANGPTL3 and PCSK9 in participants of the UK Biobank
The study design and population of the UK Biobank (UKB) have previously been reported.30 UKB includes more than 500 000 participants between 40 and 69 years old who were recruited from the United Kingdom’s National Health Service (NHS) central registers between 2006 and 2010. The assessment for diseases and serological tests for these participants were done in 22 assessment centres during the same period. Data collection included a self-report questionnaire, physical measures, and blood, urine, and saliva sample collection. Analyses in UKB were conducted under the data application number 25205. A total of 407 668 White British UKB participants were included in the current analysis. Protein-truncating variants (PTVs) in ANGPTL3 and PCSK9 (the latter used as a positive control) were extracted from the 407 668 White British participants with available genotyping data for the corresponding gene in the UKB and were identified from whole-exome sequencing, as previously described.31 We selected those for whom triglyceride levels (field number 30870) and apoB levels (field number 30640) were available. PTVs were defined as variants causing the loss of an exon or a frameshift in the translation or if they introduced a stop codon, a splice acceptor or a splice donor site.32 In order to confirm that the identified PTVs are considered as strong, we used the Ensembl variant effect predictor (VEP) tool33 to extract the Combined Annotation Dependent Depletion (CADD) Phred scores for each variant. We also used GnomAD34 to extract the predicted loss-of-function (pLoF) value for each PTV. We considered a PTV to be strong if it had either a CADD Phred score above 20 (in the top 1% most deleterious substitutions of the genome) or a high-confidence pLoF value. Non-carriers were defined as having none of the variants identified. This procedure was performed independently for ANGPTL3 and PCSK9.
Health impact of protein-truncating variants in ANGPTL3
For each PTV in ANGPTL3, we performed a linear regression on circulating triglyceride levels between non-carriers and carriers. Health effects of PTVs were investigated if PTVs were significantly associated with a reduction of triglyceride levels in both analyses (P < .05). All strong ANGPTL3 PTV carriers were then grouped together, and the impact of carrier status on CAD was determined. To define prevalent CAD, we selected the participants with ICD-10 codes for MI (I21.X, I22.X, I23.X, I24.1, or I25.2), other acute ischaemic heart diseases (IHD) (I24.0, I24.8–9), or chronic IHD (I25.0–25.1, I25.5–25.9). In order to investigate the impact of these variants on CAD, we then performed a logistic regression analysis, adjusted for age at recruitment and sex, on the CAD status between non-carriers and carriers of all strong ANGPTL3 PTVs and obtained odds ratios for the effect of the PTVs on the risk of prevalent CAD by exponentiating the logistic regression coefficient. In order to obtain a positive control for this natural experiment, the same analytical strategy was performed for PTVs in PCSK9, except that variants were considered strong if they were significantly associated with a reduction in the levels of circulating apoB instead of triglycerides.
Supplementary methods and data availability
Additional methods describing the identification of genetic variants influencing liver ANGPTL3 expression, genetic colocalisation, GWAS used for MR, MR of the health impact of liver expression and protein levels of ANGPTL3, MR on the impact of triglyceride levels on apoB levels, baseline characteristics of UKB participants, triglyceride levels, and incident acute pancreatitis in the UKB. Access to UKB data can be granted via the Access Management System of the UKB (https://www.ukbiobank.ac.uk/enable-your-research/apply-for-access). Data access permission for this study was granted under UKB application 25205. The code used to perform this analysis is available at https://github.com/LaboArsenault.
Results
Liver ANGPTL3 expression and cardiometabolic traits and diseases
In order to determine whether genetic variants influence liver expression levels of ANGPTL3, we performed RNA sequencing of liver samples obtained from 246 patients undergoing bariatric surgery (IUCPQ Obesity Biobank). One variant in the ANGPTL3 locus (rs11207978) had a strong effect on the liver ANGPTL3 expression (P = 2 × 10−6). We investigated the impact of this variant on lipoprotein-lipid levels and cardiometabolic outcomes using MR and genetic colocalisation. As presented in Figure 2 and Supplementary data online, Table S4, this SNP had a very strong effect on plasma triglyceride levels (P = 4.47 × 10−258), a more modest effect on LDL cholesterol (P = 2.20 × 10−85), and a weaker effect on apoB levels (P = 9.35 × 10−24). However, this variant was not associated with CAD and other cardiometabolic diseases.
Figure 2.
Impact of genetically predicted liver ANGPTL3 expression on lipoprotein-lipid levels and cardiometabolic diseases. (A) The effects of the strongest SNP (rs11207978) associated with the liver ANGPTL3 expression on lipoprotein-lipid levels and cardiometabolic diseases were obtained using the Wald ratio. Effect sizes (95% CI) are represented by SD or log(OR) change in the outcome per 1-SD increase in liver ANGPTL3 expression. (B) Regional association plots depicting the impact of genetically predicted liver ANGPTL3 expression on lipoprotein-lipid levels and coronary artery disease
ANGPTL3 blood levels and cardiometabolic traits and diseases
To determine whether genetic variants influencing blood levels of ANGPTL3 have an effect on cardiometabolic traits and diseases, we investigated the impact of the top variant in the ANGPTL3 locus influencing blood levels of ANGPTL3 in the deCODE cohort (rs10889352) on lipoprotein-lipid levels and cardiometabolic outcomes using MR and genetic colocalisation. This variant and the liver eQTL were in moderate linkage disequilibrium (R2 = 0.48). Figure 3 and Supplementary data online, Table S5 show the association between the top ANGPTL3 blood protein quantitative trait loci (pQTL) and lipoprotein-lipid levels and cardiometabolic diseases. This variant had a very strong effect on plasma triglyceride levels, a more modest effect on LDL cholesterol, and a weaker effect on apoB levels. This variant was not associated with CAD and other cardiometabolic diseases.
Figure 3.
Impact of genetically predicted plasma ANGPTL3 levels on lipoprotein-lipid levels and cardiometabolic diseases. (A) The effects of the strongest SNP (rs10889352) associated with the plasma ANGPTL3 levels on lipoprotein-lipid levels and cardiometabolic diseases were obtained using the Wald ratio. Effect sizes (95% CI) are represented by SD or log(OR) change in the outcome per 1-SD increase in plasma ANGPTL3 levels. (B) Regional association plots depicting the impact of genetically predicted plasma ANGPTL3 levels on lipoprotein-lipid levels and coronary artery disease
Genome-wide association studies of circulating ANGPTL3 levels
To identify and validate genetic loci that may influence the circulating levels of ANGPTL3 and that could be used in a more comprehensive MR setting, we investigated the results of the deCODE and Fenland studies, which measured ANGPTL3 levels in 35 359 and 10 078 individuals, respectively. Figure 4 presents the results of the GWAS for ANGPTL3 levels in these two cohorts. In deCODE, genetic variation at five loci including ANGPTL3, LPL (the gene encoding lipoprotein lipase), APOA5-APOA4-APOC3-APOA1, ANGPTL4, and APOE were found to influence ANGPTL3 levels at genome-wide significant level (P < 5e-08). The Fenland cohort replicated three of these loci at genome-wide level and two of these loci at P < .01. Supplementary data online, Table S6 presents the effect size and characteristics of the top variant influencing ANGPTL3 levels at these loci in both cohorts. To investigate tissue specificity of the genes influencing ANGPTL3 levels, we assessed genetic colocalisation using visceral and subcutaneous adipose tissue samples from the GTEx project as well as liver samples from the IUCPQ Obesity Biobank. The signal at the ANGPTL3 and APOE loci colocalised with liver expression of ANGPTL3, and the signal at the LPL locus colocalised with subcutaneous expression of LPL (see Supplementary data online, Table S7). The results of this analysis suggest that genes involved in triglyceride-rich lipoprotein metabolism that are expressed in the liver or subcutaneous adipose tissue influence circulating concentrations of ANGPTL3. Variants influencing plasma ANGPTL3 levels were all involved in triglyceride-rich lipoprotein metabolism. To avoid horizontal pleiotropy whereby the use of instrumental variables from genetic regions outside of ANGPTL3 could influence the outcomes via mechanisms unrelated to ANGPTL3, we have opted to only include single-nucleotide polymorphisms (SNPs) in the ANGPTL3 locus for further MR analyses. Using independent genetic instruments with adequate strength (see Supplementary data online, Table S3), we further investigated the impact of genetically predicted ANGPTL3 levels with the inverse-variance-weighted MR method. These results show that blood ANGPTL3 levels have a strong effect on plasma triglyceride levels, a more modest effect on LDL cholesterol, and a weaker effect on apoB levels (Figure 4C). Genetically predicted ANGPTL3 levels were associated with acute pancreatitis. Genetically predicted ANGPTL3 blood levels were also associated with CAD, but the association was weak and not significant after adjusting for multiple testing (P < .05/11 = .0045) (see Supplementary data online, Table S8).
Figure 4.
Genome-wide association studies of circulating angiopoietin-like protein-3 levels. Manhattan plot representing the results of the genome-wide association studies of circulating angiopoietin-like protein-3 levels in the (A) deCODE and (B) Fenland cohorts. (C) The effects of single-nucleotide polymorphisms associated with circulating ANGPTL3 levels on lipoprotein-lipid levels and cardiometabolic diseases were obtained using inverse-variance-weighted Mendelian randomisation. Effect sizes (95% CI) are represented by SD or log(OR) change in the outcome per 1-SD increase in circulating ANGPTL3 levels
Genetically predicted triglyceride levels and apolipoprotein B levels
The results of several recent MR studies and clinical trials provided evidence that lipid-lowering therapies may be effective in reducing ASCVD risk if they provide significant reductions in plasma apoB levels (regardless of their impact on other lipoprotein parameters such as triglyceride or HDL cholesterol levels). Given that variants influencing liver expression of ANGPTL3 or blood protein levels of ANGPTL3 had a strong effect on triglyceride levels but a weaker effect on apoB, we investigated if the reductions in triglycerides predicted reductions in apoB and if these reductions were differently predicted if they were produced by changes in ANGPTL3 levels. Figure 5A presents the result of an MR study on the effects of triglyceride levels on apoB levels with variants in the ANGPTL3 locus presented in red. The results of this analysis suggest a causal effect of triglycerides on apoB levels. Since variants in the ANGPTL3 region are either on the regression line or very close to it, it may be concluded that the effect of reductions in triglycerides can predict reductions in apoB levels and that reductions in triglycerides following the inhibition of ANGPTL3 levels predict reductions in apoB levels as would be expected by most triglyceride-lowering mechanisms.
Figure 5.
Impact of genetic variation at ANGPTL3 on triglyceride and apolipoprotein B levels. (A) Inverse-variance-weighted Mendelian randomisation scatterplot depicting the causal impact of triglyceride-reducing SNPs on apolipoprotein B (apoB) levels. The top liver eQTL-SNP (rs11207978) influencing ANGPTL3 hepatic gene expression levels and the top plasma pQTL-SNP (rs10889352) influencing ANGPTL3 levels are highlighted. (B) Impact of rare and common genetic variants at the ANGPTL3 locus on triglyceride and apoB reductions
Impact of ANGPTL3 protein-truncating variants on triglyceride levels
Using whole-exome sequencing data on 407 668 UKB participants, we identified 647 participants carrying at least one of four ANGPTL3 PTVs associated with lower plasma triglyceride levels (Table 1). These PTVs had a stronger effect on both triglyceride and apoB levels compared with the top eQTL, the top pQTL, and the combination of pQTLs. Carriers of PTVs also had a comparable cardiometabolic risk profile including similar body fat distribution patterns as non-carriers. As presented in Figure 5B, the top liver eQTL-SNP (rs11207978) influencing ANGPTL3 hepatic gene expression levels, the top plasma pQTL-SNP (rs10889352) influencing ANGPTL3 levels, and the combination of independent pQTLs influencing ANGPTL3 levels (per 1-SD decrease in ANGPTL3 levels) as well as ANGPTL3 PTV provide apoB reductions that can be predicted by their reduction in triglyceride levels. Despite having a notable effect on triglyceride levels [−0.37 (interquartile range 0.41) mmol/L], these variants showed a modest effect on apoB levels (−0.06 ± 0.32 g/L). Compared with non-carriers, carriers of ANGPTL3 PTVs had 25.2% reduction in triglyceride levels, 9.7% reduction in LDL cholesterol levels, and 5.9% reduction in apoB levels. Characteristics and individual estimates of ANGPTL3 PTVs associated with lower triglyceride levels in the participants of the UKB are presented in Supplementary data online, Table S9 and Supplementary data online, Table S10. Altogether, these results suggest that genetic variants at the ANGPTL3 locus are associated with small reductions in apoB levels and that the reductions in apoB are proportional to the reduction in triglyceride levels.
Table 1.
Baseline characteristics of the UK Biobank participants according to carrier status of ANGPTL3 variants linked with lower plasma triglyceride levels
| Characteristic | Protein-truncating variants (PTVs) | Liver expression quantitative trait loci (eQTL) | Plasma protein quantitative trait loci (pQTL) | |||
|---|---|---|---|---|---|---|
| Carriers | Non-carriers | Carriers | Non-carriers | Carriers | Non-carriers | |
| n (%) | 647 (0.27) | 236 342 (99.73) | 147 103 (60.19) | 97 290 (39.81) | 225 014 (90.99) | 22 289 (9.01) |
| Male, n (%) | 298 (46.06) | 108 372 (45.85) | 67 353 (45.79) | 44 646 (45.89) | 103 142 (45.84) | 10 217 (45.84) |
| Lipid-lowering therapy, n (%) | 73 (18.53) | 40 335 (26.47) | 24 888 (26.26) | 16 703 (26.6) | 38 194 (26.34) | 3898 (27.04) |
| Blood pressure-lowering therapy, n (%) | 78 (20.05) | 24 259 (17.44) | 15 117 (17.43) | 10 008 (17.52) | 23 164 (17.48) | 2262 (17.28) |
| Smoking, n (%) | 304 (46.99) | 106 890 (45.23) | 66 533 (45.23) | 44 010 (45.24) | 101 800 (45.24) | 10 005 (44.89) |
| Diabetes, n (%) | 35 (5.10) | 11 375 (4.56) | 7052 (4.54) | 4417 (4.54) | 10 812 (4.55) | 1063 (4.48) |
| Age at enrolment, years (SD) | 56.62 (±8.03) | 56.90 (±8.00) | 56.90 (±8.00) | 56.90 (±8.00) | 56.90 (±8.00) | 56.86 (±8.00) |
| Body mass index, kg/m2 (SD) | 27.37 (±4.85) | 27.40 (±4.74) | 27.39 (±4.75) | 27.42 (±4.73) | 27.39 (±4.74) | 27.45 (±4.77) |
| Triglycerides, mmol/L (IQR) | 1.10 (0.66) | 1.47 (1.08) | 1.46 (1.07) | 1.48 (1.08) | 1.46 (1.07) | 1.50 (1.12) |
| HDL cholesterol, mmol/L (IQR) | 1.28 (0.44) | 1.40 (0.50) | 1.40 (0.50) | 1.40 (0.50) | 1.40 (0.50) | 1.39 (0.50) |
| LDL cholesterol, mmol/L (IQR) | 3.18 (0.98) | 3.52 (1.16) | 3.51 (1.16) | 3.53 (1.16) | 3.52 (1.16) | 3.55 (1.17) |
| Apolipoprotein B, g/L (IQR) | 0.96 (0.26) | 1.02 (0.32) | 1.01 (0.32) | 1.02 (0.32) | 1.02 (0.32) | 1.02 (0.32) |
| Apolipoprotein A1, g/L (IQR) | 1.39 (0.32) | 1.51 (0.35) | 1.51 (0.35) | 1.51 (0.35) | 1.51 (0.35) | 1.51 (0.35) |
| Alanine aminotransferase, U/L (IQR) | 19.63 (12.04) | 20.12 (11.90) | 20.10 (11.84) | 20.14 (11.99) | 20.11 (11.87) | 20.13 (12.15) |
| Aspartate aminotransferase, U/L (IQR) | 23.80 (8.00) | 24.30 (7.70) | 24.40 (7.70) | 24.30 (7.80) | 24.30 (7.70) | 24.40 (7.90) |
| Gamma-glutamyltransferase, U/L (IQR) | 25.40 (20.50) | 26.20 (22.30) | 26.20 (22.40) | 26.20 (22.20) | 26.20 (22.40) | 26.20 (22.20) |
| Glucose, mmol/L (IQR) | 4.93 (0.72) | 4.93 (0.71) | 4.93 (0.71) | 4.93 (0.71) | 4.93 (0.71) | 4.93 (0.72) |
| Creatinine, mmol/L (IQR) | 69.10 (18.90) | 70.50 (19.40) | 70.40 (19.30) | 70.50 (19.50) | 70.40 (19.40) | 70.60 (19.60) |
| Abdominal MRI Subsample | ||||||
| n (%) | 45 (0.28) | 15 844 (99.72) | 9862 (60.02) | 6568 (39.98) | 15 161 (91.01) | 1498 (8.99) |
| Liver PDFF (%) (IQR) | 3.10 (3.74) | 2.99 (3.24) | 2.99 (3.22) | 3.00 (3.28) | 3.00 (3.25) | 2.90 (3.02) |
| Total trunk fat volume (L) (IQR) | 9.93 (5.97) | 10.06 (5.80) | 10.05 (5.79) | 10.10 (5.75) | 10.08 (5.81) | 9.83 (5.37) |
| ASAT volume (L) (IQR) | 6.01 (4.00) | 6.25 (3.76) | 6.24 (3.76) | 6.27 (3.73) | 6.25 (3.74) | 6.19 (3.68) |
| VAT volume (L) (IQR) | 3.52 (3.59) | 3.34 (3.18) | 3.31 (3.18) | 3.39 (3.19) | 3.34 (3.19) | 3.25 (3.03) |
HDL, high-density lipoprotein; IQR, interquartile range; LDL, low-density lipoprotein; MRI, magnetic resonance spectroscopy; PDFF, proton density fat fraction; ASAT, abdominal subcutaneous adipose tissue volume, VAT, visceral subcutaneous adipose. Percentages are relative to the number of participants without missing values. Smoking status includes current and past smoking.
Impact of ANGPTL3 protein-truncating variants on coronary artery disease
We finally sought to investigate whether participants of the UKB carrying at least one ANGPTL3 PTVs associated with plasma triglyceride levels had a lower rate of CAD compared with non-carriers. In this population, the CAD event rate was comparable in carriers vs. non-carriers (Figure 6). To present a positive control for this ‘natural’ experiment and demonstrate the analytical pipeline that was developed was able to detect effects on CAD, we investigated whether carriers with at least one PCSK9 PTVs associated with plasma apoB levels (n = 475) had a lower rate of CAD compared with non-carriers. As expected, carriers of sequence variants at the PCSK9 locus associated with lower apoB levels had a lower CAD event rate compared with non-carriers. Altogether, the results of this analysis suggest that despite having important lifelong reduction in triglyceride levels, carriers of ANGPTL3 PTVs have smaller lifelong reductions in apoB levels and a similar CAD risk compared with non-carriers.
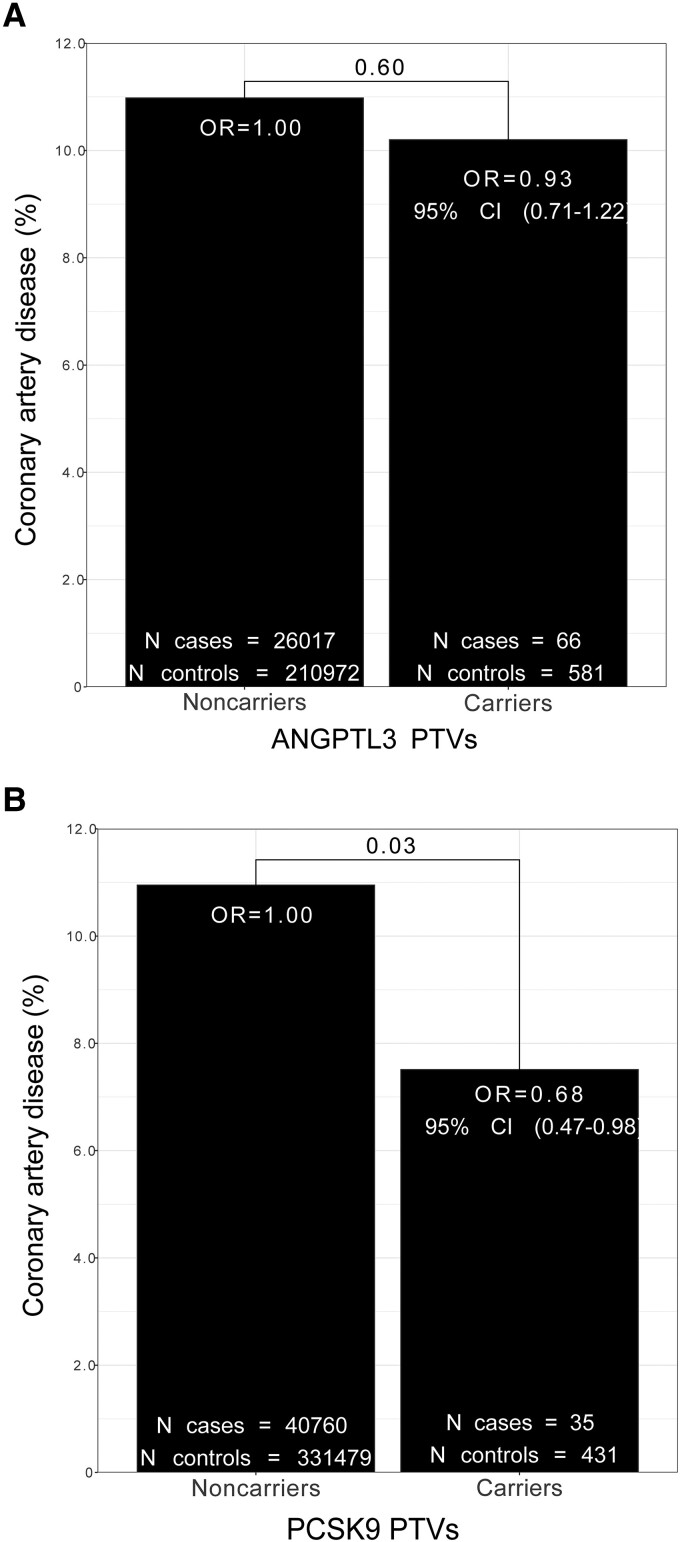
Figure 6.
Impact of ANGPTL3 protein-truncating variants on coronary artery disease rates in the participants of the UK Biobank. (A) Carriers of at least one ANGPTL3 protein-truncating variant (PTV) associated with lower plasma triglyceride levels had a comparable coronary artery disease (CAD) event rate than non-carriers. (B) Carriers of at least one PCSK9 PTV associated with lower plasma apolipoprotein B levels had a lower CAD event rate than non-carriers
Impact of higher triglyceride levels on incident acute pancreatitis in the UK Biobank
Since genetically predicted ANGPTL3 levels were associated with acute pancreatitis, we investigated the association of plasma triglyceride levels among 467 319 UKB participants without acute pancreatitis diagnosis prior to enrolment on incident acute pancreatitis (2523 incident cases). Plasma triglycerides was associated with acute pancreatitis incidence (HR 1.10 95% CI 1.06–1.14, P = 4.6e-08 per each mmol/L increase in triglyceride level). Participants in the top triglyceride level quintile had an HR of incident AP of 2.36 (95% CI 2.06–2.69, P = 6.1e-36) compared with the participants in the bottom quintile (see Supplementary data online, Table S11). The impact of triglyceride levels on incident acute pancreatitis appeared to be stronger in women than in men as women in the top triglyceride level quintile had an HR of incident acute pancreatitis of 3.16 (95% CI 2.58–3.86, P = 5.1e-29) compared with women in the bottom quintile while men in the top triglyceride level quintile had an HR of incident acute pancreatitis of 1.55 (95% CI 1.30–1.85, P = 8.7e-7) compared with men in the bottom quintile.
Discussion
In this study, we explored the potential health effects of reducing triglyceride levels through ANGPTL3 inhibition. Using multiple MR strategies, we mimicked the effect of a liver-targeted, RNA-based ANGPTL3 inhibition strategy, the effect of an antibody-based ANGPTL3 inhibition strategy, and the effect of gene-editing-based ANGPTL3 inhibition strategy. These analyses provided consistent evidence that ANGPTL3 ‘genetic inhibition’ had a strong effect on circulating triglyceride levels, a weak effect on circulating apoB levels, and no effect on CAD (Structured Graphical Abstract). Although the impact of ANGPTL3-mediated triglyceride reductions on ASCVD will ultimately need to be examined in large cardiovascular outcomes trials, our findings suggest that the long-term cardiometabolic benefits of this therapeutic strategy in patients at high residual risk of ASCVD with elevated triglyceride levels may be limited.
Cardiovascular outcome trials and MR studies have provided convincing evidence that lipid-lowering therapies may reduce long-term ASCVD risk if they provide significant reductions in plasma apoB levels and that the reduction in apoB levels is proportional to ASCVD risk reductions.4–6 Our MR findings suggest that although ANGPTL3 inhibition may profoundly reduce plasma triglyceride levels, this strategy provides small apoB reductions. This finding is consistent with the results of randomised clinical trials.14,15,35 In patients with elevated triglyceride levels, type 2 diabetes, and NAFLD, treatment with the liver-targeted antisense oligonucleotide against ANGPTL3 vupanorsen administered once every 4 weeks for 6 months provided large reductions in triglyceride levels (53%) and small reductions in apoB levels (9%) at the dose of 80 mg.14 In the TRANSLATE-TIMI 70 trial, a higher exposure to vupanorsen (160 mg administered once every 4 weeks for 24 weeks) provided large triglyceride reductions (45.9%) and also smaller apoB reductions (12.6%) in statin-treated patients with elevated triglyceride and non-HDL cholesterol levels.15 In smaller phase 1 trials performed in patients with elevated triglycerides and LDL cholesterol levels, treatment with the monoclonal antibody evinacumab provided more potent short-term effects.35 In a single escalating dose study, maximal triglyceride reduction (76.9%) was observed on day 3 with intravenous evinacumab (10 mg/kg) while the maximal triglyceride reduction (83.1%) was observed on day 2 in a multiple escalating dose study. While the corresponding reductions in apoB at these timepoints were ∼10% and 20%, respectively, more important apoB reductions (reaching a nadir of 25% and 35%, respectively) were observed in the following days. Subcutaneous evinacumab injections provided consistent but less potent effects. Altogether, our study, combined with those of randomised clinical trials of short-term ANGPTL3 inhibition with RNA-based and antibody-based therapies, suggests that very potent and near-complete ANGPTL3 reductions will likely be required to reducing significantly apoB levels and ultimately ASCVD risk. We also observed that variants influencing triglyceride levels in the ANGPTL3 region have a comparable effect on apoB reductions than the average of genetic regions influencing triglyceride levels. This finding suggested that reductions in triglycerides following the inhibition of ANGPTL3 would predict reductions in apoB levels as would be expected by most triglyceride-lowering mechanisms. Exploring other triglyceride-reduction mechanisms that provide more potent apoB reductions than what could be expected by reductions in triglyceride levels need to be investigated to provide ASCVD risk reductions in patients with high triglycerides with an elevated ASCVD residual risk.
Although our study provides evidence that ANGPTL3-mediated triglyceride reductions may have a limited impact on apoB and ASCVD, this study does not rule out potential benefits of ANGPTL3 inhibition in other patient populations such as FH. In patients with heterozygous FH with refractory hypercholesterolaemia or homozygous FH, treatment with evinacumab reduced LDL cholesterol levels by ∼50% and provided apoB reductions in the same range.16,36 Interestingly, individuals carrying a loss-of-function mutation in the ANGPTL3 gene and low LDL cholesterol levels do have a lower CAD risk compared with non-carriers.24 In our study, only one PTV was found to be associated with apoB levels, and the association of this PTV with CAD could not be explored since this PTV was found in only six individuals. Our study does not rule out the possibility that ANGPTL3 inhibitors, regardless of the inhibition strategy, could lead to important reductions in apoB levels and a reduction in ASCVD risk in patients with elevated LDL/apoB levels on optimal lipid-lowering therapy. Combined, the results from MR studies and RCTs suggest that ANGPTL3 inhibitors may be a more promising therapeutic approach in patients with very high LDL cholesterol/apoB levels as a result of apoB lowering rather than in patients at high residual risk with high triglyceride levels in whom apoB lowering may not be high enough to promote cardiovascular benefits.
Interestingly, we found an effect of circulating ANGPTL3-mediated reductions in triglyceride levels on acute pancreatitis, suggesting that antibody-based reductions in triglycerides could prevent acute pancreatitis, a disease that is highly prevalent in patients with hypertriglyceridaemia.37 Our observation that triglyceride levels do predict incident acute pancreatitis in the participants of the UKB further suggests that triglyceride levels may cause acute pancreatitis at much lower levels than previously reported. These results are in line with a previous MR study linking triglyceride levels with acute pancreatitis,38 as well as with those of the Copenhagen study reporting effects of genetic variants influencing triglyceride levels via the LPL pathway on acute pancreatitis.37 However, we could not explore whether ANGPTL3 PTVs were associated with acute pancreatitis because of low statistical power. The role of ANGPTL3 inhibitors as therapeutics to prevent acute pancreatitis may need to be further explored. We also found no associations between triglyceride-reducing ANGPTL3 variants (by any mechanism) and liver fat accumulation, liver enzymes, and NAFLD. This finding contrasts those of RNA-based therapies against ANGPTL3 that have reported increases in hepatic fat fractions in some,15 but not all studies.14 Whether increases in liver fat fraction following ANGPTL3 inhibition reflect a true biological phenomenon or an off-target effect of vupanorsen will need to be further explored in longer-term randomised clinical trials.
The results of this study also provide new information on the genetic and biological determinants of circulating ANGPTL3 levels. Through a combination of GWAS and genetic colocalisation with our liver gene expression dataset as well as that of the Genotype-Tissue Expression (GTEx), we found that several genes outside of the ANGPTL3 locus implicated in triglyceride-rich lipoprotein metabolism could influence circulating ANGPTL3 levels. For instance, signals colocalising respectively with hepatic expression of APOE and subcutaneous adipose tissue expression of LPL were found. Other signals at the ANGPTL4 as well as APOA5-APOA4-APOC3-APOA1 loci were identified. This finding may suggest a bidirectional relationship between ANGPTL3 and triglyceride-rich metabolism. Indeed, although ANGPTL3 genetic or pharmacologic modulation does influence lipoprotein metabolism, other determinants of triglyceride-rich metabolism could in turn influence plasma ANGPTL3 levels. Whether these genes influence free or lipoprotein-bound ANGPTL3 needs to be further explored. Because these variants are all involved in triglyceride-rich lipoprotein metabolism, they could introduce horizontal pleiotropy in MR analyses. Consequently, we only include SNPs in the ANGPTL3 locus in our MR analyses aiming to predict long-term effect inhibition of ANGPTL3 with antibody-based strategies.
This study has strengths and limitations. First, the ANGPTL3 eQTL was identified in a study sample of patients undergoing bariatric surgery with a high prevalence of NAFLD. While the presence of liver fat might influence gene expression levels and the effect sizes of some variants on gene expression level, the studied liver ANGPTL3 eQTL was nevertheless strongly associated with blood ANGPTL3 and triglyceride levels, making it a valid genetic instrument to investigate the effect of this eQTL on cardiometabolic outcomes. The results of this study suggest that genetic variation at ANGPTL3 may have a stronger effect on LDL cholesterol than on apoB, despite the fact that up to 80% of apoB likely resides on LDL particles in normolipidemic populations such as the UKB. Since ANGPTL3 inhibitors are currently being investigated in patients with different types of dyslipidaemia, this potential discrepancy may not be relevant to these populations. Additionally, while our study results suggest that triglyceride reductions with ‘genetic inhibition’ of ANGPTL3 may offer little cardiovascular prevention benefits, it must be acknowledged that such an effect was studied in the general population rather than in patients with elevated triglyceride levels. However, the advantage of our study design is that we are able to study almost lifelong reductions in triglyceride levels (since genetic variations are acquired at meiosis). While randomised clinical trials of triglyceride-lowering would reduce triglyceride levels by more than 25% (which is the average triglyceride reduction observed in carriers of ANGPTL3 PTVs), their effect on cardiovascular outcomes can only be investigated over a short period of time in RCTs, making our study design complementary to RCTs. For these reasons, MR studies are less susceptible to reverse causality. Still, our study design was retrospective and observational. Finally, although ANGPTL3 eQTLs and pQTLs had statistically significant effect on triglyceride levels, this lifelong effect was weaker than that of ANGPTL3 PTVs and certainly weaker than ANGPTL3 inhibitors.
In conclusion, our study is the first to use a comprehensive MR study design to predict the long-term health effects of three modes of action evaluating the possible health benefits of a therapeutic target under investigation. Our results suggest that RNA-interfering-based, antibody-based, and gene-editing-based ANGPTL3 inhibition provide important effects on circulating triglyceride levels, less potent effects on circulating apoB levels, and no effects on CAD. Therefore, ANGPTL3-mediated triglyceride reductions may have a limited impact on apoB and ASCVD risk in patients with high triglycerides at high ASCVD risk despite optimal LDL/apoB levels. Near-complete inhibition of ANGPTL3 function, however, could lower apoB levels and potentially reduce ASCVD risk in patients with lipid disorders characterised by elevated concentrations of plasma apoB who are unable to meet guideline-recommended lipoprotein-lipid levels.
Supplementary Material
Acknowledgements
The authors would like to thank all the study participants as well as all investigators of the studies that were used throughout the course of this investigation. We acknowledge the invaluable collaboration of the surgery team, bariatric surgeons, and Biobank staff and would like to thank all the study participants from the Institut universitaire de cardiologie et de pneumologie de Québec (IUCPQ).
Contributor Information
Émilie Gobeil, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada.
Jérôme Bourgault, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada.
Patricia L Mitchell, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada.
Ursula Houessou, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada.
Eloi Gagnon, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada.
Arnaud Girard, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada.
Audrey Paulin, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada.
Hasanga D Manikpurage, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada.
Valérie Côté, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada.
Christian Couture, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada.
Simon Marceau, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada; Department of Surgery, Faculty of Medicine, Université Laval, Québec, Canada.
Yohan Bossé, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada; Department of Molecular Medicine, Faculty of Medicine, Université Laval, Québec, Canada.
Sébastien Thériault, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada; Department of Molecular Biology, Medical Biochemistry and Pathology, Faculty of Medicine, Université Laval, Québec, Canada.
Patrick Mathieu, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada; Department of Surgery, Faculty of Medicine, Université Laval, Québec, Canada.
Marie-Claude Vohl, School of Nutrition, Université Laval, Québec, Canada; Centre Nutrition, santé et société, Institut sur la nutrition et les aliments fonctionnels, Université Laval, Québec, Canada.
André Tchernof, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada; School of Nutrition, Université Laval, Québec, Canada.
Benoit J Arsenault, Centre de recherche de l’Institut universitaire de cardiologie et de pneumologie de Québec—Université Laval, 2725 chemin Ste-Foy, Québec, QC G1V 4G5, Canada; Department of Medicine, Faculty of Medicine, Université Laval, 2325 Rue de l'Université, Québec, QC G1V 0A6, Canada.
Supplementary data
Supplementary data are available at European Heart Journal online.
Declarations
Disclosure of Interest
B.J.A. is a consultant for Novartis, Eli Lilly, Editas Medicine, and Silence Therapeutics and has received research contracts from Pfizer, Ionis Pharmaceuticals, Eli Lilly, and Silence Therapeutics. A.T. receives research funding from Johnson & Johnson, Medtronic, and G.I. windows for studies related to bariatric surgery as well as consulting fees from Bausch Health and Novo Nordisk. The remaining authors declare no disclosure of interest for this contribution.
Data Availability
All genome-wide association study summary statistics used are publicly available. Summary statistics for ANGPTL3 levels from deCODE genetics were downloaded from https://www.decode.com/summarydata/. The full summary statistics for genome-proteome-wide association study for ANGPTL3 from the Fenland cohort were downloaded from: http://www.omicscience.org/apps/pgwas/. All data needed to evaluate this work are present in the paper and/or the Supplementary data online, Materials. Accession number and URLs are provided in the methods section. Access to UK Biobank data can be granted via the Access Management System of the UK Biobank (https://www.ukbiobank.ac.uk/enable-your-research/apply-for-access).
Funding
Pfizer; Canadian Institutes of Health Research; and Institut universitaire de cardiologie et de pneumologie de Québec. The Genotype-Tissue Expression (GTEx) Project was supported by the Common Fund of the Office of the Director of the National Institutes of Health, and by National Cancer Institute, National Human Genome Research Institute, National Heart, Lung and Blood Institute, National Institute on Drug Abuse, National Institute of Mental Health, and National Institute of Neurological Disorders and Stroke. ÉG hold a Doctoral Research Award from the CIHR. JB holds a Masters’ Research Award from the Fonds de recherche du Québec: Santé (FRQS). EG holds a Doctoral Research Award from the FRQS. BJA holds a Senior Scholar Award from the FRQS. YB holds a Canada Research Chair in Genomics of Heart and Lung Diseases. PM holds a FRQS Research Chair on the Pathobiology of Calcific Aortic Valve Disease. MCV is holding a Canada Research Chair in Genomics applied to Nutrition and Metabolic Health.
Ethical Approval
The study was conducted in accordance with the Declaration of Helsinki and approved by the Institutional Review Board (Ethics Committee) of Institut universitaire de cardiologie et de pneumologie de Québec-Université Laval (IUCPQ-UL) (approval number 2021-3656; date of approval: 17 June 2021). For the other publicly available GWAS summary statistics, all participants provided informed consent, and study protocols were approved by their respective local ethical committee.
Pre-registered Clinical Trial Number
Not applicable.
References
- 1. Emerging Risk Factors Collaboration; Di Angelantonio E, Sarwar N, Perry P, Kaptoge S, Ray KK, et al. Major lipids, apolipoproteins, and risk of vascular disease. JAMA 2009;302:1993–2000. 10.1001/jama.2009.1619 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Arsenault BJ, Boekholdt SM, Kastelein JJP. Lipid parameters for measuring risk of cardiovascular disease. Nat Rev Cardiol 2011;8:197–206. 10.1038/nrcardio.2010.223 [DOI] [PubMed] [Google Scholar]
- 3. Sniderman AD, Navar AM, Thanassoulis G. Apolipoprotein B vs low-density lipoprotein cholesterol and non–high-density lipoprotein cholesterol as the primary measure of apolipoprotein B lipoprotein-related risk. JAMA Cardiol 2022;7:257–8. 10.1001/jamacardio.2021.5080 [DOI] [PubMed] [Google Scholar]
- 4. Ference BA, Ginsberg HN, Graham I, Ray KK, Packard CJ, Bruckert E, et al. Low-density lipoproteins cause atherosclerotic cardiovascular disease. 1. Evidence from genetic, epidemiologic, and clinical studies. A consensus statement from the European atherosclerosis society consensus panel. Eur Heart J 2017;38:2459–72. 10.1093/eurheartj/ehx144 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Richardson TG, Sanderson E, Palmer TM, Ala-Korpela M, Ference BA, Davey Smith G, et al. Evaluating the relationship between circulating lipoprotein lipids and apolipoproteins with risk of coronary heart disease: a multivariable Mendelian randomisation analysis. PLoS Med 2020;17:e1003062. 10.1371/journal.pmed.1003062 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Ference BA, Kastelein JJP, Ray KK, Ginsberg HN, Chapman MJ, Packard CJ, et al. Association of triglyceride-lowering LPL variants and LDL-C–lowering LDLR variants with risk of coronary heart disease. JAMA 2019;321:364. 10.1001/jama.2018.20045 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Bhatt DL, Steg PG, Miller M, Brinton EA, Jacobson TA, Ketchum SB, et al. Cardiovascular risk reduction with icosapent ethyl for hypertriglyceridemia. N Engl J Med 2019;380:11–22. 10.1056/nejmoa1812792 [DOI] [PubMed] [Google Scholar]
- 8. Pradhan A D, Glynn RJ, Fruchart J-C, Macfadyen JG, Zaharris ES, Everett BM, et al. Triglyceride lowering with pemafibrate to reduce cardiovascular risk. N Engl J Med 2022;387:1923–34. 10.1056/nejmoa2210645 [DOI] [PubMed] [Google Scholar]
- 9. Nicholls SJ, Lincoff AM, Garcia M, Bash D, Ballantyne CM, Barter PJ, et al. Effect of high-dose Omega-3 fatty acids vs corn oil on Major adverse cardiovascular events in patients at high cardiovascular risk. JAMA 2020;324:2268–80. 10.1001/jama.2020.22258 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Xu Y-X, Redon V, Yu H, Querbes W, Pirruccello J, Liebow A, et al. Role of angiopoietin-like 3 (ANGPTL3) in regulating plasma level of low-density lipoprotein cholesterol. Atherosclerosis 2018;268:196–206. 10.1016/j.atherosclerosis.2017.08.031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Balasubramaniam D, Schroeder O, Russell AM, Fitchett JR, Austin AK, Beyer TP, et al. An anti-ANGPTL3/8 antibody decreases circulating triglycerides by binding to a LPL-inhibitory leucine zipper-like motif. J Lipid Res 2022;63:100198. 10.1016/j.jlr.2022.100198 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Musunuru K, Pirruccello JP, Do R, Peloso GM, Guiducci C, Sougnez C, et al. Exome sequencing, ANGPTL3 mutations, and familial combined hypolipidemia. N Engl J Med 2010;363:2220–7. 10.1056/nejmoa1002926 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Willer CJ, Sanna S, Jackson AU, Scuteri A, Bonnycastle LL, Clarke R, et al. Newly identified loci that influence lipid concentrations and risk of coronary artery disease. Nat Genet 2008;40:161–9. 10.1038/ng.76 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Gaudet D, Karwatowska-Prokopczuk E, Baum SJ, Hurh E, Kingsbury J, Bartlett VJ, et al. Vupanorsen, an N-acetyl galactosamine-conjugated antisense drug to ANGPTL3 mRNA, lowers triglycerides and atherogenic lipoproteins in patients with diabetes, hepatic steatosis, and hypertriglyceridaemia. Eur Heart J 2020;41:3936–45. 10.1093/eurheartj/ehaa689 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Bergmark BA, Marston NA, Bramson CR, Curto M, Ramos V, Jevne A, et al. Effect of vupanorsen on non–high-density lipoprotein cholesterol levels in statin-treated patients with elevated cholesterol: tRANSLATE-TIMI 70. Circulation 2022;145:1377–86. 10.1161/circulationaha.122.059266 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Raal FJ, Rosenson RS, Reeskamp LF, Hovingh GK, Kastelein JJP, Rubba P, et al. Evinacumab for homozygous familial hypercholesterolemia. N Engl J Med 2020;383:711–20. 10.1056/nejmoa2004215 [DOI] [PubMed] [Google Scholar]
- 17. Rosenson RS, Gaudet D, Ballantyne CM, Baum SJ, Bergeron J, Kershaw EE, et al. Evinacumab in severe hypertriglyceridemia with or without lipoprotein lipase pathway mutations: a phase 2 randomized trial. Nat Med 2023;29:729–37. 10.1038/s41591-023-02222-w [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Chadwick AC, Evitt NH, Lv W, Musunuru K. Reduced blood lipid levels with in vivo CRISPR-cas9 base editing of ANGPTL3. Circulation 2018;137:975–7. 10.1161/circulationaha.117.031335 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Khera A, Lee R, Rohde E, Jayaram H, Kathiresan S, Bellinger A. An in vivo CRISPR base editing therapy to inactivate the ANGPTL3 gene: nomination of a development candidate for VERVE-201. Eur Heart J 2022;43:3087. 10.1093/eurheartj/ehac544.308736044989 [DOI] [Google Scholar]
- 20. Cohen JC, Boerwinkle E, Mosley TH, Hobbs HH. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N Engl J Med 2006;354:1264–72. 10.1056/nejmoa054013 [DOI] [PubMed] [Google Scholar]
- 21. Nicholls SJ, Puri R, Anderson T, Ballantyne CM, Cho L, Kastelein JJP, et al. Effect of evolocumab on progression of coronary disease in statin-treated patients. JAMA 2016;316:2373. 10.1001/jama.2016.16951 [DOI] [PubMed] [Google Scholar]
- 22. Sabatine MS, Giugliano RP, Keech AC, Honarpour N, Wiviott SD, Murphy SA, et al. Evolocumab and clinical outcomes in patients with cardiovascular disease. N Engl J Med 2017;376:1713–22. 10.1056/nejmoa1615664 [DOI] [PubMed] [Google Scholar]
- 23. Schwartz GG, Steg PG, Szarek M, Bhatt DL, Bittner VA, Diaz R, et al. Alirocumab and cardiovascular outcomes after acute coronary syndrome. N Engl J Med 2018;379:2097–107. 10.1056/nejmoa1801174 [DOI] [PubMed] [Google Scholar]
- 24. Stitziel NO, Khera AV, Wang X, Bierhals AJ, Vourakis AC, Sperry AE, et al. ANGPTL3 deficiency and protection against coronary artery disease. J Ame Coll Cardiol 2017;69:2054–63. 10.1016/j.jacc.2017.02.030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Dewey FE, Gusarova V, Dunbar RL, O’Dushlaine C, Schurmann C, Gottesman O, et al. Genetic and pharmacologic inactivation of ANGPTL3 and cardiovascular disease. N Engl J Med 2017;377:211–21. 10.1056/nejmoa1612790 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Wang Q, Oliver-Williams C, Raitakari OT, Viikari J, Lehtimäki T, Kähönen M, et al. Metabolic profiling of angiopoietin-like protein 3 and 4 inhibition: a drug-target Mendelian randomization analysis. Eur Heart J 2020;42:1160–9. 10.1093/eurheartj/ehaa972 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Brunt EM, Janney CG, Di Bisceglie AM, Neuschwander-Tetri BA, Bacon BR. Nonalcoholic steatohepatitis: a proposal for grading and staging the histological lesions. Am J Gastroenterol 1999;94:2467–74. 10.1111/j.1572-0241.1999.01377.x [DOI] [PubMed] [Google Scholar]
- 28. Ferkingstad E, Sulem P, Atlason BA, Sveinbjornsson G, Magnusson MI, Styrmisdottir EL, et al. Large-scale integration of the plasma proteome with genetics and disease. Nat Genet 2021;53:1712–21. 10.1038/s41588-021-00978-w [DOI] [PubMed] [Google Scholar]
- 29. Pietzner M, Wheeler E, Carrasco-Zanini J, Cortes A, Koprulu M, Wörheide MA, et al. Mapping the proteo-genomic convergence of human diseases. Science 2021;374:eabj1541. 10.1126/science.abj1541 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Sudlow C, Gallacher J, Allen N, Beral V, Burton P, Danesh J, et al. UK biobank: an open access resource for identifying the causes of a wide range of Complex diseases of middle and old age. PLoS Med 2015;12:e1001779. 10.1371/journal.pmed.1001779 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Backman JD, Li AH, Marcketta A, Sun D, Mbatchou J, Kessler MD, et al. Exome sequencing and analysis of 454,787 UK biobank participants. Nature 2021;599:628–34. 10.1038/s41586-021-04103-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Palmer DS, Howrigan DP, Chapman SB, Adolfsson R, Bass N, Blackwood D, et al. Exome sequencing in bipolar disorder identifies AKAP11 as a risk gene shared with schizophrenia. Nat Genet 2022;54:541–7. 10.1038/s41588-022-01034-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Cunningham F, Allen JE, Allen J, Alvarez-Jarreta J, Amode MR, Armean IM, et al. Ensembl 2022. Nucleic Acids Res 2022;50:D988–95. 10.1093/nar/gkab1049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alföldi J, Wang Q, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 2020;581:434–43. 10.1038/s41586-020-2308-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Ahmad Z, Banerjee P, Hamon S, Chan K-C, Bouzelmat A, Sasiela WJ, et al. Inhibition of angiopoietin-like protein 3 with a monoclonal antibody reduces triglycerides in hypertriglyceridemia. Circulation 2019;140:470–86. 10.1161/circulationaha.118.039107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Rosenson RS, Burgess LJ, Ebenbichler CF, Baum SJ, Stroes ESG, Ali S, et al. Evinacumab in patients with refractory hypercholesterolemia. N Engl J Med 2020;383:2307–19. 10.1056/nejmoa2031049 [DOI] [PubMed] [Google Scholar]
- 37. Hansen SEJ, Madsen CM, Varbo A, Tybjærg-Hansen A, Nordestgaard BG. Genetic variants associated with increased plasma levels of triglycerides, via effects on the lipoprotein lipase pathway, increase risk of acute pancreatitis. Clin Gastroenterol Hepatol 2021;19:1652–60.e1656. 10.1016/j.cgh.2020.08.016 [DOI] [PubMed] [Google Scholar]
- 38. Yuan S, Giovannucci EL, Larsson SC. Gallstone disease, diabetes, calcium, triglycerides, smoking and alcohol consumption and pancreatitis risk: mendelian randomization study. NPJ Genom Med 2021;6:27. 10.1038/s41525-021-00189-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All genome-wide association study summary statistics used are publicly available. Summary statistics for ANGPTL3 levels from deCODE genetics were downloaded from https://www.decode.com/summarydata/. The full summary statistics for genome-proteome-wide association study for ANGPTL3 from the Fenland cohort were downloaded from: http://www.omicscience.org/apps/pgwas/. All data needed to evaluate this work are present in the paper and/or the Supplementary data online, Materials. Accession number and URLs are provided in the methods section. Access to UK Biobank data can be granted via the Access Management System of the UK Biobank (https://www.ukbiobank.ac.uk/enable-your-research/apply-for-access).