ABSTRACT
Bacterial keratitis is a vision-threatening infection mainly caused by Gram-positive bacteria (GPB). Antimicrobial therapy is commonly empirical using broad-spectrum agents with efficacy increasingly compromised by the emergence of antimicrobial resistance. We used a combination of phenotypic tests and genome sequencing to identify the predominant lineages of GPB causing keratitis and to characterize their antimicrobial resistance patterns. A total of 161 isolates, including Staphylococcus aureus (n = 86), coagulase-negative staphylococci (CoNS; n = 34), Streptococcus spp. (n = 34), and Enterococcus faecalis (n = 7), were included. The population of S. aureus isolates consisted mainly of clonal complex 5 (CC5) (30.2%). Similarly, the population of Staphylococcus epidermidis was homogenous with most of them belonging to CC2 (78.3%). Conversely, the genetic population of Streptococcus pneumoniae was highly diverse. Resistance to first-line antibiotics was common among staphylococci, especially among CC5 S. aureus. Methicillin-resistant S. aureus was commonly resistant to fluoroquinolones and azithromycin (78.6%) and tobramycin (57%). One-third of the CoNS were resistant to fluoroquinolones and 53% to azithromycin. Macrolide resistance was commonly caused by erm genes in S. aureus, mphC and msrA in CoNS, and mefA and msr(D) in streptococci. Aminoglycoside resistance in staphylococci was mainly associated with genes commonly found in mobile genetic elements and that encode for nucleotidyltransferases like ant(4′)-Ib and ant(9)-Ia. Fluroquinolone-resistant staphylococci carried from 1 to 4 quinolone resistance-determining region mutations, mainly in the gyrA and parC genes. We found that GPB causing keratitis are associated with strains commonly resistant to first-line topical therapies, especially staphylococcal isolates that are frequently multidrug-resistant and associated with major hospital-adapted epidemic lineages.
KEYWORDS: keratitis, genomics, MRSA, Gram-positive bacteria, antimicrobial resistance, resistome
INTRODUCTION
Microbial keratitis is a costly disease and a leading cause of ocular morbidity and visual loss. It is one of the most important causes of corneal opacification, which is among the four common causes of blindness worldwide following cataracts (1). In the USA, according to a Morbidity and Mortality Weekly Report, infectious keratitis results in approximately 1 million clinical visits to health practitioners and 58,000 visits to emergency departments annually (2). It costs the US healthcare system an estimated 175 million dollars in direct health expenditures and approximately 70 million dollars in Medicare- and Medicaid-related costs (2).
Bacterial keratitis is mainly caused by Gram-positive pathogens such as Staphylococcus aureus, Staphylococcus epidermidis, and Streptococcus pneumoniae and by some Gram-negative bacteria like Pseudomonas aeruginosa and Enterobacteriaceae (3). Because this is a sight-threatening infection with rapid evolution for which culture has a very low sensitivity of pathogen detection (4), patients are commonly treated with empirical therapies at presentation to prevent complications and avoid extensive ocular tissue damage. Most schemes include broad-spectrum antimicrobial agents with good activity against Gram-positive and Gram-negative bacteria, such as fluoroquinolones (FQs), or combinations of aminoglycosides and cephalosporins (5). The choice of antibiotics can be then adapted according to clinical response, cultured organisms, and their susceptibilities.
Increasing antimicrobial resistance due to inappropriate use of antibiotics has become a major public health issue (6). The World Health Organization Global Action Plan on Antimicrobial Resistance has emphasized the importance of antibiotic resistance surveillance programs and research to strengthen the existing knowledge base and fight this growing issue (7). Antimicrobial resistance surveillance studies of ocular isolates have been performed previously on a local or nationwide scale and have shown that ocular bacteria, especially Gram-positive species that account for most cases of keratitis, are becoming increasingly resistant to first-line empirical therapies (8–10). However, these studies do not commonly involve molecular typing to characterize their population structure and the genetic elements associated with phenotypic antimicrobial resistance. To fill this gap, we sought to investigate the genomic epidemiology of Gram-positive organisms causing keratitis at Massachusetts Eye and Ear (MEE) from 2014 to 2017. We used quantitative antimicrobial susceptibility testing and whole-genome sequencing to determine the population structure and the resistome of the major genetic lineages and to identify differences in the phenotypic and genotypic resistance profiles of these isolates.
MATERIALS AND METHODS
Bacterial isolates
A total of 161 Gram-positive isolates were collected from patients with keratitis at MEE from 2014 to 2017. Protocols for obtaining discarded isolates with waived informed consent were approved by the Mass General Brigham Institutional Review Board. Primary clinical specimens collected from corneal infections were obtained by the attending ophthalmologist or resident following institutional guidelines and submitted to the clinical microbiology laboratory for processing. Species identification was performed by using the MicroScan WalkAway system (Beckman Coulter, Brea, CA, USA) following the manufacturer’s protocol. Isolates were stored at −80°C in Microbank cryopreservative tubes (ProLab Diagnostics). Frozen isolates were cultured on 5% sheep blood agar plates (BD Biosciences, San Jose, CA, USA) and incubated at 37°C for 18 to 24 h.
Clinical data collection
Demographic data and risk factors for Gram-positive keratitis infection were collected using the IRB-approved Research Electronic Data Capture (REDCap) tool, hosted by MEE and Harvard Medical School (11). General demographic data included age and sex. Ocular comorbidities, including any history of ocular surgery, ocular surface disease, eyelid disease, contact lens use, and trauma, were collected.
Antimicrobial susceptibility testing
Isolates were tested for susceptibility by broth microdilution following the Clinical and Laboratory Standards Institute (CLSI) M07-A9 document (12). Quality assurance was performed by concurrently testing CLSI-recommended strains. CLSI-approved interpretive breakpoint criteria were applied (13). Minimum inhibitory concentration (MIC)50 and MIC90 were calculated using SPSS software (version 24, IBM, Armonk, NY, USA).
Genome sequencing and assemblies
To understand the population structure of Gram-positive bacteria causing keratitis, isolates included in this study were submitted to whole-genome sequencing. Total DNA was purified from an overnight culture using the DNeasy DNA extraction kit (Qiagen, Valencia, CA, USA). DNA quality was verified on a Bio-Tek Synergy 2 microplate reader (Winooski, VT, USA) prior to quantification using a Qubit fluorometer and dsDNA High-Sensitivity assay kit (Invitrogen, Carlsbad, CA, USA). Library preparation for Illumina sequencing was carried out using the Nextera XT DNA Library Preparation Kit (Illumina, San Diego, CA, USA), according to the manufacturer’s specifications. The quality and quantity of each sample library were measured on a TapeStation instrument (Agilent Technologies, Santa Clara, CA, USA). The genomes were sequenced as 2 × 150- or 250-bp reads on an Illumina HiSeq sequencer, according to the manufacturer’s specifications, with a minimum depth of coverage of 30×. Sequence reads were assembled de novo using CLC Genomics Workbench (CLC Bio, Cambridge, MA, USA). Samples with sequence reads below a quality score of 25 at any position were re-sequenced.
Prediction of STs, antibiotic resistance genes, and serotypes
We used the Center for Genomic Epidemiology pipeline to confirm species identification and sequence types (STs). The Comprehensive Antibiotic Resistance Database (CARD) algorithm was used to identify the pool of acquired antibiotic resistance genes and quinolone resistance-determining region (QRDR) mutations in each genome (https://card.mcmaster.ca/). A cutoff of 95% identity at the nucleotide sequence level was used for the detection of antibiotic resistance genes. Confirmation of QRDR mutations was performed by extracting the topoisomerase genes from assembled genomes and aligning them against a reference sequence using Geneious (v10.1.2). Clonal complexes (CCs) were determined using the MLST data by the goeBURST algorithm (http://www.phyloviz.net/goeburst/). The genetic environment of the transposon Tn554 was analyzed using Geneious v10.1.2. Capsule serotypes in S. pneumoniae isolates were predicted from genome sequences using PneumoCaT (Pneumococcal Capsular Typing) v1.2.1. Determination of newly proposed nomenclature by the GPS project of Global Pneumococcal Sequence Clusters (GPSCs) for internationally distributed pneumococcal lineages (14) was done using PathogenWatch v12.0.4.
Phylogenetic tree
Parsnp v1.2 with default settings was used to create a whole-genome alignment of all genomes for each species (S. aureus, S. epidermidis, and S. pneumoniae) (15). One alignment was created including an outgroup, and another was created without an outgroup. Gubbins (16) v2.3.1 with default settings was used to remove recombination from each alignment and create a phylogeny using RAxML (17). The phylogenies with outgroups were used to determine the topology of the tree. The phylogeny was visualized and annotated with iTol v4 (18). cgMLST phylogenetic relationships among S. aureus isolates were performed using Pathogenwatch v12.0.4.
RESULTS
Bacterial population
A total of 161 Gram-positive bacterial isolates causing keratitis—S. aureus [n = 86, including 14 methicillin-resistant S. aureus (MRSA)], coagulase-negative staphylococci (CoNS) (n = 34, including 15 methicillin-resistant CoNS, S. epidermidis n = 23, Staphylococcus lugdunensis n = 5, Staphylococcus capitis n = 4, and Staphylococcus haemolyticus n = 2), S. pneumoniae (n = 21), viridans group streptococci (VGS) (n = 10, including Streptococcus anginosus n = 1, Streptococcus mitis n = 7, Streptococcus oralis n = 1, and Streptococcus parasanguinis n = 1), Streptococcus pyogenes (n = 1), Streptococcus agalactiae (n = 2), and Enterococcus faecalis (n = 7)—were recovered from patients treated at MEE from 2014 to 2017. Most of these patients were female (93, 59.6%) (Supplementary data 1). Age at presentation ranged from 3 to 96 years (median 60 years old). Methicillin resistance in S. aureus appeared to be more common among older patients (median of patients with MRSA keratitis was 63 years old, and median of patients with MSSA keratitis was 56 years old, not statistically significant). In our population, patients presenting with Gram-positive bacterial keratitis often had a history of glaucoma (18%), diabetes (18%), dry eyes (9.4%), previous keratitis (9.9%), corneal epithelial defect (6.7%), and contact lens wear (6.2%) (Supplementary data 1).
Antimicrobial resistance rates
To determine the rates of antibiotic resistance and the drugs that would optimize treatment outcomes of bacterial keratitis based on in vitro susceptibility profiles, we tested the susceptibility of all isolates against a panel of commonly used antibiotics. Despite the community origins of our collection, we found moderate to high rates of resistance among MSSA isolates to clinically important antibiotics that are often used for the treatment of ocular infections, such as FQs (from 12.5% to 16.7%, MIC90s ranging from 1 to 16 mg/L) and azithromycin (47.2%, MIC90 >16 mg/L) (Table 1). Resistance to these antibiotics was substantially more frequent among MRSA, with 78.6% of the isolates resistant to all FQ drugs tested (MIC90s ranging from 32 to >32 mg/L), except for besifloxacin (MIC90 4 mg/L), for which breakpoints are not available. The later-generation FQs (besifloxacin and moxifloxacin) had in general lower MIC50 and MIC90 compared with the earlier ones (ofloxacin and levofloxacin). Besifloxacin showed the lowest MIC50 and MIC90, and ofloxacin showed the highest. Resistance to azithromycin was also common among MRSA isolates (78.6%, MIC90 >16 mg/L). Resistance to tobramycin went from 6.2% among MSSA isolates (MIC90 1 mg/L) to 57.1% for MRSA (MIC90 >16 mg/L). More than 30% of CoNS were resistant to FQs (MIC90s ranging from 4 to >32 mg/L), and half of them were resistant to azithromycin (52.9%, MIC90 >16 mg/L). Tobramycin demonstrated good in vitro coverage against ocular CoNS isolates with resistance rates below 9% (MIC90 4 mg/L).
TABLE 1.
In vitro antimicrobial activity of antibiotics commonly used to treat bacterial keratitisb
| Organism/antibiotic | MICs (mg/L) | Susceptibility profile (%) | |||
|---|---|---|---|---|---|
| Range | MIC50 | MIC90 | S | NS | |
| MSSA (n = 72) | |||||
| Besifloxacin | ≤0.25–8 | ≤0.25 | 0.5 | ||
| Moxifloxacin | ≤0.25–32 | ≤0.25 | 1 | 87.5 | 12.5 |
| Levofloxacin | ≤0.25 to >32 | ≤0.25 | 2 | 84.7 | 15.3 |
| Ofloxacin | ≤0.25 to >32 | 0.5 | 16 | 83.3 | 16.7 |
| Tobramycin | ≤0.12 to >16 | 0.25 | 1 | 93.1 | 6.2 |
| Azithromycin | 0.5 to >16 | 2 | >16 | 52.8 | 47.2 |
| Vancomycin | 0.5–2 | 1 | 1 | 100 | 0 |
| MRSA (n = 14) | |||||
| Besifloxacin | ≤0.25–8 | 2 | 4 | ||
| Moxifloxacin | ≤0.25–32 | 8 | 32 | 21.4 | 78.6 |
| Levofloxacin | ≤0.25 to >32 | 32 | >32 | 21.4 | 78.6 |
| Ofloxacin | ≤0.25 to >32 | >32 | >32 | 21.4 | 78.6 |
| Tobramycin | ≤0.12 to >16 | >16 | >16 | 42.9 | 57.1 |
| Azithromycin | ≤0.12 to > 6 | >16 | >16 | 21.4 | 78.6 |
| Vancomycin | 0.25–2 | 1 | 1 | 100 | 0 |
| Coagulase negative staphylococci (n = 34) | |||||
| Besifloxacin | ≤0.25–8 | ≤0.25 | 4 | ||
| Moxifloxacin | ≤0.25–32 | ≤0.25 | 32 | 67.6 | 32.4 |
| Levofloxacin | ≤0.25 to >32 | ≤0.25 | >32 | 64.7 | 35.3 |
| Ofloxacin | ≤0.25 to >32 | 0.5 | >32 | 64.7 | 35.3 |
| Tobramycin | ≤0.12 to >16 | ≤ 0.12 | 4 | 91.2 | 8.8 |
| Azithromycin | ≤0.12 to >16 | 8 | >16 | 47.1 | 52.9 |
| Vancomycin | 0.5–4 | 2 | 2 | 100 | 0 |
| S. pneumoniae (n = 13)a | |||||
| Besifloxacin | ≤0.25 | ≤0.25 | ≤0.25 | ||
| Moxifloxacin | ≤0.25–1 | ≤0.25 | ≤0.25 | 100 | 0 |
| Levofloxacin | ≤0.25–4 | 1 | 2 | 92.3 | 7.7 |
| Ofloxacin | ≤0.25–8 | 2 | 4 | 84.6 | 15.4 |
| Azithromycin | ≤0.12 to >16 | ≤0.12 | >16 | 69.2 | 30.8 |
| Vancomycin | 0.25–0.5 | 0.25 | 0.5 | 100 | 0 |
| Penicillin | ≤0.12–1 | ≤0.12 | 0.5 | 69.2 | 30.8 |
| Ceftriaxone | ≤0.12–0.5 | ≤0.12 | ≤0.12 | 92.3 | 7.7 |
| VGS (n = 10) | |||||
| Besifloxacin | ≤0.25–1 | ≤0.25 | 0.5 | ||
| Moxifloxacin | ≤0.25–4 | ≤0.25 | 2 | ||
| Levofloxacin | ≤0.25–32 | 1 | 16 | 80 | 20 |
| Ofloxacin | ≤0.25–32 | 1 | 32 | 70 | 30 |
| Azithromycin | ≤0.12 to >16 | 1 | >16 | 40 | 60 |
| Vancomycin | 0.5 | 0.5 | 0.5 | 100 | 0 |
| Penicillin | ≤0.12–0.5 | ≤0.12 | 0.25 | 80 | 20 |
| Ceftriaxone | ≤0.12–0.5 | ≤0.12 | ≤0.12 | 100 | 0 |
| E. faecalis (n = 7) | |||||
| Besifloxacin | ≤0.25–4 | ≤0.25 | 4 | ||
| Moxifloxacin | ≤0.25–16 | ≤0.25 | 16 | ||
| Levofloxacin | 0.5 to >32 | 1 | >32 | 85.7 | 14.3 |
| Ofloxacin | 0.5 to >32 | 2 | >32 | ||
| Penicillin | 1–2 | 2 | 2 | 100 | 0 |
| Vancomycin | 1–2 | 1 | 2 | 100 | 0 |
MICs were tested for 13/21 S. pneumoniae isolates.
NS, none susceptible; S, susceptible.
We found low resistance rates (<10%) in S. pneumoniae isolates to all antibiotics tested except for ofloxacin (15.4%), azithromycin (30.8%), and penicillin (30.8%). Among VGS isolates, resistance rates were also low (below 20%) to all antibiotics tested except for azithromycin (60%). All our E. faecalis isolates were susceptible to penicillin (Table 1) and to high concentrations of aminoglycosides (data not shown). In our population, all Gram-positive isolates were vancomycin-susceptible.
Population structure of Gram-positive isolates
To understand the population structure of Gram-positive bacteria causing keratitis, our collection was first typed using core genome MLST. As determined by goeBURST analysis, our S. aureus isolates (n = 86) mainly grouped in one of the following major CCs: CC5 (n = 26, 30.2%), followed by CC30 (n = 14, 16.3%) and CC8 (n = 13, 15.1%) (Table 2). These are the major and expanding CCs found in the USA and contain community- and hospital-associated epidemic clones (19). CC2 (n = 18, 78.3%) was the main CC identified among S. epidermidis isolates (n = 23 isolates in total) (Table 2), and separation of subclusters within CC2 was used in the present study as previously described (20). The majority of S. epidermidis isolates in our population belonged to CC2 subcluster II (Table 2), which includes lineages that are widespread in the USA (21). Isolates within subcluster I, which are commonly found in hospital-associated infections in Europe (22, 23) and in about one-third of S. epidermidis infections in the USA (21), were less common in our population. S. pneumoniae isolates (n = 21) were represented by a highly diverse population of strains grouped within 16 STs (Table 2).
TABLE 2.
Species, CC, and ST of the Gram-positive bacteria causing keratitis
| Species (no. of isolates) | CC (no. of isolates) | ST (no. of isolates) |
|---|---|---|
| S. aureus (86) | CC5 (26) | ST5 (21), ST105 (3), ST840 (1), ST146 (1) |
| CC30 (14) | ST30 (14) | |
| CC8 (13) | ST8 (10), ST72 (1), ST1181 (1), ST1150 (1) | |
| CC15 (10) | ST15 (7), ST199 (1), ST333 (1), ST188 (1) | |
| CC45 (10) | ST45 (4), ST1970 (3), ST2498 (1), ST3299 (1), ST2537 (1) | |
| CC398 (3) | ST398 (3) | |
| CC1 (2) | ST1 (2) | |
| ST6 (2) | ST6 (2) | |
| CC12 (1) | ST1156 (1) | |
| Unknown (5) | Unknown (5) | |
| S. epidermidis (23) | CC2-II (14) | ST5 (7), ST73 (3), ST278 (2), ST7 (1), ST83 (1) |
| CC2-I (4) | ST16 (2), ST2 (1), ST22 (1) | |
| Unknown (5) | Unknown (5) | |
| S. pneumoniae (21) | CC199 (3) | ST199 (3) |
| CC62 (2) | ST3019 (1), ST62 (1) | |
| CC433 (2) | ST433 (2) | |
| CC63 (1) | ST11818 (1) | |
| CC66 (1) | ST66 (1) | |
| CC97 (1) | ST14056 (1) | |
| CC100 (1) | ST100 (1) | |
| CC439 (1) | ST42 (1) | |
| CC180 (1) | ST2183 (1) | |
| CC1262 (1) | ST1262 (1) | |
| CC383 (1) | ST1840 (1) | |
| CC5178 (1) | ST5178 (1) | |
| CC558 (1) | ST558 (1) | |
| CC338 (1) | ST338 (1) | |
| CC9355 (1) | ST9355 (1) | |
| Unknown (2) | Unknown (2) |
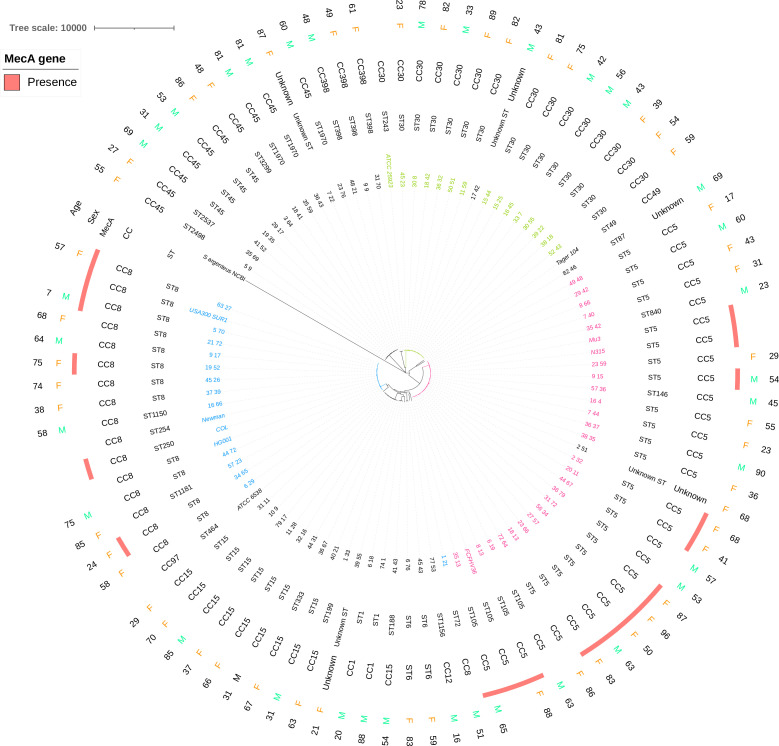
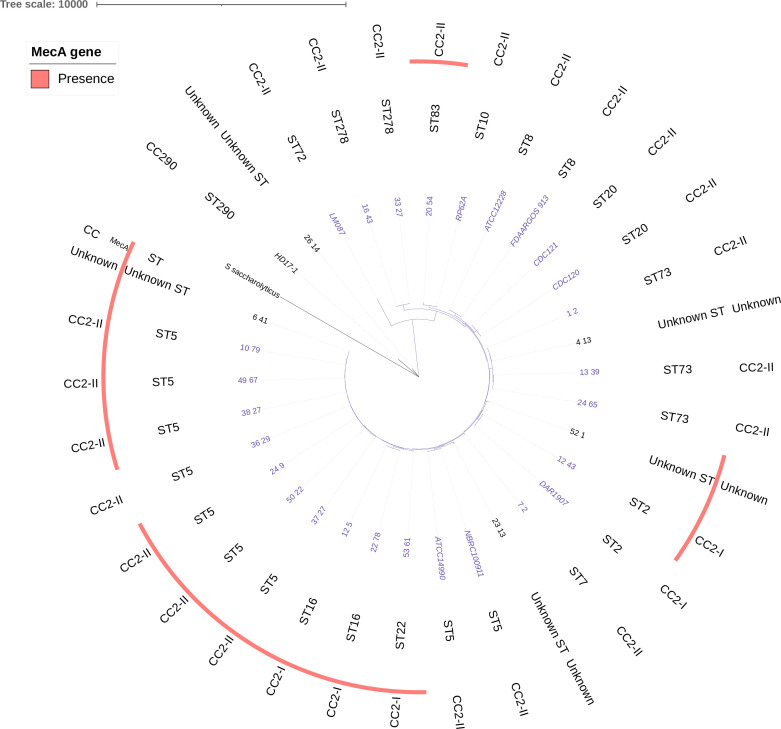
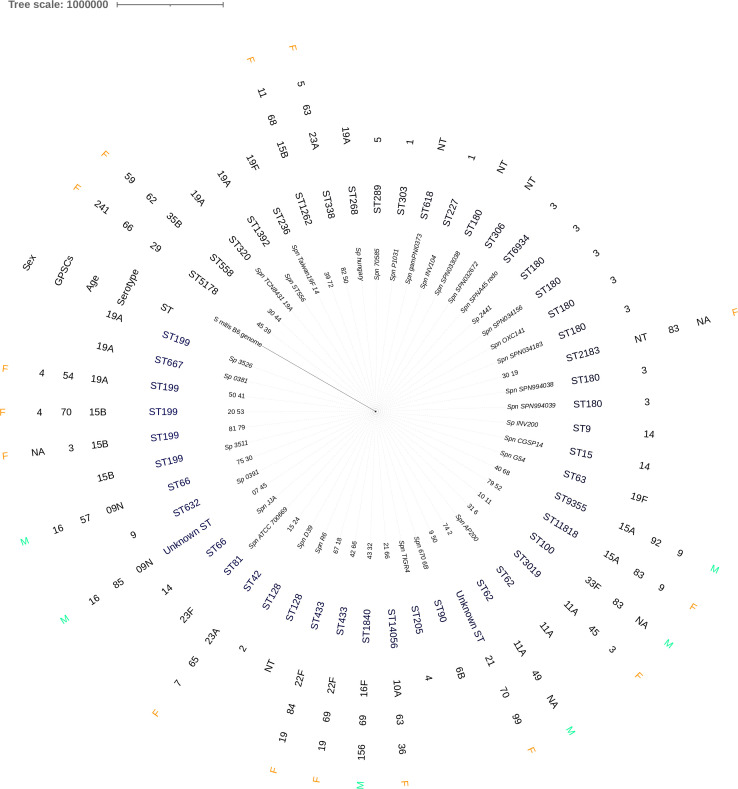
To correlate the MLST-based population structure findings with the phylogenomics of this collection, a single-nucleotide polymorphism (SNP)-based maximum-likelihood tree was generated for the most common species included in our study. S. aureus isolates belonging to CC5 (in pink), CC30 (in green), and CC8 (in blue) formed three resolved groups (except for ST72 that separated from CC8 strains; Fig. 1) as determined by phylogenomic analysis. S. epidermidis isolates belonging to CC2 (in purple) formed a deeply resolved group (Fig. 2). Lineages causing pneumococcal keratitis did not cluster together (Fig. 3); rather, they were scattered across the tree and interspersed with strains that cause infections in other body sites, confirming that they form a highly diverse population of unrelated isolates as determined by MLST. The diversity of this population was also confirmed by assigning our isolates to the GPSCs. In total, 13 unique GPSCs were found (Fig. 3). The genotypically heterogeneous population of keratitis isolates also presents with a high degree of diversity at the capsule serotype level (Fig. 3). Importantly, the SNP-based phylogenetic tree was reconstructed after the removal of recombinant DNA segments, demonstrating that this population structure was not driven by recombination.
Fig 1.
SNP-based maximum-likelihood phylogenetic tree of the S. aureus keratitis population generated from a core-genome alignment. All comparator genomes included in the tree were in italic.
Fig 2.
SNP-based maximum-likelihood phylogenetic tree of the S. epidermidis keratitis population generated from a core-genome alignment. All comparator genomes included in the tree were in italic.
Fig 3.
SNP-based maximum-likelihood phylogenetic tree of the S. pneumoniae keratitis population generated from a core-genome alignment. All comparator genomes included in the tree were in italic.
Acquired antibiotic resistance genes
To characterize the genotypic antibiotic resistance profile of Gram-positive bacteria causing keratitis, we used their draft genomes to identify the pool of acquired resistance genes carried by these isolates using the CARD database. CARD analysis revealed that antimicrobial resistance in these keratitis isolates is achieved through various mechanisms including antibiotic inactivation (aph(3′)-III, ant(4′)-Ib, ant(9)-Ia, mphC, and fosB), target alteration (erm genes and mecA), antibiotic target replacement (dfrG, dfrC, and dfrE), and drug efflux (msr(A), mef(A)-msrD, tetK, and tetM). Fourteen (16.3%) of our S. aureus isolates causing keratitis that were also phenotypically resistant to methicillin harbored the gene mecA (Table 3). The occurrence of methicillin resistance was enriched among certain lineages, with the vast majority of MRSA isolates grouped within CC5 (71.4%) and the rest of them belonging to CC8 (Fig. S1). In line with what we observed phenotypically, MRSA harbored more resistance genes than MSSA (Fig. S1). Resistance to aminoglycosides in S. aureus CC8 isolates was mainly associated with the aminoglycoside phosphotransferase gene aph(3′)-lll, whereas for S. aureus CC5 isolates, it was mostly caused by the aminoglycoside nucleotidyltransferase gene ant(9)-Ia (Fig. S1). Less frequently (10.5%), resistance to aminoglycosides in S. aureus was also caused by ant(4′)-Ib (Table 3). The aminoglycoside tobramycin is commonly used to treat keratitis. Among the acquired resistance genes found in our population, ant(4′)-Ib and aac(6′)-aph(2″) could cause tobramycin resistance whereas aph(3′)-lll and ant(9)-Ia confer resistance to kanamycin/neomycin and spectinomycin, respectively. Macrolide resistance in S. aureus was frequently associated with mphC/msr(A) genotype in CC8 strains and with ermA gene in the CC5 group (Fig. S1). Our genotypic data correlate with what we observed phenotypically as 88.9% of S. aureus harboring ant(4′)-Ib were resistant to tobramycin (Supplementary data 2). Regarding macrolide resistance, 96.7% of S. aureus harboring erm genes and 90% of S. aureus harboring mph(C)/msr(A) genotype were resistant to azithromycin. Almost three out of four S. aureus harbored the gene fosB that confers resistance to fosfomycin (Table 3); 26.7% of S. aureus co-harbored ant(9)-Ia and ermA genes that confer resistance to aminoglycosides and to macrolides, lincosamides, and streptogramin B (Table 3). By looking at the genetic environment of the genes, we found that ant(9)-Ia and ermA genes were localized in a mobile genetic element, the Tn554 transposon (24) (Fig. S2).
TABLE 3.
Acquired resistant genes in Gram-positive bacteria causing keratitis according to the speciesa
| Antibiotic classes/genes no. (%) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AS | METHI | MLSb | FOS | FA | TET | DP | ||||||||||||||
| Species (no.) | aac(6′)-aph(2″) | aph(3′)-lll | ant(4′)-lb | ant(9)-Ia | mecA | ermA | ermB | ermC | ermT | mefA | mphC | msr(A) | msr(D) | fos(B) | fusB | tet(K) | tet(M) | dfrG | dfrC | dfrE |
| S. aureus (86) | 0 (0) | 10 (11.6) | 9 (10.5) | 23 (26.7) | 14 (16.3) | 23 (26.7) | 0 (0) | 3 (3.5) | 4 (4.7) | 0 (0) | 9 (10.5) | 10 (11.7) | 0 (0) | 62 (72.1) | 0 (0) | 2 (2.3) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| CoNS (34) | 3 (8.8) | 3 (8.8) | 8 (23.5) | 1 (2.9) | 15 (44.1) | 1 (2.9) | 0 (0) | 4 (11.8) | 0 (0) | 0 (0) | 14 (41.2) | 15 (44.1) | 0 (0) | 0 (0) | 4 (11.8) | 6 (17.6) | 0 (0) | 7 (20.6) | 19 (55.8) | 0 (0) |
| S. pneumoniae (21) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 2 (9.5) | 0 (0) | 0 (0) | 4 (19.0) | 0 (0) | 0 (0) | 4 (19.0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| VGS (10) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 3 (30.0) | 0 (0) | 0 (0) | 7 (70.0) | 0 (0) | 0 (0) | 6 (60.0) | 0 (0) | 0 (0) | 0 (0) | 4 (40.0) | 0 (0) | 0 (0) | 0 (0) |
| E. faecalis (7) | 2 (28.6) | 1 (14.3) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 1 (14.3) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 7 (100) | 0 (0) | 1 (14.3) | 7 (100) |
AS, aminoglycosides; DP, diaminopyrimidine; FA, fusidic acid; FOS, fosfomycin; L, lincosamides; M, macrolides; METHI, methicillin; S, streptogramin b; TET, tetracycline.
Macrolide resistance in CoNS was frequently associated with the mphC/msr(A) genotype and less commonly with the errmC gene (Table 3). Genotypic/phenotypic correlation analysis showed that 80% of CoNS harboring erm genes and 93.3% of CoNS harboring mph(C)/msr(A) genotype were resistant to azithromycin (Supplementary data 2). CoNS were also positive for the dfrC (55.8%) and tetK genes (17.6%) that confer resistance to diaminopyrimidine and tetracycline, respectively (Table 3).
S. pneumoniae isolates often carried macrolide resistance genes (ermB and mefA/msrD) (Table 3) integrated into mobile genetic elements. mefA/msrD genes were located in the macrolide efflux genetic assembly (MEGA) cassette, and ermB was localized in the Tn-916-like transposon (25). ermB gene was also found in VGS (30%) and E. faecalis isolates (14.3%) (Table 3); 80% of Streptococcus spp. positive for the ermB gene and 72.7% harboring mefA/msrD genotype showed phenotypic resistance to azithromycin (Supplementary data 2).
Amino acid changes in the QRDR of DNA gyrase and topoisomerase IV
As FQs are antibiotics commonly used in ophthalmology for the prevention and treatment of ocular infections (5), we looked at mutations in the QRDR. We found between one and four mutations in the QRDR of the topoisomerase subunits GyrA (DNA gyrase) and ParC (topoisomerase IV) for staphylococci isolates (21.7% of S. epidermidis and 25.0% of S. aureus) (Table 4). In the gyrA gene, the substitution of a serine for a leucine at codon 84 (S84L) was found for 88% of S. aureus isolates carrying QRDR mutations (n = 25), whereas for S. epidermidis isolates, the serine was substituted either by a phenylalanine (S84F) or by a tyrosine (S84Y). In addition, a second point mutation at codon 88 (E88K) was also identified for five S. epidermidis isolates and for three S. aureus isolates.
TABLE 4.
Distribution of FQs resistance genotypes and phenotypes for S. epidermidis (n = 5) and S. aureus (n = 21) isolates
| Strains no. (%) of isolates | Genotypic resistance | Range of MICs (in mg/L) | % Phenotypica agreement | |||||
|---|---|---|---|---|---|---|---|---|
| GyrA | ParC | ParE | BESI | MOX | LEV | OFLO | ||
| S epidermidis (n = 5; 21.7% out of the total) | ||||||||
| 1 (20.0) | E88K S84F | S80F D84Y | 8 | 32 | 32 | >32 | 100 | |
| 1 (20.0) | E88K S84Y | S80F D84Y | 4 | 16 | >32 | >32 | 100 | |
| 3 (60.0) | E88K S84F | S80Y D84Y | 4–8 | 32 | >32 | >32 | 100 | |
| S. aureus (n = 21; 25% out of the total) | ||||||||
| 2 (9.5) | S84L | S80F | D434N | ≤0.25–2 | 1–8 | >32 | >32 | 100 |
| 3 (14.3) | S84L | S80F | 1–2 | 2–8 | 8 to >32 | >32 | 100 | |
| 1 (4.8) | S84L | S80Y | 1 | 2 | 8 | 16 | 100 | |
| 9 (42.9) | S84L | S80Y E84G | 0.5–4 | 1–8 | 2 to >32 | 4 to >32 | 100 | |
| 3 (14.3) | S84L E88K | S80Y E84G | 2–8 | 4–32 | 8 to >32 | >32 | 100 | |
| 2 (9.5) | S80F | ≤0.25 | ≤0.25 | 1 | 2 | 100b | ||
| 1 (4.8) | S80Y | 0.50 | 1.00 | 4.00 | 8.00 | 100 | ||
Based on ofloxacin breakpoints.
Intermediate.
In the parC gene, a single point mutation (S80F or S80Y) was found in nine S. aureus isolates carrying QRDR mutations (36%) whereas the remaining 12 isolates harbored a double point mutation S80Y and E84G (64%). S. aureus with only a single point mutation in parC demonstrated lower-level resistance to FQs with MICs ranging from 1 to 8 mg/L for ofloxacin and levofloxacin and from ≤0.25 to 1 mg/L for moxifloxacin and besifloxacin. Isolates with one mutation in parC in combination with one in gyrA had higher MICs ranging from 8 to >32 mg/L for ofloxacin and levofloxacin and from 1 to 8 mg/L for moxifloxacin and ≤0.25 to 2 mg/L for besifloxacin. All the S. epidermidis isolates carrying QRDR mutations (n = 5) harbored a double point mutation with two different combinations, including S80F and D84Y in two isolates (40%) and S80Y and D84Y in three isolates (60%).
Isolates with a double point mutation in the gyrA gene and in the parC gene had higher MICs to all the FQs tested, including besifloxacin (MICs ranged from 2 to 8 mg/L) and moxifloxacin (MICs ranged from 4 to 32 mg/L) (Table 4). Most S. aureus carrying QRDR mutations accumulated two mutations in parC while showing one mutation in gyrA (42.9%). The lack of one additional mutation in gyrA slightly lowered the MICs when compared with isolates carrying double mutations in gyrA and parC, but the MIC range was still within the resistance category and ranged from 1 to 8 mg/L for moxifloxacin, 2 to >32 mg/L for levofloxacin, and 4 to >32 mg/L for ofloxacin (Table 4). Two S. aureus isolates harbored a single point mutation in the parE gene (D434N), in combination with one SNP in gyrA and one in parC, which resulted in high-level resistance to older FQs (MICs >32 mg/L) and lower to newer molecules (MICs 0.25–8 mg/L).
We did not find any QRDR mutations in S. pneumoniae isolates that were resistant to FQs. These isolates were mostly intermediate resistant to ofloxacin and levofloxacin and susceptible to moxifloxacin. We found both a major facilitator superfamily transporter called PmrA as well as an ATP-binding-cassette transporter called PatAB that is likely to have contributed to reduced susceptibility to older FQs in these isolates (26, 27). The MICs of isolates carrying these genes ranged from ≤0.25 to 1 mg/L for moxifloxacin and besifloxacin and from ≤0.25 to 8 mg/L for levofloxacin and ofloxacin.
DISCUSSION
Microbial keratitis is a serious and potentially blinding condition that must be treated promptly to preserve vision. While this infection can be caused by many types of pathogens including bacteria, fungi, protozoa, and viruses, bacterial etiologies are most common in developed countries, led by Gram-positive species (3). In our study, S. aureus (53.4%), CoNS (21.1%), and S. pneumoniae (13%) were the leading causes of Gram-positive bacterial keratitis. This proportion is similar to what has been found in other US centers (28, 29). In a retrospective study conducted at the University of Pittsburgh that analyzed the distribution of species involved in bacterial keratitis from January 1993 to November 2012, S. aureus isolates represented 47% of the Gram-positive isolates (30). CoNS (all species) represented between 16.9% and 31.6% of the Gram-positive isolates causing keratitis in the USA (28, 29). The ARMOR study showed that S. epidermidis, S. capitis, and Staphylococcus hominis are the most common species of CoNS causing ocular infections in the USA (8). S. pneumoniae is the cause of 6%–10% of keratitis cases in the USA (14%–19% among the Gram-positive isolates) (28, 31), a prevalence similar to what we found in our population. The distribution of bacterial species causing ocular infections is slightly different in other parts of the world, especially in Asia (3, 32). In India, for example, S. pneumoniae isolates are the most commonly identified cause of bacterial keratitis (32, 33).
By using genomic approaches, we found that S. aureus isolates causing keratitis in a population of patients representative of New England were dominated by strains within the CC5, which have been found to be leading causes of antibiotic-resistant hospital-associated MRSA infections in the USA (34). Our previous and current studies suggest that CC5 S. aureus seems to be well adapted to cause infections of the wet epithelial tissues of the ocular surface, especially corneal infections (35). The predominance of CC5 in keratitis is surprising for a disease typically acquired in the community. Our results showed that despite their hospital association, infections caused by CC5 MRSA strains also occur in the community. Nasal carriage of CC5 MRSA has been identified in patients who are residents of long-term care facilities (36) and also in non-institutionalized individuals with no known risk factors for nasal colonization with these hospital-adapted clones (37, 38). These point to the existence of reservoirs for CC5 MRSA strains in community settings. Given the well-established associations between MRSA nasal carriage consequent bloodstream and wound infections (39, 40), we might also suspect nasal carriage to be a risk factor for keratitis, particularly due to anatomical proximity.
The majority of S. epidermidis isolates examined in the present study belonged mainly to CC2 subcluster II, a subcluster commonly found in US hospitals (21). Lineages within CC2-II were also found to be the most prevalent among ocular S. epidermidis isolates from Brazil (41). In Europe, the most common clones isolated from the hospital environment also belonged to CC2 but within subcluster I (23, 42, 43). On the other hand, our S. pneumoniae isolates causing keratitis were represented by a highly diverse population grouped within 16 STs. This heterogeneous population was formed by isolates grouped within a variety of CCs with no apparent clonal dominance, carried 13 different capsule operon types, and appears to mirror the current post–PCV-13 strain compositions in invasive diseases (44) and carriage (45), where PCV-13 serotypes other than serotype 3 are relatively uncommon. This greatly differs from what we have found for patients with pneumococcal conjunctivitis where most isolates are ST448 or closely related strains (25, 46). Although the population structure of Gram-positive keratitis isolates seems to be similar to non-ocular isolates from the USA, our results may not be generalizable to other regions as differences in the diversity of circulating genotypes may exist in other geographies and patient populations.
Antimicrobial-resistant bacteria causing ocular infections increase the risk of treatment failure with potentially sight-threatening outcomes (47). In our population, staphylococci isolates were commonly resistant to empirical antibiotics used to treat bacterial keratitis. FQs are the most frequently used monotherapies for bacterial keratitis. Our population of staphylococci isolates from keratitis showed different degrees of resistance to FQs. The rates of FQ resistance in our population were comparable to what has been found in other local or national studies (8, 28), which showed that up to 8% of MSSA, 90% of MRSA, and 75% of CoNS isolates from keratitis are resistant to moxifloxacin (28), one of the most commonly used FQ eyedrops to treat bacterial ocular infections. For other antibiotics, our results were also consistent with those from the ARMOR program and other local studies, which found resistance to macrolides ranging from 28% to 47.2% in MSSA, from 92.9% to 96% in MRSA, and from 61.4% to 63.2% in CoNS, whereas aminoglycoside resistance was below 10% for MSSA and CoNS and close to 40% for MRSA (8, 28).
Comparable to a large study from San Francisco (28), we found that around one out of six of our S. aureus keratitis isolates and almost half of CoNS were methicillin-resistant. Higher levels of methicillin resistance in S. aureus were found in our study among isolates from older patients, consistent with the ARMOR study (8, 10). The nature and frequency of healthcare exposures in older populations, including antibiotic use and hospitalizations, are risk factors for colonization by drug-resistant organisms (48). Unsurprisingly, MRSA isolates in our study were more frequently resistant to the FQs, macrolides, and aminoglycosides than MSSA (8, 28). This phenotypic resistance was also associated with the population structure of our isolates as most of our MRSA belonged to CC5 (71.4%), representing the most highly antibiotic-resistant MRSA clade. MRSA CC5 isolates from invasive and non-invasive infections are commonly MDR with resistance rates between 84% and 99% to macrolides, lincosamides, FQs, and tetracycline (49). S. epidermidis CC2 is also a lineage associated with multidrug resistance (23, 41, 50). In our study, these strains were commonly resistant to FQs and macrolides. A study that characterized S. epidermidis CC2 keratitis isolates from Brazil found similar results and demonstrated that >60% of their isolates were also resistant to FQs and macrolides (41).
Isolates of S. pneumoniae showed high levels of in vitro resistance (30%) to azithromycin. Resistance to macrolides in S. pneumoniae isolated in the USA is relatively common, with rates ranging from 20% to 40% (51). Other antibiotics like FQs (especially moxifloxacin) and ceftriaxone had good coverage against our ocular pneumococci population as also found by other studies (8, 28).
In line with what we observed phenotypically, the characterization of the resistome showed a variety of acquired resistance genes and QRDR mutations that confer resistance to first-line topical therapies. Resistance caused by methylation of the ribosomal target of the macrolides remains one of the most widespread mechanisms of resistance to macrolides (52). It leads to cross-resistance to macrolides, lincosamides, and streptogramins B, resulting in the MLSB phenotype. Three major different erm genes are detected in Gram-positive bacteria: ermA, ermB, and ermC (53, 54). As we found in our population, ermA and ermC are typically present in staphylococci isolates, whereas ermB is mostly spread in streptococci and enterococci isolates (53). Spreading of macrolide resistance genes between S. pneumoniae isolates is likely facilitated by horizontal gene transfer, since these mutations are found in mobile genetic elements like transposons (Tn916-like transposon) or chromosomal cassettes (MEGA cassette) (25). Elements that determine resistance to multiple drugs are common in MRSA strains such as gene cassettes with integrated transposons that carry various antimicrobial-resistant genes (55, 56). In our CC5 population, the mecA gene is integrated into a SCCmec cassette type II that often also carries a full transposon (Tn554) containing the ermA and ant(9)-Ia genes, both present in 90% of our MRSA CC5 isolates. Acquisition of this gene cassette results in resistance not only to all beta-lactam antibiotics (conferred by the mecA gene) but also to aminoglycosides due to ant(9)-Ia and MLSB phenotype that is conferred by the gene ermA. Furthermore, CC5 is the principal genetic background within S. aureus upon which full resistance to vancomycin has arisen by the acquisition of the vanA operon from Enterococcus spp. (57). S. epidermidis CC2 is also a widespread multidrug-resistant, hospital-adapted lineages harboring genes and mutations conferring resistance to last-resort antibiotics such as linezolid (23, 50). Although we found good agreement in genotype-to-phenotype correlations in general for the pool of acquired antimicrobial resistance genes found in our isolates and antimicrobial agents tested, it is relevant to note that in some instances, the presence of a given AMR gene did not predict in vitro phenotypic resistance for a small percentage of isolates (e.g., around 10% of ant(4′)-Ib-positive isolates were susceptible to tobramycin; Table S2). These discrepancies are found for non-ocular isolates as well and are commonly a result of methodological issues or are related to the specific genes being detected (e.g., gene detected but no longer functional due to genetic alterations or mechanism of resistance is inducible and not detected by broth microdilution in the absence of a drug that derepresses gene expression) (58).
Accumulation of QRDR mutations in the genes of the topoisomerases is the main mechanism of resistance to FQs in staphylococci species (59, 60). Other mechanisms include efflux pumps that result in reductions in cytoplasmic drug concentration (61–63). In our population, all FQ-resistant S. aureus and S. epidermidis isolates carried from one to four QRDR mutations mainly in the gyrA or parC genes, with FQ MIC levels correlating with the number of mutations the isolates accumulated in the QRDR regions of the topoisomerase genes. Isolates with double point mutations in one of those genes had higher MICs to all the FQs tested, including the newer molecules besifloxacin and moxifloxacin. Previous studies have also shown that a higher number of substitutions in the gyrA and parC of ocular staphylococci isolates are related to higher MICs to the latest-generation FQs (20, 64). They found similar MIC ranges for isolates carrying four QRDR mutations, with moxifloxacin MICs ranging from 4 to >32 mg/L and besifloxacin MICs ranging from 1 to 8 mg/L (20, 64). Although overexpression of efflux pumps might play a role in high-level FQ resistance, some studies suggest that efflux pumps do not play a major role in resistance to besifloxacin, moxifloxacin, gatifloxacin, and levofloxacin in staphylococci when compared to the effect of mutations in the genes encoding for DNA gyrase and topoisomerase IV (64, 65). In addition, we found that besifloxacin had greater potency against FQ-resistant strains suggesting that besifloxacin is less affected by topoisomerase mutations compared with older FQs. In the 10-year ARMOR data set, besifloxacin MIC90s were at least fourfold lower than moxifloxacin and gatifloxacin and at least 16-fold lower than ciprofloxacin and levofloxacin for Gram-positive bacteria including S. aureus, CoNS, and S. pneumoniae (8). The study conducted by Sanfilippo et al. demonstrated that as the number of mutations increases in genes encoding for DNA gyrase and topoisomerase IV, MIC values increase for all tested FQs (besifloxacin, moxifloxacin, gatifloxacin, ciprofloxacin, and levofloxacin). The magnitude of this increase for besifloxacin, however, was the smallest (128-fold between susceptible and the most resistant strains) compared to all other FQs (1,024- to 2,048-fold) (64). Unlike all other ophthalmic FQs, besifloxacin is not approved for systemic use, nor has it been used in agriculture or animal husbandry (66). Because these are two major drivers for the selection of resistance among bacteria (67), it has been suggested that the limitation to ocular use may slow the development of bacterial resistance to besifloxacin (68).
In summary, we showed that the population of S. aureus isolates recovered from patients with bacterial keratitis seen at the MEE is dominated by CC5 lineages that are commonly associated with antimicrobial resistance genes and QRDR mutations that result in high levels of phenotypic resistance to first-line topical therapies. S. epidermidis isolates causing keratitis in our patient population are dominated by the widespread and commonly MDR CC2, which was often resistant to antibiotics used empirically to treat keratitis. S. pneumoniae causing keratitis in our patient population formed a highly diverse population of strains often susceptible to first-line topical therapies. To prevent treatment failures, especially among staphylococcal species, close and continuing monitoring of antimicrobial resistance rates and tracking of high-risk clones that cause serious eye infections are important epidemiological tools that can help guide empirical treatments locally and support the development of novel approaches for prevention and treatment.
ACKNOWLEDGMENTS
The authors thank medical technologists from the Clinical Microbiology Laboratory at MEE, including Lisa Bove and Nancy Sutcliffe for their support in creating a microbial repository of strains isolated from ocular infections.
This work was supported in part by the New England Corneal Transplant Research Fund and the Massachusetts Lions Eye Research Fund (PJMB) and by NIH grant EY031600. C.A. was supported by a scholarship from Fondation pour la Recherche Médicale (FDM202006011203) and FEMS (G0-2020-186). Funding agencies had no role in study design, data analysis, decision to publish, or preparation of the manuscript.
Contributor Information
Paulo J. M. Bispo, Email: paulo_bispo@meei.harvard.edu.
Helen Boucher, Tufts University - New England Medical Center, Boston, Massachusetts, USA.
SUPPLEMENTAL MATERIAL
The following material is available online at https://doi.org/10.1128/aac.01247-23.
Demographic and clinicopathologic data for patients with bacterial keratitis at MEE, 2014-2017.
Genotypic/phenotypic resistance correlations.
cgMLST-based phylogenetic relationships among S. aureus causing keratitis.
Tn554.
ASM does not own the copyrights to Supplemental Material that may be linked to, or accessed through, an article. The authors have granted ASM a non-exclusive, world-wide license to publish the Supplemental Material files. Please contact the corresponding author directly for reuse.
REFERENCES
- 1. Bourne RRA, Jonas JB, Bron AM, Cicinelli MV, Das A, Flaxman SR, Friedman DS, Keeffe JE, Kempen JH, Leasher J, Limburg H, Naidoo K, Pesudovs K, Peto T, Saadine J, Silvester AJ, Tahhan N, Taylor HR, Varma R, Wong TY, Resnikoff S, Vision Loss Expert Group of the Global Burden of Disease Study . 2018. Prevalence and causes of vision loss in high-income countries and in Eastern and Central Europe in 2015: magnitude, temporal trends and projections. Br J Ophthalmol 102:575–585. doi: 10.1136/bjophthalmol-2017-311258 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Collier SA, Gronostaj MP, MacGurn AK, Cope JR, Awsumb KL, Yoder JS, Beach MJ, Centers for Disease Control and Prevention (CDC) . 2014. Estimated burden of keratitis — United States, 2010. MMWR Morb Mortal Wkly Rep 63:1027–1030. [PMC free article] [PubMed] [Google Scholar]
- 3. Ung L, Bispo PJM, Shanbhag SS, Gilmore MS, Chodosh J. 2019. The persistent dilemma of microbial keratitis: global burden, diagnosis, and antimicrobial resistance. Surv Ophthalmol 64:255–271. doi: 10.1016/j.survophthal.2018.12.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Zhang Z, Cao K, Liu J, Wei Z, Xu X, Liang Q. 2022. Pathogens and antibiotic susceptibilities of global bacterial keratitis: a meta-analysis. Antibiotics (Basel) 11:238. doi: 10.3390/antibiotics11020238 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Lin A, Rhee MK, Akpek EK, Amescua G, Farid M, Garcia-Ferrer FJ, Varu DM, Musch DC, Dunn SP, Mah FS, American Academy of Ophthalmology Preferred Practice Pattern Cornea and External Disease Panel . 2019. Bacterial keratitis preferred practice pattern. Ophthalmology 126:P1–P55. doi: 10.1016/j.ophtha.2018.10.018 [DOI] [PubMed] [Google Scholar]
- 6. Tan SZ, Walkden A, Au L, Fullwood C, Hamilton A, Qamruddin A, Armstrong M, Brahma AK, Carley F. 2017. Twelve-year analysis of microbial keratitis trends at a UK tertiary hospital. Eye (Lond) 31:1229–1236. doi: 10.1038/eye.2017.55 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. WHO . 2015. Global action plan on antimicrobial resistance. World Health Organization, Geneva. [DOI] [PubMed] [Google Scholar]
- 8. Asbell PA, Sanfilippo CM, Sahm DF, DeCory HH. 2020. Trends in antibiotic resistance among ocular microorganisms in the United States from 2009 to 2018. JAMA Ophthalmol 138:439–450. doi: 10.1001/jamaophthalmol.2020.0155 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Bispo PJM, Sahm DF, Asbell PA. 2022. A systematic review of multi-decade antibiotic resistance data for ocular bacterial pathogens in the United States. Ophthalmol Ther 11:503–520. doi: 10.1007/s40123-021-00449-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Thomas RK, Melton R, Asbell PA. 2019. Antibiotic resistance among ocular pathogens: current trends from the ARMOR surveillance study (2009-2016). Clin Optom (Auckl) 11:15–26. doi: 10.2147/OPTO.S189115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Harris PA, Taylor R, Thielke R, Payne J, Gonzalez N, Conde JG. 2009. Research electronic data capture (REDCap) - A metadata-driven methodology and workflow process for providing translational research informatics support. J Biomed Inform 42:377–381. doi: 10.1016/j.jbi.2008.08.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Clinical and Laboratory Standards Institute . 2018. Approved standard M07-A11. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically. 10th ed. CLSI, Wayne, PA, USA. [Google Scholar]
- 13. Clinical and Laboratory Standards Institute . 2018. CLSI supplement M100. Performance standards for antimicrobial susceptibility testing. 28th ed. CLSI, Wayne, PA. [Google Scholar]
- 14. Gladstone RA, Lo SW, Goater R, Yeats C, Taylor B, Hadfield J, Lees JA, Croucher NJ, van Tonder AJ, Bentley LJ, et al. 2020. Visualizing variation within global pneumococcal sequence clusters (GPSCs) and country population snapshots to contextualize pneumococcal isolates. Microb Genom 6:e000357. doi: 10.1099/mgen.0.000357 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Treangen TJ, Ondov BD, Koren S, Phillippy AM. 2014. The Harvest suite for rapid core-genome alignment and visualization of thousands of intraspecific microbial genomes. Genome Biol 15:524. doi: 10.1186/s13059-014-0524-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Croucher NJ, Page AJ, Connor TR, Delaney AJ, Keane JA, Bentley SD, Parkhill J, Harris SR. 2015. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res 43:e15. doi: 10.1093/nar/gku1196 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Stamatakis A. 2014. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. doi: 10.1093/bioinformatics/btu033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Letunic I, Bork P. 2019. Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47:W256–W259. doi: 10.1093/nar/gkz239 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Uhlemann A-C, Otto M, Lowy FD, DeLeo FR. 2014. Evolution of community- and healthcare-associated methicillin-resistant Staphylococcus aureus. Infect Genet Evol 21:563–574. doi: 10.1016/j.meegid.2013.04.030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Bispo PJM, Alfonso EC, Flynn HW, Miller D. 2013. Emerging 8-methoxyfluoroquinolone resistance among methicillin-susceptible Staphylococcus epidermidis isolates recovered from patients with endophthalmitis. J Clin Microbiol 51:2959–2963. doi: 10.1128/JCM.00846-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Mendes RE, Deshpande LM, Costello AJ, Farrell DJ. 2012. Molecular epidemiology of Staphylococcus epidermidis clinical isolates from U.S. hospitals. Antimicrob Agents Chemother 56:4656–4661. doi: 10.1128/AAC.00279-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Ibrahem S, Salmenlinna S, Lyytikäinen O, Vaara M, Vuopio-Varkila J. 2008. Molecular characterization of methicillin-resistant Staphylococcus epidermidis strains from bacteraemic patients. Clin Microbiol Infect 14:1020–1027. doi: 10.1111/j.1469-0691.2008.02080.x [DOI] [PubMed] [Google Scholar]
- 23. Côrtes MF, André C, Martins Simões P, Corvec S, Caillon J, Tristan A, Bes M, Vandenesch F, Figueiredo AMS, Dupieux C, Laurent F. 2022. Persistence of a multidrug-resistant worldwide-disseminated methicillin-resistant Staphylococcus epidermidis clone harbouring the cfr linezolid resistance gene in a French hospital with evidence of interspecies transfer to several Staphylococcus aureus lineages. J Antimicrob Chemother 77:1838–1846. doi: 10.1093/jac/dkac119 [DOI] [PubMed] [Google Scholar]
- 24. Murphy E, Huwyler L, de Freire Bastos M do C. 1985. Transposon Tn554: complete nucleotide sequence and isolation of transposition-defective and antibiotic-sensitive mutants. EMBO J 4:3357–3365. doi: 10.1002/j.1460-2075.1985.tb04089.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Andre C, Rouhana J, Scarpa de Mello S, Rosa da Cunha G, Van Camp AG, Gilmore MS, Bispo PJM. 2022. Population structure of ocular Streptococcus pneumoniae is highly diverse and formed by lineages that escape current vaccines. Microb Genom 8:000763. doi: 10.1099/mgen.0.000763 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Gill MJ, Brenwald NP, Wise R. 1999. Identification of an efflux pump gene, pmrA, associated with fluoroquinolone resistance in Streptococcus pneumoniae. Antimicrob Agents Chemother 43:187–189. doi: 10.1128/AAC.43.1.187 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Boncoeur E, Durmort C, Bernay B, Ebel C, Di Guilmi AM, Croizé J, Vernet T, Jault J-M. 2012. PatA and PatB form a functional heterodimeric ABC multidrug efflux transporter responsible for the resistance of Streptococcus pneumoniae to fluoroquinolones. Biochemistry 51:7755–7765. doi: 10.1021/bi300762p [DOI] [PubMed] [Google Scholar]
- 28. Peng MY, Cevallos V, McLeod SD, Lietman TM, Rose-Nussbaumer J. 2018. Bacterial keratitis: isolated organisms and antibiotic resistance patterns in San Francisco. Cornea 37:84–87. doi: 10.1097/ICO.0000000000001417 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Kowalski RP, Nayyar SV, Romanowski EG, Shanks RMQ, Mammen A, Dhaliwal DK, Jhanji V. 2020. The prevalence of bacteria, fungi, viruses, and acanthamoeba from 3,004 cases of keratitis, endophthalmitis, and conjunctivitis. Eye Contact Lens 46:265–268. doi: 10.1097/ICL.0000000000000642 [DOI] [PubMed] [Google Scholar]
- 30. Chang VS, Dhaliwal DK, Raju L, Kowalski RP. 2015. Antibiotic resistance in the treatment of Staphylococcus aureus keratitis: a 20-year review. Cornea 34:698–703. doi: 10.1097/ICO.0000000000000431 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Geevarghese A, Shah P, Lopez J, Tsui E, Raju L. 2022. Common microbes and antibiotic resistance in ocular infections at an urban public tertiary care hospital. Ocul Immunol Inflamm 30:481–486. doi: 10.1080/09273948.2020.1799033 [DOI] [PubMed] [Google Scholar]
- 32. Lalitha P, Manoharan G, Karpagam R, Prajna NV, Srinivasan M, Mascarenhas J, Das M, Porco TC, Lietman TM, Cevallos V, Keenan JD. 2017. Trends in antibiotic resistance in bacterial keratitis isolates from South India. Br J Ophthalmol 101:108–113. doi: 10.1136/bjophthalmol-2016-308487 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Leck AK, Thomas PA, Hagan M, Kaliamurthy J, Ackuaku E, John M, Newman MJ, Codjoe FS, Opintan JA, Kalavathy CM, Essuman V, Jesudasan CAN, Johnson GJ. 2002. Aetiology of suppurative corneal ulcers in Ghana and South India, and epidemiology of fungal keratitis. Br J Ophthalmol 86:1211–1215. doi: 10.1136/bjo.86.11.1211 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. McDougal LK, Steward CD, Killgore GE, Chaitram JM, McAllister SK, Tenover FC. 2003. Pulsed-field gel electrophoresis typing of oxacillin-resistant Staphylococcus aureus isolates from the United States: establishing a national database. J Clin Microbiol 41:5113–5120. doi: 10.1128/JCM.41.11.5113-5120.2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Bispo PJM, Ung L, Chodosh J, Gilmore MS. 2020. Hospital-associated multidrug-resistant MRSA lineages are trophic to the ocular surface and cause severe microbial keratitis. Front Public Health 8:204. doi: 10.3389/fpubh.2020.00204 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Hudson LO, Reynolds C, Spratt BG, Enright MC, Quan V, Kim D, Hannah P, Mikhail L, Alexander R, Moore DF, Godoy D, Bishop CJ, Huang SS. 2013. Diversity of methicillin-resistant Staphylococcus aureus strains isolated from residents of 26 nursing homes in Orange County, California. J Clin Microbiol 51:3788–3795. doi: 10.1128/JCM.01708-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Tenover FC, McAllister S, Fosheim G, McDougal LK, Carey RB, Limbago B, Lonsway D, Patel JB, Kuehnert MJ, Gorwitz R. 2008. Characterization of Staphylococcus aureus isolates from nasal cultures collected from individuals in the United States in 2001 to 2004. J Clin Microbiol 46:2837–2841. doi: 10.1128/JCM.00480-08 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Velasco V, Buyukcangaz E, Sherwood JS, Stepan RM, Koslofsky RJ, Logue CM. 2015. Characterization of Staphylococcus aureus from humans and a comparison with isolates of animal origin, in North Dakota, United States. PLoS One 10:e0140497. doi: 10.1371/journal.pone.0140497 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. von Eiff C, Becker K, Machka K, Stammer H, Peters G. 2001. Nasal carriage as a source of Staphylococcus aureus bacteremia. Study group. N Engl J Med 344:11–16. doi: 10.1056/NEJM200101043440102 [DOI] [PubMed] [Google Scholar]
- 40. Clarridge JE, Harrington AT, Roberts MC, Soge OO, Maquelin K. 2013. Impact of strain typing methods on assessment of relationship between paired nares and wound isolates of methicillin-resistant Staphylococcus aureus. J Clin Microbiol 51:224–231. doi: 10.1128/JCM.02423-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Bispo PJM, Hofling-Lima AL, Pignatari ACC, Patel R. 2014. Characterization of ocular methicillin-resistant Staphylococcus epidermidis isolates belonging predominantly to clonal complex 2 subcluster II. J Clin Microbiol 52:1412–1417. doi: 10.1128/JCM.03098-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Li M, Wang X, Gao Q, Lu Y. 2009. Molecular characterization of Staphylococcus epidermidis strains isolated from a teaching hospital in Shanghai, China. J Med Microbiol 58:456–461. doi: 10.1099/jmm.0.007567-0 [DOI] [PubMed] [Google Scholar]
- 43. Schoenfelder SMK, Lange C, Eckart M, Hennig S, Kozytska S, Ziebuhr W. 2010. Success through diversity - how Staphylococcus epidermidis establishes as a nosocomial pathogen. Int J Med Microbiol 300:380–386. doi: 10.1016/j.ijmm.2010.04.011 [DOI] [PubMed] [Google Scholar]
- 44. Beall B, Chochua S, Gertz RE, Li Y, Li Z, McGee L, Metcalf BJ, Ricaldi J, Tran T, Walker H, Pilishvili T. 2018. A population-based descriptive atlas of invasive pneumococcal strains recovered within the U.S. During 2015–2016. Front Microbiol 9:2670. doi: 10.3389/fmicb.2018.02670 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Desai AP, Sharma D, Crispell EK, Baughman W, Thomas S, Tunali A, Sherwood L, Zmitrovich A, Jerris R, Satola SW, Beall B, Moore MR, Jain S, Farley MM. 2015. Decline in pneumococcal nasopharyngeal carriage of vaccine serotypes after the introduction of the 13-valent pneumococcal conjugate vaccine in children in Atlanta, Georgia. Pediatr Infect Dis J 34:1168–1174. doi: 10.1097/INF.0000000000000849 [DOI] [PubMed] [Google Scholar]
- 46. Valentino MD, McGuire AM, Rosch JW, Bispo PJM, Burnham C, Sanfilippo CM, Carter RA, Zegans ME, Beall B, Earl AM, Tuomanen EI, Morris TW, Haas W, Gilmore MS. 2014. Unencapsulated Streptococcus pneumoniae from conjunctivitis encode variant traits and belong to a distinct phylogenetic cluster. Nat Commun 5:5411. doi: 10.1038/ncomms6411 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Bertino JS. 2009. Impact of antibiotic resistance in the management of ocular infections: the role of current and future antibiotics. Clin Ophthalmol 3:507–521. doi: 10.2147/opth.s5778 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Almagor J, Temkin E, Benenson I, Fallach N, Carmeli Y, DRIVE-AB consortium . 2018. The impact of antibiotic use on transmission of resistant bacteria in hospitals: insights from an agent-based model. PLoS One 13:e0197111. doi: 10.1371/journal.pone.0197111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Aires de Sousa M, de Lencastre H, Santos Sanches I, Kikuchi K, Totsuka K, Tomasz A. 2000. Similarity of antibiotic resistance patterns and molecular typing properties of methicillin-resistant Staphylococcus aureus isolates widely spread in hospitals in New York City and in a hospital in Tokyo, Japan. Microb Drug Resist 6:253–258. doi: 10.1089/mdr.2000.6.253 [DOI] [PubMed] [Google Scholar]
- 50. Lee JYH, Monk IR, Gonçalves da Silva A, Seemann T, Chua KYL, Kearns A, Hill R, Woodford N, Bartels MD, Strommenger B, Laurent F, Dodémont M, Deplano A, Patel R, Larsen AR, Korman TM, Stinear TP, Howden BP. 2018. Global spread of three multidrug-resistant lineages of Staphylococcus epidermidis. Nat Microbiol 3:1175–1185. doi: 10.1038/s41564-018-0230-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Niederman MS. 2015. Macrolide-resistant pneumococcus in community-acquired pneumonia. Is there still a role for macrolide therapy? Am J Respir Crit Care Med 191:1216–1217. doi: 10.1164/rccm.201504-0701ED [DOI] [PubMed] [Google Scholar]
- 52. Wilson DN. 2014. Ribosome-targeting antibiotics and mechanisms of bacterial resistance. Nat Rev Microbiol 12:35–48. doi: 10.1038/nrmicro3155 [DOI] [PubMed] [Google Scholar]
- 53. Roberts MC, Sutcliffe J, Courvalin P, Jensen LB, Rood J, Seppala H. 1999. Nomenclature for macrolide and macrolide-lincosamide-streptogramin B resistance determinants. Antimicrob Agents Chemother 43:2823–2830. doi: 10.1128/AAC.43.12.2823 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Weisblum B. 1995. Erythromycin resistance by ribosome modification. Antimicrob Agents Chemother 39:577–585. doi: 10.1128/AAC.39.3.577 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Firth N, Jensen SO, Kwong SM, Skurray RA, Ramsay JP. 2019. Staphylococcal plasmids, transposable and integrative elements, p 499–520. In Fischetti VA, Novick RP, Ferretti JJ, Portnoy DA, Braunstein M, Rood J (ed), Gram-positive pathogens, 3rd ed. American Society for Microbiology, Washington, DC. [Google Scholar]
- 56. Vestergaard M, Frees D, Ingmer H. 2019. Antibiotic resistance and the MRSA problem, p 747–765. In Fischetti VA, Novick RP, Ferretti JJ, Portnoy DA, Braunstein M, Rood J (ed), Gram-positive pathogens. American Society for Microbiology, Washington, DC, USA. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Kos VN, Desjardins CA, Griggs A, Cerqueira G, Van Tonder A, Holden MTG, Godfrey P, Palmer KL, Bodi K, Mongodin EF, Wortman J, Feldgarden M, Lawley T, Gill SR, Haas BJ, Birren B, Gilmore MS. 2012. Comparative Genomics of vancomycin-resistant Staphylococcus aureus strains and their positions within the clade most commonly associated with Methicillin-resistant S. aureus hospital-acquired infection in the United States. mBio 3:e00112-12. doi: 10.1128/mBio.00112-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Yee R, Dien Bard J, Simner PJ. 2021. The genotype-to-phenotype dilemma: how should laboratories approach discordant susceptibility results? J Clin Microbiol 59:e00138-20. doi: 10.1128/JCM.00138-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Hooper DC, Jacoby GA. 2015. Mechanisms of drug resistance: quinolone resistance. Ann N Y Acad Sci 1354:12–31. doi: 10.1111/nyas.12830 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Ferrero L, Cameron B, Manse B, Lagneaux D, Crouzet J, Famechon A, Blanche F. 1994. Cloning and primary structure of Staphylococcus aureus DNA topoisomerase IV: a primary target of fluoroquinolones. Mol Microbiol 13:641–653. doi: 10.1111/j.1365-2958.1994.tb00458.x [DOI] [PubMed] [Google Scholar]
- 61. Fournier B, Truong-Bolduc QC, Zhang X, Hooper DC. 2001. A mutation in the 5’ untranslated region increases stability of norA mRNA, encoding a multidrug resistance transporter of Staphylococcus aureus. J Bacteriol 183:2367–2371. doi: 10.1128/JB.183.7.2367-2371.2001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Yu J-L, Grinius L, Hooper DC. 2002. NorA functions as a multidrug efflux protein in both cytoplasmic membrane vesicles and reconstituted proteoliposomes. J Bacteriol 184:1370–1377. doi: 10.1128/JB.184.5.1370-1377.2002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Truong-Bolduc QC, Strahilevitz J, Hooper DC. 2006. NorC, a new efflux pump regulated by MgrA of Staphylococcus aureus. Antimicrob Agents Chemother 50:1104–1107. doi: 10.1128/AAC.50.3.1104-1107.2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Sanfilippo CM, Hesje CK, Haas W, Morris TW. 2011. Topoisomerase mutations that are associated with high-level resistance to earlier fluoroquinolones in Staphylococcus aureus have less effect on the antibacterial activity of besifloxacin. Chemotherapy 57:363–371. doi: 10.1159/000330858 [DOI] [PubMed] [Google Scholar]
- 65. Shinabarger DL, Zurenko GE, Hesje CK, Sanfilippo CM, Morris TW, Haas W. 2011. Evaluation of the effect of bacterial efflux pumps on the antibacterial activity of the novel fluoroquinolone besifloxacin. J Chemother 23:80–86. doi: 10.1179/joc.2011.23.2.80 [DOI] [PubMed] [Google Scholar]
- 66. O’Brien TP. 2012. Besifloxacin ophthalmic suspension, 0.6%: a novel topical fluoroquinolone for bacterial conjunctivitis. Adv Ther 29:473–490. doi: 10.1007/s12325-012-0027-7 [DOI] [PubMed] [Google Scholar]
- 67. Cohen ML. 1992. Epidemiology of drug resistance: implications for a post-antimicrobial era. Science 257:1050–1055. doi: 10.1126/science.257.5073.1050 [DOI] [PubMed] [Google Scholar]
- 68. McDonald M, Blondeau JM. 2010. Emerging antibiotic resistance in ocular infections and the role of fluoroquinolones. J Cataract Refract Surg 36:1588–1598. doi: 10.1016/j.jcrs.2010.06.028 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Demographic and clinicopathologic data for patients with bacterial keratitis at MEE, 2014-2017.
Genotypic/phenotypic resistance correlations.
cgMLST-based phylogenetic relationships among S. aureus causing keratitis.
Tn554.