Abstract
The Macavirus, ovine gammaherpesvirus 2 (OvGHV2), is the cause of sheep-associated malignant catarrhal fever (SA-MCF). Although SA-MCF occurs in a wide range of mammalian hosts, there are few descriptions of this disease and/or infection in goats. This report describes the findings observed in a goat that was infected by OvGHV2 and adds to the rare description of this infection in this animal species. A 6.5-year-old, female, Anglo Nubian goat, with a neurological syndrome, that was euthanized after severe esophageal obstruction was investigated to determine the cause of the brain disease. Histopathology revealed cerebral cortical edema, hemorrhagic rhombencephalitis, severe hepatic necrosis, and atrophic enteritis. An immunohistochemical (IHC) assay identified intracytoplasmic antigens of a malignant catarrhal fever virus (MCFV) within epithelial cells of the intestine, liver, lungs, and kidneys. A semi-nested PCR assay amplified the partial fragment of the OvGHV2 tegument protein gene from the intestine, confirming that the MCFV identified by IHC was OvGHV2. A qPCR assay that targeted the OvGHV2 polymerase gene revealed an elevated quantification cycle (Cq), while nanoplate-based digital PCR (dPCR) detected low viral copy load within the OvGHV2 DNA. Furthermore, the nucleic acids of several disease pathogens associated with diseases in ruminants were not amplified. However, the exact cause of the neurological syndrome remained obscure since nucleic acids of neurological disease pathogens such as bovine viral diarrhea virus, bovine alphaherpesvirus 1 and 5, Histophilus somni, and OvGHV2 were not detected from the brain. Collectively, the results of the Cq and dPCR confirmed that this goat was infected with a low viral load of OvGHV2, which probably was insufficient to induce the typical histopathological alterations and subsequent clinical manifestations associated with SA-MCF and/or infections by OvGHV2. Therefore, elevated viral loads of OvGHV2 would have been required for the development of histological lesions and/or clinical manifestations of SA-MCF in this goat. Furthermore, the dPCR methodology can be used for the efficient detection and quantification of OvGHV2 DNA in animals with or without clinical and/or histopathological evidence of SA-MCF. Additionally, since previous cases of OvGHV2 infections in goats did not have the typical clinical manifestations of SA-MCF, one wonders if this Macavirus can induce SA-MCF in goats.
Supplementary Information
The online version contains supplementary material available at 10.1007/s42770-023-01170-4.
Keywords: dPCR, Macavirus, qPCR, Pathogenesis, Sheep-associated malignant catarrhal fever, Viral load
Introduction
Malignant catarrhal fever (MCF) is a severe lymphotropic vascular disease caused by distinct members of the Macavirus genus, subfamily Gammaherpesvirinae, family Herpesviridae [1], that are collectively referred to as the MCF virus (MCFV) group within this genus [2]. The main agents of epidemiological importance associated with the development of MCF in distinct mammalian hosts are ovine gammaherpesvirus 2 (OvGHV2) and alcelaphine gammaherpesvirus 1 and 2 (AlGHV1 and AlGHV2), with sheep and wildebeest serving as reservoir hosts, respectively [3–5].
All members of the MCFV are considered unique as they share the 15A epitope [6] and have similar genetic and antigenic characteristics [7]. Furthermore, the 15A monoclonal antibody (15A-MAb) was initially used in serological assays associated with molecular investigations for the detection of OvGHV2 [8–10]. We later demonstrated that the 15A-MAb can be successfully used for the immunohistochemical (IHC) detection of OvGHV2 tissues antigens in cattle with sheep associated-MCF (SA-MCF) [11] and have since used this antibody in studies associated with the molecular amplification of OvGHV2 [12–15].
While goats are considered the natural reservoirs for caprine gammaherpesvirus (CpGHV2) and responsible for the development of MCF in specific cervids [3, 16, 17], there are few reports of infections by OvGHV2 in this ruminant species. Three cases were described in goats with MCF-like lesions, where infection was confirmed by the molecular identification of OvGHV2 DNA [18], and there are the descriptions of cutaneous alterations [19, 20] and multisystemic necrotizing vasculitis [21] associated with infections by OvGHV2 in goats. Although the pathogenesis associated with infections by OvGHV2 in cattle and sheep is well established, it remains uncertain whether SA-MCF in goats requires an elevated viral load for the demonstration of clinical disease or if there is a direct relationship between the load of OvGHV2 DNA and the development of lesions in SA-MCF [4]. This report investigated the possible relationship between viral load and the development of histopathologic alterations in a goat that was naturally infected by OvGHV2 and provides additional evidence of the unusual manifestation of OvGHV2 in this ruminant species.
Materials and methods
Clinical history, study location, and post-mortem evaluations
A 6.5-year-old, female, Anglo Nubian goat was admitted to the Large Animal Clinics, Veterinary Teaching Hospital, Universidade Estadual de Londrina (VTH-UEL) with a complaint of apathy, hyporexia, and permanent recumbency. The animal was dewormed one month prior to admission and had given birth to two kids by normal parturition two months before being admitted at the VTH-UEL. On the day before arrival at the VTH-UEL, the goat was reportedly found recumbent, lethargic, and hyperthermic and was medicated by a consulting veterinarian with florfenicol, flunixin meglumine, and a commercial solution containing calcium borogluconate, phosphorus, magnesium, and dextrose (20 mL). However, the animal was irresponsive to therapy.
This goat was from a small farm located on the outskirt of Londrina, Southern Brazil. Goats at this farm were maintained in enclosed pens containing adult female goats (n = 12) and several kids (n = 6). This farm also contained sheep (n = 22), reared approximately 200 m distant from the goats. All small ruminants at this farm were maintained on corn silage, hay and were supplemented by a commercial mineral salt; water was provided ad libitum.
The animal was examined clinically on admission at the VTH-UEL, blood and serum samples were collected for routine biochemical, hematological, and hemogasometric evaluations, while fecal samples were submitted for parasitological analysis.
The goat was initially placed on intravenous fluid therapy to correct dehydration and medicated with moxidectin (0.4 mg/kg, SC, single dose), toltrazuril (20 mg/kg, PO, single dose), and flunixin meglumine (2.2 mg/kg, IV, once a day for 3 days). However, one day after admission, the general clinical condition of the goat deteriorated, and the animal was found in permanent lateral recumbency, with hypothermia (35.7 °C), nystagmus, and vocalization. The animal was then warmed adequately, received a whole blood transfusion (350 mL in 4 h), and was placed on fluid therapy with glucose. Thiamine (20 mg/kg/IM/twice a day for 2 days) and dexamethasone (0.5 mg/kg/IV/SID for 4 days) were then included in the therapy, since polioencephalomalacia was suspected.
In the fifth day after admission, the goat had an episode of intense sialorrhea, mandibular trismus, and cough. Oral-ruminal probing was attempted but was unsuccessful since the probe was unable to pass the beginning of the thorax. A radiographic evaluation revealed cranial distension of the esophagus by gas and caudal obstruction by ingesta (Supplemental Fig. 1A). The esophageal distention extended to the thoracic esophagus, revealing an esophageal obstruction, probably due to food content. Due to the worsening of the clinical condition and low commercial value of the animal, the owner opted for euthanasia.
The carcass was then submitted for routine post-mortem evaluations soon after death to determine the possible cause of the neurological syndrome. Selected tissues of major organs were collected, routinely processed for possible histological alterations, and evaluated with the Hematoxylin and eosin stain. Specific formalin-fixed paraffin embedded (FFPE) tissue sections were used in IHC assays. Duplicate sections of specific tissues were collected and maintained at -80℃ until used in molecular assays.
Immunohistochemical detection of MCFV antigens
FFPE tissue sections of the small intestine, lungs, kidneys, and liver were used in IHC assays designed to identify intralesional antigens of MCFV by using the MAb-15A as previously described [11]. Positive controls consisted of the utilization of FFPE tissue sections of the small intestine derived from a cow known to contain intralesional antigens of OvGHV2 [12]. Negative controls consisted of the substitution of the MAb-15A with its diluent and the utilization of FFPE tissue sections known not to contain intralesional antigens of OvGHV2. Positive and negative controls were included in all IHC assays.
Molecular investigation of the principal infectious disease agents of ruminants
Nucleic acids from selected tissue fragments (cerebrum, cerebellum, liver, spleen, small intestine, and lungs) collected post-mortem were extracted as described [22, 23]. Molecular assays were done to amplify the nucleic acids of specific infectious disease agents associated with the development of respiratory, alimentary, and systemic diseases of ruminants. These molecular assays targeted specific genes of viral and bacterial agents, based on previously described protocols (Supplemental Table 1). These included OvGHV2, bovine gammaherpesvirus 6 (BoGHV6), bovine alphaherpesvirus 1 and 5 (BoAHV1 and BoAHV5), bovine viral diarrhea virus (BVDV), bovine parainfluenza virus 3 (BPIV3), bovine respiratory syncytial virus (BRSV), bovine coronavirus (BCoV), Pasteurella multocida, Histophilus somni, Mannheimia haemolytica, and Mycoplasma bovis. Positive controls consisted of the utilization of DNA/RNA from previous investigations [15, 24]; sterile, ultrapure water served as the negative control. Positive and negative controls were included in all nucleic molecular assays.
Sequencing and phylogenetic analysis of the OvGHV2 tegument protein gene
The products derived from the semi-nested OvGHV2 PCR using the Baxter primers [25] were purified (Quick Gel Extraction and PCR Purification Combo Kit; Invitrogen Life Technologies, Carlsbad, CA, USA), quantified (Qubit® Fluorometer; Invitrogen Life Technologies, Eugene, OR, USA), and sequenced in both directions with the forward and reverse primers in an ABI3500 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA).
Sequence quality analyses and consensus sequences were obtained using PHRED and CAP3 software (http://asparagin.cenargen.embrapa.br/phph), respectively. Similar nucleotides (nt) of the OvGHV2 tegument protein gene identified during this study were compared with those deposited in GenBank using the BLASTn software (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Multiple and pairwise alignments of the deduced amino acids (aa) sequences were compared with the aa reference strain of OvGHV2 and with OvGHV2 field strains available at GenBank and were performed with the MEGA software [26].
The aa identity matrix was performed using the BioEdit software [27]. The phylogenetic analysis of the OvGHV2 strain from this study was obtained using the Maximum Likelihood method based on the JTT matrix-based model [28] with the MEGA7 software [26] and compared with the reference strain of OvGHV2, an AlGHV1 strain, and OvGHV2 field strains derived from several animals in different countries.
qPCR for OvGHV2
For the OvGHV2 qPCR assay, 3 μL of the DNA samples derived from the Baxter semi-nested PCR assay [25] was added to 5 μL of TaqMan Fast Advanced Master Mix (Life Technologies, Carlsbad, CA, USA), 0.1 µM of primer forward and reverse primers, and 0.2 µM of the hydrolysis probe (Table 1). For each run, ultrapure DNase–RNase-free distilled water (ThermoFisher Scientific, Waltham, MA, USA) was included as a No Template Control (NTC), and a confirmed pool of positive samples was included as positive control (PC). The assay was specific for an 80 bp sequence targeting the OvGHV2 DNA polymerase gene [29]. The cycling conditions consisted of 50 °C for 2 min, 95 °C for 2 min, followed by 45 cycles of denaturation at 95 °C for 3 s, and annealing, amplification, and detection at 55 °C for 30 s. Samples were considered negative in the absence of amplification. The reactions were done in a 7500 Fast Real-time PCR System (Applied Biosystem, San Diego, CA, USA) in 96 well-plates (ThermoFisher Scientific, Waltham, MA, USA).
Table 1.
Primers, probe, and synthetic DNA molecule sequences used in the real-time PCR for OvGHV2
| Primer name | DNA sequence (5′-3′) probe labels | Amplicon size (bp) | References |
|---|---|---|---|
| dpol1771-F | CAC ACC CAA CTG GAG TAT GAC | 80 | Cunha et al., 2009 |
| dpol1831-R | ATG TTG TAG TGG GGC CAG TC | ||
| OvGHV2 probe | FAM- ATG TGC GCT TCG ACC CTC -BHQ1 | ||
| OvGHV2 synthetic DNA | GTACACGTGCAGCAACTTTTCTGCCCGAGCGAGTGACCACCGCGACTCCCACACGCAACTGGAGTACGACTGTGCCGTGGGTGATGTGCGCTTCGACCCTCACCGCCTGGACTGGCCCCACTACAACATCCTATCCTTTGATATAGAGTGCCTGGGCGAGTGCGGCTTCCCCACGGCCGAT | 181 | This study |
To verify the assay efficiency of the field samples and synthetic DNA, a standard curve was constructed using tenfold dilutions of a DNA pool consisted of field samples derived from the brains of cattle known to contain OvGHV2 and gBlocks® gene fragments of OvGHV2 (Integrated DNA Technologies, Coralville, IA, USA). A linear regression analysis was performed by plotting the quantification cycle (Cq) values against the logarithm of the copy number of the gene targeted.
Nanoplate-based digital PCR (dPCR) quantification of OvGHV2 DNA
The dPCR assays were carried out in single-plicate with the same primers and probe sequences used in the qPCR assay (Table 1), with 10 μL of DNA, added to 10 μL of Probe PCR Master Mix (Qiagen, Hilden, Germany) and 26,000 24-well Nanoplates (Qiagen, Hilden, Germany), to the 0.8 µM forward, 0.8 µM reverse primers and 0.4 µM probe, in a final volume of 40 μL. The nanoplate was then loaded onto the QIAcuity One dPCR (Qiagen, Hilden, Germany) and subjected to an automated workflow, cycling protocol (95 °C for 2 min for enzyme activation, 95 °C for 15 s for denaturation, and 60 °C for 30 s for annealing/extension for 40 cycles), and a final imaging step made by reading in the FAM channel. For each run, ultrapure DNase–RNase-free distilled water (ThermoFisher Scientific, Waltham, MA, USA) was included as a No Template Control (NTC). Samples were considered negative when there was no amplification in any of the partitions. All thresholds were set up manually to allow discrimination between positive and negative. Absolute quantification of PCR targets was performed using the QIAcuity Software Suite 2.1.7.182 and expressed as copy per μL.
Viral load quantification by dPCR
The absolute number of viral DNA sequences was determined by dPCR using the QIAcuity One dPCR platform (Qiagen, Hilden, Germany) with field samples and synthetic OvGHV2 DNA. Absorbance measurements at 260 nm by UV spectrophotometry in a NanoDrop ND-ONE-W spectrophotometer (ThermoScientific, Waltham, MA, USA) and fluorescence measure with PicoGreen dye in a Qubit 3.0 fluorometer (Life Technologies, Carlsbad, CA, USA) were used according to the manufacturer’s protocols to quantify DNA and the amounts of synthetic DNA molecule gBlocks® gene fragments (Integrated DNA Technologies, Coralville, IA, USA) of the tenfold serial dilutions increments (Pure to 10−6).
Results
Clinical and laboratory findings
The general physical examination of the goat on arrival at the HV-UEL revealed hypothermia (32.9 °C), moderate dehydration (8%), ruminal atony, bilateral mucous nasal discharge, pendulous abdomen, pale mucous membranes, and submandibular edema. Additionally, depressed mental status, lack of response to the threat reflex, decreased pupillary reflex, and permanent sternal recumbency were observed during the neurological evaluation.
The hematological analysis revealed severe anemia (red blood cells 3.03/mm3; hemoglobin: 4.2 g/dL, hematocrit: 14%), mild leukocytosis (13,000/mm3) with neutrophilia (10,270/mm3), and left shift (130/mm3). Serum biochemistry identified hypoproteinemia (total protein 5.6 g/dL), hypoalbuminemia (1.5 g/dL), and increased serum concentration of gamma-glutamyltransferase (140U/L). Hemogasometry revealed hyponatremia (140.6 mmol/L), hypokalemia (2.57 mmol/L), hyperchloremia (115 mmol/L) and hypocalcemia (ionized calcium 2.1 mg/dL). The coproparasitological examination diagnosed parasitism by Strongyloides spp. (3850 eggs per gram) and Eimeria sp. (4000 oocyst count per gram). Consequently, a laboratory diagnosis of verminosis and eimeriosis with nutritional deficiency was established.
Gross and histopathological findings
Grossly, the ocular and oral mucous membranes were pale with severe submandibular edema, mild hydrothorax, and ascites. Significant gross alterations were restricted to the alimentary, pulmonary, hepatobiliary, and cardiovascular systems, and to some extent, the brain. The esophagus was severely dilated and packed with large accumulations of partiality ruminated and undigested feedstuff, resulting in severe esophageal dilation and obstruction (Supplemental Fig. 1B-C). Additionally, significant alterations within the alimentary tract included moderate parasitic abomasitis due to Haemonchus contortus and mild parasitic jejunitis due to the cestode, Moniezia expansa. Pulmonary alterations were predominantly edema, with severe accumulation of a foamy fluid within the entire extension of the lumen of the trachea; the lungs were wet and heavy. The liver was slightly enlarged, very friable, with several pale areas at the capsular and section surfaces of the organ. Gelatinous atrophy of the pericardial fatty tissue was the most significant alteration to the cardiovascular system. Additionally, there was mild flattening of the sulci and narrowing of the gyri of the brain.
Histopathology revealed severe atrophic enteritis; severe, diffused pulmonary edema with patches of bronchointerstitial pneumonia; random, severe necrohemorrhagic hepatitis (Supplemental Fig. 1D), mild to moderate cerebral cortical edema with focal hemorrhagic rhombencephalitis (Supplemental Fig. 1E-F).
Immunohistochemical identification of MCFV antigens
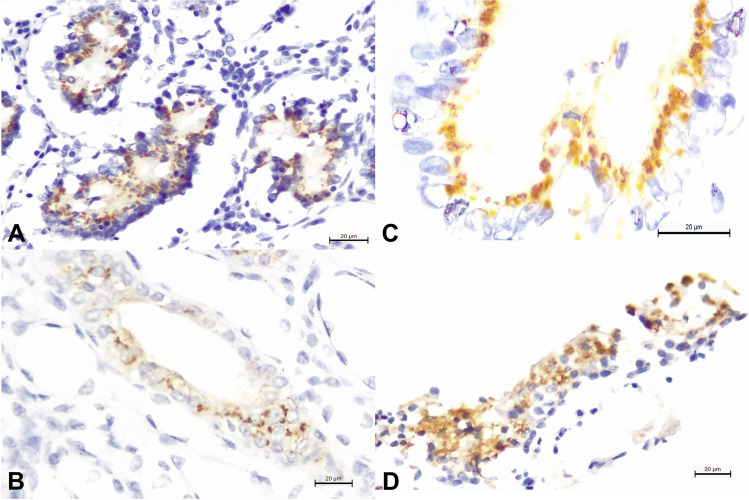
Positive intracytoplasmic immunoreactivity to the 15A-MAb was identified within epithelial cells of the lungs, small intestine, liver, and kidney (Fig. 1A–D), confirming the identification of intralesional antigens of a MCFV within these organs.
Fig. 1.
Immunohistochemical detection of the MAb-15A antigen in a goat infected with OvGHV2. There is positive, intracytoplasmic immunoreactivity of MCFV antigen within epithelial cells of the small intestine (A), kidneys (B), bile duct (C), and lungs (D). Immunoperoxidase counterstained with hematoxylin. Bar (A–D), 20 µm
Molecular characterization of infectious disease agents of ruminants
The Baxter primers [25] amplified the expected base pairs (bp) of the fragment of the OvGHV2 tegument protein gene from the small intestine; sequencing confirmed the PCR results. Furthermore, OvGHV2 DNA was not amplified from the other organs (lungs, liver, and kidney) with positive immunoreactivity for MCFV. The OvGHV2 strain identified in this goat is known as OvHV-2/BRA-UEL/PR-N56/2021 and is deposited in GenBank (Accession # OK490363). All other infectious disease pathogens evaluated were not amplified by their respective molecular assays from the target organs.
Phylogenetic genetic analysis of the partial fragment of the OvGHV2 tegument protein gene
The OvGHV2 strain identified in this investigation has 100% amino acid (aa) homology with the prototype strain of OvGHV2 (NC007646) and with OvGHV2 filed strains identified in cows from Brazil, India, Egypt, and Turkey, as well as with a sheep from India. Additionally, this strain herein described has 98.9–99.4% aa homology with several wild-type strains of OvGHV2 identified in mammalian species from various countries.
The generated phylogenetic analysis (Fig. 2) revealed that this strain herein identified clustered with the prototype strain for OvGHV2 and with similar field strains of OvGHV2 described in multiple animal species from several countries.
Fig. 2.
Phylogenetic analysis of the OvGHV2 strain identified in goat based on the tegument protein gene. The evolutionary history was inferred by using the Maximum Likelihood method based on the JTT matrix-based model. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) are shown next to the branches. The strain identified in this study is highlighted (
 ) and compared with similar strains derived from diverse animal species identified in several countries. Bovine alphaherpesvirus 1 was used as the outgroup
) and compared with similar strains derived from diverse animal species identified in several countries. Bovine alphaherpesvirus 1 was used as the outgroup
Standard Curve for OvGHV2 qPCR
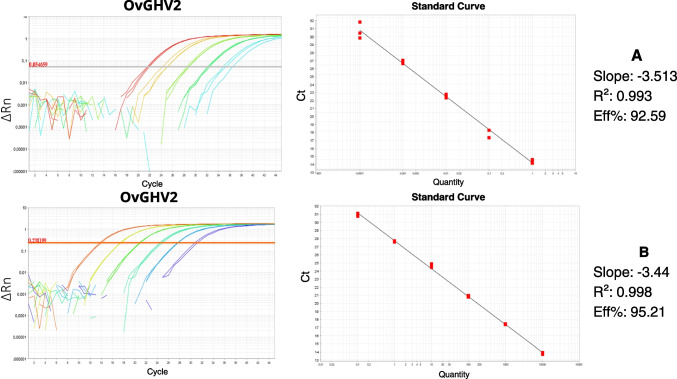
The optimized assay standard curve linearity for the pool of positive sample (Pure to 10−4) ran in triplicate for each dilution resulted in an amplification efficiency of 92.59%, correlation coefficient (R2) of 0.993, with a − 3.53 slope of (Fig. 3A). Furthermore, the standard curve for the OvGHV2 DNA molecule gBlocks® gene fragments (Integrated DNA Technologies, Coralville, IA, USA) ran in triplicate (pure to 10−5) presented an amplification efficiency of 95.214%, correlation coefficient (R2) of 0.998, and a slope of − 3.44 (Fig. 3B).
Fig. 3.
Efficiency and standard curves for qPCR of OvGHV2, based on tenfold serial dilutions (1 to 1 × 10−4) of a field sample (A) and tenfold serial dilutions (1 to 1 × 10−5) of gBlocks® (B)
Quantification of OvGHV2 DNA by dPCR
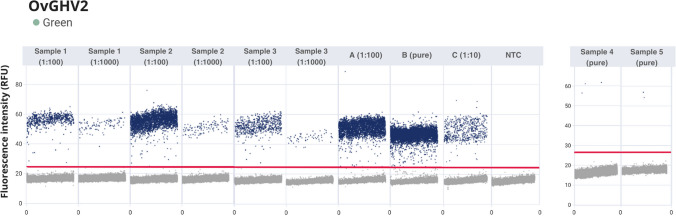
The results of the quantification cycle (Cq) for qPCR and dPCR quantification of copy number/μL for the pool of positive samples, synthetic OvGHV2 DNA, and field samples are provided in Table 2. Most samples evaluated had a Cq that was compatible with the number of viral copies identified by dPCR. The best results were obtained from DNA derived from the lungs of a cow with SA-MCF (sample #1; 14,400 copy/µL), as well as the spleen (sample #2; 75,200 copy/µL) and kidney (sample #3; 13,400 copy/µL) obtained from a cow with typical the histopathological lesions of SA-MCF (unpublished data). Furthermore, the results obtained in these three samples were consistent with those of the positive controls and the synthetic OvGHV2 DNA. However, DNA derived from the urine of an asymptomatic cow (sample #4), and the intestine of the goat from this study (sample #5), presented elevated Cq by qPCR and low viral copy numbers (1.28 copy/µL; and 0.84 copy/µL, respectively) by dPCR. A graphical representation of the quantification of OvGHV2 DNA from each sample as well as the negative control is provided in Fig. 4.
Table 2.
Quantification cycle (Cq) values for qPCR and number of copies for dPCR of OvGHV2 DNA derived from field samples, pool of positive samples, and OvGHV2 synthetic DNA
| Sample# | Sample origin | ||||
|---|---|---|---|---|---|
| qPCR | Digital droplet PCR | ||||
| Cq | Copy number/μL | Copy number/3 μL | CI (95%) | ||
| 1 | 20.12 | 14,400 | 43,200 | 7.5 | Lung, SA-MCF in cattle; Headley et al. 2012 |
| 2 | 19.05 | 75,200 | 225,600 | 3.4 | Spleen, cow with typical histological lesions of MCF; unpublished data |
| 3 | 21.94 | 13,444 | 40,332 | 7.9 | Kidney, cow with typical histological lesions of MCF; unpublished data |
| 4 | 38.45 | 1.28 | 3.84 | 106.4 | Urine, asymptomatic cow; unpublished data |
| 5 | 37.89 | 0.84 | 2.52 | 119.0 | Intestine, goat, this study |
| A | 23.55 | 81,800 | 245,400 | 2.6 | Pool of pure positive controls |
| B | 24.63 | 1422.8 | 4,268.4 | 2.6 | OvGHV2 synthetic DNA |
| C | 27.69 | 141.84 | 425.52 | 7.8 | OvGHV2 synthetic DNA; 1:10 |
Fig. 4.
One-dimensional scatterplots of digital PCR for absolute quantification of field sample, pool of positive controls, and OvGHV2 synthetic DNA. Legend: Fluorescence thresholds for the FAM channel for samples #1 (1:100); #1 (1:1000); # 2 (1:100); # 2 (1:1000); # 3 (1:100); # 3 (1:1000); A, Pool of pure positive controls, (1:100); B, OvGHV2 pure synthetic sample; C, OvGHV2 synthetic DNA, 1:10; and No Template Control (NTC) were 24.71. Fluorescence thresholds for the FAM channel for samples # 4 (pure) and #5 (pure) were 26.59
Discussion
Although this goat did not present clinical manifestations nor gross and histopathological findings suggestive of an infection by OvGHV2, namely, vasculitis and/or proliferative vascular lesions [3–5, 13, 14, 30], intralesional antigens of MCFV were identified in multiple tissues; similar IHC findings were described in cattle [12, 13, 15] and a sheep [14] infected by OvGHV2. Furthermore, conventional PCR using the Baxter primers [25] amplified the partial gene fragment of the OvGHV2 tegument protein gene, while qPCR based on the OvGH2 polymerase gene [29] detected genetic material from the positive sample, which was also quantified by dPCR, confirming that the intralesional MCFV antigens identified in several organs by IHC are those of OvGHV2. Consequently, these findings confirmed that this goat was infected by OvGHV2 and represent one the few confirmed cases of OvGHV2-related infections in this ruminant species. Additionally, the non-amplification of the nucleic acids of BoGHV6, BoAHV1, BoAHV5, BVDV, BPIV3, BRSV, BCoV, P. multocida, H. somni, M. haemolytica, and M. bovis suggests that these pathogens were not present in the target organs of this goat and, hence, did not contribute to the pathological alterations observed. Furthermore, neurological disease of sheep, such as scrapie and listeriosis, were discarded as contributing towards the brain syndrome due to the absence of typical histopathological findings. Therefore, it can be argued that since this goat did not present typical clinical manifestations nor gross and histopathologic findings associated with infection by OvGHV2 or SA-MCF, this animal was probably asymptomatic or suffered subclinical infection by this Macavirus (see below). Asymptomatic and/or subclinical infections were previously described in ruminants infected by OvGHV2 [9, 12, 13, 31, 32], so the real prevalence of OvGHV2-related infections in suspectable animal populations may be underestimated.
During this investigation, there was positive intracytoplasmic detection of MCFV antigens within the lungs, small intestine, liver, and kidney, while OvGHV2 DNA was only amplified form the small intestine. The 15A MAb IHC assay used during this investigation detects all members of the Macavirus genus known to be associated with the development of SA-MCF [6], with confirmation of the associated Macavirus requires the utilization of specific molecular assays [11]. Consequently, the non-amplification of OvGHV2 DNA from organs with positive immunoreactivity for MCFV may suggest that these organs were probably infected by another Macavirus. Simultaneous infections by BoGHV6 and OvGHV2, members of the Macavirus genus, in the same animal were previously described [13, 33]. However, in this case BoGHV6 was not identified. These findings may suggest the circulation of another undiagnosed Macavirus in ruminants from Brazil which may not need sheep as the asymptomatic host.
The conventional PCR assay amplified the partial fragment of the OvGHV2 tegument protein gene from the intestine of this goat that demonstrated positive intracytoplasmic immunoreactivity for MCFV, while sequencing and phylogenetic analyses confirmed that the amplified product had 100% aa homology with the OvGHV2 prototype strain. Alternatively, the viral load detected in this sample by dPCR was very low when compared with samples derived from an outbreak of SA-MCF [24] and cattle with typical histopathological manifestations of SA-MCF (manuscript in preparation). It was demonstrated by qPCR that the viral load within the intestine of sheep with clinical SA-MCF was 91.4 copy/µL, normal sheep 0.0148 copy/µL, and adult normal sheep as 0.0856 copy/µL [34]. However, in this case, the number of viral copies detected by dPCR (0.84 copy/µL) was probably insufficient to induce the typical histopathological manifestations of infection with subsequent demonstration of clinical SA-MCF, considering that sheep with clinical SA-MCF contained a viral load of 91.4 copy/µL [34]. Furthermore, the viral load detected in this goat was comparatively more elevated than that detected by digital droplet PCR (ddPCR) in sheep during an outbreak of SA-MCF [35]. Nevertheless, these findings correlate well with the known pathogenesis of SA-MCF, where lesion severity is directly related to viral load in the affected animal [36], and will explain the elevated viral load identified in the cow with SA-MCF and in the tissues of a cow with typical histopathological evidence of SA-MCF (manuscript in preparation). These findings may also suggest that sheep, and probably goats, must be infected with elevated viral loads of OvGHV2 for the development of clinical manifestations and/or histopathological lesions typical of SA-MCF, as was previously postulated [34]. Consequently, these findings confirmed that this goat was subclinically infected by OvGHV2 and/or was asymptomatic; asymptomatic infections by OvGHV2 seem to be the predominant manifestation of SA-MCF in goats [4].
Interestingly, the asymptomatic and/or subclinical infection by OvGHV2 in this goat contrasts the previous descriptions of OvGHV2-related diseases in goats. In one of these reports, the three affected goats demonstrated clinical manifestations of a neurological syndrome and developed typical histopathological alterations of OvGHV2 infections, with corneal being observed in one of these goats [18]; corneal edema is frequently diagnosed in cattle with SA-MCF [4, 5, 30]. In another description of OvGHV2-related infections in a goat, the animal had scaling alopecia clinically, a histopathologic diagnosis of folliculitis and vasculitis, with OvGHV2 DNA being detected from FFPE cutaneous tissues and blood samples [19]. A similar cutaneous disease was also associated with infection by OvGHV2 in a goat that had typical histopathological findings of fibrinoid vasculitis by histopathology [20]. Additionally, OvGHV2 was associated with disseminated necrotizing vasculitis and nonsuppurative encephalitis diagnosed in a pygmy goat (Capra hircus) maintained in a zoological park [21]. Further contrasting features during this investigation relative to the previous reports of OvGHV2 were the utilization of several diagnostic strategies to confirm the participation of this Macavirus in this goat and the quantification of OvGHV2 DNA. Moreover, during a seroepidemiological study done in the USA, 60.8% (177/291) of goats demonstrated positive seroreactivity to a MCFV antibody by competitive ELISA [37]. However, the ELISA was based on the 15A-MAb [6], so the real prevalence of OvGHV2 in goats could not have been estimated, considering that the 15A-MAb detects all Macavirus known to produce MCF [6]. However, a study done in Pakistan identified OvGHV2 DNA in 44% (22/50) of the blood samples collected from healthy goats [38], while 16.7% (1/6) of healthy goats from Switzerland contained OvGHV2 DNA from blood and nasal samples [39]. Consequently, additional studies must be done in goats to determine the effects and dynamics of infections by OvGHV2 on this ruminant species, since this Macavirus may not be able to induce the typical clinical findings of SA-MCF in goats.
The nanoplate digital PCR (dPCR) technology used during this investigation is a relatively new, revolutionized, third generation PCR, for the absolute quantification of target nucleic acids without the utilization of a standard curve [40–42]. The ddPCR methodology was previously used for the detection and quantification of OvGHV2 in tissues of cattle and sheep during an outbreak of SA-MCF [35]. Several animal studies have used the ddPCR technology; these include the quantification of BoAHV1 in the semen of cattle [43], and porcine circovirus 3 from clinical samples [44]. However, ddPCR is currently more frequently used to efficiently diagnose a wide range of diseases in human medicine [40, 42], and zoonotic diseases [42]. The quantification of OvGHV2 from animals with clinical SA-MCF, animals with histopathologic evidence of the classical manifestations of SA-MCF, and asymptomatic animals correlated well with the Cq identified in the respective qPCR assays; similar results were described in another study that used ddPCR to detect OvGHV2 [35]. Due to the high efficiency of the dPCR detection relative to qPCR [40, 41], this methodology will greatly improve the understanding of the pathogenesis of OvGHV2, particularly the dynamics associated with clinical and subclinical/asymptomatic infections in mammalian hosts. Additionally, the Cq obtained in all qPCR assays demonstrated that this assay can be used for the efficient detection of OvGHV2 in clinical samples of SA-MCF and in tissues with the typical histopathological lesions of SA-MCF. As far as the authors are aware, this is probably the first study to use nanoplate-based digital PCR (dPCR) for the quantification of DNA in animal studies. Although both diagnostic techniques can be used independently for the quantification of OvGHV2 DNA, we used the qPCR assays to validate the dPCR results.
The severe hepatic and brain lesions observed by histopathology, in addition to the laboratory findings, suggest that this goat suffered from nutritional deficiency with concomitant parasitic diseases, which may have been responsible for the hemodynamic alterations observed in this animal. The exact cause of the neurological syndrome remains obscure while the esophageal obstruction further hampered the wellbeing of this goat. As indicted above, common neurological diseases of ruminants were not identified, and it must be highlighted that listeriosis is not a frequent disease of ruminants in Brazil as compared to common Europe and North America [45, 46]. Collectively, these alterations probably contributed towards the debilitating immunological state of the goat and made this animal more susceptible to infection by OvGHV2. In this case, infection by OvGHV2 probably occurred via contamination during comingling with asymptomatic sheep maintained at the farm. Concomitant and contributory disease conditions were also identified in sheep infected with OvGHV2 [14, 34], suggesting that the occurrence of SA-MCF in sheep, and probably goats, may occur in these species when there is a severe concomitant debilitating condition or disease process. Notwithstanding the above, additional confirmed infections by goats are needed to understand the dynamics of OvGHV2 infections in this ruminant species, considering that only one animal was investigated during this study.
In conclusion, OvGHV2 DNA was amplified from the intestine of a goat without clinical manifestation or histopathological alterations of infection related to SA-MCF. A nanoplate-based digital PCR assay confirmed that the viral load present in the sample was very low and probably insufficient to induce clinical manifestations and/or typical histopathological lesions suggestive of SA-MCF and/or infection by OvGHV2. Consequently, these findings demonstrate that this goat was more likely subclinically infected or suffered from an asymptomatic infection by OvGHV2. Nevertheless, additional studies are need in goats to effectively evaluate the effects of OvGHV2 on the development of MCF in this ruminant species, since only one goat was evaluated during this study. Furthermore, this study has demonstrated that the 15A-MAb immunohistochemical assay can be used to detect MCFV antigens in goats, the nanoplate digital PCR methodology is an effective tool for the quantification of viral copy of OvGHV2 DNA in ruminants with and without typical histopathological manifestations of infection by this Macavirus, and the qPCR assay was efficient to identify OvGHV2 DNA from clinical samples.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplemental Figure 1 Radiological, gross, and histopathological findings observed in a goat infected by OvGHV2. The radiological evaluation demonstrated a severely distended esophagus (A). Gross evaluation revealed that the esophagus was obstructed by a mixture of ruminated and undigested leaves (B-C). Observe severe, random, hepatic necrosis (D), cerebral cortical edema (E), and acute hemorrhagic rhombencephalitis (F). D-F, Hematoxylin and eosin stain; Bar: D, 200 µm; E, 50 µm; F, 20 µm. (JPG 3105 KB)
Acknowledgements
The authors are extremely grateful to the Molecular Biology Institute of Paraná (IPBM) for the utilization of the nanoplate digital PCR (dPCR) technology and the technical support received from the research scientist, Marcel Kruchelski Tschá, and technician, Geison Eduardo Cambri, of IPBM during the realization of the dPCR analyses.
Author contribution
LES drafted the initial manuscript and contributed to the analysis and interpretation of all pathological and immunohistochemical data. AMDA and NSG participated in the realization of all conventional PCR analyses. MCR and IVS participated in the realization of the qPCR and dPCR assays and interpretations. AMDA, MCR, IVS, NSG, and AAA contributed to the analysis of all molecular data. JAK and PFVP contributed to the realization of all clinical and laboratorial analyses and interpretations. AACX participated in the realization of all histological, histochemical, and immunohistochemical stains and analyses. VRR and RFA participated in the realization of the post-mortem and histopathological evaluations. SAH contributed to the conception, design, and overall supervision of the study. All authors have read, critically analyzed, approved the final draft of this manuscript, and have agreed to be accountable for all aspects of the study in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Funding
The authors are grateful for the fellowships/funding received from the Brazilian National Council of Scientific and Technological Development (CNPq; Brazil), Coordination for the Improvement of Higher Education Personnel (CAPES; Brazil), and the National Institute of Science and Technology for Dairy Production Chain (INCT–LEITE). Xavier, A.A.C. is the recipient of graduate fellowships from the Coordination for the Improvement of Higher Education Personnel (CAPES; Brazil). Alfieri, A.A and Headley, S.A. are recipients of the Brazilian National Council of Scientific and Technological Development (CNPq; Brazil) fellowships.
Data availability
The partial nucleotide sequence of the OvGHV2 strain identified during this study is deposited in GenBank (https://www.ncbi.nlm.nih.gov/genbank/). The name of strain is OvHV-2/BRA-UEL/PR-N56/2021 (GenBank Accession # OK490363).
Declarations
Ethics approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. Moreover, permission to realize studies in sheep was obtained from the National Council for the Control of Animals in Experiments (CONCEA; Brazil) and approved by the Animal Ethics Committees for Animal Usage of the Universidade Norte do Paraná (CEUA, UNOPAR; protocol #008/20).
Conflict of interest
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.ICTV (2022) International committee on taxonomy of viruses. Subfamily: Gammaherpesvirinae, Genus: Macavirus. Retrieved 13 October, 2022, from https://ictv.global/report/chapter/herpesviridae/herpesviridae/macavirus
- 2.Crawford TB, Li H, Rosenburg SR, Norhausen RW, Garner MM. Mural folliculitis and alopecia caused by infection with goat-associated malignant catarrhal fever virus in two sika deer. J Am Vet Med Assoc. 2002;221(6):843–847. doi: 10.2460/javma.2002.221.843. [DOI] [PubMed] [Google Scholar]
- 3.Li H, Cunha CW, Taus NS, Knowles DP. Malignant catarrhal fever: inching toward understanding. Annul Rev Anim Biosci. 2014;2:209–233. doi: 10.1146/annurev-animal-022513-114156. [DOI] [PubMed] [Google Scholar]
- 4.O'Toole D, Li H. The pathology of malignant catarrhal fever, with an emphasis on ovine herpesvirus 2. Vet Pathol. 2014;51(2):437–452. doi: 10.1177/0300985813520435. [DOI] [PubMed] [Google Scholar]
- 5.Headley SA, Oliveira TES, Cunha CW. A review of the epidemiological, clinical, and pathological aspects of sheep-associated malignant catarrhal fever with emphasis on Brazil. Braz J Microbiol. 2020;51:1405–1432. doi: 10.1007/s42770-020-00273-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Li H, Shen DT, Knowles DP, Gorham JR, Crawford TB. Competitive inhibition enzyme-linked immunosorbent assay for antibody in sheep and other ruminants to a conserved epitope of malignant catarrhal fever virus. J Clin Microbiol. 1994;32(7):1674–1679. doi: 10.1128/jcm.32.7.1674-1679.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Li H, Gailbreath K, Bender LC, West K, Keller J, Crawford TB. Evidence of three new members of malignant catarrhal fever virus group in muskox (Ovibos moschatus), Nubian ibex (Capra nubiana), and gemsbok (Oryx gazella) J Wildl Dis. 2003;39(4):875–880. doi: 10.7589/0090-3558-39.4.875. [DOI] [PubMed] [Google Scholar]
- 8.Li H, Shen DT, O'Toole D, Knowles DP, Gorham JR, Crawford TB. Investigation of sheep-associated malignant catarrhal fever virus infection in ruminants by PCR and competitive inhibition enzyme-linked immunosorbent assay. J Clin Microbiol. 1995;33(8):2048–2053. doi: 10.1128/jcm.33.8.2048-2053.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Li H, McGuire TC, Muller-Doblies UU, Crawford TB. A simpler, more sensitive competitive inhibition enzyme-linked immunosorbent assay for detection of antibody to malignant catarrhal fever viruses. J Vet Diagn Invest. 2001;13(4):361–364. doi: 10.1177/104063870101300417. [DOI] [PubMed] [Google Scholar]
- 10.Müller-Doblies UU, Li H, Hauser B, Adler H, Ackermann M. Field validation of laboratory tests for clinical diagnosis of sheep-associated malignant catarrhal fever. J Clin Microbiol. 1998;36(10):2970–2972. doi: 10.1128/jcm.36.10.2970-2972.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Headley SA, Oliveira TES, Li H, Lisbôa JAN, Queiroz GR, Fritzen JTT, Flores EF, Alfieri AA, Cunha CW. Immunohistochemical detection of intralesional antigens of Ovine Gamaherpesvirus-2 in cattle with Sheep-Associated Malignant Catarrhal Fever. J Comp Path. 2020;174:86–98. doi: 10.1016/j.jcpa.2019.11.002. [DOI] [PubMed] [Google Scholar]
- 12.Headley SA, de Lemos GAA, Dall Agnol AM, Xavier AAC, Depes VCA, Yasumitsu CY, Oliveira TES, Silva LE, Faccin TC, Alfieri AA, Lisboa JAN. Ovine gammaherpesvirus 2 infections in cattle without typical manifestations of sheep-associated malignant catarrhal fever and concomitantly infected with bovine coronavirus. Braz J Microbiol. 2022;53:433–446. doi: 10.1007/s42770-021-00653-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Headley SA, Dall Agnol AM, Bessegato JA, Frucchi APS, Maturana ÉFL, Rodrigues RV, Xavier AAC, Alfieri AF, Alfieri AA. Association of ovine gammaherpesvirus 2 with an outbreak of acute respiratory disease in dairy cattle. Sci Rep. 2023;13(1):5623. doi: 10.1038/s41598-023-30133-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Headley SA, Agnol AMD, Navolar FMN, Frucchi APS, de Matos A, Pereira PFV, Xavier AAC, Dos Santos VH, Silva LE, Depes VCA, Alfieri AF, Cunha CW, Alfieri AA. Ovine gammaherpesvirus-2 infection associated with chronic interstitial pneumonia in a sheep. Microb Pathog. 2021;161(Pt A):105220. doi: 10.1016/j.micpath.2021.105220. [DOI] [PubMed] [Google Scholar]
- 15.Headley SA, Fritzen JTT, Silva DC, Xavier AAC, Yasumitsu CY, Silva FHP, Alfieri AF, Soethe AM, Alfieri AA. Histophilus somni disease conditions with simultaneous infections by ovine gammaherpesvirus 2 in cattle herds from Southern Brazil. Braz J Microbiol. 2023 doi: 10.1007/s42770-023-00915-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Li H, Cunha CW, Abbitt B, deMaar TW, Lenz SD, Hayes JR, Taus NS. Goats are a potential reservoir for the herpesvirus (MCFV-WTD), causing malignant catarrhal fever in deer. J Zoo Wildl Med. 2013;44(2):484–486. doi: 10.1638/2012-0087r.1. [DOI] [PubMed] [Google Scholar]
- 17.Li H, Keller J, Knowles DP, Crawford TB. Recognition of another member of the malignant catarrhal fever virus group: an endemic gammaherpesvirus in domestic goats. J Gen Virol. 2001;82(1):227–232. doi: 10.1099/0022-1317-82-1-227. [DOI] [PubMed] [Google Scholar]
- 18.Jacobsen B, Thies K, von Altrock A, Förster C, König M, Baumgärtner W. Malignant catarrhal fever-like lesions associated with ovine herpesvirus-2 infection in three goats. Vet Microbiol. 2007;124(3):353–357. doi: 10.1016/j.vetmic.2007.04.037. [DOI] [PubMed] [Google Scholar]
- 19.Foster AP, Twomey DF, Monie OR, Kiupel M, Hoffmann I, Willoughby K. Diagnostic exercise: generalized alopecia and mural folliculitis in a goat. Vet Pathol. 2010;47(4):760–763. doi: 10.1177/0300985810363574. [DOI] [PubMed] [Google Scholar]
- 20.Bagrinovschi B, Klang A, Turcitu MA, Möstl K, Krametter-Frötscher R. A case of malignant catarrhal fever in a goat associated with ovine herpesvirus 2 infection. Wien Tierärztl Monat - Vet Med Austria. 2014;101:142–148. [Google Scholar]
- 21.Twomey DF, Campbell I, Cranwell MP, Nettleton PF, Sayers G. Multisystemic necrotising vasculitis in a pygmy goat (Capra hircus) Vet Rec. 2006;158(25):867–869. doi: 10.1136/vr.158.25.867. [DOI] [PubMed] [Google Scholar]
- 22.Boom R, Sol CJ, Salimans MM, Jansen CL, Wertheim-van Dillen PM, van der Noordaa J. Rapid and simple method for purification of nucleic acids. J Clin Microbiol. 1990;28(3):495–503. doi: 10.1128/jcm.28.3.495-503.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Alfieri AA, Parazzi ME, Takiuchi E, Médici KC, Alfieri AF. Frequency of group A rotavirus in diarrhoeic calves in Brazilian cattle herds, 1998–2002. Trop Anim Health Prod. 2006;38(7–8):521–526. doi: 10.1007/s11250-006-4349-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Headley SA, Sousa IKF, Minervino AHH, Barros IO, BarrêtoJúnior RA, Alfieri AF, Ortolani EL, Alfieri AA. Molecular confirmation of ovine herpesvirus 2-induced malignant catarrhal fever lesions in cattle from Rio Grande do Norte, Brazil. Pesq Vet Bras. 2012;32(12):1213–1218. doi: 10.1590/S0100-736X2012001200001. [DOI] [Google Scholar]
- 25.Baxter SI, Pow I, Bridgen A, Reid HW. PCR detection of the sheep-associated agent of malignant catarrhal fever. Arch Virol. 1993;132(1–2):145–159. doi: 10.1007/BF01309849. [DOI] [PubMed] [Google Scholar]
- 26.Kumar S, Stecher G, Tamura K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33(7):1870–1874. doi: 10.1093/molbev/msw054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Hall TA. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. 1999;41:95–98. [Google Scholar]
- 28.Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16(2):111–120. doi: 10.1007/bf01731581. [DOI] [PubMed] [Google Scholar]
- 29.Cunha CW, Otto L, Taus NS, Knowles DP, Li H. Development of a multiplex real-time PCR for detection and differentiation of malignant catarrhal fever viruses in clinical samples. J Clin Microbiol. 2009;47(8):2586–2589. doi: 10.1128/jcm.00997-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Russell GC, Stewart JP, Haig DM. Malignant catarrhal fever: a review. Vet J. 2009;179(3):324–335. doi: 10.1016/j.tvjl.2007.11.007. [DOI] [PubMed] [Google Scholar]
- 31.Sausker EA, Dyer NW. Polymerase chain reaction and DNA sequencing for detection of Ovine Herpesvirus 2 in American Bison (Bison bison) J Vet Diagn Invest. 2002;14(1):40–46. doi: 10.1177/104063870201400108. [DOI] [PubMed] [Google Scholar]
- 32.O'Toole D, Li H, Sourk C, Montgomery DL, Crawford TB. Malignant catarrhal fever in a bison (Bison bison) feedlot, 1993–2000. J Vet Diagn Invest. 2002;14(3):183–193. doi: 10.1177/104063870201400301. [DOI] [PubMed] [Google Scholar]
- 33.Rosato G, Ruiz Subira A, Al-Saadi M, Michalopoulou E, Verin R, Dettwiler M, Nordgren H, Chiers K, Groβmann E, Köhler K, Suntz M, Stewart JP, Kipar A (2021) Gammaherpesvirus infections in cattle in Europe. Viruses. 13(12). 10.3390/v13122337 [DOI] [PMC free article] [PubMed]
- 34.Phillips IL, Cunha CW, Galbraith D, Highland MA, Bildfell RJ, Li H. High copy number of ovine gammaherpesvirus 2 DNA associated with malignant catarrhal fever-like syndrome in a lamb. J Vet Diagn Invest. 2018;30(4):623–627. doi: 10.1177/1040638718766976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Pinheiro de Oliveira TF, Laguardia-Nascimento M, Xavier FG, do Amaral Pinto C, Ferreira LR, de Castro Campos de Souza I, Hammerschmitt ME, Bianchi RM, Wronski JG, Etges RN, Rigon GM, Camargos MF, Júnior AVR, Fonseca Junior AA. Quantification of ovine herpesvirus 2 by digital PCR in an outbreak of malignant catarrhal fever. Arch Virol. 2019 doi: 10.1007/s00705-019-04382-y. [DOI] [PubMed] [Google Scholar]
- 36.Cunha CW, Gailbreath KL, O’Toole D, Knowles DP, Schneider DA, White SN, Taus NS, Davies CJ, Davis WC, Li H. Ovine herpesvirus 2 infection in American bison: virus and host dynamics in the development of sheep-associated malignant catarrhal fever. Vet Microbiol. 2012;159(3):307–319. doi: 10.1016/j.vetmic.2012.04.021. [DOI] [PubMed] [Google Scholar]
- 37.Li H, Shen DT, Jessup DA, Knowles DP, Gorham JR, Thorne T, O'Toole D, Crawford TB. Prevalence of antibody to malignant catarrhal fever virus in wild and domestic ruminants by competitive-inhibition ELISA. J Wildl Dis. 1996;32(3):437–443. doi: 10.7589/0090-3558-32.3.437. [DOI] [PubMed] [Google Scholar]
- 38.Riaz A, Dry I, Dalziel R, Rehman SU, Shah MA, Akhtar HMN, Yousaf A, Baig R. (2021) Molecular detection and characterization of ovine herpesvirus-2 using heminested PCR in Pakistan. J Vet Sci 22(4). 10.4142/jvs.2021.22.e51 [DOI] [PMC free article] [PubMed]
- 39.Stahel ABJ, Baggenstos R, Engels M, Friess M, Ackermann M. Two different Macaviruses, ovine herpesvirus-2 and caprine herpesvirus-2, behave differently in water buffaloes than in cattle or in their respective reservoir species. PLoS ONE. 2013;8(12):e83695. doi: 10.1371/journal.pone.0083695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kojabad AA, Farzanehpour M, Galeh HEG, Dorostkar R, Jafarpour A, Bolandian M, Nodooshan MM. Droplet digital PCR of viral DNA/RNA, current progress, challenges, and future perspectives. J Med Virol. 2021;93(7):4182–4197. doi: 10.1002/jmv.26846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kuypers J, Jerome KR. Applications of Digital PCR for clinical microbiology. J Clin Microbiol. 2017;55(6):1621–1628. doi: 10.1128/jcm.00211-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lei S, Chen S, Zhong Q. Digital PCR for accurate quantification of pathogens: principles, applications, challenges and future prospects. Int J Biol Macromol. 2021;184:750–759. doi: 10.1016/j.ijbiomac.2021.06.132. [DOI] [PubMed] [Google Scholar]
- 43.Yu Z, Zhao Z, Chen L, Yan H, Cui Q, Ju X, Yong Y, Liu X, Ma X, Zhang G. Development of a droplet digital PCR assay to detect bovine alphaherpesvirus 1 in bovine semen. BMC Vet Res. 2022;18(1):125. doi: 10.1186/s12917-022-03235-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Liu Y, Meng H, Shi L, Li L. Sensitive detection of porcine circovirus 3 by droplet digital PCR. J Vet Diagn Invest. 2019;31(4):604–607. doi: 10.1177/1040638719847686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Headley SA, Fritzen JTT, Queiroz GR, Oliveira RAM, Alfieri AF, Santis GWD, Lisbôa JAN, Alfieri AA. Molecular characterization of encephalitic bovine listeriosis from southern Brazil. Trop Anim Health Prod. 2013;46:19–25. doi: 10.1007/s11250-013-0441-0. [DOI] [PubMed] [Google Scholar]
- 46.Queiroz GR, Oliveira RAM, Flaiban KKMC, Santis GWD, Bracarense APFRL, Headley SA, Alfieri AA, Lisbôa JAN. Diagnóstico diferencial das doenças neurológicas dos bovinos no estado do Paraná. Pesq Vet Bras. 2018;38(1):1264–1277. doi: 10.1590/1678-5150-PVB-5429. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental Figure 1 Radiological, gross, and histopathological findings observed in a goat infected by OvGHV2. The radiological evaluation demonstrated a severely distended esophagus (A). Gross evaluation revealed that the esophagus was obstructed by a mixture of ruminated and undigested leaves (B-C). Observe severe, random, hepatic necrosis (D), cerebral cortical edema (E), and acute hemorrhagic rhombencephalitis (F). D-F, Hematoxylin and eosin stain; Bar: D, 200 µm; E, 50 µm; F, 20 µm. (JPG 3105 KB)
Data Availability Statement
The partial nucleotide sequence of the OvGHV2 strain identified during this study is deposited in GenBank (https://www.ncbi.nlm.nih.gov/genbank/). The name of strain is OvHV-2/BRA-UEL/PR-N56/2021 (GenBank Accession # OK490363).