Abstract
Chlamydia trachomatis is a common cause of sexually transmitted infections in humans with devastating sequelae. Understanding of disease on all scales, from molecular details to the immunology underlying pathology, is essential for identifying new ways of preventing and treating chlamydia. Infection models of various complexity are essential to understand all aspects of chlamydia pathogenesis. Cell culture systems allow for research into molecular details of infection, including characterization of the unique biphasic Chlamydia developmental cycle and the role of type III secreted effectors in modifying the host environment to allow for infection. Multi-cell type and organoid culture provide means to investigate how cells other than the infected cells contribute to the control of infection. Emerging comprehensive three-dimensional biomimetic systems may fill an important gap in current models to provide information on complex phenotypes that cannot be modeled in simpler in vitro models.
Keywords: Chlamydia, biomimetics, model systems, host-pathogen interactions, obligate intracellular bacteria
Introduction
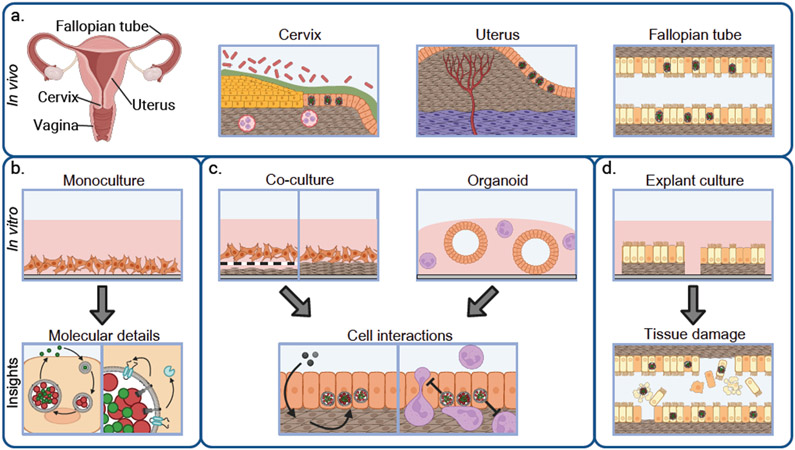
Chlamydia trachomatis is a human-specific, obligate intracellular bacterium that causes the most frequently reported bacterial sexually transmitted infection [1]. The female reproductive tract is composed of a series of connected organs that each have distinct cell composition and architecture (Figure 1a). In women, C. trachomatis infections are initiated within the lower reproductive tract (vagina and cervix) and can ascend to organs of the upper reproductive tract (uterus and fallopian tubes). In all cases, the bacteria replicates within epithelial cells. If the bacteria are not cleared by the immune system, infection induces substantial inflammation within reproductive tissue and can lead to sequelae including pelvic inflammatory disease and infertility [2], as well as a potential increased risk of developing cervical cancer [3]. Despite the availability of effective antibiotic treatments, C. trachomatis infections still represent a substantial public health problem due to the sequelae, the prevalence of asymptomatic infections, and the lack of a vaccine in humans [4].
Figure 1.
Current models of the female genital tract to study Chlamydia infections. (a) The human female reproductive tract. Organs relevant to Chlamydia infection are labeled, with overall cellular composition of the cervix, uterus, and fallopian tube lining shown at right. The cervix contains squamous (light orange, at left) and columnar (orange, at right) epithelial cells, a layer of mucus (green), the cervicovaginal microbiota (red), and a layer of fibroblasts (brown) containing blood vessels lined with endothelial cells (red) and containing various immune cells (purple). The uterus includes columnar epithelial cells (orange), endometrial stromal cells (brown), muscle cells of the myometrium (dark purple), and spiral arteries (red). The fallopian tube lining contains secretory (orange) and ciliated (yellow) epithelial cells as well as fibroblasts (brown). Chlamydia inclusions containing infectious (green) and replicative (red) bacteria are depicted in some of the epithelial cells. (b) Top panel: The most common in vitro immortalized cell culture model of the female reproductive tract, consisting of a monolayer of immortalized epithelial cells (orange) on the bottom of a plastic dish (gray) covered in cell culture media (pink). Such systems are used to gain insights into molecular details of Chlamydia infection (bottom panel): at left, the developmental cycle of Chlamydia elementary bodies (green) and reticulate bodies (red) within the lumen of the inclusion and, at right, the secretion of various type 3 secretion system (black) effectors (teal). (c) Top left panel: Common in vitro co-culture models include immortalized epithelial cells (orange), sometimes on a porous transwell membrane (black dashed line), with fibroblasts (brown) at the bottom of the plastic dish (gray), all within cell culture media (pink). Top right panel: Other co-culture models include organoid systems, containing spheroids of stem cell-derived epithelial cells (orange) within a gel matrix (pink) in a plastic dish (gray), with the potential addition of other components such as immune cells (purple). Bottom panel: Co-culture systems are useful to investigate proposed cell-cell interactions during infection, such as the role of stromal cells (brown) as intermediates in relaying signals, such as (left) hormones (grey spheres) to modulate Chlamydia infection in epithelial cells (orange) or (right) investigating how Chlamydia impedes neutrophil (purple) recruitment and activation to the site of infection. (d) Top panel: An in vitro explant model of fallopian tube, consisting of excised pieces of fallopian tube in a dish (gray) within cell culture media (pink). Bottom panel: Explant models have allowed for proposed insights into complex phenotypes, such as tissue damage, caused by infection.
There has been substantial research into chlamydia pathogenesis to understand how to prevent and treat infections and their sequelae. As with other infectious diseases, a variety of models are used to investigate Chlamydia-host interactions, ranging from simple monoculture systems in vitro to in vivo work in mice and humans. Here, we describe how different models, particularly in vitro systems, have aided in dissecting the molecular basis of chlamydia pathogenesis. We additionally provide a perspective on emerging and future models that will be valuable for human-relevant investigations of Chlamydia-host interactions, focusing on systems made possible by recent advances in three dimensional (3D) complex cell culture models.
Insights into Chlamydia-host interactions gained from immortalized cell culture systems
Insights from commonly used monoculture systems
Cancer-derived (e.g. HeLa) and artificially immortalized (e.g. COS-7) cell lines are used to provide insights into host-Chlamydia interactions at the cellular and molecular level (Figure 1b). Immortalization provides the cells with the ability to expand for generations, allowing for many replicable experiments. Typical monoculture systems are often readily genetically manipulable, which has been especially useful in Chlamydia research as Chlamydia spp. were genetically intractable until recently. Such monoculture systems are also amenable to microscopy, a cornerstone of Chlamydia research for decades, that has allowed for key insights into Chlamydia cell biology. Monoculture systems have long been, and will remain to be, attractive models for dissecting fundamental molecular aspects of Chlamydia-host interactions.
Chlamydia spp. undergo a unique biphasic developmental cycle wherein the bacterium enters the host cell as an infectious elementary body (EB), forming a membrane bound compartment termed the inclusion within which the EB transitions to the replicative, but non-infectious reticulate body (RB) (Figure 1b) [5]. After replicating, RBs transition back to EBs, which are released from the infected cell via extrusion or lysis. Transcriptomic and proteomic based studies provided a wealth of information about the distinct properties of RBs and EBs [6-8] and were instrumental in generating fluorescent reporters of Chlamydia developmental forms to aid in quantifying the kinetics of the development cycle [9-11]. While little is conclusively known about the factors that regulate transition from one developmental form to another, several models have recently been proposed. Investigations utilizing fluorescent reporters suggest a model by which EBs are the result of asymmetric division of RBs [11]. Another recently proposed model, based on 3D electron microscopy of infected HeLa cells, predicts size-dependent regulation of RB to EB transition [12]. Additional data indicate that the metabolic state of host cells may be one signal that promotes the initial EB to RB transition after infection [13]. Specific Chlamydia genes, including clpX [14,15], incS [16], and euo [17], were also recently identified as regulating development through incompletely understood mechanisms. Regulation of the unique developmental cycle is of key interest to ongoing research, with monoculture systems providing the ideal system for detailed molecular dissection of pathways which could be targeted to limit the spread of the bacteria during infection.
Many host-Chlamydia interactions are dependent on the bacterial type III secretion system (T3SS), which translocates effector proteins from within the bacteria into the inclusion membrane and the host cell [18] (Figure 1b). To date, up to 100 effectors, including 50 predicted (~ 40 confirmed) inclusion membrane-associated (Inc) proteins, have been identified [19]. The recent development of genetic tools including expression plasmids and inducible expression systems [20-22], random mutagenesis [23-25], and targeted knockout and knockdowns [26-29] for Chlamydia have been instrumental for characterizing the roles of Chlamydia effectors.
Beside promoting uptake of the bacteria [30-33], many Chlamydia effectors interface with host proteins to promote interactions, directly or indirectly, between the inclusion and host organelles to favor the intracellular Chlamydia lifestyle [19,34] (Figure 1b). Some of the latest studies show that several T3SS substrates including the inclusion membrane proteins IncD [35,36], IncV [37,38], and IncS [39] mediate interactions between the inclusion and the endoplasmic reticulum (ER), establishing ER-inclusion membrane contact sites, putatively promoting processes such as lipid acquisition [40]. Multiple effectors target the Golgi apparatus, with Cdu1 inducing Golgi fragmentation [41,42] and InaC coordinating with host ARF GTPases to control positioning of Golgi ministacks around the inclusion [23,43]. At least two effectors localize to the mitochondria, which may contribute to altered mitochondrial protein composition during infection [44]. Host cell division is substantially altered, such as Chlamydia effector CteG inducing abnormal centrosome amplification [45] and IncM inducing multinucleation and inhibiting cytokinesis [46].
Perhaps not surprisingly for an obligate intracellular pathogen with a reduced genome, several secreted effectors have multiple roles during infection, both at distinct times and sites. IncM impacts a plethora of aspects of host cell biology, altering interactions with the Golgi apparatus in addition to its roles in modulating cell division [46]. IncS promotes the EB to RB transition early in infection and maintains the integrity of the inclusion late in infection [16,39]. CTL0390, subsequently renamed GarD, inhibits both ubiquitylation and subsequent clearance of the inclusion [47], as well as interacting with the STING pathway to promote host cell lysis [48]. CpoS also interacts with the STING pathway and suppresses host interferon signaling [49], while additionally regulating formation of microdomains on the inclusion surface [50]. Moonlighting roles of effector proteins may be more common than currently appreciated, providing a valuable vein of further investigation into molecular host-Chlamydia interactions.
The emergence of more physiologically relevant cell monoculture systems
Cancer-derived cell lines exhibit a variety of defects compared to cells in vivo, including altered metabolism, chromosomal abnormalities, lack of polarization, and altered immune responses to pathogens, including Chlamydia [51]. These significant limitations have driven the identification and development of more physiologically relevant cell lines to enhance the human relevance of in vitro studies.
A2EN and End1 cells are endocervical epithelial cells immortalized by expression of the human papillomavirus genes E6 and E7 [52,53]. They closely mimic epithelial cells in terms of polarization, metabolism, expression of key cell markers and cytokines, and hormone responses [52-55], thereby providing attractive alternatives to assess phenotypes observed in less physiologically relevant cell lines, or to reveal phenotypes that differ or would not be detectable in typical, non-polarized cell lines [45,49,55,56]. For example, the Chlamydia effector protein TepP is crucial for replication in A2EN, but not in HeLa cells [56]. Tight junctions present in A2EN, but not HeLa, monolayers, were destabilized by TepP, identifying a putative role in promoting the spread of infection in vivo [57]. Although only utilized in a limited number of studies so far, newer, more relevant cell lines that provide much of the ease of use of cancer-derived cell lines, without the altered physiology, will be key as the field moves forward in elucidating basic concepts of Chlamydia-host interactions.
Insights into Chlamydia-host interactions gained from co-culture systems
In vivo, infected epithelial cells are not found in isolation. Rather, the outcome of infection depends on interactions between infected cells and the surrounding environment, including stromal cells, immune cells, and the microbiota. Whole-organism models, such as mice or humans, overcome this limitation, but also come with significant disadvantages (Box 1 and Box 2).
Box 1. Murine models of Chlamydia infection of the female reproductive tract.
Various small animal models have been used for Chlamydia research, with mouse models being the most common. Murine models have been instrumental for dissecting the contribution of various immune factors in restricting Chlamydia infections. Amongst the many insights, it has become clear that a few key host factors, including T cells and interferon gamma (IFN-γ) are especially important in controlling Chlamydia [89-91]. Additionally, with the recent advances in genetic tools for Chlamydia and the expansion of mutants, mouse models have been used to confirm that genes important for infection in vitro also matter in vivo [16,45,57,92-95].
Despite their utility, murine models of Chlamydia remain limited in their human relevance. One major limitation is the fact that intravaginal infection of a mouse with C. trachomatis does not resemble human infection, as the bacteria are rapidly cleared and do not efficiently cause pathology [91,96,97]. Rapid clearance has been linked to an increased susceptibility of C. trachomatis to murine innate immune responses, such as an inability to counteract the murine IFN-γ response [96,98]. Sustained murine infections require the utilization of a distinct Chlamydia species, C. muridarum, for which the mouse is the natural host. Like C. trachomatis, C. muridarum is species-restricted by its sensitivity to human, but not murine, IFN-γ responses [99]. Differential susceptibility to infection by the two species can be partially ameliorated by utilizing a transcervical infection route, introducing C. trachomatis directly into the mouse uterine horns and resulting in substantial pathology [91,100,101].
Box 2. Insights into Chlamydia gained from human studies.
Observational studies utilizing human subjects can overcome Chlamydia species specificity (Box 1). Such studies have identified various host genetic factors involved in controlling susceptibility to infection and pathology [102-106]. One recent study identified host genes associated with infertility, allowing more focused investigation of sequelae of Chlamydia with clinical relevance [105].
Human studies have also demonstrated key roles for the cervicovaginal microbiota in modulating susceptibility to infection [107,108]. These studies demonstrated that communities of bacteria dominated by Lactobacillus spp. within the lower reproductive tract are associated with reduced risk of Chlamydia infection and increased ability to clear infection. Lactobacillus-dominated cervicovaginal communities (community state type (CST) I, II, III, and V) are generally considered healthy, in contrast to the diverse communities of predominantly anaerobic bacteria, termed CST IV, which are associated with the dysbiotic condition bacterial vaginosis and an increased risk of Chlamydia infection [109,110]. The protective role for Lactobacillus spp. has been investigated using cell culture models [111], identifying D (−) lactic acid as a key mediator of anti-Chlamydia activity [61,112].
Further human studies identifying host and bacterial factors that may be involved in disease will likely continue to be key sources of hypotheses to be tested in controlled laboratory environments.
Integrating multiple cell types into a single cell culture dish is a convenient middle-ground to study inter-cell type interactions at the molecular level. Co-culture systems described so far typically consist of epithelial cells grown on the permeable membrane of a transwell insert with stromal cells either as a feeder layer at the bottom of the well [58-60], or on the other side of the membrane [61,62] (Figure 1c). Alternatively, stromal cells can be embedded within a collagen matrix with the epithelial cells seeded on top [63]. Such systems allowed for investigating the role of female sex hormones and inter-cell signaling on Chlamydia infection, revealing the stromal cell-dependent pro-Chlamydia effects of estrogen upon co-culture of HEC-1-B or Ishikawa endometrial cells along SHT-290 endometrial fibroblasts (Figure 1c) [58]. With increasing cellular complexity, co-culture systems will be instrumental in teasing apart how inter-cell interactions influence Chlamydia infection, in relatively simple and tractable setups.
Towards more physiologically relevant cells and structure: primary cells, organoids, and explants
Many current systems remain limited in their resemblance to in vivo conditions, due to lack of physiological cell behavior, architecture, or both. Additionally, cell lines in common use are predominantly derived from the cervix and uterus preventing modeling of infection within portions of the upper reproductive tract, such as the fallopian tube, that are relevant for clinical pathology of Chlamydia infections.
Primary cells:
Primary cells are defined as cells isolated from biopsy material or whole organs that are subsequently propagated in vitro without immortalization. Models utilizing primary cells derived from relevant sites of the reproductive tract are key to increasing the physiological relevance of in vitro models of Chlamydia infection. Commercial sources are a good alternative to the potentially challenging acquisition of patient-derived cells, but do not alleviate the limited replicative capacity and culturing requirements of primary cells. Regardless, primary cells from throughout the female reproductive tract, including cervix and fallopian tube, are available and susceptible to Chlamydia infections [47,64,65]. The benefit of using primary cells over immortalized cells was recently highlighted by the replication defect of the cteG mutant displayed in primary cervical cells and in vivo, but not in HeLa or A2EN cells [45]. These results reinforce that, in addition to validating key phenotypes in primary cells to increase confidence in human relevance, investigators should also consider primary cells in their overall experimental design.
Organoids:
Organoids are 3-dimensional simulacra of in vivo tissue generated from stem cells cultured in vitro. The added benefits of organoids, compared to primary cells, are increased self-renewal and the possible inclusion of multiple cell types. Murine organoids, despite their limitations (Box 1), can offset the major hurdle of sourcing human reproductive tract tissues. Organoids derived from several female reproductive organs including human and murine cervix [66,67], human and murine fallopian tubes [13,68,69], and murine uterine endometrium [57,70,71] have been used to successfully model Chlamydia intracellular replication, and to confirm functions of host genes, e.g. SLC1A5 [13], or Chlamydia genes, e.g. tepP [57]. Organoids may also provide a model to investigate the link between Chlamydia infections and gynecological cancers, as highlighted by the increased stemness of epithelial cells upon infection of human fallopian tube organoids [68]. Immune cells can also be incorporated into organoid culture models, which has already revealed the role of the chlamydial effector TepP in driving neutrophil infiltration [71] (Figure 1c).
Explant cultures:
Explant cultures consist of the in vitro culture of small intact portions of surgically resected tissue or organs collected from an organism, allowing for the faithful preservation of tissue architecture (Figure 1d). However, key factors such as immune cells may be lacking. Sourcing, culture conditions, and longevity are also significant limitations. Fallopian tube explant cultures are most commonly used in Chlamydia research, allowing for investigation of an organ not modeled by cell culture lines [72-76]. Explant cultures are attractive models to investigate tissue disruption during infection and were instrumental in revealing roles for Wnt and IL-1 signaling in mediating tissue damage [75,76].
Biomimetics as potential future models for Chlamydia trachomatis infection
Advantages and limitations of biomimetics:
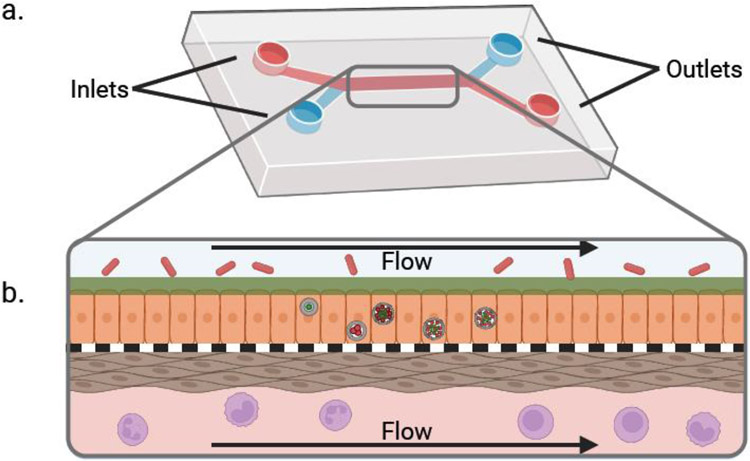
Organ-on-a-chip or biomimetic models (Figure 2) have emerged in recent years as attractive options for modeling human diseases in vitro [77]. The integration of diverse cell types into an in-house made or commercially available single microfluidic device mimics the in vivo tissue structure allowing for in-depth, human-relevant studies in a reductionist system.
Figure 2.
Key features of future biomimetic models to study Chlamydia infections. (a) A representative biomimetic chip containing two stacked channels (red and blue, respectively) separated by a porous membrane (not shown). Inlets and outlets allow for the introduction and/or removal of media to/from the channels. (b) Longitudinal side view through the center of the biomimetic chip shown in (a). An upper layer of columnar epithelial cells (orange) with mucus (green) are separated from a lower layer of fibroblasts (brown) via a porous membrane (black dashed line). Each channel can be supplied with a distinct media type (pink and blue), with additional biological or chemical alterations as needed. Here, the upper channel contains bacterial members of the microbiota (red), while the lower channel contains an assortment of immune cells (purple).
Bioengineering considerations are instrumental in creating a 3D simulacrum of tissue structure within a biomimetic device. Physical factors, such as stretch or flow, which are key modulators of tissue physiology, can be integrated to improve similarity to in vivo tissue [78]. The addition or removal of specific biological components including immune cells [79,80] and the microbiota [79,81,82], can reveal their roles in disease processes. Like any model, biomimetics are only as good as the cells that constitute them and carry the inherent limitations of each kind of cell culture as discussed above. Immortalized cancer-derived cell lines [83], primary cells [84], or organoid-derived cells [85] are used within biomimetics and should be chosen based on the goals of the study.
While biomimetic models are getting traction in various biomedical fields, the high cost of commercially available devices and the lack of technical expertise is perceived as a significant limitation, especially among scientists who do not use such platforms [86]. Current users also identify a number of technical limitations such as lack of long-term stability and insufficient complexity as prominent drawbacks of current systems [86]. Collaborations between groups with overlapping interests, and complementary expertise in bioengineering and biology, will be instrumental in overcoming perceived and technical obstacles to promote the widespread adoption of biomimetic devices.
Biomimetics and infectious diseases research:
Biomimetics of organs such as the lung, gut, and liver are making their way into infectious diseases research to dissect cellular details of host-pathogen interactions [87]. Additionally, the development of biomimetics of various organs of the female reproductive tract [88] is promising for Chlamydia research, providing potential new models of infection. Advancements such as including the microbiota and immune cells, as recently described in other culture systems for Chlamydia [62,71], will be instrumental in utilizing biomimetic technology to probe aspects of Chlamydia infections that are often not possible with current models but are key for infection in vivo. It opens the possibility of modulating immune cell populations in ways that are not readily feasible in vivo or in explant cultures, or of investigating the role of stromal cells during infection when structure more closely mimics in vivo organization than current co-culture models.
Conclusions
All models for studying a specific disease, including chlamydia, possess inherent biological and technical limitations that may decrease relevance of a model to human disease and that impede the use of these models. However, most models are still useful for studying certain aspects of a disease depending on the level of complexity required for the study at hand. Care should be given to ensure that a chosen model is both biologically and technically appropriate for the question being asked.
Monoculture of immortalized cells is best suited for interrogating molecular interactions and performing large-scale experiments, such as screens, that would be unwieldy in other models, but may require validation in primary cells or organoids. Co-culture systems allow for investigation of interactions between cell types, while explant cultures and organoid systems provide the ability to interrogate phenotypes such as tissue damage that are not found in simpler culture models. We further propose that biomimetic models are an attractive middle ground, overcoming the limitations of current mono- and co-culture models and offering the tractability and the relevance needed to further explore the molecular mechanisms underlying chlamydia pathogenesis.
Highlights.
Chlamydia infections have damaging consequences on female reproductive health
Diverse human model systems are required for a comprehensive understanding of Chlamydia-host interactions
Simple cell culture systems permit studies at the molecular level
Multi-cell systems allow insights into cell-cell interactions during infection
Biomimetic systems provide tractable models for complex host-pathogen interactions
Acknowledgements
The authors thank members of the Derré laboratory for their feedback on the manuscript. This work is supported by the National Institute of Allergy and Infectious Disease grants R01AI162758, R21AI166237, and U19AI158930 to ID. Figures were created using BioRender.com.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Declaration of Competing Interest
The authors declare no competing interests relevant to this article.
Data Availability
No data were used for the research described in the article.
References and recommended reading
Papers of particular interest, published within the period of review have been highlighted as:
* of special interest
** of outstanding interest
- 1.Kreisel KM, Weston EJ, St Cyr SB, Spicknall IH: Estimates of the Prevalence and Incidence of Chlamydia and Gonorrhea Among US Men and Women, 2018. Sex Transm Dis 2021, 48:222–231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Haggerty CL, Gottlieb SL, Taylor BD, Low N, Xu F, Ness RB: Risk of sequelae after Chlamydia trachomatis genital infection in women. J Infect Dis 2010, 201 Suppl 2:S134–155. [DOI] [PubMed] [Google Scholar]
- 3.Karim S, Souho T, Benlemlih M, Bennani B: Cervical Cancer Induction Enhancement Potential of Chlamydia trachomatis: A Systematic Review. Curr Microbiol 2018, 75:1667–1674. [DOI] [PubMed] [Google Scholar]
- 4.de la Maza LM, Darville TL, Pal S: Chlamydia trachomatis vaccines for genital infections: where are we and how far is there to go? Expert Rev Vaccines 2021, 20:421–435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Abdelrahman YM, Belland RJ: The chlamydial developmental cycle. FEMS Microbiol Rev 2005, 29:949–959. [DOI] [PubMed] [Google Scholar]
- 6.Belland RJ, Zhong G, Crane DD, Hogan D, Sturdevant D, Sharma J, Beatty WL, Caldwell HD: Genomic transcriptional profiling of the developmental cycle of Chlamydia trachomatis. Proc Natl Acad Sci U S A 2003, 100:8478–8483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Saka HA, Thompson JW, Chen YS, Kumar Y, Dubois LG, Moseley MA, Valdivia RH: Quantitative proteomics reveals metabolic and pathogenic properties of Chlamydia trachomatis developmental forms. Mol Microbiol 2011, 82:1185–1203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Skipp PJ, Hughes C, McKenna T, Edwards R, Langridge J, Thomson NR, Clarke IN: Quantitative Proteomics of the Infectious and Replicative Forms of Chlamydia trachomatis. PLoS One 2016, 11:e0149011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Cortina ME, Ende RJ, Bishop RC, Bayne C, Derré I: Chlamydia trachomatis and Chlamydia muridarum spectinomycin resistant vectors and a transcriptional fluorescent reporter to monitor conversion from replicative to infectious bacteria. PLoS One 2019, 14:e0217753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chiarelli TJ, Grieshaber NA, Omsland A, Remien CH, Grieshaber SS: Single-Inclusion Kinetics of Chlamydia trachomatis Development. mSystems 2020, 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chiarelli TJ, Grieshaber NA, Appa C, Grieshaber SS: Computational Modeling of the Chlamydial Developmental Cycle Reveals a Potential Role for Asymmetric Division. mSystems 2023, 8:e0005323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lee JK, Enciso GA, Boassa D, Chander CN, Lou TH, Pairawan SS, Guo MC, Wan FYM, Ellisman MH, Sutterlin C, et al. : Replication-dependent size reduction precedes differentiation in Chlamydia trachomatis. Nat Commun 2018, 9:45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Rajeeve K, Vollmuth N, Janaki-Raman S, Wulff TF, Baluapuri A, Dejure FR, Huber C, Fink J, Schmalhofer M, Schmitz W, et al. : Reprogramming of host glutamine metabolism during Chlamydia trachomatis infection and its key role in peptidoglycan synthesis. Nat Microbiol 2020, 5:1390–1402. [DOI] [PubMed] [Google Scholar]
- 14.Wood NA, Blocker AM, Seleem MA, Conda-Sheridan M, Fisher DJ, Ouellette SP: The ClpX and ClpP2 Orthologs of Chlamydia trachomatis Perform Discrete and Essential Functions in Organism Growth and Development. mBio 2020, 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Wood NA, Swoboda AR, Blocker AM, Fisher DJ, Ouellette SP: Tag-Dependent Substrate Selection of ClpX Underlies Secondary Differentiation of Chlamydia trachomatis. mBio 2022, 13:e0185822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Cortina ME, Bishop RC, DeVasure BA, Coppens I, Derré I: The inclusion membrane protein IncS is critical for initiation of the Chlamydia intracellular developmental cycle. PLoS Pathog 2022, 18:e1010818. * This paper defines a role for Chlamydia inclusion membrane protein IncS in promoting the differentiation from elementary body to reticulate body. Additionally, it is an example of the utility of multiple models in vitro and in vivo to confirm the importance of phenotypes in diverse systems.
- 17.Hakiem OR, Rizvi SMA, Ramirez C, Tan M: Euo is a developmental regulator that represses late genes and activates midcycle genes in. mBio 2023:e0046523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Rucks EA: Type III Secretion in Chlamydia. Microbiol Mol Biol Rev 2023:e0003423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Bugalhão JN, Mota LJ: The multiple functions of the numerous. Microb Cell 2019, 6:414–449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wang Y, Kahane S, Cutcliffe LT, Skilton RJ, Lambden PR, Clarke IN: Development of a transformation system for Chlamydia trachomatis: restoration of glycogen biosynthesis by acquisition of a plasmid shuttle vector. PLoS Pathog 2011, 7:e1002258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wickstrum J, Sammons LR, Restivo KN, Hefty PS: Conditional gene expression in Chlamydia trachomatis using the Tet system. PLoS One 2013, 8:e76743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Agaisse H, Derré I: A C. trachomatis cloning vector and the generation of C. trachomatis strains expressing fluorescent proteins under the control of a C. trachomatis promoter. PLoS One 2013, 8:e57090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kokes M, Dunn JD, Granek JA, Nguyen BD, Barker JR, Valdivia RH, Bastidas RJ: Integrating chemical mutagenesis and whole-genome sequencing as a platform for forward and reverse genetic analysis of Chlamydia. Cell Host Microbe 2015, 17:716–725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.LaBrie SD, Dimond ZE, Harrison KS, Baid S, Wickstrum J, Suchland RJ, Hefty PS: Transposon Mutagenesis in Chlamydia trachomatis Identifies CT339 as a ComEC Homolog Important for DNA Uptake and Lateral Gene Transfer. mBio 2019, 10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.O'Neill CE, Skilton RJ, Forster J, Cleary DW, Pearson SA, Lampe DJ, Thomson NR, Clarke IN: An inducible transposon mutagenesis approach for the intracellular human pathogen Chlamydia trachomatis. Wellcome Open Res 2021, 6:312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mueller KE, Wolf K, Fields KA: Gene Deletion by Fluorescence-Reported Allelic Exchange Mutagenesis in Chlamydia trachomatis. mBio 2016, 7:e01817–01815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Johnson CM, Fisher DJ: Site-specific, insertional inactivation of incA in Chlamydia trachomatis using a group II intron. PLoS One 2013, 8:e83989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ouellette SP, Blay EA, Hatch ND, Fisher-Marvin LA: CRISPR Interference To Inducibly Repress Gene Expression in Chlamydia trachomatis. Infect Immun 2021, 89:e0010821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Keb G, Hayman R, Fields KA: Floxed-Cassette Allelic Exchange Mutagenesis Enables Markerless Gene Deletion in Chlamydia trachomatis and Can Reverse Cassette-Induced Polar Effects. J Bacteriol 2018, 200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Keb G, Ferrell J, Scanlon KR, Jewett TJ, Fields KA: Chlamydia trachomatis TmeA Directly Activates N-WASP To Promote Actin Polymerization and Functions Synergistically with TarP during Invasion. mBio 2021, 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Scanlon KR, Keb G, Wolf K, Jewett TJ, Fields KA: TmeB antagonizes actin polymerization via direct interference with Arp2/3 activity. Front Cell Infect Microbiol 2023, 13:1232391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Faris R, McCullough A, Andersen SE, Moninger TO, Weber MM: The Chlamydia trachomatis secreted effector TmeA hijacks the N-WASP-ARP2/3 actin remodeling axis to facilitate cellular invasion. PLoS Pathog 2020, 16:e1008878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Romero MD, Carabeo RA: Distinct roles of the Chlamydia trachomatis effectors TarP and TmeA in the regulation of formin and Arp2/3 during entry. J Cell Sci 2022, 135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Bastidas RJ, Valdivia RH: The emerging complexity of Chlamydia trachomatis interactions with host cells as revealed by molecular genetic approaches. Curr Opin Microbiol 2023, 74:102330. * This review summarizes recent insights into the roles of many C. trachomatis effectors during infection, providing further description of the molecular details of Chlamydia-host interactions.
- 35.Agaisse H, Derré I: Expression of the effector protein IncD in Chlamydia trachomatis mediates recruitment of the lipid transfer protein CERT and the endoplasmic reticulum-resident protein VAPB to the inclusion membrane. Infect Immun 2014, 82:2037–2047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Derré I, Swiss R, Agaisse H: The lipid transfer protein CERT interacts with the Chlamydia inclusion protein IncD and participates to ER-Chlamydia inclusion membrane contact sites. PLoS Pathog 2011, 7:e1002092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Stanhope R, Flora E, Bayne C, Derré I: IncV, a FFAT motif-containing Chlamydia protein, tethers the endoplasmic reticulum to the pathogen-containing vacuole. Proc Natl Acad Sci U S A 2017, 114:12039–12044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ende RJ, Murray RL, D'Spain SK, Coppens I, Derré I: Phosphoregulation accommodates Type III secretion and assembly of a tether of ER-. Elife 2022, 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Cortina ME, Derré I: Homologues of the Chlamydia trachomatis and Chlamydia muridarum Inclusion Membrane Protein IncS Are Interchangeable for Early Development but Not for Inclusion Stability in the Late Developmental Cycle. mSphere 2023, 8:e0000323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Elwell CA, Jiang S, Kim JH, Lee A, Wittmann T, Hanada K, Melancon P, Engel JN: Chlamydia trachomatis co-opts GBF1 and CERT to acquire host sphingomyelin for distinct roles during intracellular development. PLoS Pathog 2011, 7:e1002198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Auer D, Hugelschaffer SD, Fischer AB, Rudel T: The chlamydial deubiquitinase Cdu1 supports recruitment of Golgi vesicles to the inclusion. Cell Microbiol 2020, 22:e13136. [DOI] [PubMed] [Google Scholar]
- 42.Pruneda JN, Bastidas RJ, Bertsoulaki E, Swatek KN, Santhanam B, Clague MJ, Valdivia RH, Urbe S, Komander D: A Chlamydia effector combining deubiquitination and acetylation activities induces Golgi fragmentation. Nat Microbiol 2018, 3:1377–1384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wesolowski J, Weber MM, Nawrotek A, Dooley CA, Calderon M, St Croix CM, Hackstadt T, Cherfils J, Paumet F: Chlamydia Hijacks ARF GTPases To Coordinate Microtubule Posttranslational Modifications and Golgi Complex Positioning. mBio 2017, 8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Dimond Z, Bauler LD, Zhang Y, Carmody A, Hackstadt T: Chlamydia trachomatis Alters Mitochondrial Protein Composition and Secretes Effector Proteins That Target Mitochondria. mSphere 2022, 7:e0042322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Steiert B, Icardi CM, Faris R, McCaslin PN, Smith P, Klingelhutz AJ, Yau PM, Weber MM: The Chlamydia trachomatis type III-secreted effector protein CteG induces centrosome amplification through interactions with centrin-2. Proc Natl Acad Sci U S A 2023, 120:e2303487120. * This paper identifies the Chlamydia effector CteG as a key regulator of centrosome amplification during infection, which may contribute to increased risk of gynecological cancer following Chlamydia infection. Additionally, it highlights the importance of using more complex systems, such as primary cells and in vivo experiments, in assessing phenotypes as the importance of CteG to infection is not seen in immortalized cells.
- 46. Luis MP, Pereira IS, Bugalhao JN, Simoes CN, Mota C, Romao MJ, Mota LJ: The Chlamydia trachomatis IncM Protein Interferes with Host Cell Cytokinesis, Centrosome Positioning, and Golgi Distribution and Contributes to the Stability of the Pathogen-Containing Vacuole. Infect Immun 2023, 91:e0040522. * This study demonstrates multifaceted impacts of the Chlamydia effector IncM on multiple host organelles and provides evidence that these diverse impacts may hinge on interactions between IncM and host microtubules. This work highlights the importance of considering multiple roles of individual effectors and provides key mechanistic insights into how one such effector functions.
- 47.Walsh SC, Reitano JR, Dickinson MS, Kutsch M, Hernandez D, Barnes AB, Schott BH, Wang L, Ko DC, Kim SY, et al. : The bacterial effector GarD shields Chlamydia trachomatis inclusions from RNF213-mediated ubiquitylation and destruction. Cell Host Microbe 2022, 30:1671–1684 e1679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Bishop RC, Derré I: The Chlamydia trachomatis Inclusion Membrane Protein CTL0390 Mediates Host Cell Exit via Lysis through STING Activation. Infect Immun 2022, 90:e0019022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Sixt BS, Bastidas RJ, Finethy R, Baxter RM, Carpenter VK, Kroemer G, Coers J, Valdivia RH: The Chlamydia trachomatis Inclusion Membrane Protein CpoS Counteracts STING-Mediated Cellular Surveillance and Suicide Programs. Cell Host Microbe 2017, 21:113–121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Meier K, Jachmann LH, Türköz G, Babu Sait MR, Pérez L, Kepp O, Valdivia RH, Kroemer G, Sixt BS: The Chlamydia effector CpoS modulates the inclusion microenvironment and restricts the interferon response by acting on Rab35. mBio 2023, 14:e0319022. ** This study identifies inclusion membrane protein CpoS as a critical bacterial regulator of proper localization of other inclusion membrane proteins, highlighting the importance of not just effector-host protein interactions but also effector-effector interactions in regulating Chlamydia infection.
- 51. Tang C, Liu C, Maffei B, Niragire B, Cohen H, Kane A, Donnadieu AC, Levy-Zauberman Y, Vernay T, Hugueny J, et al. : Primary ectocervical epithelial cells display lower permissivity to Chlamydia trachomatis than HeLa cells and a globally higher pro-inflammatory profile. Sci Rep 2021, 11:5848. * This paper provides a comparison of differences between primary and immortalized cells in the context of Chlamydia infection, highlighting the importance of selecting a proper cell model for in vitro experiments.
- 52.Fichorova RN, Rheinwald JG, Anderson DJ: Generation of papillomavirus-immortalized cell lines from normal human ectocervical, endocervical, and vaginal epithelium that maintain expression of tissue-specific differentiation proteins. Biol Reprod 1997, 57:847–855. [DOI] [PubMed] [Google Scholar]
- 53.Buckner LR, Schust DJ, Ding J, Nagamatsu T, Beatty W, Chang TL, Greene SJ, Lewis ME, Ruiz B, Holman SL, et al. : Innate immune mediator profiles and their regulation in a novel polarized immortalized epithelial cell model derived from human endocervix. J Reprod Immunol 2011, 92:8–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Buckner LR, Lewis ME, Greene SJ, Foster TP, Quayle AJ: Chlamydia trachomatis infection results in a modest pro-inflammatory cytokine response and a decrease in T cell chemokine secretion in human polarized endocervical epithelial cells. Cytokine 2013, 63:151–165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.N'Gadjaga MD, Perrinet S, Connor MG, Bertolin G, Millot GA, Subtil A: Chlamydia trachomatis development requires both host glycolysis and oxidative phosphorylation but has only minor effects on these pathways. J Biol Chem 2022, 298:102338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Carpenter V, Chen YS, Dolat L, Valdivia RH: The Effector TepP Mediates Recruitment and Activation of Phosphoinositide 3-Kinase on Early Chlamydia trachomatis Vacuoles. mSphere 2017, 2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Dolat L, Carpenter VK, Chen YS, Suzuki M, Smith EP, Kuddar O, Valdivia RH: Chlamydia repurposes the actin-binding protein EPS8 to disassemble epithelial tight junctions and promote infection. Cell Host Microbe 2022, 30:1685–1700 e1610. ** This paper describes a role for the Chlamydia effector TepP in dissembling tight junctions to promote infection. This work specifically demonstrates the utility of combining a variety of models, including immortalized cell culture, organoid culture, and murine infections to fully characterize the importance of Chlamydia genes in host-pathogen interactions at multiple scales.
- 58.Kintner J, Schoborg RV, Wyrick PB, Hall JV: Progesterone antagonizes the positive influence of estrogen on Chlamydia trachomatis serovar E in an Ishikawa/SHT-290 co-culture model. Pathog Dis 2015, 73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Hall JV, Schell M, Dessus-Babus S, Moore CG, Whittimore JD, Sal M, Dill BD, Wyrick PB: The multifaceted role of oestrogen in enhancing Chlamydia trachomatis infection in polarized human endometrial epithelial cells. Cell Microbiol 2011, 13:1183–1199. [DOI] [PubMed] [Google Scholar]
- 60.Kintner J, Moore CG, Whittimore JD, Butler M, Hall JV: Inhibition of Wnt Signaling Pathways Impairs Chlamydia trachomatis Infection in Endometrial Epithelial Cells. Front Cell Infect Microbiol 2017, 7:501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Edwards VL, Smith SB, McComb EJ, Tamarelle J, Ma B, Humphrys MS, Gajer P, Gwilliam K, Schaefer AM, Lai SK, et al. : The Cervicovaginal Microbiota-Host Interaction Modulates Chlamydia trachomatis Infection. mBio 2019, 10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Edwards VL, McComb E, Gleghorn JP, Forney L, Bavoil PM, Ravel J: Three-dimensional models of the cervicovaginal epithelia to study host-microbiome interactions and sexually transmitted infections. Pathog Dis 2022, 80. ** This paper describes a three-dimensional model of the cervicovaginal epithelium, including epithelial cells and fibroblasts along with members of the cervicovaginal microbiota, to study Chlamydia infection. It represents a significant advancement in the in vitro models used within the field.
- 63.Nogueira AT, Braun KM, Carabeo RA: Characterization of the Growth of Chlamydia trachomatis in In Vitro-Generated Stratified Epithelium. Front Cell Infect Microbiol 2017, 7:438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.McQueen BE, Kiatthanapaiboon A, Fulcher ML, Lam M, Patton K, Powell E, Kollipara A, Madden V, Suchland RJ, Wyrick P, et al. : Human Fallopian Tube Epithelial Cell Culture Model To Study Host Responses to Chlamydia trachomatis Infection. Infect Immun 2020, 88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Zadora PK, Chumduri C, Imami K, Berger H, Mi Y, Selbach M, Meyer TF, Gurumurthy RK: Integrated Phosphoproteome and Transcriptome Analysis Reveals Chlamydia-Induced Epithelial-to-Mesenchymal Transition in Host Cells. Cell Rep 2019, 26:1286–1302.e1288. [DOI] [PubMed] [Google Scholar]
- 66.Gurumurthy RK, Koster S, Kumar N, Meyer TF, Chumduri C: Patient-derived and mouse endo-ectocervical organoid generation, genetic manipulation and applications to model infection. Nat Protoc 2022, 17:1658–1690. [DOI] [PubMed] [Google Scholar]
- 67.Koster S, Gurumurthy RK, Kumar N, Prakash PG, Dhanraj J, Bayer S, Berger H, Kurian SM, Drabkina M, Mollenkopf HJ, et al. : Modelling Chlamydia and HPV co-infection in patient-derived ectocervix organoids reveals distinct cellular reprogramming. Nat Commun 2022, 13:1030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Kessler M, Hoffmann K, Fritsche K, Brinkmann V, Mollenkopf HJ, Thieck O, Teixeira da Costa AR, Braicu EI, Sehouli J, Mangler M, et al. : Chronic Chlamydia infection in human organoids increases stemness and promotes age-dependent CpG methylation. Nat Commun 2019, 10:1194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Vollmuth N, Schlicker L, Guo Y, Hovhannisyan P, Janaki-Raman S, Kurmasheva N, Schmitz W, Schulze A, Stelzner K, Rajeeve K, et al. : c-Myc plays a key role in IFN-gamma-induced persistence of Chlamydia trachomatis. Elife 2022, 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Bishop RC, Boretto M, Rutkowski MR, Vankelecom H, Derré I: Murine Endometrial Organoids to Model Chlamydia Infection. Front Cell Infect Microbiol 2020, 10:416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Dolat L, Valdivia RH: An endometrial organoid model of interactions between Chlamydia and epithelial and immune cells. J Cell Sci 2021, 134. ** This paper describes an organoid-neutrophil co-culture system for Chlamydia studies, confirming a role for CPAF and TepP in recruiting neutrophils to sites of infection. This demonstrates the utility of organoids in modeling complex interactions between Chlamydia and various host cells, leading to insights that would not be possible in simpler cell culture models.
- 72.Graspeuntner S, Koethke K, Scholz C, Semmler L, Lupatsii M, Kirchhoff L, Herrmann J, Rox K, Wittstein K, Kading N, et al. : Sorangicin A Is Active against Chlamydia in Cell Culture, Explanted Fallopian Tubes, and Topical In Vivo Treatment. Antibiotics (Basel) 2023, 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Jerchel S, Knebel G, Konig P, Bohlmann MK, Rupp J: A human fallopian tube model for investigation of C. trachomatis infections. J Vis Exp 2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Roth A, Konig P, van Zandbergen G, Klinger M, Hellwig-Burgel T, Daubener W, Bohlmann MK, Rupp J: Hypoxia abrogates antichlamydial properties of IFN-gamma in human fallopian tube cells in vitro and ex vivo. Proc Natl Acad Sci U S A 2010, 107:19502–19507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Hvid M, Baczynska A, Deleuran B, Fedder J, Knudsen HJ, Christiansen G, Birkelund S: Interleukin-1 is the initiator of Fallopian tube destruction during Chlamydia trachomatis infection. Cell Microbiol 2007, 9:2795–2803. [DOI] [PubMed] [Google Scholar]
- 76.Kessler M, Zielecki J, Thieck O, Mollenkopf HJ, Fotopoulou C, Meyer TF: Chlamydia trachomatis disturbs epithelial tissue homeostasis in fallopian tubes via paracrine Wnt signaling. Am J Pathol 2012, 180:186–198. [DOI] [PubMed] [Google Scholar]
- 77.Leung CM, de Haan P, Ronaldson-Bouchard K, Kim G-A, Ko J, Rho HS, Chen Z, Habibovic P, Jeon NL, Takayama S, et al. : A guide to the organ-on-a-chip. Nature Reviews Methods Primers 2022, 2:33. [Google Scholar]
- 78.Thompson CL, Fu S, Knight MM, Thorpe SD: Mechanical Stimulation: A Crucial Element of Organ-on-Chip Models. Front Bioeng Biotechnol 2020, 8:602646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Kim HJ, Li H, Collins JJ, Ingber DE: Contributions of microbiome and mechanical deformation to intestinal bacterial overgrowth and inflammation in a human gut-on-a-chip. Proc Natl Acad Sci U S A 2016, 113:E7–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Si L, Bai H, Rodas M, Cao W, Oh CY, Jiang A, Moller R, Hoagland D, Oishi K, Horiuchi S, et al. : A human-airway-on-a-chip for the rapid identification of candidate antiviral therapeutics and prophylactics. Nat Biomed Eng 2021, 5:815–829. ** This study describes a complex biomimetic device of the airway mucosa that is used for in-depth studies of potential treatments against respiratory viruses. It is an excellent model for the varieties of studies that would be feasible for Chlamydia research in a biomimetic system.
- 81.Jalili-Firoozinezhad S, Gazzaniga FS, Calamari EL, Camacho DM, Fadel CW, Bein A, Swenor B, Nestor B, Cronce MJ, Tovaglieri A, et al. : A complex human gut microbiome cultured in an anaerobic intestine-on-a-chip. Nat Biomed Eng 2019, 3:520–531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Shah P, Fritz JV, Glaab E, Desai MS, Greenhalgh K, Frachet A, Niegowska M, Estes M, Jager C, Seguin-Devaux C, et al. : A microfluidics-based in vitro model of the gastrointestinal human-microbe interface. Nat Commun 2016, 7:11535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Villenave R, Wales SQ, Hamkins-Indik T, Papafragkou E, Weaver JC, Ferrante TC, Bahinski A, Elkins CA, Kulka M, Ingber DE: Human Gut-On-A-Chip Supports Polarized Infection of Coxsackie B1 Virus In Vitro. PLoS One 2017, 12:e0169412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Ortega-Prieto AM, Skelton JK, Wai SN, Large E, Lussignol M, Vizcay-Barrena G, Hughes D, Fleck RA, Thursz M, Catanese MT, et al. : 3D microfluidic liver cultures as a physiological preclinical tool for hepatitis B virus infection. Nat Commun 2018, 9:682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Hashimoto R, Takahashi J, Shirakura K, Funatsu R, Kosugi K, Deguchi S, Yamamoto M, Tsunoda Y, Morita M, Muraoka K, et al. : SARS-CoV-2 disrupts respiratory vascular barriers by suppressing Claudin-5 expression. Sci Adv 2022. 8:eabo6783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Busek M, Aizenshtadt A, Amirola-Martinez M, Delon L, Krauss S: Academic User View: Organ-on-a-Chip Technology. Biosensors (Basel) 2022, 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Yokoi F, Deguchi S, Takayama K: Organ-on-a-chip models for elucidating the cellular biology of infectious diseases. Biochim Biophys Acta Mol Cell Res 2023, 1870:119504. [DOI] [PubMed] [Google Scholar]
- 88.Frances-Herrero E, Lopez R, Hellstrom M, de Miguel-Gomez L, Herraiz S, Brannstrom M, Pellicer A, Cervello I: Bioengineering trends in female reproduction: a systematic review. Hum Reprod Update 2022, 28:798–837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Perry LL, Feilzer K, Caldwell HD: Immunity to Chlamydia trachomatis is mediated by T helper 1 cells through IFN-gamma-dependent and - independent pathways. J Immunol 1997, 158:3344–3352. [PubMed] [Google Scholar]
- 90.Helble JD, Gonzalez RJ, von Andrian UH, Starnbach MN: Gamma Interferon Is Required for. mBio 2020, 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Gondek DC, Olive AJ, Stary G, Starnbach MN: CD4+ T cells are necessary and sufficient to confer protection against Chlamydia trachomatis infection in the murine upper genital tract. J Immunol 2012, 189:2441–2449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Yang Z, Tang L, Shao L, Zhang Y, Zhang T, Schenken R, Valdivia R, Zhong G: The Chlamydia-Secreted Protease CPAF Promotes Chlamydial Survival in the Mouse Lower Genital Tract. Infect Immun 2016, 84:2697–2702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Rajeeve K, Das S, Prusty BK, Rudel T: Chlamydia trachomatis paralyses neutrophils to evade the host innate immune response. Nat Microbiol 2018, 3:824–835. [DOI] [PubMed] [Google Scholar]
- 94.Yang C, Kari L, Sturdevant GL, Song L, Patton MJ, Couch CE, Ilgenfritz JM, Southern TR, Whitmire WM, Briones M, et al. : Chlamydia trachomatis ChxR is a transcriptional regulator of virulence factors that function in in vivo host-pathogen interactions. Pathog Dis 2017, 75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Weber MM, Lam JL, Dooley CA, Noriea NF, Hansen BT, Hoyt FH, Carmody AB, Sturdevant GL, Hackstadt T: Absence of Specific Chlamydia trachomatis Inclusion Membrane Proteins Triggers Premature Inclusion Membrane Lysis and Host Cell Death. Cell Rep 2017, 19:1406–1417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Nelson DE, Virok DP, Wood H, Roshick C, Johnson RM, Whitmire WM, Crane DD, Steele-Mortimer O, Kari L, McClarty G, et al. : Chlamydial IFN-gamma immune evasion is linked to host infection tropism. Proc Natl Acad Sci U S A 2005, 102:10658–10663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Sturdevant GL, Caldwell HD: Innate immunity is sufficient for the clearance of Chlamydia trachomatis from the female mouse genital tract. Pathog Dis 2014, 72:70–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Perry LL, Su H, Feilzer K, Messer R, Hughes S, Whitmire W, Caldwell HD: Differential sensitivity of distinct Chlamydia trachomatis isolates to IFN-gamma-mediated inhibition. J Immunol 1999, 162:3541–3548. [PubMed] [Google Scholar]
- 99.Haldar AK, Piro AS, Finethy R, Espenschied ST, Brown HE, Giebel AM, Frickel EM, Nelson DE, Coers J: Chlamydia trachomatis Is Resistant to Inclusion Ubiquitination and Associated Host Defense in Gamma Interferon-Primed Human Epithelial Cells. mBio 2016, 7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Lijek RS, Helble JD, Olive AJ, Seiger KW, Starnbach MN: Pathology after Chlamydia trachomatis infection is driven by nonprotective immune cells that are distinct from protective populations. Proc Natl Acad Sci U S A 2018, 115:2216–2221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Pal S, Tifrea DF, Zhong G, de la Maza LM: Transcervical Inoculation with Chlamydia trachomatis Induces Infertility in HLA-DR4 Transgenic and Wild-Type Mice. Infect Immun 2018, 86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Liu C, Hufnagel K, O'Connell CM, Goonetilleke N, Mokashi N, Waterboer T, Tollison TS, Peng X, Wiesenfeld HC, Hillier SL, et al. : Reduced Endometrial Ascension and Enhanced Reinfection Associated With Immunoglobulin G Antibodies to Specific Chlamydia trachomatis Proteins in Women at Risk for Chlamydia. J Infect Dis 2022, 225:846–855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Liu C, Mokashi NV, Darville T, Sun X, O'Connell CM, Hufnagel K, Waterboer T, Zheng X: A Machine Learning-Based Analytic Pipeline Applied to Clinical and Serum IgG Immunoproteome Data To Predict Chlamydia trachomatis Genital Tract Ascension and Incident Infection in Women. Microbiol Spectr 2023, 11:e0468922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Poston TB, Lee DE, Darville T, Zhong W, Dong L, O'Connell CM, Wiesenfeld HC, Hillier SL, Sempowski GD, Zheng X: Cervical Cytokines Associated With Chlamydia trachomatis Susceptibility and Protection. J Infect Dis 2019, 220:330–339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105. Zheng X, Zhong W, O'Connell CM, Liu Y, Haggerty CL, Geisler WM, Anyalechi GE, Kirkcaldy RD, Wiesenfeld HC, Hillier SL, et al. : Host Genetic Risk Factors for Chlamydia trachomatis-Related Infertility in Women. J Infect Dis 2021, 224:S64–S71. ** This paper identifies more than 100 human genetic risk factors associated with the development of infertility following Chlamydia infection in a large cohort of women, emphasizing the limitations of in vitro systems in studying phenotypes like infertility that can only be seen in organismal studies, as well as the strength of human cohort studies.
- 106.Zhong W, Kollipara A, Liu Y, Wang Y, O'Connell CM, Poston TB, Yount K, Wiesenfeld HC, Hillier SL, Li Y, et al. : Genetic susceptibility loci for Chlamydia trachomatis endometrial infection influence expression of genes involved in T cell function, tryptophan metabolism and epithelial integrity. Front Immunol 2022, 13:1001255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Mott PD, Taylor CM, Lillis RA, Ardizzone CM, Albritton HL, Luo M, Calabresi KG, Martin DH, Myers L, Quayle AJ: Differences in the Genital Microbiota in Women Who Naturally Clear. Front Cell Infect Microbiol 2021, 11:615770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Wiesenfeld HC, Hillier SL, Krohn MA, Landers DV, Sweet RL: Bacterial vaginosis is a strong predictor of Neisseria gonorrhoeae and Chlamydia trachomatis infection. Clin Infect Dis 2003, 36:663–668. [DOI] [PubMed] [Google Scholar]
- 109.France MT, Ma B, Gajer P, Brown S, Humphrys MS, Holm JB, Waetjen LE, Brotman RM, Ravel J: VALENCIA: a nearest centroid classification method for vaginal microbial communities based on composition. Microbiome 2020, 8:166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Ravel J, Gajer P, Abdo Z, Schneider GM, Koenig SS, McCulle SL, Karlebach S, Gorle R, Russell J, Tacket CO, et al. : Vaginal microbiome of reproductive-age women. Proc Natl Acad Sci U S A 2011, 108 Suppl 1:4680–4687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Mastromarino P, Di Pietro M, Schiavoni G, Nardis C, Gentile M, Sessa R: Effects of vaginal lactobacilli in Chlamydia trachomatis infection. Int J Med Microbiol 2014, 304:654–661. [DOI] [PubMed] [Google Scholar]
- 112.Gong Z, Luna Y, Yu P, Fan H: Lactobacilli inactivate Chlamydia trachomatis through lactic acid but not H2O2. PLoS One 2014, 9:e107758. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
No data were used for the research described in the article.