Abstract
Backgrounds
The resurgence of Anopheles funestus, a dominant vector of human malaria in western Kenya was partly attributed to insecticide resistance. However, evidence on the molecular basis of pyrethroid resistance in western Kenya is limited. Noncoding RNAs (ncRNAs) form a vast class of RNAs that do not code for proteins and are ubiquitous in the insect genome. Here, we demonstrated that multiple ncRNAs could play a potential role in An. funestusresistance to pyrethroid in western Kenya.
Materials and Methods
Anopheles funestus mosquitoes were sampled by aspiration methods in Bungoma, Teso, Siaya, Port Victoria and Kombewa in western Kenya. The F1 progenies were exposed to deltamethrin (0.05%), permethrin (0.75%), DDT (4%) and pirimiphos-methyl (0.25%) following WHO test guidelines. A synergist assay using piperonyl butoxide (PBO) (4%) was conducted to determine cytochrome P450s’ role in pyrethroid resistance. RNA-seq was conducted on a combined pool of specimens that were resistant and unexposed, and the results were compared with those of the FANG susceptible strain. This approach aimed to uncover the molecular mechanisms underlying pyrethroid resistance.
Results
Pyrethroid resistance was observed in all the sites with an average mortality rate of 57.6%. Port Victoria had the highest level of resistance to permethrin (MR=53%) and deltamethrin (MR=11%) pyrethroids. Teso had the lowest level of resistance to permethrin (MR=70%) and deltamethrin (MR=87%). Resistance to DDT was observed only in Kombewa (MR=89%) and Port Victoria (MR=85%). A full susceptibility to P-methyl (0.25%) was observed in all the sites. PBO synergist assay revealed high susceptibility (>98%) to the pyrethroids in all the sites except for Port Victoria (MR=96%, n=100). Whole transcriptomic analysis showed that most of the gene families associated with pyrethroid resistance comprised non-coding RNAs (67%), followed by imipenemase (10%), cytochrome P450s (6%), cuticular proteins (5%), olfactory proteins (4%), glutathione S-transferases (3%), UDP-glycosyltransferases (2%), ATP-binding cassettes (2%) and carboxylesterases(1%).
Conclusions
This study unveils the molecular basis of insecticide resistance in An. funestus in western Kenya, highlighting for the first time the potential role of non-coding RNAs in pyrethroid resistance. Targeting non-coding RNAs for intervention development could help in insecticide resistance management.
Keywords: Anopheles funestus, insecticide resistance, non-coding RNAs, western Kenya, RNA-seq, pyrethroid, PBO, DDT
1.0. Introduction
Vector control particularly the use of bed nets treated with pyrethroids has had an impact on entomological parameters, such as reducing infection rate in the vector population, vector abundance, and parity rate [1, 2] leading to a decline in malaria morbidity and mortality in sub-Saharan Africa as a result of a decline in vectorial capacity [3–5]. Despite these successes, malaria resurgence and outbreaks have been reported in various transmission settings in sub-Saharan Africa where ITNs were deployed [6–9]. Hence, the effectiveness of the primary vector control methods with regard to insecticide resistance needs continuous monitoring and probing of the resistance mechanisms.
In contrast to other major vectors, Anopheles funestus sensu stricto (hereafter An. funestus) has received very scant attention owing to the difficulty in colonizing this species under laboratory conditions. An. funestus is distributed throughout Africa similar to the distributed union of An. gambiae. After developing resistance and exhibiting behavioural adaptability, An. funestus has a higher ability to colonize a niche [10, 11]. It is one of the most ubiquitous and efficient malaria vectors in the world; highly susceptible to the P. falciparum parasite, highly anthropophagic and endophilic [12–14]. The significance of studying this mosquito is highlighted by its versatility in ecological adaptation and the emergence of resistance to recommended public health insecticides for vector control [10, 15].
Increased resistance to pyrethroids used for bed net impregnation has led to low efficacy of conventional LLINs against An. funestus [16]. Resistance monitoring focuses on transmission foci, hotspots of localized outbreaks, or after spikes in disease cases in pre-elimination and elimination settings [17]. For effective insecticide resistance management, it is essential to genetically characterize insecticide resistance profiles and mechanisms in the vector populations. Metabolic resistance poses the biggest threat to the control of malaria vectors [18]. Cytochrome P450s, Glutathione S-transferases (GSTs) and carboxylesterases (COEs) are well-established enzyme families in malaria vectors known to confer resistance to pyrethroids [19, 20]. These detoxification genes are pivotal in the molecular mechanism of insecticide resistance.
Non-coding RNAs (ncRNAs) form a vast class of RNAs that do not code for protein. Examples of ncRNAs include transfer RNA (tRNA), ribosomal RNA (rRNA), small nuclear RNA (snRNA), small nucleolar RNA (snoRNA), microRNA (miRNA), PIWI-interacting RNA (piRNA), endogenous small interfering RNA (siRNA), circular RNA (circRNA), long non-coding RNA (lncRNA), protein functional effector small ncRNA (pfeRNA), and other ncRNAs whose functions remain unknown [21, 22]. They can control the expression of genes at the chromosomal, transcriptional, post-transcriptional, and translational levels and play a role in the entire developmental process. ncRNAs have been demonstrated in studies on arthropods to be essential for several physiological and developmental processes, including molting, reproduction, immunity, wing development, and insecticide resistance [23]. ncRNAs can modify signalling pathways involved in these biological processes by targeting both DNA and RNA substrates. Sequences of regulatory ncRNAs can also help establish epigenetic alterations such as histone acetylation/deacetylation, DNA/histone methylation, etc. within the nucleus by bringing in chromatin remodelling agents that are known to change transcriptional activity [24, 25]. Based on their length, ncRNAs are arbitrarily divided into two groups: small ncRNAs (scnRNAs, < 200 nts) and long ncRNAs (lncRNAs, > 200 nts)[26]. Depending on where they are in relation to genes that code for proteins, lncRNAs can also be categorized as sense, antisense, intronic, or intergenic [27]. With regards to insecticide resistance in insects, lncRNAs that were found to be differentially expressed during the larval stage development of resistant Plutella xylostella genotypes [28] and uniquely differentially expressed during the egg to adult moth stages in Bt-toxin resistant strains of the same insect [29]. Similarly, the expression of the lncRNAs in P. xylostella was linked to the expression of the cytochrome P450, the ATP-binding cassette (ABC) transporter and the esterase genes involved in resistance to chlorantraniliprole insecticide [27]. Moreover, some long intergenic non-coding RNAs were overexpressed in deltamethrin-resistant larvae of Plutella xylostella exposed to deltamethrin [28]. ncRNAs are intriguing candidates to study when organisms are exposed to insecticides and other toxicants since they are involved in pathways linked to responses to cellular stress [28, 30]. The genes for ribosomal proteins, such as L39 [31], S4 [32], L22 [33], and S29 [34], have been found to be associated with the resistance mechanism of Culex mosquitoes.
In the malaria-endemic region of western Kenya, there has been a resurgence of endophilic An. funestus and increased 20-fold over a decade ago [9, 35]. The resurgence of this vector was partly attributed to resistance to pyrethroids used in ITN impregnation [36]. As the country is aiming to achieve the malaria elimination goal by 2030, it is very crucial to have a comprehensive understanding of the resistant profile of this important, re-emerged vector to inform stakeholders of the right choice of control strategy to adopt. To date, there have been few investigations on An. funestus susceptibility to insecticides in Kenya. The initial study on An. funestus susceptibility to insecticides from two study areas in western Kenya was reported in 2007 [37] and even though the species were not identified using molecular techniques, previously identified Anopheles species from the same areas revealed that only An. funestus was present [38]. Later in western Kenya, seven adults An. funestus were sampled and their F1 progenies’ susceptibility to insecticides revealed that they were susceptible to DDT but resistant to permethrin [39]. Further study in Kisumu in the lowland area of western Kenya has shown that An. funestus is resistant to pyrethroids (deltamethrin and permethrin) with overexpression CYP6P9a and CYP6P9b responsible for pyrethroid resistance [15]. A recent study in the same Kisumu, using microarray for transcriptome analysis has revealed that overexpression of cytochrome P450s notably, CYP4H18, CYP6M7, CYP9K1, CYP4C36 and CYP4H17 in pyrethroid-resistant An. funestus population [40]. The use of microarrays can only be used with the gene families that have been identified on the array, and they only give information on relative expression levels. The RNA-seq technology offers single nucleotide level resolution, absolute rather than relative gene expression profile, and a comprehensive view of the transcriptome in a specific state [41].
In this study, we examined the insecticide resistance profile of An. funestus across five sites in four counties in western Kenya and elucidated the molecular mechanisms of resistance using RNA-seq. Our results provide new novel insights into insecticide resistance at the molecular level in this important malaria vector, which has received limited attention, and could help in designing effective control strategies.
2.0. Materials and Methods
2.1. Sampling of indoor-resting Anopheles mosquitoes
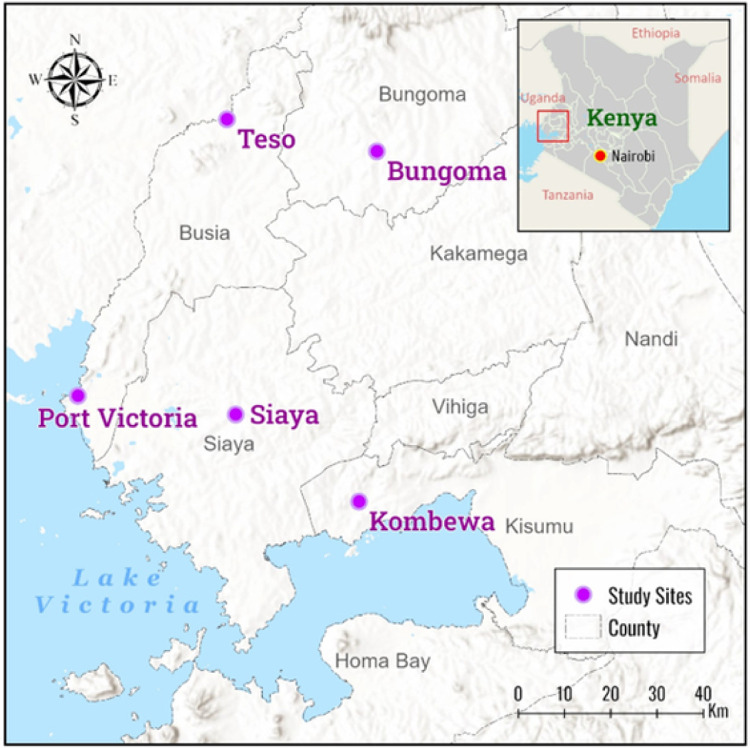
Anopheles mosquitoes were sampled from five sites: Bungoma [00.54057°N, 034.56410°E, 1386–1,545m above sea level (asl)] (highland), Teso (0°43’0” N. 34°21’0” E, 1357–1,500m asl) (highland), Siaya (0.0626° N, 34.2878° E, 1,140–1,400m asl) (lowland), Port Victoria (0° 6’ 0” N / 33° 58’ 0” E, 1,149 asl) (lowland) and Kombewa (00 07’N, 340 30’E, 1150–1300 m asl) (lowland) in western Kenya (Fig. 1). These sites are malaria-endemic areas predominated by An. funestus mosquitoes. Adult An. funestus population were sampled from the indoor living room using mouth and prokopack aspiration methods after informed consent was sought and provided by the owners of the households.
Figure 1. Map of study sites where mosquitoes were sampled in western Kenya.
The software ArcGIS Pro 2.6 was used to create the map. Map sources: USGS, ESRI, and CGIAR (www.esri.com)
2.2. Mosquito sorting and identification
Live mosquitoes were sorted by separating male mosquitoes from the females and Culex spp from Anopheles. Later Anopheles mosquitoes were morphologically identified as An. funestus s.l and An. gambiae s.l following morphological and taxonomic keys [42, 43].
2.3. Raising of F1 progenies.
An. funestus blood-fed, gravid and half gravid were put into cages to lay eggs on wet filter papers. The F0 females in these physiological states were fed on 10% sugar solutions soaked in cotton wool and laying pads/Petri dishes. After laying the eggs, the eggs were allowed to hatch into larvae. The larvae were put in a pan containing spring water and were fed with the larvae feed, tetramin until they matured to a pupal stage where they were transferred into cages to emerge into adults.
2.4. Insecticide susceptibility tests
The F1 adult female mosquitoes aged between 3–5 days old were used for the bioassay. The insecticide susceptibility test was carried out following the standard insecticide tube test method developed by the WHO [17]. The mortality was scored 24 hours post-exposure after maintaining under standard laboratory conditions at a temperature of 27 °C ± 2 °C and a relative humidity of 75% ± 10%.
2.5. PBO synergist bioassays
After establishing pyrethroid resistance in the An. funestus population, a synergist bioassay was conducted with PBO-impregnated papers to determine the role of P450 monooxygenases in pyrethroid resistance. The PBO inhibits these enzymes’ activity in insects including mosquitoes. The female mosquito samples (F1 progenies) were pre-exposed to 4% PBO for 1 hour before they were immediately exposed to the pyrethroids (0.05% deltamethrin and 0.75% permethrin).
2.6. Preparation of samples for molecular and transcriptome analysis
Surviving resistant An. funestus samples after exposure to the insecticides (permethrin and deltamethrin) and unexposed (An. funestus F1 progenies samples that were not exposed to any insecticides) were killed immediately by keeping them in a deep freezer for about 10 minutes until they were completely knockdown. Samples were immediately stored in 0.5 ml Eppendorf tubes with RNALater and were immediately frozen at −80°C for subsequent molecular and whole transcriptome analysis.
2.7. DNA extraction and molecular identification of species
DNA was extracted from the legs of each stored mosquito specimen using the Chelex®-100 method [44] and was transferred into pre-labelled 1.5ml storage vials and stored at −20°C for molecular analysis. An. funestus-specific PCR was conducted to confirm species using the species-specific primers (ITS2A/FUN) in the internal transcribed spacer region (ITS2) on the ribosomal DNA [45, 46]. Species-specific primers for An. funestus (5 − GCATCGATGGGTT ∀TCATG − 3) and universal primer (5 – TGTG ∀CTGCAGGACACAT − 3) were used. A final volume of 12.5 μl of PCR mixture containing 1μl of genomic DNA, 6.5 μl DreamTaq Green PCR Master Mix (2x), 0.5 μl of each of the primers and 4.0 μl of PCR water. Genomic DNA amplification was performed using a T100 thermal cycler (Biorad). The PCR conditions include initial denaturation at 95 °C for 3 seconds, denaturation of 94 °C for 30 seconds, annealing at 55 °C for 30 seconds for 34 cycles, extension at 72 °C for 45 seconds and final extension at 72 °C for 6 seconds. The DNA bands were visualized using the agarose gel electrophoresis.
2.8. RNA extraction
Total RNA was extracted from a pool of ten mosquitoes for each group (pyrethroids-resistant group and unexposed to pyrethroids). Total RNA was isolated and purified from the whole mosquito using the ZYMO Quick-RNA miniprep kit [47]. The details of the pooled samples is shown in Table 1.
Table 1.
RNA-seq read filtering and mapping statistics.
| Sample ID | Site | Phenotype | Pool | Raw reads | Clean reads | Mapped reads | % Mapped |
|---|---|---|---|---|---|---|---|
| Kr01 | Kombewa | Resistant | 10 mosquitoes | 69034333 | 68982973 | 68982973 | 100% |
| Kr02 | Kombewa | Resistant | 10 mosquitoes | 66278265 | 66237148 | 66237148 | 100% |
| Kr03 | Kombewa | Resistant | 10 mosquitoes | 82298618 | 82251274 | 82251274 | 100% |
| Ku07 | Kombewa | Unexposed | 10 mosquitoes | 68512143 | 68463374 | 68463374 | 100% |
| Pr08 | Port Victoria | Resistant | 10 mosquitoes | 81542127 | 81505195 | 81505195 | 100% |
| Sr10 | Siaya | Resistant | 10 mosquitoes | 96640964 | 96603256 | 96603256 | 100% |
| Su12 | Siaya | Unexposed | 10 mosquitoes | 72322576 | 72294283 | 72294283 | 100% |
| Tr13 | Teso | Resistant | 10 mosquitoes | 92317683 | 92276651 | 92276651 | 100% |
| ERR981209 | FANG colony | Susceptible | 61161962 | 60881678 | 60881678 | 100% | |
| ERR981210 | FANG colony | Susceptible | 44384316 | 44042916 | 44042916 | 100% | |
| ERR981211 | FANG colony | Susceptible | 55329478 | 55063614 | 55063614 | 100% |
2.9. cDNA library preparation and RNA Sequencing
The quality of each total RNA sample was assessed using High Sensitivity RNA Tapestation (Agilent Technologies Inc., California, USA) and quantified with Qubit 3.0 RNA HS assay (ThermoFisher, Massachusetts, USA). Ribosomal RNA was depleted with Ribo-Zero Plus rRNA Removal Kit (Illumina Inc., California, USA). Samples were then heated, fragmented and randomly primed according to the manufacturer’s recommendation. The first strand was synthesized with the Protoscript II Reverse Transcriptase with a longer extension period, approximately 30 minutes at 42 C. All remaining steps for library construction were performed according to the NEBNext® Ultra™ II Directional RNA Library Prep Kit for Illumina® (New England BioLabs Inc., Massachusetts, USA). Final libraries quantity was assessed by Qubit 3.0 (ThermoFisher, Massachusetts, USA) and quality was assessed by TapeStation D1000 ScreenTape (Agilent Technologies Inc., California, USA). The final library size was about 350bp with an insert size of about 200bp. Illumina® 8-nt dual indices were used. Equimolar pooling of libraries was performed based on QC values and sequenced on an Illumina® NovaSeq platform (Illumina, California, USA) with a read length configuration of 150 PE for [120M PE] reads per sample (60M in each direction).
2.10. Bioassay data analysis
The mortality rate of the sample tested was expressed as the total number of dead An. funestus mosquitoes of all the replicates exposed to a particular insecticide and expressed this as the percentage of all the population exposed to that insecticide. Abbott’s formula was used to correct mortality if the mortality at 24 hours in the control tube was between 5% and 20%. Following the WHO criteria [17] for determining insecticide resistance in the malaria vector population, a population is classified as susceptible when the mortality is between 98–100%, resistant when mortality is less than 90% and suspected resistance when mortality is between 90–97%.
2.11. Quality control and differential expression analysis
Upon obtaining paired-end sequence reads from the sequencing centre (range 46,981,760–96640964 total reads), they were checked for quality using FASTQC (v0.11.5) [48] and cleaned to remove adapters (Table 1). Trimmomatic module (v.0.39) [49] was used to remove the Illumina adapters (TruSeq3-PE-2) that were used to construct the library, which resulted in the selection of reads that were more than 50 bp and a Phred-Quality-Score greater than 20 for downstream analysis. The resulting reads were confirmed to be of acceptable quality by running FASTQC (v0.11.3) before they were aligned to the reference genome. The Anopheles funestus FUMOZ genome from VectorBase (AfunF3.53) was used as the reference genome and this was aligned using the HISAT2 v2.2.1, which involved building the HISAT2 index file for the genome [50]. To reduce the size of the output SAM tools from the alignment output, they were converted to BAM files, sorted, and indexed using SAMtools v1.10 (Li et al., 2009). The sorted and indexed files were used as input for the Htseq-count reads, which were created using the module htseq-count (v.0.6.1) as described [52]. A reference gene transfer format was used to count the number of alignment mapping to each gene based on union and intersection-strict [52]. Gene expression values were normalized using Relative Log Expression (RLE) from DESeq2. Expression abundance between different treatments (pyrethroids-resistant group vs pyrethroid susceptible FANG colony of An. funestus mosquitoes [53] and unexposed vs the susceptible FANG colony) for the study sites (Teso, Port Victoria, Siaya and Kombewa) was determined using DESeq2 (v.1.18.0) [54]. The susceptible FANG colony raw sequence data was retrieved from the GenBank (accession: ERR981209–ERR981211). A correlation of gene expression between biological replicates was calculated by Pearson’s correlation as suggested before [55], while the Benjamini-Hochberg method was applied in calibrating the p-value to decrease chances of false positives [56]. Therefore, differential expression between treatment groups was considered significant if the p-value < 0.05 and the fold change (FC) > 1.5 [57]. Hierarchical clustering analysis was applied to cluster genes exhibiting similar expression patterns/levels, while the Gene Ontology (GO) from the GO database (http://geneontology.org/) was utilized for functional analysis of the differentially expressed genes to establish their biological profiles (52).
3.0. Results
3.1. Phenotypic resistance profile in western Kenya
Pyrethroid resistance was observed in all the sites with an average mortality rate (MR) of 57.6%. Port Victoria had the highest level of resistance to permethrin (MR = 53%) and deltamethrin (MR = 11%) pyrethroids. Teso had the lowest level of resistance to permethrin (MR = 70%) and deltamethrin (MR = 87%). Resistance to DDT was observed only in Kombewa (MR = 89%) and Port Victoria (MR = 85%). However, after samples were pre-exposed to the synergist, PBO, high susceptibility (> 98%) to the pyrethroids (deltamethrin and permethrin) was observed in all the sites except for Port Victoria where suspected resistance (96%) was observed for PBO + deltamethrin. In addition to the pyrethroid resistance, resistance to DDT was observed in Kombewa (89%) and Port Victoria (85%) (Table 2). Notwithstanding, a suspected resistance to DDT was observed in Siaya (93%) and Teso (92%). An. funestus was, however, fully susceptible to pirimiphos- methyl (0.25%) in all the sites.
Table 2.
Mortality rate of An. funestus exposed to different insecticides and synergists (24-hr post-exposure).
| Study site | Type and % of insecticide/chemical used | N | % Mortality (24hr) |
|---|---|---|---|
| Kombewa | Permethrin (0.75%) | 280 | 54 |
| Permethrin (0.75%) + PBO (4%) | 300 | 99 | |
| Deltamethrin (0.05%) | 180 | 59 | |
| Deltamethrin (0.05%) + PBO (4%) | 100 | 100 | |
| DDT (4%) | 100 | 89 | |
| Pirimiphos methyl (0.25%) | 180 | 99 | |
| Siaya | Permethrin (0.75%) | 100 | 78 |
| Deltamethrin (0.05%) | 133 | 52 | |
| 0.75% permethrin + PBO (4%) | 100 | 100 | |
| Deltamethrin (0.05%) + PBO (4%) | 100 | 100 | |
| DDT (4%) | 100 | 93 | |
| Pirimiphos methyl (0.25%) | 100 | 100 | |
| Teso | Deltamethrin (0.05%) | 100 | 70 |
| Permethrin (0.75%) | 100 | 87 | |
| 0.75% permethrin + 4% PBO | 100 | 100 | |
| Deltamethrin (0.05%) + PBO (4%) | 100 | 100 | |
| DDT (4%) | 300 | 92 | |
| Pirimiphos methyl (0.25%) | 100 | 100 | |
| Port Victoria | Permethrin (0.75%) | 217 | 53 |
| Deltamethrin (0.05%) | 100 | 11 | |
| Deltamethrin (0.05%) + PBO (4%) | 100 | 96 | |
| DDT (4%) | 100 | 85 | |
| Bungoma | Permethrin (0.75%) | 100 | 69 |
| Deltamethrin (0.05%) | 100 | 43 |
N: number of mosquitoes exposed to the insecticide
3.2. Differentially expressed genes between groups.
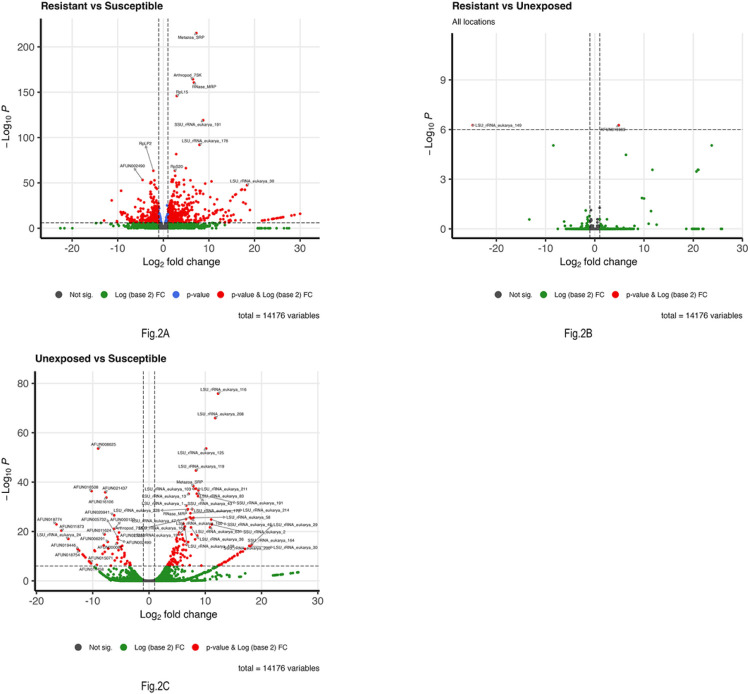
After quality control and the elimination of genes with low read counts, differential expression analysis was carried out on the transcripts. Three pairwise comparisons were performed: resistant versus susceptible, resistant versus unexposed (control) and unexposed versus susceptible. The resistant versus unexposed comparison helps to account for the induction of transcription during the pyrethroid exposure; genes were filtered by analyzing their expression profiles in the susceptible An. funestus population, under the assumption that constitutive resistance genes will be significantly differentially expressed between both survivors of the bioassay and the unexposed field F1 progenies when compared to the susceptible FANG colony. The volcano plots (Fig. 2) showed the downregulation and upregulation of the genes between resistant vs susceptible (Fig. 2A), resistant vs unexposed (Fig. 2B) and unexposed vs susceptible (Fig. 2C). There was a clear distinction between overexpression of genes in resistant vs susceptible (Fig. 2A) and unexposed vs susceptible (Fig. 2C) group comparisons. However, there was no difference in the gene overexpression between the resistant and unexposed group comparisons.
Figure 2. Volcano plot indicating upregulation and downregulation for resistant vs susceptible (A), resistant vs unexposed (B) and unexposed (control) vs susceptible (C).
The X-axis indicates the log2 fold-change- positive and negative values are up and down-regulated respectively relative to the susceptible group in A and C. The Y-axis indicates −log10 of the adjusted P-value (FDR) (−log10FDR values >200 for A, > 9 for B and > 80 for C). In each volcano plot, genes that are overexpressed in the population are >0 on the x-axis. P-values of < 0.05 are indicated by the horizontal line, while 2-fold expression differences are indicated by vertical dotted lines.
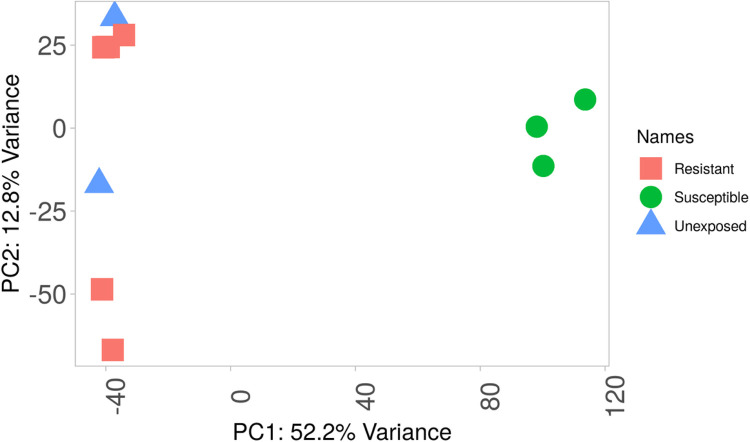
To summarize the expression pattern of genes in the sample groups, principal component analysis (PCA) and heatmap were used. The PCA plots indicate a representation of differences in the sample groups (resistant, unexposed and susceptible). The samples in the resistant and unexposed groups clustered together to the left-hand side away from the susceptible counterparts indicating the similarity between them (Fig. 3). The susceptible FANG group clusters towards the left side away from the resistant and the unexposed groups (Fig. 3).
Figure 3.
A principal component analysis showing the gene expression pattern of the sample groups relative to the susceptible group.
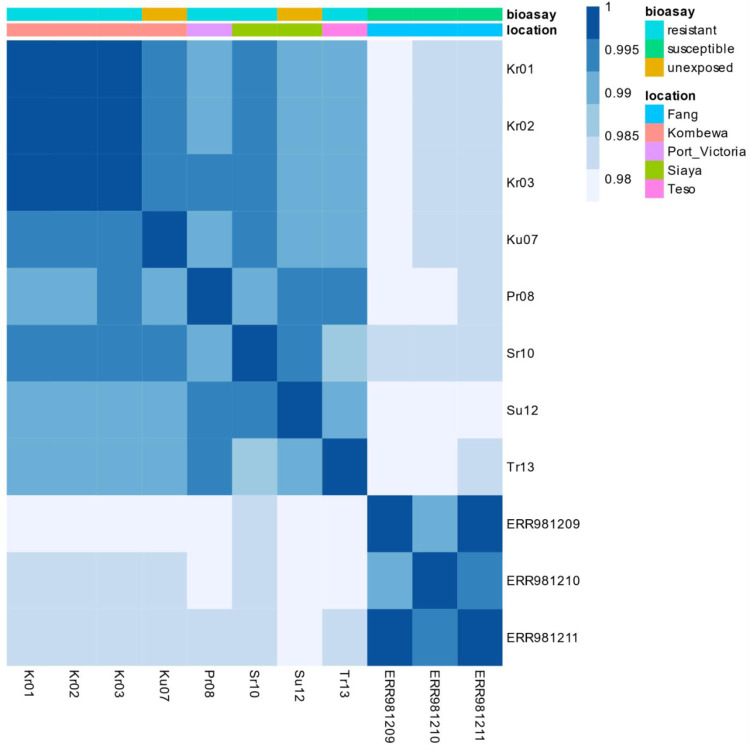
The heatmap revealed that there was an obvious grouping of the samples into resistant, unexposed and susceptible. Dissimilarity in the gene expression levels was noticed between the groups. The overall gene expression profile indicates a higher level of expression in the resistant and unexposed sample groups compared to the susceptible (Fig. 4). Moreover, most of the genes were highly expressed in the Kombewa-resistant (Kr01, Kr02 and Kr03) samples. This was followed by the resistant samples from Port Victoria, Siaya and Teso. However, low levels of gene expression were observed in the susceptible samples (ERR981209, ERR981210 and ERR981211).
Figure 4.
Heatmap indicating the expression of genes in the sample groups relative to the susceptible group.
Comparison using the Venn diagrams, 33 genes (n = 14176) were differentially expressed in all the comparisons [resistant vs susceptible (R-S), resistant vs unexposed (R-C) and vs unexposed and susceptible (C-S)] (Fig. 5A). However, 953, 35 and 455 common genes were differentially expressed in only R-S, R-C and C-S respectively. More genes (1597) were significantly differentially expressed between R-S and C-S comparisons compared to the other comparisons. This was followed by 87 differentially expressed genes observed between C-S and R-C comparisons and 43 differentially expressed genes between R-S and R-C comparisons (Fig. 5A). Most of the downregulated genes were found in the R-S and this was followed by C-S (Fig. 5B). Nine Hundred and forty-nine (949) genes were downregulated between R-S and C-S comparisons (Fig. 5B).
Figure 5. Venn diagram comparing upregulated and downregulated genes between-group comparisons.
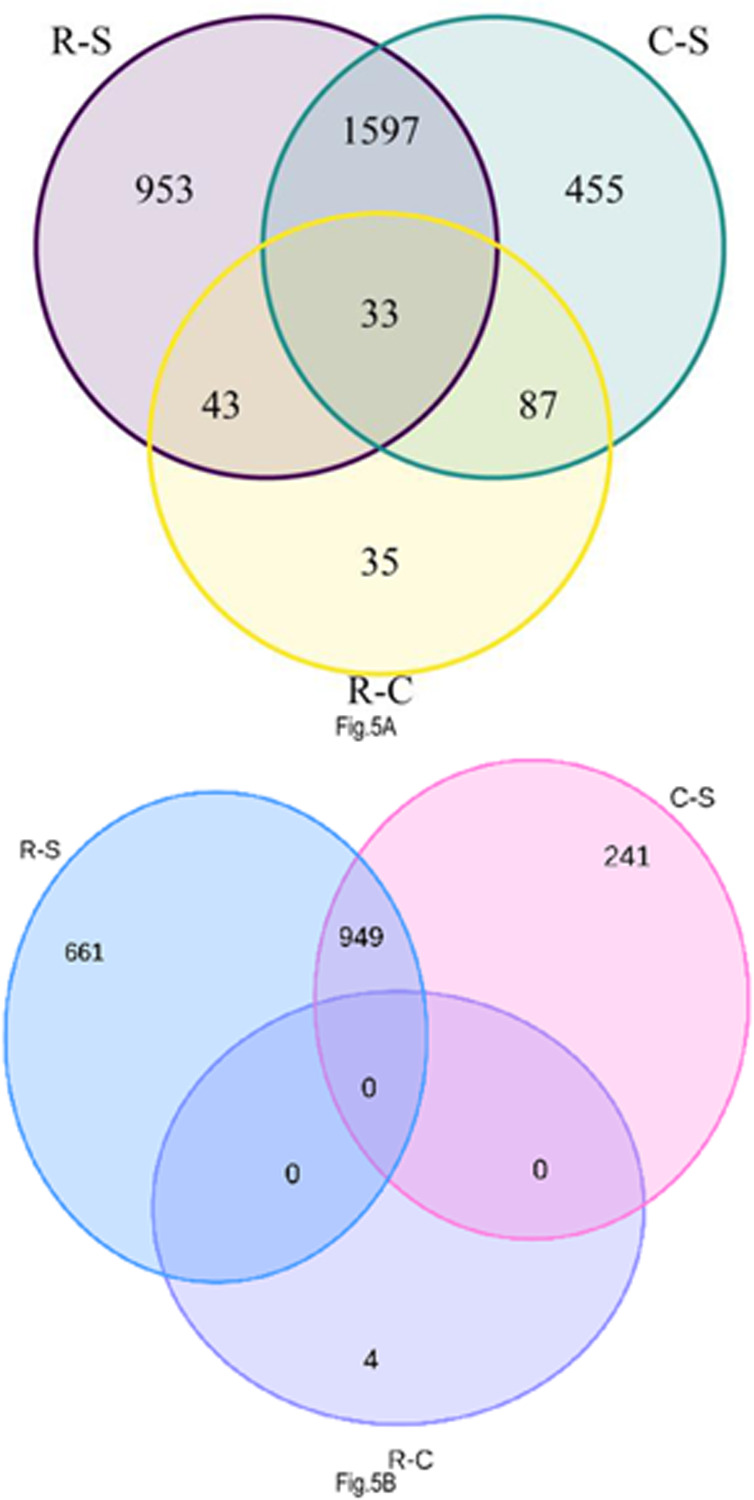
A indicates upregulated genes between the groups and B indicates downregulated genes between groups. R-S: field-resistant population that survived pyrethroid exposure vs susceptible colony, R-C: field-resistant population that survived pyrethroid exposure vs unexposed (control) field population and C-S: unexposed (control) field population vs susceptible colony.
3.3. Differentially expressed ncRNAs linked to pyrethroid Resistance.
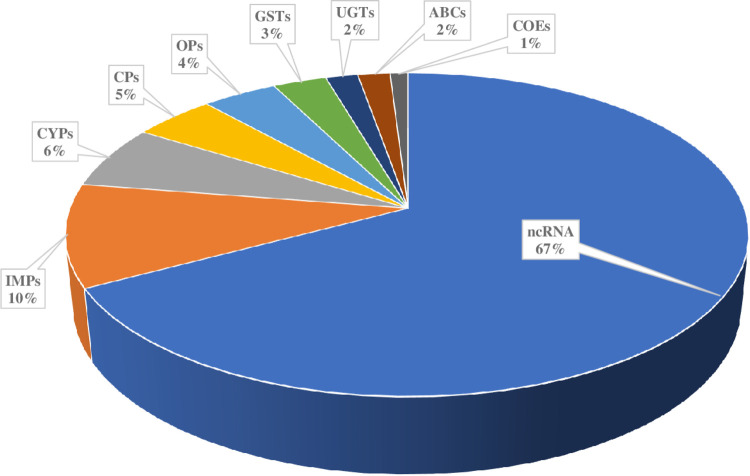
Differentially expressed ncRNAs between resistant vs susceptible (R-S) and unexposed vs susceptible (C-S) were determined by a fold change FC > 1.5 and FDR < 0.05. The whole transcriptome analysis shows that ncRNAs constituted 67%, the highest proportion of the gene families involved in pyrethroid resistance (Fig. 6). This was followed by IMPs (10%), CYPs (6%), CPs (5%), OPs (4%), GSTs (3%), UGTs (2%), ABCs (2%) and COEs (1%) (Fig. 6).
Figure 6. Pie chart showing the proportion of gene family involving pyrethroid resistance.
IMPs: Imipenemase, CYPs: Cytochrome P450s, CPs: cuticular proteins, OPs: olfactory proteins, GSTs: Glutathione S-transferases, UGTs: UDP-glycosyltransferases, ABCs:ATP-binding cassettes, COEs: carboxylesterases and ncRNA: non-coding RNA
The main ncRNAs that were overexpressed are in the resistant vs susceptible (R-S) and unexposed vs susceptible (C-S) are Metazoa_SR, RNaseP_nu, U3_1, Arthropod_7S, LSU_rRNA_eukarya_, SSU_rRNA_eukarya_2, LSU_rRNA_eukarya_13, SSU_rRNA_eukarya_46, LSU_rRNA_eukarya_2, LSU_rRNA_eukarya_3, SSU_rRNA_eukarya_15, LSU_rRNA_eukarya_5, LSU_rRNA_eukarya_6, SSU_rRNA_eukarya_164, SSU_rRNA_eukarya_19, SSU_rRNA_eukarya_200, LSU_rRNA_eukarya_155, LSU_rRNA_eukarya_17, LSU_rRNA_eukarya_17, LSU_rRNA_eukarya_214 and RNase_MRP (Table 3).
3.4. Differentially expressed metabolic genes associated with pyrethroid resistance.
Similarly, to identify the main genes in the enzyme families responsible for high pyrethroid metabolic resistance, FDR < 0.05 and a fold change FC > 1.5 were used. The main enzyme families identified are the cytochrome P450s, GSTs, salivary gland proteins, Peptidase S1 domain-containing proteins, UGTs and sulfotransferases (Table 4). However, most of these genes were moderately differentially expressed. The findings indicate that in western Kenya, different genes within these enzyme families were responsible for resistance (Table 4). The top cytochrome P450 enzymes are moderately overexpressed in the An. funestus in western Kenya were CYP6P9, CYP6P9, CYP6N, CYP6N, CYP9J, CYP49A, CYP6P, AFUN02089, AFUN01936, CYP9K, CYP304B. These genes were overexpressed in resistant vs susceptible and unexposed/control vs susceptible group comparisons. However, CYP304C1 and CYP315A1 were overexpressed only in the resistant vs susceptible comparison (Table 4). Among the GSTs, the overexpressed genes in resistant vs susceptible and unexposed/control vs susceptible groups are GSTD, GSTT, GSTE, GSTD, and GSTD3 were overexpressed only in the resistant vs susceptible comparison (Table 4). AFUN02142, AFUN021428 and AFUN019106 were the only cuticular proteins that were overexpressed in the resistant vs susceptible comparison. The differential expression analyses revealed that some of these UGTs were overexpressed in the resistant vs susceptible and unexposed/control vs susceptible groups comparisons (Table 4). These include UGT302A, UGT310B, UGT308D, UGT306A3 and AFUN003620. AFUN016205 and AFUN016207 were the sulfotransferases that were overexpressed in the An. funestus population from western Kenya (Table 4). The summary of the RNA-seq data set for the FC and P-values of each gene is presented in Additional file 1.
Table 4.
List of the top genes of immunity, metabolic, cuticle and olfactory
| Gene ID | Symbol | Chr. | FC (R vs S) | FC (C vs S) | Resistant (read count) | Unexposed (read count) | Susceptible (read count) | Group |
|---|---|---|---|---|---|---|---|---|
| AFUN008117 | AFUN008117 | 2 | 2.4 | 2.1 | 120.9 | 145.2 | 178.5 | Cytochrome |
| AFUN015889 | CYP6P9b | 2 | 6.2 | 4.7 | 946.4 | 1423.4 | 1519.4 | Cytochrome |
| AFUN015792 | CYP6P9A | 2 | 3.5 | 2.5 | 568.3 | 893.9 | 925.1 | Cytochrome |
| AFUN010918 | CYP6N1 | 2 | 3.4 | 3.9 | 341.5 | 306.3 | 726.8 | Cytochrome |
| AFUN001383 | CYP9J5 | 3 | 2.1 | 2 | 215.7 | 469.7 | 293.4 | Cytochrome |
| AFUN015735 | CYP49A1 | 3 | 3.4 | 2.9 | 160.5 | 113.2 | 218.2 | Cytochrome |
| AFUN005715 | CYP315A1 | X | 2.3 | NS | 138.6 | 167.1 | 168.2 | Cytochrome |
| AFUN015888 | CYP6P5 | 2 | 6.3 | 9.5 | 129.3 | 220.2 | 288.1 | Cytochrome |
| AFUN020895 | AFUN020895 | 2 | 6.7 | 5.6 | 394.7 | 604.3 | 506.2 | Cytochrome |
| AFUN019365 | AFUN019365 | 2 | 10.4 | 10.7 | 267.8 | 333.2 | 412.2 | Cytochrome |
| AFUN007549 | CYP9K1 | X | 10.5 | 10.5 | 3310.9 | 5502.8 | 4501.7 | Cytochrome |
| AFUN015938 | CYP9M1 | 2 | 2.1 | NS | 384.6 | 452.2 | 414.9 | Cytochrome |
| AFUN015956 | CYP304B1 | 2 | 4.5 | 7 | 261.2 | 109.1 | 373.3 | Cytochrome |
| AFUN015957 | CYP304C1 | 2 | 2.5 | NS | 230.7 | 341.1 | 241.2 | Cytochrome |
| AFUN016010 | GSTD1 | 2 | 3.4 | 3.3 | 4552.7 | 4424.7 | 5215.8 | Glutathione s-transferase |
| AFUN007291 | GSTT2 | X | 2.5 | 2.2 | 143.4 | 116.3 | 164.3 | Glutathione s-transferase |
| AFUN011410 | GSTD7 | 2 | 2.1 | NS | 143 | 128.1 | 214.9 | Glutathione s-transferase |
| AFUN015767 | GSTD11 | 2 | 2.7 | NS | 13.3 | 20.1 | 20.3 | Glutathione s-transferase |
| AFUN015839 | GSTD3 | 2 | 2.8 | NS | 241.9 | 515.5 | 281.2 | Glutathione s-transferase |
| AFUN016008 | GSTE6 | 2 | 9.7 | 6.6 | 99.1 | 122.4 | 106.9 | Glutathione s-transferase |
| AFUN004194 | Or42 | 2 | 2.5 | NS | 6 | 15 | 13.7 | Odorant receptor |
| AFUN018482 | AFUN018482 | 3 | 7.5 | 9.9 | 9.3 | 0 | 4.3 | Peptidase S1 domain-containing protein |
| AFUN018981 | AFUN018981 | 3 | 3.1 | 5.3 | 9 | 26.1 | 22 | Peptidase S1 domain-containing protein |
| AFUN018580 | AFUN018580 | 3 | 11.5 | 11 | 210.4 | 127.4 | 212.8 | Peptidase S1 domain-containing protein |
| AFUN019220 | AFUN019220 | 2 | 5.2 | 5.7 | 390.5 | 458.9 | 577.3 | ABC transporter |
| AFUN015896 | AFUN015896 | 2 | 2.1 | NS | 142.7 | 112 | 151.9 | CLIP-domain serine protease |
| AFUN021427 | AFUN021427 | 2 | 2.3 | NS | 11.6 | 11 | 8.3 | Cuticular protein |
| AFUN021428 | AFUN021428 | 2 | 2.7 | NS | 15.6 | 7 | 8.3 | Cuticular protein |
| AFUN019106 | AFUN019106 | 3 | 3.1 | NS | 8 | 8.7 | 20.4 | Cuticular protein |
| AFUN019845 | UGT302A3 | 3 | 3.2 | 2 | 421.3 | 681.5 | 634.9 | UDP-glycosyltransferases |
| AFUN011266 | UGT310B2 | 2 | NS | 3.1 | 16.7 | 16 | 38.6 | UDP-glycosyltransferases |
| AFUN020198 | UGT308D2 | 3 | 2 | NS | 22.3 | 46 | 49.1 | UDP-glycosyltransferases |
| AFUN016302 | UGT306A3 | 3 | NS | 2 | 279.6 | 318.2 | 391.4 | UDP-glycosyltransferases |
| AFUN003620 | 2 | 2 | 2.1 | 320.2 | 325.8 | 437.8 | UDP-glycosyltransferases | |
| AFUN016205 | 3 | NS | 3.8 | 250.6 | 123.4 | 403.2 | sulfotransferase | |
| AFUN016207 | 3 | 2.1 | 2.4 | 170.8 | 193.5 | 234.9 | sulfotransferase |
R: resistant field mosquito population that survived the pyrethroid exposure, S: susceptible FANG colony, C: unexposed/control field mosquito population, FC: fold change, NS: not significant
3.5. Gene ontology analysis of the differentially expressed genes
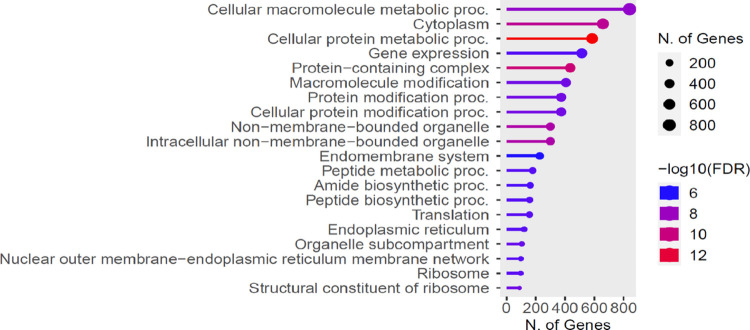
GO term annotation pathways analysis was employed to elucidate the biological functions and signalling pathways that may be regulated by the differentially expressed genes in An. funestus. Our findings revealed that these genes were engaged in a wide variety of biological functions and signalling pathways. Detailed GO enrichment for the differentially expressed genes in ontologies of cellular components, biological processes, and molecular function is represented in Fig. 7. By GO annotation, the differentially expressed genes in pyrethroid-resistant An. funestus were enriched mostly in cellular macromolecule metabolic processes, cytoplasm, cellular protein metabolic processes and gene expression (Fig. 7).
Figure 7. Gene Ontology (GO) enrichment analysis of the differentially expressed genes.
The x-axis indicates the gene count/number of genes while the y-axis indicates the enriched terms. The colour is used to distinguish at different levels.
4.0. Discussion
The successful implementation and development of insecticide resistance management measures depends on elucidating the mechanisms underlying resistance in malaria vectors. In this study, we have characterized the phenotypic resistance profile of An. funestus and the molecular basis of pyrethroid resistance in western Kenya. This is one of the most comprehensive studies on the An. funestus susceptibility status to pyrethroids and DDT in western Kenya.
Our study revealed a high level of pyrethroid resistance across western Kenya although resistance levels vary from site to site. In addition, resistance to DDT has been detected in Kombewa and Port Victoria. This confirmed a previous study in East Africa including western Kenya which reported widespread pyrethroid resistance in the An. funestus population [15]. The rise of multiple resistance of An. funestus was also confirmed in a previous study in western Kenya [15], Benin, west Africa [59] and Malawi, southern Africa [60]. An. funestus was, however, fully susceptible to pirimiphos methyl, the organophosphate in all the study sites. This is congruent with a previous study in Tanzania where a full susceptibility of this vector to pirimiphos methyl in Tanzania was reported [12]. This is an indication that this insecticide can still be maintained for IRS programs in western Kenya. The preexposure of samples to the synergist, PBO has shown that An. funestus was fully susceptible to the pyrethroids in all the study sites except Port Victoria where 96% mortality was observed for the PBO + deltamethrin. This implies that the metabolic resistance mechanism (cytochrome P450 monoxygenases) was fully involved in insecticide resistance in the An. funestus in these sites but partially involved in Port Victoria [17]. Other mechanism(s) might be contributing to pyrethroid resistance in Port Victoria leading to that site having the highest level of resistance compared to the other sites.
An. funestus has no kdr markers for resistance [61] hence metabolic resistance mechanism through overexpression of detoxification genes plays a crucial role in insecticide resistance [62, 63]. In this study, we have identified the top twenty ncRNAs that were differentially expressed in resistant and unexposed field populations of An. funestus from western Kenya. Although their mechanisms of pyrethroid resistance in An. funestus is unknown, they could be playing a role in regulating the expression of pyrethroid-resistant metabolic genes in the An. funestus resistant populations. Our findings add up to a body of evidence which hypothesised that ncRNAs play roles in insecticide resistance development [28]. In general, the biological roles of ncRNAs in detoxification and insecticide resistance pathways are poorly understood. However, few studies have reported that some ncRNAs (notably microRNAs) interfered with the expression of insecticide-detoxifying enzymes. For instance, MiR-2b-3p has been proposed to potentially suppress the cytochrome P450 9f2 (CYP9F2) gene’s transcriptional activity, which would impede the larvae of P. xylostella from progressing through developmental detoxification pathways [64]. Furthermore, it has been observed that an overabundance of miR-13664 reduced the cytochrome P450 314A1 (CpCYP314A1) gene’s mRNA expression levels, increasing Culex pipiens pallens (Diptera: Culicidae) susceptibility to deltamethrin [65]. Given that a few proportions of the metabolic gene families (IMP, cytochrome P450, CP, OP, GST, UGTs, ABCs and COE.) were identified in this study to be involved in pyrethroid resistance and were mostly moderately overexpressed, this large proportion of highly overexpressed ncRNAs may play a crucial role in regulating their expression.
Studies have established those genes belonging to the esterase, cytochrome P450, esterases, GSTs, UGTs, cuticular proteins and ABC transporter families are implicated in insecticide resistance [19, 40, 66–68]; as a result, ncRNAs could be considered for designing RNAi-based control systems. This will, however, require a deeper understanding of the molecular mechanisms underlying RNAi-based control systems in mosquitoes since there already existing gaps in understanding this technology in controlling other insects [69]. Furthermore, high-throughput sequencing techniques have recently yielded important new information about the functions of ncRNAs in insect development and the evolution of insecticide resistance [69]. Non-coding rRNAs constitute over 80% of the total cellular RNA in mosquitoes [70]. In this study, the majority of rRNA was removed using the RiboZero Plus kit. Typically, the literature indicates that rRNA comprises anywhere from 1 to 20% of the final rRNA-depleted sequencing libraries [71]. Our results demonstrate that our RNA-seq method can effectively detect and quantify both coding and non-coding RNA.
Two different ncRNA-based insect management approaches have been proposed following these findings: (a) using biodegradable ncRNA-insecticide solutions to control insects, [72, 73] and (b) using metabolic engineering techniques to find and take advantage of target species’ ncRNA-associated signalling pathways [74]. However, the molecular mechanisms behind the functioning of ncRNAs in detoxification and insecticide resistance signalling pathways are still not clear, despite the mounting body of evidence suggesting these molecules are significant regulators of insect development [28, 69, 75]. This is because research in this area is still in its early stages. The straightforward CRISPR-Cas9 genome editing technique has the potential to generate novel understandings of the roles of regulatory ncRNA sequences as well as ncRNA-based techniques targeted at managing insects including disease vectors. Moreover, using inhibitors to target specific ncRNAs might interfere with the expression levels and reduce or reverse insecticide resistance. Thus, ncRNAs could be potential targets for vector control in the future. Recently, Oberemok et al. [76] proposed an innovative strategy to tackle insecticide resistance and create safer compounds. Their method employs synthetic DNA oligomers to disrupt gene expression by targeting ribosomal RNA (rRNA) rather than messenger RNA (mRNA). Because rRNA makes up 80% of cellular RNA and is more plentiful and stable than mRNA, it represents a promising target for DNA antisense oligonucleotide (ASO) interventions. This strategy seeks to offer a more efficient and enduring approach to combating insecticide resistance.
An. funestus is a notorious vector of human malaria in Africa and has contributed to over 90% of all malaria transmission in some parts of eastern and southern African regions [12, 13]. It is noteworthy that the outcome of our study represents an advancement in the molecular basis of insecticide resistance in An. funestus population. Our comprehension of the functional importance of ncRNAs in insecticide resistance pathways could enable the creation of ncRNA-based vector control techniques to control An. funestus.
Conclusions
An. funestus population is highly resistant to pyrethroids in western Kenya with Port Victoria recording the highest levels of resistance to the type I and type II pyrethroids. However, preexposure to PBO synergists recorded high susceptibility to the pyrethroids except in Port Victoria. We have shown for the first time that insecticide resistance in An. funestus is linked to the expression of ncRNAs hence a better understanding of these molecular events could help to develop resistance management strategies for future malaria control.
Acknowledgements:
Sincere thanks to the heads of households in the Kombewa, Teso, Bungoma, Port Victoria and Siaya communities who permitted us to collect mosquitoes inside their living rooms. Special thanks to the staff International Centre of Excellence for Malaria Research, Tom Mboya University, Homabay, Kenya and Mr Arnaud Tepa of Centre for Research in Infectious Diseases (CRID), Cameron for his inputs and advice on the data analysis.
Funding:
This study was supported by grants from the US National Institute of Health (R01 A1123074, U19 AI129326, R01 AI050243, and D43 TW001505). There was no additional external funding received for this study. The funders, however, did not play any role in designing, data collection, and manuscript writing.
List Of Abbreviations
- ABCs
ATP-binding cassettes
- COEs
carboxylesterases
- CPs
cuticular proteins
- CYPs
cytochrome P450s
- DDT
dichlorodiphenyltrichloroethane
- FC
fold change
- FDR
false discovery rate
- GO
gene ontology
- GSTs
glutathione S-transferases
- IMPs
imipenemase
- ncRNAs
non-coding RNAs
- OPs
olfactory proteins
- PBO
piperonyl butoxide
- PCA
principal component analysis
- UGTs
UDP-glycosyltransferases
Funding Statement
This study was supported by grants from the US National Institute of Health (R01 A1123074, U19 AI129326, R01 AI050243, and D43 TW001505). There was no additional external funding received for this study. The funders, however, did not play any role in designing, data collection, and manuscript writing.
Footnotes
Ethics approval and consent: This study was approved by Maseno University’s Ethics Review Committee (MUERC/00778/19). Verbal consent was sought from owners of households before mosquitoes were collected inside the living rooms.
Competing interests: The authors declare no conflict of interest.
Table 3 is available in the Supplementary Files section.
Supplementary Files
Contributor Information
Isaiah Debrah, University of Ghana.
Daibin Zhong, University of California, Irvine.
Maxwell G. Machani, Kenya Medical Research Institute
Godfrey Nattoh, Kenya Medical Research Institute.
Kevin O. Ochwedo, Sub-Saharan African International Centre of Excellence for Malaria Research, Tom Mboya University
Collins M. Morang’a, University of Ghana
Ming-Chieh Lee, University of California, Irvine.
Linda E. Amoah, University of Ghana
Andrew K. Githeko, Kenya Medical Research Institute
Yaw A. Afrane, University of Ghana Medical School, University of Ghana
Guiyun Yan, University of California, Irvine.
Availability of data and materials:
The RNA-Seq raw reads of the An. funestus data obtained were submitted to the NCBI Sequence Read Archive (SRA) under the BioSample accession number: SAMN39976597-SAMN39976605 and bioproject accession number: PRJNA1077587.
References
- 1.Ototo EN, Mbugi JP, Wanjala CL, Zhou G, Githeko AK, Yan G. Surveillance of malaria vector population density and biting behaviour in western Kenya. Malar J. 2015;14:244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Trape J-F, Tall A, Sokhna C, Ly AB, Diagne N, Ndiath O, et al. The rise and fall of malaria in a west African rural community, Dielmo, Senegal, from 1990 to 2012: a 22 year longitudinal study. Lancet Infect Dis. 2014;14:476–88. [DOI] [PubMed] [Google Scholar]
- 3.Bhatt S, Weiss DJ, Cameron E, Bisanzio D, Mappin B, Dalrymple U, et al. The effect of malaria control on Plasmodium falciparum in Africa between 2000 and 2015. Nature. 2015;526:207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Eisele TP, Larsen DA, Walker N, Cibulskis RE, Yukich JO, Zikusooka CM, et al. Estimates of child deaths prevented from malaria prevention scale-up in Africa 2001–2010. Malar J. 2012;11:1–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.O’Meara WP, Mangeni JN, Steketee R, Greenwood B. Changes in the burden of malaria in sub-Saharan Africa. Lancet Infect Dis. 2010;10:545–55. [DOI] [PubMed] [Google Scholar]
- 6.Mwesigwa J, Okebe J, Affara M, Di Tanna GL, Nwakanma D, Janha O, et al. On-going malaria transmission in The Gambia despite high coverage of control interventions: a nationwide cross-sectional survey. Malar J. 2015;14:1–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Nabatanzi M, Ntono V, Kamulegeya J, Kwesiga B, Bulage L, Lubwama B, et al. Malaria outbreak facilitated by increased mosquito breeding sites near houses and cessation of indoor residual spraying, Kole district, Uganda, January-June 2019. BMC Public Health. 2022;22:1–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Trape J-F, Tall A, Diagne N, Ndiath O, Ly AB, Faye J, et al. Malaria morbidity and pyrethroid resistance after the introduction of insecticide-treated bednets and artemisinin-based combination therapies: a longitudinal study. Lancet Infect Dis. 2011;11:925–32. [DOI] [PubMed] [Google Scholar]
- 9.Zhou G, Afrane YA, Vardo-Zalik AM, Atieli H, Zhong D, Wamae P, et al. Changing patterns of malaria epidemiology between 2002 and 2010 in Western Kenya: the fall and rise of malaria. PloS One. 2011;6:e20318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Dia I, Guelbeogo MW, Ayala D. Advances and Perspectives in the Study of the Malaria Mosquito Anopheles funestus. Anopheles Mosquitoes-New Insights Malar Vectors. 2013;10:55389. [Google Scholar]
- 11.Debrah I, Afrane YA, Amoah LE, Ochwedo KO, Mukabana WR, Zhong D, et al. Larval ecology and bionomics of Anopheles funestus in highland and lowland sites in western Kenya. PLOS ONE. 2021;16:e0255321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kaindoa EW, Matowo NS, Ngowo HS, Mkandawile G, Mmbando A, Finda M, et al. Interventions that effectively target Anopheles funestus mosquitoes could significantly improve control of persistent malaria transmission in south-eastern Tanzania. PloS One. 2017;12:e0177807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mapua SA, Hape EE, Kihonda J, Bwanary H, Kifungo K, Kilalangongono M, et al. Persistently high proportions of plasmodium-infected Anopheles funestus mosquitoes in two villages in the Kilombero valley, South-Eastern Tanzania. Parasite Epidemiol Control. 2022;:e00264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Sougoufara S, Diédhiou SM, Doucouré S, Diagne N, Sembène PM, Harry M, et al. Biting by Anopheles funestus in broad daylight after use of long-lasting insecticidal nets: a new challenge to malaria elimination. Malar J. 2014;13:125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Mulamba C, Riveron JM, Ibrahim SS, Irving H, Barnes KG, Mukwaya LG, et al. Widespread pyrethroid and DDT resistance in the major malaria vector Anopheles funestus in East Africa is driven by metabolic resistance mechanisms. PloS One. 2014;9:e110058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Akoton R, Tchigossou GM, Djègbè I, Yessoufou A, Atoyebi MS, Tossou E, et al. Experimental huts trial of the efficacy of pyrethroids/piperonyl butoxide (PBO) net treatments for controlling multi-resistant populations of Anopheles funestus s.s. in Kpomè, Southern Benin. Wellcome Open Res. 2018;3:71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.WHO. Test procedures for insecticide resistance monitoring in malaria vector mosquitoes. 2016.
- 18.Riveron JM, Yunta C, Ibrahim SS, Djouaka R, Irving H, Menze BD, et al. A single mutation in the GSTe2 gene allows tracking of metabolically based insecticide resistance in a major malaria vector. Genome Biol. 2014;15:1–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Messenger LA, Impoinvil LM, Derilus D, Yewhalaw D, Irish S, Lenhart A. A whole transcriptomic approach provides novel insights into the molecular basis of organophosphate and pyrethroid resistance in Anopheles arabiensis from Ethiopia. Insect Biochem Mol Biol. 2021;139:103655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wondji CS, Morgan J, Coetzee M, Hunt RH, Steen K, Black WC, et al. Mapping a quantitative trait locus (QTL) conferring pyrethroid resistance in the African malaria vector Anopheles funestus. BMC Genomics. 2007;8:34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cech TR, Steitz JA. The noncoding RNA revolution—trashing old rules to forge new ones. Cell. 2014;157:77–94. [DOI] [PubMed] [Google Scholar]
- 22.Mei Y, Khan H, Shishikura M, Ishiyama S, Khan A, Orita H, et al. pfeRNAs-A Novel Class of Small Non-Coding RNAs With Real Translational Potential. J Surg Res. 2023;284:237–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Qiao J, Du Y, Yu J, Guo J. MicroRNAs as Potential Biomarkers of Insecticide Exposure: A Review. Chem Res Toxicol. 2019;32:2169–81. [DOI] [PubMed] [Google Scholar]
- 24.Catalanotto C, Cogoni C, Zardo G. MicroRNA in control of gene expression: an overview of nuclear functions. Int J Mol Sci. 2016;17:1712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci. 2009;106:11667–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, et al. RNA Maps Reveal New RNA Classes and a Possible Function for Pervasive Transcription. Science. 2007;316:1484–8. [DOI] [PubMed] [Google Scholar]
- 27.Zhu B, Xu M, Shi H, Gao X, Liang P. Genome-wide identification of lncRNAs associated with chlorantraniliprole resistance in diamondback moth Plutella xylostella (L.). BMC Genomics. 2017;18:380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Etebari K, Furlong MJ, Asgari S. Genome-wide discovery of long intergenic non-coding RNAs in Diamondback moth (Plutella xylostella) and their expression in insecticide resistant strains. Sci Rep. 2015;5:14642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Liu F, Guo D, Yuan Z, Chen C, Xiao H. Genome-wide identification of long non-coding RNA genes and their association with insecticide resistance and metamorphosis in diamondback moth, Plutella xylostella. Sci Rep. 2017;7:15870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Chen L-L, Carmichael GG. Decoding the function of nuclear long non-coding RNAs. Curr Opin Cell Biol. 2010;22:357–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Tan W, Sun L, Zhang D, Sun J, Qian J, Hu X, et al. Cloning and overexpression of ribosomal protein L39 gene from deltamethrin-resistant Culex pipiens pallens. Exp Parasitol. 2007;115:369–78. [DOI] [PubMed] [Google Scholar]
- 32.Hu X, Wang W, Zhang D, Jiao J, Tan W, Sun Y, et al. Cloning and characterization of 40S ribosomal protein S4 gene from Culex pipiens pallens. Comp Biochem Physiol B Biochem Mol Biol. 2007;146:265–70. [DOI] [PubMed] [Google Scholar]
- 33.He J, Sun H, Zhang D, Sun Y, Ma L, Chen L, et al. Cloning and characterization of 60S ribosomal protein L22 (RPL22) from Culex pipiens pallens. Comp Biochem Physiol B Biochem Mol Biol. 2009;153:216–22. [DOI] [PubMed] [Google Scholar]
- 34.Sun H, Sun L, He J, Shen B, Yu J, Chen C, et al. Cloning and characterization of ribosomal protein S29, a deltamethrin resistance associated gene from Culex pipiens pallens. Parasitol Res. 2011;109:1689–97. [DOI] [PubMed] [Google Scholar]
- 35.McCann RS, Ochomo E, Bayoh MN, Vulule JM, Hamel MJ, Gimnig JE, et al. Reemergence of Anopheles funestus as a Vector of Plasmodium falciparum in Western Kenya after Long-Term Implementation of Insecticide-Treated Bed Nets. Am J Trop Med Hyg. 2014;90:597–604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhou G, Lee M-C, Githeko AK, Atieli HE, Yan G. Insecticide-treated net campaign and malaria transmission in western Kenya: 2003–2015. Front Public Health. 2016;4:153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kamau L, Agai D, Matoke D, Wachira L, Gikandi G, Vulule JM. Status of insecticide susceptibility in Anopheles gambiae sensu lato and Anopheles funestus mosquitoes from western Kenya. J Insect Sci. 2008;8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kamau L, Munyekenye GO, Koekemoer LL, Hunt RH, Coetzee M. A survey of the Anopheles funestus (Diptera: Culicidae) group of mosquitoes from 10 sites in Kenya with special emphasis on population genetic structure based on chromosomal inversion karyotypes. J Med Entomol. 2003;40:664–71. [DOI] [PubMed] [Google Scholar]
- 39.Kawada H, Dida GO, Ohashi K, Komagata O, Kasai S, Tomita T, et al. Multimodal pyrethroid resistance in malaria vectors, Anopheles gambiae ss, Anopheles arabiensis, and Anopheles funestus ss in western Kenya. PloS One. 2011;6:e22574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Sandeu MM, Mulamba C, Weedall GD, Wondji CS. A differential expression of pyrethroid resistance genes in the malaria vector Anopheles funestus across Uganda is associated with patterns of gene flow. Plos One. 2020;15:e0240743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Crawford JE, Guelbeogo WM, Sanou A, Traoré A, Vernick KD, Sagnon N, et al. De Novo Transcriptome Sequencing in Anopheles funestus Using Illumina RNA-Seq Technology. PLoS ONE. 2010;5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Coetzee M. Key to the females of Afrotropical Anopheles mosquitoes (Diptera: Culicidae). Malar J. 2020;19:1–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Gillies MT, Coetzee M. A supplement to the Anophelinae of Africa South of the Sahara. Publ Afr Inst Med Res. 1987;55:1–143. [Google Scholar]
- 44.Musapa M, Kumwenda T, Mkulama M, Chishimba S, Norris DE, Thuma PE, et al. A simple Chelex protocol for DNA extraction from Anopheles spp. JoVE J Vis Exp. 2013;:e3281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Koekemoer LL, Kamau L, Hunt RH, Coetzee M. A cocktail polymerase chain reaction assay to identify members of the Anopheles funestus (Diptera: Culicidae) group. Am J Trop Med Hyg. 2002;66:804–11. [DOI] [PubMed] [Google Scholar]
- 46.Cohuet A, Simard F, Toto J-C, Kengne P, Coetzee M, Fontenille D. SPECIES IDENTIFICATION WITHIN THE ANOPHELES FUNESTUS GROUP OF MALARIA VECTORS IN CAMEROON AND EVIDENCE FOR A NEW SPECIES. Am J Trop Med Hyg. 2003;69:200–5. [PubMed] [Google Scholar]
- 47.Vitucci EC, McCullough SD. Isolation of Total RNA with the Zymo Quick-RNA Miniprep Kit. 2022.
- 48.Andrews S. FastQC: a quality control tool for high throughput sequence data. 2010. 2017.
- 49.Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Anders S, Pyl PT, Huber W. HTSeq—a Python framework to work with high-throughput sequencing data. bioinformatics. 2015;31:166–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Wondji CS, Hearn J, Irving H, Wondji MJ, Weedall G. RNAseq-based gene expression profiling of the Anopheles funestus pyrethroid-resistant strain FUMOZ highlights the predominant role of the duplicated CYP6P9a/b cytochrome P450s. G3. 2022;12:jkab352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Anders S, Huber W. Differential expression analysis for sequence count data. Nat Preced. 2010;:1–1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Schulze SK, Kanwar R, Gölzenleuchter M, Therneau TM, Beutler AS. SERE: single-parameter quality control and sample comparison for RNA-Seq. BMC Genomics. 2012;13:1–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Ferreira JA, Zwinderman AH. On the benjamini–hochberg method. Ann Stat. 2006;34:1827–49. [Google Scholar]
- 57.Wu Y-J, Wu Y-J, Luo X, Shen X-L, Zhao D-G. Identification of differentially expressed genes that potentially confer pest resistance in transgenic ChIFN-γ tobacco. Gene. 2014;543:181–9. [DOI] [PubMed] [Google Scholar]
- 58.Alexa A, Rahnenfuhrer J. topGO: enrichment analysis for gene ontology. R Package Version. 2010;2:2010. [Google Scholar]
- 59.Djouaka R, Riveron JM, Yessoufou A, Tchigossou G, Akoton R, Irving H, et al. Multiple insecticide resistance in an infected population of the malaria vector Anopheles funestus in Benin. Parasit Vectors. 2016;9:453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Riveron JM, Chiumia M, Menze BD, Barnes KG, Irving H, Ibrahim SS, et al. Rise of multiple insecticide resistance in Anopheles funestus in Malawi: a major concern for malaria vector control. Malar J. 2015;14:344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Menze BD, Riveron JM, Ibrahim SS, Irving H, Antonio-Nkondjio C, Awono-Ambene PH, et al. Multiple Insecticide Resistance in the Malaria Vector Anopheles funestus from Northern Cameroon Is Mediated by Metabolic Resistance Alongside Potential Target Site Insensitivity Mutations. PloS One. 2016;11:e0163261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Hemingway J. The molecular basis of two contrasting metabolic mechanisms of insecticide resistance. Insect Biochem Mol Biol. 2000;30:1009–15. [DOI] [PubMed] [Google Scholar]
- 63.Hemingway J, Ranson H. Insecticide resistance in insect vectors of human disease. Annu Rev Entomol. 2000;45:371–91. [DOI] [PubMed] [Google Scholar]
- 64.Etebari K, Afrad MH, Tang B, Silva R, Furlong MJ, Asgari S. Involvement of microRNA miR-2b-3p in regulation of metabolic resistance to insecticides in Plutella xylostella. Insect Mol Biol. 2018;27:478–91. [DOI] [PubMed] [Google Scholar]
- 65.Sun XH, Xu N, Xu Y, Zhou D, Sun Y, Wang WJ, et al. A novel miRNA, miR-13664, targets CpCYP314A1 to regulate deltamethrin resistance in Culex pipiens pallens. Parasitology. 2019;146:197–205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Riveron JM, Tchouakui M, Mugenzi L, Menze BD, Chiang M-C, Wondji CS. Insecticide resistance in malaria vectors: an update at a global scale. In: Towards malaria elimination-a leap forward. IntechOpen; 2018. [Google Scholar]
- 67.Riveron JM, Yunta C, Ibrahim SS, Djouaka R, Irving H, Menze BD, et al. A single mutation in the GSTe2 gene allows tracking of metabolically based insecticide resistance in a major malaria vector. Genome Biol. 2014;15:R27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Al-Yazeedi T, Muhammad A, Irving H, Ahn S-J, Hearn J, S. Wondji C. Overexpression and nonsynonymous mutations of UDP-glycosyltransferases potentially associated with pyrethroid resistance in Anopheles funestus. bioRxiv. 2023;:2023–08. [DOI] [PubMed] [Google Scholar]
- 69.Vaschetto LM, Beccacece HM. The emerging importance of noncoding RNAs in the insecticide tolerance, with special emphasis on Plutella xylostella (Lepidoptera: Plutellidae). WIREs RNA. 2019;10:e1539. [DOI] [PubMed] [Google Scholar]
- 70.Gale K, Crampton J. The ribosomal genes of the mosquito, Aedes aegypti. Eur J Biochem. 1989;185:311–7. [DOI] [PubMed] [Google Scholar]
- 71.Potemkin N, Cawood SM, Treece J, Guévremont D, Rand CJ, McLean C, et al. A method for simultaneous detection of small and long RNA biotypes by ribodepleted RNA-Seq. Sci Rep. 2022;12:621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Kolliopoulou A, Swevers L. Recent progress in RNAi research in Lepidoptera: Intracellular machinery, antiviral immune response and prospects for insect pest control. Curr Opin Insect Sci. 2014;6:28–34. [DOI] [PubMed] [Google Scholar]
- 73.Powell M, Pyati P, Cao M, Bell H, Gatehouse JA, Fitches E. Insecticidal effects of dsRNA targeting the Diap1 gene in dipteran pests. Sci Rep. 2017;7:15147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Buchman A, Marshall JM, Ostrovski D, Yang T, Akbari OS. Synthetically engineered Medea gene drive system in the worldwide crop pest Drosophila suzukii. Proc Natl Acad Sci. 2018;115:4725–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Bavithra CML, Murugan M, Pavithran S, Naveena K. Enthralling genetic regulatory mechanisms meddling insecticide resistance development in insects: role of transcriptional and post-transcriptional events. Front Mol Biosci. 2023;10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Oberemok VV, Laikova KV, Gal’chinsky NV, Useinov RZ, Novikov IA, Temirova ZZ, et al. DNA insecticide developed from the Lymantria dispar 5.8 S ribosomal RNA gene provides a novel biotechnology for plant protection. Sci Rep. 2019;9:6197. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The RNA-Seq raw reads of the An. funestus data obtained were submitted to the NCBI Sequence Read Archive (SRA) under the BioSample accession number: SAMN39976597-SAMN39976605 and bioproject accession number: PRJNA1077587.