Graphical abstract
Keywords: Extracellular matrix, Collagen, Collagen PTMs, PTM identification, PTM quantitation, Knowledge base
Highlights
-
•
First dedicated knowledge base, ColPTMScape for collagen PTMs.
-
•
Information on the site-specificity and occupancy of collagen prolyl 3-hydroxylation in different tissue and species.
-
•
Consolidated site-specific knowledge of collagen lysyl-hydroxylation and O-glycosylation.
-
•
ColPTMScape would contribute to the larger matrix biology community.
Abstract
Collagen is a key component of the extracellular matrix (ECM). In the remodeling of ECM, a remarkable variation in collagen post-translational modifications (PTMs) occurs. This makes collagen a potential target for understanding extracellular matrix remodeling during pathological conditions. Over the years, scientists have gathered a huge amount of data about collagen PTM during extracellular matrix remodeling. To make such information easily accessible in a consolidated space, we have developed ColPTMScape (https://colptmscape.iitmandi.ac.in/), a dedicated knowledge base for collagen PTMs. The identified site-specific PTMs, quantitated PTM sites, and PTM maps of collagen chains are deliverables to the scientific community, especially to matrix biologists. Through this knowledge base, users can easily gain information related to the difference in the collagen PTMs across different tissues in different organisms.
Introduction
The extracellular matrix (ECM) forms a meshwork composed of different macromolecules such as elastin, collagen, proteoglycans, and others to support the tissues as well as help in cell proliferation, differentiation, and apoptosis [1]. The ECM remodels according to the cell behavior, which is also observed in pathological conditions such as fibrosis [2]. In fibrosis, excessive collagen deposits in the ECM (a hallmark) [3]. Collagen is a fibrous structural protein, which is most abundant in the ECM [4]. Owing to the structural integrity of collagen, it provides mechanical strength to the tissues. Collagen has a right-handed triple helical structure, comprising three left-handed polyproline type II (PPII)-like polypeptide strands. A common repetitive tripeptide motif, Gly-Xaa-Yaa, is required for the formation of a collagen triple helix. Prevalently, proline and hydroxyproline occupy the Xaa and Yaa positions, respectively [5]. There are two broader families of collagen based on the organization, which are, fibrillar and non-fibrillar in nature. Collagen I and collagen IV are the abundant types in fibrillar and basement-membrane forming collagen families, respectively. The presence and variations of types of collagens are tissue-specific. Collagen has a plethora of post-translational modifications (PTMs) such as hydroxylation, glycosylation, phosphorylation etc. [6], [7]. The levels of PTMs vary in types of collagen, for instance, collagen type IV has a higher number (10–15 residues per 1000 amino acids) of 3-hydroxyproline (3-Hyp), whereas only 1–2 residues in a collagen chain of collagen type I [8], [9], [10], [11]. Two of the PTMs that are well-studied are hydroxylation of proline and lysine and glycosylation of hydroxylysine (Fig. 1). PTMs of collagen are critical for the maintenance of tissue homeostasis. Perturbed PTMs of collagen culminate in different disorders, such as 3-Hyp deficiency in collagen I and IV causes osteogenesis imperfecta [12], [13], platelet aggregation [14], and poor eye tissue development [15], respectively. Similarly, the deficiency of 4-Hyp, a PTM that provides thermal stability to collagen, can cause the development of musculoskeletal diseases, myopia, etc. [16]. Lysine can be hydroxylated and further o-glycosylated to form hydroxylysine and galactosyl-hydroxylysine (Gal-Hyl) or glucosyl-galactosyl-hydroxylysine (GluGal-Hyl), respectively [17], [18], [19]. The lysine or hydroxylysine residues in the telopeptide region become allysine (Lysald) or hydroxyallysine (Hylald), respectively, to go into a condensation reaction to form crosslinks between collagen molecules, which are essential to forming a collagen fibril [17], [20], [21]. These modifications stabilize the collagen fibril through the crosslink formation [17], [21]. The alteration in the levels of hydroxylysine (Hyl) and o-glycosylations of Hyl is implicated in cerebral small vessel disease [22], musculoskeletal defects [23], connective tissue disorder [24], Bruck syndrome [25], [26], fibrosis [27], [28], connective tissue disorder [29], [30], [31], high myopia [15], Ehlers Danlos syndrome type VI A [32], cancer [33], [34], [35], etc. The levels of collagen as well as its PTMs alter during the ECM remodeling [6], [36]. Therefore, the knowledge of the alteration in collagen and its PTMs is significant to understanding the undergoing changes in the diseased conditions. However, the characterization of collagen PTMs has remained a challenge for the scientific community.
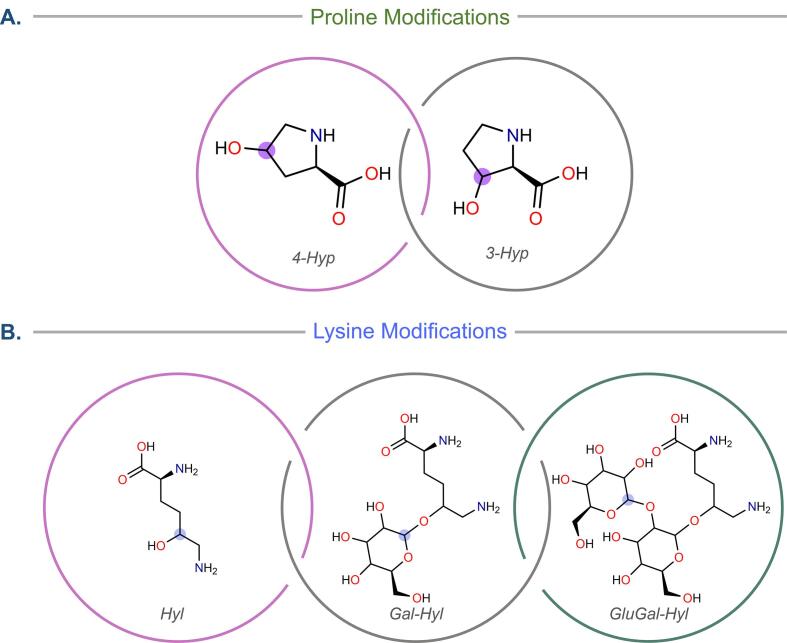
Fig. 1.
Modifications on proline and lysine. A. Two types of prolyl hydroxylations, 4-hydroxyproline (4-Hyp) and 3-hydroxyproline (3-Hyp). B. Lysine can be hydroxylated to form hydroxylysine (Hyl) and o-glycosylated to form galactosyl-hydroxylysine (Gal-Hyl) and glucosylgalactosyl-hydroxylysine (GluGal-Hyl).
After the discovery of collagen structure, gaining information about amino acid sequence of collagen has become of paramount importance. Initially, the amino acid analysis was performed by hydrolysing the collagen chains [37], [38], [39], which provided qualitative information. However, site-specific quantitation of PTMs remained elusive. With the development of high-resolution mass spectrometry (MS)-based proteomics it has become feasible now to identify and quantitate site-specific collagen PTMs [6], [22], [40], [41], [42], [43], [44], [45]. Different research groups including our lab have been using MS-based proteomics approach to characterize the matrisome proteins and site-specific collagen PTMs [40], [45], [46], [47], [48]. The knowledge base for the matrix biology community has been created (MatrisomeDB and MatrixDB) [49], [50], [51], [52], [53]. However, the resolution of different types of hydroxylations and glycosylations on collagen at a site-specific level were not properly addressed. Such information could be immensely useful for matrix biologists, clinicians, and basic scientists to understand the molecular changes in collagen during disease development. This has inspired us to develop a dedicated knowledge base solely for collagen PTMs, named ColPTMScape (https://colptmscape.iitmandi.ac.in/). ColPTMScape is uniquely different from the other large extracellular matrix-related databases. It is dedicated to only collagen PTMs and highlights the data in a map that is already available from different research studies. Hence, it is less complicated, loads faster, and is easy to use. ColPTMScape has incorporated the occupancy level of site-specific PTMs of collagen. Downloading the site-specific PTMs and copying the sequence of a collagen chain is available. However, it has some limitations. Currently, the database does not show sequence coverage, does not allow a user to download the highlighted PTM map of a collagen chain, and does not have data on abnormal or disease conditions. This paves the way for future incorporation of additional features.
With ColPTMScape, we will bring the site-specifically identified and quantified collagen (mainly I and IV) PTMs to one consolidated space. Characterization of specific hydroxylation and glycosylation was performed using the in-house developed in-silico approach. Data for 6 different organisms from several published studies [26], [30], [31], [35], [40], [41], [45], [46], [47], [48], [54], [55], [56] has been used to develop the first version of this knowledge base. We have highlighted the following PTMs based on the availability of data: hydroxyproline (Hyp), 3-Hyp, 4-hydroxyproline (4-Hyp), Lysald/Hylald, Hyl, galactosyl-hydroxylysine (Gal-Hyl), and glucosylgalactosyl-hydroxylysine (GluGal-Hyl) in the knowledge base. With new discoveries in collagen PTM field, ColPTMScape will be updated on a regular basis. This knowledge base would lay the consolidated foundation on collagen PTMs in a tissue and species-specific manner and would contribute to the larger matrix biology community.
Results and discussion
ColPTMScape search request
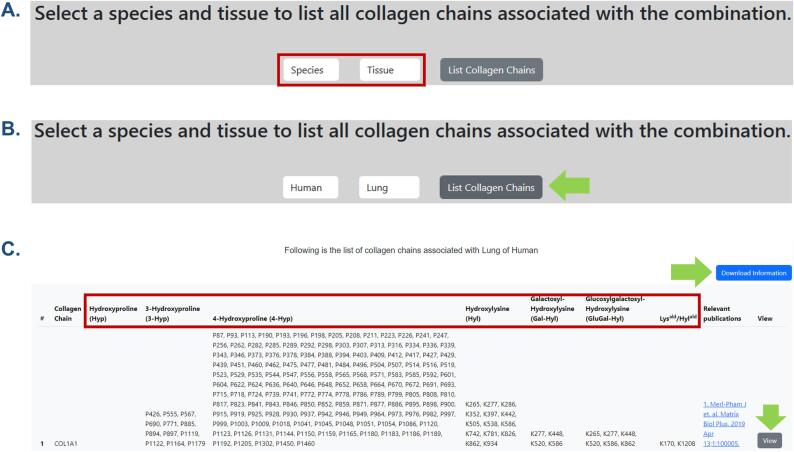
ColPTMScape consists of a user-friendly interface to search for the collagen chains in the desired tissue of an organism. We provided the drop-down option to the user to select an organism as well as a tissue in that organism (Fig. 2A). Then, the user needs to select the “List Collagen Chains” button (Fig. 2B). The button directs the user to the page where collagen chains in the selected tissue of the organism are listed.
Fig. 2.
Interaction with initial pages of the knowledge base. A. Drop-down options are highlighted with the red box. B. For getting collagen chains for a particular tissue, the list collagen chains button directs to the respective page. C. All the PTMs are highlighted with the red box. A user can view these PTMs on the PTM map using the view button. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
Listing collagen chains
After selecting the organism and the tissue, the user will see the page with all collagen chains that were analyzed for site-specific PTM identification and quantitation. For each collagen chain, the user finds positions of Hyp, 3-Hyp, 4-Hyp, Hyl, Gal-Hyl, GluGal-Hyl, and Lysald/Hylald mentioned on the page. Moreover, by selecting the “View” button, the user is directed to the page where the collagen map of a particular chain is available (Fig. 2C). The collagen PTM map contains information on the key PTMs. Furthermore, there is a “Download Information” button on the page where the list of all collagen chains and site-specific posttranslational modifications (PTMs) is present for any combination of organism and tissue in the form of an excel sheet. (Fig. 2C).
Collagen PTM map
As a result of searching through ColPTMScape, users find a site-specific collagen PTM map. This page contains crucial information. On the top middle of the page (highlighted in the red box in Fig. 3A), the information bar provides initially the length of the sequence of the collagen chain. Post-selecting the modifications to be displayed on the map, the box on the top of the page shows the number of modified amino acid residues. Moreover, on the top-left corner of the page, the toggle buttons allow the user to select an option, for instance, the 3-Hyp modification. The 3-Hyp modified residues are highlighted (Fig. 3B). Similarly, the user can select other modifications.
Fig. 3.
COL1A1 PTM map of the human lung. A. The length of the collagen chain is highlighted in the red box. On the top-left side of the page, all modifications and signal peptides are given as toggle buttons. On the top right-side of the page, buttons connecting to Uniprot and to copy sequence are available. B. 3-Hyp is selected to be highlighted on the collagen PTM map. The number of 3-Hyp on the information bar are highlighted with the red box. 3-Hyp modifications are highlighted with blue color. The position and occupancy of a particular 3-Hyp can be seen by moving the cursor to the modification. Underline in black represents propeptides, and underline in red represents telopeptides. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
One toggle button is provided for highlighting the signal peptide in a sequence of a particular collagen chain (Fig. 3A). ColPTMScape numbers amino acids in the sequence from the N-terminal of the signal peptides for any collagen chain. The amino acids in sequences are numbered in multiples of 10 as it allows a user to easily locate the modifications. Moreover, the N- and C-terminal propeptides are highlighted through solid black underline, whereas the N- and C-telopeptides through the solid red underline. To avoid overcrowding in a single page and for a better visualization, ColPTMScape does not highlight the helical domain. The domain that is not highlighted using a button or a type of underline is a helical domain in the map. On the top right side, there is a button named “Copy Sequence”, which allows a user to copy the sequence without any spaces between amino acids. Also, there is an option to go to a Uniprot ID from where the sequence is used for generating the map.
On hovering over the highlighted modified amino acid residues, a small data box pops up. In the box, the position of the modified residue and the occupancy levels of each modification are displayed (Fig. 3B). The information displayed in the box follows the same pattern for every toggle button except one that is the “Lysald/Hylald” button. When a user clicks on this button, lysine in the telopeptides will get highlighted. On hovering over such lysine residues, the occupancy of it being lysine (Lys) or Hyl will be displayed. The information can be used to understand the variations in the position and occupancy level of PTMs in normal and diseased conditions.
Compare conserved sites
At the home page of the ColPTMScape, a user finds a section named “Compare Conserved Sites” at the top of the page (Fig. 4A). We have developed a comparison tool that compares collagen chains of different organisms. Currently, we have only listed the conserved 3-Hyp and crosslinking lysine sites, both the telopeptidyl and helical lysine or hydroxylysine sites, of collagen 1 α1 and α2 chains. After selecting the collagen chain and modification, a user needs to select one or more organisms according to the requirement and then click on “Get Results” (Fig. 4A). The conserved PTM sites will be presented in a table format (Fig. 4C), which can be downloaded in a csv file format. A hyphen (-) in comparison is used if there is no evidence of the conserved PTM site detected in an organism. Moreover, this comparison tool does not show any results for an organism for which there is no mass spectrometry (MS)-based experimental evidence available.
Fig. 4.
A web page to compare conserved sites. A. Compare conserved sites link is present at the top of the home page. B. COL1A1 and 3-Hyp sites are selected for organisms Human and Mouse. C. 3-Hyp sites of COL1A1 chain that are conserved in organisms Human and Mouse are presented in a table. P1-P14 are abbreviations for position 1 to position 14.
Conclusions
In summary, we have developed a dedicated knowledge base, ColPTMScape for collagen PTMs. It provides information on the site-specific PTMs with the occupancy level of each modification. Along with the tabular information, the knowledge base can make a collagen PTM map, highlighting the selected PTMs as per the requirement. Moreover, information regarding the position of site-specific modification and occupancy level is available for users in a consolidated box. Also, the user can download the site-specific PTMs of collagen chains corresponding to a tissue of an organism. In the future, we would like to include data from different organisms and tissues. Moreover, we would also like to show the conservative nature of the site-specific PTMs, other than 3-Hyp, across different organisms and the variations of occupancy levels of PTMs in different diseased conditions.
Methodology
The project aimed to develop a web-based PTM (Post-Translational Modification) site visualizer for the matrix-biology community. An in-house in silico approach was developed to identify and quantify the site-specific collagen PTM using MS data [45]. Different datasets have been utilized as a source of MS data for different organisms. PXD011627 and PXD010092 for zebrafish myocardium ECM [57], PXD028908 for human’s heart ECM [58], PXD002488 for mice’s heart ECM [59], PXD005726 for pig’s BES and DES-induced neointima ECM [60], PXD008802 for fibrillar collagen in mice’s skin were utilized [61]. Using an in-house developed pipeline, our lab could identify and quantify collagen PTM sites more than previously known for these tissues. The information gathered through these studies was used as an input for knowledge base curation. We used collagen sequence, number of PTMs, and sites of PTMs to develop this first version of the knowledge base. The primary objective was to create a knowledge base and web tool that would display PTM sites of proteins from various species. PTM data for various tissues and proteins were collected from available sources. The backend of the web application was developed using Python-Flask, a microweb framework, to handle data processing and communication with the knowledge base Flask-Python was chosen for the backend due to its flexibility and simplicity. It allowed for efficient data handling, routing, and API development. The front end was designed using Bootstrap 5.0, providing a responsive and user-friendly interface for data visualization. Bootstrap, a popular frontend framework, was employed for designing the web interface. Its responsive design ensured compatibility with various devices and screen sizes. A comprehensive knowledge base was designed to store PTM data, ensuring that it could accommodate a wide range of proteins, tissues, and PTM types. One of the key features of the tool is its scalability. It can efficiently handle a growing number of entries and PTMs as the knowledge base expands over time. The web tool's user interface was developed to provide a user-friendly experience, allowing researchers to easily access and visualize PTM sites in proteins from different species. The knowledge base is optimized for the Google Chrome and Mozilla Firefox browser.
Author contributions
AJ and AN worked on development, design, manuscript draft preparation, and figure creation. LNM developed the initial framework of the browser. BM and TB conceptualized the overall structure of the study and finalized the manuscript.
CRediT authorship contribution statement
Ashutosh Joshi: Writing – original draft, Formal analysis, Data curation, Conceptualization. Ayush Nigam: Formal analysis, Data curation. Lalit Narayan Mudgal: Formal analysis, Data curation. Bhaskar Mondal: Writing – review & editing, Conceptualization. Trayambak Basak: Writing – review & editing, Writing – original draft, Supervision, Investigation, Funding acquisition, Formal analysis, Conceptualization.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgements
The Science and Engineering Research Board (SERB) funded core-research grant “Decoding the dynamics of cardiac ECM Matrisome during post-MI (myocardial infarction) remodeling (CRG/2022/006204; IITM/SERB/TB/332)” to TB is acknowledged for this work. AJ also acknowledges the HTRA fellowship (MoE, Govt. of India) for the doctoral program.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.mbplus.2024.100144.
Contributor Information
Bhaskar Mondal, Email: bhaskarmondal@iitmandi.ac.in.
Trayambak Basak, Email: trayambak@iitmandi.ac.in.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
Data availability
Data will be made available on request.
References
- 1.Oxford J.T., Reeck J.C., Hardy M.J. Extracellular matrix in development and disease. Int. J. Mol. Sci. 2019;20 doi: 10.3390/ijms20010205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Wynn T.A., Ramalingam T.R. Mechanisms of fibrosis: therapeutic translation for fibrotic disease. Nat. Med. 2012;18:1028–1040. doi: 10.1038/nm.2807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Cowling R.T., Kupsky D., Kahn A.M., Daniels L.B., Greenberg B.H. Mechanisms of cardiac collagen deposition in experimental models and human disease. Transl. Res. 2019;209:138–155. doi: 10.1016/j.trsl.2019.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Di Lullo G.A., Sweeney S.M., Körkkö J., Ala-Kokko L., San Antonio J.D. Mapping the ligand-binding sites and disease-associated mutations on the most abundant protein in the human, type I collagen. J. Biol. Chem. 2002;277:4223–4231. doi: 10.1074/jbc.M110709200. [DOI] [PubMed] [Google Scholar]
- 5.Shoulders M.D., Raines R.T. Collagen structure and stability. Annu. Rev. Biochem. 2009;78:929–958. doi: 10.1146/annurev.biochem.77.032207.120833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Onursal C., Dick E., Angelidis I., Schiller H.B., Staab-Weijnitz C.A. Collagen biosynthesis, processing, and maturation in lung ageing. Front. Med. 2021;8 doi: 10.3389/FMED.2021.593874/BIBTEX. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ishikawa Y., Bächinger H.P. A molecular ensemble in the rER for procollagen maturation. Biochim. Biophys. Acta - Mol. Cell Res. 1833;2013:2479–2491. doi: 10.1016/J.BBAMCR.2013.04.008. [DOI] [PubMed] [Google Scholar]
- 8.Kefalides N.A. Structure and biosynthesis of basement membranes. Int. Rev. Connect. Tissue Res. 1973;6:63–104. doi: 10.1016/B978-0-12-363706-2.50008-8. [DOI] [PubMed] [Google Scholar]
- 9.Gryder R.M., Lamon M., Adams E. Sequence position of 3 hydroxyproline in basement membrane collagen. isolation of glycyl 3 hydroxyprolyl 4 hydroxyproline from swine kidney. J. Biol. Chem. 1975;250:2470–2474. doi: 10.1016/s0021-9258(19)41624-4. [DOI] [PubMed] [Google Scholar]
- 10.Kresina T.F., Miller E.J. Isolation and characterization of basement membrane collagen from human placental tissue. Evidence for the Presence of Two Genetically Distinct Collagen Chains. Biochemistry. 1979;18:3089–3097. doi: 10.1021/bi00581a028. [DOI] [PubMed] [Google Scholar]
- 11.Weis M.A., Hudson D.M., Kim L., Scott M., Wu J.J., Eyre D.R. Location of 3-hydroxyproline residues in collagen types I, II, III, and V/XI implies a role in fibril supramolecular assembly. J. Biol. Chem. 2010;285:2580–2590. doi: 10.1074/jbc.M109.068726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Morello R., Bertin T.K., Chen Y., Hicks J., Tonachini L., Monticone M., Castagnola P., Rauch F., Glorieux F.H., Vranka J., Bächinger H.P., Pace J.M., Schwarze U., Byers P.H., Weis M.A., Fernandes R.J., Eyre D.R., Yao Z., Boyce B.F., Lee B. CRTAP is required for prolyl 3- hydroxylation and mutations cause recessive osteogenesis imperfecta. Cell. 2006;127:291–304. doi: 10.1016/J.CELL.2006.08.039. [DOI] [PubMed] [Google Scholar]
- 13.W.A. Cabral, W. Chang, A.M. Barnes, M. Weis, M.A. Scott, S. Leikin, E. Makareeva, N. V. Kuznetsova, K.N. Rosenbaum, C.J. Tifft, D.I. Bulas, C. Kozma, P.A. Smith, D.R. Eyre, J.C. Marini, Prolyl 3-hydroxylase 1 deficiency causes a recessive metabolic bone disorder resembling lethal/severe osteogenesis imperfecta, Nat. Genet. 2007 393. 39 (2007) 359–365. https://doi.org/10.1038/ng1968. [DOI] [PMC free article] [PubMed]
- 14.Pokidysheva E., Boudko S., Vranka J., Zientek K., Maddox K., Moser M., Fässler R., Ware J., Bächinger H.P. Biological role of prolyl 3-hydroxylation in type IV collagen. Proc. Natl. Acad. Sci. U. S. A. 2014;111:161–166. doi: 10.1073/PNAS.1307597111/SUPPL_FILE/PNAS.201307597SI.PDF. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hudson D.M., Joeng K.S., Werther R., Rajagopal A., Weis M., Lee B.H., Eyre D.R. Post-translationally abnormal collagens of prolyl 3-hydroxylase-2 null mice offer a pathobiological mechanism for the high myopia linked to human LEPREL1 mutations. J. Biol. Chem. 2015;290:8613–8622. doi: 10.1074/jbc.M114.634915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Salo A.M., Myllyharju J. Prolyl and lysyl hydroxylases in collagen synthesis. Exp. Dermatol. 2021;30:38–49. doi: 10.1111/exd.14197. [DOI] [PubMed] [Google Scholar]
- 17.Yamauchi M., Sricholpech M. Lysine post-translational modifications of collagen. Essays Biochem. 2012;52:113–133. doi: 10.1042/BSE0520113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Schegg B., Hülsmeier A.J., Rutschmann C., Maag C., Hennet T. Core glycosylation of collagen is initiated by two β(1-O)Galactosyltransferases. Mol. Cell. Biol. 2009;29:943–952. doi: 10.1128/MCB.02085-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Sricholpech M., Perdivara I., Nagaoka H., Yokoyama M., Tomer K.B., Yamauchi M. Lysyl hydroxylase 3 glucosylates galactosylhydroxylysine residues in type I collagen in osteoblast culture. J. Biol. Chem. 2011;286:8846–8856. doi: 10.1074/JBC.M110.178509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Trackman P.C. Enzymatic and non-enzymatic functions of the lysyl oxidase family in bone. Matrix Biol. 2016;52–54:7–18. doi: 10.1016/j.matbio.2016.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Piersma B., Bank R.A. Collagen cross-linking mediated by lysyl hydroxylase 2: an enzymatic battlefield to combat fibrosis. Essays Biochem. 2019;63:377–387. doi: 10.1042/EBC20180051. [DOI] [PubMed] [Google Scholar]
- 22.Miyatake S., Schneeberger S., Koyama N., Yokochi K., Ohmura K., Shiina M., Mori H., Koshimizu E., Imagawa E., Uchiyama Y., Mitsuhashi S., Frith M.C., Fujita A., Satoh M., Taguri M., Tomono Y., Takahashi K., Doi H., Takeuchi H., Nakashima M., Mizuguchi T., Takata A., Miyake N., Saitsu H., Tanaka F., Ogata K., Hennet T., Matsumoto N. Biallelic COLGALT1 variants are associated with cerebral small vessel disease. Ann. Neurol. 2018;84:843–853. doi: 10.1002/ANA.25367. [DOI] [PubMed] [Google Scholar]
- 23.Geister K.A., Lopez-Jimenez A.J., Houghtaling S., Ho T.H., Vanacore R., Beier D.R. Loss of function of Colgalt1 disrupts collagen post-translational modification and causes musculoskeletal defects. Dis. Model. Mech. 2019;12 doi: 10.1242/dmm.037176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Salo A.M., Cox H., Farndon P., Moss C., Grindulis H., Risteli M., Robins S.P., Myllylä R. A connective tissue disorder caused by mutations of the lysyl hydroxylase 3 gene. Am. J. Hum. Genet. 2008;83:495–503. doi: 10.1016/J.AJHG.2008.09.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gistelinck C., Witten P.E., Huysseune A., Symoens S., Malfait F., Larionova D., Simoens P., Dierick M., Van Hoorebeke L., De Paepe A., Kwon R.Y., Weis M.A., Eyre D.R., Willaert A., Coucke P.J. Loss of type I collagen telopeptide lysyl hydroxylation causes musculoskeletal abnormalities in a zebrafish model of Bruck syndrome. J. Bone Miner. Res. 2016;31:1930–1942. doi: 10.1002/jbmr.2977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gistelinck C., Weis M.A., Rai J., Schwarze U., Niyazov D., Song K.M., Byers P.H., Eyre D.R. Abnormal bone collagen cross-linking in osteogenesis Imperfecta/Bruck syndrome caused by compound heterozygous PLOD2 mutations. JBMR plus. 2021;5:1–15. doi: 10.1002/jbm4.10454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Van der Slot A.J., Zuurmond A.M., Bardoel A.F.J., Wijmenga C., Pruijs H.E.H., Sillence D.O., Brinckmann J., Abraham D.J., Black C.M., Verzijl N., DeGroot J., Hanemaaijer R., TeKoppele J.M., Huizinga T.W.J., Bank R.A. Identification of PLOD2 as telopeptide lysyl hydroxylase, an important enzyme in fibrosis. J. Biol. Chem. 2003;278:40967–40972. doi: 10.1074/jbc.M307380200. [DOI] [PubMed] [Google Scholar]
- 28.Van Der Slot-Verhoeven A.J., Van Dura E.A., Attema J., Blauw B., DeGroot J., Huizinga T.W.J., Zuurmond A.M., Bank R.A. The type of collagen cross-link determines the reversibility of experimental skin fibrosis. Biochim. Biophys. Acta - Mol. Basis Dis. 2005;1740:60–67. doi: 10.1016/j.bbadis.2005.02.007. [DOI] [PubMed] [Google Scholar]
- 29.Cabral W.A., Perdivara I., Weis M.A., Terajima M., Blissett A.R., Chang W., Perosky J.E., Makareeva E.N., Mertz E.L., Leikin S., Tomer K.B., Kozloff K.M., Eyre D.R., Yamauchi M., Marini J.C. Abnormal type I collagen post-translational modification and crosslinking in a cyclophilin B KO mouse model of recessive osteogenesis imperfecta. PLoS Genet. 2014;10 doi: 10.1371/journal.pgen.1004465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Terajima M., Taga Y., Chen Y., Cabral W.A., Hou-Fu G., Srisawasdi S., Nagasawa M., Sumida N., Hattori S., Kurie J.M., Marini J.C., Yamauchi M. Cyclophilin-B modulates collagen cross-linking by differentially affecting lysine hydroxylation in the helical and telopeptidyl domains of tendon type I collagen. J. Biol. Chem. 2016;291:9501–9512. doi: 10.1074/jbc.M115.699470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Saito T., Terajima M., Taga Y., Hayashi F., Oshima S., Kasamatsu A., Okubo Y., Ito C., Toshimori K., Sunohara M., Tanzawa H., Uzawa K., Yamauchi M. Decrease of lysyl hydroxylase 2 activity causes abnormal collagen molecular phenotypes, defective mineralization and compromised mechanical properties of bone. Bone. 2022;154 doi: 10.1016/j.bone.2021.116242. [DOI] [PubMed] [Google Scholar]
- 32.Takaluoma K., Hyry M., Lantto J., Sormunen R., Bank R.A., Kivirikko K.I., Myllyharju J., Soininen R. Tissue-specific changes in the hydroxylysine content and cross-links of collagens and alterations in fibril morphology in lysyl hydroxylase 1 knock-out mice. J. Biol. Chem. 2007;282:6588–6596. doi: 10.1074/jbc.M608830200. [DOI] [PubMed] [Google Scholar]
- 33.Chen Y., Terajima M., Yang Y., Sun L., Ahn Y.H., Pankova D., Puperi D.S., Watanabe T., Kim M.P., Blackmon S.H., Rodriguez J., Liu H., Behrens C., Wistuba I.I., Minelli R., Scott K.L., Sanchez-Adams J., Guilak F., Pati D., Thilaganathan N., Burns A.R., Creighton C.J., Martinez E.D., Zal T., Grande-Allen K.J., Yamauchi M., Kurie J.M. Lysyl hydroxylase 2 induces a collagen cross-link switch in tumor stroma. J. Clin. Invest. 2015;125:1147–1162. doi: 10.1172/JCI74725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Saito T., Uzawa K., Terajima M., Shiiba M., Amelio A.L., Tanzawa H., Yamauchi M. Aberrant collagen cross-linking in human Oral squamous cell carcinoma. J. Dent. Res. 2019;98:517–525. doi: 10.1177/0022034519828710. [DOI] [PubMed] [Google Scholar]
- 35.Terajima M., Taga Y., Brisson B.K., Durham A.C., Sato K., Uzawa K., Saito T., Hattori S., Sørenmo K.U., Yamauchi M., Volk S.W. Collagen molecular phenotypic switch between non-neoplastic and neoplastic canine mammary tissues. Sci. Rep. 2021;11:1–15. doi: 10.1038/s41598-021-87380-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhou Y., Horowitz J.C., Naba A., Ambalavanan N., Atabai K., Balestrini J., Bitterman P.B., Corley R.A., Sen Ding B., Engler A.J., Hansen K.C., Hagood J.S., Kheradmand F., Lin Q.S., Neptune E., Niklason L., Ortiz L.A., Parks W.C., Tschumperlin D.J., White E.S., Chapman H.A., Thannickal V.J. Extracellular matrix in lung development, homeostasis and disease. Matrix Biol. 2018;73:77–104. doi: 10.1016/j.matbio.2018.03.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Miller E.J., Piez K.A. An accelerated single-column procedure for the automatic analysis of amino acids in collagen and elastin hydrolyzates. Anal. Biochem. 1966;16:320–326. doi: 10.1016/0003-2697(66)90161-8. [DOI] [PubMed] [Google Scholar]
- 38.Hulmes D.J.S., Miller A., Parry D.A.D., Piez K.A., Woodhead-Galloway J. Analysis of the primary structure of collagen for the origins of molecular packing. J. Mol. Biol. 1973;79:137–148. doi: 10.1016/0022-2836(73)90275-1. [DOI] [PubMed] [Google Scholar]
- 39.Miller E.J., Narkates A.J., Niemann M.A. Amino acid analysis of collagen hydrolysates by reverse-phase high-performance liquid chromatography of 9-fluorenylmethyl chloroformate derivatives. Anal. Biochem. 1990;190:92–97. doi: 10.1016/0003-2697(90)90139-Z. [DOI] [PubMed] [Google Scholar]
- 40.Basak T., Vega-Montoto L., Zimmerman L.J., Tabb D.L., Hudson B.G., Vanacore R.M. Comprehensive characterization of glycosylation and hydroxylation of basement membrane collagen IV by high-resolution mass spectrometry. J. Proteome Res. 2016;15:245–258. doi: 10.1021/acs.jproteome.5B00767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Terajima M., Taga Y., Nakamura T., Guo H.F., Kayashima Y., Maeda-Smithies N., Parag-Sharma K., Kim J.S., Amelio A.L., Mizuno K., Kurie J.M., Yamauchi M. Lysyl hydroxylase 2 mediated collagen post-translational modifications and functional outcomes. Sci. Rep. 2022;12:1–19. doi: 10.1038/s41598-022-18165-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ishikawa Y., Taga Y., Zientek K., Mizuno N., Salo A.M., Semenova O., Tufa S.F., Keene D.R., Holden P., Mizuno K., Gould D.B., Myllyharju J., Bächinger H.P. Type I and type V procollagen triple helix uses different subsets of the molecular ensemble for lysine posttranslational modifications in the rER. J. Biol. Chem. 2021;296 doi: 10.1016/j.jbc.2021.100453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Terajima M., Taga Y., Sricholpech M., Kayashima Y., Sumida N., Maeda N., Hattori S., Yamauchi M. Role of glycosyltransferase 25 domain 1 in type i collagen glycosylation and molecular phenotypes. Biochemistry. 2019;58:5040–5051. doi: 10.1021/acs.biochem.8b00984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sricholpech M., Perdivara I., Yokoyama M., Nagaoka H., Terajima M., Tomer K.B., Yamauchi M. Lysyl hydroxylase 3-mediated glucosylation in type I collagen: molecular loci and biological significance. J. Biol. Chem. 2012;287:22998–23009. doi: 10.1074/jbc.M112.343954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sarohi V., Srivastava S., Basak T. Comprehensive mapping and dynamics of site-specific prolyl-hydroxylation, lysyl-hydroxylation and lysyl O-glycosylation of collagens deposited in ECM during zebrafish heart regeneration. Front. Mol. Biosci. 2022;9:1–17. doi: 10.3389/fmolb.2022.892763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Terajima M., Perdivara I., Sricholpech M., Deguchi Y., Pleshko N., Tomer K.B., Yamauchi M. Glycosylation and cross-linking in bone type I collagen. J. Biol. Chem. 2014;289:22636–22647. doi: 10.1074/JBC.M113.528513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Sarohi V., Basak T. Perturbed post-translational modification (PTM) network atlas of collagen I during stent-induced neointima formation. J. Proteomics. 2023;276 doi: 10.1016/j.jprot.2023.104842. [DOI] [PubMed] [Google Scholar]
- 48.Sarohi V., Basak T. Decoding the comprehensive substrate-specificity and evidence of altered O-glycosylation in P4ha1 and P4ha2 deleted mutant mice. BioRxiv. 2023 [Google Scholar]
- 49.Chautard E., Fatoux-Ardore M., Ballut L., Thierry-Mieg N., Ricard-Blum S. MatrixDB, the extracellular matrix interaction database. Nucleic Acids Res. 2011;39:D235. doi: 10.1093/nar/gkq830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Naba A., Clauser K.R., Ding H., Whittaker C.A., Carr S.A., Hynes R.O. The extracellular matrix: tools and insights for the “omics” era. Matrix Biol. 2016;49:10–24. doi: 10.1016/J.MATBIO.2015.06.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Shao X., Taha I.N., Clauser K.R., Gao Y. (Tom), Naba A. MatrisomeDB: The ECM-protein knowledge database. Nucleic Acids Res. 2020;48:D1136–D1144. doi: 10.1093/nar/gkz849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Shao X., Gomez C.D., Kapoor N., Considine J.M., Grams C., Gao Y., Naba A. MatrisomeDB 2.0: updates to the ECM-protein knowledge database. Nucleic Acids Res. 2023;51(2023):D1519–D1530. doi: 10.1093/NAR/GKAC1009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Wilson R., Diseberg A.F., Gordon L., Zivkovic S., Tatarczuch L., Mackie E.J., Gorman J.J., Bateman J.F. Comprehensive profiling of cartilage extracellular matrix formation and maturation using sequential extraction and label-free quantitative proteomics. Mol. Cell. Proteomics. 2010;9:1296–1313. doi: 10.1074/mcp.M000014-MCP201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Terajima M., Taga Y., Cabral W.A., Liu Y., Nagasawa M., Sumida N., Kayashima Y., Chandrasekaran P., Han L., Maeda N., Perdivara I., Hattori S., Marini J.C., Yamauchi M. Cyclophilin B control of lysine post-translational modifications of skin type I collagen. PLoS Genet. 2019;15:1–26. doi: 10.1371/journal.pgen.1008196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Song E., Mechref Y. LC-MS/MS identification of the o-glycosylation and hydroxylation of amino acid residues of collagen α-1 (II) chain from bovine cartilage. J. Proteome Res. 2013;12:3599–3609. doi: 10.1021/PR400101T/SUPPL_FILE/PR400101T_SI_003.PDF. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Merl-Pham J., Basak T., Knüppel L., Ramanujam D., Athanason M., Behr J., Engelhardt S., Eickelberg O., Hauck S.M., Vanacore R., Staab-Weijnitz C.A. Quantitative proteomic profiling of extracellular matrix and site-specific collagen post-translational modifications in an in vitro model of lung fibrosis. Matrix Biol. plus. 2019;1 doi: 10.1016/J.MBPLUS.2019.04.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Garcia-Puig A., Mosquera J.L., Jiménez-Delgado S., García-Pastor C., Jorba I., Navajas D., Canals F., Raya A. Proteomics analysis of extracellular matrix remodeling during zebrafish heart regeneration. Mol. Cell. Proteomics. 2019;18:1745–1755. doi: 10.1074/mcp.RA118.001193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Barallobre-Barreiro J., Radovits T., Fava M., Mayr U., Lin W.Y., Ermolaeva E., Martínez-López D., Lindberg E.L., Duregotti E., Daróczi L., Hasman M., Schmidt L.E., Singh B., Lu R., Baig F., Siedlar A.M., Cuello F., Catibog N., Theofilatos K., Shah A.M., Crespo-Leiro M.G., Doménech N., Hübner N., Merkely B., Mayr M. Extracellular matrix in heart failure: role of ADAMTS5 in proteoglycan remodeling. Circulation. 2021;144:2021–2034. doi: 10.1161/CIRCULATIONAHA.121.055732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Padmanabhan Iyer R., Chiao Y.A., Flynn E.R., Hakala K., Cates C.A., Weintraub S.T., de Castro Brás L.E. Matrix metalloproteinase-9-dependent mechanisms of reduced contractility and increased stiffness in the aging heart. Proteomics - Clin. Appl. 2016;10:92–107. doi: 10.1002/prca.201500038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Suna G., Wojakowski W., Lynch M., Barallobre-Barreiro J., Yin X., Mayr U., Baig F., Lu R., Fava M., Hayward R., Molenaar C., White S.J., Roleder T., Milewski K.P., Gasior P., Buszman P.P., Buszman P., Jahangiri M., Shanahan C.M., Hill J., Mayr M. Extracellular matrix proteomics reveals interplay of aggrecan and aggrecanases in vascular remodeling of stented coronary arteries. Circulation. 2018;137:166–183. doi: 10.1161/CIRCULATIONAHA.116.023381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Sipila K.H., Drushinin K., Rappu P., Jokinen J., Salminen T.A., Salo A.M., Käpyla J., Myllyharju J., Heino J. Proline hydroxylation in collagen supports integrin binding by two distinct mechanisms. J. Biol. Chem. 2018;293:7645–7658. doi: 10.1074/jbc.RA118.002200. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data will be made available on request.