ABSTRACT
Here, we report the annotated genome of enterococcal phage G01. The G01 genome is 41,189 bp in length and contains 67 predicted open reading frames. Host range analysis revealed G01 can infect 28.6% (6/21) of Enterococcus faecalis strains tested and appears to not require the enterococcal phage infection protein PIPEF.
KEYWORDS: Enterococcus, bacteriophages, genomics
ANNOUNCEMENT
Enterococci are minority members of the human microbiota; however, during antibiotic therapy, multidrug-resistant (MDR) enterococci can outgrow and cause opportunistic infection (1–6). MDR enterococcal infections are difficult to treat, leading to renewed interest in bacteriophages (phages) as a therapeutic option (7–12). Isolation and characterization of novel enterococcal phages will inform the rational development of targeted phage therapeutics.
Phage G01 was isolated from river water collected near the Adi Ganga Ghat, Kolkata, West Bengal, India (22.520498, 88.340097). The water was filtered (0.45 µm), and isolation was performed through serial passage on Enterococcus faecalis OG1RF in top agar, as previously described (13). G01 phages were amplified by infecting a culture of E. faecalis OG1RF and purified using cesium chloride gradient centrifugation (14). DNA was extracted using phenol:chloroform:isoamyl alcohol extraction (13). The same sample of G01 DNA was used for Illumina and Oxford Nanopore sequencing. Libraries were prepared with the Illumina DNA prep kit and IDT 10 bp UDI indices (Illumina) or the PCR-free Oxford Nanopore Technologies Ligation Sequencing Kit with NEBNext Companion Module (Oxford Nanopore). Illumina paired-end 150 bp sequencing was performed on a NextSeq 2000. Demultiplexing, quality control, and adapter trimming were performed with bcl-convert v3.9.3. Single-end Nanopore sequencing was performed on a MinION. Super-accurate basecalling, demultiplexing, and adapter removal were done using Guppy v6.4.6. Illumina and Nanopore sequencing reads were utilized for a hybrid assembly using SPAdes v3.15.5 (15, 16). Assembly resulted in one 38,867 bp contig and one 2,399 bp contig with coverages of 5572.3× and 6525.5×, respectively. The assembly also yielded 310 contigs of less than 283 bp with coverages less than 1.5×. Analysis proceeded with the two largest, highest coverage contigs. These contigs were mapped against a single 41,883 bp nanopore sequencing read using CLC Genomics Workbench v20.0.4 to determine orientation. Using PCR primers facing outward from each end of the assembled sequence, it was determined that the G01 genome is circularly permuted during replication (17, 18). Open reading frames were predicted using CPT Galaxy PAP Structural Workflow v2023.1 and annotated with RASTtk and InterProScan (19–23). All tools were run with default parameters.
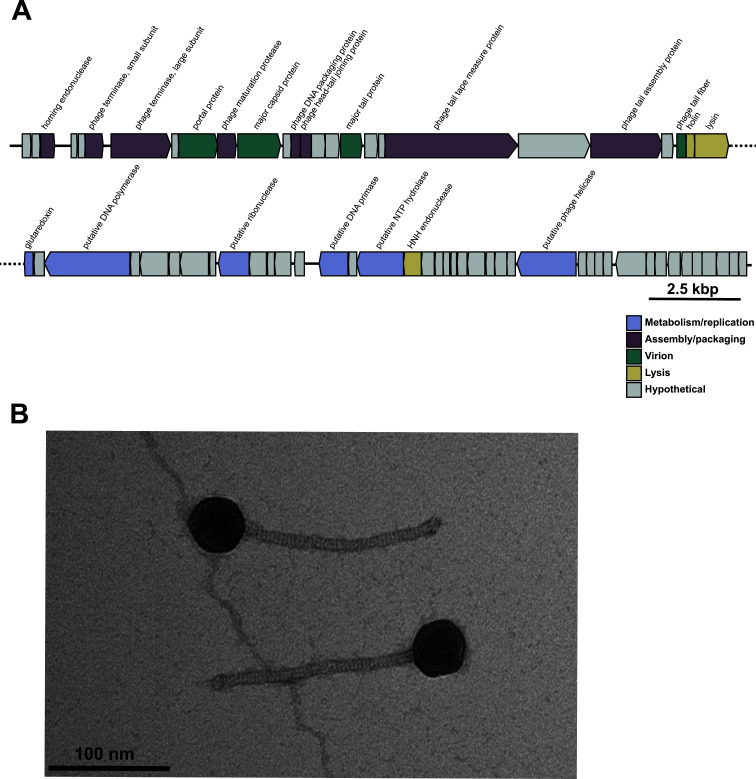
The G01 genome is 41,189 bp in length with 34% GC content and 67 predicted open reading frames (Fig. 1A). G01 shares 95.56% nucleotide identity over 75% of its genome with the phage MSF2 (accession number MK982307), the most similar complete phage genome identified by BLAST and a member of genus Efquatrovirus (24). Transmission electron microscopy determined G01 is a non-contractile tailed phage with siphophage morphology (Fig. 1B). Host range analysis was performed on E. faecalis strains embedded in top agar as described previously (25). Of the 21 strains tested, G01 formed plaques on 28.6% of E. faecalis strains tested and caused growth inhibition at low dilutions for an additional six strains (Table 1). G01 apparently infects E. faecalis OG1RF independent of the phage infection protein PIPEF, a previously identified E. faecalis phage receptor that determines phage tropism (Table 1) (13). However, infection appears to depend on the enterococcal polysaccharide antigen that is required for phage adsorption (25).
Fig 1.
(A) The G01 genome contains 67 predicted open reading frames. Annotations are indicated above the open reading frame, if available, and colored by the predicted function. (B) Purified G01 virions were stained with 0.75% uranyl formate and subjected to transmission electron microscopy on a ThermoFisher Tecnai G2 12 BioTwin equipped with a side-mount XR80 (8mp) CCD AMT camera and a low-mount NS15B (15mp) sCMOS AMT camera, with an accelerating voltage of 80 kV. Micrographs of the G01 virion reveal it has siphophage morphology with a tail approximately 169 nm long and a capsid approximately 45 nm in diameter. Measurements reflect analysis of 17 individual virions using ImageJ (26).
TABLE 1.
Dilutions of high titer G01 lysate were plated on E. faecalis strains for host range analysisa
| E. faecalis bacterial strain | Infection phenotype | Reference |
|---|---|---|
| ARO1 | None | (27) |
| ATCC 29212 | None | (28) |
| ATCC 4200 | 2.7 × 1010 PFU/mL | (29) |
| CH188 | None | (27) |
| Com6 | 1.1 × 1011 PFU/mL | (30) |
| D6 | 6 × 107 PFU/mL | (27) |
| DS16 | 10−1 | (31) |
| DS5 | 2.6 × 1010 PFU/mL | (27) |
| E1Sol | None | (27) |
| HIP11704 | 10−1 | (27) |
| JH2-2 | 100 | (32) |
| Merz96 | None | (27) |
| OG1RF | 3 × 1010 PFU/mL | (33) |
| OG1RF∆epaOX | None | (34) |
| OG1RF∆epaR | None | (35) |
| OG1RF∆pip | 5 × 1010 PFU/mL | (13) |
| SF28073 | None | (36) |
| T1 | 10−3 | (37) |
| T11 | None | (27) |
| T2 | 6 × 1010 PFU/mL | (27) |
| T3 | None | (27) |
| T8 | 100 | (27) |
| V583 | None | (38) |
| X98 | 100 | (39) |
If plaques formed on a given strain, the titer of the lysate on that strain is listed in the second column. If the strain showed growth inhibition at high dilutions without ever forming individual plaques, the lowest dilution in which clearing was seen is listed.
ACKNOWLEDGMENTS
Illumina and Nanopore library preparation and sequencing were performed by SeqCenter, Pittsburg, PA. Electron microscopy was performed by the University of Colorado Anschutz Medical Campus Electron Microscopy Core Facility, Aurora, CO.
Contributor Information
Breck A. Duerkop, Email: breck.duerkop@cuanschutz.edu.
Simon Roux, DOE Joint Genome Institute, USA.
DATA AVAILABILITY
Data are available at NCBI GenBank under the accession number OR797478. DNA sequencing reads have been deposited in the European Nucleotide Archive (ENA) under project accession number PRJEB68156. Illumina reads are deposited under run accession number ERR12205061. Nanopore reads are deposited under run accession number ERR12257447.
REFERENCES
- 1. Gilmore MS, Lebreton F, van Schaik W. 2013. Genomic transition of enterococci from gut commensals to leading causes of multidrug-resistant hospital infection in the antibiotic era. Curr Opin Microbiol 16:10–16. doi: 10.1016/j.mib.2013.01.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Donskey CJ, Chowdhry TK, Hecker MT, Hoyen CK, Hanrahan JA, Hujer AM, Hutton-Thomas RA, Whalen CC, Bonomo RA, Rice LB. 2000. Effect of antibiotic therapy on the density of vancomycin-resistant enterococci in the stool of colonized patients. N Engl J Med 343:1925–1932. doi: 10.1056/NEJM200012283432604 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Ostergaard L, Voldstedlund M, Bruun NE, Bundgaard H, Iversen K, Kober N, Christensen JJ, Rosenvinge FS, Jarlov JO, Moser C, Andersen CO, Coia J, Marmolin ES, Sogaard KK, Lemming L, Kober L, Fosbol EL. 2022. Temporal changes, patient characteristics, and mortality, according to microbiological cause of infective endocarditis: a nationwide study. J Am Heart Assoc 11:e025801. doi: 10.1161/JAHA.122.025801 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Vogkou CT, Vlachogiannis NI, Palaiodimos L, Kousoulis AA. 2016. The causative agents in infective endocarditis: a systematic review comprising 33,214 cases. Eur J Clin Microbiol Infect Dis 35:1227–1245. doi: 10.1007/s10096-016-2660-6 [DOI] [PubMed] [Google Scholar]
- 5. Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M, Gill SR, Nelson KE, Relman DA. 2005. Diversity of the human intestinal microbial flora. Science 308:1635–1638. doi: 10.1126/science.1110591 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Hayashi H, Takahashi R, Nishi T, Sakamoto M, Benno Y. 2005. Molecular analysis of jejunal, Ileal, caecal and recto-sigmoidal human colonic microbiota using 16S rRNA gene libraries and terminal restriction fragment length polymorphism. J Med Microbiol 54:1093–1101. doi: 10.1099/jmm.0.45935-0 [DOI] [PubMed] [Google Scholar]
- 7. Rubalskii E, Ruemke S, Salmoukas C, Boyle EC, Warnecke G, Tudorache I, Shrestha M, Schmitto JD, Martens A, Rojas SV, Ziesing S, Bochkareva S, Kuehn C, Haverich A. 2020. Bacteriophage therapy for critical infections related to cardiothoracic surgery. Antibiotics (Basel) 9:232. doi: 10.3390/antibiotics9050232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Suh GA, Lodise TP, Tamma PD, Knisely JM, Alexander J, Aslam S, Barton KD, Bizzell E, Totten KMC, Campbell JL, Chan BK, Cunningham SA, Goodman KE, Greenwood-Quaintance KE, Harris AD, Hesse S, Maresso A, Nussenblatt V, Pride D, Rybak MJ, Sund Z, van Duin D, Van Tyne D, Patel R, Antibacterial Resistance Leadership Group . 2022. Considerations for the use of phage therapy in clinical practice. Antimicrob Agents Chemother 66:e0207121. doi: 10.1128/AAC.02071-21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Anonymous . 2022. 2021 antibacterial agents in clinical and preclinical development: an overview and analysis. World Health Organization, Geneva. [Google Scholar]
- 10. Fiore E, Van Tyne D, Gilmore MS. 2019. Pathogenicity of enterococci. Microbiol Spectr 7. doi: 10.1128/microbiolspec.GPP3-0053-2018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Miller WR, Murray BE, Rice LB, Arias CA. 2016. Vancomycin-resistant enterococci: therapeutic challenges in the 21st century. Infect Dis Clin North Am 30:415–439. doi: 10.1016/j.idc.2016.02.006 [DOI] [PubMed] [Google Scholar]
- 12. Chiang H-Y, Perencevich EN, Nair R, Nelson RE, Samore M, Khader K, Chorazy ML, Herwaldt LA, Blevins A, Ward MA, Schweizer ML. 2017. Incidence and outcomes associated with infections caused by vancomycin-resistant enterococci in the United States: systematic literature review and meta-analysis. Infect Control Hosp Epidemiol 38:203–215. doi: 10.1017/ice.2016.254 [DOI] [PubMed] [Google Scholar]
- 13. Duerkop BA, Huo W, Bhardwaj P, Palmer KL, Hooper LV. 2016. Molecular basis for lytic bacteriophage resistance in enterococci. mBio 7:e01304-16. doi: 10.1128/mBio.01304-16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Maniatis T, Fritsch EF, Sambrook J. 1982. Isolation of Bacteriophage Λ and Plasmid DNA, p 75–96. In Molecular cloning: A laboratory manual. Vol. 1. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY. [Google Scholar]
- 15. Antipov D, Korobeynikov A, McLean JS, Pevzner PA. 2016. An algorithm for hybrid assembly of short and long reads. Bioinformatics 32:1009–1015. doi: 10.1093/bioinformatics/btv688 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Prjibelski A, Antipov D, Meleshko D, Lapidus A, Korobeynikov A. 2020. Using SPAdes de novo assembler. Curr Protoc Bioinformatics 70:e102. doi: 10.1002/cpbi.102 [DOI] [PubMed] [Google Scholar]
- 17. Casjens SR, Gilcrease EB. 2009. Determining DNA packaging strategy by analysis of the termini of the chromosomes in tailed-bacteriophage virions. Methods Mol Biol 502:91–111. doi: 10.1007/978-1-60327-565-1_7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Kalinski A, Black LW. 1986. End structure and mechanism of packaging of bacteriophage T4 DNA. J Virol 58:951–954. doi: 10.1128/JVI.58.3.951-954.1986 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, et al. 2008. The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75. doi: 10.1186/1471-2164-9-75 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Overbeek R, Olson R, Pusch GD, Olsen GJ, Davis JJ, Disz T, Edwards RA, Gerdes S, Parrello B, Shukla M, Vonstein V, Wattam AR, Xia F, Stevens R. 2014. The SEED and the rapid annotation of microbial genomes using subsystems technology (RAST). Nucleic Acids Res 42:D206–D214. doi: 10.1093/nar/gkt1226 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Brettin T, Davis JJ, Disz T, Edwards RA, Gerdes S, Olsen GJ, Olson R, Overbeek R, Parrello B, Pusch GD, Shukla M, Thomason JA, Stevens R, Vonstein V, Wattam AR, Xia F. 2015. RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep 5:8365. doi: 10.1038/srep08365 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Jones P, Binns D, Chang H-Y, Fraser M, Li W, McAnulla C, McWilliam H, Maslen J, Mitchell A, Nuka G, Pesseat S, Quinn AF, Sangrador-Vegas A, Scheremetjew M, Yong S-Y, Lopez R, Hunter S. 2014. Interproscan 5: genome-scale protein function classification. Bioinformatics 30:1236–1240. doi: 10.1093/bioinformatics/btu031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Rasche H, Bretaudeau A, Boudet M. 2019. Galaxy apollo tools
- 24. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. 1990. Basic local alignment search tool. J Mol Biol 215:403–410. doi: 10.1016/S0022-2836(05)80360-2 [DOI] [PubMed] [Google Scholar]
- 25. Chatterjee A, Johnson CN, Luong P, Hullahalli K, McBride SW, Schubert AM, Palmer KL, Carlson PE, Jr, Duerkop BA. 2019. Bacteriophage resistance alters antibiotic-mediated intestinal expansion of enterococci. Infect Immun 87:e00085-19. doi: 10.1128/IAI.00085-19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Schneider CA, Rasband WS, Eliceiri KW. 2012. NIH image to imageJ: 25 years of image analysis. Nat Methods 9:671–675. doi: 10.1038/nmeth.2089 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Palmer KL, Carniol K, Manson JM, Heiman D, Shea T, Young S, Zeng Q, Gevers D, Feldgarden M, Birren B, Gilmore MS. 2010. High-quality draft genome sequences of 28 Enterococcus sp. isolates. J Bacteriol 192:2469–2470. doi: 10.1128/JB.00153-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Kim EB, Kopit LM, Harris LJ, Marco ML. 2012. Draft genome sequence of the quality control strain Enterococcus faecalis ATCC 29212. J Bacteriol 194:6006–6007. doi: 10.1128/JB.01423-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Birkhaug KE. 1927. Bacteriologic studies in acute rheumatic fever with reference to soluble toxin production. Proc Soc Exp Biol Med 24:541–545. doi: 10.3181/00379727-24-3450 [DOI] [Google Scholar]
- 30. McBride SM, Fischetti VA, Leblanc DJ, Moellering RC, Gilmore MS. 2007. Genetic diversity among Enterococcus faecalis. PLOS ONE 2:e582. doi: 10.1371/journal.pone.0000582 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Tomich PK, An FY, Damle SP, Clewell DB. 1979. Plasmid-related transmissibility and multiple drug resistance in Streptococcus faecalis subsp. zymogenes strain DS16. Antimicrob Agents Chemother 15:828–830. doi: 10.1128/AAC.15.6.828 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Jacob AE, Hobbs SJ. 1974. Conjugal transfer of plasmid-borne multiple antibiotic resistance in Streptococcus faecalis var. zymogenes. J Bacteriol 117:360–372. doi: 10.1128/jb.117.2.360-372.1974 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Bourgogne A, Garsin DA, Qin X, Singh KV, Sillanpaa J, Yerrapragada S, Ding Y, Dugan-Rocha S, Buhay C, Shen H, et al. 2008. Large scale variation in Enterococcus faecalis illustrated by the genome analysis of strain OG1RF. Genome Biol 9:R110. doi: 10.1186/gb-2008-9-7-r110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Dale JL, Cagnazzo J, Phan CQ, Barnes AMT, Dunny GM. 2015. Multiple roles for Enterococcus faecalis glycosyltransferases in biofilm-associated antibiotic resistance, cell envelope integrity, and conjugative transfer. Antimicrob Agents Chemother 59:4094–4105. doi: 10.1128/AAC.00344-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Ho K, Huo W, Pas S, Dao R, Palmer KL. 2018. Loss-of-function mutations in epaR confer resistance to ϕNPV1 infection in Enterococcus faecalis OG1RF. Antimicrob Agents Chemother 62:e00758-18. doi: 10.1128/AAC.00758-18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Oprea SF, Zaidi N, Donabedian SM, Balasubramaniam M, Hershberger E, Zervos MJ. 2004. Molecular and clinical epidemiology of vancomycin-resistant Enterococcus faecalis. J Antimicrob Chemother 53:626–630. doi: 10.1093/jac/dkh138 [DOI] [PubMed] [Google Scholar]
- 37. Maekawa S, Yoshioka M, Kumamoto Y. 1992. Proposal of a new scheme for the serological typing of Enterococcus faecalis strains. Microbiol Immunol 36:671–681. doi: 10.1111/j.1348-0421.1992.tb02070.x [DOI] [PubMed] [Google Scholar]
- 38. Paulsen IT, Banerjei L, Myers GSA, Nelson KE, Seshadri R, Read TD, Fouts DE, Eisen JA, Gill SR, Heidelberg JF, et al. 2003. Role of mobile DNA in the evolution of vancomycin-resistant Enterococcus faecalis. Science 299:2071–2074. doi: 10.1126/science.1080613 [DOI] [PubMed] [Google Scholar]
- 39. Wheeler SM, Foley GE. 1945. Studies on the streptococci ("Enterococci") of Lancefield group D: II. Recovery of Lancefield group D streptococci from antemortem and postmortem cultures from infants and young children. Am J Dis Child 70:207–213. doi: 10.1001/archpedi.1945.02020220008002 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Data are available at NCBI GenBank under the accession number OR797478. DNA sequencing reads have been deposited in the European Nucleotide Archive (ENA) under project accession number PRJEB68156. Illumina reads are deposited under run accession number ERR12205061. Nanopore reads are deposited under run accession number ERR12257447.