ABSTRACT
The Gram-negative bacterium Coxiella burnetii is the causative agent of query fever in humans and coxiellosis in livestock. C. burnetii infects a variety of cell types, tissues, and animal species including mammals and arthropods, but there is much left to be understood about the molecular mechanisms at play during infection in distinct species. Human stimulator of interferon genes (STING) induces an innate immune response through the induction of type I interferons (IFNs), and IFN promotes or suppresses C. burnetii replication, depending on tissue type. Drosophila melanogaster contains a functional STING ortholog (Sting) which activates NF-κB signaling and autophagy. Here, we sought to address the role of D. melanogaster Sting during C. burnetii infection to uncover how Sting regulates C. burnetii infection in flies. We show that Sting-null flies exhibit higher mortality and reduced induction of antimicrobial peptides following C. burnetii infection compared to control flies. Additionally, Sting-null flies induce lower levels of oxidative stress genes during infection, but the provision of N-acetyl-cysteine (NAC) in food rescues Sting-null host survival. Lastly, we find that reactive oxygen species levels during C. burnetii infection are higher in Drosophila S2 cells knocked down for Sting compared to control cells. Our results show that at the host level, NAC provides protection against C. burnetii infection in the absence of Sting, thus establishing a role for Sting in protection against oxidative stress during C. burnetii infection.
KEYWORDS: STING, ROS, bacteria, pathogenesis, fly
INTRODUCTION
The Gram-negative bacterium Coxiella burnetii is the causative agent of query (Q) fever in humans and coxiellosis in livestock (1). Coxiellosis can lead to spontaneous abortions and stillbirths, and outbreaks are controlled through animal vaccination where available and segregation of periparturient animals (2). In humans, Q fever is largely transmitted via inhalation of pathogen-contaminated aerosols, such as from birth products of diseased animals (3, 4). Although transmission of C. burnetii to humans occurs most often through inhalation of pathogen-contaminated aersolos from diseased animals, arthropods are an important reservoir of the bacterium in the wild (1, 5). The ability of C. burnetii to infect a variety of hosts including arthropods indicates there are host-specific pathogen interactions that remain to be explored.
Our lab previously demonstrated that both Drosophila melanogaster Schneider 2 (S2) cells and adult flies are susceptible to the BSL2 Nine Mile phase II (NMII) strain of C. burnetii, and that this strain can replicate in adult Drosophila (6). Since then, we have built on the D. melanogaster model to dissect host immune responses to C. burnetii infection both in gene-specific and genome-wide studies (7, 8). Importantly, one of the advances of using the live D. melanogaster model to study innate immunity has led to the identification of Drosophila Sting and its relevance to human disease (8). Specifically, this study showed that cyclic dinucleotides (CDNs) and Listeria monocytogenes infection activate a Sting-mediated innate immune response through the immune deficiency (IMD) pathway in Drosophila (8–11). CDNs activate the human cGAS-stimulator of interferon genes (STING) signaling pathway, a central DNA-sensing defense mechanism against viral and bacterial infections, where cGAS produces CDNs upon recognition of double-stranded cytosolic DNA, which then bind to STING and activate a downstream interferon (IFN) signaling response (12, 13). The role of STING in DNA recognition has been studied extensively in the context of viral infections such as for adenovirus, which has evolved a variety of mechanisms to subvert STING-mediated IFN signaling (14–16). During bacterial infections, Chlamydia trachomatis and Staphylococcus aureus also induce a type I IFN response via STING (17–19). In contrast, induction of IFN genes is dependent on cell-type during C. burnetii infection (20). We hypothesized that the role of STING during C. burnetii infection may also be host dependent, specifically in our Drosophila model.
C. burnetii thrives in a broad range of tissues in part by evading host immune responses. Some of these evasion mechanisms include avoiding apoptosis activation (21–23), avoiding Toll-like receptor recognition (24–26), and suppressing cell proliferation (27, 28). C. burnetii also avoids host-mediated oxidative stress by inhibiting NADPH oxidase in neutrophils (21, 29), but the effect of oxidative stress in vivo during infection has not been explored. We investigated oxidative stress because several studies have shown that STING is involved with reactive oxygen species (ROS) during pathogenic infections. For example, murine gamma herpesvirus 68 induces an oxidative stress response that increases host ROS in vivo, which subsequently oxidizes murine STING at cysteine residue 147 to benefit the virus (30). Similarly, the Mycobacterium abscessus-R strain induces mitochondrial ROS in macrophages, which exerts a pro-bacterial effect via cGAS-STING-dependent IFN I induction (31). Oxidative stress is important in human health because a balance between ROS production and clearance is necessary for immunity (32). For example, diabetic patients have reduced ROS production, which causes a weakened immune response against intracellular pathogens; however, diabetic patients can also experience reduced clearance of ROS, which leads to prolonged inflammation and contributes further to weakened immunity (33). Although STING is known to affect infection outcomes, little is known about its role during C. burnetii infection.
In this study, we used a Drosophila Sting-null fly to determine the role of Sting during C. burnetii infection in vivo. We demonstrate that Sting-null flies have reduced ability to survive infection and induce antimicrobial peptide or antioxidant enzyme genes. At the cellular level we find that ROS increases during infection in Sting-knockdown S2 cells. We then show that feeding the antioxidant N-acetyl-cysteine (NAC) in Sting-null flies rescues survival, which indicates that Sting helps protect against increased cellular ROS during C. burnetii infection. Altogether, our results indicate that Sting is necessary for reducing bystander ROS damage during infection and to support bacterial growth.
MATERIALS AND METHODS
Drosophila melanogaster and C. burnetii infections
The D. melanogaster strains used in this study are w1118 (referred to as control) and Sting∆RG3 (Flybase ID: FBal0340352, referred to as Sting-null) (8). Fly stocks were maintained in standard meal agar fly food at 25°C and 65% humidity. Adult flies (2–7 days old) were injected with live 105 genome equivalent (GE)/fly as described in reference (6) with wild-type C. burnetii clone 4 RSA439. C. burnetii stocks were propagated in ACCM-2 media as in reference (34). Flies were anesthetized with CO2 and injected at the ventrolateral surface of the fly thorax with 23 nL of C. burnetii or PBS using a pulled 0.53-mm glass needle and an automatic nanoliter injector (Drummond Scientific, Broomall, PA). After injections, flies were either counted every day for survival experiments or collected at different times post-infection for quantifying bacterial load or gene expression. For survival experiments, flies were maintained at 25°C and 68% humidity, and food vials were changed every 2 days.
Measurement of bacterial load via GE
Quantification of C. burnetii from fly samples was performed as described in reference (35). Briefly, bacterial DNA was released from three flies per condition in a single tube containing Lysing D beads (MP Biomedicals) and 0.3 mL of molecular-grade water. Samples were homogenized once at 6 m/s for 20 s and centrifuged for 10 min at 500 rpm. The resulting supernatant was transferred to a fresh tube containing 0.1-mm zirconia beads, homogenized three times at 5.0 m/s for 30 s, and centrifuged for 1 min at 10,000 rpm. A 1:10 dilution was made for all samples in molecular-grade water for quantitative PCR quantification.
Gene expression
The relative expression of each antimicrobial peptide (AMP) gene was determined in adult flies by quantitative reverse transcription PCR (qRT-PCR) as described in reference (6). Briefly, five biological replicates of five flies each or six biological replicates of S2 cells per condition were lysed in Trizol Reagent (ThermoFisher 15596), and RNA was isolated by column purification (ZymoResearch R2050). Samples were treated with DNaseI (Invitrogen), and cDNA was synthesized using the iScript Reverse Transcriptase kit (Bio-Rad, Hercules, CA). qRT-PCR was performed using SYBR green PCR master mix (Bio-Rad). The following primer sequences were used for each gene: AttacinA (5-CACAATGTGGTGGGTCAGG-3 and 5-GGCACCATGACCAGCATT-3), CecropinA1 (5-GGACAATCGGAAGCTGGTT-3 and 5-TGTGCTGACCAACACGTTC-3), Drosocin (5-GCACAATGAAGTTCACCATCGT-3 and 5-CCACACCCATGGCAAAAAC-3), Defensin (5-GCCAGAACGCAGCCACAT-3 and 5-CGGTGTGGTTCCAGTTCCA-3), Drosomycin (5-CGTGAGAACCTTTTCCAATATGATG-3 and 5-TCCCAGGACCACCAGCAT-3), hid (5-ACGGCCATCCGAATCCGAAC-3 and 5-TGCTGCTGCCGGAAGAAGAAGTT-3), Eiger (5-GATGGTCTGGATTCCATTGC-3 and 5-TAGTCTGCGCCAACATCATC-3), Sod1 (5-GGAGTCGGTGATGTTGACCT-3 and 5-GTTCGGTGACAACACCAATG-3), Catalase (5-TTCTGGTTATCCCGTTGAGC-3 and 5-GGTAATGGCACCAGGAGAA-3), and RpII (5-TTGACGTAAGCATCACCTG-3 and 5-GAAGCGTTTCTCCAAACGAG-3).
D. melanogaster S2 cells and ROS assay
S2 cells (2.5 × 105) were plated as previously described (6). Cells were infected with mCherry-expressing C. burnetii at a multiplicity of infection (MOI) of 100 GE/cell. At the indicated times post-infection, three biological replicates per condition were collected and centrifuged 5 min at 300 × g to pellet the cells. All cells were stained in 10-µM dichlorodihydrofluorescein diacetate (DCFH-DA) for 30 min in the dark, washed with PBS, and re-plated into a clear-bottom black-well 96-well plate at 2.5 × 105 cells/well and imaged using a DMi8 Leica fluorescence microscope 1 h after plating. Fluorescence levels from the micrographs were quantified using ImageJ software (36, 37). Specifically, dichlorofluorescein (DCF) fluorescence levels (emitting 524 nm) were quantified from at least six images, and quantified fluorescence levels of infected cells were normalized to mock-infected cells (as presented in Fig. 3). At the indicted times post-infection, cells were transfected with non-targeting dsRNA or dsRNA targeting Sting using Cellfectin II (ThermoFisher) as previously described (8). Forty-eight hours post-transfection, cells were collected and imaged as described above. DCF and mCherry (emitting 610 nm) fluorescence was quantified by counting the number of double-positive cells (as presented in Fig. 5) from at least nine images per condition.
NAC fly food
The protocol for preparing fly food containing N-acetyl-cysteine was adapted from references (38, 39). Briefly, a fresh stock of NAC (Sigma, A7250-5G) at 1-g/mL concentration was prepared at the same time as preparing the standard meal agar fly food. A 1:10 dilution of the NAC stock was added directly to the fly food for a final NAC concentration of 10 mg/mL. The food was mixed thoroughly and aliquoted into standard plastic fly food vials. NAC food vials were stored at 4°C and used to feed flies during survival data collection, changing vials every 2 days as for non-NAC food.
Quantification and statistical analyses
Results shown are representative of at least three independent experiments. Circles in dot plots represent a biological replicate of a pool of five flies (Fig. 1 and 2), fluorescence quantification (Fig. 3), a well of cells (Fig. 4), or double-positive events (Fig. 5). Statistical analyses were completed using GraphPad Prism. Two-tailed unpaired t-tests assuming unequal variance were utilized to compare normally distributed pairwise quantitative data. Two-way analysis of variance with Tukey’s correction for multiple comparisons was used to compare multivariate data. All error bars represent standard error of the mean. For qRT-PCR data, statistical outliers were calculated using the interquartile range equation and removed (40). Survival curves represent three replicate experiments containing 40–50 flies per condition pooled together and analyzed by the log-rank (Mantel-Cox) test using GraphPad Prism to determine P values between infected genotypes as previously described (6, 7, 41).
RESULTS
Drosophila sting mediates C. burnetii infection
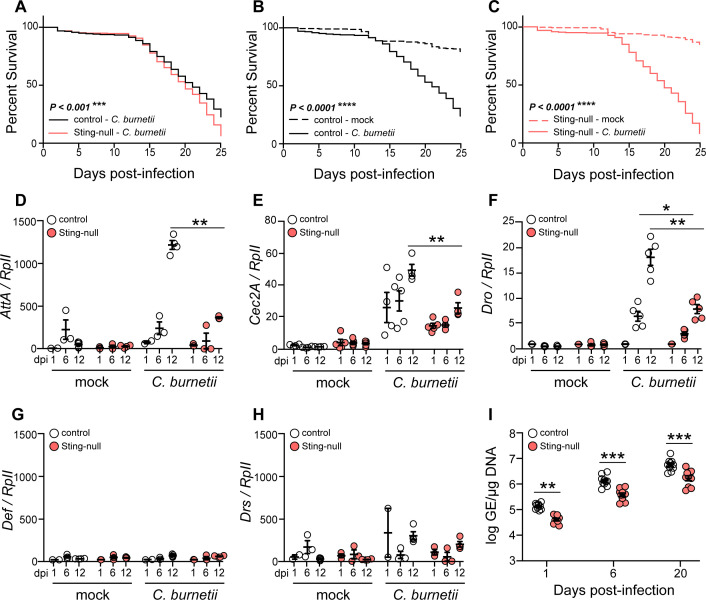
Previously, our lab showed that D. melanogaster can be used as a live animal model to study C. burnetii infection (6). Later, we demonstrated that Sting activates the NF-κB ortholog Relish during Listeria monocytogenes infection in D. melanogaster (8). Therefore, we hypothesized that Sting-null flies could be used to identify a role for Sting during C. burnetii infection. We found that Sting-null flies are more susceptible to C. burnetii infection than control flies (Fig. 1A through C). To test if Sting-null flies can induce an immune response against C. burnetii, we measured induction of AMP genes and found that Sting-null flies express the IMD-mediated AMPs Attacin, Cecropin, and Drosocin significantly less than control flies at 12 dpi (Fig. 1D through F), and significantly less than control flies at 6 dpi for Drosocin only (Fig. 1F). Unlike Attacin, Cecropin, and Drosocin, there is no difference in expression of the Toll-mediated AMPs Defensin and Drosomycin (Fig. 1G and H) during infection. To test if bacterial replication is the cause of host death, we measured bacterial load and found that C. burnetii replicates significantly less in Sting-null flies compared to control flies at 1, 6, and 20 dpi (Fig. 1I). Together, these results show that C. burnetii infection induces an IMD-mediated immune response through Sting. Interestingly, Sting-null flies have lower bacterial load compared to control, suggesting that Sting supports general health and maintenance of the host cell.
Fig 1.
Sting-null flies succumb to C. burnetii infection. (A–C) Adult Drosophila flies were mock-infected or infected with C. burnetii, and survival was monitored for 25 days, comparing the infected flies of each genotype (A) or mock- and C. burnetii-infected flies for each genotype (B and C). Expression of Attacin (D), Cecropin (E), Drosocin (F), Defensin (G), and Drosomycin (H) in control or Sting-null flies were determined at 1, 6, and 12 days post-infection (dpi) by reverse transcriptase quantitative real-time PCR, and the results were normalized to Drosophila RpII transcripts. (I) Bacterial loads were quantified at 1, 6, and 20 dpi by quantitative PCR. Asterisks denote significance: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Error bars indicate standard error of the mean. Each survival curve represents three independent experiments of 40–50 flies that were combined for a final survival curve and statistical analysis. For qRT-PCR, each circle represents an individual biological replicate of a pooled collection of five flies, and results are representative of three independent experiments.
Sting-null flies cannot induce cell death and oxidative stress genes
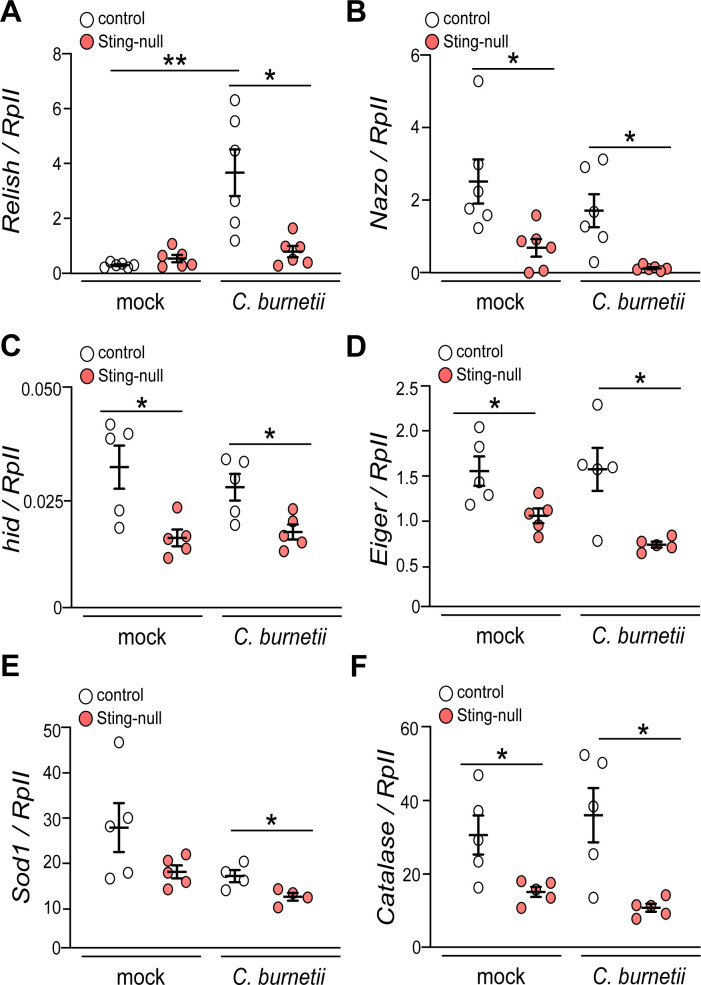
To test whether Sting is required for expression of genes related to cell death or oxidative stress during C. burnetii infection, we examined the induction of a variety of host genes including the Sting-mediated genes Relish (8, 42) and Nazo (43), the pro-apoptotic gene hid, and markers of increased ROS including oxidative stress genes Superoxide dismutase 1 (Sod1) and Catalase (44–47). While we observe Sting-mediated Relish induction during C. burnetii infection (Fig. 2A), infection does not induce Nazo, even though Nazo levels are lower in Sting-null flies (Fig. 2B). This observation is expected since Nazo has been shown to be induced during viral infections (43, 48). Regarding cell death genes, we find that expression of the pro-apoptotic gene hid is significantly reduced in Sting-null flies during C. burnetii infection compared to control flies at 12 dpi (Fig. 2C). The JNK pathway gene marker, Eiger, is also known to induce cell death in Drosophila (49, 50), and we observe significantly reduced expression of Eiger in Sting-null flies during C. burnetii infection compared to control flies (Fig. 2D). These data show that Sting-null flies are not inducing apoptosis or Eiger-mediated cell death pathways when flies begin succumbing to infection, suggesting that flies are not dying from these cell death pathways. Regarding the oxidative stress-related genes, we observe that expressions of Sod1 (Fig. 2E) and Catalase (Fig. 2F) are also significantly less induced in Sting-null flies during C. burnetii infection compared to control flies. Sod1 and Catalase are both important gene markers for oxidative stress homeostasis (47, 51, 52), indicating that during C. burnetii infection, Sting-null flies could be unable to reduce oxidative stress.
Fig 2.
Sting-null flies cannot induce cell death and oxidative stress genes. Expression of Relish (A), Nazo (B), hid (C), Eiger (D), Sod1 (E), and Catalase (F) at 12 dpi was determined in control and Sting-null flies by reverse transcriptase quantitative real time PCR, and the results were normalized to Drosophila RpII transcripts. Asterisks denote significance: *P < 0.05, **P < 0.01. Error bars indicate standard error of the mean. Each circle represents an individual biological replicate of a pooled collection of five flies, and results are representative of three independent experiments.
ROS are generated during C. burnetii infection and are higher in Sting-knockdown S2 cells
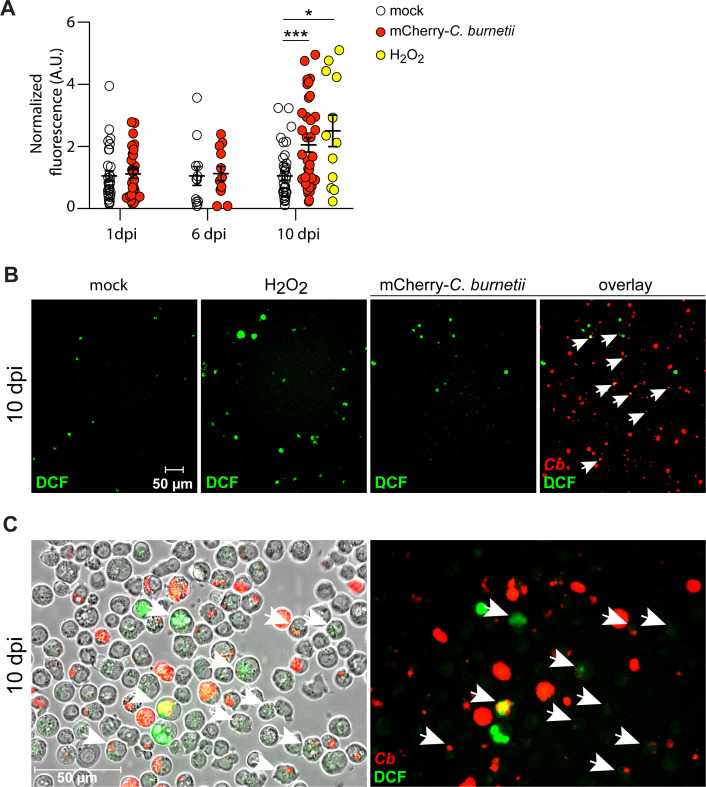
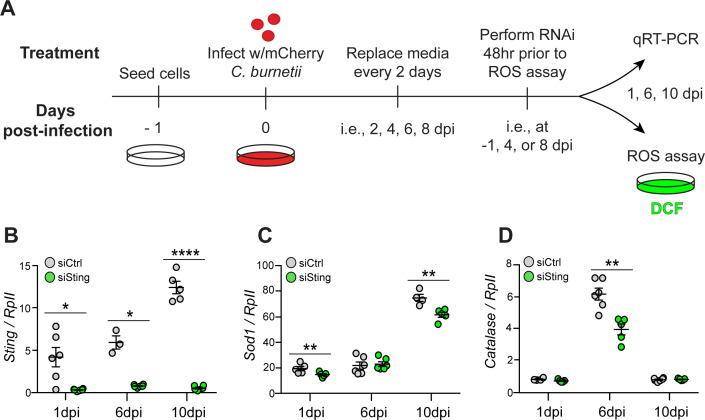
Since Sting-null flies exhibit reduced induction of antioxidant genes during C. burnetii infection, we hypothesized there is higher oxidative stress in the absence of Sting, which dysregulates the host’s ability to survive infection. To test this hypothesis, we first determined if ROS was produced during C. burnetii infection in S2 cells. We found that ROS levels are increased in C. burnetii-infected cells by 10 dpi (Fig. 3A). Representative images at 10 dpi show cells positive for ROS (green) and infection (mCherry-C. burnetii), indicated with white arrows (Fig. 3B). At higher magnification, double-positive cells, exhibiting both mCherry and DCF fluorescence, are observed clearly and are indicated with white arrows (Fig. 3C). We then knocked down Sting to measure gene expression and ROS at late times post-infection using RNAi (Fig. 4A and B). Sod1 expression is significantly lower in Sting-knockdown cells during C. burnetii infection at 1 and 10 dpi compared to control-knockdown cells (Fig. 4C). Similarly, Catalase expression is significantly lower in Sting-knockdown cells at 6 dpi (Fig. 4D). Compensatory signaling mechanisms could explain why the genes are not induced at the same time in infected cells. At the timepoints tested, we observe reduced expression of these two antioxidant enzyme genes in Sting-knockdown cells, which corroborates results in the fly.
Fig 3.
ROS is generated during C. burnetii infection in S2 cells. S2 cells were infected with mCherry-C. burnetii (100 GE/cell), and at 1, 6, and 10 dpi, cells were collected for DCFH-DA staining. Cells were imaged, and fluorescence intensity was quantified using ImageJ, where fluorescence of infected cells was normalized to mock (A). H2O2 was used as positive control to induce ROS in non-infected cells. Asterisks denote significance: *P < 0.05, ***P < 0.001. Error bars indicate standard error of the mean. Representative images of data quantification from 10 dpi are shown in panel B, where DCF fluorescence is shown for mock, H2O2, mCherry-C. burnetii, and overlay channels of infected cells, where white arrows point at double-positive cells (×200 magnification), and ×400 magnification (C). Data are representative of six individual images from three independent experiments.
Fig 4.
Expression of Sod1 is reduced in Sting-knockdown S2 cells during C. burnetii infection. Schematic of experimental design (A). At the indicated dpi, cells are transfected with control dsRNA (siCtrl) or dsRNA targeting Sting (siSting). Two days post-transfection, the ROS assay is performed. Expression of Sting (B), Sod1 (C), and Catalase (D) at 1, 6, and 10 dpi was determined in control and Sting-knockdown cells by reverse transcriptase quantitative real-time PCR, and the results were normalized to Drosophila RpII transcripts. Asterisks denote significance: *P < 0.05, **P < 0.01, ****P < 0.0001. Error bars indicate standard errors of the mean. Each circle represents an individual biological replicate of a well of cells, and results are representative of three independent experiments.
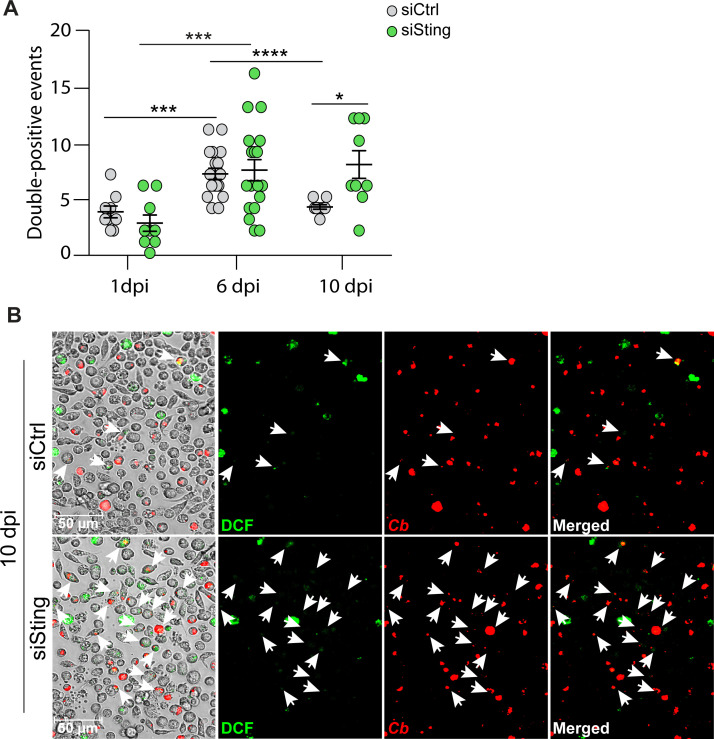
Lower expression of Sod1 and Catalase in Sting-knockdown cells suggest that ROS levels could be dysregulated. To test this hypothesis, we quantified the number of cells that were positive for both ROS and C. burnetii infection in control and Sting-knockdown cells and found that ROS double-positive cells are higher in Sting-knockdown cells compared to control knockdown at 10 dpi (Fig. 5A). While the number of double-positive cells increased between 1 and 6 dpi for both control and Sting-knockdown cells, we only observed a decrease in double-positive cells between 6 and 10 dpi in control cells and a sustained high level of ROS in Sting-knockdown cells at 10 dpi. This suggests that expression of Sting in C. burnetii-infected cells is required to protect against ROS levels in the cells. Representative images of double-positive cells are indicated with white arrows at 10 dpi for both control and Sting-knockdown cells (Fig. 5B). Taken together, our data show that ROS is higher in the absence of Sting during C. burnetii infection, suggesting that the mechanism of host death occurs through dysregulation of oxidative stress. We hypothesize that we could rescue Sting-null fly survival during C. burnetii infection by targeting host oxidative stress.
Fig 5.
ROS is higher in Sting-knockdown S2 cells during C. burnetii infection. The number of double-positive S2 cells with both mCherry-C. burnetii and DCF fluorescence was quantified during siCtrl or siSting knockdown at 1, 6, and 10 dpi (A). Asterisks denote significance: *P < 0.05, ***P < 0.001, ****P < 0.0001. Error bars indicate standard error of the mean. Representative images of siCtrl and siSting S2 cells at ×400 magnification at 10 dpi showing (from left to right) phase contrast image overlay, DCF fluorescence, mCherry-C. burnetii fluorescence, and merged panels (B), where white arrows point at double-positive cells. Data are representative of six individual images from three independent experiments.
NAC protects Sting-null flies against C. burnetii infection
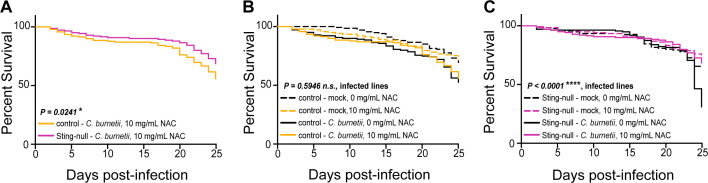
NAC can extend the lifespan Drosophila (38) and modulate ROS-induced gene expression (53). NAC is a powerful antioxidant and scavenger of free radicals that has been used to improve inflammation and oxidative stress in pneumonia patients (54) and reduce ROS-mediated damage during Heliobacter pylori infection (55). We infected flies with C. burnetii and continuously fed flies an experimental NAC-containing fly food throughout infection. We observed that Sting-null flies fed food containing NAC survive better during infection compared to control flies (Fig. 6A). The presence of NAC did not alter the survival of mock-infected flies of either genotype (Fig. 6B and C), as indicated by the dotted lines. We found that NAC did not alter survival during C. burnetii infection in control flies (Fig. 6B), but survival was increased in C. burnetii-infected Sting-null flies (Fig. 6C), as indicated by the respective solid lines. These results suggest that the ROS scavenger NAC is effective in rescuing host survival in the absence of Sting and that the presence of Sting reduces ROS during C. burnetii infection. Taken together, our results suggest that Sting has an important role in preserving host fitness and protecting against ROS-induced bystander damage during C. burnetii infection.
Fig 6.
NAC rescues host survival to C. burnetii infection in Sting-null hosts. Adult flies were mock- or C. burnetii-infected. Following infection, flies were placed on control or NAC-containing food, and survival was monitored for 25 days. Survival curves compare infected NAC-fed flies of both fly genotypes (A), control genotypes under all conditions (B), or Sting-null genotypes under all conditions (C). Asterisks denote significance: *P < 0.05, ****P < 0.0001. Each survival curve represents three independent experiments of 40–50 flies that were combined for a final survival curve and statistical analysis.
DISCUSSION
In this study, we demonstrate that Drosophila Sting plays an important role during C. burnetii infection. We show that in the absence of Sting, flies exhibit increased mortality to C. burnetii infection and have reduced ability to induce an IMD-specific innate immune response, and that bacterial load is lower compared to control flies. Interestingly, C. burnetii load was reduced even at one dpi, before the CCV is established. This suggests that Sting’s role in autophagy (42, 56) and phagosome formation (57) may reduce the ability of cells in vivo to be infected with C. burnetii via phagocytosis. At the cellular level, we show that ROS is generated during C. burnetii infection and find that the level of ROS is higher at late times post-infection in Sting-knockdown cells. Specifically, we observe that in control cells, the level of ROS is reduced between 6 and 10 dpi, while in Sting-knockdown cells, ROS levels remain unchanged. Similarly, in C. burnetii-infected Sting-null flies or Sting-knockdown cells, there is reduced induction of Sod1 and Catalase compared to control, suggesting a correlative mechanistic role for the protein products of these genes in reducing ROS during infection. Together, these results suggest that in the absence of Sting, ROS clearance is dysregulated and contributes to host death during C. burnetii infection. This hypothesis is corroborated by the rescuing of Sting-null fly death during infection when fed the antioxidant NAC, which is known for its broad ROS-scavenging properties (58). Moreover, the survival of NAC-fed Sting-null flies during C. burnetii infection was greater than control flies. We hypothesize, based on our in vitro data, that Sting-null flies have higher ROS levels than control flies during infection. NAC is a precursor for intracellular glutathione, which acts as a substrate of ROS scavenging enzymes (59). Additionally, NAC itself has reducing properties via thiol-disulfide exchange (60). The final product of ROS reduction is water (61), which may improve host survival in NAC-fed flies with higher initial ROS levels. Finally, we observe that during C. burnetii infection, Sting induces an IMD-mediated induction of AMP genes. These results corroborate our previous studies which showed that C. burnetii induces the IMD innate immune pathway in D. melanogaster (6) and that Sting functions primarily in the IMD-Relish-mediated axis to induce AMP gene expression (8). This is the first study to show that C. burnetii infection induces IMD-mediated AMPs in a Sting-dependent manner and that Sting is important for regulating ROS during C. burnetii infection in D. melanogaster.
Elevated ROS levels and impaired antioxidant defense systems are known to collectively cause oxidative stress (62). Additionally, ROS plays an important role in innate immunity because they can directly damage invading pathogens or elicit innate immune activation (63, 64). C. burnetii is known to evade oxidative stress by delaying phagolysosomal fusion and inhibiting NADPH oxidase (29, 65, 66), but also by expressing ROS-degrading enzymes (67, 68). Other mechanisms of defense against oxidative stress are reviewed in detail in Mertens and Samuel (69). It is generally accepted that ROS is beneficial to the host in clearing pathogenic bacteria such as Salmonella enterica serovar Typhimurium, where host-generated ROS induces cytoskeletal changes that promote antibacterial autophagy (70). In contrast, C. burnetii and other vacuole-forming pathogens like Chlamydia trachomatis sabotage ROS-induced autophagy to promote their own growth (71, 72). Lower bacterial burdens in the Sting-null fly suggest that one or more downstream cellular processes may be affected by the dysregulation of ROS clearance. In this manner, our Drosophila model provides a starting point to inspect a variety of signaling pathways at the ROS-STING interface during C. burnetii infection in mammalian systems.
Recently, ROS was shown to oxidize mouse STING and to impair IFN-mediated antiviral response (30), and ROS can activate human STING to promote antitumor immune responses (73). In Drosophila, Sting activates the IMD innate immune pathway through Relish (8, 42, 43), the human homolog of NF-κB. During infection, Relish inhibits c-Jun N-terminal kinase (JNK) signaling, and Relish was recently shown to inhibit JNK signaling in hematopoiesis in larval development, further connecting the IMD and JNK pathway in immunity (74). Importantly, Relish-mediated activation of IMD and JNK pathways are implicated in oxidative stress (75–77), where ROS activates JNK signaling to induce lipid droplets in fly mutants, and this phenotype is reduced when flies are fed N-acetyl amide (78). It is possible that JNK signaling plays additional roles during C. burnetii infection and remains to be investigated. Furthermore, dysfunctional Relish signaling resulted in greater fly mortality when infected with ROS-resistant pathogens (79), and C. burnetii employs multiple mechanisms to evade oxidative stress (69). Since Sting activates Relish [Fig. 2A and references (8, 42, 43)], our study further highlights the interplay of different signaling pathways that mediate ROS during infection.
The Drosophila IMD pathway is most closely related to the human tumor necrosis factor receptor pathway (80), and the only known homolog of TNF in Drosophila is Eiger, which can also be activated in response to infection and inflammation (49, 81–83). Eiger mutants have tolerance to C. burnetii infection (6), and here, we further show that Eiger is not induced during infection. We propose that maintaining a proper basal expression is necessary for protecting against infection, and this likely occurs through other mechanisms that remain to be identified. Eiger induces ROS production as a result of pyruvate dehydrogenase (PDH), where PDH regulates Eiger-mediated non-apoptotic cell death downstream of the JNK pathway (49, 84, 85). Since ROS readily activates JNK signaling (86, 87), it is likely that ROS production or clearance during C. burnetii infection will include the JNK and other metabolic pathways in addition to Sting-mediated IMD signaling.
In this study, we use D. melanogaster to uncover a new role for Sting in regulating oxidative stress during C. burnetii infection; however, it will be interesting to test if ROS or other oxidative stress processes are affected in the absence of STING in mammalian systems. We expect that results from mammalian studies will corroborate some phenotypes while also elucidating species-specific differences. Future studies can address how NAC or catalase modulates ROS levels or affects bacterial load in the absence of Sting. In our fly system, the reduced bacterial burden in the absence of Sting suggests that host factors are necessary for protecting the host against death but also for providing a healthy replicative niche for C. burnetii. Altogether, our study shows that in the absence of Sting, dysregulation of ROS is detrimental for both host and C. burnetii replication.
ACKNOWLEDGMENTS
The Coxiella burnetii clone 4 RSA439 used in this study was a gift from Dr. Robert A. Heinzen (Rocky Mountain Laboratories, National Institutes of Health [NIH], Hamilton, MT). We thank Dr. Laura Ahlers for the review of the manuscript and Dr. Manish Chauhan for helpful discussion of concepts and molecular recommendations throughout the project.
This research was supported by NIH/National Institute of Allergy and Infectious Diseases (NIAID) (grant R01 AI139051 to A.G.G.).
Conceptualization by R.M.G. and A.G.G.; methodology by R.M.G., A.G.G., A.O., and D.K.S.; fly food preparation by N.G.S. and Z.L., in consultation with R.M.G.; fly maintenance by R.M.G., O.M.H., and M.L.; reagents provided by H.Y. and A.O.; experiments, optimization, software, and statistics completed by R.M.G., M.L., and C.A.O.; figures made by R.M.G., in consultation with A.G.G.; writing by R.M.G. and revised by A.G.G., D.K.S., and A.O; visualization by R.M.G., D.K.S., and A.G.G.
Contributor Information
Alan G. Goodman, Email: alan.goodman@wsu.edu.
Sunny Shin, University of Pennsylvania Perelman School of Medicine, Philadelphia, Pennsylvania, USA.
REFERENCES
- 1. Maurin M, Raoult D. 1999. Q fever. Clin Microbiol Rev 12:518–553. doi: 10.1128/CMR.12.4.518 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Plummer PJ, McClure JT, Menzies P, Morley PS, Van den Brom R, Van Metre DC. 2018. Management of Coxiella burnetii infection in livestock populations and the associated zoonotic risk: a consensus statement. J Vet Intern Med 32:1481–1494. doi: 10.1111/jvim.15229 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Mori M, Roest HJ. 2018. Farming, Q fever and public health: agricultural practices and beyond. Arch Public Health 76:2. doi: 10.1186/s13690-017-0248-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Waag DM. 2007. Coxiella burnetii: host and bacterial responses to infection. Vaccine 25:7288–7295. doi: 10.1016/j.vaccine.2007.08.002 [DOI] [PubMed] [Google Scholar]
- 5. Duron O, Sidi-Boumedine K, Rousset E, Moutailler S, Jourdain E. 2015. The importance of ticks in Q fever transmission: what has (and has not) been demonstrated? Trends Parasitol 31:536–552. doi: 10.1016/j.pt.2015.06.014 [DOI] [PubMed] [Google Scholar]
- 6. Bastos RG, Howard ZP, Hiroyasu A, Goodman AG. 2017. Host and bacterial factors control susceptibility of Drosophila melanogaster to Coxiella burnetii infection. Infect Immun 85:e00218-17. doi: 10.1128/IAI.00218-17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Guzman RM, Howard ZP, Liu Z, Oliveira RD, Massa AT, Omsland A, White SN, Goodman AG. 2021. Natural genetic variation in Drosophila melanogster reveals genes associated with Coxiella burnetii infection. Genetics 217:iyab005. doi: 10.1093/genetics/iyab005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Martin M, Hiroyasu A, Guzman RM, Roberts SA, Goodman AG. 2018. Analysis of Drosophila STING reveals an evolutionarily conserved antimicrobial function. Cell Rep 23:3537–3550. doi: 10.1016/j.celrep.2018.05.029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Govind S. 2008. Innate immunity in Drosophila: pathogens and pathways. Insect Sci 15:29–43. doi: 10.1111/j.1744-7917.2008.00185.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Hoffman HM, Mueller JL, Broide DH, Wanderer AA, Kolodner RD. 2001. Mutation of a new gene encoding a putative pyrin-like protein causes familial cold autoinflammatory syndrome and Muckle-Wells syndrome. Nat Genet 29:301–305. doi: 10.1038/ng756 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Pandey UB, Nichols CD. 2011. Human disease models in Drosophila melanogaster and the role of the fly in therapeutic drug discovery. Pharmacol Rev 63:411–436. doi: 10.1124/pr.110.003293 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Barber GN. 2015. STING: infection, inflammation and cancer. Nat Rev Immunol 15:760–770. doi: 10.1038/nri3921 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Matz KM, Guzman RM, Goodman AG. 2019. The role of nucleic acid sensing in controlling microbial and autoimmune disorders. Int Rev Cell Mol Biol 345:35–136. doi: 10.1016/bs.ircmb.2018.08.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Anghelina D, Lam E, Falck-Pedersen E. 2016. Diminished innate antiviral response to adenovirus vectors in cGAS/STING-deficient mice minimally impacts adaptive immunity. J Virol 90:5915–5927. doi: 10.1128/JVI.00500-16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Lam E, Falck-Pedersen E. 2014. Unabated adenovirus replication following activation of the cGAS/STING-dependent antiviral response in human cells. J Virol 88:14426–14439. doi: 10.1128/JVI.02608-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Lam E, Stein S, Falck-Pedersen E. 2014. Adenovirus detection by the cGAS/STING/TBK1 DNA sensing cascade. J Virol 88:974–981. doi: 10.1128/JVI.02702-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Barker JR, Koestler BJ, Carpenter VK, Burdette DL, Waters CM, Vance RE, Valdivia RH. 2013. STING-dependent recognition of cyclic di-AMP mediates type I interferon responses during Chlamydia trachomatis infection. mBio 4:e00018-13. doi: 10.1128/mBio.00018-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Gries CM, Bruger EL, Moormeier DE, Scherr TD, Waters CM, Kielian T. 2016. Cyclic di-AMP released from Staphylococcus aureus biofilm induces a macrophage type I interferon response. Infect Immun 84:3564–3574. doi: 10.1128/IAI.00447-16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Zhang Y, Yeruva L, Marinov A, Prantner D, Wyrick PB, Lupashin V, Nagarajan UM. 2014. The DNA sensor, cyclic GMP-AMP synthase, is essential for induction of IFN-β during Chlamydia trachomatis infection . J Immunol 193:2394–2404. doi: 10.4049/jimmunol.1302718 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Hedges JF, Robison A, Kimmel E, Christensen K, Lucas E, Ramstead A, Jutila MA. 2016. Type I interferon counters or promotes Coxiella burnetii replication dependent on tissue. Infect Immun 84:1815–1825. doi: 10.1128/IAI.01540-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Howe D, Mallavia LP. 2000. Coxiella burnetii exhibits morphological change and delays phagolysosomal fusion after internalization by J774A.1 cells. Infect Immun 68:3815–3821. doi: 10.1128/IAI.68.7.3815-3821.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Lührmann A, Roy CR. 2007. Coxiella burnetii inhibits activation of host cell apoptosis through a mechanism that involves preventing cytochrome c release from mitochondria. Infect Immun 75:5282–5289. doi: 10.1128/IAI.00863-07 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Shannon JG, Howe D, Heinzen RA. 2005. Virulent Coxiella burnetii does not activate human dendritic cells: role of lipopolysaccharide as a shielding molecule. Proc Natl Acad Sci U S A 102:8722–8727. doi: 10.1073/pnas.0501863102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Conti F, Boucherit N, Baldassarre V, Trouplin V, Toman R, Mottola G, Mege JL, Ghigo E. 2014. Coxiella burnetii lipopolysaccharide blocks p38α-MAPK activation through the disruption of TLR-2 and TLR-4 Association. Front Cell Infect Microbiol 4:182. doi: 10.3389/fcimb.2014.00182 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Meghari S, Honstettre A, Lepidi H, Ryffel B, Raoult D, Mege JL. 2005. TLR2 is necessary to inflammatory response in Coxiella burnetii infection. Ann N Y Acad Sci 1063:161–166. doi: 10.1196/annals.1355.025 [DOI] [PubMed] [Google Scholar]
- 26. Zamboni DS, Campos MA, Torrecilhas ACT, Kiss K, Samuel JE, Golenbock DT, Lauw FN, Roy CR, Almeida IC, Gazzinelli RT. 2004. Stimulation of toll-like receptor 2 by Coxiella burnetii is required for macrophage production of pro-inflammatory cytokines and resistance to infection. J Biol Chem 279:54405–54415. doi: 10.1074/jbc.M410340200 [DOI] [PubMed] [Google Scholar]
- 27. Damrow TA, Williams JC, Waag DM. 1985. Suppression of in vitro lymphocyte proliferation in C57BL/10 ScN mice vaccinated with phase I Coxiella burnetii. Infect Immun 47:149–156. doi: 10.1128/iai.47.1.149-156.1985 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Koster FT, Williams JC, Goodwin JS. 1985. Cellular immunity in Q fever: specific lymphocyte unresponsiveness in Q fever endocarditis. J Infect Dis 152:1283–1289. doi: 10.1093/infdis/152.6.1283 [DOI] [PubMed] [Google Scholar]
- 29. Siemsen DW, Kirpotina LN, Jutila MA, Quinn MT. 2009. Inhibition of the human neutrophil NADPH oxidase by Coxiella burnetii. Microbes Infect 11:671–679. doi: 10.1016/j.micinf.2009.04.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Tao L, Lemoff A, Wang G, Zarek C, Lowe A, Yan N, Reese TA. 2020. Reactive oxygen species oxidize STING and suppress interferon production. Elife 9:e57837. doi: 10.7554/eLife.57837 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Kim B-R, Kim B-J, Kook Y-H, Kim B-J. 2020. Mycobacterium abscessus infection leads to enhanced production of type 1 interferon and NLRP3 inflammasome activation in murine macrophages via mitochondrial oxidative stress. PLoS Pathog 16:e1008294. doi: 10.1371/journal.ppat.1008294 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Pizzino G, Irrera N, Cucinotta M, Pallio G, Mannino F, Arcoraci V, Squadrito F, Altavilla D, Bitto A. 2017. Oxidative stress: harms and benefits for human health. Oxid Med Cell Longev 2017:1–13. doi: 10.1155/2017/8416763 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Dowey R, Iqbal A, Heller SR, Sabroe I, Prince LR. 2021. A bittersweet response to infection in diabetes; targeting neutrophils to modify inflammation and improve host immunity. Front Immunol 12:678771. doi: 10.3389/fimmu.2021.678771 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Omsland A, Cockrell DC, Howe D, Fischer ER, Virtaneva K, Sturdevant DE, Porcella SF, Heinzen RA. 2009. Host cell-free growth of the Q fever bacterium Coxiella burnetii. Proc Natl Acad Sci U S A 106:4430–4434. doi: 10.1073/pnas.0812074106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Sanchez SE, Goodman AG, Omsland A. 2021. Metabolic plasticity aids amphotropism of Coxiella burnetii. Infect Immun 89:e0013521. doi: 10.1128/IAI.00135-21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Chen W, Luan X, Yan Y, Wang M, Zheng Q, Chen X, Yu J, Fang J. 2020. CG8005 mediates transit-amplifying spermatogonial divisions via oxidative stress in Drosophila testes. Oxid Med Cell Longev 2020:2846727. doi: 10.1155/2020/2846727 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Graham C, Stefanatos R, Yek AEH, Spriggs RV, Loh SHY, Uribe AH, Zhang T, Martins LM, Maddocks ODK, Scialo F, Sanz A. 2022. Mitochondrial ROS signalling requires uninterrupted electron flow and is lost during ageing in flies. Geroscience 44:1961–1974. doi: 10.1007/s11357-022-00555-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Brack C, Bechter-Thüring E, Labuhn M. 1997. N-acetylcysteine slows down ageing and increases the life span of Drosophila melanogaster. Cell Mol Life Sci 53:960–966. doi: 10.1007/pl00013199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Sun X, Komatsu T, Lim J, Laslo M, Yolitz J, Wang C, Poirier L, Alberico T, Zou S. 2012. Nutrient-dependent requirement for SOD1 in lifespan extension by protein restriction in Drosophila melanogaster. Aging Cell 11:783–793. doi: 10.1111/j.1474-9726.2012.00842.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Seo S. 2006. A review and comparison of methods for detecting outliers in univariate data sets
- 41. Ahlers LRH, Trammell CE, Carrell GF, Mackinnon S, Torrevillas BK, Chow CY, Luckhart S, Goodman AG. 2019. Insulin potentiates JAK/STAT signaling to broadly inhibit flavivirus replication in insect vectors. Cell Rep 29:1946–1960. doi: 10.1016/j.celrep.2019.10.029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Liu Y, Gordesky-Gold B, Leney-Greene M, Weinbren NL, Tudor M, Cherry S. 2018. Inflammation-induced, STING-dependent autophagy restricts zika virus infection in the Drosophila brain. Cell Host Microbe 24:57–68. doi: 10.1016/j.chom.2018.05.022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Goto A, Okado K, Martins N, Cai H, Barbier V, Lamiable O, Troxler L, Santiago E, Kuhn L, Paik D, Silverman N, Holleufer A, Hartmann R, Liu J, Peng T, Hoffmann JA, Meignin C, Daeffler L, Imler JL. 2018. The kinase IKKβ regulates a STING- and NF-κB-dependent antiviral response pathway in Drosophila. Immunity 49:225–234. doi: 10.1016/j.immuni.2018.07.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Boriushkin E, Wang JJ, Zhang SX. 2014. Role of p58IPK in endoplasmic reticulum stress-associated apoptosis and inflammation. J Ophthalmic Vis Res 9:134–143. [PMC free article] [PubMed] [Google Scholar]
- 45. Chow CY, Wolfner MF, Clark AG. 2013. Using natural variation in Drosophila to discover previously unknown endoplasmic reticulum stress genes . Proc Natl Acad Sci U S A 110:9013–9018. doi: 10.1073/pnas.1307125110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Chow CY, Avila FW, Clark AG, Wolfner MF. 2015. Induction of excessive endoplasmic reticulum stress in the Drosophila male accessory gland results in infertility. PLoS One 10:e0119386. doi: 10.1371/journal.pone.0119386 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Hill VM, O’Connor RM, Sissoko GB, Irobunda IS, Leong S, Canman JC, Stavropoulos N, Shirasu-Hiza M. 2018. A bidirectional relationship between sleep and oxidative stress in Drosophila. PLoS Biol 16:e2005206. doi: 10.1371/journal.pbio.2005206 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Cai H, Holleufer A, Simonsen B, Schneider J, Lemoine A, Gad HH, Huang J, Huang J, Chen D, Peng T, Marques JT, Hartmann R, Martins NE, Imler J-L. 2020. 2'3'-cGAMP triggers a STING- and NF-κB-dependent broad antiviral response in Drosophila. Sci Signal 13:eabc4537. doi: 10.1126/scisignal.abc4537 [DOI] [PubMed] [Google Scholar]
- 49. Igaki T, Kanda H, Yamamoto-Goto Y, Kanuka H, Kuranaga E, Aigaki T, Miura M. 2002. Eiger, a TNF superfamily ligand that triggers the Drosophila JNK pathway. EMBO J 21:3009–3018. doi: 10.1093/emboj/cdf306 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Moreno E, Yan M, Basler K. 2002. Evolution of TNF signaling mechanisms: JNK-dependent apoptosis triggered by Eiger, the Drosophila homolog of the TNF superfamily. Curr Biol 12:1263–1268. doi: 10.1016/s0960-9822(02)00954-5 [DOI] [PubMed] [Google Scholar]
- 51. Missirlis F, Phillips JP, Jäckle H. 2001. Cooperative action of antioxidant defense systems in Drosophila. Curr Biol 11:1272–1277. doi: 10.1016/s0960-9822(01)00393-1 [DOI] [PubMed] [Google Scholar]
- 52. Wang Y, Branicky R, Noë A, Hekimi S. 2018. Superoxide dismutases: dual roles in controlling ROS damage and regulating ROS signaling. J Cell Biol 217:1915–1928. doi: 10.1083/jcb.201708007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Niraula P, Kim MS. 2019. N-acetylcysteine extends lifespan of Drosophila via modulating ROS scavenger gene expression. Biogerontology 20:533–543. doi: 10.1007/s10522-019-09815-4 [DOI] [PubMed] [Google Scholar]
- 54. Zhang Q, Ju Y, Ma Y, Wang T. 2018. N-acetylcysteine improves oxidative stress and inflammatory response in patients with community acquired pneumonia: a randomized controlled trial. Medicine 97:e13087. doi: 10.1097/MD.0000000000013087 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Xie C, Yi J, Lu J, Nie M, Huang M, Rong J, Zhu Z, Chen J, Zhou X, Li B, Chen H, Lu N, Shu X. 2018. N-acetylcysteine reduces ROS-mediated oxidative DNA damage and PI3K/AKT pathway activation induced by Helicobacter pylori infection. Oxid Med Cell Longev 2018:1874985. doi: 10.1155/2018/1874985 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Gui X, Yang H, Li T, Tan X, Shi P, Li M, Du F, Chen ZJ. 2019. Autophagy induction via STING trafficking is a primordial function of the cGAS pathway. Nature 567:262–266. doi: 10.1038/s41586-019-1006-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Chen T, Feng Y, Sun W, Zhao G, Wu H, Cheng X, Zhao F, Zhang L, Zheng Y, Zhan P, Zhao W, Liu B, Gao C. 2023. The nucleotide receptor STING translocates to the phagosomes to negatively regulate anti-fungal immunity. Immunity 56:1727–1742. doi: 10.1016/j.immuni.2023.06.002 [DOI] [PubMed] [Google Scholar]
- 58. Zhitkovich A. 2019. N-acetylcysteine: antioxidant, aldehyde scavenger, and more. Chem Res Toxicol 32:1318–1319. doi: 10.1021/acs.chemrestox.9b00152 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Circu ML, Aw TY. 2008. Glutathione and apoptosis. Free Radic Res 42:689–706. doi: 10.1080/10715760802317663 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Zafarullah M, Li WQ, Sylvester J, Ahmad M. 2003. Molecular mechanisms of N-acetylcysteine actions. Cell Mol Life Sci 60:6–20. doi: 10.1007/s000180300001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Juan CA, Pérez de la Lastra JM, Plou FJ, Pérez-Lebeña E. 2021. The chemistry of reactive oxygen species (ROS) revisited: outlining their role in biological macromolecules (DNA, lipids and proteins) and induced pathologies. Int J Mol Sci 22:4642. doi: 10.3390/ijms22094642 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Sies H. 1997. Oxidative stress: oxidants and antioxidants. Exp Physiol 82:291–295. doi: 10.1113/expphysiol.1997.sp004024 [DOI] [PubMed] [Google Scholar]
- 63. Kohchi C, Inagawa H, Nishizawa T, Soma G-I. 2009. ROS and innate immunity. Anticancer Res 29:817–821. [PubMed] [Google Scholar]
- 64. Paiva CN, Bozza MT. 2014. Are reactive oxygen species always detrimental to pathogens? Antioxid Redox Signal 20:1000–1037. doi: 10.1089/ars.2013.5447 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Bitew MA, Hofmann J, De Souza DP, Wawegama NK, Newton HJ, Sansom FM. 2020. SdrA, an NADP(H)-regenerating enzyme, is crucial for Coxiella burnetii to resist oxidative stress and replicate intracellularly. Cell Microbiol 22:e13154. doi: 10.1111/cmi.13154 [DOI] [PubMed] [Google Scholar]
- 66. Brennan RE, Russell K, Zhang G, Samuel JE. 2004. Both inducible nitric oxide synthase and NADPH oxidase contribute to the control of virulent phase I Coxiella burnetii infections. Infect Immun 72:6666–6675. doi: 10.1128/IAI.72.11.6666-6675.2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Heinzen RA, Frazier ME, Mallavia LP. 1990. Nucleotide sequence of Coxiella burnetii superoxide dismutase. Nucleic Acids Res 18:6437. doi: 10.1093/nar/18.21.6437 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Heinzen RA, Frazier ME, Mallavia LP. 1992. Coxiella burnetii superoxide dismutase gene: cloning, sequencing, and expression in Escherichia coli. Infect Immun 60:3814–3823. doi: 10.1128/iai.60.9.3814-3823.1992 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Mertens K, Samuel JE. 2012. Defense mechanisms against oxidative stress in Coxiella burnetii: adaptation to a unique intracellular niche. Adv Exp Med Biol 984:39–63. doi: 10.1007/978-94-007-4315-1_3 [DOI] [PubMed] [Google Scholar]
- 70. Huang J, Canadien V, Lam GY, Steinberg BE, Dinauer MC, Magalhaes MAO, Glogauer M, Grinstein S, Brumell JH. 2009. Activation of antibacterial autophagy by NADPH oxidases. Proc Natl Acad Sci U S A 106:6226–6231. doi: 10.1073/pnas.0811045106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Gutierrez MG, Vázquez CL, Munafó DB, Zoppino FCM, Berón W, Rabinovitch M, Colombo MI. 2005. Autophagy induction favours the generation and maturation of the Coxiella-replicative vacuoles. Cell Microbiol 7:981–993. doi: 10.1111/j.1462-5822.2005.00527.x [DOI] [PubMed] [Google Scholar]
- 72. Yasir M, Pachikara ND, Bao X, Pan Z, Fan H. 2011. Regulation of chlamydial infection by host autophagy and vacuolar ATPase-bearing organelles. Infect Immun 79:4019–4028. doi: 10.1128/IAI.05308-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Hu Z, Teng XL, Zhang T, Yu X, Ding R, Yi J, Deng L, Wang Z, Zou Q. 2021. SENP3 senses oxidative stress to facilitate STING-dependent dendritic cell antitumor function. Molecular Cell 81:940–952. doi: 10.1016/j.molcel.2020.12.024 [DOI] [PubMed] [Google Scholar]
- 74. Ramesh P, Dey NS, Kanwal A, Mandal S, Mandal L. 2021. Relish plays a dynamic role in the niche to modulate Drosophila blood progenitor homeostasis in development and infection. Elife 10:e67158. doi: 10.7554/eLife.67158 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Maitra U, Scaglione MN, Chtarbanova S, O’Donnell JM. 2019. Innate immune responses to paraquat exposure in a Drosophila model of Parkinson’s disease. Sci Rep 9:12714. doi: 10.1038/s41598-019-48977-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Nakano H, Nakajima A, Sakon-Komazawa S, Piao JH, Xue X, Okumura K. 2006. Reactive oxygen species mediate crosstalk between NF-κB and JNK. Cell Death Differ 13:730–737. doi: 10.1038/sj.cdd.4401830 [DOI] [PubMed] [Google Scholar]
- 77. Park JM, Brady H, Ruocco MG, Sun H, Williams D, Lee SJ, Kato T, Richards N, Chan K, Mercurio F, Karin M, Wasserman SA. 2004. Targeting of TAK1 by the NF-κB protein relish regulates the JNK-mediated immune response in Drosophila. Genes Dev 18:584–594. doi: 10.1101/gad.1168104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Liu L, Zhang K, Sandoval H, Yamamoto S, Jaiswal M, Sanz E, Li Z, Hui J, Graham BH, Quintana A, Bellen HJ. 2015. Glial lipid droplets and ROS induced by mitochondrial defects promote neurodegeneration. Cell 160:177–190. doi: 10.1016/j.cell.2014.12.019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Ryu JH, Ha EM, Oh CT, Seol JH, Brey PT, Jin I, Lee DG, Kim J, Lee D, Lee WJ. 2006. An essential complementary role of NF-κB pathway to microbicidal oxidants in Drosophila gut immunity. EMBO J 25:3693–3701. doi: 10.1038/sj.emboj.7601233 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Hoffmann JA. 2003. The immune response of Drosophila. Nature 426:33–38. doi: 10.1038/nature02021 [DOI] [PubMed] [Google Scholar]
- 81. Brandt SM, Dionne MS, Khush RS, Pham LN, Vigdal TJ, Schneider DS. 2004. Secreted bacterial effectors and host-produced Eiger/TNF drive death in a Salmonella-infected fruit fly. PLoS Biol 2:e418. doi: 10.1371/journal.pbio.0020418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Mabery EM, Schneider DS. 2010. The Drosophila TNF ortholog Eiger is required in the fat body for a robust immune response. J Innate Immun 2:371–378. doi: 10.1159/000315050 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Schneider DS, Ayres JS, Brandt SM, Costa A, Dionne MS, Gordon MD, Mabery EM, Moule MG, Pham LN, Shirasu-Hiza MM. 2007. Drosophila eiger mutants are sensitive to extracellular pathogens. PLoS Pathog 3:e41. doi: 10.1371/journal.ppat.0030041 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Igaki T, Miura M. 2014. The Drosophila TNF ortholog Eiger: emerging physiological roles and evolution of the TNF system. Semin Immunol 26:267–274. doi: 10.1016/j.smim.2014.05.003 [DOI] [PubMed] [Google Scholar]
- 85. Kanda H, Igaki T, Okano H, Miura M. 2011. Conserved metabolic energy production pathways govern Eiger/TNF-induced nonapoptotic cell death. Proc Natl Acad Sci U S A 108:18977–18982. doi: 10.1073/pnas.1103242108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Shen H-M, Liu Z. 2006. JNK signaling pathway is a key modulator in cell death mediated by reactive oxygen and nitrogen species. Free Radic Biol Med 40:928–939. doi: 10.1016/j.freeradbiomed.2005.10.056 [DOI] [PubMed] [Google Scholar]
- 87. Ueda S, Masutani H, Nakamura H, Tanaka T, Ueno M, Yodoi J. 2002. Redox control of cell death. Antioxid Redox Signal 4:405–414. doi: 10.1089/15230860260196209 [DOI] [PubMed] [Google Scholar]