Abstract
Chromosomal abnormalities are the most common causes of early pregnancy losses (EPLs). In this study, we aimed to evaluate the incidence and spectrum of chromosomal abnormalities in EPLs and correlate them with different clinical characteristics. We performed Quantitative Fluorescent PCR (QF-PCR), followed by subtelomeric Multiplex Ligation Probe Amplification (MLPA) analysis to detect chromosomal abnormalities in 900 products of conceptions (POCs) from EPLs collected over a period of 10 years.
Chromosomal abnormalities were present in 56.25% of uncontaminated EPLs, with significantly higher incidence in women ≥36 years (71.37%, p<0.0001) in comparison to women ≤30 years of age (43.40%). Trisomies were also more common in women ≥36 years (79.68%, p<0.0001) than in those ≤30 years of age (48.70%). In contrast, triploidy and monosomies were more prevalent in women ≤30 years of age (26.09%, p<0.0001 and 16.52%, p=0.0066 respectively) than in women ≥36 years of age (6.42% and 6.42% respectively). Trisomy 16 was more common in women ≤30 (39.29%, p=0.0009) than in those ≥36 years of age (16.78%), while trisomy 22 was predominant among women ≥36 (23.49%, p=0.013), and was not present in the group of women ≤30 years of age. The frequency of chromosomal abnormalities in POCs from women with sporadic (61.19%) was higher than in those with recurrent EPLs (55.21%). This difference, however, was not statistically significant (p=0.164). Although some differences in the chromosomal aneuploidy rates among women with different ABO blood groups, as well as among 6–8 and 9–11 gestational week EPLs were observed, further larger studies are required to confirm these findings.
In conclusion, our study enriches the knowledge about chromosomal abnormalities as a cause of EPLs and confirms the higher incidence of foetal chromosomal abnormalities in EPLs in women of older reproductive age. Furthermore, it shows that using QF-PCR and MLPA methodologies, a high detection rate of chromosomal abnormalities in EPLs can be reached.
Keywords: Chromosomal abnormality, early pregnancy loss (EPL), QF-PCR, MLPA
INTRODUCTION
Human reproduction is characterized by a high rate of abnormal conceptions, most of which are spontaneously eliminated before the pregnancy is clinically recognized. Pregnancy loss (PL), spontaneous abortion or miscarriage refers to the spontaneous (unintended) loss of pregnancy before the foetus reaches viability, i.e. before twenty weeks of gestation [1]. Early pregnancy loss (EPL) represents a spontaneous loss of pregnancy before the 12th week of gestation (first trimester). In the United States, recurrent pregnancy loss (RPL) is defined as having two or more consecutive failed clinical pregnancies, documented by ultrasound or histopathology, while in the United Kingdom it is defined as having three or more consecutive early pregnancy losses [2,3]. Up to 15% of all clinically recognized pregnancies are miscarried, and nearly 2% of the couples that are trying to conceive experience RPL.
The aetiology of EPL can include various factors, such as maternal endocrine dysregulation, anatomical abnormalities of the uterus, implantation factors, various infections during the pregnancy and foetal chromosomal abnormalities. About 40–65% of the miscarried foetuses are associated with various chromosomal abnormalities, the most common of which are chromosomal trisomies, followed by polyploidies and monosomy X [4,5,6].
Some studies have detected nearly equal frequencies in sporadic and recurrent EPLs, while others show a lower rate of chromosomal abnormalities in RPLs [7, 8]. Due to age-related oogenesis errors, advanced maternal age represents a considerable risk factor for EPL [9]. Indeed, many studies have confirmed a higher incidence of POCs with chromosomal abnormalities in women with advanced age. Published data indicate that foetal triploidies and monosomies are more common in younger women, while trisomies are more prevalent in older women [10, 11]. A limited number of data provide insight into the distribution of chromosomal abnormalities in POCs in reference to the week of gestation. Up to now, there is no published study that correlates the maternal ABO blood groups and Rhesus factor with foetal chromosomal abnormality, even though the available data indicate that incompatible mating and adverse pregnancy outcomes may correlate with ABO blood groups [12].
Chromosomal karyotyping has been the gold standard for studying the chromosomes for many decades, but this method is hampered by a high culture failure or maternal cell contamination. Currently, chromosomal microarrays represent a first-tier method for chromosomal abnormality investigations, however due to the high price, it is rarely used for EPLs. On the other side, quantitative fluorescent PCR (QF-PCR) and multiplex ligation probe amplification (MLPA) methods have emerged to determine any chromosomal abnormalities in POCs. This is due to their lower cost, faster reporting times, and accurate results [13, 14].
Here we present the results of our 10 year study of EPLs using QF-PCR, followed by subtlomeric MLPA, including the distribution of foetal chromosomal abnormalities in relation to the clinical characteristics of women experiencing EPLs.
MATERIALS AND METHODS
Study subjects
Our study included 900 POC samples from women who experienced EPLs (gestational age ≤12 weeks). POC samples, previously selected by a gynaecologist/pathologist [15] and accompanied with maternal whole blood sample, were referred for analysis of chromosomal abnormalities to the Research Centre for Genetic Engineering and Biotechnology “Georgi D. Efremov”, at the Macedonian Academy of Sciences and Arts, Skopje. Signed informed consent was obtained from all participants in this study. The study has been approved by the ethical committee of the Macedonian Academy of Sciences and Arts (09-1047/6 from 04.05.2016).
Table 1 displays the clinical characteristics of the women with EPLs, including maternal age, ethnic origin, history of previous EPL, previous live birth, maternal ABO blood group, Rhesus (Rh) factor, and the gestational week (gw) of the EPLs studied. The samples were categorized into three groups according to the maternal age: ≤30, 31–35, and ≥36 years. The majority of patients were of Macedonian (n=528) ethnic origin as well as Albanian (n=208). Gestational ages of the POCs consisted of samples from gw=6 to gw=11 (mean gestational age 8.5 weeks).
Table 1.
Characteristics of the study groups
| Characteristic | Study groups | All cases | Macedonian | Albanian | Other | ||||
|---|---|---|---|---|---|---|---|---|---|
| n (%) | MA±SD | n (%) | MA±SD | n (%) | MA±SD | n (%) | MA±SD | ||
| n=768 | 32.9±5.35 | n=528 | 34.14±4.90 | n=208 | 33.16±5.46 | n=32 | 31.90±5.68 | ||
| Maternal age | ≤30 | 265 (34.50) | 27.19±2.56 | 131 (24.81) | 27.82±2.18 | 121 (58.17) | 26.53±2.72 | 13 (40.62) | 26.92±3.22 |
| 31–35 | 241 (31.37) | 32.87±1.39 | 178 (33.71) | 32.89±1.40 | 52 (25.00) | 32.82±1.41 | 11 (34.37) | 32.27±0.90 | |
| ≥36 | 262 (34.11) | 38.96±2.39 | 219 (41.47) | 38.94±2.31 | 35 (16.82) | 39.02±2.61 | 8 (25.00) | 39.5±3.46 | |
| ND | / | / | / | / | / | / | / | / | |
| History of PL | Sporadic | 268 (34.89) | 33.16±5.26 | 202 (38,25) | 33.95±4.78 | 48 (23,07) | 29.60±5.55 | 18 (56,25) | 33.61±5.81 |
| RPL | 259 (33,72) | 33.1±5.24 | 164 (31,06) | 34.67±4.71 | 86 (41,34) | 30.56±5.11 | 9 (28,12) | 28.77±4.14 | |
| ND | 241 (31,38) | / | 162 (30,68) | / | 74 (35,57) | / | 5 (15,62) | / | |
| Previous live birth | Yes | 142 (18,48) | 34.66±4.75 | 95 (17,99) | 35.76±4.48 | 41 (19,71) | 32.82±4.64 | 6 (18,75) | 29.66±2.25 |
| No | 383 (49,86) | 32.57±5.32 | 269 (50,94) | 33.77±4.76 | 93 (44,71) | 29.07±5.15 | 21 (65,62) | 32.66±6.26 | |
| ND | 243 (31,64) | / | 164 (31,06) | / | 74 (35,57) | / | 5 (15,62) | / | |
| Maternal ABO group | 0 | 183 (23,821) | 33.09±5.52 | 123 (23,29) | 34.73±4.98 | 55 (26,44) | 29.38±4.98 | 5 (15,62) | 33.60±4.72 |
| A | 183 (23,82) | 33.61±4.95 | 138 (26,13) | 34.41±4.60 | 40 (19,23) | 31.32±4.99 | 5 (15,62) | 30.00±7.68 | |
| B | 67 (8,72) | 32.82±5.03 | 46 (8,71) | 33.91±4.63 | 15 (7,21) | 30.46±5.99 | 6 (18,75) | 30.33±2.42 | |
| AB | 36 (4,68) | 31.63±5.57 | 21 (3,97) | 33.33±5.38 | 9 (4,32) | 28.11±5.46 | 6 (18,75) | 31.00±4.28 | |
| ND | 299 (38,93) | / | 200 (37,87) | / | 89 (42,78) | / | 10 (31,25) | / | |
| Maternal RhD status | RhD+ | 410 (53,38) | 33.19±5.26 | 288 (54,54) | 34.4±4.81 | 102 (49,03) | 30.14±5.23 | 20 (62,5) | 31.30±5.00 |
| RhD− | 59 (7,68) | 32.91±5.21 | 40 (7,57) | 34.3±4.75 | 17 (8,17) | 29.64±5.08 | 2 (6,25) | 30.00±2.82 | |
| ND | 299 (38,93) | / | 200 (37,87) | / | 89 (42,78) | / | 10 (31,25) | / | |
| Gestational week | 6 | 36 (4,68) | 33.02±4.64 | 24 (4,54) | 34.66±3.71 | 11 (5,28) | 29.90±4.90 | 1 (3,12) | 28.00±0.00 |
| 7 | 80 (10,41) | 34.27±5.29 | 58 (10,98) | 34.75±4.98 | 16 (7,69) | 31.93±5.37 | 6 (18,75) | 35.83±7.02 | |
| 8 | 102 (13,28) | 32.89±5.43 | 71 (13,44) | 34.45±5.05 | 26 (12,5) | 29.53±4.53 | 5 (15,62) | 28.20±5.21 | |
| 9 | 65 (8,46) | 32.21±5.28 | 46 (8,71) | 32.91±4.83 | 17 (8,17) | 30.23±5.99 | 2 (6,25) | 33.00±8.48 | |
| 10 | 33 (4,29) | 33.72±5.51 | 24 (4,54) | 34.91±4.97 | 7 (3,36) | 31.14±6.54 | 2 (6,25) | 28.5±3.53 | |
| 11 | 39 (5,07) | 30.74±4.98 | 26 (4,92) | 31.65±4.31 | 11 (5,28) | 29.27±6.35 | 2 (6,25) | 27.00±1.41 | |
| ND | 413 (53,77) | / | 279 (52,84) | / | 120 (57,69) | / | 14 (43,75) | / | |
| Fetal sex | Male | 379 (49.34) | 33.03±5.26 | 264 (50.00) | 34.09±4.74 | 101 (48.55) | 30.21±5.46 | 14 (43.75) | 33.42±6.01 |
| Female | 389 (50.65) | 32.93±5.44 | 264 (50.00) | 34.19±5.06 | 107 (51.44) | 30.20±5.27 | 18 (56.25) | 30.72±5.28 | |
| Total | 768 (100.00) | / | 528 (100.00) | / | 208 (100.00) | / | 32 (100.00) | / | |
MA, mean age; SD, standard deviation; ND, no data
Methods
DNA extraction from the POC samples, as well as from maternal blood was performed using the standard phenol/chloroform method or the automated magnetic bead-based protocol using the MagCore Super instrument (RBC Bioscience). The study primarily used the quantitative fluorescent (QF)-PCR method with STR markers on chromosomes 13, 18, 21 and sex chromosomes. Three markers were located on chromosome 13, four on chromosomes 18 and 21 each and six on the sex chromosomes. Except aneuploidies on the given chromosomes, this method also allowed for the determination of triploid samples, and exclusion of maternal DNA contamination in the analysed samples. This method is described in detail by Noveski et al. [16]. All results for chromosomal aneuploidies obtained by the QF-PCR analyses were confirmed by the subsequent subtelomere MLPA analyses.
Chromosomal gains and losses were detected with the Multiplex Ligation Probe Amplification (MLPA) method, using the SALSA MLPA P036 Subtelomeres mix 1 and SALSA MLPA P070 Subtelomeres mix 2B (MRC-Holland). Each MLPA kit contains two probes for each chromosome. For metacentric chromosomes the two probes were located subtelomerically, while for the acrocentric chromosomes, one probe was located subcentromeric, while the other was subtelomeric. The detailed MLPA protocol as well as the chromosomal location of each probe contained in the kits is available on the MRC-Holland site. Capillary electrophoresis was performed on the AB3500 Genetic Analyser (Life Technologies), and the obtained results were analysed and interpreted using the Coffalyzer software (MRC-Holland). Mean values, standard deviations, percentages, odds ratios, and p-values were determined where appropriate, using the MedCalc software (MedCalc Software Ltd. https://www.medcalc.org/ Version 22.016; accessed November 17, 2023). A p-value below 0.05 was considered statistically significant.
RESULTS
The QF-PCR analysis performed on 900 POC samples showed maternal DNA contamination in 14.66%. These samples were excluded from further investigation and MLPA analyses were performed on a total of 768 POC samples.
Chromosomal abnormalities - overall incidence
Chromosomal abnormalities were present in 56.25% of uncontaminated POC samples. Chromosomal trisomies were detected in 66.20%, triploidy in 15.28%, and monosomies were present in 10.88% of the positive cases. The group of monosomies consisted of monosomy X which was predominant (91.49%) and monosomy 21 was present in 8.51%.
All chromosomes, apart from chromosomes 1 and 19, were affected by trisomy; the most common was trisomy 16, present in 26.57% of all trisomy samples, followed by trisomy 22 (15.73%), 21 (10.13%) and 15 (9.09%) (Figure 1).
Figure 1.
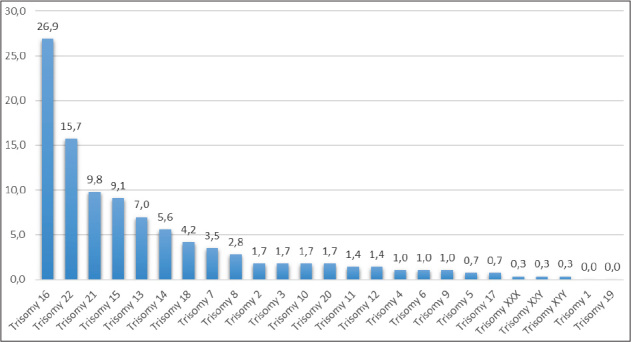
Distribution of chromosomal trisomies in EPLs as a percentage of all detected trisomies (n=286).
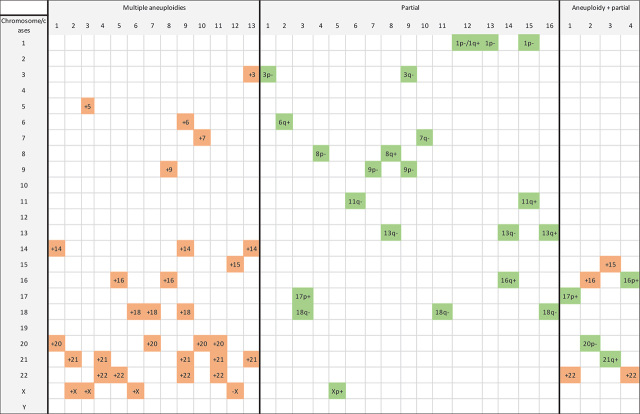
Partial chromosomal abnormalities (terminal deletions and duplications), multiple chromosomal aneuploidies, and chromosomal aneuploidies accompanied by partial chromosomal abnormality were found in 3.70%, 3.01%, and 0.93% of all positive samples (Table 2). A detailed description of the later three groups of chromosomal abnormalities is provided in Figure 2. Chromosomes 20, 21, 22 and X were most involved in multiple aneuploidies, chromosomes 1, 13 and 18 in partial aneuploidies, while chromosomes 15, 16 and 22 were found in combination with a partial aneuploidy. In cases of partial chromosomal abnormalities, parental karyotyping could not be performed at the time.
Table 2.
Frequency of chromosomal abnormalities in EPLs according to different clinical characteristics.
| Clinical characteristics | Study groups | Total | Chromosomal abnormalities | Trisomy n (%) | Triploidy n (%) | Monosomy n (%) | Multiple aneuploidies n (%) | Partial n (%) | Aneuploidy + partial n (%) | Total n (%) |
|---|---|---|---|---|---|---|---|---|---|---|
| n | n (%) | |||||||||
| Total | 768 | 432 (56.25) | 286 (66.20) | 66 (15.28) | 47 (10.88) | 13 (3.01) | 16 (3.70) | 4 (0.93) | 432 (100) | |
| Age group | ≤30 | 265 | 115 (43.40) | 56 (48.70) | 30 (26.09) | 19 (16.52) | 2 (1.74) | 7 (6.09) | 1 (0.87) | 115 (100) |
| 31–35 | 241 | 130 (53.94) | 81 (62.31) | 24 (18.46) | 16 (12.31) | 0 (0.00) | 7 (5.38) | 2 (1.54) | 130 (100) | |
| ≥36 | 262 | 187 (71.37) | 149 (79.68) | 12 (6.42) | 12 (6.42) | 11 (5.88) | 2 (1.07) | 1 (0.53) | 187 (100) | |
| Ethnic origin | Macedonian | 528 | 309 (58.52) | 216 (69.90) | 38 (12.30) | 29 (9.39) | 13 (4.21) | 10 (3.24) | 3 (0.97) | 309 (100) |
| Albanian | 208 | 106 (50.96) | 58 (54.72) | 27 (25.47) | 16 (15.09) | 0 (0.00) | 3 (2.83) | 2 (1.89) | 106 (100) | |
| Other | 32 | 17 (53.13) | 12 (70.59) | 1 (5.88) | 2 (11.76) | 0 (0.00) | 2 (11.76) | 0 (0.00) | 17 (100) | |
| History of PL | Sporadic | 268 | 164 (61.19) | 109 (66.46) | 22 (13.41) | 19 (11.59) | 6 (3.66) | 7 (4.27) | 1 (0.61) | 164 (100) |
| RPL (≥2) | 259 | 143 (55.21) | 94 (65.73) | 20 (13.99) | 17 (11.89) | 4 (2.80) | 5 (3.50) | 3 (2.10) | 143 (100) | |
| Previous live birth | Yes | 142 | 88 (61.97) | 60 (68.18) | 11 (12.50) | 14 (15.91) | 2 (2.27) | 1 (1.14) | 0 (0.00) | 88 (100) |
| No | 383 | 218 (56.92) | 143 (65.60) | 31 (14.22) | 21 (9.63) | 8 (3.67) | 11 (5.05) | 4 (1.83) | 218 (100) | |
| Maternal Blood Group | 0 | 183 | 103 (56.28) | 63 (61.17) | 14 (13.59) | 15 (14.56) | 5 (4.85) | 4 (3.88) | 2 (1.94) | 103 (100) |
| A | 183 | 115 (62.84) | 77 (66.96) | 16 (13.91) | 12 (10.43) | 4 (3.48) | 4 (3.48) | 2 (1.74) | 115 (100) | |
| B | 67 | 34 (50.75) | 26 (76.47) | 4 (11.76) | 3 (8.82) | 0 (0.00) | 1 (2.94) | 0 (0.00) | 34 (100) | |
| AB | 36 | 23 (63.89) | 16 (69.57) | 3 (13.04) | 3 (13.04) | 0 (0.00) | 1 (4.35) | 0 (0.00) | 23 (100) | |
| Maternal RhD status | RhD + | 410 | 244 (59.51) | 163 (66.80) | 33 (13.52) | 31 (12.70) | 6 (2.46) | 7 (2.87) | 4 (1.64) | 244 (100) |
| RhD− | 59 | 31 (52.54) | 20 (64.52) | 4 (12.90) | 3 (9.68) | 1 (3.23) | 3 (9.68) | 0 (0.00) | 31 (100) | |
| Gestational week | 6 | 36 | 19 (52.78) | 14 (73.68) | 2 (10.53) | 1 (5.26) | 0 (0.00) | 1 (5.26) | 0 (0.00) | 19 (100) |
| 7 | 80 | 45 (56.25) | 38 (84.44) | 5 (11.11) | 0 (0.00) | 0 (0.00) | 2 (4.44) | 0 (0.00) | 45 (100) | |
| 8 | 102 | 56 (54.90) | 38 (67.86) | 8 (14.29) | 5 (8.93) | 1 (1.79) | 3 (5.36) | 1 (1.79) | 56 (100) | |
| 9 | 65 | 41 (63.08) | 24 (58.54) | 9 (21.95) | 4 (9.76) | 3 (7.32) | 1 (2.44) | 0 (0.00) | 41 (100) | |
| 10 | 33 | 22 (66.67) | 10 (45.45) | 3 (13.64) | 8 (36.36) | 0 (0.00) | 0 (0.00) | 1 (4.55) | 22 (100) | |
| 11 | 39 | 21 (53.85) | 12 (57.14) | 5 (23.81) | 4 (19.05) | 0 (0.00) | 0 (0.00) | 0 (0.00) | 21 (100) | |
| Fetal sex | Male | 379 | 203 (53.56) | 141 (69.45) | 45 (22.16) | 3 (1.47) | 3 (1.47) | 8 (3.94) | 3 (1.47) | 203 (100) |
| Female | 389 | 229 (58.86) | 146 (63.75) | 22 (9.60) | 45 (19.65) | 8 (3.49) | 7 (3.05) | 1 (0.43) | 229 (100) |
Figure 2.
Multiple aneuploidies, partial chromosomal abnormalities, aneuploidies accompanied by partial abnormality detected among the studied EPLs.
Chromosomal abnormalities and maternal age
As shown in Table 2 the overall frequency of chromosomally abnormal POCs increased with maternal age (43.40% in the women ≤30 years of age, 53.94% in women 31–35 years and 71.37% in woman ≥36 years). The odds for chromosomally abnormal EPLs were 1.52 higher in the group of women 31–35 years of age (95% CI: 1.07–2.16; p=0.018) and 3.25 higher in women ≥36 years (95% CI: 2.26–4.66; p<0.0001), when compared to the group of women ≤30 years of age (Table 3).
Table 3.
Comparisons of the incidence of chromosomal abnormalities between the three groups according to the maternal age (Odds ratios and p-values).
| ≤30 (n) | 31–35 (n) | OR (95% CI) | p | >36 (n) | OR (95% CI) | p | |
|---|---|---|---|---|---|---|---|
| Abnormal | 115 | 130 | 1.52 (1.07–2.16) | 0.018 | 187 | 3,25 (2.26–4.66) | < 0.0001 |
| Trisomy | 56 | 81 | 1.88 (1.26–2.81) | 0.0017 | 149 | 4.92 (3.35–7.21) | < 0.0001 |
| Triploidy | 30 | 24 | 0.64 (0.24–1,17) | 0.152 | 12 | 0.19 (0.09–0.39) | < 0.0001 |
| Monosomy | 19 | 16 | 0.89 (0.43–1.85) | 0.771 | 12 | 0.34 (0.16–0.74) | 0.0066 |
| Trisomy 16 | 22 | 25 | 0.86 (0.43–1.74) | 0.678 | 29 | 0.31 (0.15–0.62) | 0.0009 |
| Trisomy 22 | 0 | 10 | 16.59 (0.95–289.31) | 0.054 | 35 | 35.03 (2.11–581.6) | 0.013 |
| Trisomy 21 | 7 | 9 | 0.87 (0.30–2.50) | 0.803 | 13 | 0.66 (0.25–1.77) | 0.419 |
| Trisomy 15 | 0 | 4 | 6.56 (0.34–124.34) | 0.210 | 22 | 19.94 (1.18–334.53) | 0.037 |
| Trisomy 13 | 7 | 7 | 0.66 (0.21–2.00) | 0.465 | 6 | 0.29 (0.09–0.91) | 0.034 |
| Trisomy 14 | 7 | 4 | 0.36 (0.10–1.30) | 0.121 | 5 | 0.24 (0.07–0.80) | 0.020 |
| Other abnormalities | 10 | 10 | 1.10 (0.45–2.70) | 0.828 | 14 | 1.43 (0.62–3.30) | 0.389 |
The spectrum of the chromosomal abnormalities differed in the three groups of women according to their age. In the group ≥36 years, the trisomies were predominant (79.68%), while triploidy and monosomies were present in small portions (6.42% each). In the group of women ≤30 years of age, a more even distribution of the chromosomal abnormalities was observed, with trisomies present in 48.70%, triploidy in 26.09% and monosomies in 16.52%. The odds for EPLs with trisomy were 1.88 times higher in the group of women 31–35 years of age (95% CI: 1.26–2.81; p=0.0017) and 4.92 higher in women ≥36 years (95% CI: 3.35–7.21; p<0.0001), when compared to the group of women ≤30 years of age (Table 3).
Multiple chromosomal trisomies were more common among the group ≥36 years (5.88%), while partial chromosomal aneuploidies were more frequent in women ≤35 (Table 2).
Considering the individual trisomies, trisomy 16 had the highest incidence in the group ≤30 years (39.29%), and lowest in the group ≥36 years of age (16.78%). Furthermore, trisomies 13 and 14 were also more common among the group ≤30 years (12.50% and 12.50% each), than in the other two groups. By contrast, trisomy 22 was more common in the group ≥36 years (23.49%) than in the group 31–35 years (12.35%). It was entirely absent in the youngest group of women (≤30 years of age). Similarly, the trisomy 15 was most common in the group of women ≥36 (14.77%) than in other groups (0.00% and 4.94% in groups of women ≤30 and 31–35 years, respectively) (Table 4). The odds of EPLs with trisomy 16 were 0.31 lower in the group of women ≥36 years (95% CI: 0.15–0.62; p=0.0009) while for trisomy 22, the odds were 16.59 higher in the group of women 31–35 years of age (95% CI: 0.95–289.31; p=0.054) and 35.03 higher in women ≥36 years (95% CI: 2.11–581.6; p=0.013) when compared to the group of women ≤30 years of age. The odds for trisomy 15 were higher (OR=19.94, 95% CI: 1.18–334.53; p=0.037), while for trisomies 13 and 14 were lower in women ≥36 years (OR=0.29, 95% CI: 0.09–0.91; p=0.034 and OR=0.24, 95% CI: 0.07–0.80; p=0.02, respectively) when compared to the group of women ≤30 years of age (Table 3).
Table 4.
The most common chromosomal trisomies in the different studied groups shown in table with number of cases as well as percentage.
| Clinical characteristics | Study groups | Trisomy 16 | Trisomy 22 | Trisomy 21 | Trisomy 15 | Trisomy 13 | Trisomy 14 | Other | Total | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| n | % | n | % | n | % | n | % | n | % | n | % | n | % | n | % | ||
| Age group | ≤30 | 22 | 39.29 | 0 | 0.00 | 7 | 12.50 | 0 | 0.00 | 7 | 12.50 | 7 | 12.50 | 13 | 23.21 | 56 | 100.00 |
| 31–35 | 29 | 35.80 | 10 | 12.35 | 9 | 11.11 | 4 | 4.94 | 7 | 8.64 | 4 | 4.94 | 18 | 22.22 | 81 | 100.00 | |
| ≥36 | 25 | 16.78 | 35 | 23.49 | 13 | 8.72 | 22 | 14.77 | 6 | 4.03 | 5 | 3.36 | 43 | 28.86 | 149 | 100.00 | |
| Ethnic origin | MK | 53 | 24.54 | 40 | 18.52 | 23 | 10.65 | 18 | 8.33 | 11 | 5.09 | 12 | 5.56 | 59 | 27.31 | 216 | 100.00 |
| AL | 19 | 32.76 | 4 | 6.90 | 5 | 8.62 | 7 | 12.07 | 8 | 13.79 | 3 | 5.17 | 12 | 20.69 | 58 | 100.00 | |
| History of miscarriage | Sporadic | 25 | 22.94 | 21 | 19.27 | 9 | 8.26 | 12 | 11.01 | 7 | 6.42 | 5 | 4.59 | 30 | 27.52 | 109 | 100.00 |
| RPL | 24 | 25.53 | 17 | 18.09 | 5 | 5.32 | 11 | 11.70 | 7 | 7.45 | 6 | 6.38 | 24 | 25.53 | 94 | 100.00 | |
| Previous live birth | Yes | 9 | 15.00 | 11 | 18.33 | 2 | 3.33 | 11 | 18.33 | 3 | 5.00 | 3 | 5.00 | 21 | 35.00 | 60 | 100.00 |
| No | 40 | 27.97 | 27 | 18.88 | 12 | 8.39 | 12 | 8.39 | 10 | 6.99 | 8 | 5.59 | 34 | 23.78 | 143 | 100.00 | |
| Maternal Blood Group | 0 | 20 | 31.75 | 10 | 15.87 | 6 | 9.52 | 11 | 17.46 | 4 | 6.35 | 3 | 4.76 | 9 | 14.29 | 63 | 100.00 |
| A | 19 | 24.68 | 14 | 18.18 | 3 | 3.90 | 9 | 11.69 | 4 | 5.19 | 3 | 3.90 | 25 | 32.47 | 77 | 100.00 | |
| B | 6 | 23.08 | 5 | 19.23 | 0 | 0.00 | 4 | 15.38 | 2 | 7.69 | 1 | 3.85 | 8 | 30.77 | 26 | 100.00 | |
| AB | 4 | 25.00 | 7 | 43.75 | 0 | 0.00 | 1 | 6.25 | 0 | 0.00 | 1 | 6.25 | 3 | 18.75 | 16 | 100.00 | |
| Maternal RhD status | RhD+ | 43 | 26.38 | 29 | 17.79 | 7 | 4.29 | 22 | 13.50 | 8 | 4.91 | 8 | 4.91 | 46 | 28.22 | 163 | 100.00 |
| RhD− | 6 | 30.00 | 3 | 15.00 | 2 | 10.00 | 3 | 15.00 | 2 | 10.00 | 0 | 0.00 | 4 | 20.00 | 20 | 100.00 | |
| Gestational week | 6 | 5 | 35.71 | 2 | 14.29 | 1 | 7.14 | 0 | 0.00 | 0 | 0.00 | 1 | 7.14 | 5 | 35.71 | 14 | 100.00 |
| 7 | 13 | 34.21 | 7 | 18.42 | 1 | 2.63 | 0 | 0.00 | 3 | 7.89 | 4 | 10.53 | 10 | 26.32 | 38 | 100.00 | |
| 8 | 8 | 21.05 | 7 | 18.42 | 1 | 2.63 | 4 | 10.53 | 2 | 5.26 | 3 | 7.89 | 13 | 34.21 | 38 | 100.00 | |
| 9 | 3 | 12.50 | 4 | 16.67 | 4 | 16.67 | 5 | 20.83 | 3 | 12.50 | 1 | 4.17 | 4 | 16.67 | 24 | 100.00 | |
| 10 | 3 | 30.00 | 1 | 10.00 | 0 | 0.00 | 3 | 30.00 | 0 | 0.00 | 0 | 0.00 | 3 | 30.00 | 10 | 100.00 | |
| 11 | 3 | 25.00 | 2 | 16.67 | 3 | 25.00 | 1 | 8.33 | 1 | 8.33 | 0 | 0.00 | 2 | 16.67 | 12 | 100.00 | |
| Foetal sex | Male | 40 | 28.36 | 19 | 13.47 | 13 | 9.21 | 10 | 7.09 | 10 | 7.09 | 10 | 7.09 | 19 | 13.47 | 141 | 100.00 |
| Female | 37 | 25.34 | 26 | 17.80 | 15 | 10.27 | 16 | 10.95 | 9 | 6.16 | 6 | 4.10 | 37 | 25.34 | 146 | 100.00 | |
Although not statistically significant, a higher incidence of chromosomally abnormal POCs were detected in the Macedonian ethnic group (58.52%) in comparison to the Albanian group (50.96%; p=0.06) (Table 2). Triploidy and chromosomal monosomies were significantly more common among the Albanian ethnic group (25.47% and 15.09% respectively; p=0.0001) than in the Macedonian group (12.30% and 9.39% respectively). In contrast, chromosomal trisomies were more prevalent in POCs of Macedonian ethnic origin (69.90%) than in the Albanian group (54.72), (p=0.0044).
Regarding the individual trisomies, the most evident difference was observed in trisomy 16, being more common among Albanians (32.76%, p=0.207) than among Macedonians (24.54%). Trisomy 22 was found to be more frequent in the Macedonian group compared to the Albanian (18.52% vs. 6.90%, respectively, p=0.035) (Table 4). However, these differences can be explained by the age distribution of samples with different ethnicity. Namely, most Macedonians (41.47%) belonged to the ≥36 years group and only 24.81% were in the group ≤30, while the majority of Albanians (58.17%) belonged to the group ≤30, and only 16.82% were in the group ≥36. This is in accordance with the tendency of earlier childbearing among Albanian women.
Chromosomal abnormalities and history of EPLs
The frequency of chromosomal abnormalities in POCs from women with sporadic EPLs was higher than in those with RPLs (61.19% vs. 55.21%, respectively, p=0.164), this difference, however, was not statistically significant (Table 2). No evident difference was observed between these two groups regarding the presence of different classes of chromosomal abnormalities and chromosomal trisomies (Tables 2 and 4).
Chromosomal abnormalities and previous live birth
In the group of women with a previous live birth, a slightly higher, but not significantly different, incidence of chromosomally abnormal POCs was found compared to the group of women without a live birth (61.97% vs. 56.92%, respectively, p=0.297) (Table 2). However, this is most probably due to the higher mean age of the women with a live birth (34.66 years) in comparison to that of women with no live birth (32.57 years). Triploidy and trisomies did not differ much among these groups, while monosomies were more common in the group with (15.91%, p=0.119) compared to the group without live births (9.63%), but without statistical significance.
Trisomy 16 and 21 were more prevalent in the group without live births (27.97% and 8.39%, respectively) than in the group with live births (15.00%, p=0.049 and 3.33%, p=0.195; respectively), while trisomy 15 was more common in the group with live births (18.33%, p=0,041) vs those without (8.39%) (Table 4). These findings were statistically significant for trisomies 16 and 15, but not for trisomy 21.
Chromosomal abnormalities and maternal blood groups/RhD status
The highest incidence of chromosomally abnormal POCs was detected in women with the AB blood group (63.89%), followed by A (62.84%), O (56.28%) and B (50.75%) (Table 2). The O blood group had the highest incidence of monosomies (14.56%) in contrast to the B with the lowest rate of monosomies in our study (8.82%) (p=0.312). The opposite was observed concerning the trisomies, where B had the highest, and O the lowest incidence (76.47% and 61.17%, respectively) (p=0.176).
Higher presence of trisomy 22 was observed among women of the AB blood group (43.75%), in comparison to women of other (combined B, A and O) blood groups (17.46%) (p=0.012). Furthermore, trisomy 21 was more frequently observed among women in the O blood group (9.52%; 6/63), compared to other groups (2.52%) (p=0.034) (Table 4).
Women with positive RhD positive (RhD+) had slightly higher incidence of abnormal POCs (59.51%) than the RhD negative (RhD−) women (52.54) (p=0.310).
Chromosomal abnormalities according to the gestational age
Overall, increased incidence of chromosomal abnormalities was present in advanced gestational age. The POCs eliminated in gestational week (gw) 10 showed the highest incidence of chromosomal abnormalities (66.67%), followed by gw 9 (63.08%), while the lowest rate was observed in the POCs eliminated in the gw 6 (52.78%).
Higher incidences of triploidy and monosomies were observed in POCs eliminated between gw 9 and gw 11 (p=0.135 and p=0.0015, respectively), while trisomies were more common among studied POCs from gw 6 to 8 (p=0.0026) (Table 2).
Chromosomal abnormalities and POCs sex
Among the 768 POCs, 379 were male sex (49.34%) and 389 were of female sex (50.65%). A slightly lower number of chromosomally abnormal male POCs was observed (53.56%) in comparison to the female POCs (58.86%). Triploidies were more common among male POCs in contrast to the females (25.56% vs. 13.75%, respectively; p=0.026). Chromosomal monosomies were more prevalent in female than in male POCs (28.12% vs. 1.7%; p<0.0001), but this was expected since monosomy X was predominant (95.55%) in this group, and there were only 2 cases with monosomy 21 (4.44%). Regarding the chromosomal trisomies, no significant difference was observed between males and females (Tables 2 and 4). In the group of RPL, a higher percentage of male POCs were euploid in comparison to the female POCs (50% vs. 39.85%), but the difference was not significant (p=0.1).
DISCUSSION
Foetal chromosomal abnormalities have been recognized as a major cause of EPLs [4]. Several methods have been used for the detection of chromosomal abnormalities with different detection rates [17,18,19,20,21]. Using a combination of QF-PCR and MLPA analyses, the chromosomal abnormality rate in our study was 56.25%, which agrees with the majority of previously published data. Chromosomal trisomies were the most frequent abnormalities (66.20%), followed by triploidy (15.28%) and monosomies (10.88%). Multiple chromosomal aneuploidies and partial chromosomal abnormalities were also identified. Thus, our approach could detect most of the chromosomal abnormalities known to cause EPLs.
In comparison to the conventional cytogenetic methods, our approach does not require viable tissue material, lowering the rate of unsuccessful analyses and has a potential to detect maternal cell contamination. Indeed, in our study 14.67% of the POC samples showed maternal contamination and were excluded from further analysis. Furthermore, the use of QF-PCR enables not only detection of maternal cell contamination, but also detection of foetal triploidy, which could not be possible by using only MLPA analysis. Although QF-PCR/MLPA approach could not accurately detect other polyploidies, such as tetraploidy, these abnormalities rarely occur and represent only less than 3% of all chromosomal abnormalities in EPLs [19]. Some disadvantages of the QF-PCR and MLPA methods used in present study include the inability to detect balanced structural chromosomal rearrangements, as well as inter-stitial deletions and duplications, ring chromosomes, and inversions. Some of these variations could be detected with other molecular approaches such as aCGH and NGS-based methodologies, but because of their higher price, they are not widely used in the investigation of EPLs samples [5, 6].
In our study, a significantly higher frequency of chromosomally abnormal POCs was observed in the group of women ≥36 years (71.37%) compared to the groups ≤30 (43.4%; OR=3.25; p<0.0001) and 31–35 years (53.9%; OR=1.52, p=0.018). Thus, our study confirms previous observations of increased EPL aneuploidy rate in women with advanced age [13, 22]. Extensive research has been performed with the aim to explain the reason behind the increased rate of aneuploidy with advanced maternal age. Some of the leading causes identified include recombination failure, cohesion and spindle deregulation, abnormalities in post-translational modification of histones and tubulin, and mitochondrial dysfunction [23, 24].
The high trisomy rate in our study (66%), with trisomy 16 being the most frequent, followed by trisomies 22, 21, 13 and 15 is in accordance with the data from previous published studies [21, 25,26,27]. The trisomy rate increased with maternal age, being the highest (79.68%) in women ≥36 years of age and lowest in women ≤30 years (48.70%). On the other hand, our study showed higher rates of monosomies and triploidy in younger women, which has been also shown by other authors [26, 28].
The distribution of individual trisomies differed among the three age groups. The frequency of trisomy 16 (calculated on all studied samples) was similar in the three groups, while trisomies 22, 21, 15 as well as other rare trisomies showed the highest frequencies in the group ≥36 years (Table 5). Many previous studies have also shown that trisomies 21 and 22 are more common in women with advanced age and have shown that they occur mostly because of maternal non-disjunction in meiosis I. Both chromosomes 21 and 22 are acrocentric and, on average, each is held by a single chiasma, which can be commonly lost during meiosis [29]. Thus, our study confirms the findings that trisomies 22, 21 and 15 occur due to age-related factors, but also imply that other, non-age-related factors might be involved in the occurrence of trisomy 16.
Table 5.
Distribution of the chromosomal trisomies among all studied EPLs
| Trisomy | ≤30 n (%) | 31–35 n (%) | ≥36 n (%) |
|---|---|---|---|
| 16 | 22 (8.30) | 29 (12.03) | 25 (9.54) |
| 22 | 0 (0.00) | 10 (4.15) | 35 (13.36) |
| 21 | 7 (7.64) | 9 (3.73) | 13 (4.96) |
| 15 | 0 (0.00) | 4 (1.66) | 22 (8.40) |
| 13 | 7 (2.64) | 7 (2.90) | 6 (2.29) |
| 14 | 7 (2.64) | 4 (1.66) | 5 (1.91) |
| Other | 13 (4.91) | 18 (7.47) | 43 (16.41) |
| Total trisomy | 56 (48.71) | 81 (62.3) | 149 (79.68) |
| Total POC | 265 | 241 | 262 |
Our results favour the findings of lower chromosomal aneuploidy rate in recurrent compared to the sporadic EPLs [7], however with a similar distribution of the chromosomal abnormalities and trisomies 16 and 22 being the most common in both groups [30]. The differences observed among the women with different ethnic origins as well as those with and without live births could be related to the maternal age. Although, we have observed some differences in the chromosomal aneuploidy rates among women with different ABO blood groups and RhD status (with higher rates in groups A and AB, as well as in RhD positive women) further larger studies are required to confirm our findings.
Our results show that triploidy and monosomies occur more frequently in POCs eliminated in gw 9–11, while trisomies are more common in gw 6–8 POCs, findings observed also by other studies [31]. Thus, our study suggests that chromosomal trisomies have a more negative effect on POCs, causing earlier elimination during the pregnancy in contrast to the triploidies and monosomies.
We have observed a slight predominance of chromosomally abnormal POC in females than in males (58.86% vs. 53.56%), although without statistical significance. The female predominance was also observed by other authors indicating relative weakness in female embryo formation and development, supported by an animal model where, male embryos development was favoured in comparison to female development [32, 33].
CONCLUSION
Our study furthers the understanding regarding chromosomal abnormalities as a cause of EPLs and confirms the higher incidence of foetal chromosomal abnormalities in EPLs in women of older reproductive age. Furthermore, it shows that by using QF-PCR and MLPA methodologies, a high detection rate of chromosomal abnormalities in EPLs can be found.
ACKNOWLEDGMENT
The study was supported by grant No. 08-707/2023 by the Macedonian Academy of Sciences and Arts to DPK.
Footnotes
CONFLICT OF INTEREST
None declared.
REFERENCES
- 1.Macklon NS, Geraedts JPM, Fauser BCJM. Conception to ongoing pregnancy: the “black box” of early pregnancy loss. Hum Reprod Update. 2002;8:333–343. doi: 10.1093/humupd/8.4.333. [DOI] [PubMed] [Google Scholar]
- 2.Pillarisetty LS, Mahdy H. Recurrent Pregnancy Loss. [Updated 2022 Sep 6]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2023 Jan-. Available from: https://www.ncbi.nlm.nih.gov/books/NBK554460/ ) [PubMed]
- 3.Ford HB, Schust DJ. Recurrent pregnancy loss: etiology, diagnosis, and therapy. Reviews in obstetrics and gynecology. 2009;2(2):76. [PMC free article] [PubMed] [Google Scholar]
- 4.Goddijn M, Leschot NJ. Genetic aspects of miscarriage. Best Practice & Research Clinical Obstetrics & Gynaecology. 2000 NaN 1;14(5):855–65. doi: 10.1053/beog.2000.0124. [DOI] [PubMed] [Google Scholar]
- 5.Gou L, Liu T, Wang Y, Wu Q, Hu S, Dong B, Wang C, Zhang Y, Shan X, Wang X, Suo F. Clinical utilization of chromosomal microarray analysis for the genetic analysis in subgroups of pregnancy loss. The Journal of Maternal-Fetal & Neonatal Medicine. 2022 NaN 17;35(22):4404–11. doi: 10.1080/14767058.2020.1849126. [DOI] [PubMed] [Google Scholar]
- 6.Tamura Y, Santo M, Araki Y, Matsubayashi H, Takaya Y, Kitaya K, Doshida M, Yamaguchi K, Mizuta S, Takahashi C, Kim N. Chromosomal copy number analysis of products of conception by conventional karyotyping and next-generation sequencing. Reproductive Medicine and Biology. 2021 NaN;20(1):71–5. doi: 10.1002/rmb2.12351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lei D, Zhang XY, Zheng PS. Recurrent pregnancy loss: fewer chromosomal abnormalities in products of conception? a meta-analysis. Journal of Assisted Reproduction and Genetics. 2022 NaN;39(3):559–72. doi: 10.1007/s10815-022-02414-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Smits MA, van Maarle M, Hamer G, Mastenbroek S, Goddijn M, van Wely M. Cytogenetic testing of pregnancy loss tissue: a meta-analysis. Reproductive BioMedicine Online. 2020 NaN 1;40(6):867–79. doi: 10.1016/j.rbmo.2020.02.001. [DOI] [PubMed] [Google Scholar]
- 9.Andersen AM, Wohlfahrt J, Christens P, Olsen J, Melbye M. Maternal age and fetal loss: population based register linkage study. Bmj. 2000 NaN 24;320(7251):1708–12. doi: 10.1136/bmj.320.7251.1708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gu C, Li K, Li R, Li L, Li X, Dai X, He Y. Chromosomal aneuploidy associated with clinical characteristics of pregnancy loss. Frontiers in Genetics. 2021 NaN 15;12:667697. doi: 10.3389/fgene.2021.667697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zhang X, Fan J, Chen Y, Wang J, Song Z, Zhao J, Li Z, Wu X, Hu Y. Cytogenetic analysis of the products of conception after spontaneous abortion in the first trimester. Cytogenetic and Genome Research. 2021 NaN 21;161(3–4):120–31. doi: 10.1159/000514088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Clarke CA. Practical effects of blood group incompatibility between mother and fetus. British Medical Journal. 1972 NaN 4;2(5805):90. doi: 10.1136/bmj.2.5805.90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Donaghue C, Mann K, Docherty Z, Mazzaschi R, Fear C, Ogilvie C. Combined QF-PCR and MLPA molecular analysis of miscarriage products: an efficient and robust alternative to karyotype analysis. Prenatal Diagnosis: Published in Affiliation With the International Society for Prenatal Diagnosis. 2010 NaN;30(2):133–7. doi: 10.1002/pd.2424. [DOI] [PubMed] [Google Scholar]
- 14.Saxena D, Agarwal M, Gupta D, Agrawal S, Das V, Phadke SR. Utility and limitations of multiplex ligation-dependent probe amplification technique in the detection of cytogenetic abnormalities in products of conception. Journal of postgraduate medicine. 2016 NaN;62(4):239. doi: 10.4103/0022-3859.192664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kubelka-Sabit K, Bozinovski G, Jasar D, Filipovski V, Lazarevski S, Ivanovski M, Plaseska-Karanfilska D. Detection of placental chromosomal aberrations in early spontaneous abortions in correlation with the histologic findings. Macedonian Medical Review. 2017;71(1):64–71. [Google Scholar]
- 16.Noveski P, Terzic M, Vujovic M, Kuzmanovska M, Sukarova Stefanovska E, Plaseska-Karanfilska D. Multilevel regression modeling for aneuploidy classification and physical separation of maternal cell contamination facilitates the QF-PCR based analysis of common fetal aneuploidies. Plos one. 2019 NaN 20;14(8):e0221227. doi: 10.1371/journal.pone.0221227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Pylyp LY, Spynenko LO, Verhoglyad NV, Mishenko AO, Mykytenko DO, Zukin VD. Chromosomal abnormalities in products of conception of first-trimester miscarriages detected by conventional cytogenetic analysis: a review of 1000 cases. Journal of assisted reproduction and genetics. 2018 NaN;35:265–71. doi: 10.1007/s10815-017-1069-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shearer BM, Thorland EC, Carlson AW, Jalal SM, Ketterling RP. Reflex fluorescent in situ hybridization testing for unsuccessful product of conception cultures: a retrospective analysis of 5555 samples attempted by conventional cytogenetics and fluorescent in situ hybridization. Genetics in Medicine. 2011 NaN 1;13(6):545–52. doi: 10.1097/GIM.0b013e31820c685b. [DOI] [PubMed] [Google Scholar]
- 19.Lin SB, Xie YJ, Chen Z, Zhou Y, Wu JZ, Zhang ZQ, Shi SS, Chen BJ, Fang Q. Improved assay performance of single nucleotide polymorphism array over conventional karyotyping in analyzing products of conception. Journal of the Chinese Medical Association. 2015 NaN 1;78(7):408–13. doi: 10.1016/j.jcma.2015.03.010. [DOI] [PubMed] [Google Scholar]
- 20.Kowalczyk K, Smyk M, Bartnik-Głaska M, Plaskota I, Wiśniowiecka-Kowalnik B, Bernaciak J, Chojnacka M, Paczkowska M, Niemiec M, Dutkiewicz D, Kozar A. Application of array comparative genomic hybridization (aCGH) for identification of chromosomal aberrations in the recurrent pregnancy loss. Journal of Assisted Reproduction and Genetics. 2022 NaN;39(2):357–67. doi: 10.1007/s10815-022-02400-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gug C, Rațiu A, Navolan D, Drăgan I, Groza IM, Păpurică M, Vaida MA, Mozoș I, Jurcă MC. Incidence and spectrum of chromosome abnormalities in miscarriage samples: a retrospective study of 330 cases. Cytogenetic and Genome Research. 2019 NaN 24;158(4):171–83. doi: 10.1159/000502304. [DOI] [PubMed] [Google Scholar]
- 22.Cimadomo D, Fabozzi G, Vaiarelli A, Ubaldi N, Ubaldi FM, Rienzi L. Impact of maternal age on oocyte and embryo competence. Frontiers in endocrinology. 2018 NaN 29;9:327. doi: 10.3389/fendo.2018.00327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Mikwar M, MacFarlane AJ, Marchetti F. Mechanisms of oocyte aneuploidy associated with advanced maternal age. Mutation Research/Reviews in Mutation Research. 2020 NaN 1;785:108320. doi: 10.1016/j.mrrev.2020.108320. [DOI] [PubMed] [Google Scholar]
- 24.Vlachadis N, Papadopoulou T, Vrachnis D, Manolakos E, Loukas N, Christopoulos P, Pappa K, Vrachnis N. Incidence and Types of Chromosomal Abnormalities in First Trimester Spontaneous Miscarriages: a Greek Single-Center Prospective Study. Maedica. 2023 NaN;18(1):35. doi: 10.26574/maedica.2023.18.1.35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jia CW, Wang L, Lan YL, Song R, Zhou LY, Yu L, Yang Y, Liang Y, Li Y, Ma YM, Wang SY. Aneuploidy in early miscarriage and its related factors. Chinese medical journal. 2015 NaN 20;128(20):2772–6. doi: 10.4103/0366-6999.167352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gomez R, Hafezi N, Amrani M, Schweiger S, Dewenter MK, Thomas P, Lieb C, Hasenburg A, Skala C. Genetic findings in miscarriages and their relation to the number of previous miscarriages. Archives of gynecology and obstetrics. 2021 NaN;303:1425–32. doi: 10.1007/s00404-020-05859-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Popescu-Hobeanu G, Riza AL, Streață I, Tudorache Ș, Comănescu A, Tănase F, Drăgușin RC, Pascu C, Dijmărescu AL, Cara ML, Dorobanțu Ș. Cytogenetic Analysis of Sporadic First-Trimester Miscarriage Specimens Using Karyotyping and QF-PCR: A Retrospective Romanian Cohort Study. Genes. 2022 NaN 29;13(12):2246. doi: 10.3390/genes13122246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hall HE, Surti U, Hoffner L, Shirley S, Feingold E, Hassold T. The origin of trisomy 22: evidence for acrocentric chromosome-specific patterns of nondisjunction. American Journal of Medical Genetics Part A. 2007 NaN 1;143(19):2249–55. doi: 10.1002/ajmg.a.31918. [DOI] [PubMed] [Google Scholar]
- 29.Lovrečić L, Pereza N, Jaklič H, Ostojić S, Peterlin B. Combination of QF-PCR and aCGH is an efficient diagnostic strategy for the detection of chromosome aberrations in recurrent miscarriage. Molecular Genetics & Genomic Medicine. 2019 NaN;7(12):e980. doi: 10.1002/mgg3.980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kajii T, Ferrier A, Niikawa N, Takahara H, Ohama K, Avirachan S. Anatomic and chromosomal anomalies in 639 spontaneous abortuses. Human Genetics. 1980 NaN;55(1):87–98. doi: 10.1007/BF00329132. [DOI] [PubMed] [Google Scholar]
- 31.Soler A, Morales C, Mademont-Soler I, Margarit E, Borrell A, Borobio V, Muñoz M, Sánchez A. Overview of chromosome abnormalities in first trimester miscarriages: a series of 1,011 consecutive chorionic villi sample karyotypes. Cytogenetic and genome research. 2017 NaN 1;152(2):81–9. doi: 10.1159/000477707. [DOI] [PubMed] [Google Scholar]
- 32.Cheng HH, Ou CY, Tsai CC, Chang SD, Hsiao PY, Lan KC, Hsu TY. Chromosome distribution of early miscarriages with present or absent embryos: female predominance. Journal of assisted reproduction and genetics. 2014 NaN;31:1059–64. doi: 10.1007/s10815-014-0261-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Nedambale TL, Dinnyés A, Yang X, Tian XC. Bovine blastocyst development in vitro: timing, sex, and viability following vitrification. Biology of reproduction. 2004 NaN 1;71(5):1671–6. doi: 10.1095/biolreprod.104.027987. [DOI] [PubMed] [Google Scholar]