Summary
Epigenetic modifications have emerged as key regulators of metabolism-related complex diseases including the alcoholic fatty liver disease (AFLD) prevalent chronic liver disorder with significant economic implications. Building upon previous research that emphasizes ten-eleven translocation (TET) proteins' involvement in adipocyte insulin sensitization and fatty acid oxidation, we explored the role of TET2 protein in AFLD pathogenesis which catalyzes 5-methylcytosine into 5-hydroxymethylcytosine in DNA/RNA. Our findings revealed that TET2 deficiency exacerbates AFLD progression. And TET2 influenced the expression and activity of sterol regulatory element binding protein 1 (SREBP1), a key regulator of hepatic lipid synthesis, by modulating Srebp1 mRNA retention. Employing RIP-qPCR and bisulfite sequencing techniques, we provided evidence of TET2-mediated epigenetic modifications on Srebp1 mRNA, thereby affecting lipid metabolism. Through elucidating the role of methylation in RNA nuclear retention via paraspeckles, our study enhances understanding of AFLD pathogenesis from an epigenetic perspective, paving the way for identifying potential therapeutic targets.
Subject areas: Classification Description: Molecular biology, Molecular mechanism of gene regulation, Epigenetics
Graphical abstract

Highlights
-
•
TET2 deficiency promotes alcoholic fatty liver formation
-
•
TET2 interacts with NONO in paraspeckles
-
•
TET2 regulates the nuclear retention of Srebp1 mRNA in paraspeckles
-
•
TET2 demethylates Srebp1 mRNA 3′UTR
Classification Description: Molecular biology; Molecular mechanism of gene regulation; Epigenetics
Introduction
Currently liver disease is the eleventh-leading cause of death, among which alcoholic fatty liver disease (AFLD) is one of the most common chronic liver diseases worldwide.1,2 Excessive alcohol consumption increases fat production and lipid mobilization, while reducing liver lipid catabolism, resulting in lipid accumulation in hepatocytes.3 The early stages of AFLD, which manifests as simple hepatic steatosis, are reversible; AFLD may further develop into progressive alcoholic steatohepatitis (ASH), cirrhosis, and even hepatocellular carcinoma (HCC) in some cases.4,5 Furthermore, obesity and metabolic disturbance are common in patients with AFLD, suggesting that excessive alcohol use and metabolic phenotypes may interact in AFLD progression.6 Excessive drinking promotes the occurrence of obesity,7 meanwhile obesity also increases the morbidity and mortality of patients with alcoholic hepatitis.8 Like many other complex diseases, fatty liver disease is affected by heredity and environment.9 Recent studies indicate the presence of tissue-specific epigenetic modifications in both nuclear and mitochondrial genomes in fatty liver disease.10,11 Despite significant advancements in understanding epigenetic modifications, the exploration of their relationship and application in the context of fatty liver is still at an early stage. Considering the substantial burden and limited treatment options linked to AFLD, adopting an epigenetic perspective is crucial, especially in the era of personalized and precision medicine.
5-hydroxymethylcytosine (5hmC) is an ancient but recently rediscovered and understood epigenetic modification in DNA and RNA, exhibiting significant potential for various biological processes.12 The distribution of 5hmC in the genome appears to be tissue-specific and highly influenced by the cellular state, enabling responses to environmental stimuli and metabolic disturbances.13,14 For instance, 5hmC plays a regulatory role in the phenotype of non-alcoholic fatty liver disease (NAFLD) by modulating the transcriptional activity of liver mitochondrial biogenesis and peroxisome proliferative activated receptor gamma coactivator 1 alpha (PPARGC1A).12 TET family proteins, including TET1, TET2 and TET3, can catalyze the conversion of 5-methyl-cytosine (5mC) to 5hmC.15,16 Recent studies have revealed that changes in DNMT1/TET2-mediated Cmip methylation regulate the development of NAFLD by controlling the guanylate binding protein 2 (GBP2)-PPARG-CD36 axis.17 Nevertheless, the regulatory effect of TET2 on AFLD remains unknown.
In mammals, the nucleus not only retains the non-splicing mRNA, but also maintains high-level reserves for various mature mRNAs.18,19 Under different stimuli such as stress, differentiation, virus infection, DNA damage, or neuronal activation, specific retained mRNAs are released into the cytoplasm, which enables rapid and massive translation of the corresponding proteins to assist the organism in responding to the stimuli.20 The crucial structure for mRNA nuclear retention is the paraspeckle, located in the interchromosome region of the nucleus.21 Paraspeckles consist of long noncoding RNA Neat1 (Lnc NEAT1) and proteins such as non-POU domain-containing octamer-binding protein (NONO), polypyrimidine tract-binding protein-associated splicing factor (PSF), paraspeckle component 1 (PSPC1), among others. NONO, as a core protein of paraspeckles, has been proven to bind to both double-stranded and single-stranded RNA and DNA.22 In the case of Ctn mRNA, which interacts with NONO and localizes to paraspeckles, it can be post-transcriptionally cleaved to release a translation-competent mRNA upon cellular stress.23 The nuclear retention or regulated selective output of mRNAs represents a new mechanism for controlling gene expression in mammalian cells while limited knowledge is currently available on this topic. Researchers discovered that methylation of mRNA is crucial for its nuclear export. Edens BM et al. demonstrated that N6-methyladenosine (m6A) can be recognized by fragile X mental retardation protein (FMRP), promoting nuclear export of methylated mRNA targets during neural differentiation.24 Recent reports have highlighted TET2’s capability to demethylate RNA.25,26 Is the 5mC level of RNA associated with the nuclear retention of mRNA? Moreover, due to the absence of RNA binding domains in TET2,16,27 the mechanisms underlying mRNA demethylation by TET2 remain poorly understood.
In this study, taking TET2 as the starting point, we’ve proven that NONO recruits TET2 into paraspeckles through direct interaction to fuse into the phase separation structure. Our findings indicate that TET2 regulates the nuclear retention of sterol regulatory element binding protein 1 (Srebp1) mRNA by affecting its methylation level, thus impacting the development of alcoholic fatty liver. The primary objective of this study is to investigate the molecular mechanism by which TET2 post-transcriptionally regulates the nuclear retention of mRNA related to lipid metabolism in paraspeckles, and how this subsequently affects lipid metabolism in mice. This research aims to identify effective targets and provide a theoretical basis for the clinical treatment of obesity-related diseases, while also revealing a new post-transcriptional regulatory mechanism of mRNA.
Results
TET2 knockout aggravates alcoholic fatty liver in mice
Alcohol abuse is associated with an elevated risk of multiple liver diseases. Previous studies have demonstrated that alcohol exposure promotes the progression of HCC and decreases TET2 expression in clinical HCC specimens and HCC cell lines.28 To assess the impact of TET2 on AFLD, we established a mouse model of AFLD by alcohol liquid feeding for 6 weeks. As shown in Figure 1A, the liver exposed to alcohol leads to liver swelling and yellowish appearance and Tet2 knockout aggravates this phenomenon (Figure 1A). The knockout efficiency of Tet2 gene is depicted in Figure S1A. During the induction of AFLD, the body weight of Tet2−/− mice showed a notable increase (Figure 1B). After alcohol liquid feeding for 6 weeks, the serum levels of alanine transaminase (ALT), aspartate aminotransferase (AST) and gamma-glutamyl transferase (γ-GT) exhibited a substantial increase in mice, aligning with the clinical diagnosis of alcoholic fatty liver.29,30 Moreover, these serum biochemical indexes in Tet2−/− mice were significantly higher than those in WT mice (Figure S1B). Compared with WT mice, the ratio of liver weight to body weight of Tet2−/− mice increased significantly after alcohol induction (Figure S1C). Histological analysis of liver tissue sections revealed Tet2 knockout aggravated the formation of alcohol-induced vacuoles (Figure 1C). Consistently, the size of lipid droplets was significantly greater in the Tet2−/− mice group compared to the WT mice group upon alcohol induction, indicating that Tet2 knockout promotes the formation of lipid droplets in alcoholic fatty liver as evidenced by oil red O staining (Figure 1D). Moreover, the measurement of triglyceride (TG) content confirmed that Tet2 knockout promotes the accumulation of TG in the alcohol-induced livers (Figure 1E, left panel). Hyperinsulinemia and insulin resistance are frequently observed pathophysiological mechanisms and prominent risk factors in various metabolic disorders. Specifically, they are strongly associated with fatty liver and other metabolic disorders characterized by lipid accumulation.31 Therefore, we examined the impact of Tet2 knockout on insulin level and insulin resistance. The results showed that Tet2 knockout markedly increased the level of serum insulin in mice regardless of alcohol exposure (Figure 1E, right panel). Oral glucose tolerance test (OGTT) and insulin tolerance test (ITT) assays revealed that Tet2 knockout significantly aggravated the impaired glucose tolerance and insulin resistance induced by alcohol in mice (Figures 1F and 1G).
Figure 1.
TET2 deficiency exacerbates alcohol-induced hepatic steatosis in mice
8- to 10-week-old WT mice and Tet2−/− mice were fed with an isocaloric diet or a liquid diet containing alcohol (37% of total calorie) for 6 weeks, then livers and blood were collected for analysis.
(A) Gross liver images obtained from WT mice and Tet2−/− mice fed with an isocaloric diet or a liquid diet containing alcohol for 6 weeks.
(B) Body weight changes observed during the development of alcoholic fatty liver by liquid diet or isocaloric diet containing alcohol treatment.
(C) Representative H&E staining in wild type mice liver and Tet2−/− mice liver after alcohol liquid diet or isocaloric diet treatment. Scale bar represents 50 μm.
(D) Representative Oil red O staining in WT mice liver and Tet2−/− mice liver after alcohol liquid diet or isocaloric diet treatment. Scale bars represent 100 μm (top row) and 50 μm (bottom row).
(E) Quantification of hepatic TG content and insulin levels in mice with fatty liver induced by alcohol.
(F) Assessment of blood glucose levels through oral glucose tolerance test (OGTT) and insulin tolerance test (ITT).
(G) The area under the curve (AUC) in OGTT and ITT.
(H) Representative Oil red O staining in primary hepatocytes isolated from both WT and Tet2−/− mice and were treated with 100 mM alcohol and indicated concentrations of FFAs for 24 h. Scale bars represent 20 μm.
(I) Measurement of triglyceride content in primary hepatocytes isolated from WT and Tet2−/− mice exposed to both 100 mM alcohol and 2 mM FFAs. Data are expressed as means ± SD, n ≥ 5 mice per group, representative of two experiments, ∗p < 0.05, ∗∗p < 0.01 and ∗∗∗p < 0.001 were determined by the one-way ANOVA.
To further investigate the impact and mechanisms of TET2 on fatty liver formation in vitro, we isolated primary mouse hepatocytes and simulated fatty liver conditions through alcohol and free fatty acids (FFAs) treatment. Oil Red O staining revealed significantly larger lipid droplets in Tet2−/− hepatocytes compared to WT mouse hepatocytes under the indicated stimulation of different FFAs concentrations (Figure 1H). The results of TG content in primary hepatocytes exhibited a similar trend (Figure 1I). The results of periodic acid Schiff (PAS) glycogen staining revealed that there were significant differences in glycogen accumulation between primary hepatocytes of Tet2−/− mice and WT mice under different glucose concentrations (Figure S1D). This suggests that Tet2 knockdown also leads to abnormal glycogen accumulation in hepatocytes. Excessive glycogen accumulation not only disrupts glucose metabolism, but also enhances the conversion of glycogen to triglycerides, which in turn aggravates fatty liver.32 Therefore, Tet2 knockout aggravates the formation of alcoholic fatty liver and metabolic disorder and the accumulation of fat in hepatocytes.
Conversely, whether alcoholic fatty liver affect TET2 expression? Although TET2 was down-regulated by alcohol in HCC liver tissue and HCC cell line,28 there was no significant change in TET2 expression in rats with alcoholic liver disease.33 Initially, we examined TET2 expression in WT mouse livers of AFLD model using qPCR and western blotting. Our findings indicated that alcohol exposure did not significant alter TET2 RNA and protein levels (Figures S1E and S1F). To further investigate potential alterations in TET2 expression specifically in hepatocytes—the primary cell type affected by AFLD, we conducted immunohistochemistry (IHC) on serial liver sections. IHC was combined with staining for mature hepatocytes marker (hepatocyte nuclear factor-4 alpha, HNF4α) and TET2. The results revealed that alcohol exposure did not exert a notable effect on TET2 expression in hepatocytes in the mouse liver (Figure S1G). Given TET2’s role as a widely expressed DNA/RNA demethylase involved in various biological processes, it appears to be minimally affected by alcohol in our AFLD model.
TET2 regulates the formation of alcoholic fatty liver by SREBP1
Previous studies have reported that acetaldehyde, which is produced by ethanol metabolism, increases the synthesis of mature SREBP1 and promotes liver adipogenesis, leading to the occurrence of fatty liver.34 Consistent with these findings, our study demonstrates that alcohol exposure leads to the upregulation of SREBP1 (Figures 2A and 2B). SREBP1 is a pivotal transcription factor in the de novo fatty acid synthesis pathway, a critical metabolic pathway for fat in the liver.35 Subsequently, we explored the effect of TET2 on SREBP1 protein expression in mouse liver and HepG2 cell lines. The results demonstrated a significant increase of SREBP1 protein expression in Tet2−/− liver, while overexpression of Tet2 resulted in a decrease of SREBP1 protein in HepG2 cells, as shown in Figures 2A and 2B. The precursor of SREBP1 undergoes a two-step proteolysis process, releasing SREBP1 (N-SREBP1) in the form of N-terminal transcriptional activity (∼70Kd). The released N- SREBP1 is transferred to the nucleus, stimulating the expression of adipogenic genes. IHC staining revealed that alcohol treatment enhanced the nuclear localization of SREBP1, with a significantly higher nuclear signal observed in the Tet2−/− -Alcohol group compared to the WT alcohol group (Figure 2C). These results suggest that knockout of Tet2 not only elevates the protein expression of SREBP1, but also enhances the activation of SREBP1.
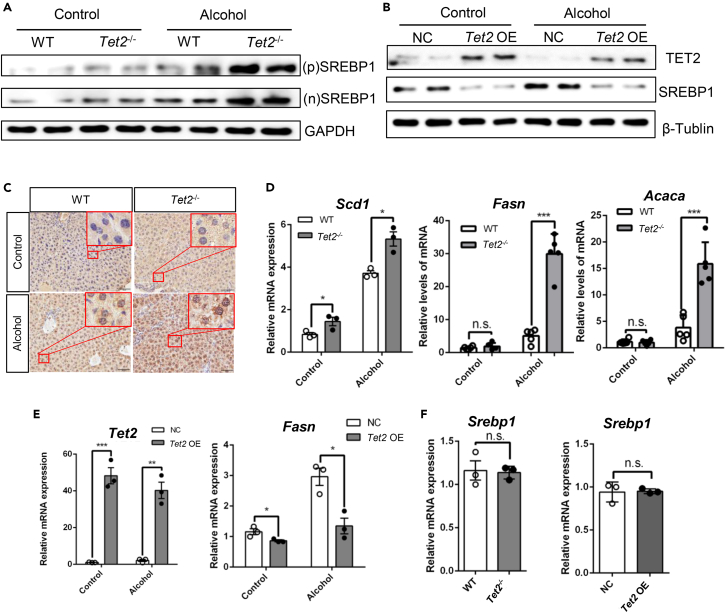
Figure 2.
TET2 facilitates the formation of alcoholic fatty liver by SREBP1
(A) Expression of SREBP1 was measured at the protein level in WT and Tet2−/− mouse livers after alcohol liquid diet or isocaloric diet treatment for 6 weeks.
(B) Protein levels of TET2 and SREBP1 in HepG2 cells transfected with Tet2 or pcDNA3.1 plasmid for 48 h.
(C) IHC staining for SREBP1 in WT and Tet2−/− mice liver after alcohol liquid diet treatment or isocaloric diet treatment for 6 weeks. Scale bar represents 50 μm.
(D) Hepatic mRNA relative expression levels of Scd1, Fasn, and Acaca were determined by qPCR analysis in WT and Tet2−/− mice after alcohol liquid diet or isocaloric diet treatment for 6 weeks.
(E) qPCR analysis of Tet2 and Fasn expression in HepG2 cells transfected with pcDNA3.1-Tet2 or pcDNA3.1 plasmid for 48 h.
(F) qPCR analysis of Srebp1 in mouse livers after alcohol liquid diet or isocaloric diet treatment for 6 weeks and HepG2 cells transfected with pcDNA3.1-Tet2 or pcDNA3.1 plasmid for 48 h; Data are expressed as means ± SD, n ≥ 3 mice per group, representative of two experiments, ∗p < 0.05, ∗∗p < 0.01 and ∗∗∗p < 0.001 were determined by the one-way ANOVA.
As a transcription factor, SREBP1 governs numerous genes involved in lipid synthesis, including Fasn, Acaca, and Scd1.36 Therefore, we examined the expression of Fasn, Acaca, and Scd1. The results showed that Tet2 knockout significantly upregulated transcription of Fasn, Acaca, and Scd1 in mice, while Tet2 overexpression led to a significant reduction of the mRNA of Scd1 in HepG2 cells (Figures 2D and 2E). However, neither overexpression nor knockdown of Tet2 had a significant impact on the transcript level of Srebp1 (Figure 2F). These results indicate that TET2 affects hepatic fat metabolism by regulating the translation of Srebp1 rather than its transcription.
The absence of Tet2 exacerbates alcohol-induced lipid accumulation in mouse liver. Besides increased fatty acid synthesis, is it also related to increased fatty acid uptake in the liver? As we know, CD36 is responsible for fatty acid uptake as a fatty acid transporter enzyme.37 To explore whether the hepatic fat accumulation observed in Tet2−/− mice was due to increased hepatic de novo lipogenesis or enhanced hepatic fatty acid uptake or both, we examined serum TG levels, as well as hepatic CD36 expression in mice. As depicted in Figure S2A, no significant difference was observed between WT and Tet2 knockout mice under control treatment conditions. However, alcohol-induced Tet2 knockout mice markedly elevated serum TG levels. Both qPCR and western blotting results consistently demonstrated a significant upregulation of CD36 in response to both TET2 knockout and alcohol treatment. Notably, no significant difference in hepatic CD36 expression was observed between Tet2 knockout and WT mice following alcohol treatment (Figures S2B and S2C). In the control groups, Tet2 knockout led to a slight up-regulation of CD36 without altering serum TG levels. Although alcohol treatment significantly increased CD36 expression, there was no discernible difference between Tet2 knockout mice and WT mice. This suggests that while alcohol treatment enhanced the liver’s ability to uptake fatty acids, the phenotypic difference in fatty liver between the Tet2−/−-Alcohol group and WT-Alcohol group was primarily attributed to the up-regulation of fatty acid synthesis.
TET2 promotes the distribution of Srebp1 mRNA in nuclear
To investigate how TET2 affected the translational level of SREBP1, we conducted the nuclear-cytoplasmic separation experiments. We validated our nucleoplasmic separation experiments' reliability by detecting the expression of CoxⅡ and U6 in the cytoplasmic and nuclear fractions, respectively. CoxⅡ, as a mitochondrial gene, has its mRNA mainly distributed in the cytoplasm, while U6 snRNA is mainly distributed in the nucleus. The results demonstrated predominant CoxⅡ expression in the cytoplasmic fraction and U6 expression in the nuclear fraction, thereby confirming the success of the nucleoplasmic separation process (Figure S3D). The result revealed a significant increase in Srebp1 mRNA within the nucleus, accompanied by a decrease in the cytoplasm after Tet2 overexpression in HepG2 cells (Figure 3A). And Tet2 knockout resulted in a decrease of Srebp1 mRNA in the nucleus and an increase in the cytoplasm (Figure 3B). Based on this observation, we formulated a hypothesis that TET2 modulates the nucleoplasmic distribution of Srebp1 mRNA. To validate this hypothesis, we performed in situ hybridization experiments. As shown in Figure 3C, Srebp1 mRNA was predominantly retained in the cytoplasm in Tet2−/− mouse hepatocytes compared to WT mouse hepatocytes. Interestingly, the in-situ hybridization displayed Srebp1 mRNA scattered in the cytoplasm, but gathered bright spots in the nucleus (Figure 3C). Currently, it is widely acknowledged that the 3′ untranslated region (UTR) of mRNA affects mRNA translation by controlling its nuclear export.38,39 Hence, we developed an RNA imaging system utilizing fluorescence to enable real-time tracking of Srebp1 mRNA. The underlying principles of this system are illustrated in Figure S3A. Previous studies have utilized 24 copies of a short hairpin sequence inserted into the 3′UTR, along with co-expression of the PP7 phage coat protein, to fluorescently label mRNA. The protein binds to hairpin sequence with high affinity and fused with three copies of mCherry (PP7-mCherry), enabling real-time tracking of RNA in living cells.40 On this basis, we inserted NLS sequence into the N-terminal of mCherry protein, so that it can actively enter the nucleus for Srebp1 mRNA binding, and exit the nucleus with the nuclear output of RNA. In this system, a lot of 3∗mCherry can bind to 24∗PP7 bind sites on Srebp1 mRNA, which can form red bright spots in the field of vision (Figure S3B). Schematic overview of the plasmids used for RNA imaging system were displayed in Figure S3C. By utilizing this imaging system, we observed that Srebp1 mRNA decreased in the cytoplasm, while increased significantly in the nucleus after overexpression of Tet2 in HepG2 cells (Figure 3D). To further verify the regulatory effect of TET2 on Srebp1 mRNA, we generated a mutant plasmid for the Srebp1 3′UTR mRNA to investigate whether the disruption of TET2’s effect on 3′UTR methylation promotes the nuclear export of Srebp1 mRNA (Figure 3E). WT and 3′UTR methylation site mutant Srebp1 plasmids were separately transfected into HepG2 cells. Following nuclear and cytoplasmic separation, qPCR was employed to assess whether Tet2 overexpression affected the nuclear and cytoplasmic distribution of mutant Srebp1 mRNA. The results indicated that overexpression of Tet2 in cells transfected with WT Srebp1 still significantly reduced the amount of Srebp1 mRNA in the cytoplasm while elevating its level in the nucleus (Figure 3F). However, overexpression of Tet2 in cells transfected with 3′UTR mutant Srebp1 had no significant effect on RNA nuclear retention (Figure 3G). These consistent findings provide evidence that TET2 promotes the nuclear retention of Srebp1 mRNA.
Figure 3.
TET2 facilitates nuclear localization of Srebp1 mRNA in hepatocytes
(A) HepG2 cells transfected with pcDNA3.1-Tet2 or pcDNA3.1 plasmid for 48 h. Then, cells were separated the nuclear and cytoplasmic fractions and isolated RNA, respectively. qPCR analysis of Srebp1 in different cellular fractions.
(B) Primary hepatocytes isolated from WT and Tet2−/− mice exposed to alcohol and FFAs for 48 h. Then, cells were separated the nuclear and cytoplasmic fractions and isolated RNA, respectively. qPCR analysis of Srebp1 in different cellular fractions.
(C) Representative images from RNAscope ISH assays for Srebp1 on mouse liver paraffin section treated with alcohol liquid diet or isocaloric diet treatment for 6 weeks. Green signal represents Srebp1, blue shows DAPI. Scale bars, 20 μm.
(D) Evaluation of the impact of TET2 on the localization of Srebp1 mRNA by PP7-mCherry RNA imaging system in HEK293T cells. Red spot represents Srebp1, blue shows DAPI. Scale bars, 10 μm.
(E) Schematic overview of the plasmids of pcDNA3.1- Srebp1 WT and pcDNA3.1-Srebp1▵3′UTR.
(F) HepG2 cells transfected with pcDNA3.1- Srebp1 WT and pcDNA3.1-Tet2 or pcDNA3.1 plasmid for 48 h. Then, cells were separated the nuclear and cytoplasmic fractions and isolated RNA, respectively. qPCR analysis of Srebp1 in different cellular fractions.
(G) HepG2 cells transfected with pcDNA3.1-Srebp1▵3′UTR and pcDNA3.1-Tet2 or pcDNA3.1 plasmid for 48 h. Then, cells were separated the nuclear and cytoplasmic fractions and isolated RNA, respectively. qPCR analysis of Srebp1 in different cellular fractions.Data are expressed as means ± SD, n ≥ 3 mice per group, representative of two experiments, ∗∗p < 0.01 and ∗∗∗p < 0.001 were determined by the one-way ANOVA.
NONO mediates recruiting of TET2 to the paraspeckles
Nuclear retention of mRNA is a novel and previously underestimated form of gene regulation that enables cells to respond rapidly to stress, viral infection, differentiation signals, or changing environmental conditions.41 Nuclear retention of mRNA is one of the most classical functions of paraspeckles.42 It has been reported that TET2 can be recruited to the transcriptional active site by PSPC1.25 Considering that both PSPC1 and NONO function as core proteins in paraspeckles, we speculated that TET2 might interact with paraspeckles. Co-immunoprecipitation (Co-IP) experiments employing TET2 and NONO antibodies confirmed that TET2 and NONO proteins bind to each other (Figure 4A). The expression of TET2-mCherry fusion protein demonstrated that most TET2 co-located with NONO fluorescence in HepG2 cells (Figure 4B). Furthermore, the immunofluorescence signals of TET2 and NONO exhibited co-localization (Figure 4C), providing additional evidence supporting their interaction and potential functional relationship. Moreover, combined the PP7-mCherry RNA imaging system and immunofluorescence staining of TET2 and NONO in HepG2 cells revealed co-localization of Srebp1 with TET2 and NONO (Figure 4D). To validate the pivotal role of NONO in TET2, we employed siRNA-mediated knockdown of NONO. We designed siRNAs targeting three sites within the Nono CDS region and assessed knockdown efficiency via quantitative PCR (qPCR). Among these, siRNA-299 exhibited the highest efficacy, and we consequently selected siRNA-299 for all subsequent experiments (Figure 4E). Subsequent evaluation by qPCR and Western Blot revealed a significant reduction in TET2 expression (Figures 4F and 4G). Additionally, fluorescence co-localization analysis demonstrated a notable decrease in the overlap between TET2 and NONO spots, and TET2 distribution within the nucleus became more diffuse following NONO knockdown (Figure 4H). These findings imply that NONO plays a crucial role in mediating the regulatory function of TET2 within paraspeckles. Considering that paraspeckles are liquid-liquid phase separation, we speculated that TET2 may interact with the paraspeckles in a similar manner. As shown in Figure 4I, the TET2-mCherry fluorescence recovered rapidly after bleaching, indicating that TET2 forms liquid-liquid phase separation. These results illustrate that NONO recruits TET2 to paraspeckles.
Figure 4.
NONO mediates recruiting of TET2 to paraspeckles
(A) Co-immunoprecipitation (Co-IP) assay detected the interaction between TET2 and NONO in WT mouse livers. Immunoprecipitation was performed using anti-TET2 and anti-NONO antibody, followed by immunoblotting for TET2 and NONO proteins.
(B) HepG2 cells transfected with Tet2-mCherry plasmid for 48 h, followed by immunofluorescence staining with NONO. Red represents TET2-mCherry, green represents NONO, blue shows DAPI. Scale bars, 20 μm.
(C) Representative images from immunofluorescence (IF) staining of TET2 and NONO in HepG2 cells. Red represents TET2, green represents NONO, blue shows DAPI. Scale bars, 20 μm.
(D) Representative images from the PP7-mCherry RNA imaging system combined with IF for TET2 and NONO. White represents Srebp1, red represents TET2, green represents NONO, blue shows DAPI. Scale bars, 5 μm.
(E) HepG2 cells were transfected with Nono siRNAs for 48 h. Nono knockout efficiency detected by qPCR.
(F) qPCR analysis of Tet2 expression in HepG2 cells transfected with Nono siRNA for 48 h.
(G) Protein levels of TET2 and SREBP1 in HepG2 cells transfected with Nono-siRNA for 5 days.
(H) HepG2 cells transfected with Nono-siRNA for 5 days, followed by IF staining with NONO and TET2. Red represents TET2, green represents NONO, blue shows DAPI. Scale bars, 2 μm.
(I) Fluorescence Recovery After Photo bleaching (FRAP) analysis in HepG2 cells overexpressing TET2-mCherry, assessing the dynamics of TET2 localization. Scale bars, 10 μm.Data are expressed as means ± SD, n ≥ 3 mice per group, representative of two experiments, ∗p < 0.01 and ∗∗p < 0.001 were determined by the one-way ANOVA.
TET2 affects the methylation level of Srebp1 mRNA and regulates the nuclear retention in paraspeckles
We next investigated how TET2 affected the nuclear retention of Srebp1 mRNA. Results from RNA immunoprecipitation using TET2 antibody followed by qPCR confirmed the interaction between TET2 and Srebp1 mRNA (Figure 5A). Immunohistochemical staining of 5hmC in the liver revealed a significantly lower 5hmC signal in the Tet2−/− group compared to the WT group (Figure 5B). Subsequent sequencing of bisulfite-treated mouse liver Srebp1 mRNA showed that a portion of the methylation sites in the 3′UTR of Srebp1 mRNA could not be demethylated in the Tet2−/− mice (Figure 5C), suggesting that TET2 binds to Srebp1 mRNA and demethylates its 3′UTR. To explore whether TET2 regulates the nuclear retention of Srebp1 mRNA through paraspeckles, we performed immunofluorescence and in situ hybridization on NONO and Srebp1 mRNA in WT mouse livers. As shown in Figure 5D, most of NONO signals and bright spots of nuclear Srebp1 mRNA were co-located, indicating that this portion of Srebp1 mRNA was retained in the paraspeckles. Furthermore, we observed that TET2 also influences the number of paraspeckles. The protein expression of NONO in the liver of Tet2−/− mice was significantly decreased, whereas TET2 overexpression in cells led to a significant increase in NONO levels (Figures 5E and 5F). Regardless of alcohol exposure, Tet2 knockout resulted in a notable decrease of the paraspeckles skeleton Lnc Neat1 (Figure 5G). Considering that stability of RNA is affected by the methylation state, we assessed the stability of Srebp1 mRNA and found that Tet2 overexpression caused a certain degree of reduction in the stability of Srebp1 mRNA (Figure 5H). These results indicate that TET2 influences the methylation level of Srebp1 mRNA and regulates its nuclear retention in paraspeckles.
Figure 5.
TET2 modulates methylation levels of Srebp1 mRNA and regulates nuclear retention in paraspeckles
(A) qPCR analysis of SREBP1 RNA immunoprecipitation (RIPed) samples from WT mouse liver treated with alcohol liquid diet for 6 weeks, normalized to the 1% input after incubation.
(B) IHC staining for 5hmC in WT and Tet2−/− mice liver after alcohol liquid diet treatment for 6 weeks. Scale bars, 100 μm.
(C) RNA bisulfite conversion sequencing assay was performed in WT and Tet2−/− mice liver after alcohol liquid diet treatment for 6 weeks demonstrating demethylation sites on the Srebp1 mRNA 3′ UTR.
(D) Representative images from RNAscope ISH assays for Srebp1 combined with IF for NONO was performed on livers paraffin sections of WT mice treated with alcohol liquid diet for 6 weeks. Green represents Srebp1, red represents NONO, blue shows DAPI. Scale bars, 50 μm.
(E) Expression of NONO was measured at the protein level in WT and Tet2−/− mouse livers after alcohol liquid diet or isocaloric diet treatment for 6 weeks.
(F) Protein levels of TET2 and NONO in HepG2 cells transfected with Tet2 or pcDNA3.1 plasmid for 48 h.
(G) Relative lncRNA NEAT1 levels were determined by qPCR analysis in WT and Tet2−/− mice liver after alcohol liquid diet treatment for 6 weeks.
(H) HepG2 cells transfected with Tet2 or pcDNA3.1 plasmid for 48 h, followed by ActD treatment for different time. Assessment of the effect of TET2 on RNA stability of Srebp1 mRNA by qPCR analysis.Data are expressed as means ± SD, n ≥ 3 mice per group, representative of two experiments, ∗p < 0.05 and ∗∗p < 0.01 were determined by the one-way ANOVA.
Taken together, our data demonstrate that TET2 plays a crucial role in the regulation of alcoholic fatty liver formation through the nuclear retention of Srebp1 mRNA in paraspeckles.
Discussion
Given the growing prevalence of fatty liver disease,43 it is of paramount importance to explore the underlying research and mechanisms pertaining to hepatic fat metabolism. In this study, we highlight a robust association between Tet2 and alcohol fatty liver and identified an innovative mechanism through which TET2 modulates fat metabolism in the liver. Our findings demonstrate that TET2 interacts with Srebp1 mRNA via NONO, facilitating its nuclear retention. Consequently, TET2 regulates the protein expression of SREBP1, which in turn affects the formation of fatty liver.
Considerable efforts have been devoted to exploring the relationship between epigenetics and cancer.44,45 However, understanding of epigenetics in fatty liver and its associated metabolic syndrome remains limited. Specifically, the epigenetic modification 5hmC has recently garnered research attention, with TET family proteins identified as its regulators.46,47 Alcohol consumption can induce cellular damage and alter the epigenetic landscape.48 A previous study found that chronic ethanol exposure leads to hepatocyte apoptosis, accompanied by diminished TET1 levels and heightened 5hmC formation.33 This study is the first to show that TET2 can also affect the development of AFLD, mainly including lipid droplet accumulation and insulin resistance. Notably, Tet2−/− mice exhibited higher serum insulin concentrations compared to WT mice. This observation implies the presence of insulin resistance, with elevated insulin levels representing a compensatory response to maintain blood glucose homeostasis. However, this compensatory mechanism also places an increased burden on pancreatic islet β cells.49
The liver and gastrointestinal tract play a central role in alcohol metabolism. Ethanol is initially oxidized to acetaldehyde by alcohol dehydrogenase (ADH) Acetaldehyde facilitates the translocation of SREBP1 from endoplasmic reticulum to Golgi, protein hydrolysis and maturation in Golgi to its active form.50 Once activated, SREBP1 upregulates the expression of downstream factors such as FASN, ACC, and SCD1. This leads to increased synthesis and accumulation of TG, resulting in the ectopic deposition of lipids in the liver and the development of fatty liver.35 Our results align with these findings. Alcohol exposure promotes the expression and maturation of Srebp1. Regardless of Tet2 knockout and overexpression, the protein expression of Srebp1 precursor form and mature form increased significantly under the stimulation of alcohol. Moreover, compared to the control group, the transcriptional levels of Srebp1 target genes-Fasn, Acaca, and Scd1, show a significant increase after alcohol stimulation. This indicates that alcohol promotes the transcriptional activity of Srebp1. Additionally, we observed a significant increase in Srebp1 mRNA signals in the livers of mice exposed to alcohol.
Our study put forward that paraspeckles play a crucial role in mediating the interaction between TET2 and RNA, thereby influencing its nuclear retention. Although previous studies have reported a potential role for TET proteins in mediating the oxidation of 5mC to 5hmC in RNA,51 it is unclear how TET2 binds RNA due to the absence of an RNA-binding structural domain in TET2. It was reported that TET2 interacting with NONO in mass spectrometry data in an article detailing the interaction between TET2 and PSPC1.25 NONO, recognized as an RNA binding protein and a core component of paraspeckles, is known to participate in the nuclear retention of mRNA within paraspeckles. Motivated by this information, we selected NONO as the primary focus for studying the regulation of mRNA nucleoplasmic distribution. Our results suggest that TET2 can bind to Srebp1 mRNA via NONO protein, presenting a new perspective on the RNA-related actions of TET2. Many studies have shown that 5mC is involved in a variety of RNA metabolism, including mRNA export,52 RNA stabilization,53 and translation.54 And The 5mC modification of mRNA is enriched around 5′UTR and 3′UTR.55 The 3′UTR of mRNA is involved in many processes that regulate mRNA, such as mRNA localization, mRNA stability, and translation.56 Shen et al. found that Tet2 knockdown caused a significant increase in 5mC located in the mRNA 3′UTR, further confirming the regulatory effect of TET2 on the mRNA 3′UTR.57 Based on these findings, our study focused on the regulatory role of TET2 affecting 3′UTR methylation. Further, our data suggests that TET2 regulates its nuclear retention by modulating the methylation level of the Srebp1 mRNA 3′UTR. Paraspeckles, which are substructures within the cell nucleus, act as sponges or platforms to adsorb or recruit various RNAs, proteins, or DNA, thus performing diverse biological functions. One of the most classical functions is the nuclear retention of mRNAs containing inverted repeat sequences in the 3′UTR region.58 NONO specifically binds to these mRNAs and retains them in the nucleus to regulate their translation, a process that is dynamic and finely regulated.59 Our data revealed that TET2 not only interacts with NONO, but also co-localizes with the paraspeckle. This implies that TET2 can affect not only Srebp1 mRNA retention, but also may affect other biological processes in the paraspeckle.
Additionally, we developed an imaging system to enable visual tracking of Srebp1 mRNA in living cells. This system utilized orthogonal bacteriophage PP7 and MS2 stem-loops to label the transcript within both the coding sequence (PP7) and the 3′UTR (MS2) with fluorescent proteins that emit distinct spectra.60 By incorporating multiple recognition sites on a single mRNA, the pp7-mCherry molecules formed clusters, resulting in the visualization of bright red dots representing mCherry-labeled mRNA molecules.61 However, the previously reported PP7-mCherry system lacked the ability to label RNA in the nucleus due to the restricted nuclear entry of the 3∗mCherry protein, which has a molecular weight exceeding 50 kDa.62 To overcome this limitation, we inserted the SV40 NLS nuclear localization sequence into the N-terminal of 3∗mCherry protein. This modification facilitated active nuclear entry of the 3∗mCherry protein, allowing our imaging system to label RNA within the nucleus.63 Therefore, our imaging system is able to label RNA in the nucleus and the 3∗mCherry protein is able to exit the nucleus along with the nuclear export of RNA. Using the PP7-NLS-3∗mCherry imaging system, we observed that TET2 promoted the nuclear retention of Srebp1 mRNA, consistent with the previous results.
In summary, this study reported for the first time the regulatory effect of TET2 on alcoholic fatty liver. We discovered that TET2 interacts with paraspeckles to demethylate Srebp1 mRNA 3′UTR, thus influencing the nuclear retention in paraspeckles, and subsequently modulating the expression of SREBP1 to regulate hepatic lipid production. Therefore, comprehending these epigenetic modifications to target the primary cause of the disease might prove a rational strategy to prevent the disease and develop innovative therapeutic interventions. This investigation establishes a connection between epigenetic modifications and fat metabolism, thus enhancing our understanding of the mechanisms underlying the formation and progression of AFLD. In another prevalent fatty liver disease, NAFLD, SREBP1 also holds significance in regulating the development of fatty liver.64 We hypothesize that TET2 may similarly influence the nuclear and cytoplasmic distribution of Srebp1 mRNA in NAFLD. The widely accepted "Multiple-hit" theory constitutes the classical pathogenesis of NAFLD, indicating a complex interplay of factors influencing its formation.65 Whether the regulatory impact of TET2 on RNA can exert a predominant influence amid numerous contributing factors, leading to a significant phenotype, necessitates further experimental investigation. Furthermore, the influence of TET2 on RNA methylation and its subsequent impact on nuclear and cytoplasmic distribution may extend to a broader spectrum of genes, warranting comprehensive exploration in future studies.
Limitations of the study
Our study shows that TET2 knockout causes changes in Srebp1 mRNA 3′UTR methylation and then affects its nuclear export. However, how changes in methylation affect the nuclear export of Srebp1 mRNA requires further exploration. In addition, although nuclear retention of mRNA is strongly correlated with the 3′UTR,66,67 we do not have sufficient evidence to demonstrate whether the same methylation changes and effects exist in other regions of the Srebp1 mRNA. The molecular mechanisms by which the methylation levels of different regions of mRNA affect its nucleoplasmic distribution may be revealed in future studies.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Polyclonal mouse anti-TET2 | Active motif | Cat#61389; RRID: AB_2736901 |
| Polyclonal rabbit anti-NONO | Abcam | Cat#ab70335; RRID: AB_1269576 |
| Polyclonal rabbit anti-SREBP1 | Bioss | Cat#bs-1402R; RRID: AB_10854434 |
| Polyclonal rabbit anti-5hmC | Active motif | Cat#40900; RRID: AB_01419007 |
| Polyclonal mouse anti-GAPDH | Proteintech | Cat#60004-I-Ig; RRID: AB_2737588 |
| Polyclonal rabbit anti-β-Tublin | Abmart | Cat#M30109S; RRID: AB_2916070 |
| HRP-conjugated Goat Anti-Rabbit IgG(H + L) | Proteintech | Cat#SA00001-2; RRID:AB_2722565 |
| HRP-conjugated Goat Anti-Mouse IgG(H + L) | Proteintech | Cat#SA00001-1; RRID: AB_2722564 |
| Monoclonal rabbit anti-CD36 | Huabio | Cat#SA00001-1 |
| Mouse IgG | Millipore | Cat#CS200621 |
| Rabbit IgG | Millipore | Cat# PP64B |
| Goat anti-Rabbit IgG (H + L) Secondary Antibody, Alexa Fluor 555 | Invitrogen | Cat#A-21428; RRID: AB_2535849 |
| Goat anti-Mouse IgG (H + L) Secondary Antibody, Alexa Fluor 555 | Invitrogen | Cat#A-21422; RRID: AB_2535844 |
| Goat anti-Rabbit IgG (H + L) Secondary Antibody, Alexa Fluor 488 | Invitrogen | Cat#A11008; RRID: AB_2534069 |
| Chemicals, peptides, and recombinant proteins | ||
| DAPI | Abcam | Cat#104139 |
| Oil Red O | Sigma | Cat#1320-06-5 |
| Actinomycin D | MCE | Cat#HY-17559 |
| Lipofectamine 3000 | Invitrogen | Cat#L3000015 |
| Critical commercial assays | ||
| Magna RIP™ RNA-Binding Protein Immunoprecipitation Kit | Merk | Cat#17-700 |
| RNAscope Multiplex Fluorescent Reagent Kit v.2 | Advanced Cell Diagnostics | Cat#323110 |
| RNAscope H202 and Protease Reagents | Advanced Cell Diagnostics | Cat#322381 |
| Hematoxylin and Eosin Staining Kit | Beyotime | Cat#C0105S |
| Triglyceride Quantification kit | Nanjing jiancheng | Cat#A110-1-1 |
| EZ RNA Methylation Kit | Zymo | Cat#R5001 |
| GOT/AST kit | Nanjing jiancheng | Cat#C010-2 |
| GPT/ALT kit | Nanjing jiancheng | Cat#C009-2 |
| γ- Glutamyl transferase Assay Kit | Nanjing jiancheng | Cat#C017-2-1 |
| Experimental models: Cell lines | ||
| HepG2 | ATCC | Cat#HB-8065 |
| 293T | ATCC | Cat#CRL-3216; RRID: CVCL_0045 |
| Experimental models: Organisms/strains | ||
| Tet2−/− mice | Shanghai Model Organisms | strain name: B6; 129S-Tet2tm1Smoc, stock number: NM-KO-00127 |
| C57BL/6 mice | Laboratory Animal Center | N/A |
| Oligonucleotides | ||
| See Tables S1 and S2 | ||
| Recombinant DNA | ||
| pcDNA3.1(+)-Tet2-mCherry plasmid | This paper | N/A |
| pHR-NLS-tdPP7-3∗mCherry | This paper | N/A |
| PcDNA4TO-Srebp1 3′UTR-24∗PP7 | This paper | N/A |
| pcDNA3.1(+)-Srebp1 ▵3′UTR plasmid | This paper | N/A |
| pcDNA3.1(+)-Srebp1 WT plasmid | This paper | N/A |
| Software and algorithms | ||
| GraphPad Prism 8 | GraphPad Prism | https://www.graphpad.com/ |
| Zen 2.3 lite | Carl Zeiss Microscopy | https://www.zeiss.com/microscopy/int/products/microscope-software.html |
| Other | ||
| Lieber-DeCarli Ethanol and Control liquid diet | Dyets | Cat#710028 |
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact, Lisheng Zhang (lishengzhang@mail.hzau.edu.cn).
Materials availability
This study did not generate new unique reagents.
Data and code availability
Data: All data reported in this paper will be shared by the lead contact upon request.
Code: This paper does not report original code.
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Experimental model and study participant details
Animals
WT C57BL/6 mice (female, 8 to 10-week-old) were obtained from Experimental Animal Center of Huazhong Agricultural University (Wuhan). Tet2−/− mice on a C57BL/6 background were provided by Shanghai Model Organisms, and back-crossed to the C57BL/6 background. WT mice and Tet2−/− mice were acclimated by feeding for 1 week. Then mice were fed with an isocaloric diet or a Lieber-DeCarli ethanol liquid diet (37% of total calorie) for 6 weeks68,69 Mice were humanely killed, and livers and blood were collected for further analysis. All animal procedures were approved by the animal ethics and welfare committee of Huazhong Agricultural University. The ethics number of the experimental animal research is HZAUMO-2020-0099.
Method details
Cell and plasmid
HepG2 and HEK293 cell lines were cultured in high glucose DMEM (Hyclone, SH30243) supplied with 10% (v/v) fetal bovine serum (FBS; Gibco, 10099) and 1% (v/v) penicillin-streptomycin (Hyclone, sv30010). Cells were cultured in a carbon dioxide incubator (Thermo Scientific, V531205) with 5% CO2 at 37°C siRNAs targeting Nono were obtained from Genechem. The oligo sequences are listed in Table S3. Transfection was carried out with Lipofectamine 3000 (Invitrogen, L3000001) according to the manufacturers’ protocols. To establish an AFLD model in vitro, we induced cells with 100 mM alcohol and 2 mM FFAs.
For isolation of primary mouse hepatocytes, the WT and Tet2−/− mice (female, 6 to 8-week-old) were anesthetized with Avertin (Sigma, T48402, 240 mg/kg body weight) by intraperitoneal injection. The liver perfusion was done by injecting needle into the portal vein and providing the following solutions sequentially: 50 mL of EBSS (Sigma, E6276) supplemented with 0.5 mM EGTA (Sigma, E3889) and then 50 mL of HBSS (Sigma, H1641) supplemented with 100 U/mL of Collagenase IV (Invitrogen, 17101-015), and 0.05 mg/mL of Trypsin inhibitor (Sigma, T2011). The perfused liver was carefully taken out, put onto a Petri dish, added 25 mL of hepatocyte wash media (Invitrogen,17704-024), and massaged with two cell scrapers until the liver has become apart with only connective tissue left behind. Dissociated cells were passed through funnel with mesh into 50 mL of centrifugal tube. After centrifugation at 900 rpm for 5 min, cell pellet was resuspended in hepatocyte wash media, which were carefully overlaid percoll (Sigma, P4937) solution (50%). After centrifugation at 900 rpm for 10 min, harvested cell pellet was washed twice with hepatocyte wash media, and then suspended in Williams’ E medium (Invitrogen, 12551–032) supplemented with 10% FBS, 1% penicillin/streptomycin and were seeded into 6-well, 24-well and 48-well.
OGTT and ITT
At 6 weeks after ethanol exposure, we performed oral glucose tolerance tests (OGTTs) and insulin tolerance tests (ITTs). For the OGTT, mice were fasted overnight and gavaged with glucose (i.g. injection of a 20% solution, 1 mg/g body weight), and blood glucose was measured using a Bayer Contour glucometer after 0, 15, 30, 60, 90 and 120 min. The ITT was performed approximately 2 days after the OGTT. After fasting for 4 h and recording basal blood glucose, i.p. injection of insulin was performed at a dose of 0.75 IU/kg body weight for each mouse, and blood glucose was measured using a Bayer Contour glucometer after 15, 30, 60, 90 and 120 min.
H&E and immunohistochemistry staining
Mouse livers were fixed in 4% paraformaldehyde for at least 48 h. The tissue for paraffin-embedded sections was dehydrated, embedded, and cut in 5 μm-thick sections. After dewaxing and hydration, paraffin sections were stained with 7 min in hematoxylin, and dehydrated after reverse blue with ammonia. Then stain them in eosin solution for 1 min. After dehydration, sections were cleared in xylene and mount in resinous medium. As for immunohistochemistry, the sections were immunostained with antibodies against SREBP1 (1:200 dilution, Bioss, bs-1402R) and 5hmC (1:200 dilution, Active Motif, 40900) and a secondary antibody conjugated with horseradish peroxidases (1:1000 dilution, Proteintech SA00001-1; 1:1000 dilution, Proteintech SA00001-2). They were stained with DAB reagent (Gene Tech, GK347011). H&E and immunohistochemistry staining were observed using a light microscope(Mshot, MSX2).
Nuclear and cytoplasmic RNA fractionation, RNA isolation and qPCR
Cell pellets were suspended by gentle pipetting in 200 μL of lysis buffer (10 mM Tris at pH 8.0, 140 mM NaCl, 1.5 mM MgCl2, 0.5% Igepal, 2 mM vanadyl ribonucleoside complex [VRC]) and incubated for 5 min on ice. One-fifth of the lysate was saved as total RNA. The rest of the lysate was centrifuged at 1000g for 3 min at 4°C to pellet the nuclei, and the supernatant was the cytoplasmic fraction. To obtain pure cytoplasmic RNA, the supernatant fraction was further centrifuged at 13,000 rpm for 10 min at 4°C and then collected carefully to a new tube, and RNA was extracted with RNAiso Plus (Takara, 9109). To obtain pure nuclear RNA, the nuclear pellets were subjected to two additional washes with 160 μL of lysis buffer and one additional wash by adding 0.5% deoxycholic acid into the lysis buffer. Finally, the purified nuclei were resuspended in 100 μL of lysis buffer followed by extraction with RNAiso Plus. The RNAiso Plus was used to isolate total RNA. Then the first-strand cDNA was synthesized using the PrimeScript RT Reagent Kit with gDNA Eraser (Takara, RR047A). Real-time PCR was performed using the MonAmp SYBR Green qPCR Mix (MQ10201S, Low ROX). Real-time PCR was measured using QuantStudio3 Real-Time PCR Instrument (Applied Biosystems, USA). The relative levels were calculated using the comparative-Ct method (2-▵▵Ct method). The primer sequences are listed in Table S1.
Western blotting
Mouse livers and hepatocytes were extracted with protein lysis buffer (Beyotime, P0013) supplemented with protease inhibitor cocktail. BCA Kit (Beyotime, P0009) was used to assess the protein concentration. Proteins (40 μg) were separated on 8–10% polyacrylamide precast SDS gels followed by blotting on PVDF membranes (Millipore Billerica, MA, USA). The membranes were blocked in 1×PBS 1% Casein Blocker (BioRad) diluted 1:10 for 1 h at room temperature and then incubated with antibodies against NONO (1:1000 dilution, Abcam, ab70335), TET2 (1:1000 dilution, Active motif, 61389), SREBP1(1:1000 dilution, Bioss, bs-1402R), CD36 (1:1000 dilution, Bioss, ET1701-24) and GAPDH (1:3000 dilution, Proteintech, 60004). HRP conjugate anti-Mouse IgG (1:10000 dilution, Proteintech, SA00001-1) and HRP conjugate anti-Rabbit IgG (1:10000 dilution, Proteintech, SA00001-2) were used as secondary antibody. Finally, the membranes were visualized with enhanced chemiluminescence (Bio-Rad, USA).
Immunoprecipitation
Livers were harvested and suspended in immunoprecipitation buffer (50 mM HEPES at pH 7.6, 250 mM NaCl, 5 mM EDTA at pH 8.0, 0.1% NP-40, 1 mM PMSF, protease inhibitor cocktail) followed by sonication. After centrifuging at 13,000 rpm for 15 min at 4°C, the supernatant was transferred into a new tube and precleared with 10 μL of protein A/G magnetic beads (MCE, HY-K0202). Next, the precleared supernatant was incubated with 20 μL of beads G with antibodies for NONO (rabbit polyclonal, Abcam) or IgG (Millipore, CS200621) for 4 h at 4°C followed by washing with immunoprecipitation buffer. To harvest the protein complex, 50 μL of 1×SDS loading buffer (62.4 mM Tris at pH 6.8, 10% glycerol, 2% SDS, 0.0012% bromophenol blue) was added, incubated for 10 min at 95°C, and analyzed by Western blotting.
RNA ISH and immunofluorescence microscopy
Fluorescence ISH of Srebp1 was carried out using the RNAscope ISH Assay (ACD,323100), following the manufacturer’s instructions. Concisely, prepared cryosections were fixed in formaldehyde for 15 min at 4°C, dehydrated, and pre-treated in hydrogen peroxide for 10 min, followed by 0.5 h digestion in protease III. Prepared formalin-fixed, paraffin-embedded liver sections were baked, deparaffinized and then pre-treated in hydrogen peroxide, followed by target retrieval and protease digestion. Subsequently, slices were pre-amplified and amplified in terms of the directions. The resultant sections were counterstained using mounting medium with DAPI. For colocalization studies, Liver sections were co-stained with rabbit anti-NONO and/or mouse anti-TET2. The nuclei were counterstained with DAPI. Alexa Fluor 488 conjugate anti-Mouse IgG (Invitrogen, A-21422) and Alexa Fluor 555 conjugate anti-Rabbit IgG (Invitrogen, A-21428) were used as secondary antibody. Images were taken with a Zeiss LSM 510 microscope or an Nicon IX70 Delta Vision RT deconvolution system microscope. The probe information for RNAscope assay is listed in Table S2.
Oil Red O staining
Briefly, mice were sacrificed and the liver was perfused with ice-cold 1×PBS to remove the excess of blood. Pieces from the main liver lobes were embedded in optimal cutting temperature (O.C.T.) compound (Scigen, 4583) and frozen at −80°C. For cryosection, the embedded liver was equilibrated at −20°C in a cryos tat chamber and 14 μm thick sections were cut. Unfixed tissue slides were stained with Oil red O and images were taken with a Zeiss VivaTome microscope at a 20× magnification.
Liver triglycerides quantification
Livers were collected from ad libitum fed mice. The livers were perfused with 2 mL of ice-cold PBS through the spleen and immediately collected. For triglycerides quantification, Triglyceride Quantification kit (Nanjing jiancheng, A110-1-1) was used, following manufacturer’s instructions. Briefly, 100 mg of liver powder were homogenized in 5% NP-40 using a Dounce homogenizer. Samples were then heated to 80°C–100°C for 2–5 min and cooled down to room temperature. The heating step was repeated one more time and samples were centrifuged at top speed for 2 min. Supernatants were diluted 1:10 before proceeding with the assay.
RNA-immunoprecipitation (RIP)
The livers were washed with 2 mL of ice-cold PBS and immediately collected. RNA immunoprecipitations (RIPs) were performed using Magna RIP RNA-binding protein immunoprecipitation kits (Millipore, 17–700) according to the manufacturer’s recommendations. Lysate was centrifuged and protein concentration in the supernatant was measured with Pierce BCA assay. 0.5 mg of protein were incubated with 40 μl of protein A agarose beads pre-coated with 10 μg of polyclonal mouse anti-TET2 antibody or normal mouse IgG for 2 h at 4°C. Before the incubation, 1/10 of the supernatant was put aside to be used as input. Immunoprecipitated RNA and input RNA were extracted using Trizol reagent (Takara, 9109).
Bisulfite-PCR sequencing
RNAs isolated from liver were bisulfite-converted using an EZ RNA Methylation Kit, with modifications described in bisulfite sequencing (Zymo, R5001) according to the manufacturer’s instructions. The RNAs were treated as in the RIP assay. The treated RNA was reverse transcribed using random hexamers and PrimeScript RT Reagent Kit with gDNA Eraser (Takara, RR047A). PCR was used to amplify the target regions. The amplicons were subcloned, and ten clones were selected and sequenced to calculate the methylation rate of each cytosine.
Quantification and statistical analysis
Data are expressed as the means ± SD and were analyzed using Prism 8 (GraphPad). Statistical details of the experiments can be found in the results and figure legends. One-way ANOVA or Student’s t-tests was used to determine statistical significance differences between groups. Statistical significance was presented at the level of ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001.
Acknowledgments
We would like to appreciate our workmate, Mi Chen, who gave us their help on discussion. The project is supported by National Natural Science Foundation of China 32071143, National Key R&D Plan No. 2017YFA0103200 and 2017YFA0103202.
Author contributions
Conceptualization, Q.L. and L.Z.; Methodology, Q.L.; Investigation, Q.L., Y.P., J.Z., B.H., D.Q., S.L., and N.C. Data curation, Q.L. and L.Z.; Formal analysis, Q.L.; Validation, L.Z.; Visualization, Q.L. and L.Z.; Resources, L.Z.; Writing–Original Draft, Q.L.; Writing–Review and Editing, L.Z.; Supervision, L.Z.; Funding acquisition, L.Z.; Project administration, L.Z..
Declaration of interests
The authors declare that they have no conflict of interest.
Published: February 20, 2024
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2024.109278.
Supplemental information
References
- 1.Nomura T., Ono M., Kobayashi K., Akaiwa Y., Ayaki M., Ogi T., Ogi M., Takahashi H., Ishikawa K., Morishita A., et al. Validation of fatty liver index as a predictor of hepatic steatosis in Asian populations: Impact of alcohol consumption and sex. Hepatol. Res. 2023;53:968–977. doi: 10.1111/hepr.13935. [DOI] [PubMed] [Google Scholar]
- 2.Devarbhavi H., Asrani S.K., Arab J.P., Nartey Y.A., Pose E., Kamath P.S. Global burden of liver disease: 2023 update. J. Hepatol. 2023;79:516–537. doi: 10.1016/j.jhep.2023.03.017. [DOI] [PubMed] [Google Scholar]
- 3.European Association for the Study of Liver EASL clinical practical guidelines: management of alcoholic liver disease. J. Hepatol. 2012;57:399–420. doi: 10.1016/j.jhep.2012.04.004. [DOI] [PubMed] [Google Scholar]
- 4.Johnston M.P., Patel J., Byrne C.D. Causes of Mortality in Non-Alcoholic Fatty Liver Disease (NAFLD) and Alcohol Related Fatty Liver Disease (AFLD) Curr. Pharmaceut. Des. 2020;26:1079–1092. doi: 10.2174/1381612826666200128094231. [DOI] [PubMed] [Google Scholar]
- 5.Zhang P., Wang W., Mao M., Gao R., Shi W., Li D., Calderone R., Sui B., Tian X., Meng X. Similarities and Differences: A Comparative Review of the Molecular Mechanisms and Effectors of NAFLD and AFLD. Front. Physiol. 2021;12 doi: 10.3389/fphys.2021.710285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Singh A., Amin H., Garg R., Gupta M., Lopez R., Alkhouri N., MCCullough A. Increased Prevalence of Obesity and Metabolic Syndrome in Patients with Alcoholic Fatty Liver Disease. Dig. Dis. Sci. 2020;65:3341–3349. doi: 10.1007/s10620-020-06056-1. [DOI] [PubMed] [Google Scholar]
- 7.Traversy G., Chaput J.P. Alcohol Consumption and Obesity: An Update. Curr. Obes. Rep. 2015;4:122–130. doi: 10.1007/s13679-014-0129-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Parker R., Kim S.J., Im G.Y., Nahas J., Dhesi B., Vergis N., Sinha A., Ghezzi A., Rink M.R., McCune A., et al. Obesity in acute alcoholic hepatitis increases morbidity and mortality. EBioMedicine. 2019;45:511–518. doi: 10.1016/j.ebiom.2019.03.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zou Z.Y., Wong V.W.S., Fan J.G. Epidemiology of nonalcoholic fatty liver disease in non-obese populations: Meta-analytic assessment of its prevalence, genetic, metabolic, and histological profiles. J. Dig. Dis. 2020;21:372–384. doi: 10.1111/1751-2980.12871. [DOI] [PubMed] [Google Scholar]
- 10.Zhang R.N., Pan Q., Zheng R.D., Mi Y.Q., Shen F., Zhou D., Chen G.Y., Zhu C.Y., Fan J.G. Genome-wide analysis of DNA methylation in human peripheral leukocytes identifies potential biomarkers of nonalcoholic fatty liver disease. Int. J. Mol. Med. 2018;42:443–452. doi: 10.3892/ijmm.2018.3583. [DOI] [PubMed] [Google Scholar]
- 11.Pirola C.J., Gianotti T.F., Burgueño A.L., Rey-Funes M., Loidl C.F., Mallardi P., Martino J.S., Castaño G.O., Sookoian S. Epigenetic modification of liver mitochondrial DNA is associated with histological severity of nonalcoholic fatty liver disease. Gut. 2013;62:1356–1363. doi: 10.1136/gutjnl-2012-302962. [DOI] [PubMed] [Google Scholar]
- 12.Pirola C.J., Scian R., Gianotti T.F., Dopazo H., Rohr C., Martino J.S., Castaño G.O., Sookoian S. Epigenetic Modifications in the Biology of Nonalcoholic Fatty Liver Disease: The Role of DNA Hydroxymethylation and TET Proteins. Medicine (Baltim.) 2015;94 doi: 10.1097/MD.0000000000001480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Nestor C.E., Ottaviano R., Reddington J., Sproul D., Reinhardt D., Dunican D., Katz E., Dixon J.M., Harrison D.J., Meehan R.R. Tissue type is a major modifier of the 5-hydroxymethylcytosine content of human genes. Genome Res. 2012;22:467–477. doi: 10.1101/gr.126417.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Thomson J.P., Hunter J.M., Lempiäinen H., Müller A., Terranova R., Moggs J.G., Meehan R.R. Dynamic changes in 5-hydroxymethylation signatures underpin early and late events in drug exposed liver. Nucleic Acids Res. 2013;41:5639–5654. doi: 10.1093/nar/gkt232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.He Y.F., Li B.Z., Li Z., Liu P., Wang Y., Tang Q., Ding J., Jia Y., Chen Z., Li L., et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science. 2011;333:1303–1307. doi: 10.1126/science.1210944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hu L., Li Z., Cheng J., Rao Q., Gong W., Liu M., Shi Y.G., Zhu J., Wang P., Xu Y. Crystal structure of TET2-DNA complex: insight into TET-mediated 5mC oxidation. Cell. 2013;155:1545–1555. doi: 10.1016/j.cell.2013.11.020. [DOI] [PubMed] [Google Scholar]
- 17.Lee J., Song J.H., Park J.H., Chung M.Y., Lee S.H., Jeon S.B., Park S.H., Hwang J.T., Choi H.K. Dnmt1/Tet2-mediated changes in Cmip methylation regulate the development of nonalcoholic fatty liver disease by controlling the Gbp2-Ppargamma-CD36 axis. Exp. Mol. Med. 2023;55:143–157. doi: 10.1038/s12276-022-00919-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shalgi R., Hurt J.A., Lindquist S., Burge C.B. Widespread inhibition of posttranscriptional splicing shapes the cellular transcriptome following heat shock. Cell Rep. 2014;7:1362–1370. doi: 10.1016/j.celrep.2014.04.044. [DOI] [PubMed] [Google Scholar]
- 19.Godet A.C., Roussel E., David F., Hantelys F., Morfoisse F., Alves J., Pujol F., Ader I., Bertrand E., Burlet-Schiltz O., et al. Long non-coding RNA Neat1 and paraspeckle components are translational regulators in hypoxia. Elife. 2022;11 doi: 10.7554/eLife.69162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Yap K., Lim Z.Q., Khandelia P., Friedman B., Makeyev E.V. Coordinated regulation of neuronal mRNA steady-state levels through developmentally controlled intron retention. Genes Dev. 2012;26:1209–1223. doi: 10.1101/gad.188037.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Fox A.H., Lam Y.W., Leung A.K.L., Lyon C.E., Andersen J., Mann M., Lamond A.I. Paraspeckles: a novel nuclear domain. Curr. Biol. 2002;12:13–25. doi: 10.1016/s0960-9822(01)00632-7. [DOI] [PubMed] [Google Scholar]
- 22.Simko E.A.J., Liu H., Zhang T., Velasquez A., Teli S., Haeusler A.R., Wang J. G-quadruplexes offer a conserved structural motif for NONO recruitment to NEAT1 architectural lncRNA. Nucleic Acids Res. 2020;48:7421–7438. doi: 10.1093/nar/gkaa475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Prasanth K.V., Prasanth S.G., Xuan Z., Hearn S., Freier S.M., Bennett C.F., Zhang M.Q., Spector D.L. Regulating gene expression through RNA nuclear retention. Cell. 2005;123:249–263. doi: 10.1016/j.cell.2005.08.033. [DOI] [PubMed] [Google Scholar]
- 24.Edens B.M., Vissers C., Su J., Arumugam S., Xu Z., Shi H., Miller N., Rojas Ringeling F., Ming G.L., He C., et al. FMRP Modulates Neural Differentiation through m(6)A-Dependent mRNA Nuclear Export. Cell Rep. 2019;28:845–854.e5. doi: 10.1016/j.celrep.2019.06.072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Guallar D., Bi X., Pardavila J.A., Huang X., Saenz C., Shi X., Zhou H., Faiola F., Ding J., Haruehanroengra P., et al. RNA-dependent chromatin targeting of TET2 for endogenous retrovirus control in pluripotent stem cells. Nat. Genet. 2018;50:443–451. doi: 10.1038/s41588-018-0060-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhou L., Ren M., Zeng T., Wang W., Wang X., Hu M., Su S., Sun K., Wang C., Liu J., et al. TET2-interacting long noncoding RNA promotes active DNA demethylation of the MMP-9 promoter in diabetic wound healing. Cell Death Dis. 2019;10:813. doi: 10.1038/s41419-019-2047-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Leddin E.M., Cisneros G.A. Comparison of DNA and RNA substrate effects on TET2 structure. Adv. Protein Chem. Struct. Biol. 2019;117:91–112. doi: 10.1016/bs.apcsb.2019.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chen D., Yan Y., Wang X., Li S., Liu Y., Yu D., He Y., Deng R., Liu Y., Xu M., et al. Chronic alcohol exposure promotes HCC stemness and metastasis through beta-catenin/miR-22-3p/TET2 axis. Aging (Albany NY) 2021;13:14433–14455. doi: 10.18632/aging.203059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Niemelä O., Alatalo P. Biomarkers of alcohol consumption and related liver disease. Scand. J. Clin. Lab. Invest. 2010;70:305–312. doi: 10.3109/00365513.2010.486442. [DOI] [PubMed] [Google Scholar]
- 30.Wang X., Yu H., Xing R., Li P. Hepatoprotective Effect of Oyster Peptide on Alcohol-Induced Liver Disease in Mice. Int. J. Mol. Sci. 2022;23 doi: 10.3390/ijms23158081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ghadieh H.E., Russo L., Muturi H.T., Ghanem S.S., Manaserh I.H., Noh H.L., Suk S., Kim J.K., Hill J.W., Najjar S.M. Hyperinsulinemia drives hepatic insulin resistance in male mice with liver-specific Ceacam1 deletion independently of lipolysis. Metabolism. 2019;93:33–43. doi: 10.1016/j.metabol.2019.01.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Allende D.S., Gawrieh S., Cummings O.W., Belt P., Wilson L., Van Natta M., Behling C.A., Carpenter D., Gill R.M., Kleiner D.E., et al. Glycogenosis is common in nonalcoholic fatty liver disease and is independently associated with ballooning, but lower steatosis and lower fibrosis. Liver Int. 2021;41:996–1011. doi: 10.1111/liv.14773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ji C., Nagaoka K., Zou J., Casulli S., Lu S., Cao K.Y., Zhang H., Iwagami Y., Carlson R.I., Brooks K., et al. Chronic ethanol-mediated hepatocyte apoptosis links to decreased TET1 and 5-hydroxymethylcytosine formation. Faseb. J. 2019;33:1824–1835. doi: 10.1096/fj.201800736R. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Purohit V., Gao B., Song B.J. Molecular mechanisms of alcoholic fatty liver. Alcohol Clin. Exp. Res. 2009;33:191–205. doi: 10.1111/j.1530-0277.2008.00827.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.You M., Crabb D.W. Molecular mechanisms of alcoholic fatty liver: role of sterol regulatory element-binding proteins. Alcohol. 2004;34:39–43. doi: 10.1016/j.alcohol.2004.07.004. [DOI] [PubMed] [Google Scholar]
- 36.Xu H.F., Luo J., Zhao W.S., Yang Y.C., Tian H.B., Shi H.B., Bionaz M. Overexpression of SREBP1 (sterol regulatory element binding protein 1) promotes de novo fatty acid synthesis and triacylglycerol accumulation in goat mammary epithelial cells. J. Dairy Sci. 2016;99:783–795. doi: 10.3168/jds.2015-9736. [DOI] [PubMed] [Google Scholar]
- 37.Hames K.C., Vella A., Kemp B.J., Jensen M.D. Free fatty acid uptake in humans with CD36 deficiency. Diabetes. 2014;63:3606–3614. doi: 10.2337/db14-0369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Conne B., Stutz A., Vassalli J.D. The 3' untranslated region of messenger RNA: A molecular 'hotspot' for pathology? Nat. Med. 2000;6:637–641. doi: 10.1038/76211. [DOI] [PubMed] [Google Scholar]
- 39.Dangelmaier E.A., Li X.L., Hartford C.C.R., King J.C., Zibitt M.S., Chari R., Grammatikakis I., Lal A. An Evolutionarily Conserved AU-Rich Element in the 3' Untranslated Region of a Transcript Misannotated as a Long Noncoding RNA Regulates RNA Stability. Mol. Cell Biol. 2022;42 doi: 10.1128/mcb.00505-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yan X., Hoek T.A., Vale R.D., Tanenbaum M.E. Dynamics of Translation of Single mRNA Molecules In Vivo. Cell. 2016;165:976–989. doi: 10.1016/j.cell.2016.04.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Wegener M., Müller-McNicoll M. Nuclear retention of mRNAs - quality control, gene regulation and human disease. Semin. Cell Dev. Biol. 2018;79:131–142. doi: 10.1016/j.semcdb.2017.11.001. [DOI] [PubMed] [Google Scholar]
- 42.Zhou T., Yang M., Zhang G., Kang L., Yang L., Guan H. Long non-coding RNA nuclear paraspeckle assembly transcript 1 protects human lens epithelial cells against H(2)O(2) stimuli through the nuclear factor kappa b/p65 and p38/mitogen-activated protein kinase axis. Ann. Transl. Med. 2020;8:1653. doi: 10.21037/atm-20-7365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Fouda S., Pappachan J.M. Metabolic-Associated Fatty Liver Disease: A Disastrous Human Health Challenge. Endocrinol Metab. Clin. N. Am. 2023;52 doi: 10.1016/j.ecl.2023.03.001. xv–xvi. [DOI] [PubMed] [Google Scholar]
- 44.Wang Y., Wang Y., Patel H., Chen J., Wang J., Chen Z.S., Wang H. Epigenetic modification of m(6)A regulator proteins in cancer. Mol. Cancer. 2023;22:102. doi: 10.1186/s12943-023-01810-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Lian Y., Meng L., Ding P., Sang M. Epigenetic regulation of MAGE family in human cancer progression-DNA methylation, histone modification, and non-coding RNAs. Clin. Epigenet. 2018;10:115. doi: 10.1186/s13148-018-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Ma X., Yang B., Li X., Miao Z. Tet Enzymes-Mediated DNA 5hmC Modification in Cerebral Ischemic and Hemorrhagic Injury. Neurotox. Res. 2022;40:884–891. doi: 10.1007/s12640-022-00505-7. [DOI] [PubMed] [Google Scholar]
- 47.Ito S., D'Alessio A.C., Taranova O.V., Hong K., Sowers L.C., Zhang Y. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature. 2010;466:1129–1133. doi: 10.1038/nature09303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Choudhary N.S., Duseja A. Genetic and epigenetic disease modifiers: non-alcoholic fatty liver disease (NAFLD) and alcoholic liver disease (ALD) Transl. Gastroenterol. Hepatol. 2021;6:2. doi: 10.21037/tgh.2019.09.06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Reaven G.M. Compensatory hyperinsulinemia and the development of an atherogenic lipoprotein profile: the price paid to maintain glucose homeostasis in insulin-resistant individuals. Endocrinol Metab. Clin. N. Am. 2005;34:49–62. doi: 10.1016/j.ecl.2004.12.001. [DOI] [PubMed] [Google Scholar]
- 50.Donohue T.M., Jr. Alcohol-induced steatosis in liver cells. World J. Gastroenterol. 2007;13:4974–4978. doi: 10.3748/wjg.v13.i37.4974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Delatte B., Wang F., Ngoc L.V., Collignon E., Bonvin E., Deplus R., Calonne E., Hassabi B., Putmans P., Awe S., et al. RNA biochemistry. Transcriptome-wide distribution and function of RNA hydroxymethylcytosine. Science. 2016;351:282–285. doi: 10.1126/science.aac5253. [DOI] [PubMed] [Google Scholar]
- 52.Yang X., Yang Y., Sun B.F., Chen Y.S., Xu J.W., Lai W.Y., Li A., Wang X., Bhattarai D.P., Xiao W., et al. 5-methylcytosine promotes mRNA export - NSUN2 as the methyltransferase and ALYREF as an m(5)C reader. Cell Res. 2017;27:606–625. doi: 10.1038/cr.2017.55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Chen X., Li A., Sun B.F., Yang Y., Han Y.N., Yuan X., Chen R.X., Wei W.S., Liu Y., Gao C.C., et al. 5-methylcytosine promotes pathogenesis of bladder cancer through stabilizing mRNAs. Nat. Cell Biol. 2019;21:978–990. doi: 10.1038/s41556-019-0361-y. [DOI] [PubMed] [Google Scholar]
- 54.Janin M., Ortiz-Barahona V., de Moura M.C., Martínez-Cardús A., Llinàs-Arias P., Soler M., Nachmani D., Pelletier J., Schumann U., Calleja-Cervantes M.E., et al. Epigenetic loss of RNA-methyltransferase NSUN5 in glioma targets ribosomes to drive a stress adaptive translational program. Acta Neuropathol. 2019;138:1053–1074. doi: 10.1007/s00401-019-02062-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Chen Y.S., Yang W.L., Zhao Y.L., Yang Y.G. Dynamic transcriptomic m(5) C and its regulatory role in RNA processing. Wiley Interdiscip Rev RNA. 2021;12:e1639. doi: 10.1002/wrna.1639. [DOI] [PubMed] [Google Scholar]
- 56.Mayr C. What Are 3' UTRs Doing? Cold Spring Harbor Perspect. Biol. 2019;11 doi: 10.1101/cshperspect.a034728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Shen Q., Zhang Q., Shi Y., Shi Q., Jiang Y., Gu Y., Li Z., Li X., Zhao K., Wang C., et al. Tet2 promotes pathogen infection-induced myelopoiesis through mRNA oxidation. Nature. 2018;554:123–127. doi: 10.1038/nature25434. [DOI] [PubMed] [Google Scholar]
- 58.Pisani G., Baron B. Nuclear paraspeckles function in mediating gene regulatory and apoptotic pathways. Noncoding. RNA Res. 2019;4:128–134. doi: 10.1016/j.ncrna.2019.11.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Torres M., Becquet D., Blanchard M.P., Guillen S., Boyer B., Moreno M., Franc J.L., François-Bellan A.M. Circadian RNA expression elicited by 3'-UTR IRAlu-paraspeckle associated elements. Elife. 2016;5 doi: 10.7554/eLife.14837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Hocine S., Raymond P., Zenklusen D., Chao J.A., Singer R.H. Single-molecule analysis of gene expression using two-color RNA labeling in live yeast. Nat. Methods. 2013;10:119–121. doi: 10.1038/nmeth.2305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Ruijtenberg S., Hoek T.A., Yan X., Tanenbaum M.E. Imaging Translation Dynamics of Single mRNA Molecules in Live Cells. Methods Mol. Biol. 2018;1649:385–404. doi: 10.1007/978-1-4939-7213-5_26. [DOI] [PubMed] [Google Scholar]
- 62.Halstead J.M., Lionnet T., Wilbertz J.H., Wippich F., Ephrussi A., Singer R.H., Chao J.A. Translation. An RNA biosensor for imaging the first round of translation from single cells to living animals. Science. 2015;347:1367–1671. doi: 10.1126/science.aaa3380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Nagano T., Itoh N., Ebisutani C., Takatani T., Miyoshi T., Nakanishi T., Tanaka K. The transport mechanism of metallothionein is different from that of classical NLS-bearing protein. J. Cell. Physiol. 2000;185:440–446. doi: 10.1002/1097-4652(200012)185:3<440::AID-JCP15>3.0.CO;2-N. [DOI] [PubMed] [Google Scholar]
- 64.Han J., Li E., Chen L., Zhang Y., Wei F., Liu J., Deng H., Wang Y. The CREB coactivator CRTC2 controls hepatic lipid metabolism by regulating SREBP1. Nature. 2015;524:243–246. doi: 10.1038/nature14557. [DOI] [PubMed] [Google Scholar]
- 65.Buzzetti E., Pinzani M., Tsochatzis E.A. The multiple-hit pathogenesis of non-alcoholic fatty liver disease (NAFLD) Metabolism. 2016;65:1038–1048. doi: 10.1016/j.metabol.2015.12.012. [DOI] [PubMed] [Google Scholar]
- 66.Wang J., Rajbhandari P., Damianov A., Han A., Sallam T., Waki H., Villanueva C.J., Lee S.D., Nielsen R., Mandrup S., et al. RNA-binding protein PSPC1 promotes the differentiation-dependent nuclear export of adipocyte RNAs. J. Clin. Invest. 2017;127:987–1004. doi: 10.1172/JCI89484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Hu S.B., Xiang J.F., Li X., Xu Y., Xue W., Huang M., Wong C.C., Sagum C.A., Bedford M.T., Yang L., et al. Protein arginine methyltransferase CARM1 attenuates the paraspeckle-mediated nuclear retention of mRNAs containing IRAlus. Genes Dev. 2015;29:630–645. doi: 10.1101/gad.257048.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Bertola A., Mathews S., Ki S.H., Wang H., Gao B. Mouse model of chronic and binge ethanol feeding (the NIAAA model) Nat. Protoc. 2013;8:627–637. doi: 10.1038/nprot.2013.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Guo F., Zheng K., Benedé-Ubieto R., Cubero F.J., Nevzorova Y.A. The Lieber-DeCarli Diet-A Flagship Model for Experimental Alcoholic Liver Disease. Alcohol Clin. Exp. Res. 2018;42:1828–1840. doi: 10.1111/acer.13840. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data: All data reported in this paper will be shared by the lead contact upon request.
Code: This paper does not report original code.
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.