Abstract
Fuzzy rough entropy established in the notion of fuzzy rough set theory, which has been effectively and efficiently applied for feature selection to handle the uncertainty in real-valued datasets. Further, Fuzzy rough mutual information has been presented by integrating information entropy with fuzzy rough set to measure the importance of features. However, none of the methods till date can handle noise, uncertainty and vagueness simultaneously due to both judgement and identification, which lead to degrade the overall performances of the learning algorithms with the increment in the number of mixed valued conditional features. In the current study, these issues are tackled by presenting a novel intuitionistic fuzzy (IF) assisted mutual information concept along with IF granular structure. Initially, a hybrid IF similarity relation is introduced. Based on this relation, an IF granular structure is introduced. Then, IF rough conditional and joint entropies are established. Further, mutual information based on these concepts are discussed. Next, mathematical theorems are proved to demonstrate the validity of the given notions. Thereafter, significance of the features subset is computed by using this mutual information, and corresponding feature selection is suggested to delete the irrelevant and redundant features. The current approach effectively handles noise and subsequent uncertainty in both nominal and mixed data (including both nominal and category variables). Moreover, comprehensive experimental performances are evaluated on real-valued benchmark datasets to demonstrate the practical validation and effectiveness of the addressed technique. Finally, an application of the proposed method is exhibited to improve the prediction of phospholipidosis positive molecules. RF(h2o) produces the most effective results till date based on our proposed methodology with sensitivity, accuracy, specificity, MCC, and AUC of 86.7%, 90.1%, 93.0% , 0.808, and 0.922 respectively.
Keywords: Rough set, Granular structure, Intuitionisitic fuzzy relation, Intuitionistic Fuzzy Set, Mutual information
Subject terms: Computational biology and bioinformatics, Health care, Molecular medicine
Introduction
The current trend of accumulation of huge amount of data in different databases pertaining to different domains has given rise to the unique opportunity of knowledge discovery/extraction using a plethora of data mining techniques1. These techniques2 can be explored in three ways namely knowledge types, architecture types, and analysis types along with their powerful applications in distinct research and practical domains to solve the interesting real-world problems. Data Mining plays a vital role in establishing smart agriculture application tools to accomplish real-time data analysis with large volume of data. Data mining tasks3 offer essential hidden patterns, correlation, and knowledge from the various applications of bioinformatics datasets, viscous dissipation, and activation energy4,5. Machine learning methods provide a set of techniques that can be used to create prediction/discriminatory models and subsequent knowledge extraction, which may facilitate in decision making or for better understanding of the concerned domain6,7. The “curse of dimensionality” plagues the effectiveness of various machine learning algorithms, but the development of dimensionality reduction methods8 have considerably impacted in reducing the effects of redundancy present in high dimensional datasets. In the fields of data mining, signal processing, biomedical imaging, agriculture, industrial engineering, and bioinformatics, researchers frequently face obstacles due to “curse of dimensionality” as it leads to enlarge the cost of data storage and extensive computing9. Moreover, this issue directly affects both the efficiency and accuracy to cope with different problems10. Dimensionality reduction process can easily eliminate redundancy and/or irrelevancy, noise, minimize the complexity of machine learning methods, and enhance the overall accuracy of classification process, and can be identified as an essential and key phase in pattern recognition scheme11.
Redundant features affects negatively to the various machine learning algorithms mostly resulting in high computation time and less accurate predictive models12. It also complicates the model interpretation. Feature reduction methods can be used to mitigate the negative effects of high dimensional data by facilitating the selection of low dimensional non-redundant subset of features. Feature reduction methods have been found to be very effective in a wide variety of research areas, including biological domain13,14.
Most popular methods of feature reduction algorithms fall under filter and wrapper methods. While wrapper methods are classifier dependent for the evaluation of features15,16, filter methods use classifier independent feature selection criterion and are generally less computationally intensive17.
Previously rough set theory18,19 have been applied very promisingly in feature selection20. Although classical rough set theory based feature selection methods21,22 can only be used on discrete features, which makes it mandatory for discretization of continuous features23,24. There is a fair chance of information loss during the process of discretization25.
The combination of fuzzy26 and rough sets27 effectively deals with uncertainty, vague and incomplete data. Rough set theory has been competently employed to produce the most informative features from a dataset consisted of discretized conditional attribute values. This informative feature subset is produced from the original features set with minimum information loss, and termed as reduct. Rough set deals with vagueness, whilst fuzzy set handles uncertainty. Fuzzy set theory ensures that real-valued datasets can be handled without any further discretization. By combining fuzzy set with rough set, information loss due to discretization can be effectively avoided as fuzzy rough set (FRS) can handle real-valued information system (dataset) directly. FRS can be effectively used for mitigating the effects of information loss as a consequence of discretization of features by using fuzzy similarity measures to tackle the continuous feature values28. Broadly, FRS aided dimensionality reduction29 methods can be categorized into two types30,31 which are based on discernibility matrix and dependency function32. Discernibility matrix assisted approaches provide numerous reduct sets33, whilst dependency function leads to a single feature subset34.
In FRS aided dimensionality reduction theory, a similarity relation is incorporated between the data points to construct lower and upper approximations. By taking union of the computed lower approximations, we obtain the positive region of decision. Here, the wider is the obtained membership to positive region; greater is the plausibility of instance belonging to an individual category35. Based on dependency function, we compute significance of a subset of features. Moreover, the conditional entropy measure is employed in to calculate reduct set for both homogeneous and heterogeneous information system respectively36–38. However, it may lead to misclassification of samples when there is a large degree of imbricate between diverse categories of data. Also, it can cope with only with membership of data point to a set, where uncertainty cannot be handled due to both identification and justification. Hence, there is an essential and utmost requirement of distinct kind of mathematical model that can both fit data, and at the same moment it can tackle uncertainty emerging due to identification39.
Intuitionistic fuzzy (IF) set40,41 is step ahead that offers two degree of freedom by taking into consideration both membership and non-membership, which can cope with uncertainty that emerges both in judgement and identification42. It has been successfully exercised in decision making43, image segmentation, rule generation, and machine learning44,45. In the recent few years, the assemblage of IF46 and rough sets47 are employed to establish numerous IF rough set models48,49 to effectively handle later uncertainty and vagueness in the data50,51. Huang et al.52 proposed a ranking based model for selecting the neighbourhood of objects53,54 and presented a Dominant IF Decision Table (DIFDT)55 by using discernibility matrix and assisted discernibility function23. They developed IFRS based reduction technique for knowledge extraction from given information system. Huang et al.56 presented the IF multigranulation rough set (IFMGRS) model and studied different reduction techniques to eliminate redundant granules by introducing reducts for three different types of IFMGRSs in 2014. Tan et al.57 used the concept of granular structure to introduce an IF rough set model58 and employed it for feature selection. Tiwari et al.59 discussed an IF tolerance relation, which was applied to establish IF rough set aided feature selection. Shreevastava et al.60 addressed different similarity relation assisted technique to deal with both supervised and semi-supervised data. Tiwari et al.61–63 and Shreevastava et al.64,65 elaborated different issues related to feature selection technique and presented several lower and upper approximations by using various mathematical ideas. A feature selection to track multiple samples was presented by Li et al.66 by using IF clustering notion. IF quantifier was introduced by Singh et al.67 to construct IF rough set model and its application to feature reduction. Jain et al.39 tried to minimize noise in the data by using the concept of IF granules and incorporated different types of IF relations to introduce feature selection both robust and non-robust. From the recent published articles, it is conspicuous that the use of IF set theory assisted notion for feature selection is still in its incipient stage. Uncertainty is measured in terms of entropy and has its origin in the telecommunications domain68,69. Mutual information (MI)70 aims to measure the relationship between feature and the target. Further, it can be stated that mutual information (MI)71 is an interesting quantity that evaluates the dependence between conditional features and has been repeatedly employed to solve an extensive diverse problems. Feature selection techniques can be converted into effective one by incorporating information entropy estimation notion for attribute extraction based on MI72 and the conventional feature selection approaches on the basis of class seperability. Broadly MI measures the amount of information that can be deduced from a random variable/vector about another random variable/vector73,74.
Max-relevance-minimum-redundancy method75,76 is based on the concept of MI and has been relevant in a number of previous studies. It deduces the target MI with minimum redundancy10,77 among the selected features. A number of MI based feature selection algorithms have been in practice in various domains72,74. Fuzzy rough entropy was effectively used to avoid the limitation of rough entropy to handle the real-valued feature data78,79, but fuzzy rough entropy leads to lessening monotonically with the rise of the dimensions of data, which can promptly reflect the roughness of information systems. This issue was resolved up to certain extent by presenting the extension of fuzzy rough based information entropy with conditional entropy, joint entropy, and mutual information. However, none of the works has handled the noise, vagueness, and uncertainty due to both identification and judgement simultaneously, which is frequently appearing in the current era of high-dimensional datasets due to advancement of internet based technologies. In the current study, a new IFRS based joint entropy, conditional entropy, and mutual information based on a new IF hybrid relation and IF granular structure to handle the different issues such as later uncertainty, vagueness, and imprecision available in the large volume of high dimensional datasets that may degrade the performances of learning algorithms. Firstly, a novel hybrid IF similarity relation is presented. Secondly, joint and conditional entropies are established in IF rough framework. Thirdly, IF rough mutual information is introduced. Then, lower and upper approximations are computed by using presented hybrid IF similarity relation. Thereafter, dependency function is computed by using the defined lower approximation. Next, significance of feature subset is computed by using IF rough mutual information. Further, a heuristic feature selection algorithm is discussed by using both significance and dependency function. IF rough mutual information are employed to measure the later uncertainty and the correlation between features and class. Next, this algorithm is applied on benchmark datasets, and the reduct is computed. The effectiveness of the proposed algorithm is further explained by measuring the performances of seven widely used learning techniques on reduced data produced by our method and four existing approaches. Finally, the proposed method is applied to enhance the overall prediction to discriminate the phsopholipidosis80 positive (PL+) and phsopholipidosis negative (PL-) molecules. Phospholipidosis is a condition when there is an abnormal buildup of phospholipids in various tissues due to the usage of cationic amphiphilic pharmaceuticals. Phsopholipidosis (PPL) is a reversible condition, and phospholipidosis levels revert to normal once the cationic amphiphilic medications are stopped81. Computational prediction of possible inducing characteristics utilizing structure-activity relationship (SAR) can enhance the traditional high throughput screening and drug development pipelines because to its rapidity and cost-effectiveness82.The main contributions of the entire study can be highlighted as follows:
Major contributions of the study
This study establishes a new hybrid IF similarity relation that can deal with both nominal and numerical features.
An IF granular structure is presented to handle the noise in mixed data.
IF rough entropy, joint entropy, and conditional entropy is given to handle the later uncertainty with information entropy.
Further, the idea of an If rough mutual information is discussed.
Moreover, this If rough mutual information is employed to evaluate both uncertainty and the correlation between conditional feature and decision class.
Then, a feature selection approach is introduced by using this IF rough mutual information concept.
Finally, a framework is designed based on our proposed methods to enhance the prediction of phospholipidosis positive molecules.
Theoretical background
In this segment, few essential basic notions about IF set, IF relation, IF information system, and mutual information is reviewed. These concepts can be explained/described as follows:
Definition 2.1
IF set An IF set X in is well defined collection of samples/objects having the form
| 1 |
where, portrays the set of data points/samples/objects. Moreover, along with , which holds the essential condition . Here, and are depicted as the imperative membership and non-membership grades for a given element . Further, portrays the hesitancy grade of . Additionally, we have , . Thus, the obtained ordered pair is depicted as a requisite IF value.
Definition 2.2
IF information system An IF information system (IFIS) can be exemplified by a quadruple ( ,, where, we have , which is comprised of all IF values. Further, we have a mapping, which can be portrayed by IF : , in such a way that ,, .
Definition 2.3
IF relation Let be an IF binary relation induced on the system. is IF similarity relation if it satisfies :
- Reflexivity: For any given i and j,
2 - Symmetry: For any given i and j,
3
Definition 2.4
Mutual information Mutual information (MI) can be expresserd based on broadely depicted entropy and well-known conditional entropy by using the following given equation
| 4 |
where, , H(D) and H(D|P) depict information entropy and conditional entropy respectively. Decrease of uncertainty about D gernerated by P is evaluated by mutual information and its inverse is computed in the same way. Mutual information is employed to calculate either volume of information of P enclosed in D or D included in P. H(P) is amount of information contained in P about itself which means I(P;P)=H(P)
Definition 2.5
Significance of conditional feature For a given IFIS and , if we have an arbitrary conditional dimension/feature , then its significance can be illustrated by the following equation
| 5 |
and , , which is a MI between conditional dimension/feature b and decision feature D. If the calculated value of SGF(b, B, D) is greater, then it insinuates that under the known condition of feature subset B, dimension b is found to be more potential for the available decision feature D.
Proposed work
In the underway segment, we demonstrate a hybrid IF similarity relation, granular structure, and MI. Based on these concepts, a feature selection procedure is introduced to discard irrelevancy and redundancy available in the high-dimensional information systems.
IF Relation: For all , and , the hybrid similarity between and with respect to any given a can be defined by:
| 6 |
where, is depicted as an adaptive IF radius. The IF relation and IF relation matrix enticed by a are and , where .
If we have , then,
| 7 |
Proof
-
Reflexive: If we take a case when , then, proposed relation follows only two cases, which are first and third. Moreover, other two cases are rejected by default.
Case 1. if where a is a nominal , then we obtain ==1
Case 2. If a is numerical and and ,then
Now,if we put , we get the folllowing results:
, therefore, we get as refelxive
- Symmetry:
8
Now, it can be identified that9
So , is symmetric
Since, is both reflexive and symmetric. Hence, we can obviously conclude that is an IF similarity relation.
Granular structure
The IF granule is elicited by as follows:
| 10 |
,
further,
| 11 |
is subset of C and
By using IF granulation structure, rough entropy can be discussed into IF rough framework, and IF rough entropy of a feature can be described by:
Definition 3.1
The IF rough entropy of can be given as:
| 12 |
It is obvious to identify that iff so . In this suit all the sample pairs are found to be identical. Therefore, the obtained granulation space is found to be the largest at this time, on the contrary which indicates . Therefore, . Now,the granulation space is instituated as the smallest one.
Definition 3.2
The IF joint rough entropy of and can be expressed by :
| 13 |
Definition 3.3
The IF rough conditional entropy of relative to can be addressed by the following equation :
| 14 |
Definition 3.4
The IF rough mutual information of and can be computed as follows;
| 15 |
Definition 3.5
The IF rough mutual information between D and can be illustrated by the equation:
| 16 |
By using this equation, IF rough mutual information considers as the correlation between and decision feature D . If the obtained value of IF rough mutual information between D and is higher, then, we get more correlated value between and D.
Proposition 3.6
If , then
Proof
As discussed by the aforesaid definition 3.1, , and
Now, ;
Proposition 3.7
If , then .
Proof
For a given , we have . Now, we can simply write
Therefore, we detect the result by using the definition 3.1 as .
Proposition 3.8
IF then
Proof
For any given , we have the following expression based on the Proposition 3.6,
. Moreover, by using Proposition 3.7, we can conclude the following result:
Proposition 3.8 depcits that IF rough entropy reduces when feature subset accquire larger size, whilst,it grows in case of features subset procures smaller size . It can be easily observed that IF rough entropy definition can evaluate the uncertainty of IF approximation space.
Proposition 3.9
Suppose , then
Proof
Since and and By Proposition 3.2, we have and
Proposition 3.10
IF , then
Proof
Since , hence, by using the Proposition 3.6, we get
So,
According to the Proposition 3.10, when there are two IF granules produced by two potential feature subsets respectively, then IF joint rough entropy of the calculated two potential feature subsets is equal to the IF rough entropy of the feature subsets corresponding to relatively smaller IF granulation.
Proposition 3.11
.
Proof
Based on the Definition 3.3, we have
Proposition 3.12
If , then
Proof
Since, , hence, based on the Proposition 3.6, we can conclude that. Therefore, , , furthermore, , , now, based on the Definition 3.3, we have
IF rough mutual information can’t only be used to measure the uncertainty of IF approximation space but also can be applied to evaluate the correlation between conditional feature and decision class.
Proposition 3.13
Proof
Based on the Proposition 3.9, we have Similarly, we can get
Proposition 3.14
Proof
Obviously satisfies based on the Definitions 3.1, 3.4, and 3.5. Now, we obtain the following results:
Definition 3.15
For a given IFIS, let P be subset of conditional dimensions/features(C).Thereafter, is found to be the significance as , which can be further given by:
| 17 |
, and can be outlined as, , which depicts the MI of IF conditional feature T and the decision feature D. If the value of increases, then IF conditional dimension/feature T is obtained to be more relevant for a given decision feature D.
Algorithm 1.
Feature selection alogrithm based on IF mutual information (FSIFMI)
Experimentation
In the current experimental section, the performance of the proposed method is evaluated and compared with the existing fuzzy and IF sets assisted techniques. All the pre-processing concepts are implemented in Matlab 202383 and learning algorithms are implemented in WEKA84. Firstly, fuzzification and intuitionistic fuzzification of the real valued data is performed by using the methods proposed by Jensen et al.6 and Tan et al.57 respectively. Secondly, the reduced datasets are obtained by the previously presented approaches. Thirdly, different threshold parameters values are adjusted for our established method to produce the reduct. Then, reduced datasets are generated by discarding the noise to the maximum level. The reduct is computed by changing the value of from 0.1 to 0.8 in small interval, and the value of providing the maximum performance measures in the experiment is selected as the final one. To perform the entire experimental study, the following setup is exercised to conduct the comprehensive experiments:
Dataset
Ten benchmark datasets are taken from widely discussed University of California from Irvine based Machine Learning Repository85 to conduct the entire experiments. The required details of these datasets are outlined in Table 1. The dimension and size of these datasets depict that these are small to large datasets as number of data points range from 62 to 4521 and features range from 9 to 10000.
Table 1.
Dataset characteristics and reduct size.
| Dataset | Instances | Features | Reduct size | ||||
|---|---|---|---|---|---|---|---|
| FSFrMI | GIFRFS | TIFRFS | FRFS | IFRFSMI | |||
| Bank marketing | 4521 | 16 | 10 | 12 | 15 | 15 | 14 |
| Breast cancer | 699 | 9 | 8 | 9 | 9 | 8 | 8 |
| Dbworld-bodies | 64 | 4702 | 97 | 128 | 187 | 88 | 8 |
| Arcene | 200 | 10000 | 453 | 287 | 303 | 268 | 169 |
| Colon | 62 | 32 | 24 | 27 | 21 | 18 | 8 |
| Gsar-biodegradation | 1055 | 41 | 31 | 36 | 29 | 33 | 25 |
| Fertility diagnosis | 100 | 9 | 8 | 6 | 8 | 7 | 7 |
| Thyroid- hypothyroid | 3163 | 25 | 11 | 17 | 19 | 15 | 12 |
| Heart disease | 294 | 13 | 11 | 10 | 10 | 12 | 9 |
| Wdbc | 569 | 21 | 17 | 14 | 18 | 10 | 8 |
Classifiers
Seven different learning methods86 are applied to demonstrate the performance measures on the reduced datasets obtained from different feature selection techniques. RealAdaBoost with random forest as base classifier (RARF) and IBK are employed for the objective of evaluating overall classification accuracies with standard deviation by using diverse validation techniques for ten benchmark reduced datasets. Moreover, we applied naive bayes, SMO, IBK, RARF, PART, JRip,J48, and random forest (RF) to evaluate the performances based on various evaluation metrics for the reduced Nath et al.87 dataset for evaluating the effectiveness of the proposed technique when compared to existing method for discriminating PL+ and PL- molecules.
Dataset split: Feature selection process is carried out over complete information system. After production of reduced datasets, individual learning algorithm is evaluated based on percentage split of 66:34 and kd-fold cross validation. In percentage split technique, dataset is randomly divided into two parts, where training is done on 66% of the entire dataset, while 34% of the dataset is employed to perform testing. In kd-fold cross validation, whole dataset is randomly separated into kd subsets, where kd-1 parts form training set, whilst one is employed to form testing set. After kd such repetitions, average value of different evaluation metrics is considered as final performance. In the current study, the value of kd is taken as 10.
Performance evaluation metrics
The prediction performance measures of the seven learning algorithms from different categories are evaluated using both broadly elaborated threshold-dependent and threshold- independent assessment parameters. These assessment parameters are ascertained based on the calculated values of true positive (TRP), true negative (TRN), false positive (FLP), and false negative (FLN). TRP is computed number of correctly predicted positive data points; TRN is calculated number of correctly predicted negative data points. FLN is representation for the number of incorrectly predicted positive samples, while FLP is depiction for the number of incorrectly predicted negative samples. We employ different parameters namely: Sensitivity (Sn), Specificity (Sp), Accuracy (Ac), AUC, and MCC to measure the overall performances of the individual learning algorithms. Now, these evaluation parameters can be mathematically discussed as follows:
Sn: This calculates the overall percentage of correctly classified PPL+, which is specified by:
| 18 |
Sp: This includes the efficacious percentage of correctly classified PPL−, which is produced by:
| 19 |
Ac: The percentage of required overall correctly classified PPL+ and PPL− , which can be stated as:
| 20 |
AUC: It is applied to observe the important and required area under the receiver operating characteristic curve (ROC), the more tends its count towards 1, the better will be the obtained predictor.
MCC: Mathew’s correlation coefficient is a very much potential and the most awaited parameters, which is computed with the help of following equation:
| 21 |
This parameter is applied not only to clarify the effectiveness of the binary classifications but also to justify its efficiency. An MCC value tends towards 1 to specify that the predictor is the promising one.
Results and discussion
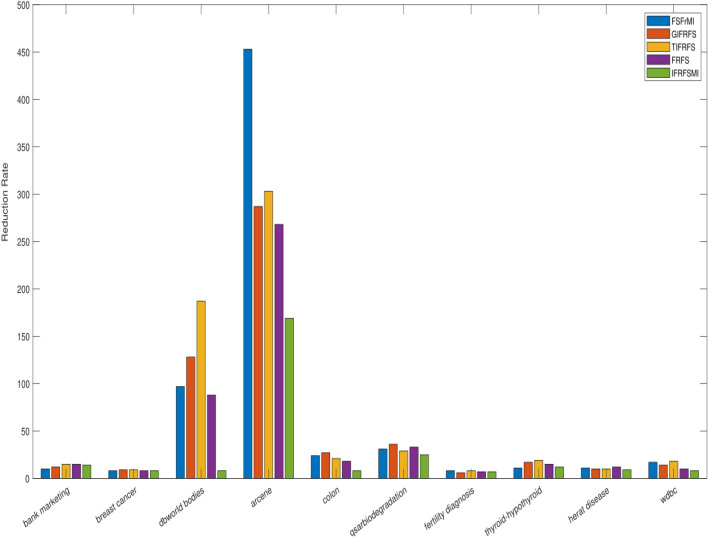
The details of the ten benchmark datasets along with the reduct as produced by four existing as well as presented methods is depicted in Table 1. Real-valued datasets are converted into fuzzy and IF values by using widely discussed Jensen et al.6 and Tan et al.57 concepts. Entire reduction process is accomplished over complete data by using both fuzzy and IF aided techniques. FSFrMI72, GIFRFS57, TIFRFS59, and FRFS6 are the earlier efficacious and effective techniques, which are incorporated to perform the comparative results (Table 2). Our proposed method produced reduct set range from 7 to 169, where reduct size is smaller when compared to reduct size by earlier approaches, except bank marketing and thyroid-hypothyroid datasets. For bank marketing dataset, FSFrM and GIFRFS resulted in relatively less size data, whilst smaller size is produced by FSFrMI and FRFS for thyroid-hypothyroid and fertility diagnosis datasets respectively in contrast with IFRFSMI. Moreover, for breast cancer, FSFrM and FRFS provide the similar size, whilst, for fertility diagnosis dataset FRFS produce similar size of the data when compared to the results presented by proposed method. From the recorded reduct in Table 1, it can be observed that our proposed technique is generating more reduced dimensions for most of the cases related to all the ten datasets rather than recently established powerful methods. We have presented the visualization of reduction process based on different methods in Fig. 1, which clearly indicates that our proposed method produces high percentage of overall feature elimination with the increment of total conditional features. Then, IBK and RARF are chosen to show the learning performances in terms of standard deviation with overall accuracies for the reduced datasets generated by four existing and our proposed techniques, where 10-fold cross validation is employed to avoid the overfitting. These results are reported in Table 2, where the ranks are outlined in the superscript of all the individual results. From the results available in Table 2, it is obvious that our proposed method is dispensing the better results in contrast with the results of other previous approaches regardless of reduced data produced by previous approaches, except the outcome for breast cancer and heart disease datasets. For breast cancer dataset, TIFRFS presents better outcome when compared to IFRFSMI by using both IBK and RARF, while, for heart disease dataset TIFRFS gave the best result with RARF. For colon and heart disease datasets, GIFRFS and TIFRFS leads to identical results as compared to IFRFSMI based results by IBK. Similar results are presented by RARF for fertility diagnosis and wdbc datasets based on the reduced datasets produced by FSFrMI and GIFRFS respectively in contrast with proposed method based reduced datasets. Entire results can be visualized by Figs. 2 and 3. These figures depict that proposed concept are very much effective for both low and high-dimensional datasets as the reduced datasets produced by this method always leads to increment of overall accuracies of the different learning algorithms regardless of their dimensionality size.
Table 2.
Comparison of overall accuracies with standard deviation for the datasets produced by FSFrMI GIFRFS, TIFRFS, FRFS, and IFRFSMI by using 10-fold cross validation.
| Dataset | Classifier | FSFrMI | GIFRFS | TIFRFS | FRFS | IFRFSMI |
|---|---|---|---|---|---|---|
| Bank | IBK | 84.75 | 83.79 | 83.01 | 81.21 | 86.28 |
| Marketing | RARF | 87.23 | 86.18 | 87.59 | 83.18 | 89.37 |
| Breast | IBK | 81.11 | 86.24 | 96.11 | 84.29 | 95.67 |
| Cancer | RARF | 89.34 | 93.34 | 97.12 | 88.66 | 96.04 |
| Dbworld | IBK | 89.16 | 90.86 | 91.89 | 88.89 | 94.74 |
| Bodies | RARF | 90.25 | 92.19 | 93.55 | 90.55 | 97.21 |
| Arcene | IBK | 71.47 | 70.72 | 72.09 | 71.09 | 74.00 |
| RARF | 75.69 | 74.69 | 77.55 | 72.35 | 83.45 | |
| Colon | IBK | 75.88 | 78.12 | 79.06 | 73.06 | 78.12 |
| RARF | 79.21 | 80.41 | 81.17 | 77.17 | 82.81 | |
| Qsarbio-degradation | IBK | 78.27 | 77.69 | 79.51 | 75.87 | 82.09 |
| RARF | 80.28 | 81.33 | 82.06 | 79.16 | 86.74 | |
| Fertility diagnosis | IBK | 83.21 | 81.41 | 83.17 | 80.17 | 84.30 |
| Thyroid- hypothyroid | RARF | 87.20 | 83.69 | 85.23 | 82.87 | 87.20 |
| IBK | 92.33 | 91.23 | 95.16 | 88.33 | 97.87 | |
| RARF | 95.21 | 93.41 | 97.17 | 92.17 | 99.11 | |
| Heart disease | IBK | 79.26 | 78.46 | 81.16 | 76.25 | 81.16 |
| RARF | 81.27 | 80.38 | 83.69 | 78.98 | 82.74 | |
| Wdbc | IBK | 95.68 | 93.46 | 95.16 | 89.33 | 96.06 |
| RARF | 96.41 | 97.73 | 97.69 | 91.26 | 97.73 | |
| Average | IBK | 3.20 | 3.55 | 2.15 | 4.80 | 1.30 |
| Rank | RARF | 3.45 | 3.35 | 2.00 | 4.90 | 1.30 |
| F statistics | IBK | 23.09 | ||||
| RARF | 32.38 |
Figure 1.
Comparison of overall reduction for different daasets by previous and proposed methods.
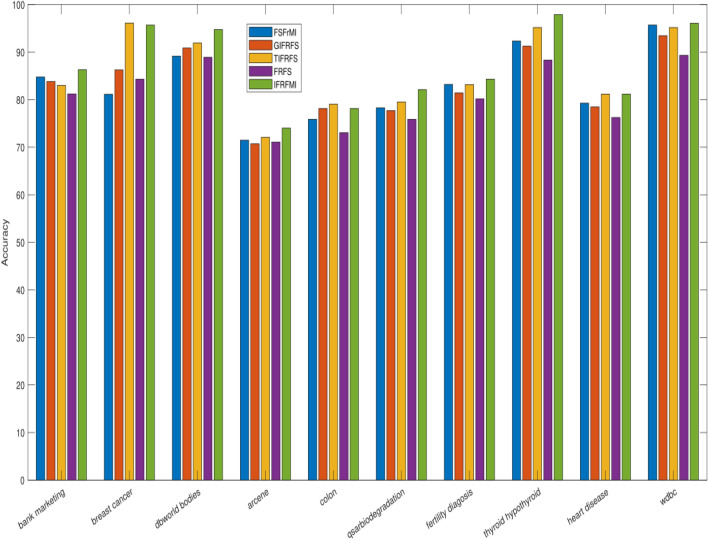
Figure 2.
Comparison of average accuracies by IBK for different reduced datasets as produced by existing and proposed methods.
Figure 3.
Comparison of average accuracies by RARF for different reduced datasets as produced by existing and proposed methods.
Our assumptions to verify the significance of our proposed method are as follows:
Null Hypothesis: All the employed methods are equivalent.
Alternate Hypothesis: There is significant difference among the employed methods.
Two widely accepted testing approaches namely Freidman test88 and Bonferoni Dunn test89 are applied to validate the significance of the presented method. Freidman test is used to perform comparative study of multiple models. Further, Bonferoni Dunn is employed to obtain which method is significantly different from proposed technique. The null hypothesis can be rejected at level of significance if the values between their average ranks is higher rather than critical distance value. In the current study, average ranks by both IBK and RARF based on our proposed method are recorded as the minimum value (Table II). These values are clearly depicting the superiority of our established models. Moreover, F-statistics computed values based on IFRFSMI are obtained larger for both IBK and RARF when compared to F-tabular value. F-statistics computed values for IBK and RARF are 23.09 and 32.38 (Table II), whilst F-tabular value is 2.634 (F(4,36) = 2.634 at 5% level of significance). Therefore, based on Dunn Test our proposed method is found as significantly different.
Case study: an application to discriminate PL+ and PL- molecules
One of the prime applications of machine learning based methods in cheminformatics is the reduction of enormous chemical space with respect to some property of interest. The reduced chemical space can then be validated using wet lab based experiments, thus making the fidelity of machine learning methods of outmost importance.
One of the hallmarks of phospholipidosis is the accumulation of phospholipids in the various types of tissues for eg. kidneys, eyes etc. mostly caused by cationic amphiphilic molecules. Highly accurate machine learning prediction models can facilitate in screening of phospholipidosis inducing compounds in early stages of drug discovery workflows, thereby reducing the cost and time associated with wet lab based experiments (Fig. 4).
Figure 4.

ROC for the RF algorithm on phospholipidosis dataset.
The present methodology can open new possibilities for further research in early screening of phospholipidosis inducing molecules.
Now, our proposed approach is applied to Nath et al.87 dataset to produce the effective reduced form by minimizing noise, uncertainty, imprecision available in the data along with removal of redundant, and irrelevant attributes. Thereafter, seven classifiers from different categories are investigated to evaluate their performances over this reduced dataset based on sensitivity, AUC, Specificity, MCC, and accuracy, which have reported in Tables 3, 4, 5 and 6. Moreover, for original and reduced data, a commodious approach to represent theoverall performance measures of all the seven classifiers at the best decision threshold can be given by Receiver Operating Characteristic (ROC) curve, which furnishes a visual explanation of the classifiers performance. Figures 5 and 6 depict ROC curves for original and reduced dataset based on 10-fold cross validation. These figures indicate that RARF algorithm achieved the best AUC in comparison to all the other algorithms().
Table 3.
Performance evaluation metrics of eight classifiers for original dataset consisting of PL+ and PL- molecules based on 10-fold cross validation.
| Classifiers | Sensitivity | Specificity | Accuracy | AUC | MCC |
|---|---|---|---|---|---|
| Davie Bayes | 75.5 | 81.4 | 78.4 | 0.828 | 0.570 |
| SMO | 81.4 | 85.3 | 83.3 | 0.833 | 0.667 |
| IBK | 82.4 | 80.4 | 81.4 | 0.806 | 0.628 |
| RARF | 81.4 | 85.3 | 83.3 | 0.908 | 0.667 |
| PART | 75.5 | 73.5 | 74.5 | 0.718 | 0.490 |
| JRip | 66.7 | 69.6 | 68.1 | 0.723 | 0.363 |
| RandomForest | 83.3 | 82.4 | 82.8 | 0.893 | 0.657 |
| J48 | 74.5 | 74.5 | 74.5 | 0.769 | 0.510 |
Table 4.
Performance evaluation metrics of eight classifiers for reduced dataset generated by proposed approach consisting of PL+ and PL- molecules based on 10-fold cross validation.
| Classifiers | Sensitivity | Specificity | Accuracy | AUC | MCC |
|---|---|---|---|---|---|
| Navie Bayes | 85.3 | 70.6 | 77.9 | 0.846 | 0.565 |
| SMO | 81.4 | 68.6 | 75.0 | 0.750 | 0.504 |
| IBK | 87.3 | 87.3 | 87.3 | 0.811 | 0.745 |
| RARF | 88.2 | 84.3 | 86.3 | 0.925 | 0.726 |
| PART | 71.6 | 72.5 | 72.1 | 0.778 | 0.441 |
| JRip | 74.5 | 80.4 | 77.5 | 0.811 | 0.550 |
| RandomForest | 84.3 | 84.3 | 84.3 | 0.915 | 0.686 |
| J48 | 74.5 | 75.5 | 75.0 | 0.752 | 0.500 |
Table 5.
Performance evaluation metrics of eight classifiers for original dataset consisting of PL+ and PL- molecules based on percentage split of 66:34.
| Classifiers | Sensitivity | Specificity | Accuracy | AUC | MCC |
|---|---|---|---|---|---|
| Navie Bayes | 70.3 | 84.4 | 76.8 | 0.831 | 0.548 |
| SMO | 70.3 | 87.5 | 78.3 | 0.789 | 0.581 |
| IBK | 75.7 | 84.4 | 79.7 | 0.789 | 0.599 |
| RARF | 78.4 | 81.3 | 79.7 | 0.893 | 0.595 |
| PART | 56.8 | 81.3 | 68.1 | 0.700 | 0.388 |
| JRip | 78.4 | 65.6 | 72.5 | 0.733 | 0.445 |
| RandomForest | 75.7 | 81.3 | 78.3 | 0.868 | 0.568 |
| J48 | 70.3 | 71.9 | 71.0 | 0.735 | 0.420 |
Table 6.
Performance evaluation metrics of eight classifiers for reduced dataset generated by proposed approach consisting of PL+ and PL- molecules based on percentage split of 66:34.
| Classifiers | Sensitivity | Specificity | Accuracy | AUC | MCC |
|---|---|---|---|---|---|
| Navie Bayes | 86.5 | 71.9 | 79.7 | 0.851 | 0.593 |
| SMO | 73.0 | 81.3 | 76.8 | 0.771 | 0.541 |
| IBK | 81.1 | 84.4 | 82.6 | 0.834 | 0.653 |
| RARF | 91.9 | 87.5 | 89.9 | 0.903 | 0.796 |
| PART | 78.4 | 84.4 | 81.2 | 0.890 | 0.626 |
| LRip | 54.1 | 93.8 | 72.5 | 0.735 | 0.512 |
| RandomForest | 81.1 | 87.5 | 84.1 | 0.904 | 0.684 |
| J48 | 83.8 | 87.5 | 85.5 | 0.842 | 0.711 |
Figure 5.
ROC curve for orginal dataset for various machine learing algorithms.
Figure 6.
ROC curve for reduced dataset by various machine learing algorithms.
To compare with the performance evaluation metrics for the phospholipidosis dataset, we used the same package in R (https://https://cran.r-project.org/web/packages/h2o/index.html)as used in the original work (Nath et al.87). We used a grid search strategy to obtain the best hyperparameters for the random forest algorithm Hyperpaprametersntrees = c(20,50,100,500),max depth = c(20,40,60,80),sample rate = c(0.2,1,0.01). Further, we used the same of features (JOELib+Structural alerts), which are calculated using the ChemMine tools webserver (https://chemminetools.ucr.edu/). The dataset consisted of 102 phospholipidosis inducing compounds (positive samples) and 83 phospholipidosis non-inducing compounds (negative samples), thus constituting a total of 185 molecules. Schematic representation for entire process is given by Fig. 7. In the current methodology, we start the process with a dataset consisted of phospholipidosis positive molecules and phospholipidosis negative molecules. Then, descriptor generator converts the initial data into target data. Further, SMOTE is applied to obtain the balanced dataset. Next, this dataset is converted into intuitionistic fuzzy information system by using Tan et al.57 approach. Thereafter, our proposed feature subset selection method is applied to remove noise, vagueness, irrelevancy, redundancy, and uncertainty to obtain reduced dataset. Moreover, several classifiers are used to discriminate positive and negative classes. Finally, RARF is identified as the best performer.
Figure 7.
Schematic representation for generating classifier for phospholipidosis.
The performance evaluation metrics for the current method and the previous ensemble based method are presented in Table 7. The dataset preprocessing introduced in the current work resulted in enhanced performance evaluation metrics for the RF algorithm in comparison to the previously published results. Notably a 2 percent rise on overall accuracy is observed. As the dataset is slightly imbalanced, a rise in MCC for the current method proves the usefulness of the dataset preprocessing step. The ROC plot for the RF(h2o) model is presented in Fig. 4. An AUC value of 0.922 indicates an acceptable prediction model for phospholipidosis inducing molecules. In the end of the entire study, the list of abrreviations, signs, and symbols are presented in Table 8.
Table 7.
Perfomance evaluation metrics for the RF algorithm with previous method.
| Classifiers | Sensitivity | Specificity | Accuracy | AUC | MCC |
|---|---|---|---|---|---|
| RF(h2o) | 86.7 | 93.0 | 90.1 | 0.922 | 0.808 |
| Nath et. al87 | 86.2 | 90.1 | 88.2 | 0.896 | 0.725 |
Table 8.
The list of Abbreviations,Symbols, and Signs.
| Abbreviations/symbols/signs | Explanation |
|---|---|
| IFS | Intuitionistic Fuzzy Set |
| FRS | Fuzzy rough set |
| DIFDT | Dominant intuitionistic fuzzy decision table |
| IFRS | Intuitionistic fuzzy rough set |
| IFMGRS | Intuitionistic fuzzy multigranulation rough set |
| MI | Mutual information |
| PL+ | Phsopholipidosis positive |
| PL- | Phsopholipidosis negative |
| IFIS | Intuitionistic fuzzy information system |
| RARF | RealAdaBoost random forest |
| TRP | True positive |
| TRN | True negative |
| FLP | False positive |
| FLN | False negative |
| Sn | Sensitivity |
| Sp | Specificity |
| Ac | Accuracy |
| AUC | Area under curve |
| MCC | Mathews correlation coefficient |
| ROC | Receiver operating characteristic |
| SMO | Sequential minimal optimization |
| IBK | Instance based learner |
| FSFrMI | Feature selection based on fuzzy rough mutual information |
| GIFRFS | Granular structure based intuitionistic fuzzy rough feature selection |
| TIFRFS | Tolrence based fuzzy rough feature selection |
| FRFS | Fuzzy rough feature selection |
| IFRFSMI | Intuitionistic fuzzy rough feature selection based on mutual information |
| Membership grade | |
| Non-membership grade | |
| Hesitancy grade | |
| Hybrid similarity relation | |
| Adaptive intuitionistic fuzzy radius | |
| ET | Entropy |
| I | Mutual information |
| Summation | |
| Union | |
| Intersection | |
| Epsilon | |
| Forall | |
| Belong | |
| Significance |
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conclusion
Dimensionality reduction broadly aims to obtain a feature subset from existing original feature set by using certain powerful evaluation criterion. Since dimensionality reduction can produce efficient feature subset, where feature selection has found as an interesting central technique for data pre-processing in various beneficial and interesting data mining tasks. Conventional fuzzy rough set frequently incorporates dependency function as an evaluation criterion of feature subset selection. However, this method only maintained the maximum membership grade of a data point to one decision class and found to be unable in discarding later uncertainty and noise up to certain extent, which cannot characterize the classification error. To avoid these issues, we presented a novel intuitionistic fuzzy aided technique, where feature selection method is established by integrating information entropy with IF rough set concept.
Initially, we established a hybrid IF similarity relation, which is further employed to present a novel IF rough joint and conditional entropies.
Then, IF granular structure was introduced based on the proposed hybrid similarity relation.
Thereafter, IF rough set model was described by using the aforesaid relation.
Based on these entropies and granular structure, we suggested a mutual information idea to compute the significance of the feature subset for a decision class.
Next, mathematical theorems are validated to justify the correctness of the proposed ideas.
By using the significance notion a heuristic IF rough feature selection algorithm is represented. Then, we apply this heuristic algorithm on ten benchmark datasets to illustrate extensive experiments.
Finally, proposed method is successfully employed to enhance the prediction performance for identifying PL+ and PL- molecules.
For dbworld-bodies dataset, our method has eliminated 99.83% features. Moreover, performance measures of learning algorithms were evaluated based on the reduced data produced by four existing and our proposed methods, where results clearly indicate superiority of the proposed technique. For thyroid- hypothyroid dataset, RARF has reported an accuracy of 99.11% and standard deviation of 0.46% for IFRFSMI based reduced dataset. For the discrimination of PL+ and PL- molecules, the best sensitivity is achieved based on 66:34 validation technique with 91.9%. The best overall result was obtained by RF(h2o) with sensitivity, specificity, accuracy, AUC, and MCC of 86.7%, 93.0%, 90.1%, 0.922, and 0.808 respectively.
The advantages of our proposed methodology can be outlined as bellow:
This study presents a new hybrid similarity relation that can handle mixed data in intuitionistic fuzzy framework.
Adaptive radius is computed in the recursive way from relation itself, which ensures the information loss.
IF granular structure is implemented to deal with noise in mixed data as it is based on our proposed hybrid relation.
IF rough mutual information is implemented to cope with noise and later uncertainty based on the proposed IF granular structure.
This study presents a new methodology to discriminate PL+ and PL- molecules in an efficient and efficacious way.
In future, the proposed hybrid similarity relation can be improved by providing a more effective definition of adaptive radius. Further, inner and outer significance can be computed by assembling mutual information in robust IF rough framework to establish efficient approach to calculate the correlation between feature subset and class.
Author contributions
A.K.T.: Conceptualization, Problem formulation, Methodology, Original draft preparation, Reviewing and Editing, and Final drafting. R.S.: Numerical analysis, Programming, Mathematical Modelling. A.N.: Data curation, Programming, Simulation, Validation, Numerical analysis, Visualization, and System set-up. P.S.: Mathematical Modelling, Visualization, and Investigation. M.A.S.: Supervision, Problem formulation, Programming, Validation, Writing, Reviewing, and Editing.
Funding
The authors did not receive support from any organization for the submitted work.
Data availability
The data supporting this study’s findings are available from the corresponding author (Mohd Asif Shah) upon reasonable request.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Rajat Saini, Email: afcatrajat@gmail.com.
Mohd Asif Shah, Email: drmohdasifshah@kdu.edu.et.
References
- 1.Issad HA, Aoudjit R, Rodrigues JJ. A comprehensive review of data mining techniques in smart agriculture. Eng. Agric. Environ. Food. 2019;12(4):511–525. doi: 10.1016/j.eaef.2019.11.003. [DOI] [Google Scholar]
- 2.Li J, Cheng K, Wang S, Morstatter F, Trevino RP, Tang J, Liu H. Feature selection: A data perspective. ACM Comput. Surv. (CSUR) 2017;50(6):1–45. doi: 10.1145/3136625. [DOI] [Google Scholar]
- 3.Papakyriakou D, Barbounakis IS. Data mining methods: A review. Int. J. Comput. Appl. 2022;183(48):5–19. [Google Scholar]
- 4.Awais M, Salahuddin T. Radiative magnetodydrodynamic cross fluid thermophysical model passing on parabola surface with activation energy. Ain Shams Eng. J. 2024;15(1):102282. doi: 10.1016/j.asej.2023.102282. [DOI] [Google Scholar]
- 5.Awais, M. & Salahuddin, T. Variable thermophysical properties of magnetohydrodynamic cross fluid model with effect of energy dissipation and chemical reaction. Int. J. Mod. Phys. B, 2450197 (2023).
- 6.Jensen R, Shen Q. Semantics-preserving dimensionality reduction: Rough and fuzzy-rough-based approaches. IEEE Trans. Knowl. Data Eng. 2004;16(12):1457–1471. doi: 10.1109/TKDE.2004.96. [DOI] [Google Scholar]
- 7.Awais M, Salahuddin T, Muhammad S. Effects of viscous dissipation and activation energy for the MHD Eyring-powell fluid flow with Darcy-Forchheimer and variable fluid properties. Ain Shams Eng. J. 2024;15(2):102422. doi: 10.1016/j.asej.2023.102422. [DOI] [Google Scholar]
- 8.Chauhan, D. & Mathews, R. Review on dimensionality reduction techniques. In Proceeding of the International Conference on Computer Networks, Big Data and IoT (ICCBI-2019) 356–362 (Springer International Publishing, 2020).
- 9.Hu J, Chen H, Heidari AA, Wang M, Zhang X, Chen Y, Pan Z. Orthogonal learning covariance matrix for defects of grey wolf optimizer: Insights, balance, diversity, and feature selection. Knowl.-Based Syst. 2021;213:106684. doi: 10.1016/j.knosys.2020.106684. [DOI] [Google Scholar]
- 10.Jia W, Sun M, Lian J, Hou S. Feature dimensionality reduction: A review. Complex Intell. Syst. 2022;8(3):2663–2693. doi: 10.1007/s40747-021-00637-x. [DOI] [Google Scholar]
- 11.Tubishat M, Idris N, Shuib L, Abushariah MA, Mirjalili S. Improved Salp Swarm Algorithm based on opposition based learning and novel local search algorithm for feature selection. Expert Syst. Appl. 2020;145:113122. doi: 10.1016/j.eswa.2019.113122. [DOI] [Google Scholar]
- 12.Chandrashekar G, Sahin F. A survey on feature selection methods. Comput. Electr. Eng. 2014;40(1):16–28. doi: 10.1016/j.compeleceng.2013.11.024. [DOI] [Google Scholar]
- 13.Remeseiro B, Bolon-Canedo V. A review of feature selection methods in medical applications. Comput. Biol. Med. 2019;112:103375. doi: 10.1016/j.compbiomed.2019.103375. [DOI] [PubMed] [Google Scholar]
- 14.Saeys Y, Inza I, Larranaga P. A review of feature selection techniques in bioinformatics. Bioinformatics. 2007;23(19):2507–2517. doi: 10.1093/bioinformatics/btm344. [DOI] [PubMed] [Google Scholar]
- 15.Bommert A, Sun X, Bischl B, Rahnenführer J, Lang M. Benchmark for filter methods for feature selection in high-dimensional classification data. Comput. Stat. Data Anal. 2020;143:106839. doi: 10.1016/j.csda.2019.106839. [DOI] [Google Scholar]
- 16.Cai J, Luo J, Wang S, Yang S. Feature selection in machine learning: A new perspective. Neurocomputing. 2018;300:70–79. doi: 10.1016/j.neucom.2017.11.077. [DOI] [Google Scholar]
- 17.Dash M, Liu H. Feature selection for classification. Intell. Data Anal. 1997;1(1–4):131–156. doi: 10.3233/IDA-1997-1302. [DOI] [Google Scholar]
- 18.Pawlak Z. Rough sets. Int. J. Comput. Inf. Sci. 1982;11:341–356. doi: 10.1007/BF01001956. [DOI] [Google Scholar]
- 19.Pawlak Z, Grzymala-Busse J, Slowinski R, Ziarko W. Rough sets. Commun. ACM. 1995;38(11):88–95. doi: 10.1145/219717.219791. [DOI] [Google Scholar]
- 20.Sivasankar E, Selvi C, Mahalakshmi S. Rough set-based feature selection for credit risk prediction using weight-adjusted boosting ensemble method. Soft. Comput. 2020;24(6):3975–3988. doi: 10.1007/s00500-019-04167-0. [DOI] [Google Scholar]
- 21.Bania RK, Halder A. R-HEFS: Rough set based heterogeneous ensemble feature selection method for medical data classification. Artif. Intell. Med. 2021;114:102049. doi: 10.1016/j.artmed.2021.102049. [DOI] [PubMed] [Google Scholar]
- 22.Thangavel K, Pethalakshmi A. Dimensionality reduction based on rough set theory: A review. Appl. Soft Comput. 2009;9(1):1–12. doi: 10.1016/j.asoc.2008.05.006. [DOI] [Google Scholar]
- 23.Campagner A, Ciucci D, Hüllermeier E. Rough set-based feature selection for weakly labeled data. Int. J. Approx. Reason. 2021;136:150–167. doi: 10.1016/j.ijar.2021.06.005. [DOI] [Google Scholar]
- 24.Jensen, R. Rough set-based feature selection: A review. In Rough Computing: Theories, Technologies and Applications 70–107 (2008).
- 25.Raza MS, Qamar U. Understanding and Using Rough Set Based Feature Selection: Concepts, Techniques and Applications. Springer; 2017. [Google Scholar]
- 26.Zadeh LA. Fuzzy sets. Inf. Control. 1965;8(3):338–353. doi: 10.1016/S0019-9958(65)90241-X. [DOI] [Google Scholar]
- 27.Dubois D, Prade H. Putting rough sets and fuzzy sets together. In: Slowinski R, editor. Intelligent Decision Support: Handbook of Applications and Advances of the Rough Sets Theory. Springer; 1992. pp. 203–232. [Google Scholar]
- 28.Chen J, Mi J, Lin Y. A graph approach for fuzzy-rough feature selection. Fuzzy Sets Syst. 2020;391:96–116. doi: 10.1016/j.fss.2019.07.014. [DOI] [Google Scholar]
- 29.Qiu Z, Zhao H. A fuzzy rough set approach to hierarchical feature selection based on Hausdorff distance. Appl. Intell. 2022;52(10):11089–11102. doi: 10.1007/s10489-021-03028-4. [DOI] [Google Scholar]
- 30.Sang B, Yang L, Chen H, Xu W, Zhang X. Fuzzy rough feature selection using a robust non-linear vague quantifier for ordinal classification. Expert Syst. Appl. 2023;230:120480. doi: 10.1016/j.eswa.2023.120480. [DOI] [Google Scholar]
- 31.Yin T, Chen H, Li T, Yuan Z, Luo C. Robust feature selection using label enhancement and -precision fuzzy rough sets for multilabel fuzzy decision system. Fuzzy Sets Syst. 2023;461:108462. doi: 10.1016/j.fss.2022.12.018. [DOI] [Google Scholar]
- 32.Wang C, Huang Y, Ding W, Cao Z. Attribute reduction with fuzzy rough self-information measures. Inf. Sci. 2021;549:68–86. doi: 10.1016/j.ins.2020.11.021. [DOI] [Google Scholar]
- 33.Zhang X, Mei C, Chen D, Yang Y. A fuzzy rough set-based feature selection method using representative instances. Knowl.-Based Syst. 2018;151:216–229. doi: 10.1016/j.knosys.2018.03.031. [DOI] [Google Scholar]
- 34.Wang C, Huang Y, Shao M, Fan X. Fuzzy rough set-based attribute reduction using distance measures. Knowl.-Based Syst. 2019;164:205–212. doi: 10.1016/j.knosys.2018.10.038. [DOI] [Google Scholar]
- 35.Wang C, Wang Y, Shao M, Qian Y, Chen D. Fuzzy rough attribute reduction for categorical data. IEEE Trans. Fuzzy Syst. 2019;28(5):818–830. doi: 10.1109/TFUZZ.2019.2949765. [DOI] [Google Scholar]
- 36.Yang X, Chen H, Li T, Luo C. A noise-aware fuzzy rough set approach for feature selection. Knowl.-Based Syst. 2022;250:109092. doi: 10.1016/j.knosys.2022.109092. [DOI] [Google Scholar]
- 37.Yang X, Chen H, Li T, Zhang P, Luo C. Student-t kernelized fuzzy rough set model with fuzzy divergence for feature selection. Inf. Sci. 2022;610:52–72. doi: 10.1016/j.ins.2022.07.139. [DOI] [Google Scholar]
- 38.Yuan Z, Chen H, Xie P, Zhang P, Liu J, Li T. Attribute reduction methods in fuzzy rough set theory: An overview, comparative experiments, and new directions. Appl. Soft Comput. 2021;107:107353. doi: 10.1016/j.asoc.2021.107353. [DOI] [Google Scholar]
- 39.Jain P, Tiwari AK, Som T. A fitting model based intuitionistic fuzzy rough feature selection. Eng. Appl. Artif. Intell. 2020;89:103421. doi: 10.1016/j.engappai.2019.103421. [DOI] [Google Scholar]
- 40.Annamalai, C. Intuitionistic fuzzy sets: New approach and applications (2022).
- 41.Dan S, Kar MB, Majumder S, Roy B, Kar S, Pamucar D. Intuitionistic type-2 fuzzy set and its properties. Symmetry. 2019;11(6):808. doi: 10.3390/sym11060808. [DOI] [Google Scholar]
- 42.Atanassov KT, Stoeva S. Intuitionistic fuzzy sets. Fuzzy Sets Syst. 1986;20(1):87–96. doi: 10.1016/S0165-0114(86)80034-3. [DOI] [Google Scholar]
- 43.Cornelis C, De Cock M, Kerre EE. Intuitionistic fuzzy rough sets: At the crossroads of imperfect knowledge. Expert Syst. 2003;20(5):260–270. doi: 10.1111/1468-0394.00250. [DOI] [Google Scholar]
- 44.Zhan J, Masood Malik H, Akram M. Novel decision-making algorithms based on intuitionistic fuzzy rough environment. Int. J. Mach. Learn. Cybern. 2019;10:1459–1485. doi: 10.1007/s13042-018-0827-4. [DOI] [Google Scholar]
- 45.Zhang Z. Attributes reduction based on intuitionistic fuzzy rough sets. J. Intell. Fuzzy Syst. 2016;30(2):1127–1137. doi: 10.3233/IFS-151835. [DOI] [Google Scholar]
- 46.Atanassov KT, Atanassov KT. Intuitionistic Fuzzy Sets. Springer; 1999. [Google Scholar]
- 47.Tseng T-LB, Huang C-C. Rough set-based approach to feature selection in customer relationship management. Omega. 2007;35(4):365–383. doi: 10.1016/j.omega.2005.07.006. [DOI] [Google Scholar]
- 48.Zhang X, Zhou B, Li P. A general frame for intuitionistic fuzzy rough sets. Inf. Sci. 2012;216:34–49. doi: 10.1016/j.ins.2012.04.018. [DOI] [Google Scholar]
- 49.Zhou L, Wu W-Z. On generalized intuitionistic fuzzy rough approximation operators. Inf. Sci. 2008;178(11):2448–2465. [Google Scholar]
- 50.Jain P, Som T. Multigranular rough set model based on robust intuitionistic fuzzy covering with application to feature selection. Int. J. Approx. Reason. 2023;156:16–37. doi: 10.1016/j.ijar.2023.02.004. [DOI] [Google Scholar]
- 51.Liu Y, Lin Y. Intuitionistic fuzzy rough set model based on conflict distance and applications. Appl. Soft Comput. 2015;31:266–273. doi: 10.1016/j.asoc.2015.02.045. [DOI] [Google Scholar]
- 52.Huang B, Zhuang Y-L, Li H-X, Wei D-K. A dominance intuitionistic fuzzy-rough set approach and its applications. Appl. Math. Model. 2013;37(12–13):7128–7141. doi: 10.1016/j.apm.2012.12.009. [DOI] [Google Scholar]
- 53.Wang C, Huang Y, Shao M, Hu Q, Chen D. Feature selection based on neighborhood self-information. IEEE Trans. Cybern. 2019;50(9):4031–4042. doi: 10.1109/TCYB.2019.2923430. [DOI] [PubMed] [Google Scholar]
- 54.Xu J, Shen K, Sun L. Multi-label feature selection based on fuzzy neighborhood rough sets. Complex Intell. Syst. 2022;8(3):2105–2129. doi: 10.1007/s40747-021-00636-y. [DOI] [Google Scholar]
- 55.Huang B, Li H, Feng G, Zhou X. Dominance-based rough sets in multi-scale intuitionistic fuzzy decision tables. Appl. Math. Comput. 2019;348:487–512. [Google Scholar]
- 56.Huang B, Guo C-X, Zhuang Y-L, Li H-X, Zhou X-Z. Intuitionistic fuzzy multigranulation rough sets. Inf. Sci. 2014;277:299–320. doi: 10.1016/j.ins.2014.02.064. [DOI] [Google Scholar]
- 57.Tan A, Wu W-Z, Qian Y, Liang J, Chen J, Li J. Intuitionistic fuzzy rough set-based granular structures and attribute subset selection. IEEE Trans. Fuzzy Syst. 2018;27(3):527–539. doi: 10.1109/TFUZZ.2018.2862870. [DOI] [Google Scholar]
- 58.Zhou L, Wu W-Z, Zhang W-X. On characterization of intuitionistic fuzzy rough sets based on intuitionistic fuzzy implicators. Inf. Sci. 2009;179(7):883–898. doi: 10.1016/j.ins.2008.11.015. [DOI] [Google Scholar]
- 59.Tiwari AK, Shreevastava S, Som T, Shukla KK. Tolerance-based intuitionistic fuzzy-rough set approach for attribute reduction. Expert Syst. Appl. 2018;101:205–212. doi: 10.1016/j.eswa.2018.02.009. [DOI] [Google Scholar]
- 60.Shreevastava, S., Tiwari, A. & Som, T. Feature subset selection of semi-supervised data: An intuitionistic fuzzy-rough set-based concept. In Proceedings of International Ethical Hacking Conference 2018: eHaCON 2018, Kolkata, India (2019).
- 61.Tiwari AK, Shreevastava S, Subbiah K, Som T. An intuitionistic fuzzy-rough set model and its application to feature selection. J. Intell. Fuzzy Syst. 2019;36(5):4969–4979. doi: 10.3233/JIFS-179043. [DOI] [Google Scholar]
- 62.Tiwari AK, Shreevastava S, Shukla KK, Subbiah K. New approaches to intuitionistic fuzzy-rough attribute reduction. J. Intell. Fuzzy Syst. 2018;34(5):3385–3394. doi: 10.3233/JIFS-169519. [DOI] [Google Scholar]
- 63.Tiwari AK, Shreevastava S, Subbiah K, Som T. An intuitionistic fuzzy-rough set model and its application to feature selection. J. Intell. Fuzzy Syst. 2019;36(5):4969–4979. doi: 10.3233/JIFS-179043. [DOI] [Google Scholar]
- 64.Shreevastava S, Singh S, Tiwari A, Som T. Different classes ratio and Laplace summation operator based intuitionistic fuzzy rough attribute selection. Iran. J. Fuzzy Syst. 2021;18(6):67–82. [Google Scholar]
- 65.Shreevastava S, Tiwari AK, Som T. Intuitionistic fuzzy neighborhood rough set model for feature selection. Int. J. Fuzzy Syst. Appl. (IJFSA) 2018;7(2):75–84. [Google Scholar]
- 66.Li LQ, Wang XL, Liu ZX, Xie WX. A novel intuitionistic fuzzy clustering algorithm based on feature selection for multiple object tracking. Int. J. Fuzzy Syst. 2019;21:1613–1628. doi: 10.1007/s40815-019-00645-7. [DOI] [Google Scholar]
- 67.Singh S, Shreevastava S, Som T, Jain P. Intuitionistic fuzzy quantifier and its application in feature selection. Int. J. Fuzzy Syst. 2019;21:441–453. doi: 10.1007/s40815-018-00603-9. [DOI] [Google Scholar]
- 68.Sun L, Wang L, Ding W, Qian Y, Xu J. Feature selection using fuzzy neighborhood entropy-based uncertainty measures for fuzzy neighborhood multigranulation rough sets. IEEE Trans. Fuzzy Syst. 2020;29(1):19–33. doi: 10.1109/TFUZZ.2020.2989098. [DOI] [Google Scholar]
- 69.Sun L, Zhang X, Qian Y, Xu J, Zhang S. Feature selection using neighborhood entropy-based uncertainty measures for gene expression data classification. Inf. Sci. 2019;502:18–41. doi: 10.1016/j.ins.2019.05.072. [DOI] [Google Scholar]
- 70.Fang L, Zhao H, Wang P, Yu M, Yan J, Cheng W, Chen P. Feature selection method based on mutual information and class separability for dimension reduction in multidimensional time series for clinical data. Biomed. Signal Process. Control. 2015;21:82–89. doi: 10.1016/j.bspc.2015.05.011. [DOI] [Google Scholar]
- 71.Fernandes AD, Gloor GB. Mutual information is critically dependent on prior assumptions: Would the correct estimate of mutual information please identify itself? Bioinformatics. 2010;26(9):1135–1139. doi: 10.1093/bioinformatics/btq111. [DOI] [PubMed] [Google Scholar]
- 72.Wang Z, Chen H, Yuan Z, Yang X, Zhang P, Li T. Exploiting fuzzy rough mutual information for feature selection. Appl. Soft Comput. 2022;131:109769. doi: 10.1016/j.asoc.2022.109769. [DOI] [Google Scholar]
- 73.Xie L, Lin G, Li J, Lin Y. A novel fuzzy-rough attribute reduction approach via local information entropy. Fuzzy Sets Syst. 2023;473:108733. doi: 10.1016/j.fss.2023.108733. [DOI] [Google Scholar]
- 74.Xu F, Miao D, Wei L. Fuzzy-rough attribute reduction via mutual information with an application to cancer classification. Comput. Math. Appl. 2009;57(6):1010–1017. doi: 10.1016/j.camwa.2008.10.027. [DOI] [Google Scholar]
- 75.Fang H, Tang P, Si H. Feature selections using minimal redundancy maximal relevance algorithm for human activity recognition in smart home environments. J. Healthc. Eng. 2020;2020:1–13. [Google Scholar]
- 76.Xie S, Zhang Y, Lv D, Chen X, Lu J, Liu J. A new improved maximal relevance and minimal redundancy method based on feature subset. J. Supercomput. 2023;79(3):3157–3180. doi: 10.1007/s11227-022-04763-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Maji P, Garai P. On fuzzy-rough attribute selection: Criteria of max-dependency, max-relevance, min-redundancy, and max-significance. Appl. Soft Comput. 2013;13(9):3968–3980. doi: 10.1016/j.asoc.2012.09.006. [DOI] [Google Scholar]
- 78.Zhang X, Mei C, Chen D, Li J. Feature selection in mixed data: A method using a novel fuzzy rough set-based information entropy. Pattern Recogn. 2016;56:1–15. doi: 10.1016/j.patcog.2016.02.013. [DOI] [Google Scholar]
- 79.Zhang X, Mei C, Chen D, Yang Y, Li J. Active incremental feature selection using a fuzzy-rough-set-based information entropy. IEEE Trans. Fuzzy Syst. 2019;28(5):901–915. doi: 10.1109/TFUZZ.2019.2959995. [DOI] [Google Scholar]
- 80.Anderson N, Borlak J. Drug-induced phospholipidosis. FEBS Lett. 2006;580(23):5533–5540. doi: 10.1016/j.febslet.2006.08.061. [DOI] [PubMed] [Google Scholar]
- 81.Breiden B, Sandhoff K. Emerging mechanisms of drug-induced phospholipidosis. Biol. Chem. 2020;401(1):31–46. doi: 10.1515/hsz-2019-0270. [DOI] [PubMed] [Google Scholar]
- 82.Shayman JA, Abe A. Drug induced phospholipidosis: An acquired lysosomal storage disorder. Biochim. Biophys. Acta (BBA)-Mol. Cell Biol. Lipids. 2013;1831(3):602–611. doi: 10.1016/j.bbalip.2012.08.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Salahuddin T. Numerical Techniques in MATLAB: Fundamental to Advanced Concepts. CRC Press; 2023. [Google Scholar]
- 84.Frank E, Hall M, Trigg L, Holmes G, Witten IH. Data mining in bioinformatics using Weka. Bioinformatics. 2004;20(15):2479–2481. doi: 10.1093/bioinformatics/bth261. [DOI] [PubMed] [Google Scholar]
- 85.Asuncion, A. & Newman, D. UCI machine learning repository. In: Irvine, CA, USA (2007).
- 86.Hall M, Frank E, Holmes G, Pfahringer B, Reutemann P, Witten IH. The WEKA data mining software: An update. ACM SIGKDD Explor. Newsl. 2009;11(1):10–18. doi: 10.1145/1656274.1656278. [DOI] [Google Scholar]
- 87.Nath A, Sahu GK. Exploiting ensemble learning to improve prediction of phospholipidosis inducing potential. J. Theor. Biol. 2019;479:37–47. doi: 10.1016/j.jtbi.2019.07.009. [DOI] [PubMed] [Google Scholar]
- 88.Friedman M. A comparison of alternative tests of significance for the problem of m rankings. Ann. Math. Stat. 1940;11(1):86–92. doi: 10.1214/aoms/1177731944. [DOI] [Google Scholar]
- 89.Dunn OJ. Multiple comparisons among means. J. Am. Stat. Assoc. 1961;56(293):52–64. doi: 10.1080/01621459.1961.10482090. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The data supporting this study’s findings are available from the corresponding author (Mohd Asif Shah) upon reasonable request.