Abstract
The skeletal system is crucial for supporting bodily functions, protecting vital organs, facilitating hematopoiesis, and storing essential minerals. Skeletal homeostasis, which includes aspects such as bone density, structural integrity, and regenerative processes, is essential for normal skeletal function. Autophagy, an intricate intracellular mechanism for degrading and recycling cellular components, plays a multifaceted role in bone metabolism. It involves sequestering cellular waste, damaged proteins, and organelles within autophagosomes, which are then degraded and recycled. Autophagy’s impact on bone health varies depending on factors such as regulation, cell type, environmental cues, and physiological context. Despite being traditionally considered a cytoplasmic process, autophagy is subject to transcriptional and epigenetic regulation within the nucleus. However, the precise influence of epigenetic regulation, including DNA methylation, histone modifications, and non-coding RNA expression, on cellular fate remains incompletely understood. The interplay between autophagy and epigenetic modifications adds complexity to bone cell regulation. This article provides an in-depth exploration of the intricate interplay between these two regulatory paradigms, with a focus on the epigenetic control of autophagy in bone metabolism. Such an understanding enhances our knowledge of bone metabolism-related disorders and offers insights for the development of targeted therapeutic strategies.
Keywords: epigenetics, autophagy, bone homeostasis, bone metabolism, miRNA, bone-related diseases
Graphical Abstract
Graphical Abstract.
Introduction
The skeletal system, characterized by its dense and robust tissues, serves as a cornerstone in supporting bodily functions, safeguarding vital organs, facilitating hematopoiesis, and storing essential minerals.1–3 Maintaining bone homeostasis is paramount for ensuring the normal functionality of the skeleton, encompassing critical aspects such as bone density, structural integrity, and regenerative processes.4
Autophagy sequesters cellular waste, damaged proteins, and organelles within autophagosomes, subsequently subject to degradation and recycling.5 The outcome of autophagy in bone metabolism depends on various factors, including autophagy regulation, cell type, environmental cues, and physiological context.6
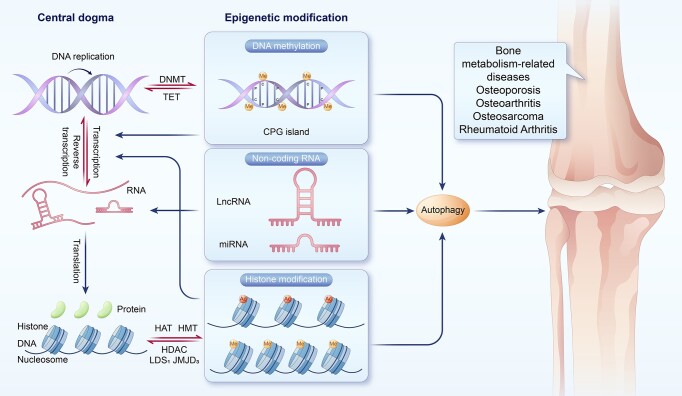
While traditionally perceived as a cytoplasmic process,7 autophagy is subject to transcriptional and epigenetic regulation within the nucleus.8–10 However, the precise impact of epigenetic regulation, encompassing DNA methylation, histone modifications, and non-coding RNA (ncRNA) expression, on cellular fate remains incompletely elucidated.8 The interplay between autophagy and epigenetic modifications adds a layer of complexity to bone cell regulation.
This article explores the intricate interplay between these two regulatory paradigms, shedding light on the epigenetic control of autophagy within the realm of bone metabolism. This exploration enhances our comprehension of bone metabolism-related disorders and pondered the future and challenges of epigenetic regulation of autophagy in bone metabolism.
Overview of Epigenetic Modifications
Epigenetics broadly pertains to inheritable modifications in phenotype that do not involve modifications in DNA sequences.11,12 While the genotype remains unchanged, hereditary alterations occur in the phenotype. Among the most well-established epigenetic mechanisms are DNA methylation, post-translational modifications of histones, and ncRNAs.13–15 These mechanisms serve as key regulators of gene expression, functioning either through the modulation of chromatin structure and gene transcription or, in the case of ncRNAs, through post-transcriptional control of protein translation.16,17 Epigenetics plays a regulatory role in various biological processes, including the specific expression of genes in different tissues, the inactivation of chromosomes, genomic imprinting, and the differentiation of cells.18,19 With the advancement of research techniques, epigenetic abnormalities have been implicated in causing malignancies, metabolic disorders, somatic diseases, and autoimmune diseases.20 Epigenetics highlights the intricate interplay between genetic and environmental factors and plays a crucial role in regulating bone metabolism,21 particularly in the differentiation of bone cells, providing insights into the study of bone metabolic disorders and their treatment directions.
DNA Methylation
DNA methylation stands as one of the most extensively studied and common epigenetic modifications in mammals.22 This entails methylating the 5′ carbon of cytosine, leading to the formation of 5-methylcytosine (5-mC).23,24 Cytosine methylation predominantly occurs within specific dinucleotide sequences known as CpG sites. Importantly, these CpG sites are not randomly distributed across the genome but are concentrated in regions called CpG islands, often situated at the 5′ ends of genes. High methylation levels in CpG islands are commonly associated with gene silencing, while low methylation levels are linked to gene activation.25,26
The enzymes responsible for transferring methyl groups from S-adenosylmethionine (SAM) to DNA include DNA methyltransferase 1 (DNMT1), DNMT3a, DNMT3b, and DNMT3L.27 Demethylation of DNA can occur either passively following DNA replication, leading to reduced maintenance methylation, or actively.28 The precise molecular mechanisms underlying active demethylation are not yet fully understood. For instance, in the context of bone biology, Interferon Regulatory Factor 8 (IRF8) transcription factor inhibits osteoclast formation.29 Experimental evidence has revealed that DNMT3a plays a significant role in inhibiting the activity of IRF8 by increasing methylation at distal regulatory elements associated with the IRF8 gene. Elevated levels of SAM can enhance this methylation, ultimately promoting osteoclast differentiation and bone resorption.30 Deleting DNMT3a in osteoclasts (OC) or using the inhibitor TF-3 in mice protects against bone loss after ovarian removal.31 Moreover, in multiple myeloma patients, alterations in bone resorption have been associated with elevated IRF8 methylation, induced by myeloma cells’ release of thymidine phosphorylase (TP), leading to decreased IRF8 expression and enhanced bone resorption.32
The transcription factors runt-related transcription factor 2 (Runx2) and Osterix (OSX) play critical roles in regulating osteoblast (OB) differentiation and bone matrix synthesis. Runx2 activates the promoter of OSX, indicating Runx2’s position as an upstream transcription factor in osteogenesis.33 During mesenchymal stem cell differentiation into osteoblasts, there is a reduction in the methylation level of Runx2, underscoring the significant regulatory function of Runx2 methylation during osteoblast differentiation.29 Additionally, epigenetic regulation of OSX is vital for guiding mesenchymal stem cells (MSCs) into osteoblast differentiation.34
The Ten-Eleven-Translocation (TET) family of enzymes are recognized for their ability to remove methyl groups from DNA, converting 5-mC into 5-hydroxymethylcytosine (5-hmC). Demethylation mediated by TET enzymes enhances chromatin accessibility for target genes controlled by Runx2, facilitating transcriptional regulation. TET proteins engage with Runx2 via their catalytic domain, influencing cytosine methylation patterns around the Runx2 binding region.35 Disruptions in the promoter methylation of key bone-related genes, such as bone morphogenetic protein 2 (BMP2), may lead to irregularities in bone formation.29
Post-Translational Histone Modifications
Inside the eukaryotic nucleus, a segment of double-stranded DNA spanning 147 base pairs coils around eight histone proteins, which include two copies of H2A, H2B, H3, and H4, creating nucleosomes. These nucleosomes are linked by DNA segments and organize into structured chromatin formations called chromatin, which subsequently compact into chromosomes.36–38 The N-terminal amino acid tails of H3 and H4 undergo post-translational modifications that profoundly affect both chromatin structure and DNA-associated activities.39 Research has unveiled that euchromatin, representing a relaxed and actively transcribed DNA state, is marked by elevated levels of acetylation and trimethylation at H3K4, H3K36, and H3K79. In contrast, reduced acetylation levels and heightened methylation at H3K9, H3K27, and H4K20 indicate a more condensed, transcriptionally inactive heterochromatin structure.23,40 Histone modification is the most intricate mode of regulation among the three epigenetic modifications.41 Currently, the most extensively studied histone modifications in bone metabolism are histone acetylation and histone methylation.
Histone Acetylation
Histone acetylation involves the addition of acetyl groups to histones and is governed by histone acetyltransferases (HATs). Conversely, histone deacetylases (HDACs) regulate the removal of these acetyl groups, leading to histone deacetylation.42 This balance between HATs and HDACs maintains a delicate equilibrium.
Histone acetyltransferases facilitate the attachment of acetyl-CoA molecules onto histones, promoting a relaxed nucleosome structure.43 This structural change activates the transcriptional machinery, enhancing gene expression. In cases where HAT activity is hindered or inhibited, the repair of damaged DNA may be compromised, potentially resulting in cellular apoptosis or programmed cell death. This underscores the crucial role of HATs in DNA repair and cell survival.44 Unlike HATs, HDACs remove acetyl groups from histones, causing histones to tightly bind to negatively charged DNA. This leads to dense chromatin compaction and the inhibition of gene transcription.45
Histone acetyltransferases are classified into distinct subfamilies based on their catalytic domains structural and functional similarities, represented as HAT domains. These subfamilies include p300/CBP, HAT1, MYST, PCAF/Gcn5, and Rtt109.46 Conversely, HDACs are categorized into 4 classical classes according to their sequence similarities: Class I, Class II, Class III, and Class IV.47 The Class I, II, and IV HDACs contain a conventional deacetylase domain, while Class III HDACs feature a NAD+-dependent catalytic domain (Table 1).48
Table 1.
Histone acetyltransferase, histone deacetylase, and their role in bone metabolism.
| HATs/HDACs | Target genes | Function | Reference |
|---|---|---|---|
| P300/CBP | Runx2 | Promotes the transcriptional activity of Runx2 to promote osteogenic differentiation | 50 |
| NFATC1 | RANKL promotes osteoclast differentiation through NFATc1 acetylation | 61 | |
| PCAF | Runx2 | Promotes osteoblast differentiation | 62 |
| CXCL12 | Promotes osteogenic differentiation of MSCs | 63 | |
| NFATC1 | Promotes osteogenic differentiation of MSCs | 61 | |
| GCN5 | Wnt, NF-kB | Enhances osteogenic differentiation ability of BMSCs | 64,65 |
| HDAC1 | FoxO3a | Promotes MSC osteogenic differentiation | 53 |
| HDAC4 | HDAC4 acts as an oncogene in osteosarcoma cancer to promote cell proliferation and inhibit apoptosis and autophagy | 66 | |
| Runx2 | Deacetylates and degrades Runx2, leading to reduced osteoblast function | 67 | |
| SIRT1 | Runx2 | Promotes osteogenic differentiation of MSCs | 68 |
| FoxO3 | Reduces FoxO acetylation levels in BMMs, scavenges ROS, and inhibits bone resorption | 69,70 |
The transcriptional activity of the Runx2 gene is modulated by the acetyltransferase P300 and nicotinamide phosphoribosyltransferase (Nampt). P300 enhances osteogenic differentiation in MSCs through H3K14 acetylation, while Nampt does so in MC3T3-E1 cells via H3K9 acetylation.49,50 Other acetyltransferases, GCN5 and PCAF, which acetylate histone H3K9, are known to enhance osteogenic differentiation and bone formation by acetylating H3K9 loci in the promoters of Wnt and BMP genes. However, their levels have been found to decline significantly in mice with ovariectomy-induced osteoporosis (OVX).51,52 In contrast to acetyltransferases, 2 HDACs, HDAC1 and Sirtuin1 (SIRT1), play roles in osteogenesis. HDAC1 promotes MSC osteogenesis through the deacetylation of Forkhead box O3a (FoxO3a), while SIRT1 inhibits osteogenic differentiation in Bone Marrow Stromal Cells (BMSCs) by deacetylating JAGGED1 (JAG1).53,54
In terms of bone health, numerous studies indicate that HDACs, particularly SIRT, play a significant role in bone development. They influence processes such as bone formation, repair, and regeneration.55 SIRT1, a representative of Class III HDACs, activation induces autophagy during cellular stress, directly deacetylating autophagy-related proteins (ATG5, ATG7, LC-3) to initiate autophagy.56,57 SIRT1 also deacetylates FoxO3, a transcription factor for autophagy-related genes.58 SIRT1 plays a pivotal role in driving MSC differentiation toward osteoblasts.59,60
Histone Methylation
Histone methylation mainly happens at lysine (K) and arginine (R) residues found at histones’ N-terminal regions. In contrast to acetylation, methylation events at these sites contribute to both transcriptional activation and inhibition. For instance, trimethylation of H3K4, H3K36, and H3K79 is associated with gene activation, while trimethylation of H3K27, H3K9, and H4K20 is linked to gene repression.71,72
Histone methylation is dynamically regulated by methyltransferases and demethylases.73,74 Methylation enzymes such as Suppressor of Variegation 3-9 Homolog 1 (SUV39H1), G9a, and Enhancer of Zeste Homolog 2 (EZH2) add methyl groups, while demethylation enzymes such as Lysine-specific demethylase 1 (LSD1) and JmjC domain (JMJD)-containing proteins remove methyl groups.
Methyltransferases and demethylases play pivotal roles in regulating gene expression in osteoblasts and osteoclasts, thereby influencing the functioning of related genes. (Table 2) For instance, SUV39H1/2 methyltransferases predominantly regulate the abundance of trimethylated H3K9. Knockdown of SUV39H1 leads to a reduction in H3K9me3 levels, enhancing DNA repair capabilities and delaying cellular senescence in progeroid cells.75
Table 2.
Common histone methyltransferases and demethylases involved in histone methylation modification, as well as their target sites, genes, and their roles in bone metabolism.
| Histone methyltransferases/ demethylases | Target histone sites | Target genes | Function | Reference |
|---|---|---|---|---|
| SUV39H1 | H3K9me2/3 | Runx2 | Delays osteoblast differentiation | 83 |
| G9a | H3K9me2 | Runx2 | Regulates proliferation and differentiation of cranial bone cells | 84 |
| H3K27me1 | MMP-9 | Induces expression of osteoclastogenesis-related genes and promotes osteoclast differentiation (24). |
85 | |
| EZH2 | H3K27me3 | Wnt4, Foxo1 | Enhances both osteogenesis and osteoclastogenesis | 86 |
| H3K27me3 | Wnt1, Wnt6, Wnt10a | BMMCs hinder bone formation and promote adipogenesis. | 77,78 | |
| SETD2 | H3K36me3 | LBP | Guiding mesenchymal stem cells to commit to osteogenic destiny, diminishing their transformation into adipocytes | 79 |
| LSD1 | H3K4me1 | Runx2 | Inhibits osteoblast differentiation of C2C12 cells | 87 |
| H3K4me2 | Wnt7b, BMP2 | Inhibits osteogenic differentiation of BMSCs | 88 | |
| JMJD2B (KDM4B) | H3K9me3 | Runx2, CCND1 | Promotes osteogenic differentiation of BMSCs and maintains bone-fat balance | 89,90 |
| JMJD3 (KDM6B) | H3K27me3 | Runx2, OSX | Regulates osteoblast differentiation | 81 |
| NFATC1 | Promotes osteoclast differentiation | 80 |
Excessive EZH2 activity leads to elevated H3K27me3 levels, causing a shift in the lineage commitment of BMMSCs toward adipocytes during osteoporosis.76 Further investigations reveal significant enrichment of both EZH2 and H3K27me3 in the promoters of Wnt1, Wnt6, and Wnt10a within BMMSCs of mice subjected to ovariectomy.77,78 Notably, EZH2 reduces the enrichment of H3K27me3 on these promoters, consequently suppressing the expression of Wnt genes. Overexpressing EZH2 results in heightened H3K27me3 levels at the transcription start sites (TSS) of Runx2 and Bglap, pivotal factors initiating osteogenesis.
The histone methyltransferase SET-domain-containing 2 (SETD2) catalyzes the modification of H3K36 trimethylation. It facilitates the binding of trimethylated histones to promoters associated with lipopolysaccharide-binding protein (LBP), thus influencing the specification of adipogenic and osteogenic pathways. Disrupting SETD2 through knockout shifts the fate of mesenchymal stem cells toward adipocyte formation, impairing their potential to differentiate into osteoblasts. In mice lacking Setd2 specifically in osteoprogenitor cells, there is a notable decrease in trabecular volume and bone formation rate, accompanied by an excessive accumulation of marrow fat.79
JMJD3 functions as a demethylase targeting H3K27. JMJD3 plays a regulatory role in influencing the expression of genes related to bone health, such as nuclear factor of activated T-cells cytoplasmic 1 (NFATC1),80 Runx2, OSX, osteopontin, bone sialoprotein (BSP), and osteocalcin (OCN).79,81 In vitro experiments reveal that inhibiting LSD1 through knockdown using shRNA or pharmacological inhibitors suppresses osteoblast function and differentiation. When LSD1 activity is inhibited in vivo, it leads to a reduction in both osteoblast count and activity, consequently resulting in osteopenia. Selective elimination of LSD1 from mesenchymal cells also results in osteopenia and disturbs the structure of the growth plate.82
Non-Coding RNAs
Non-coding RNAs constitute a significant portion of the human genome, with approximately 2% consisting of protein-coding genes. The majority of the genome is made up of non-protein-coding RNAs,91,92 involving long non-coding RNAs (lncRNAs), circular RNAs (circRNAs), and small non-coding RNAs (sncRNAs). Within the category of sncRNAs, there are further subdivisions, such as microRNAs (miRNAs), piwi-interacting RNAs (piRNAs), small-interfering RNAs (siRNAs), and more. These ncRNAs play crucial roles in gene expression, cell differentiation, development, and various diseases, making them a prominent area of study in modern biology.93
Long Non-Coding RNAs
Long Non-Coding RNAs (LncRNAs) are ncRNAs that consist of more than 200 nucleotides in length. They have emerged as non-canonical regulators participating directly in various pathophysiological processes, including autophagy.94
As research advances, increasing evidence suggests that lncRNAs, as key regulators of gene expression, play pivotal roles in the proliferation, differentiation, apoptosis, and activity of osteoblasts and osteoclasts (Table 3). Disruptions in their expression patterns have been linked to numerous diseases, such as aging, cancer, metabolic disorders, and osteoporosis (OP).
Table 3.
LncRNA participates in bone metabolism by functioning as a miRNA sponge or by modulating the activity of transcription factors and signaling pathways.
| LncRNA | Bind miRNA | Target or signaling pathway | Effect on bone cell differentiation | References |
|---|---|---|---|---|
| PGC1β-OT1 | miR-148a-3p | KDM6B | Promotes osteogenic differentiation | 95 |
| OGRU | miR-320-3p | Hoxa10 protein | Promotes osteoblast differentiation | 96 |
| AK0 | — | PI3K/AKT | Regulates the proliferation of osteoblasts | 97 |
| Linc02349 | miR-25-3p and miR-33b-5p | Smad 5, Wnt 10b | Promotes osteogenic differentiation | 98 |
| KCNQ1OT1 | miR-701-3p | FGFR3 | Promotes the proliferation, migration, and survival of osteoblasts | 99 |
| LOC100506178 | miR-214-5p | BMP2 | Promotes the differentiation of hBMSCs into osteoblasts | 112 |
| TUG1 | miR-545-3p | CNR2 | Promotes the proliferation and differentiation of osteoblast precursor cells hFOB1.19 | 113 |
| GAS5 | miR-135a-3p | FoxO1 | Promotes osteoblast differentiation | 114 |
| MALAT1 | miR-30 | Runx2 | Promotes Osteoblast Differentiation of hADMSCs | 100 |
| Rhno1 | miR-6979-5p | BMP2 | Promotes osteogenic differentiation | 101 |
| MCF2L-AS1 | miR-33a | Runx2 | Stimulates osteogenic differentiation in hBMSCs | 102 |
| H19 | miR-149 | SDF-1 | Promotes osteoblast differentiation | 103 |
| MEG3 | — | BMP4 | Promotes the osteogenic differentiation of MSCs | 115 |
| H19 | miR-185-5p | IGF-1 | Promotes mineralization in osteoblasts | 116 |
| PRNCR1 | miR-211p-5p | CXCR4 | Inhibits osteogenic differentiation | 117 |
| ANCER | — | Wnt | Inhibits the osteogenic differentiation of hPLSCs | 118 |
| HOTAIR | — | Wnt/β-catenin | Inhibits osteogenic differentiation of BMSCs | 105 |
| ODIR1 | — | FBXO25/H2BK120ub/ H3K4me3/OSX Axis | Inhibits osteogenic differentiation of HUC-MSCs | 106 |
| SNHG1 | miR-101 | — | Inhibits osteogenesis differentiation | 107 |
| UCA1 | — | BPM2/Smad1/5/8 | Inhibits osteoblast proliferation and differentiation | 108 |
| MIRG | miR-1897 | NFATc1 | Promotes osteoclast production | 109 |
| NEAT1 | miR-7 | PTK2 | Increases expression of osteoclast marker genes | 110 |
| Bmncr | — | RANKL | Inhibits osteoclast differentiation | 119 |
| NRON | — | NFATC1 | Inhibits osteoclast differentiation | 111 |
Several lncRNAs have been identified as positive regulators of osteoblastogenesis, including PGC1β-OT1,95 OGRU,96 AK0,97 LINC02349,98 KCNQ10T1,99 MALAT1,100 Rhno1,101 MCF2L-AS1,102 H19,103 GAS5,104 and more. These lncRNAs promote osteogenic differentiation by various mechanisms, such as acting as sponges for miRNAs or regulating key osteogenic factors. On the other hand, some lncRNAs negatively regulate osteogenic differentiation, including HOTAIR,105 ODIR1,106 SNHG1,107 UCA1,108 and more. These lncRNAs inhibit osteogenesis by suppressing specific signaling pathways or inhibiting the expression of osteogenic marker genes.
Long non-coding RNAs also play dual roles in osteoclastogenesis. For example, MIRG has been shown to have a positive regulatory role,109 while NEAT1110, Bmncr,110 NRON,111 and others inhibit osteoclast formation by various mechanisms. Overall, lncRNAs represent a diverse class of regulatory molecules with complex roles in the regulation of osteoblasts and osteoclasts, influencing bone health and associated diseases.
MicroRNAs
MicroRNAs (miRNAs) are short, ncRNA molecules typically composed of 19 to 25 nucleotides in length. They are highly conserved across species and play a crucial role in post-transcriptional gene regulation.120 MicroRNAs do not code for proteins themselves but instead regulate gene expression by binding to complementary sequences within the target mRNA, leading to the degradation of the target mRNA or translational repression.121 The intricate interplay between miRNAs and their target mRNAs is influenced by various factors, including the strength of their interaction, target mRNA abundance, and intracellular localization of both miRNA and mRNA.122
In the context of osteogenesis and bone cell differentiation, miRNAs have several key roles: regulation of bone formation, diverse functions and mechanisms, targeting multiple genes and potential therapeutic targets (Table 4).
Table 4.
The role of miRNA in osteogenesis and bone cell differentiation.
| Function | Functional characteristics | Phenotype | Reference |
|---|---|---|---|
| Regulation of bone formation | Regulates osteogenic differentiation signaling pathways | MiR-335-5p promotes bone formation and regeneration by activating Wnt signaling and specifically downregulating the expression of DKK1 | 123 |
| MiR-196a promotes hASC osteogenic differentiation through BMP/Smad signaling pathway | 124 | ||
| MiR-210 positively regulates osteoblast differentiation through TGF-β/activin signaling pathway | 125 | ||
| Diverse functions and mechanisms | A single miRNA can serve multiple functions within bone cells, and they operate through various mechanisms | MiR-21, miR-140 and miR-214 respectively affect fracture healing through multiple mechanisms and have been identified as potential biomarkers of fracture healing. | 126 |
| Engage in complex signaling networks, including both feedforward and feedback mechanisms, to modulate the functionality of bone cells | MiR-21 regulates osteoclastogenesis through the positive feedback loop of c-Fos/miR-21/PDCD4 | 127 | |
| MiR-125a regulates osteoclast differentiation through a TRAF6/NFATc1/miR-125a negative feedback loop | 128 | ||
| Targeting multiple genes | MiRNAs have multifaceted effects on bone cell differentiation by targeting multiple genes | The miR-29 family is involved in bone cell differentiation, with individual miRNA members targeting different genes. miR-29b suppresses Secreted Frizzled-Related Protein 1 (SFRP1) and collagen expression in mature osteoblasts, while miR-29a targets Dickkopf-Related Protein 1 (DKK1) and bone sialoprotein. | 129 |
| MiR-422a is up-regulated in osteoporosis, and it simultaneously inhibits 5 genes: CBL, CD226, IGF1, PAG1, and TOB2 | 130 | ||
| Potential therapeutic targets | MiRNAs may be concentrated in cell populations associated with functions linked to specific cell phenotypes, making them potential drug targets (miR-146a, miR-29b, miR-124 possible target for drugs to treat osteoporosis) | MiR-146a inhibits TNF-a/RANKL-induced osteoclastogenesis in human PBMCs via TRAF6 | 131 |
| MiR-29b inhibits M-CSF and RANKL-induced osteoclastogenesis in CD14 hematopoietic stem cells via c-Fos | 132 | ||
| MiR-124 modulates osteoclast formation in mouse bone marrow macrophages (BMMs) by inhibiting NFATc1 expression. | 133 |
Overall, miRNAs are essential regulators of osteogenesis and bone cell differentiation, contributing to the maintenance of bone tissue and the equilibrium between bone formation and resorption. Understanding the roles and regulatory networks of miRNAs in bone biology is crucial for advancing our knowledge of bone-related diseases and potential therapeutic interventions.
Autophagy in Bone Metabolism
In mammals, three specific types of autophagy have been recognized: macroautophagy, microautophagy, and chaperone-mediated autophagy. Of these, macroautophagy is the most prevalent and is intricately intertwined with cellular physiology, biological functions, and the development of diseases within the context of bone.134 In this review, we will primarily focus on macroautophagy when referring to autophagy.
Autophagy typically progresses through four key stages: the creation of autophagosome precursors, followed by the formation of autophagosomes, then the development of autolysosomes, and finally, the degradation phase. During the degradation phase, sizable cellular molecules are disintegrated into amino acids, lipids, nucleotides, and energy, facilitating both the cell's metabolic necessities and the rejuvenation of specific organelles.135
Currently, there are more than 40 ATG genes known to regulate autophagy.136 Each gene plays distinct roles at specific stages. ULK1, a key protein, forms the ULK1 complex (ULK1-ATG13-FIP200-ATG101) initiating autophagy. During starvation, mTOR inhibition and AMPK activation prompt ULK1 phosphorylation, initiating autophagy. Beclin1, similar to yeast ATG6, interacts with proteins to form the PI3K complex, crucial for autophagy initiation. ATG14 and UVRAG play roles in autophagosome elongation and autophagosome-lysosome fusion, respectively. Proteins such as ATG12, ATG5, and LC-3 control autophagosome formation. LC3-II, a key autophagy marker, encapsulates materials for degradation. Finally, the autophagosome fuses with lysosomes to form autolysosomes, where degradation provides cells with energy against stress.137
Importance of Autophagy in Bone Homeostasis and Remodeling
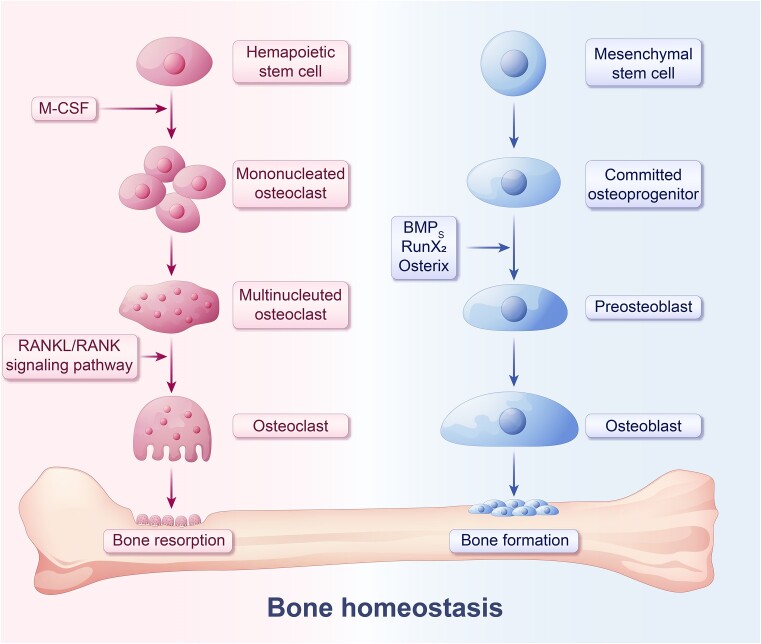
Bone homeostasis refers to the delicate balance maintained within the skeletal system through a dynamic interplay between bone-forming cells (osteoblasts), bone-resorbing cells (osteoclasts), and bone matrix.138 This intricate equilibrium is crucial for preserving the overall health, strength, and functionality of bones (Figure 1).
Figure 1.
Bone resorption and bone formation maintain the dynamic balance of bone homeostasis. Mesenchymal stem cells in the bone marrow gradually differentiate into osteoblasts, with key transcription factors such as BMPS, Runx2, and OSX playing critical roles in regulating osteoblast differentiation and bone matrix synthesis. Hematopoietic stem cells differentiate into mononucleated osteoclasts under the influence of M-CSF secreted by osteoblasts. Upon activation by RANKL-RANK signaling, these cells further differentiate into mononuclear resorbing cells and subsequently fuse into multinucleated osteoclasts.
Bone remodeling involves three cell types: MSCs differentiate into osteoblasts (OB) on the bone surface, secreting bone matrix. This matrix, with OB, transforms into osteocytes, forming a vital mechanosensory network in bones, essential for signaling; simultaneously, multinucleated osteoclasts (OC) derived from hematopoietic stem cells constantly break down and absorb the neighboring bone matrix.138,139 Generally, the balance between bone formation and resorption is continually coordinated. In this manner, the quality, structure, and function of bone tissue can be influenced by internal or external stimuli. Autophagy helps OB, OC, and chondrocytes cope with stress and nutrient deficiencies, promoting survival in harsh hypoxic and hypertonic conditions. Autophagy can also enable the long-term terminal differentiation of osteocytes. The process of autophagy encompasses not only the osteoclastic resorption process but also the acquisition of energy sources during osteoblast differentiation.140 Altered autophagy can disrupt bone cell balance, potentially causing various diseases.
Role of Autophagy in Mesenchymal Stem Cells, Osteoblasts, Chondrocytes, Osteoclasts, and Osteoclasts
Autophagy in BMSCs
As a rare and diverse subset of stromal cells, BMSCs exhibit the capacity for both self-renewal and differentiation. These cells are capable of undergoing differentiation into various lineages, not limited solely to mesenchymal lineages such as osteocytes, chondrocytes, and adipocytes,141,142 autophagy is vital in controlling the roles, differentiation, and survival of BMSCs. Modulating autophagic activity in these stem cells could have implications for enhancing bone regeneration, treating bone-related diseases, and addressing age-related bone health issues.
Autophagy-regulated redox state participates in determining the fate of BMSC differentiation. It has been reported that elevated ROS levels in BMSCs can promote adipogenesis while inhibiting osteogenic differentiation. When excessive ROS is generated within cells, the autophagic mechanism is activated to reduce ROS levels, thereby restoring osteogenic differentiation of BMSCs.143 Additionally, research has demonstrated that administering the neuropeptide substance P (SP) to rats enhances BMSC autophagic activity through the AMPK and mammalian target of rapamycin (mTOR) pathways, concurrently reducing ROS production and facilitating osteogenic differentiation.144 Rapamycin (RAPA), a well-known inhibitor of the mTOR, triggers autophagy by binding to mTOR and activating the mTOR signaling pathway. Research has indicated that RAPA enhances autophagy and influences the osteogenic differentiation of MSCs.145
Autophagy in Osteoblast
Osteoblasts are central to the processes of bone growth, repair, and remodeling. They synthesize collagen proteins and other bone matrix molecules, providing structural support to the skeleton. Osteoblasts also release bone formation-related proteins, hormones, and cytokines, such as alkaline phosphatase, osteocalcin, and others, which play significant roles in regulating bone metabolism and maintaining bone homeostasis.146
Autophagic proteins such as Beclin1, ATG5, and ATG7 play an essential role in facilitating the mineralization of osteoblast cell lines. Osteoblast autophagy deficiency can reduce its mineralization ability, leading to a low bone mass phenotype.147 Insufficient or deficient autophagy in osteoblasts leads to an increase in oxidative stress, which in turn elevates the production of TNFSF11. This further enhances the differentiation of osteoclasts, ultimately resulting in a phenotype resembling osteoporosis.148 Transcription factors FoxO and ATF4 have been extensively studied in the regulation of autophagy during the differentiation and function of osteoblasts. FoxO binds to the promoter regions of autophagy genes, increasing autophagic activity. This activation may promote the differentiation of MSCs into osteoblasts and inhibit fat generation.149 Additionally, under endoplasmic reticulum (ER) stress and amino acid deficiency conditions, ATF4 can promote autophagosome formation and autophagic flux by regulating the expression of autophagy initiation-related genes, contributing to the maintenance of osteoblast homeostasis.150
Autophagy in Osteoclasts
Osteoclasts primarily engage in bone resorption in the body and play a role in bone homeostasis. When their activity is excessively high, it can lead to osteoporosis, while conversely, decreased activity can result in increased bone formation.151 The microenvironment where osteoclasts are distributed, such as the sealing zone and the interior of bone trabeculae, is characterized by low oxygen levels, which support the survival and maturation of osteoclasts.152 Under low oxygen conditions, the expression of Hypoxia-inducible factor 1 alpha (HIF-1α) and its downstream signaling molecule BNIP3 increases, leading to elevated levels of autophagy-related proteins such as ATG5, ATG12, and Beclin1. Consequently, LC-3 is recruited to autophagosomes, enhancing the expression of bone resorption factors such as RANKL, matrix metalloproteinase K (MMP), tissue protease K, NFATc1. This, in turn, leads to increased osteoclast formation.153,154 Research has provided evidence that the targeted removal of ATG7 in osteoclast precursors in mice resulted in the improvement of bone loss and the excessive activation of osteoclasts triggered by glucocorticoids or ovariectomy.155
During the adhesion and migration of osteoclast precursor cells, chemotactic factors CXCL12 and S1P play roles in this process. Specifically, S1P binds to its receptor S1PR on the membrane of osteoclast precursor cells, participating in the migration process.156 S1P also modulates autophagy through mTOR, serving as a link between autophagy and the accumulation of osteoclasts.157 RANKL and RANK (receptor activator of NF-κB) play essential roles in osteoclast differentiation and maturation. RANKL induces autophagy activation through pathways such as MAPK and NF-κB during this process.158
Autophagy in Osteocytes
Osteocytes establish an extensive interconnected network throughout the entire skeleton. They do this through multiple branching processes that resemble dendrites, allowing them to connect with other types of bone cells such as osteoblasts, bone lining cells, and stromal cells.159 This network spans from the innermost bone regions to the blood vessel linings.160 Osteocytes play multifaceted roles: they regulate mineral metabolism and the remodeling of the perilacunar matrix, while also serving as mechanosensory cells.161
Osteocytes play a central role in regulating bone remodeling in response to mechanical loading.162 Autophagy is a vital mechanism that ensures the survival of osteocytes and enhances their capacity for mechanotransduction in the context of bone remodeling. Recent investigations have unveiled autophagy’s responsiveness to mechanical cues in osteocytes. Mechanistically induced autophagy contributes to the preservation of adenosine triphosphate (ATP) and fosters osteocyte survival. Furthermore, given the unique features of terminal differentiation, the prolonged lifespan of bone cells, and their oxygen and nutrient-deprived environment, autophagy becomes a key player in the regular physiological processes of these cells.140,163 Unsurprisingly, osteocytes exhibit a notable baseline level of autophagy both in laboratory settings and within living organisms. Selectively deleting the autophagy-related gene ATG7 in osteocytes inhibits autophagy, causing decreased bone formation and reduced bone mass in young adult mice, resembling the effects of aging on the skeletal system.164 Dysregulated autophagy in osteocytes, as seen in Ephrin B2 deficiency or triggered by substances such as pinocembrin, can affect bone health and apoptotic processes.165,166 Beyond its role as a degradative mechanism, recent evidence has spotlighted the involvement of autophagy in protein secretion, referred to as secretory autophagy, bridging intracellular autophagy with the extracellular microenvironment.167 This phenomenon may offer insights into understanding the impacts of osteocyte autophagy triggered by physical forces exerted on osteoblasts and osteoclasts.168
Autophagy in Chondrocytes
MSCs in the bone marrow differentiate into osteoprogenitor cells, which further differentiate into chondrocytes that form the cartilage primordia.169,170 Conversely, they develop into endochondral bone, determining the rate and length of longitudinal bone growth. Most skeletal growth is achieved through the ossification of cartilage in the epiphyseal growth plate of bones. Due to the low regenerative capacity of chondrocytes and limited vascularity within the growth plate, chondrocytes are prone to hypoxia and nutrient deficiency.171 It has been demonstrated that inflammatory mediators including reactive oxygen species (ROS), IL-1β, nitric oxide (NO), Fas, and tumor necrosis factor-alpha (TNF-α) are strongly associated with chondrocyte apoptosis.172
Autophagy, a cellular degradation mechanism responsible for maintaining cellular energy metabolism homeostasis, possesses the ability to restore impaired chondrocyte functionality.173 When cellular ATP levels decrease, AMPK activates and triggers autophagy, restoring nutrients and ATP, thus maintaining cellular energy balance. Autophagy also impacts protein and lipid metabolism in chondrocytes through the mTOR pathway.174
It has been demonstrated that chondrocyte degeneration and apoptosis are considered primary factors in the development of osteoarthritis (OA).175 Consequently, numerous experiments have been designed to enhance chondrocyte autophagy as a means to alleviate OA. For instance, Interventions such as vitamin D and tofacitinib exhibit potential in protecting against OA by promoting autophagy and preventing chondrocyte degeneration.176,177 Proteins such as PGRN and G protein-coupled receptor family C group 5 member B (GPRC5B) play roles in maintaining chondrocyte health through their involvement in autophagy.178,179 Moderate mechanical strain can promote the restoration of metabolic homeostasis by inhibiting inflammation and excessive autophagy.
Epigenetic Regulation of Autophagy in Bone Metabolism
Epigenetic modifications, alterations in gene expression without DNA sequence changes,11 intersect significantly with autophagy, a process vital for cellular stress response and recycling. This intricate relationship profoundly influences bone metabolism and homeostasis.
Epigenetic Modifications and Autophagy
DNA methylation, a heritable epigenetic modification, impacts autophagy regulation in diverse cancers and stem cells.173 Hypermethylation of LC-3A174 and Beclin1175 inhibits autophagy, promoting tumorigenesis. Conversely, demethylation enhances autophagy, showing therapeutic potential in cancer treatments.176,178,179 Additionally, in conditions like osteoporosis and osteoarthritis, abnormal DNA methylation affects autophagy, highlighting its role in bone health.180
Histone modifications such as methylation and acetylation regulate autophagy-related genes. G9a-mediated histone methylation inhibits autophagy, while its dissociation activates this process, crucial in non-alcoholic fatty liver disease.181 Enzymes such as JMJD3182,183 and SIRT184,185 family members modulate histone demethylation and deacetylation, respectively, impacting autophagy regulation in cellular processes.
Non-coding RNAs, including miRNAs and lncRNAs, regulate autophagy at different stages in various diseases.186 MiRNAs influence autophagy-related genes, affecting cancer cell survival and response to treatments.187–190 Long non-coding RNAs, such as H19191,192 and GAS5,193,194 exhibit both positive and negative regulatory effects on autophagy, impacting bone-related disorders such as osteoarthritis.
Epigenetic Regulation of Autophagy in Bone Metabolism-Related Diseases
Epigenetic mechanisms are crucial in regulating autophagy processes, influencing its transcriptional and post-translational regulation.195,196 These epigenetic mechanisms can be further influenced by external stimuli, alterations in phenotypic states, or pathological environmental conditions.197 Epigenetic modifications and autophagy interact in bone metabolism, regulating gene expression, cellular clearance, and stress responses, crucial for maintaining bone tissue homeostasis and function. The convergence of genetic and environmental factors, causing disruption in the epigenetic regulation of autophagy, has the potential to detrimentally affect bone metabolism, thereby contributing to the onset and advancement of bone-related disorders (Figure 2).
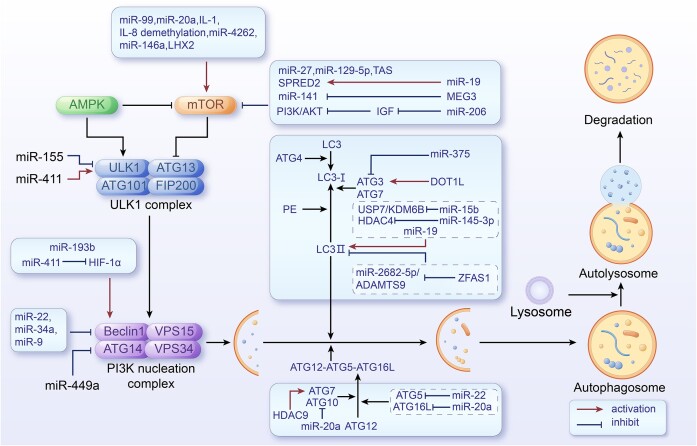
Figure 2.
Under the influence of genetic or pathogenic environmental conditions, histone modifications and ncRNAs promote or inhibit autophagy by targeting autophagy-related genes and proteins. Disordered autophagy leads to the occurrence and progression of bone metabolism-related diseases.
Osteoporosis
Osteoporosis is a degenerative bone disorder marked by reduced bone mineral density (BMD), degraded bone microstructure, increased bone fragility, and a higher susceptibility to fractures.198,199 Fundamentally, it arises from an imbalance where bone resorption surpasses bone formation, resulting in a loss of bone tissue.200 As the population ages, the rates of osteoporosis-related fragility fractures, disability, and mortality are increasing year by year.201
Numerous studies have confirmed that autophagy increases osteoclast formation under in vitro oxidative stress, low oxygen conditions, and microgravity conditions, leading to an imbalance where bone resorption surpasses bone formation, ultimately triggering osteoporosis (Figure 3).202,203 Several studies have demonstrated that miRNAs play a role in regulating bone resorption and bone formation in the process of osteoporosis formation by mediating autophagy-related factors (Figure 2). Research on RAW 264.7 cells under low oxygen conditions revealed that miR-20a directly targets the 3'UTR of ATG16L1, suppressing autophagy by reducing the levels of autophagy-related proteins LC-3 and ATG16L1, which is favorable for osteoclast differentiation. Furthermore, during osteoclast differentiation induced by hypoxia, HIF-1α can regulate miR-20a, and the HIF-1α-miR20a-ATG16L1 axis plays a significant role.204 MiR-99 may play a role in fine-tuning and integrating the mTOR signaling pathway to promote optimal osteoclast differentiation.205,206 Histone demethylases KDM4B and KDM6B play a crucial role in osteogenic commitment of MSCs by removing H3K9me3 and H3K27me3 marks.207 Upregulation of USP7 enhances osteogenic differentiation in human adipose-derived stem cells (hASCs) to suppress osteogenesis progression.208 USP7 promotes the expression of KDM6B by enhancing its stability.90 Elevated levels of miR-15b in osteoporosis inhibit the USP7/KDM6B axis, thereby suppressing osteoblast differentiation and autophagy, exacerbating osteoporosis.209 DOT1L, as a histone methyltransferase, can methylate H3k79. Experiments by Gao et al. confirmed that inhibiting DOT1L in vitro increased autophagosome assembly proteins (ATG3 and ATG8-like Gabarapl2) and autophagy receptors (Sqstm1), activated autophagy, increased pre-osteoclast migration, and increased bone density. Absorption increases and osteoporosis occurs.210 A research report suggests that age-related upregulation of HDAC9 accelerates bone loss in mice by promoting damaged-induced autophagy.211 However, administering an HDAC9 inhibitor to elderly mice can restore mesenchymal osteoblastic function and recover bone mass.212
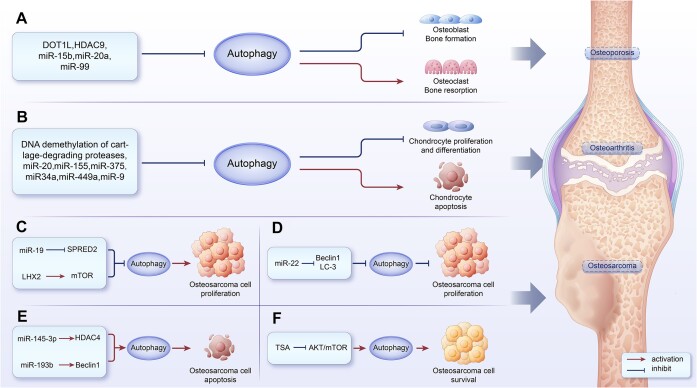
Figure 3.
Epigenetic modifications regulate autophagy in bone metabolic diseases. (A) Histone modification and miRNA's inhibition of autophagy results in enhanced osteoclast generation and impedes osteoblast differentiation, causing increased bone resorption relative to bone formation, ultimately leading to osteoporosis. (B) DNA methylation modification and miRNA's inhibition of autophagy lead to increased chondrocyte apoptosis, hindered chondrocyte proliferation, and differentiation, leading to the onset of osteoarthritis. (C-F), Autophagy demonstrates a dual role in osteosarcoma. Its inhibition can either promote or impede the proliferation of osteosarcoma cells. Simultaneously, inducing autophagy can trigger cell apoptosis or elevate the survival rate of osteosarcoma cells.
Osteoarthritis
Osteoarthritis is a degenerative condition characterized by cartilage degeneration, subchondral bone remodeling, and synovial inflammation. The development of OA is typically associated with factors such as joint stress, genetic predisposition, obesity, aging, and metabolic or hormonal changes.213 Additionally, inflammatory mediators such as interleukin (IL)-1, IL-6, and TNF are excessively produced in chondrocytes and matrix cells.214 Excessive accumulation of ROS resulting from oxidative stress leads to cartilage degradation.215
In the early stages of human OA, the level of autophagy increases, contributing to the maintenance of chondrocyte homeostasis.216 However, in late-stage OA, when chondrocytes endure prolonged stress, autophagy weakens, leading to further deterioration of cartilage (Figure 3).217
Disruption of normal age-related epigenetic patterns could contribute to age-related conditions such as OA.218 Aberrant gene activation in osteoarthritis may be associated with epigenetic derepression that leads to inflammation and catabolic metabolic phenotypes in chondrocytes. Proinflammatory cytokines decrease methylation at crucial CpG sites in the IL-1β promoter, resulting in prolonged induction of this cytokine.219 Stimulation of human chondrocytes with IL-1β leads to decreased methylation at CpG sites in the IL-8 promoter and notably triggers this chemokine in chondrocytes affected by OA.220 In the study investigating the relationship between autophagy in rat articular chondrocytes and the PI3K/AKT/mTOR signaling pathway in OA, it was found that inflammation can inhibit the proliferation of rat chondrocytes, disrupt the cell cycle, and reduce the rate of autophagy.221 Furthermore, DNA demethylation is linked to the upregulation of essential cartilage-degrading proteases such as MMP-3, MMP-9, MMP-13, and ADAMTS-4, as well as iNOS.222 Multiple epigenetic modifications have been found to cooperate in regulating gene expression within the context of osteoarthritic lesions. For example, reduced SOX9 expression in hip OA is likely due to a combination of factors, including elevated DNA methylation, heightened gene-inactivating histone marks (H3K9 and H3K27) methylation, and decreased histone acetylation at the SOX9 promoter. Furthermore, miRNA-145 has also been confirmed to inhibit the expression of SOX9.223
Multiple experiments have also confirmed that miRNAs play a significant role in regulating autophagy in the development of OA (Figure 2). MiR-20 is a member of the miR-17-92 cluster located on the chromosome.224 In OA, the expression of miR-20 is increased. It can target ATG10 through the PI3K/AKT/mTOR signaling pathway, leading to its reduction, thereby inhibiting chondrocyte proliferation and autophagy.225 In the cartilage tissue of OA, miR-375 is found to be overexpressed. It can target the ATG2B–3′ UTR and inhibit its expression in chondrocytes, suppressing autophagy and promoting endoplasmic reticulum stress (ERs), thereby exacerbating cartilage damage.226 The experiment indicated that in chondrocytes treated with IL-1β, miR-27a expression is upregulated. It targets the 3′-UTR of the PI3K gene, leading to its downregulation. Through the PI3K-AKT-mTOR pathway, miR-27a activates autophagy and inhibits the proliferation of chondrocytes treated with IL-1β, providing a protective effect against OA.227 In addition, multiple experiments have confirmed that miR-155,228 miR-34a,229 and miR-449a230 inhibit chondrocyte autophagy, aggravate chondrocyte apoptosis and cause osteoarthritis.
Osteosarcoma
Osteosarcoma is a malignant tumor that originates from mesenchymal tissue, typically found at the metaphysis of long bones such as the femur, tibia, and humerus. Although it primarily affects children and adolescents, it can also occur in adults.231 The standard treatment for osteosarcoma involves a combination of preoperative and postoperative chemotherapy along with surgical tumor removal, which may even necessitate amputation in severe cases.66 Despite aggressive treatment, the prognosis for osteosarcoma remains relatively poor, with a 5-year survival rate of 60%-70% post-surgery. Metastasis, particularly to the lungs, significantly contributes to this unfavorable prognosis, with roughly 20% of patients experiencing metastasis and a subsequent 5-year survival rate of only 30%.232
Autophagy plays a dual role at different stages of osteosarcoma (Figure 3). In one study, miR-22 was shown to reduce the expression of Beclin1, LC-3, MTDH, and ATG5 mRNA by targeting MTDH, thereby inhibiting autophagy and suppressing osteosarcoma cell proliferation.233 Conversely, inhibiting autophagy was found to promote osteosarcoma cell proliferation in another experiment. In this case, miR-19 was upregulated in osteosarcoma cells, targeting SPRED2 and reducing its expression, which suppressed autophagy, ultimately promoting the proliferation and malignant transformation of osteosarcoma cells.234
Epigenetic regulation plays a significant role in autophagy within the context of osteosarcoma (Figure 2). Trichostatin A (TSA), a histone deacetylase inhibitor (HDACi), induces autophagy in osteosarcoma cells by suppressing the AKT-mTOR signaling pathway and activating FoxO1, thereby enhancing the survival of osteosarcoma cells. However, inhibiting autophagy significantly enhances TSA-induced cell death in osteosarcoma.235 Additionally, miR-145-3p, which is downregulated in human osteosarcoma cell lines, targets the 3′-UTR region of HDAC4, increasing HDAC4 levels. This, in turn, promotes apoptosis and autophagy in osteosarcoma cells.236 Another study involving osteosarcoma cells showed that LHX2 overexpression upregulated mTOR expression, which negatively regulated autophagy through the activation of the mTOR pathway, contributing to the progression of osteosarcoma. MiR-129-5p directly targeted LHX2 3′-UTR to downregulate LHX2, making the miR-129-5p/LHX2/mTOR axis a potential target for osteosarcoma treatment.237 Additionally, DANCR is a lncRNA that acts as a ceRNA by sequestering miR-335-5p and miR-1927 in osteosarcoma, promoting ROCK1-mediated proliferation and metastasis.238 Another study on osteosarcoma reveals that miR-193b directly targets the 3′-UTR of FEN1, negatively regulating the expression of FEN1, increasing the expression of Beclin 1 and the LC3-II/I ratio, activating autophagy, and inducing cell apoptosis.239
Rheumatoid Arthritis
Rheumatoid Arthritis (RA) is a chronic autoimmune inflammatory disorder marked by synovial hyperplasia, persistent inflammation, cartilage degradation, and bone erosion.240–242 This condition leads to joint deformities and is characterized by swelling and pain. As RA progresses, it can also affect organs and systems outside the joints, such as the heart.243 The treatment of RA typically involves a comprehensive approach, including medication, physical therapy, and lifestyle management, aimed at alleviating symptoms, controlling inflammation, and maintaining the patient’s quality of life.244 Early diagnosis and treatment are essential for managing the condition and preventing joint damage.245
Epigenetic mechanisms, such as DNA demethylation and hypomethylation, play significant roles in altering DNA methylation patterns during B cell to plasma cell differentiation and in RA. These alterations impact disease progression and the expression of key genes.246,247 Histone modifications are also involved in RA, with histone H3 in the promoter region of the IL-6 gene being highly acetylated in fibroblast-like synoviocytes (FLS) from RA patients, leading to increased IL-6 expression and disease progression. Inhibitors of HAT, such as curcumin, can reduce IL-6 secretion.248
Autophagy in RA has a dual role, exerting both positive and negative effects, depending on specific cellular and molecular regulatory mechanisms.249,250 Autophagy plays a crucial role in osteoclast differentiation and maturation under hypoxic conditions, leading to increased bone resorption and accelerated progression of RA.251 Additionally, conditions such as nutrient deficiency and endoplasmic reticulum stress promote autophagy, which acts as a self-protective mechanism, allowing RA cells to evade apoptosis and prolong their lifespan.252
Research by Li et al. demonstrated that the lncRNA MEG3 is upregulated in synovial tissues of RA patients. MEG3 targets miR-141, and their expression is negatively correlated. By inhibiting the AKT/mTOR pathway and activating autophagy, MEG3 suppresses inflammation, promotes chondrocyte proliferation, and inhibits RA progression.253 In experiments by Zhou et al. using a CIA rat model, it was confirmed that treatment with WJR (Wenhua Juanbi Recipe) may inhibit autophagy by affecting the PI3K/AKT/mTOR pathway mediated by miRNA-146a. This inhibition of autophagy leads to the suppression of cell apoptosis and FLS proliferation.254 Moreover, in a study by Yang et al., it was found that ZFAS1 (lncRNA ZNFX1 antisense RNA) plays a regulatory role in FLS-RA through the miR-2682-5p/ADAMTS9 axis. Knockdown of ZFAS1 significantly inhibits FLS-RA cell proliferation, inflammatory response, autophagy, and promotes cell apoptosis (Figure 2).255
Future Directions and Challenges
Currently, the group of epigenetic drugs includes256,257 DNMTis (such as decitabine used in the treatment of AML and high-risk myelodysplastic syndromes,258 and 5-azacitidine used in high-risk myelodysplastic syndromes259), HDAC inhibitors (like TSA mentioned earlier, activating autophagy in osteosarcoma235), HAT inhibitors (curcumin can reduce IL-6 secretion in RA248), histone methyltransferase inhibitors (BIX01294 inhibits G9a in multiple myeloma260), and various miRNA-based molecular targeted therapy drugs. Despite their role in disease treatment, these epigenetic drugs lack specificity in target therapy and often exhibit toxicity. Future research directions in the epigenetic regulation of autophagy for bone metabolism-related diseases aim to minimize these drugs’ side effects and identify specific epigenetic modifications involved in autophagy for bone metabolism-related diseases.
The currently well-researched histone modifications in bone metabolism-related diseases include histone methylation and acetylation. Other histone modifications, such as histone phosphorylation,261 ADP-ribosylation,262 ubiquitination,263 SUMOylation,264 glutamylation,265 glycosylation,266 hydroxylation,267 and isomerization,268 are also covalent modifications that potentially regulate gene expression by altering chromatin structural states and functions, or affecting the affinity between transcription factors and gene promoters. Despite extensive research in fields such as tumorigenesis, energy metabolism, and cellular aging, limited reporting exists on their role in bone metabolism-related diseases. Future comprehensive investigations into the regulation of these histone modifications in autophagy concerning bone metabolism-related diseases might unveil new targets for preventing and treating these conditions and guide the development of novel epigenetic drugs.
Research on miRNA regulation of autophagy in bone metabolism-related diseases is burgeoning and provides potential targets for treating these conditions. A crucial aspect in precise treatment for bone-related diseases involves transferring target miRNAs or anti-miRNAs to target cells without being degraded by endogenous RNA or causing off-target effects. Presently, experimental delivery systems include aptamers, single-stranded DNA or RNA,269,270 8-repeat aspartic acid sequences (D-Asp8),271 and bacteriophage MS2 virus-like particles (MS2 VLPs).272 Future advancements should emphasize developing delivery systems that enhance miRNA stability and cellular uptake efficiency. MiRNAs, capable of mimicking or inhibiting the expression of target genes, have the potential to regulate bone biology processes such as bone formation, resorption, and regeneration by activating or inhibiting autophagy.
Conclusion
In this paper, we delved into the critical roles of genetic regulation and autophagy in bone metabolism and highlighted key insights relevant to bone health. Through the culmination of our research, we draw the following conclusions:
Firstly, genetic regulation and autophagy play pivotal roles in bone metabolism. Genetic regulation, by modulating gene expression, determines the differentiation, proliferation, and function of bone cells. Autophagy, on the other hand, is a self-regulating cellular process indispensable for clearing aged or damaged cells, maintaining intracellular homeostasis, and facilitating bone tissue repair. These two mechanisms intricately intertwine and cooperate to uphold bone health.
Secondly, research indicates that abnormalities in genetic regulation and autophagy are closely associated with the onset and progression of various bone metabolic disorders and bone-related diseases. Not only do genetic mutations lead to the hereditary transmission of certain bone diseases, but environmental factors, lifestyles, and aging impact the functionality of autophagy, thereby exerting adverse effects on bone health. Consequently, understanding the roles and dysregulation of these regulatory mechanisms is paramount for preventing, diagnosing, and treating bone disorders.
Lastly, we emphasize the significance of further research in propelling advancements in the field of bone health. By delving deeper into the molecular mechanisms of epigenetic regulation and autophagy, we can identify novel therapeutic targets and develop more effective treatment strategies. Simultaneously, with the aid of precision medicine and personalized treatment approaches, we can better cater to the diverse needs of individual patients, elevating the management of bone diseases.
In conclusion, this paper underscores the importance of genetic regulation and autophagy in maintaining bone health and their close connections with bone metabolic disorders and related diseases. We encourage future research to continue delving into these domains, with the aim of providing innovative solutions for preventing and treating bone disorders, ultimately enhancing the quality of life and health of patients. Through collaboration and exploration of new frontiers, we can collectively drive progress in the field of bone health, paving the way for new possibilities in future medical and clinical practices.
Acknowledgement
We express our gratitude to Jiang Pei, Director of the Translational Pharmacy Laboratory at Jining First People’s Hospital, for providing valuable guidance and support throughout this study. The graphical abstract was created using BioRender.
Contributor Information
Yazhou Zhang, Department of Foot and Ankle Surgery, Tengzhou Central People’s Hospital, Tengzhou 277500, China.
Qianqian Wang, Department of Pediatric Intensive Care Unit, Tengzhou Central People’s Hospital, Tengzhou 277500, China.
Hongjia Xue, Department of Computer Science, University College London, London, WC1E 6BT, UK.
Yujin Guo, Institute of Clinical Pharmacy & Pharmacology, Jining First People’s Hospital, Jining 272000, China.
Shanshan Wei, Department of Pharmacy, Shandong Provincial Hospital Affiliated to Shandong First Medical University, Jinan 250000, China; Department of Graduate, Shandong Academy of Medical Sciences, Shandong First Medical University, Jinan 250000, China.
Fengfeng Li, Department of Neurosurgery, Tengzhou Central People’s Hospital, Tengzhou 277500, China.
Linqiang Gong, Department of Gastroenterology, Tengzhou Central People's Hospital, Tengzhou 277500, China.
Weiliang Pan, Department of Foot and Ankle Surgery, Tengzhou Central People’s Hospital, Tengzhou 277500, China.
Pei Jiang, Translational Pharmaceutical Laboratory, Jining First People’s Hospital, Shandong First Medical University, Jining 272000, China; Institute of Translational Pharmacy, Jining Medical Research Academy, Jining 272000, China.
Author Contributions
Y.Z. wrote and edited the manuscript. Q.W. conceived the manuscript. H.X., Y.G., and S.W. revised and edited the manuscrip. F.L. and L.G. designed and provide the graphic image. W.P. and P.J. provided significant assistance. All authors have read and approved the final manuscript. All authors edited and approved the final manuscript.
Funding
The study was supported by the National Natural Science Foundation of China [81602846; 82272253] and the Natural Science Foundation of Shandong Province [ZR2021MH145, ZR2020MH375], the Traditional Chinese Medicine Science and Technology Development Plan of Shandong Province [M-2022066],China International Medical Foundation [No. Z-2018-35-2002].
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Data Availability
The data involved in this review article are available from the corresponding author on reasonable request.
References
- 1. Stieglitz J, Beheim BA, Trumble BC, Madimenos FC, Kaplan H, Gurven M. Low mineral density of a weight-bearing bone among adult women in a high fertility population. American J Phys Anthropol. 2015;156(4):637–648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Desai S, Jayasuriya CT. Implementation of endogenous and exogenous mesenchymal progenitor cells for skeletal tissue regeneration and repair. Bioeng. 2020;7(3):86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Sheehy EJ, Kelly DJ, O'Brien FJ. Biomaterial-based endochondral bone regeneration: a shift from traditional tissue engineering paradigms to developmentally inspired strategies. Materials Today Bio. 2019;3:100009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Omata Y, Okada H, Uebe S et al. Interspecies single-cell RNA-seq analysis reveals the novel trajectory of osteoclast differentiation and therapeutic targets. JBMR Plus. 2022;6(7):e10631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Valdor R, Macian F. Autophagy and the regulation of the immune response. Pharmacol Res. 2012;66(6):475–483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Conciatori F, Bazzichetto C, Falcone I et al. Role of mTOR signaling in tumor microenvironment: an overview. Int J Mol Sci. 2018;19(8):2453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Gomes LR, Menck CFM, Leandro GS. Autophagy roles in the modulation of DNA repair pathways. Int J Mol Sci. 2017;18(11):2351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Ghavami S, Zamani M, Ahmadi M et al. Epigenetic regulation of autophagy in gastrointestinal cancers. Biochimica et Biophysica Acta (BBA)—Molecular Basis of Disease. 2022;1868(11):166512. [DOI] [PubMed] [Google Scholar]
- 9. Yu YS, Shin HR, Kim D et al. Pontin arginine methylation by CARM1 is crucial for epigenetic regulation of autophagy. Nat Commun. 2020;11(1):6297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Yu YS, Kim H, Kim KI, Baek SH. Epigenetic regulation of autophagy by histone-modifying enzymes under nutrient stress. Cell Death Differ. 2023;30(6):1430–1436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Liu MM, Chan CC, Tuo J. Epigenetics in ocular diseases. CG. 2013;14(3):166–172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Castro-Muñoz LJ, Ulloa EV, Sahlgren C et al. Modulating epigenetic modifications for cancer therapy (Review). Oncol Rep. 2023;49(3):doi: 10.3892/or.2023.8496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Loughlin J. Genetic contribution to osteoarthritis development: current state of evidence. Curr Opin Rheumatol. 2015;27(3):284–288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Jeffries MA. Epigenetic editing: how cutting-edge targeted epigenetic modification might provide novel avenues for autoimmune disease therapy. Clin Immunol. 2018;196:49–58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Núñez-Carro C, Blanco-Blanco M, Villagrán-Andrade KM, Blanco FJ, de Andrés MC. Epigenetics as a Therapeutic Target in Osteoarthritis. Pharmaceuticals. 2023;16(2):156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Reddy MA, Park JT, Natarajan R. Epigenetic modifications and diabetic nephropathy. Kidney Res Clinical Practice. 2012;31(3):139–150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Furey TS, Sethupathy P, Sheikh SZ. Redefining the IBDs using genome-scale molecular phenotyping. Nat Rev Gastroenterol Hepatol. 2019;16(5):296–311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Olsson AH, Volkov P, Bacos K et al. Genome-wide associations between genetic and epigenetic variation influence mRNA expression and insulin secretion in human pancreatic islets. PLoS Genet. 2014;10(11):e1004735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. de Melo FHM, Oliveira JS, Sartorelli VOB, Montor WR. Cancer chemoprevention: classic and epigenetic mechanisms inhibiting tumorigenesis. what have we learned so far?. Front Oncol. 2018;8:644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Rodenhiser D, Mann M. Epigenetics and human disease: translating basic biology into clinical applications. Can Med Assoc J. 2006;174(3):341–348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Marini F, Cianferotti L, Brandi ML. Epigenetic mechanisms in bone biology and osteoporosis: can they drive therapeutic choices?. Int J Mol Sci. 2016;17(8):1329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Tang J, Xiong Y, Zhou HH, Chen XP. DNA methylation and personalized medicine. J Clin Pharm Ther. 2014;39(6):621–627. [DOI] [PubMed] [Google Scholar]
- 23. Curtis EM, Fuggle NR, Cooper C, Harvey NC. Epigenetic regulation of bone mass. Best Practice Res Clinical Endocrinol Metabolism. 2022;36(2):101612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Ma J, Joehanes R, Liu C et al. Elucidating the genetic architecture of DNA methylation to identify promising molecular mechanisms of disease. Sci Rep. 15 2022;12(1):19564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Zhang X, Hu M, Lyu X, Li C, Thannickal VJ, Sanders YY. DNA methylation regulated gene expression in organ fibrosis. Biochimica et Biophysica Acta (BBA)—Molecular Basis Disease. 2017;1863(9):2389–2397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Clabaut A, Grare C, Rolland-Valognes G et al. Adipocyte-induced transdifferentiation of osteoblasts and its potential role in age-related bone loss. PLoS One. 2021;16(1):e0245014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Loren P, Saavedra N, Saavedra K, Zambrano T, Moriel P, Salazar LA. Epigenetic Mechanisms Involved in Cisplatin-Induced Nephrotoxicity: an Update. Pharmaceuticals. 2021;14(6):491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Chen ZX, Riggs AD. DNA methylation and demethylation in mammals. J Biol Chem. 2011;286(21):18347–18353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Oton-Gonzalez L, Mazziotta C, Iaquinta MR et al. Genetics and epigenetics of bone remodeling and metabolic bone diseases. Int J Mol Sci. 2022;23(3):1500, [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Niu Q, Gao J, Wang L, Liu J, Zhang L. Regulation of differentiation and generation of osteoclasts in rheumatoid arthritis. Front Immunol. 2022;13:1034050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Park-Min KH. Epigenetic regulation of bone cells. Connect Tissue Res. 2017;58(1):76–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Liu H, Liu Z, Du J et al. Thymidine phosphorylase exerts complex effects on bone resorption and formation in myeloma. Sci Transl Med. 2016;8(353):353ra113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Li L, Qu Y, Jin X et al. Protective effect of salidroside against bone loss via hypoxia-inducible factor-1α pathway-induced angiogenesis. Sci Rep. 2016;6:(1):32131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Farshdousti Hagh M, Noruzinia M, Mortazavi Y et al. Different Methylation Patterns of RUNX2, OSX, DLX5 and BSP in Osteoblastic Differentiation of Mesenchymal Stem Cells. Cell J. Spring. 2015;17(1):71–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Wang L, You X, Ruan D et al. TET enzymes regulate skeletal development through increasing chromatin accessibility of RUNX2 target genes. Nat Commun. 2022;13(1):4709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Franklin S, Vondriska TM. Genomes, proteomes, and the central dogma. Circ Cardiovasc Genet. 2011;4(5):576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Lelli KM, Slattery M, Mann RS. Disentangling the many layers of eukaryotic transcriptional regulation. Annu Rev Genet. 2012;46:(1):43–68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Hajheidari M, Koncz C, Bucher M. Chromatin evolution-key innovations underpinning morphological complexity. Front Plant Sci. 2019;10:454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Clapier CR, Cairns BR. The biology of chromatin remodeling complexes. Annu Rev Biochem. 2009;78:(1):273–304. [DOI] [PubMed] [Google Scholar]
- 40. Portela A, Esteller M. Epigenetic modifications and human disease. Nat Biotechnol. 2010;28(10):1057–1068. [DOI] [PubMed] [Google Scholar]
- 41. Li CJ, Li RW. Bioinformatic Dissecting of TP53 Regulation Pathway Underlying Butyrate-induced Histone Modification in Epigenetic Regulation. Genet Epigenet. 2014;6:1–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Mittal R, Bencie N, Liu G et al. Recent advancements in understanding the role of epigenetics in the auditory system. Gene. 2020;761:144996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Liu R, Wu J, Guo H et al. Post-translational modifications of histones: mechanisms, biological functions, and therapeutic targets. MedComm. 2023;4(3):e292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Arora I, Sharma M, Tollefsbol TO. Combinatorial epigenetics impact of polyphenols and phytochemicals in cancer prevention and therapy. Int J Mol Sci. 2019;20(18):4567, [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Tong W, Zhang L. Fetal hypoxia and programming of matrix metalloproteinases. Drug Discovery Today. 2012;17(3-4):124–134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Marmorstein R, Zhou MM. Writers and readers of histone acetylation: structure, mechanism, and inhibition. Cold Spring Harb Perspect Biol. 2014;6(7):a018762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Seto E, Yoshida M. Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harb Perspect Biol. 2014;6(4):a018713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Wong JK, Zou H. Reshaping the chromatin landscape after spinal cord injury. Front Biol. 2014;9(5):356–366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Ling M, Huang P, Islam S et al. Epigenetic regulation of Runx2 transcription and osteoblast differentiation by nicotinamide phosphoribosyltransferase. Cell Biosci. 2017;7(1):27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Tang Z, Xu T, Li Y, Fei W, Yang G, Hong Y. Inhibition of CRY2 by STAT3/miRNA-7-5p Promotes Osteoblast Differentiation through Upregulation of CLOCK/BMAL1/P300 Expression. Molecular Therapy—Nucleic Acids. 2020;19:865–876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Jing H, Su X, Gao B et al. Epigenetic inhibition of Wnt pathway suppresses osteogenic differentiation of BMSCs during osteoporosis. Cell Death Dis. 2018;9(2):176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Zhang P, Liu Y, Jin C et al. Histone H3K9 acetyltransferase PCAF is essential for osteogenic differentiation through bone morphogenetic protein signaling and may be involved in osteoporosis. Stem Cells. 2016;34(9):2332–2341. [DOI] [PubMed] [Google Scholar]
- 53. Lin CH, Li NT, Cheng HS, Yen ML. Oxidative stress induces imbalance of adipogenic/osteoblastic lineage commitment in mesenchymal stem cells through decreasing SIRT1 functions. J Cellular Molecular Medi. 2018;22(2):786–796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Wang J, Wang CD, Zhang N et al. Mechanical stimulation orchestrates the osteogenic differentiation of human bone marrow stromal cells by regulating HDAC1. Cell Death Dis. 2016;7(5):e2221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Tian Q, Gao S, Zhou X, Zheng L, Zhou Y. Histone acetylation in the epigenetic regulation of bone metabolism and related diseases. Stem Cells International. 2021;2021:1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Orogo AM, Gustafsson ÅB. Therapeutic targeting of autophagy: potential and concerns in treating cardiovascular disease. Circ Res. 2015;116(3):489–503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Kume S, Koya D. Autophagy: a novel therapeutic target for diabetic nephropathy. Diabetes Metab J. 2015;39(6):451–460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Rajendran R, Garva R, Krstic-Demonacos M, Demonacos C. Sirtuins: molecular traffic lights in the crossroad of oxidative stress, chromatin remodeling, and transcription. J Biomed Biotechnol. 2011;2011:1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Ou X, Lee MR, Huang X, Messina-Graham S, Broxmeyer HE. SIRT1 positively regulates autophagy and mitochondria function in embryonic stem cells under oxidative stress. Stem Cells. 2014;32(5):1183–1194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Yang G, Collins JM, Rafiee R et al. SIRT1 gene SNP rs932658 is associated with medication-related osteonecrosis of the jaw. J Bone Mineral Res. 2021;36(2):347–356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Kim JH, Kim K, Youn BU et al. RANKL induces NFATc1 acetylation and stability via histone acetyltransferases during osteoclast differentiation. Biochem J. 2011;436(2):253–262. [DOI] [PubMed] [Google Scholar]
- 62. Wang CY, Yang SF, Wang Z et al. PCAF acetylates Runx2 and promotes osteoblast differentiation. J Bone Miner Metab. 2013;31(4):381–389. [DOI] [PubMed] [Google Scholar]
- 63. Lian WS, Ko JY, Chen YS et al. MicroRNA-29a represses osteoclast formation and protects against osteoporosis by regulating PCAF-mediated RANKL and CXCL12. Cell Death Dis. 2019;10(10):705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Liu Y, Cheng W, Zhao Y et al. Cyclic mechanical strain regulates osteoblastic differentiation of mesenchymal stem cells on TiO(2) nanotubes through GCN5 and Wnt/β-Catenin. Front Bioeng Biotechnol. 2021;9:735949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Lu W, Zhang L, Ji K, Ding L, Wu G. Regulatory mechanisms of GCN5 in osteogenic differentiation of MSCs in periodontitis. Clinical & Exp Dental Res. 2023;9(3):464–471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Misaghi A, Goldin A, Awad M, Kulidjian AA. Osteosarcoma: a comprehensive review. SICOT-J. 2018;4:12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Li J, Liu C, Li Y et al. TMCO1-mediated Ca(2+) leak underlies osteoblast functions via CaMKII signaling. Nat Commun. 2019;10(1):1589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Shakibaei M, Shayan P, Busch F et al. Resveratrol mediated modulation of Sirt-1/Runx2 promotes osteogenic differentiation of mesenchymal stem cells: potential role of Runx2 deacetylation. PLoS One. 2012;7(4):e35712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Kim HN, Han L, Iyer S et al. Sirtuin1 suppresses osteoclastogenesis by deacetylating FoxOs. Mol Endocrinol. 2015;29(10):1498–1509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Yan S, Miao L, Lu Y, Wang L. Sirtuin 1 inhibits TNF-α-mediated osteoclastogenesis of bone marrow-derived macrophages through both ROS generation and TRPV1 activation. Mol Cell Biochem. 2019;455(1-2):135–145. [DOI] [PubMed] [Google Scholar]
- 71. Alelú-Paz R, Ashour N, González-Corpas A, Ropero S. DNA methylation, histone modifications, and signal transduction pathways: a close relationship in malignant gliomas pathophysiology. J Signal Transduct. 2012;2012:1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Lu C, Thompson CB. Metabolic regulation of epigenetics. Cell Metab. 2012;16(1):9–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Ge K. Epigenetic regulation of adipogenesis by histone methylation. Biochimica et Biophysica Acta (BBA)—Gene Regulatory Mechanisms. 2012;1819(7):727–732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Wei Y, Gañán-Gómez I, Salazar-Dimicoli S, McCay SL, Garcia-Manero G. Histone methylation in myelodysplastic syndromes. Epigenomics. 2011;3(2):193–205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Sidler C, Kovalchuk O, Kovalchuk I. Epigenetic regulation of cellular senescence and aging. Front Genet. 2017;8:138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Basta MD, Petruk S, Mazo A, Walker JL. Fibrosis-the tale of H3K27 histone methyltransferases and demethylases. Front Cell Dev Biol. 2023;11:1193344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Sun Y, Zhang H, Qiu T, Liao L, Su X. Epigenetic regulation of mesenchymal stem cell aging through histone modifications. Genes Diseases. 2023;10(6):2443–2456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Cakouros D, Gronthos S. Epigenetic Regulators of Mesenchymal Stem/Stromal Cell Lineage Determination. Curr Osteoporos Rep. 2020;18(5):597–605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Chen YS, Lian WS, Kuo CW et al. Epigenetic regulation of skeletal tissue integrity and osteoporosis development. Int J Mol Sci. 2020;21(14):4923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Yasui T, Hirose J, Tsutsumi S, Nakamura K, Aburatani H, Tanaka S. Epigenetic regulation of osteoclast differentiation: possible involvement of Jmjd3 in the histone demethylation of Nfatc1. J Bone Miner Res. 2011;26(11):2665–2671. [DOI] [PubMed] [Google Scholar]
- 81. Yang D, Okamura H, Nakashima Y, Haneji T. Histone demethylase Jmjd3 regulates osteoblast differentiation via transcription factors Runx2 and osterix. J Biol Chem. 2013;288(47):33530–33541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Rummukainen P, Tarkkonen K, Dudakovic A et al. Lysine-specific Demethylase 1 (LSD1) epigenetically controls osteoblast differentiation. PLoS One. 2022;17(3):e0265027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Kim HJ, Park JW, Lee KH et al. Plant homeodomain finger protein 2 promotes bone formation by demethylating and activating Runx2 for osteoblast differentiation. Cell Res. 2014;24(10):1231–1249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Kim K, Shin Y, Kim J, Ulmer TS, An W. H3K27me1 is essential for MMP-9-dependent H3N-terminal tail proteolysis during osteoclastogenesis. Epigenetics Chromatin. 2018;11(1):23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Ideno H, Nakashima K, Komatsu K et al. G9a is involved in the regulation of cranial bone formation through activation of Runx2 function during development. Bone. 2020;137:115332. [DOI] [PubMed] [Google Scholar]
- 86. Cao X, He W, Rong K et al. DZNep promotes mouse bone defect healing via enhancing both osteogenesis and osteoclastogenesis. Stem Cell Res Ther. 2021;12(1):605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Qi Q, Wang Y, Wang X et al. Histone demethylase KDM4A regulates adipogenic and osteogenic differentiation via epigenetic regulation of C/EBPα and canonical Wnt signaling. Cell Mol Life Sci. 2020;77(12):2407–2421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88. Munehira Y, Yang Z, Gozani O. Systematic Analysis of Known and Candidate Lysine Demethylases in the Regulation of Myoblast Differentiation. J Mol Biol. 2017;429(13):2055–2065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Deng P, Yuan Q, Cheng Y et al. Loss of KDM4B exacerbates bone-fat imbalance and mesenchymal stromal cell exhaustion in skeletal aging. Cell Stem Cell. 2021;28(6):1057–1073..e7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Ye L, Fan Z, Yu B et al. Histone demethylases KDM4B and KDM6B promote osteogenic differentiation of human MSCs. Cell Stem Cell. 2018;23(6):898–899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Li J, Xuan Z, Liu C. Long non-coding RNAs and complex human diseases. Int J Mol Sci. 2013;14(9):18790–18808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Vicentini C, Galuppini F, Corbo V, Fassan M. Current role of non-coding RNAs in the clinical setting. Non-coding RNA Research. 2019;4(3):82–85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Rincón-Riveros A, Morales D, Rodríguez JA, Villegas VE, López-Kleine L. Bioinformatic tools for the analysis and prediction of ncRNA interactions. Int J Mol Sci. 2021;22(21):11397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Yang L, Wang H, Shen Q, Feng L, Jin H. Long non-coding RNAs involved in autophagy regulation. Cell Death Dis. 2017;8(10):e3073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Yuan H, Xu X, Feng X et al. A novel long noncoding RNA PGC1β-OT1 regulates adipocyte and osteoblast differentiation through antagonizing miR-148a-3p. Cell Death Differ. 2019;26(10):2029–2045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Wang K, Wang Y, Hu Z et al. Bone-targeted lncRNA OGRU alleviates unloading-induced bone loss via miR-320-3p/Hoxa10 axis. Cell Death Dis. 2020;11(5):382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Wang H, Zhao W, Tian QJ, Xin L, Cui M, Li YK. Effect of lncRNA AK023948 on rats with postmenopausal osteoporosis via PI3K/AKT signaling pathway. Eur Rev Med Pharmacol Sci. 2020;24(5):2181–2188. [DOI] [PubMed] [Google Scholar]
- 98. Cao L, Liu W, Zhong Y et al. Linc02349 promotes osteogenesis of human umbilical cord-derived stem cells by acting as a competing endogenous RNA for miR-25-3p and miR-33b-5p. Cell Prolif. 2020;53(5):e12814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99. Chen L, Xiong Y, Yan C et al. LncRNA KCNQ1OT1 accelerates fracture healing via modulating miR-701-3p/FGFR3 axis. FASEB J. 2020;34(4):5208–5222. [DOI] [PubMed] [Google Scholar]
- 100. Yi J, Liu D, Xiao J. LncRNA MALAT1 sponges miR-30 to promote osteoblast differentiation of adipose-derived mesenchymal stem cells by promotion of Runx2 expression. Cell Tissue Res. 2019;376(1):113–121. [DOI] [PubMed] [Google Scholar]
- 101. Xiong Y, Chen L, Yan C, Endo Y, Mi B, Liu G. The lncRNA Rhno1/miR-6979-5p/BMP2 axis modulates osteoblast differentiation. Int J Biol Sci. 2020;16(9):1604–1615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102. Chen Q, Wang M, Wu S. The lncRNA MCF2L-AS1 controls osteogenic differentiation by regulating miR-33a. Cell Cycle. 2020;19(9):1059–1065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103. Li G, Yun X, Ye K et al. Long non-coding RNA-H19 stimulates osteogenic differentiation of bone marrow mesenchymal stem cells via the microRNA-149/SDF-1 axis. J Cellular Molecular Medi. 2020;24(9):4944–4955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104. Yang Q, Han Y, Liu P et al. Long noncoding RNA GAS5 promotes osteogenic differentiation of human periodontal ligament stem cells by regulating GDF5 and p38/JNK signaling pathway. Front Pharmacol. 2020;11:701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105. Shen JJ, Zhang CH, Chen ZW et al. LncRNA HOTAIR inhibited osteogenic differentiation of BMSCs by regulating Wnt/β-catenin pathway. Eur Rev Med Pharmacol Sci. 2019;23(17):7232–7246. [DOI] [PubMed] [Google Scholar]
- 106. He S, Yang S, Zhang Y et al. LncRNA ODIR1 inhibits osteogenic differentiation of hUC-MSCs through the FBXO25/H2BK120ub/H3K4me3/OSX axis. Cell Death Dis. 2019;10(12):947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107. Yan L, Liao L, Su X. Role of mechano-sensitive non-coding RNAs in bone remodeling of orthodontic tooth movement: recent advances. Prog Orthod.. 2022;23(1):55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108. Zhang RF, Liu JW, Yu SP et al. LncRNA UCA1 affects osteoblast proliferation and differentiation by regulating BMP-2 expression. Eur Rev Med Pharmacol Sci. 2019;23(16):6774–6782. [DOI] [PubMed] [Google Scholar]
- 109. Ling L, Hu HL, Liu KY, Ram YI, Gao JL, Cao YM. Long noncoding RNA MIRG induces osteoclastogenesis and bone resorption in osteoporosis through negative regulation of miR-1897. Eur Rev Med Pharmacol Sci. 2019;23(23):10195–10203. [DOI] [PubMed] [Google Scholar]
- 110. Zhang Y, Chen XF, Li J, He F, Li X, Guo Y. lncRNA Neat1 stimulates osteoclastogenesis via sponging miR-7. J Bone Mineral Res. 2020;35(9):1772–1781. [DOI] [PubMed] [Google Scholar]
- 111. Zhang R, Li J, Li G et al. LncRNA Nron regulates osteoclastogenesis during orthodontic bone resorption. Int J Oral Sci. 2020;12(1):14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112. Li L, Fang J, Liu Y, Xiao L. LncRNA LOC100506178 promotes osteogenic differentiation via regulating miR-214-5p-BMP2 axis in human bone marrow mesenchymal stem cells. PeerJ. 2020;8:e8909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113. Liu SC, Sun QZ, Qiao XF et al. LncRNA TUG1 influences osteoblast proliferation and differentiation through the Wnt/β-catenin signaling pathway. Eur Rev Med Pharmacol Sci. 2019;23(11):4584–4590. [DOI] [PubMed] [Google Scholar]
- 114. Wang X, Zhao D, Zhu Y, Dong Y, Liu Y. Long non-coding RNA GAS5 promotes osteogenic differentiation of bone marrow mesenchymal stem cells by regulating the miR-135a-5p/FOXO1 pathway. Mol Cell Endocrinol. 2019;496:110534. [DOI] [PubMed] [Google Scholar]
- 115. Zhuang W, Ge X, Yang S et al. Upregulation of lncRNA MEG3 promotes osteogenic differentiation of mesenchymal stem cells from multiple myeloma patients by targeting BMP4 transcription. Stem Cells. 2015;33(6):1985–1997. [DOI] [PubMed] [Google Scholar]
- 116. Wu Y, Jiang Y, Liu Q, Liu CZ. lncRNA H19 promotes matrix mineralization through up-regulating IGF1 by sponging miR-185-5p in osteoblasts. BMC Mol Cell Biol. 2019;20(1):48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117. Gong ZM, Tang ZY, Sun XL. LncRNA PRNCR1 regulates osteogenic differentiation in osteolysis after hip replacement by targeting miR-211-5p. Biosci Rep. 2018; doi: 10.1042/bsr20180042 [DOI] [PubMed] [Google Scholar]
- 118. Peng W, Deng W, Zhang J, Pei G, Rong Q, Zhu S. Long noncoding RNA ANCR suppresses bone formation of periodontal ligament stem cells via sponging miRNA-758. Biochem Biophys Res Commun. 2018;503(2):815–821. [DOI] [PubMed] [Google Scholar]
- 119. Chen RS, Zhang XB, Zhu XT, Wang CS. LncRNA Bmncr alleviates the progression of osteoporosis by inhibiting RANML-induced osteoclast differentiation. Eur Rev Med Pharmacol Sci. 2019;23(21):9199–9206. [DOI] [PubMed] [Google Scholar]
- 120. Rachagani S, Macha MA, Heimann N et al. Clinical implications of miRNAs in the pathogenesis, diagnosis and therapy of pancreatic cancer. Adv Drug Deliv Rev. 2015;81:16–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121. Pelucchi P, Tria V, Martino V et al. A versatile tool for stable inhibition of microRNA activity. Biology. 2013;2(3):861–871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122. O'Brien J, Hayder H, Zayed Y, Peng C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front Endocrinol. 2018;9:402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123. Zhang J, Tu Q, Bonewald LF et al. Effects of miR-335-5p in modulating osteogenic differentiation by specifically downregulating Wnt antagonist DKK1. J of Bone & Mineral Res. 2011;26(8):1953–1963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124. Kim YJ, Bae SW, Yu SS, Bae YC, Jung JS. miR-196a regulates proliferation and osteogenic differentiation in mesenchymal stem cells derived from human adipose tissue. J Bone Mineral Res. 2009;24(5):816–825. [DOI] [PubMed] [Google Scholar]
- 125. Mizuno Y, Tokuzawa Y, Ninomiya Y et al. miR-210 promotes osteoblastic differentiation through inhibition of AcvR1b. FEBS Lett. 2009;583(13):2263–2268. [DOI] [PubMed] [Google Scholar]
- 126. Wang X, Omar O, Vazirisani F, Thomsen P, Ekström K. Mesenchymal stem cell-derived exosomes have altered microRNA profiles and induce osteogenic differentiation depending on the stage of differentiation. PLoS One. 2018;13(2):e0193059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127. Sugatani T, Vacher J, Hruska KA. A microRNA expression signature of osteoclastogenesis. Blood. 2011;117(13):3648–3657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128. Guo LJ, Liao L, Yang L, Li Y, Jiang TJ. MiR-125a TNF receptor-associated factor 6 to inhibit osteoclastogenesis. Exp Cell Res. 2014;321(2):142–152. [DOI] [PubMed] [Google Scholar]
- 129. Ponzetti M, Rucci N. Osteoblast differentiation and signaling: established concepts and emerging topics. Int J Mol Sci. 2021;22(13):6651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130. Cao Z, Moore BT, Wang Y et al. MiR-422a as a potential cellular microRNA biomarker for postmenopausal osteoporosis. PLoS One. 2014;9(5):e97098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131. Kagiya T, Nakamura S. Expression profiling of microRNAs in RAW264.7 cells treated with a combination of tumor necrosis factor alpha and RANKL during osteoclast differentiation. J Periodontal Research. 2013;48(3):373–385. [DOI] [PubMed] [Google Scholar]
- 132. Rossi M, Pitari MR, Amodio N et al. miR-29b negatively regulates human osteoclastic cell differentiation and function: implications for the treatment of multiple myeloma-related bone disease. J Cellular Physiology. 2013;228(7):1506–1515. [DOI] [PubMed] [Google Scholar]
- 133. Lee Y, Kim HJ, Park CK et al. MicroRNA-124 regulates osteoclast differentiation. Bone. 2013;56(2):383–389. [DOI] [PubMed] [Google Scholar]
- 134. Yin X, Zhou C, Li J et al. Autophagy in bone homeostasis and the onset of osteoporosis. Bone Res. 2019;7(1):28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135. Zhang QY, Jin HF, Chen S et al. Hydrogen sulfide regulating myocardial structure and function by targeting cardiomyocyte autophagy. Chin Med J (Engl). 2018;131(7):839–844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136. Yin XM. Autophagy in liver diseases: a matter of what to remove and whether to keep. Liver Research. 2018;2(3):109–111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137. Luo S, Li X, Zhang Y et al. Cargo recognition and function of selective autophagy receptors in plants. Int J Mol Sci. 2021;22(3):1013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138. Plotkin LI, Stains JP. Connexins and pannexins in the skeleton: gap junctions, hemichannels and more. Cell Mol Life Sci. 2015;72(15):2853–2867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139. Santos L, Elliott-Sale KJ, Sale C. Exercise and bone health across the lifespan. Biogerontology. 2017;18(6):931–946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140. Li Z, Li D, Su H, Xue H, Tan G, Xu Z. Autophagy: an important target for natural products in the treatment of bone metabolic diseases. Front Pharmacol. 2022;13:999017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141. Polisetty N, Fatima A, Madhira SL, Sangwan VS, Vemuganti GK. Mesenchymal cells from limbal stroma of human eye. Mol Vis. 2008;14:431–442. [PMC free article] [PubMed] [Google Scholar]
- 142. Di GH, Liu Y, Lu Y, Liu J, Wu C, Duan HF. IL-6 secreted from senescent mesenchymal stem cells promotes proliferation and migration of breast cancer cells. PLoS One. 2014;9(11):e113572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143. Chen H, Liu X, Chen H et al. Role of SIRT1 and AMPK in mesenchymal stem cells differentiation. Ageing Res Rev. 2014;13:55–64. [DOI] [PubMed] [Google Scholar]
- 144. Geng W, Shi H, Zhang X, Tan W, Cao Y, Mei R. Substance P enhances BMSC osteogenic differentiation via autophagic activation. Mol Med Report. 2019;20(1):664–670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145. Sotthibundhu A, McDonagh K, von Kriegsheim A et al. Rapamycin regulates autophagy and cell adhesion in induced pluripotent stem cells. Stem Cell Res Ther. 2016;7(1):166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146. Rodan GA, Noda M. Gene expression in osteoblastic cells. Crit Rev Eukaryot Gene Expr. 1991;1(2):85–98. [PubMed] [Google Scholar]
- 147. Wang J, Zhang Y, Cao J et al. The role of autophagy in bone metabolism and clinical significance. Autophagy. 2023;19(9):2409–2427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148. Bartelt A, Behler-Janbeck F, Beil FT et al. Lrp1 in osteoblasts controls osteoclast activity and protects against osteoporosis by limiting PDGF-RANKL signaling. Bone Res. 2018;6(1):4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149. Dowell P, Otto TC, Adi S, Lane MD. Convergence of peroxisome proliferator-activated receptor gamma and Foxo1 signaling pathways. J Biol Chem. 2003;278(46):45485–45491. [DOI] [PubMed] [Google Scholar]
- 150. Yang X, Karsenty G. ATF4, the osteoblast accumulation of which is determined post-translationally, can induce osteoblast-specific gene expression in non-osteoblastic cells. J Biol Chem. 2004;279(45):47109–47114. [DOI] [PubMed] [Google Scholar]
- 151. Jacome-Galarza CE, Percin GI, Muller JT et al. Developmental origin, functional maintenance and genetic rescue of osteoclasts. Nature. 2019;568(7753):541–545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152. Rachner TD, Khosla S, Hofbauer LC. Osteoporosis: now and the future. Lancet North Am Ed. 2011;377(9773):1276–1287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153. Zhao Y, Chen G, Zhang W et al. Autophagy regulates hypoxia-induced osteoclastogenesis through the HIF-1α/BNIP3 signaling pathway. J Cellular Physiol. 2012;227(2):639–648. [DOI] [PubMed] [Google Scholar]
- 154. Montaseri A, Giampietri C, Rossi M, Riccioli A, Del Fattore A, Filippini A. The role of autophagy in osteoclast differentiation and bone resorption function. Biomolecules. 2020;10(10):1398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155. Arai A, Kim S, Goldshteyn V et al. Beclin1 modulates bone homeostasis by regulating osteoclast and chondrocyte differentiation. J Bone Mineral Res. 2019;34(9):1753–1766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156. Ishii T, Shimazu Y, Nishiyama I, Kikuta J, Ishii M. The role of sphingosine 1-phosphate in migration of osteoclast precursors; an application of intravital two-photon microscopy. Mol Cells. 2011;31(5):399–404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157. Hsu LC, Reddy SV, Yilmaz Ö, Yu H. Sphingosine-1-phosphate receptor 2 controls podosome components induced by RANKL affecting osteoclastogenesis and bone resorption. Cells. 2019;8(1):17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158. Ivashkiv LB. Metabolic-epigenetic coupling in osteoclast differentiation. Nat Med. 2015;21(3):212–213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159. Fujita K, Xing Q, Khosla S, Monroe DG. Mutual enhancement of differentiation of osteoblasts and osteocytes occurs through direct cell-cell contact. J Cell Biochem. 2014;115(11). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160. Bellido T, Saini V, Pajevic PD. Effects of PTH on osteocyte function. Bone. 2013;54(2):250–257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161. Adachi T, Aonuma Y, Taira K, Hojo M, Kamioka H. Asymmetric intercellular communication between bone cells: propagation of the calcium signaling. Biochem Biophys Res Commun. 2009;389(3):495–500. [DOI] [PubMed] [Google Scholar]
- 162. Riquelme MA, Gu S, Hua R, Jiang JX. Mechanotransduction via the coordinated actions of integrins, PI3K signaling and Connexin hemichannels. Bone Res. 2021;9(1):8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163. Xu H, Xia M, Sun L, Wang H, Zhang WB. Osteocytes enhance osteogenesis by autophagy-mediated FGF23 secretion under mechanical tension. Front Cell Dev Biol. 2021;9:782736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164. Zhang B, Hou R, Zou Z et al. Mechanically induced autophagy is associated with ATP metabolism and cellular viability in osteocytes in vitro. Redox Biol. 2018;14:492–498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165. Vrahnas C, Blank M, Dite TA et al. Author correction: increased autophagy in EphrinB2-deficient osteocytes is associated with elevated secondary mineralization and brittle bone. Nat Commun. 2019;10(1):5073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166. Wang XY, Gong LJ, Huang JM, Jiang C, Yan ZQ. Pinocembrin alleviates glucocorticoid-induced apoptosis by activating autophagy via suppressing the PI3K/Akt/mTOR pathway in osteocytes. Eur J Pharmacol. 2020;880:173212. [DOI] [PubMed] [Google Scholar]
- 167. Manjithaya R, Subramani S. Autophagy: a broad role in unconventional protein secretion?. Trends Cell Biol. 2011;21(2):67–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168. Denry I, Goudouri OM, Fredericks DC, Akkouch A, Acevedo MR, Holloway JA. Strontium-releasing fluorapatite glass-ceramic scaffolds: structural characterization and in vivo performance. Acta Biomater. 2018;75:463–471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169. Luo P, Gao F, Niu D et al. The role of autophagy in chondrocyte metabolism and osteoarthritis: a comprehensive research review. Biomed Res Int. 2019;2019:1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170. Gao M, Chen C, Zhang Q, Bian J, Qin L, Bao L. Research progress on the antiosteoarthritic mechanism of action of natural products. Evid-Based Complement Altern Med. 2021;2021:1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171. Huang Y, Seitz D, König F, Müller PE, Jansson V, Klar RM. Induction of articular chondrogenesis by chitosan/hyaluronic-acid-based biomimetic matrices using human adipose-derived stem cells. Int J Mol Sci. 2019;20(18):4487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172. Xiao SQ, Cheng M, Wang L et al. The role of apoptosis in the pathogenesis of osteoarthritis. International Orthopaedics (SICOT). 2023;47(8):1895–1919. [DOI] [PubMed] [Google Scholar]
- 173. Li J, Jiang M, Yu Z et al. Artemisinin relieves osteoarthritis by activating mitochondrial autophagy through reducing TNFSF11 expression and inhibiting PI3K/AKT/mTOR signaling in cartilage. Cell Mol Biol Lett. 2022;27(1):62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174. Jiang LB, Lee S, Wang Y, Xu QT, Meng DH, Zhang J. Adipose-derived stem cells induce autophagic activation and inhibit catabolic response to pro-inflammatory cytokines in rat chondrocytes. Osteoarthritis Cartilage. 2016;24(6):1071–1081. [DOI] [PubMed] [Google Scholar]
- 175. Lian LP, Xi XY. Long non-coding RNA XIST protects chondrocytes ATDC5 and CHON-001 from IL-1β-induced injury via regulating miR-653-5p/SIRT1 axis. J Biol Regul Homeost Agents. 2020;34(2):379–391. [DOI] [PubMed] [Google Scholar]
- 176. Wang Y, Huang M, Xu W, Li F, Ma C, Tang X. Calcitriol-enhanced autophagy in gingival epithelium attenuates periodontal inflammation in rats with type 2 diabetes mellitus. Front. Endocrinol.. 2022;13:1051374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177. Zhang P, Xiao J, Luo C et al. The effect of JAK inhibitor tofacitinib on chondrocyte autophagy. Inflammation. 2023;46(5):1764–1776. [DOI] [PubMed] [Google Scholar]
- 178. Pan Y, Yang Y, Fan M et al. Progranulin regulation of autophagy contributes to its chondroprotective effect in osteoarthritis. Genes Diseases. 2023;10(4):1582–1595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179. He L, Xu Z, Niu X et al. GPRC5B protects osteoarthritis by regulation of autophagy signaling. Acta Pharmaceutica Sinica B. 2023;13(7):2976–2989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180. Li S, Gao L, Zhang W et al. MiR-152-5p suppresses osteogenic differentiation of mandible mesenchymal stem cells by regulating ATG14-mediated autophagy. Stem Cell Res Ther. 2022;13(1):359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181. Gravina GL, Festuccia C, Marampon F et al. Biological rationale for the use of DNA methyltransferase inhibitors as new strategy for modulation of tumor response to chemotherapy and radiation. Mol Cancer. 2010;9(1):305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182. Nandy D, Rajam SM, Dutta D. A three layered histone epigenetics in breast cancer metastasis. Cell Biosci. 2020;10(1):52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183. Pargol M, Akbari M et al. Investigation the role of autophagy in non-small cell lung cancer. Asian Pac J Cancer Prev. 2021;22(3):947–955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 184. Biga PR, Latimer MN, Froehlich JM, Gabillard JC, Seiliez I. Distribution of H3K27me3, H3K9me3, and H3K4me3 along autophagy-related genes highly expressed in starved zebrafish myotubes. Biol Open. 2017;6(11):1720–1725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185. Mushtaq A, Ashraf NU, Altaf M. The mTORC1-G9a-H3K9me2 axis negatively regulates autophagy in fatty acid-induced hepatocellular lipotoxicity. J Biol Chem. 2023;299(3):102937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186. Li Y, Zhang M, Sheng M et al. Therapeutic potential of GSK-J4, a histone demethylase KDM6B/JMJD3 inhibitor, for acute myeloid leukemia. J Cancer Res Clin Oncol. 2018;144(6):1065–1077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 187. Shogren-Knaak M, Ishii H, Sun JM, Pazin MJ, Davie JR, Peterson CL. Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science. 2006;311(5762):844–847. [DOI] [PubMed] [Google Scholar]
- 188. Puri D, Subramanyam D. Stress—(self) eating: epigenetic regulation of autophagy in response to psychological stress. FEBS J. 2019;286(13):2447–2460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189. Fabian MR, Sonenberg N, Filipowicz W. Regulation of mRNA translation and stability by microRNAs. Annu Rev Biochem. 2010;79(1):351–379. [DOI] [PubMed] [Google Scholar]
- 190. Dweep H, Kubikova N, Gretz N, Voskarides K, Felekkis K. Homo sapiens exhibit a distinct pattern of CNV genes regulation: an important role of miRNAs and SNPs in expression plasticity. Sci Rep. 2015;5(1):12163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191. Tan S, Shi H, Ba M et al. miR-409-3p sensitizes colon cancer cells to oxaliplatin by inhibiting Beclin-1-mediated autophagy. Int J Mol Med. 2016;37(4):1030–1038. [DOI] [PubMed] [Google Scholar]
- 192. Zhang F, Wang B, Long H et al. Decreased miR-124-3p Expression Prompted Breast Cancer Cell Progression Mainly by Targeting Beclin-1. Clin Lab. 2016;62(06/2016):1139–1145. [DOI] [PubMed] [Google Scholar]
- 193. Kovaleva V, Mora R, Park YJ et al. miRNA-130a targets ATG2B and DICER1 to inhibit autophagy and trigger killing of chronic lymphocytic leukemia cells. Cancer Res. 2012;72(7):1763–1772. [DOI] [PubMed] [Google Scholar]
- 194. Zhang J, Chen LM, Zou Y, Zhang S, Xiong F, Wang CY. Implication of epigenetic factors in the pathogenesis of type 1 diabetes. Chin Med J (Engl). 2021;134(9):1031–1042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195. Shu F, Xiao H, Li QN et al. Epigenetic and post-translational modifications in autophagy: biological functions and therapeutic targets. Sig Transduct Target Ther. 2023;8(1):32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196. Silwal P, Paik S, Jeon SM, Jo EK. Nuclear receptors as autophagy-based antimicrobial therapeutics. Cells. 2020;9(9):1979, [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197. Reddy MA, Natarajan R. Recent developments in epigenetics of acute and chronic kidney diseases. Kidney Int. 2015;88(2):250–261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198. Morris HA, Turner AG, Anderson PH. Vitamin-D regulation of bone mineralization and remodelling during growth. Front Biosci. 2012;E4(1):677–689. [DOI] [PubMed] [Google Scholar]
- 199. Siris ES, Adler R, Bilezikian J et al. The clinical diagnosis of osteoporosis: a position statement from the National Bone Health Alliance Working Group. Osteoporos Int. 2014;25(5):1439–1443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200. Gkastaris K, Goulis DG, Potoupnis M, Anastasilakis AD, Kapetanos G. Obesity, osteoporosis and bone metabolism. J Musculoskelet Neuronal Interact. 2020;20(3):372–381. [PMC free article] [PubMed] [Google Scholar]
- 201. Letarouilly JG, Broux O, Clabaut A. New insights into the epigenetics of osteoporosis. Genomics. 2019;111(4):793–798. [DOI] [PubMed] [Google Scholar]
- 202. Wang K, Niu J, Kim H, Kolattukudy PE. Osteoclast precursor differentiation by MCPIP via oxidative stress, endoplasmic reticulum stress, and autophagy. J Mol Cell Biol. 2011;3(6):360–368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 203. Sambandam Y, Townsend MT, Pierce JJ et al. Microgravity control of autophagy modulates osteoclastogenesis. Bone. 2014;61:125–131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204. Sun KT, Chen MY, Tu MG, Wang IK, Chang SS, Li CY. MicroRNA-20a regulates autophagy related protein-ATG16L1 in hypoxia-induced osteoclast differentiation. Bone. 2015;73:145–153. [DOI] [PubMed] [Google Scholar]
- 205. Jin Y, Tymen SD, Chen D et al. MicroRNA-99 family targets AKT/mTOR signaling pathway in dermal wound healing. PLoS One. 2013;8(5):e64434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206. Wei F, Liu Y, Guo Y et al. miR-99b-targeted mTOR induction contributes to irradiation resistance in pancreatic cancer. Mol Cancer. 2013;12(1):81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207. Jin Q, Martinez CA, Arcipowski KM et al. USP7 cooperates with NOTCH1 to drive the oncogenic transcriptional program in T-Cell leukemia. Clin Cancer Res. 2019;25(1):222–239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 208. Zhou F, Li F, Fang P et al. Ubiquitin-specific protease 4 antagonizes osteoblast differentiation through dishevelled. J Bone Mineral Res. 2016;31(10):1888–1898. [DOI] [PubMed] [Google Scholar]
- 209. Lu X, Zhang Y, Zheng Y, Chen B. The miRNA-15b/USP7/KDM6B axis engages in the initiation of osteoporosis by modulating osteoblast differentiation and autophagy. J Cellular Molecular Medi. 2021;25(4):2069–2081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 210. Gao Y, Ge W. The histone methyltransferase DOT1L inhibits osteoclastogenesis and protects against osteoporosis. Cell Death Dis. 2018;9(2):33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211. Zhang L, Qi M, Chen J et al. Impaired autophagy triggered by HDAC9 in mesenchymal stem cells accelerates bone mass loss. Stem Cell Res Ther. 2020;11(1):269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 212. Behera J, Ison J, Tyagi A, Mbalaviele G, Tyagi N. Mechanisms of autophagy and mitophagy in skeletal development, diseases and therapeutics. Life Sci. 2022;301:120595. [DOI] [PubMed] [Google Scholar]
- 213. Herrero-Beaumont G, Roman-Blas JA, Castañeda S, Jimenez SA. Primary osteoarthritis no longer primary: three subsets with distinct etiological, clinical, and therapeutic characteristics. Semin Arthritis Rheum. 2009;39(2):71–80. [DOI] [PubMed] [Google Scholar]
- 214. Bijlsma JW, Berenbaum F, Lafeber FP. Osteoarthritis: an update with relevance for clinical practice. Lancet North Am Ed. 2011;377(9783):2115–2126. [DOI] [PubMed] [Google Scholar]
- 215. Liu-Bryan R, Terkeltaub R. Emerging regulators of the inflammatory process in osteoarthritis. Nat Rev Rheumatol. 2015;11(1):35–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 216. Caramés B, Taniguchi N, Otsuki S, Blanco FJ, Lotz M. Autophagy is a protective mechanism in normal cartilage, and its aging-related loss is linked with cell death and osteoarthritis. Arthritis Rheumatism. 2010;62(3):791–801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217. Shapiro IM, Layfield R, Lotz M, Settembre C, Whitehouse C. Boning up on autophagy: the role of autophagy in skeletal biology. Autophagy. 2014;10(1):7–19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 218. Hollander W, Meulenbelt I. DNA Methylation in Osteoarthritis. Curr Genomics. 2015;16(6):419–426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 219. Hashimoto K, Oreffo RO, Gibson MB, Goldring MB, Roach HI. DNA demethylation at specific CpG sites in the IL1B promoter in response to inflammatory cytokines in human articular chondrocytes. Arthritis Rheumatism. 2009;60(11):3303–3313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 220. Takahashi A, de Andrés MC, Hashimoto K, Itoi E, Oreffo RO. Epigenetic regulation of interleukin-8, an inflammatory chemokine, in osteoarthritis. Osteoarthritis Cartilage. 2015;23(11):1946–1954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 221. Xue JF, Shi ZM, Zou J, Li XL. Inhibition of PI3K/AKT/mTOR signaling pathway promotes autophagy of articular chondrocytes and attenuates inflammatory response in rats with osteoarthritis. Biomed Pharmacother. 2017;89:1252–1261. [DOI] [PubMed] [Google Scholar]
- 222. de Andrés MC, Imagawa K, Hashimoto K et al. Loss of methylation in CpG sites in the NF-κB enhancer elements of inducible nitric oxide synthase is responsible for gene induction in human articular chondrocytes. Arthritis Rheumatism. 2013;65(3):732–742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 223. Martinez-Sanchez A, Dudek KA, Murphy CL. Regulation of human chondrocyte function through direct inhibition of cartilage master regulator SOX9 by microRNA-145 (miRNA-145). J Biol Chem. 2012;287(2):916–924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 224. He L, Thomson JM, Hemann MT et al. A microRNA polycistron as a potential human oncogene. Nature. 2005;435(7043):828–833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225. He W, Cheng Y. Inhibition of miR-20 promotes proliferation and autophagy in articular chondrocytes by PI3K/AKT/mTOR signaling pathway. Biomed Pharmacother. 2018;97:607–615. [DOI] [PubMed] [Google Scholar]
- 226. Li H, Li Z, Pi Y et al. MicroRNA-375 exacerbates knee osteoarthritis through repressing chondrocyte autophagy by targeting ATG2B. Aging. 2020;12(8):7248–7261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 227. Cai C, Min S, Yan B et al. MiR-27a promotes the autophagy and apoptosis of IL-1β treated-articular chondrocytes in osteoarthritis through PI3K/AKT/mTOR signaling. Aging. 2019;11(16):6371–6384. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 228. D'Adamo S, Alvarez-Garcia O, Muramatsu Y, Flamigni F, Lotz MK. MicroRNA-155 suppresses autophagy in chondrocytes by modulating expression of autophagy proteins. Osteoarthritis Cartilage. 2016;24(6):1082–1091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 229. Wang J, Li X, Guo X et al. MicroRNA-34a-5p promotes the progression of osteoarthritis secondary to developmental dysplasia of the hip by restraining SESN2-induced autophagy. J Orthopaedic Res. 2024;42(1):66–77. [DOI] [PubMed] [Google Scholar]
- 230. Ni Z, Kuang L, Chen H et al. The exosome-like vesicles from osteoarthritic chondrocyte enhanced mature IL-1β production of macrophages and aggravated synovitis in osteoarthritis. Cell Death Dis. 2019;10(7):522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 231. Taran SJ, Taran R, Malipatil NB. Pediatric osteosarcoma: an updated review. Indian J Med Paediatric Oncol. 2017;38(01):33–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 232. Lindsey BA, Markel JE, Kleinerman ES. Osteosarcoma overview. Rheumatol Ther. 2017;4(1):25–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 233. Wang P, Zhao ZQ, Guo SB et al. Roles of microRNA-22 in suppressing proliferation and promoting sensitivity of osteosarcoma cells via metadherin-mediated autophagy. Orthopaedic Surgery. 2019;11(2):285–293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 234. Xie C, Liu S, Wu B et al. miR-19 promotes cell proliferation, invasion, migration, and EMT by inhibiting SPRED2-mediated autophagy in osteosarcoma cells. Cell Transplant. 2020;29:096368972096246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 235. Bai Y, Chen Y, Chen X et al. Trichostatin A activates FOXO1 and induces autophagy in osteosarcoma. aoms. 2019;15(1):204–213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 236. Wu G, Yu W, Zhang M, Yin R, Wu Y, Liu Q. MicroRNA-145-3p suppresses proliferation and promotes apotosis and autophagy of osteosarcoma cell by targeting HDAC4. Artificial Cells Nanomedicine Biotechnol. 2018;46(sup2):579–586. [DOI] [PubMed] [Google Scholar]
- 237. Song H, Liu J, Wu X et al. LHX2 promotes malignancy and inhibits autophagy via mTOR in osteosarcoma and is negatively regulated by miR-129-5p. Aging. 2019;11(21):9794–9810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 238. Wang Y, Zeng X, Wang N et al. Long noncoding RNA DANCR, working as a competitive endogenous RNA, promotes ROCK1-mediated proliferation and metastasis via decoying of miR-335-5p and miR-1972 in osteosarcoma. Mol Cancer. 2018;17(1):89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 239. Dong S, Xiao Y, Ma X et al. miR-193b increases the chemosensitivity of osteosarcoma cells by promoting FEN1-mediated autophagy. Onco Targets Ther. 2019;12:10089–10098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 240. Mellado M, Martínez-Muñoz L, Cascio G, Lucas P, Pablos JL, Rodríguez-Frade JM. T cell migration in rheumatoid arthritis. Front Immunol. 2015;6:384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241. Scott DL. Evidence for early disease-modifying drugs in rheumatoid arthritis. Arthritis Res Ther. 2004;6(1):15–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 242. Matteson EL, Weyand CM, Fulbright JW, Christianson TJ, McClelland RL, Goronzy JJ. How aggressive should initial therapy for rheumatoid arthritis be? Factors associated with response to ‘non-aggressive’ DMARD treatment and perspective from a 2-yr open label trial. Rheumatology (Oxford). 2004;43(5):619–625. [DOI] [PubMed] [Google Scholar]
- 243. Grassi W, De Angelis R, Lamanna G, Cervini C. The clinical features of rheumatoid arthritis. Eur J Radiol. 1998;27(Suppl 1):S18–S24. [DOI] [PubMed] [Google Scholar]
- 244. Smolen JS, Aletaha D, Gruben D, Zwillich SH, Krishnaswami S, Mebus C. Brief report: remission rates with tofacitinib treatment in rheumatoid arthritis: a comparison of various remission criteria. Arthritis Rheumatol. 2017;69(4):728–734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 245. Mok CC, Tam LS, Chan TH, Lee GK, Li EK. Management of rheumatoid arthritis: consensus recommendations from the Hong Kong Society of Rheumatology. Clin Rheumatol. 2011;30(3):303–312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 246. Bottini N, Firestein GS. Epigenetics in rheumatoid arthritis: a primer for rheumatologists. Curr Rheumatol Rep. 2013;15(11):372. [DOI] [PubMed] [Google Scholar]
- 247. Ribeiro ML, Reyes-Garau D, Armengol M, Fernández-Serrano M, Roué G. Recent advances in the targeting of epigenetic regulators in B-cell non-hodgkin lymphoma. Front Genet. 2019;10:986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 248. Wada TT, Araki Y, Sato K et al. Aberrant histone acetylation contributes to elevated interleukin-6 production in rheumatoid arthritis synovial fibroblasts. Biochem Biophys Res Commun. 2014;444(4):682–686. [DOI] [PubMed] [Google Scholar]
- 249. BJ, arthritis R: Autophagy: a dual role in the life and death of RASFs. Nat Rev Rheumatol. 2013;9(11):637. [DOI] [PubMed] [Google Scholar]
- 250. Kato M, Ospelt C, Gay RE, Gay S, Klein K. Dual role of autophagy in stress-induced cell death in rheumatoid arthritis synovial fibroblasts. Arthritis Rheumatology. 2014;66(1):40–48. [DOI] [PubMed] [Google Scholar]
- 251. Kukita A, Kukita T, Nagata K et al. The transcription factor FBI-1/OCZF/LRF is expressed in osteoclasts and regulates RANKL-induced osteoclast formation in vitro and in vivo. Arthritis Rheumatism. 2011;63(9):2744–2754. [DOI] [PubMed] [Google Scholar]
- 252. Vomero M, Barbati C, Colasanti T et al. Autophagy and rheumatoid arthritis: current knowledges and future perspectives. Front Immunol. 2018;9:1577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 253. Li G, Liu Y, Meng F et al. LncRNA MEG3 inhibits rheumatoid arthritis through miR-141 and inactivation of AKT/mTOR signalling pathway. J Cellular Molecular Medi. 2019;23(10):7116–7120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 254. Zhou H, Huang L, Zhan K, Liu X. Wenhua Juanbi recipe attenuates rheumatoid arthritis via inhibiting miRNA-146a-mediated autophagy. Biomed Res Int. 2022;2022:1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 255. Yang S, Yin W, Ding Y, Liu F. Lnc RNA ZFAS1 regulates the proliferation, apoptosis, inflammatory response and autophagy of fibroblast-like synoviocytes via miR-2682-5p/ADAMTS9 axis in rheumatoid arthritis. Biosci Rep. 2020;40(8). doi: 10.1042/bsr20201273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 256. Rodríguez-Paredes M, Esteller M. Cancer epigenetics reaches mainstream oncology. Nat Med. 2011;17(3):330–339. [DOI] [PubMed] [Google Scholar]
- 257. Yen CY, Huang HW, Shu CW et al. DNA methylation, histone acetylation and methylation of epigenetic modifications as a therapeutic approach for cancers. Cancer Lett. 2016;373(2):185–192. [DOI] [PubMed] [Google Scholar]
- 258. Dhillon S. Decitabine/Cedazuridine: first Approval. Drugs. 2020;80(13):1373–1378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 259. Suwanawiboon B, Sumida KN. 5-azacitidine: an alternative treatment of myelodysplastic syndromes in patient with refractory response to hematopoietic growth factor, a case report and review of literatures. Hawaii Med J. 2004;63(1):14–16., 25. [PubMed] [Google Scholar]
- 260. De Smedt E, Devin J, Muylaert C et al. G9a/GLP targeting in MM promotes autophagy-associated apoptosis and boosts proteasome inhibitor-mediated cell death. Blood Adv. 2021;5(9):2325–2338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 261. Rossetto D, Avvakumov N, Côté J. Histone phosphorylation: a chromatin modification involved in diverse nuclear events. Epigenetics. 2012;7(10):1098–1108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 262. Hottiger MO. Nuclear ADP-ribosylation and its role in chromatin plasticity, cell differentiation, and epigenetics. Annu Rev Biochem. 2015;84:(1):227–263. [DOI] [PubMed] [Google Scholar]
- 263. Zhang Y. Transcriptional regulation by histone ubiquitination and deubiquitination. Genes Dev. 2003;17(22):2733–2740. [DOI] [PubMed] [Google Scholar]
- 264. Shiio Y, Eisenman RN. Histone sumoylation is associated with transcriptional repression. Proc Natl Acad Sci USA. 2003;100(23):13225–13230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 265. Christophorou MA, Castelo-Branco G, Halley-Stott RP et al. Citrullination regulates pluripotency and histone H1 binding to chromatin. Nature. 2014;507(7490):104–108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 266. Fujiki R, Hashiba W, Sekine H et al. GlcNAcylation of histone H2B facilitates its monoubiquitination. Nature. 2011;480(7378):557–560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 267. Tsukada Y. Hydroxylation mediates chromatin demethylation. J Biochem. 2012;151(3):229–246. [DOI] [PubMed] [Google Scholar]
- 268. Howe FS, Boubriak I, Sale MJ et al. Lysine acetylation controls local protein conformation by influencing proline isomerization. Mol Cell. 2014;55(5):733–744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 269. Li CJ, Cheng P, Liang MK et al. MicroRNA-188 regulates age-related switch between osteoblast and adipocyte differentiation. J Clin Invest. 2015;125(4):1509–1522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 270. Šmuc T, Ahn IY, Ulrich H. Nucleic acid aptamers as high affinity ligands in biotechnology and biosensorics. J Pharm Biomed Anal. 2013;81-82:210–217. [DOI] [PubMed] [Google Scholar]
- 271. Liu J, Dang L, Li D et al. A delivery system specifically approaching bone resorption surfaces to facilitate therapeutic modulation of microRNAs in osteoclasts. Biomaterials. 2015;52:148–160. [DOI] [PubMed] [Google Scholar]
- 272. Yao Y, Jia T, Pan Y et al. Using a novel microRNA delivery system to inhibit osteoclastogenesis. Int J Mol Sci. 2015;16(12):8337–8350. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The data involved in this review article are available from the corresponding author on reasonable request.