Abstract
Human immunodeficiency virus type 1 (HIV-1) may be studied by molecular or immunological approaches. Most analyses have been performed by genetic comparison of isolates and have led to the definition of clades or subtypes within the major (M) group of HIV-1. Five subtypes (A to E) were initially identified by comparison of genomic sequences. Four new subtypes (F to I) were identified more recently. Amino acid differences in the immunogenic V3 loop between isolates have also been studied, leading to a phenetic classification of at least 14 clusters (1 to 14) of sequences (B. T. M. Korber, K. McInnes, R. F. Smith, and G. Myers, J. Virol. 68:6730–6744, 1994). In this study, we compared the antigenicity of the V3 consensus sequences defined by phylogenetic analysis to the antigenicity of those defined by phenetic analysis. We used a recently developed subtype-specific enzyme immunoassay (SSEIA) that uses the principle of blocking with an excess of peptide in the liquid phase. Two SSEIAs were performed, the first with five V3 sequences defined by phylogenetic analysis and the second with 14 V3 sequences defined by phenetic analysis. A total of 168 HIV-1 sera taken from seropositive individuals from seven different countries or regions were studied. Experimental and statistical data, including correlation matrix and cluster analyses, demonstrated associations between the genetic subtypes and phenetically associated groups. Most of these were predicted by Korber et al. (J. Virol. 68:6730–6744, 1994) by theoretical analysis. We also found that V3 sequences can be grouped into between three and five antigenically unrelated categories. Residues that may be responsible for major antigenic differences were identified at the apex of the V3 loop, within the octapeptide xIGPGxxx, where x represents the critical positions. Our study provides evidence that there is a limited number of V3 serotypes which could be easily monitored by serological assays to study the diversity and dynamics of HIV-1 strains.
The diversity of human immunodeficiency virus type 1 (HIV-1) is a major problem in the development of an effective vaccine against AIDS. Many HIV-1 sequences are now available, and phylogenetic analysis resulting in a continuously developing classification into subtypes or clades is possible (45). HIV-1 isolates are classified into the M group (for major) or O group (for outlier). The O group contains only a few variants, all from a limited area of Africa (19, 27, 50). The M group includes variants responsible for the present AIDS pandemic. It contains at least five subtypes (A to E), to which have been added more recently four other subtypes (F to I) (23, 28, 34, 36, 37). Subtypes A, C, D, G, and H are common in Africa (21, 35, 37, 38). Subtype B is the most common in America and Europe (24, 26, 51). Subtype E occurs mainly in Asia (25, 30, 41), and subtype F has been detected in Brazil and Romania (3, 28, 34). These distributions are not restrictive. Subtype C is also present in Asia (India and China), and subtype G is also present in Russia (7, 12, 29). The African subtypes (A, C, and D) and the Asian subtype (E) have also been identified in North America and in European countries (9, 13, 14, 32, 48). All the subtypes are present in Africa, including B (detected in West Africa), E (Central African Republic), and F (Cameroon) (1, 35, 38). Analysis of the genetic diversity of HIV-1 is becoming more difficult due to the increasing frequency of coinfections and recombinations (15, 20, 44).
Phylogenetic trees have been generated with gag, env, or tat nucleotide sequences. Shorter DNA sequences encoding the functionally important V3 region of the envelope protein are most frequently used to provide reliable subtype designations (37). The diversity of the immunogenic V3 loop has also been studied by comparing the amino acids of different isolates, leading to a phenetic classification of at least 14 clusters of sequences, each one characterized by a consensus sequence based on the most common amino acid in a given position (22).
The heterogeneity of HIV-1 strains is studied mostly by molecular characterization of genomic sequences. This involves sequencing fragments amplified by the PCR or the use of the heteroduplex mobility assay (10, 11). However, although these methods allow direct subtype classification, they are time-consuming and expensive and require highly trained workers. Serotyping of HIV-1 by antibody (Ab) binding to the V3 region has been suggested as an alternative approach (8, 40, 49, 51). Such an approach may make it possible to identify subtypes based on antigenic rather than genetic properties. This immunological information about antigenic diversity might be of value in vaccine development. We recently developed a subtype-specific enzyme immunoassay (SSEIA) which gave results consistent with those of genotyping (4, 48). This assay used V3 consensus sequences defined by genetic classification, so we wanted to compare the antigenicity of these V3 consensus sequences to the antigenicity of those defined by phenetic analysis. The phenetic clustering of V3 loop amino acid sequences is not always consistent with phylogenetic analysis. Our results suggested that a limited number of serotypes may exist and identified amino acids at the tip of the V3 loop that may be responsible for serological discrimination.
MATERIALS AND METHODS
Peptides.
Nineteen peptides whose sequences corresponded to sequences in the V3 region of HIV-1 were synthesized (Table 1). These peptides corresponded to the consensus sequences of the five major subtypes of HIV-1 group M (A to E) (36) or to the consensus sequences of the 14 phenetic clusters described by Korber et al. in 1994 (22). The peptides were synthesized by Merrifield’s solid-phase procedure with an automated peptide synthesizer (Applied Biosystems 431A), 9-fluorenylmethoxycarbonyl-protected amino acids, and hydroxymethylphenoacetic-polystyrene resin (31). The resin support and side-chain-protecting groups were removed with trifluoroacetic acid after synthesis, and distilled water, phenol, ethanedithiol, and thioanisole were used as scavengers. The peptides were cleaved and purified by reverse-phase chromatography on C8 columns (Aquapore octyl, 20 μm, 100 by 10 mm; Applied Biosystems). The purity of the preparations was checked both by observation of a single sharp peak in high-pressure liquid chromatography analysis with C8 columns (Aquapore octyl RP-300, 7 μm, 220 by 4.6 mm; Applied Biosystems) and by amino acid analysis. Peptide compositions were as planned.
TABLE 1.
Sequences of synthetic peptides used in the assays
| Peptide | V3 consensus sequenceb |
|---|---|
| Genetic subtypea | 310 311 315 316 319 325 326 330 335 339 |
| A | N N T ·c· R K S V · H I · · G P G Q A F Y · · · A T G D I I G D I R Q A H C |
| B | N N T · · R K S I · H I · · G P G R A F Y · · · T T G E I I G D I R Q A H C |
| C | N N T · · R K S I · R I · · G P G Q T F Y · · · A T G D I I G D I R Q A H C |
| D | N N T · · R Q R T · H I · · G P G Q A L Y · · · T T · R I I G D I R Q A H C |
| Ed | N N T · · R T S I · T I · · G P G Q V F Y · · · R T G D I I G D I R K A Y C |
| Phenetic groupe | 310 311 315 316 319 325 326 330 335 339 |
| 1 | N N T · · R K S I · H I · · G P G R A F Y · · · T T G D I I G D I R Q A H C |
| 2 | N N T · · R K S I · H I · · G P G R A F Y · · · A T G E I I G D I R Q A H C |
| 3 | N N T · · S K R I · S I · · G P G R A F Y · · · A T G K I I G D I R Q A H C |
| 4 | N N T · · R K S I · R I · · G P G Q A F Y · · · A T G D I I G D I R Q A H C |
| 5 | N N T · · R K S I · H I · · G P G R A F Y · · · A T G D I I G D I R Q A H C |
| 6 | K N T R · R R I H · · I · · G P G R A F Y · · · T T K Q I I G D I R Q A H C |
| 7 | N N T · · R T S I · T I · · G P G Q V F Y · · · R T G D I I G D I R K A Y C |
| 8 | K K M · · R T S A · R I · · G P G R V F H · · · K T G S I T G D I R K A Y C |
| 9 | N N T · · R Q S T · H I · · G P G Q A L Y · · · T T · K I I G D I R Q A H C |
| 10 | N N T · · R Q G T · H I · · G P G R A Y Y · · · T T · N I I G D I R Q A H C |
| 11 | N N T · · R Q R T · S I · · G Q G Q A L Y · · · T T · E I R G D I R Q A Y C |
| 12 | D N I K · I Q R T · P I · · G Q G Q A L Y · · · T T R I T G Y I · G Q A H C |
| 13 | Q N T · · R Q R T · P I · · G L G Q S L Y · · · T T R S R S · I I G Q A H C |
| 14 | N N K · · R Q G T · P I · · G L G Q A L Y · · · T T R · I K G D I R K A H C |
V3 consensus sequences of the five subtypes initially described by Myers et al. (36).
Residues are numbered according to Ratner et al. (42).
The dots represent insertions.
Peptide E is similar to peptide 7.
Consensus sequences of the 14 V3 phenetic groups initially described by Korber et al. (22) and truncated at the four N-terminal amino acids.
Sera.
A total of 168 HIV-1 Ab-positive sera collected in seven different countries or regions were used (24 sera per country). The regions were selected so that the widest antigenic diversity was obtained. They were France and West Indies (subtype B), Thailand (subtype E), and four African countries (Burundi, Burkina-Faso, Congo, and Côte d’Ivoire) (subtypes A, C, and D).
Immunoassays.
We have shown that cross-reactivity between V3 sequences is both very strong and very frequent when analyzed by indirect enzyme immunoassays (EIA) (2). We developed an SSEIA that uses the principle of blocking with an excess of peptide in the liquid phase (4). The SSEIA is more discriminating than normal indirect EIA, probably because it is more dependent than indirect EIA on Ab affinity. Two SSEIAs were performed, the first with 5 V3 sequences defined by phylogenetic analysis (SSEIAgen) and the second with 14 V3 sequences defined by phenetic analysis (SSEIAphen).
For the SSEIAgen, wells of polyvinyl microtiter plates (Falcon) were coated with an equimolar mixture of the five V3 peptides (0.5 μg/ml each in 0.05 M bicarbonate buffer [pH 9.6]; 100 μl per well) by incubation for 20 h at 37°C. The wells were washed twice with phosphate-buffered saline containing 0.5% Tween 20 (PBS-TW), and the unoccupied sites of the wells were saturated with phosphate-buffered saline containing 2% newborn calf serum by incubation for 45 min at 37°C. Serum samples were diluted 1:100 in 0.01 M sodium phosphate buffer (pH 7.4) containing 0.75 M NaCl, 10% newborn calf serum, and 0.05% Tween 20 (PBS-TW-NBCS). Each sample was tested in seven wells in the presence of various blocking solutions. Preliminary assays were used to select the optimal concentrations of peptides to be used for both coating solutions and blocking solutions (4). Ten microliters of a 100-μg/ml solution (in PBS-TW-NBCS) of V3 peptide of the A, B, C, D, or E subtype was added to the wells (A to well 1, B to well 2, and so forth). Ten microliters of a 100-μg/ml solution of an equimolar mixture of the five peptides (theoretically 100% blocking) was added to well 6. Ten microliters of PBS-TW-NBCS (theoretically 0% blocking) was added to well 7. One hundred microliters of diluted serum was added to each well and incubated for 30 min at room temperature, and the wells were washed four times with PBS-TW. Peroxidase-conjugated goat F(ab′)2 anti-human immunoglobulin (TAGO, Burlingame, Calif.; 100 μl of a 1:5,000 dilution in PBS-TW-NBCS) was added, and the mixture was incubated for 30 min at room temperature. The wells were washed four times with PBS-TW and incubated with hydrogen peroxide–o-phenylenediamine for 15 min at room temperature. Color development was stopped with 2 N H2SO4, and the absorbance value (optical density [OD]) was read at 492 nm.
The percent inhibition of binding induced by each of the five peptides for each serum sample was calculated with the following formula: {(OD without blocking [well 7] − OD in the presence of peptide)/(OD without blocking [well 7] − OD in the presence of the five peptides (well 6)]} ×100.
The SSEIAgen indicates, for each serum sample, the immunodominant subtype (the peptide with the strongest blocking) as well as a serological profile defined by the five values of inhibition (percent inhibition by peptide A, peptide B, peptide C, peptide D, and peptide E).
The same experimental conditions were used for the SSEIAphen, except that wells were coated with an equimolar mixture of the 14 V3 peptides (0.2 μg/ml each in 0.05 M bicarbonate buffer [pH 9.6]; 100 μl per well). Each serum sample diluted 1:100 was tested in 16 wells in the presence of various blocking solutions. Ten microliters of a 100-μg/ml solution (in PBS-TW-NBCS) each of peptides 1 to 14 was added to wells 1 to 14 (peptide 1 to well 1, peptide 2 to well 2, and so forth). Ten microliters of a 100-μg/ml solution of an equimolar mixture of the 14 peptides (theoretically 100% blocking) was added to well 15. Ten microliters of PBS-TW-NBCS (theoretically 0% blocking) was added to well 16. The remaining steps were identical to those for SSEIAgen. The percent inhibition of binding induced by each of the 14 peptides for each serum sample was calculated with the following formula: [OD without blocking (well 16) − OD in the presence of peptide]/[OD without blocking (well 16) − OD in the presence of the 14 peptides (well 15)] × 100.
The SSEIAphen indicates, for each serum sample, the immunodominant subtype (the peptide with the strongest blocking) as well as a serological profile defined by the 14 values of inhibition.
Statistical analysis.
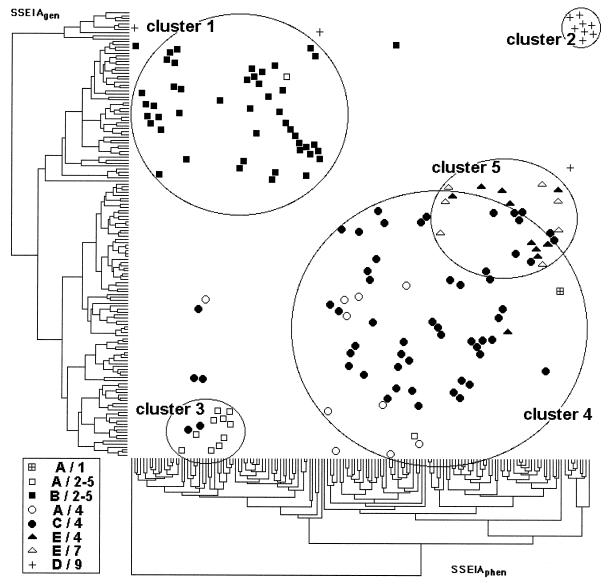
The antigenic relationship between the various peptides was studied by calculating the Pearson correlation matrix with Systat statistical software (Deltasoft, Meylan, France). For each SSEIAgen and SSEIAphen, specimens were grouped according to a similar serological profile as defined above. We used the cluster analysis of PCSM statistical software (Deltasoft). The serological profiles of all the reactive samples for each assay were entered into a computer. We used agglomerative hierarchical clustering of observations, taking into account the euclidean distance and the average linkage. The results were displayed as dendrograms. The two dendrograms obtained were drawn on x (SSEIAphen) and y (SSEIAgen) axes (see Fig. 2). Each sample was plotted at the intersection of its x and y positions (at the intersection of its positions within each dendrogram).
FIG. 2.
Result of the cluster analysis. Each sample, represented by a symbol, is located at the intersection between the two dendrograms. The symbols correspond to the dominant reactivities in both assays. Clusters correspond to samples with similar dominant reactivities in both classifications, with only a few exceptions.
RESULTS
Antigenicity of the V3 peptides.
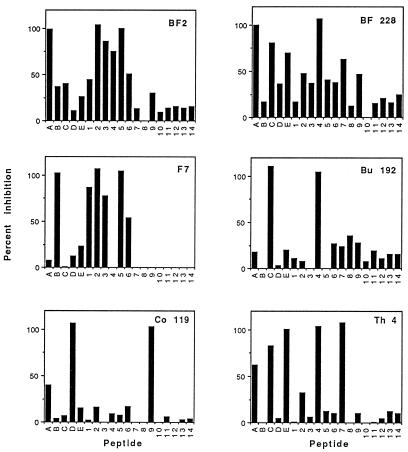
Among the 168 sera positive for the HIV-1 antibody, 154 were reactive with V3 peptides (Table 2). All the sera that reacted in the SSEIAgen were also reactive in the SSEIAphen. The nonreactive samples were negative in both assays. The serum samples collected in France or the West Indies were reactive to peptide B, most of those collected in Thailand were reactive to peptide E, and various reactivities were observed with African samples. Of the 86 reactive African samples, 22 (25.6%) were most reactive with peptide A, 1 (1.2%) was most reactive with peptide B, 53 (61.6%) were most reactive with peptide C, and 10 (11.6%) were most reactive with peptide D. The most striking observation with the SSEIAphen was the limited number of dominant reactivities. Indeed, although 14 peptides were included, only 6 were bound preferentially by the tested sera. The dominant reactivities involved peptides 1 (1 case), peptides 2 and 5 (59 cases), peptide 4 (76 cases), peptide 7 (7 cases), and peptide 9 (11 cases) (Table 3). All the B-reactive serum samples were reactive predominantly to peptides 2 and 5, the same intensity of binding being observed with these two very similar sequences. Of the 47 B- or peptide 2- or 5-reactive samples, 23 cross-reacted with peptides 1, 3, and 6 but not with other peptides. An example is shown in Fig. 1 (sample F7). All the C-reactive samples reacted with peptide 4, with no cross-reactivity. All the D-reactive samples reacted with peptide 9, with no cross-reactivity. The E-reactive samples reacted with either peptide 4 (10 cases) or peptide 7 (7 cases). The most diverse reactivities were observed with A-reactive samples, which bound preferentially to peptide 1 (1 case), 2 or 5 (12 cases), or 4 (10 cases), with various degrees of cross-reactivity (Table 3). Representative serological profiles of serum samples are shown in Fig. 1.
TABLE 2.
Reactivity of serum samples to subtype-specific V3 consensus sequences in SSEIAgen according to geographical origin
| Peptide | No. of serum samples reactive ina:
|
||||||
|---|---|---|---|---|---|---|---|
| WI | Fr | CI | BF | Co | Bu | Th | |
| A | 2 | 9 | 10 | 1 | 1 | ||
| B | 24 | 22 | 1 | ||||
| C | 20 | 7 | 7 | 19 | 3 | ||
| D | 2 | 7 | 1 | 1 | |||
| E | 17 | ||||||
| NRb | 2 | 7 | 3 | 2 | |||
WI, West Indies; Fr, metropolitan France; BF, Burkina-Faso; CI, Côte d’Ivoire; Co, Congo; Bu, Burundi; Th, Thailand.
NR, nonreactive.
TABLE 3.
Correlation between SSEIAgen and SSEIAphen
| Peptide in SSEIAgen | No. of sera reactive to peptidea in SSEIAphen
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 (None) | 2 or 5 (None) | 2 or 5 (1, 3, or 6) | 2 or 5 (4) | 4 (None) | 4 (2 or 5) | 4 (7) | 7 (None) | 7 (4) | 9 (None) | Total | |
| A | 1 | 11 | 1 | 8 | 2 | 23 | |||||
| B | 24 | 23 | 47 | ||||||||
| C | 56 | 56 | |||||||||
| D | 11 | 11 | |||||||||
| E | 8 | 2 | 5 | 2 | 17 | ||||||
Dominant reactivity (cross-reactivity). Dominant reactivity produces 100% blocking. Cross-reactivity occurs when blocking is between 95 and 100%.
FIG. 1.
Serological profiles in the two SSEIAs of typical HIV-1-positive sera from various regions. Shown are two A-reactive sera, BF2 reacting predominantly to peptides 2 and 5 and BF 228 reacting predominantly to peptide 4; a B-reactive sample (F7); a C-reactive sample (Bu 192); a D-reactive sample (Co 119); and an E-reactive sample (Th 4).
The antigenic relationships among these peptides were further analyzed by a statistical approach. The serological profiles of every serum sample, defined as the percentages of binding inhibition by each of the 19 peptides, were entered into a computer, and a Pearson correlation matrix was established (Table 4). This analysis showed positive correlations between sequence B and sequences 1, 2, 3, 5, and 6 (r, 0.69 to 0.85), between sequences C and 4 (r, 0.66), between sequences D and 9 (r, 0.67), and between sequences E and 7 (r, 0.67). No significant correlation was observed for A-reactive serum samples, consistent with the heterogeneity of responses for these samples. The matrix also yielded negative correlation coefficients, showing that reactivity to some sequences systematically excluded reactivity to others. These results show the value of SSEIA for serological discrimination. For example, reactivity to peptide 1 strongly correlated with reactivity to peptides 2 (r, 0.87), 3 (r, 0.75), 5 (r, 0.89), 6 (r, 0.89), and B (r, 0.85) and with no or low reactivity to peptides 4 (r, −0.45), 7 (r, −0.55), 9 (r, −0.37), C (r, −0.35), and E (r, −0.33).
TABLE 4.
Pearson correlation matrix for the antigenicities of the 19 V3 sequences
| Peptide |
ra for SSEIAphen peptide
|
|||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | |
| SSEIAphen | ||||||||||||||
| 1 | 1 | |||||||||||||
| 2 | 0.87a | 1 | ||||||||||||
| 3 | 0.75 | 0.79 | 1 | |||||||||||
| 4 | −0.45 | −0.36 | −0.23 | 1 | ||||||||||
| 5 | 0.89 | 0.96 | 0.79 | −0.36 | 1 | |||||||||
| 6 | 0.89 | 0.88 | 0.74 | −0.42 | 0.90 | 1 | ||||||||
| 7 | −0.55 | −0.58 | −0.46 | 0.70 | −0.55 | −0.49 | 1 | |||||||
| 8 | 0.14 | 0.10 | 0.01 | 0.02 | 0.08 | 0.15 | −0.03 | 1 | ||||||
| 9 | −0.37 | −0.44 | −0.41 | −0.03 | −0.42 | −0.34 | 0.09 | 0.01 | 1 | |||||
| 10 | 0.24 | 0.15 | 0.21 | −0.09 | 0.18 | 0.22 | −0.06 | 0.45 | 0.09 | 1 | ||||
| 11 | −0.16 | −0.24 | −0.13 | 0.14 | −0.21 | −0.15 | 0.29 | 0.16 | 0.41 | 0.42 | 1 | |||
| 12 | −0.30 | −0.32 | −0.23 | 0.30 | −0.30 | −0.27 | −0.41 | 0.15 | 0.34 | 0.19 | 0.77 | 1 | ||
| 13 | −0.29 | −0.33 | −0.22 | 0.30 | −0.31 | −0.26 | 0.35 | 0.03 | 0.34 | 0.18 | 0.73 | 0.73 | 1 | |
| 14 | −0.16 | −0.14 | −0.16 | 0.19 | −0.15 | −0.10 | 0.28 | 0.17 | 0.36 | 0.22 | 0.63 | 0.70 | 0.70 | 1 |
| SSEIAgen | ||||||||||||||
| A | 0.20 | 0.20 | 0.08 | −0.11 | 0.17 | 0.16 | −0.11 | −0.08 | −0.03 | 0.00 | −0.11 | −0.11 | −0.01 | 0.04 |
| B | 0.85 | 0.79 | 0.69 | −0.58 | 0.81 | 0.81 | −0.62 | 0.06 | −0.39 | 0.19 | −0.25 | −0.35 | −0.34 | −0.29 |
| C | −0.35 | −0.19 | −0.07 | 0.66 | −0.21 | −0.28 | 0.47 | 0.07 | −0.32 | −0.13 | 0.05 | 0.10 | 0.11 | 0.08 |
| D | 0.02 | −0.04 | −0.11 | −0.50 | −0.03 | 0.10 | −0.28 | −0.01 | 0.67 | 0.02 | 0.16 | 0.09 | 0.08 | 0.16 |
| E | −0.33 | −0.31 | −0.23 | 0.38 | −0.33 | −0.21 | 0.67 | 0.09 | −0.04 | −0.03 | 0.21 | 0.28 | 0.27 | 0.28 |
Correlations in bold were significant (P < 0.05) after Bonferroni correction.
Cluster analysis.
Groupings among the 154 reactive samples were identified by cluster analysis. Results are displayed as two dendrograms corresponding to results obtained with SSEIAgen and SSEIAphen (Fig. 2). This analysis demonstrated a correlation between the two classifications and identified five serological clusters, two of which overlap. Cluster 1 contains samples from France and the West Indies reactive with peptides B, 2, and 5. Cluster 2 contains African samples reactive with peptides D and 9. Cluster 3 contains African samples reactive mostly with peptides A, 2, and 5. Clusters 4 and 5, which overlap, contain samples mainly from Africa reactive mostly with peptides C and 4 and samples mainly from Thailand reactive with peptides E and 7, respectively. The overlapping area contains samples with high cross-reactivity with peptides C and E and/or peptides 4 and 7. Cluster 4 also contains A-reactive samples reactive mostly with peptide 4.
Amino acids responsible for subtype-specific serological identification.
We compared the amino acid sequences of the 19 peptides used in the study. The comparison was based on both antigenicity in SSEIA, i.e., positive or negative correlation as described above, and biochemical properties, i.e., conservative or nonconservative substitutions. We identified three major groups of independent (antigenically unrelated) reactive sequences from the SSEIA data (Table 5). The first comprised peptides B, 2, and 5. The second comprised peptides A, C, E, 4, and 7. The third comprised peptides D and 9. The major difference between the first group and the others is the presence at the tip of the loop of an arginine residue at position 322 in the first group. This arginine is replaced by glutamine in the others. The major difference between the second and the third groups occurs at position 324, where there is a leucine residue in peptides D and 9 but a more hydrophobic residue, phenylalanine, in peptides A, C, E, 4, and 7 (5). There is another, smaller difference between the second group and the two others at position 315. A histidine residue in the first and third groups is replaced by an arginine or a threonine residue in the second group, except in the consensus sequence of subtype A, which conserves this histidine.
TABLE 5.
Contribution of individual residues to serotype specificity
| Parameter | Result for sequencea
|
||||||||
|---|---|---|---|---|---|---|---|---|---|
| 315 | 316 | 319 | 320 | 321 | 322 | 323 | 324 | 325 | |
| x | I | G | P | G | x | x | x | Y | |
| Genetic subtype/phenetic group(s)b | |||||||||
| B/2–5 | H | —d | — | — | — | R | A | F | — |
| A/4 | H/R | — | — | — | — | Q | A | F | — |
| C | R | — | — | — | — | Q | T | F | — |
| E/7 | T | — | — | — | — | Q | V | F | — |
| D/9 | H | — | — | — | — | Q | A | L | — |
| No. of Van der Waals contactsc | 0 | 3 | 1 | 14 | 4 | 14 | 9 | 17 | 0 |
| No. of hydrogen bondsc | 0 | 0 | 1 | 0 | 1 | 2 | 1 | 0 | 0 |
| No. of salt bridgesc | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 |
| Peptide surface area buried (Å2)c | 6 | 52 | 28 | 77 | 23 | 86 | 53 | 93 | 2 |
DISCUSSION
Studies on HIV-1 diversity led to the identification of five major genetic subtypes within group M and at least 14 phenetically associated groups (22, 36). The analysis of genetic relationships between isolates is complicated, as many isolates are mosaic viruses (15, 19, 44). Serotyping of HIV-1 infections based on V3 seroreactivity would be of great value in the study of HIV-1 diversity and would provide important information for both epidemiology and future vaccine composition. This work studied the antigenic relationships between consensus sequences defined by two different approaches. The aim was to investigate the extent of the V3 serotypes and to identify, at least in part, the molecular determinants of the serotype-associated antigenic specificity. Our experimental data confirmed most of the associations between the genetic subtypes and phenetic groups that were predicted by theoretical analysis (22). However, our data showed that only a limited number of V3 serotypes can be identified due to the immunodominance of a few sequences. Korber et al. suggested that genetic subtype B was associated mainly with phenetic groups 1, 2, 3, 5, and 6 (22). Our results are consistent with these results but showed that the consensus sequences of groups 2 and 5 are immunodominant. Serological discrimination of these two sequences was almost impossible due to a single synonymous substitution (glutamic acid→aspartic acid at position 329; Table 1). We also found associations between genetic subtype C and phenetic group 4, genetic subtype D and phenetic group 9, and genetic subtype E and phenetic group 7. However, our data also suggested that consensus sequence 7 is immunodominant for subtype E and that consensus sequence 9 is immunodominant for subtype D. The phenogram of HIV-1 V3 loop protein similarities indicated that for subtype A, the association between genetic subtype and phenetic group was weaker (22). Our results are also consistent with this observation. Indeed, some subtype A-reactive samples bound to peptides containing the consensus sequences of phenetic group 2 or 5, whereas others bound to peptides containing the consensus sequences of phenetic group 4. This may be because most isolates of subtype A have the sequence GPGQ at the tip of the V3 loop but others possess the sequence GPGR. Genetic subtypes C and E appear distantly related, but cross-reactivity between peptides containing their V3 consensus sequences has been reported (4, 8, 40). We also found cross-reactivity between subtypes C and E based on data obtained with consensus sequences of phenetic groups 4 and 7.
V3 consensus sequences can be grouped into three to five major antigenically unrelated categories on the basis of our statistical data, including correlation matrix and cluster analyses. A comparison of protein sequences in these categories identified residues at positions 322 to 324 that may be responsible for major antigenic differences (Table 5). This result is consistent with previous data showing that human antibodies to the V3 region are mostly directed to the central area of V3 (51). The crystal structure of a complex between a V3 peptide and the Fab fragment of a neutralizing antibody, 59.1, showed that amino acids at the these three positions are involved in Fab binding (16). Table 5 shows the sequences responsible for serotype specificity and the contribution of individual residues to antibody interactions according to Ghiara et al. (16). Two-thirds of the hydrogen bonds, salt bridges, and Van der Waals contacts between V3 and Fab involve residues 322 to 324. Sherefa et al. used substitution peptide analogs to show that the lack of cross-reactivity between subtype B and subtype C peptides was due to an arginine-322→glutamine-322 substitution (46). They also found that the phenylalanine at position 324 was also essential for subtype specificity and showed that an alanine-323→threonine-323 substitution, which would not cause any major structural changes in the peptide backbone, might explain the cross-reactivities among peptides A, C, and 4. In another study, Sherefa et al. compared V3 serotyping and genotyping by V3 sequencing (47). They obtained results similar to ours. Amino acids present upstream at positions 315 and 316, which are part of the core of another V3 epitope recognized by human sera, might play a complementary role (6, 17, 18, 43). This fact may explain why some subtype A-reactive samples bound preferentially to peptide 1, 2 or 5, or 4 with various degrees of cross-reactivity. Peptides A, 1, 2, and 5 have a histidine residue at position 315, whereas peptide 4 has an arginine. Thus, the subtype-specific signature sequences are located at the tip of the V3 loop and involve the octapeptide xIGPGxxx. The immunodominance of some peptides within a “serogroup” may be due to slight modifications within the flanking residues that affect the conformation (Table 1).
Our study did not include samples from patients in whom the genotype of the infecting strain had been identified. We believed that this information was not relevant to the present analysis, as previous studies have shown a strong correlation between SSEIAgen and genotyping. In three independent studies involving 285 samples, the specificities of SSEIAgen were between 0.96 and 0.98 for subtype A, 0.84 and 0.98 for subtype B, 0.74 and 0.81 for subtype C, and 0.99 and 1 for subtype D; the specificity was 1 for subtype E (6a, 48).
Multivariate analysis of HIV-1 neutralization data shows the existence of a small number of neutralization clusters not correlated with the genetic clades (39). The neutralization epitopes responsible for this clustering have not been identified and may be independent of the V3 region, but this previous study and our study, addressing different questions and using different approaches, have both suggested the existence of a limited number of serotypes despite the genetic diversity of HIV-1.
Surveys of the diversity and dynamics of HIV-1 strains are an important challenge in the current AIDS pandemic and for the future. There are at least two reasons for typing: (i) surveillance of diversity, particularly its dynamics for epidemiological studies, and (ii) understanding the biological significance of this diversity. Both are essential for public health. Genotyping is probably the reference method for epidemiological surveillance of HIV-1 diversity, but current molecular methods cannot be used for large populations. Alternative methods, such as serotyping with simple immunoassays, would be of great value. Our study provides evidence for the existence of a small number of V3 serotypes. The statistical analysis indicated strong positive and negative correlations between the different sequences. This result indicates the discriminative power and reliability of SSEIA. The biological significance of HIV-1 diversity is unknown, but immunity in vivo is probably the most important property. For many viruses, such as poliovirus or influenza viruses, protection in vivo is associated with seroneutralization in vitro. Immunity in vitro does not correlate with protection in vivo for HIV-1, but a few authors have studied the association between neutralization serotypes and genotypes. Moore et al. clearly showed that genetic clades do not correlate with neutralization serotypes, indicating that genotyping may not be the ideal method for studying HIV-1 diversity for vaccine development purposes (33). It is therefore still unclear what assays or tools might be most adaptable for predicting the antigenic composition of future vaccines. The biological relevance of the V3 serotypes is unknown. However, it is unclear which methods are best suited, and all approaches to studying diversity require further evaluation.
ACKNOWLEDGMENTS
This work was supported by funds from the Agence Nationale de Recherche sur le SIDA (Paris, France) and the Institut Universitaire de France.
We thank J. Hoebeke for helpful discussion of the manuscript and R. Chout, F. Denis, and V. Vithayasai for providing serum samples.
REFERENCES
- 1.Ariyoshi K, Cheingsong-Popov R, Wilkins A, Corrah T, Weber J, Whittle H. HIV-1 subtype B in West Africa. Lancet. 1996;347:328. doi: 10.1016/s0140-6736(96)90503-6. . (Letter.) [DOI] [PubMed] [Google Scholar]
- 2.Baillou A, Brand D, Denis F, M’Boup S, Chout R, Goudeau A, Barin F. High antigenic cross-reactivity of the V3 consensus sequences of HIV-1 gp120. AIDS Res Hum Retroviruses. 1994;9:1209–1215. doi: 10.1089/aid.1993.9.1209. [DOI] [PubMed] [Google Scholar]
- 3.Bandea C I, Ramos A, Pieniazek D, Pascu R, Tanuri A, Schochetman G, Rayfield M A. Epidemiologic and evolutionary relationships between Romanian and Brazilian HIV-1 subtype F strains. Emerg Infect Dis. 1995;1:91–93. doi: 10.3201/eid0103.950305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Barin F, Lahbabi Y, Buzelay L, Lejeune B, Baillou-Beaufils A, Denis F, Mathiot C, M’Boup S, Vithayasai V, Dietrich U, Goudeau A. Diversity of antibody binding to V3 peptides representing consensus sequences of HIV type 1 genotypes A to E: an approach for HIV type 1 serological subtyping. AIDS Res Hum Retroviruses. 1996;12:1279–1289. doi: 10.1089/aid.1996.12.1279. [DOI] [PubMed] [Google Scholar]
- 5.Black S D, Mould D R. Development of hydrophobicity parameters to analyze proteins which bear post- or co-translational modifications. Anal Biochem. 1991;193:72–82. doi: 10.1016/0003-2697(91)90045-u. [DOI] [PubMed] [Google Scholar]
- 6.Catasti P, Fontenot J D, Bradbury E M, Gupta G. Local and global structural properties of the HIV-MN V3 loop. J Biol Chem. 1995;270:2224–2232. doi: 10.1074/jbc.270.5.2224. [DOI] [PubMed] [Google Scholar]
- 6a.Cheingsong-Popov, R. Unpublished data.
- 7.Cheingsong-Popov R, Bobkov A, Garaev M M, Kaleebu P, Callow P, Rzhiminova A, Saukhat S R, Burdajev N P, Kolomijets N D, Weber J. Identification of human immunodeficiency virus type 1 subtypes and their distribution in the Commonwealth of Independent States (former Soviet Union) by serologic V3 peptide-binding assays and V3 sequence analysis. J Infect Dis. 1993;168:292–297. doi: 10.1093/infdis/168.2.292. [DOI] [PubMed] [Google Scholar]
- 8.Cheingsong-Popov R, Lister S, Callow D, Kaleebu P, Beddows S, Weber J the WHO Network for HIV Isolation and Characterization. Serotyping HIV type 1 by antibody binding to the V3 loop: relationship to viral genotype. AIDS Res Hum Retroviruses. 1994;11:1379–1386. doi: 10.1089/aid.1994.10.1379. [DOI] [PubMed] [Google Scholar]
- 9.Clewley J P, Arnold C, Barlow K L, Grant P R, Parry J V. Diverse HIV-1 genetic subtypes in UK. Lancet. 1996;347:1487. doi: 10.1016/s0140-6736(96)91724-9. . (Letter.) [DOI] [PubMed] [Google Scholar]
- 10.Delwart E L, Shpaer E G, McCutchan F E, Louwagie J, Grez M, Rübsamen-Waigmann H, Mullins J I. Genetic relationships determined by a heteroduplex mobility assay: analysis of HIV env genes. Science. 1993;262:1257–1261. doi: 10.1126/science.8235655. [DOI] [PubMed] [Google Scholar]
- 11.Delwart E L, Sheppard H W, Walker B D, Goudsmit J, Mullins J I. Human immunodeficiency virus type 1 evolution in vivo tracked by DNA heteroduplex mobility assays. J Virol. 1994;68:6672–6683. doi: 10.1128/jvi.68.10.6672-6683.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dietrich U, Grez M, von Briesen H, Panhans B, Geissendörfer M, Kuhnel H, Maniar J, Manhambre G, Becker W B, Bechler M L B, Rübsamen-Waigmann H. Human immunodeficiency virus type 1 strains from India are highly divergent from prototypic African and U.S./European strains but are linked to a South African isolate. AIDS. 1993;7:23–27. doi: 10.1097/00002030-199301000-00003. [DOI] [PubMed] [Google Scholar]
- 13.Fransen K, Buvé A, Nkengasong J N, Laga M, van der Groen G. Longstanding presence in Belgians of multiple non-B HIV-1 subtypes. Lancet. 1996;347:1403. doi: 10.1016/s0140-6736(96)91042-9. . (Letter.) [DOI] [PubMed] [Google Scholar]
- 14.Gao F, Yue L, Hill S C, Robertson D L, Graves A H, Saag M S, Shaw G M, Sharp P M, Hahn B H. HIV-1 sequence subtype D in the United States. AIDS Res Hum Retroviruses. 1994;10:625–627. doi: 10.1089/aid.1994.10.625. [DOI] [PubMed] [Google Scholar]
- 15.Gao F, Robertson D L, Morrison S G, Hui H, Craig S, Decker J, Fultz P N, Girard M, Shaw G M, Hahn B H, Sharp P M. The heterosexual human immunodeficiency virus type 1 epidemic in Thailand is caused by an intersubtype (A/E) recombinant of African origin. J Virol. 1996;70:7013–7029. doi: 10.1128/jvi.70.10.7013-7029.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ghiara J B, Stura E A, Stanfield R L, Profy A T, Wilson I A. Crystal structure of the principal neutralization site of HIV-1. Science. 1994;264:82–85. doi: 10.1126/science.7511253. [DOI] [PubMed] [Google Scholar]
- 17.Gorny M K, Xu J-Y, Gianakakos V, Karwoswka S, Williams C, Sheppard H W, Hanson C V, Zolla-Pazner S. Production of site-selected neutralizing human monoclonal antibodies against the third variable domain of the human immunodeficiency virus type 1 envelope glycoprotein. Proc Natl Acad Sci USA. 1991;88:3238–3242. doi: 10.1073/pnas.88.8.3238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Gorny M K, Conley A J, Karwoswka S, Buchbinder A, Xu J-Y, Emini E A, Koenig S, Zolla-Pazner S. Neutralization of diverse human immunodeficiency virus type 1 variants by an anti-V3 human monoclonal antibody. J Virol. 1992;66:7538–7542. doi: 10.1128/jvi.66.12.7538-7542.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Gürtler L G, Hauser P H, Eberle J, von Brunn A, Knapp S, Zekeng L, Tsague J M, Kaptue L. A new subtype of human immunodeficiency virus type 1 (MVP-5180) from Cameroon. J Virol. 1994;68:1581–1585. doi: 10.1128/jvi.68.3.1581-1585.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hahn B H, Robertson D L, Sharp P M. Intersubtype recombination in HIV-1 and HIV-2. In: Myers G, Korber B, Wain-Hobson S, Jeang K-T, Henderson L E, Pavlakis G N, editors. Human retroviruses and AIDS 1995: a compilation and analysis of nucleic acid and amino acid sequences. Los Alamos, N.Mex: Los Alamos National Laboratory; 1995. pp. III-22–III-29. [Google Scholar]
- 21.Janssens W, Heyndrickx L, Fransen K, Motte J, Peeters M, Nkengasong J N, Ndumbe P M, Delaporte E, Perret J L, Atende C, Piot P, van der Groen G. Genetic and phylogenetic analysis of env subtypes G and H in Central Africa. AIDS Res Hum Retroviruses. 1994;10:877–879. doi: 10.1089/aid.1994.10.877. [DOI] [PubMed] [Google Scholar]
- 22.Korber B T M, McInnes K, Smith R F, Myers G. Mutational trends in V3 loop protein sequences observed in different genetic lineages of human immunodeficiency virus type 1. J Virol. 1994;68:6730–6744. doi: 10.1128/jvi.68.10.6730-6744.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kostrikis L G, Bagdades E, Cao Y, Zhang L, Dimitriou D, Ho D D. Genetic analysis of human immunodeficiency virus type 1 strains from patients in Cyprus: identification of a new subtype designated subtype I. J Virol. 1995;69:6122–6130. doi: 10.1128/jvi.69.10.6122-6130.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kuiken C L, Zwart G, Baan E, Coutinho R A, van den Hoek J A R, Goudsmit J. Increasing antigenic and genetic diversity of the V3 variable domain of the human immunodeficiency virus envelope protein in the course of the AIDS epidemic. Proc Natl Acad Sci USA. 1993;90:9061–9065. doi: 10.1073/pnas.90.19.9061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kunanusont C, Foy H M, Kreiss J K, Rersk-Ngarm S, Phanuphak P, Raktham S, Pau C P, Young N. HIV-1 subtypes and male-to-female transmission in Thailand. Lancet. 1995;345:1078–1083. doi: 10.1016/s0140-6736(95)90818-8. [DOI] [PubMed] [Google Scholar]
- 26.LaRosa G J, Davide J P, Weinhold K, Waterbury J A, Profy A T, Lewis J A, Langlois A J, Dreesman G R, Boswell R N, Shadduck P, Holley L H, Karplus M, Bolognesi D P, Matthews T J, Emini E A, Putney S. Conserved sequence and structural elements in the HIV-1 principal neutralization determinant. Science. 1990;249:932–935. doi: 10.1126/science.2392685. [DOI] [PubMed] [Google Scholar]
- 27.Loussert-Ajaka I, Chaix M L, Korber B, Letourneur F, Gomas E, Allen E, Ly T D, Brun-Vézinet F, Simon F, Saragosti S. Variability of human immunodeficiency virus type 1 group O strains isolated from Cameroonian patients living in France. J Virol. 1995;69:5640–5649. doi: 10.1128/jvi.69.9.5640-5649.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Louwagie J, Delwart E L, Mullins J I, McCutchan F E, Eddy G, Burke D S. Genetic analysis of HIV-1 isolates from Brazil reveals presence of two distinct genetic subtypes. AIDS Res Hum Retroviruses. 1994;10:561–567. doi: 10.1089/aid.1994.10.561. [DOI] [PubMed] [Google Scholar]
- 29.Luo C C, Tian C, Hu D J, Kai M, Dondero T J, Zheng X. HIV-1 subtype C in China. Lancet. 1995;345:1051–1052. doi: 10.1016/s0140-6736(95)90792-0. [DOI] [PubMed] [Google Scholar]
- 30.McCutchan F E, Hegerich P A, Brennan T P, Phanuphak P, Singharaj P, Jugudee A, Berman P W, Gray A M, Fowler A K, Burke D S. Genetic variants of HIV-1 in Thailand. AIDS Res Hum Retroviruses. 1992;8:1887–1895. doi: 10.1089/aid.1992.8.1887. [DOI] [PubMed] [Google Scholar]
- 31.Merrifield R B. Solid-phase peptide synthesis. I. The synthesis of a tetrapeptide. J Am Chem Soc. 1963;85:2149–2154. [Google Scholar]
- 32.Montpetit M. HIV-1 subtype in Canada. AIDS Res Hum Retroviruses. 1995;11:1421–1422. doi: 10.1089/aid.1995.11.1421. [DOI] [PubMed] [Google Scholar]
- 33.Moore J P, Cao Y, Leu J, Qui L, Korber B, Ho D D. Inter- and intraclade neutralization of human immunodeficiency virus type 1: genetic clades do not correspond to neutralization serotypes but partially correspond to gp120 antigenic serotypes. J Virol. 1996;70:427–444. doi: 10.1128/jvi.70.1.427-444.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Morgado M G, Sabino E C, Shpaer E G, Bongertz V, Brigido L, Guimaraes M D C, Castilho E A, Galvao-Castro B, Mullins J I, Hendry R M, Mayer A. V3 region polymorphisms in HIV-1 from Brazil: prevalence of subtype B strains divergent from North American/European prototype and detection of subtype F. AIDS Res Hum Retroviruses. 1994;10:569–576. doi: 10.1089/aid.1994.10.569. [DOI] [PubMed] [Google Scholar]
- 35.Murphy E E, Korber B, Georges-Courbot M C, You B, Pinter A, Cook D, Kieny M P, Georges A, Mathiot C, Barré-Sinoussi F, Girard M. Diversity of V3 region sequences of human immunodeficiency viruses type 1 from the Central African Republic. AIDS Res Hum Retroviruses. 1993;9:997–1006. doi: 10.1089/aid.1993.9.997. [DOI] [PubMed] [Google Scholar]
- 36.Myers G, Korber B, Wain-Hobson S, Smith R F, Pavlakis G N. Human retroviruses and AIDS 1992. Los Alamos, NM: a compilation and analysis of nucleic acid and amino acid sequences. Los Alamos, N.Mex: Los Alamos National Laboratory; 1992. [Google Scholar]
- 37.Myers G, Korber B, Wain-Hobson S, Smith R F, Pavlakis G N. Human retroviruses and AIDS 1994. Los Alamos, NM: a compilation and analysis of nucleic acid and amino acid sequences. Los Alamos, N.Mex: Los Alamos National Laboratory; 1994. [Google Scholar]
- 38.Nkengasong J N, Janssens W, Heyndrickx L, Fransen K, Ndumbe P M, Motte J, Leonaers A, Ngolle M, Ayuk J, Piot P, van der Groen G. Genotypic subtypes of HIV-1 in Cameroon. AIDS. 1994;8:1405–1412. doi: 10.1097/00002030-199410000-00006. [DOI] [PubMed] [Google Scholar]
- 39.Nyambi P N, Nkengasong J N, Lewi P, Andries K, Janssens W, Fransen K, Heyndrickx L, Piot P, van der Groen G. Multivariate analysis of human immunodeficiency virus type 1 neutralization data. J Virol. 1996;70:6235–6243. doi: 10.1128/jvi.70.9.6235-6243.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Pau C P, Kai M, Holloman-Candal D L, Luo C C, Kalish M L, Schochetman G, Byers B, George J R the WHO Network for HIV Isolation and Characterization. Antigenic variation and serotyping of HIV type 1 from four World Health Organisation-sponsored HIV vaccine sites. AIDS Res Hum Retroviruses. 1994;11:1369–1377. doi: 10.1089/aid.1994.10.1369. [DOI] [PubMed] [Google Scholar]
- 41.Pfützner A, Dietrich U, von Eichel U, von Briesen H, Brede H D, Maniar J K, Rübsamen-Waigmann H. HIV-1 and HIV-2 infections in a high-risk population in Bombay, India: evidence for the spread of HIV-2 and presence of a divergent HIV-1 subtype. J Acquired Immune Defic Syndr. 1992;5:972–977. [PubMed] [Google Scholar]
- 42.Ratner L, Haseltine W, Patarca R, Livak K J, Starcich B, Josephs S F, Doran E R, Rafalski J A, Whitehorn E A, Baumeister K, Ivanoff L, Petteway S R, Jr, Pearson M L, Lautenberger J A, Papas T S, Ghrayeb J, Chang N T, Gallo R C, Wong-Staal F. Complete nucleotide sequence of the AIDS virus, HTLV-III. Nature. 1985;313:277–284. doi: 10.1038/313277a0. [DOI] [PubMed] [Google Scholar]
- 43.Rini J M, Stanfield R L, Stura E A, Salinas P A, Profy A T, Wilson I A. Crystal structure of a human immunodeficiency virus type 1 neutralizing antibody, 50.1, in complex with its V3 loop peptide antigen. Proc Natl Acad Sci USA. 1993;90:6325–6329. doi: 10.1073/pnas.90.13.6325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sabino E C, Shpaer E G, Morgado M G, Korber B T M, Diaz R S, Bongertz V, Calvacante S, Galvao-Castro B, Mullins J I, Mayer A. Identification of human immunodeficiency virus type 1 envelope gene recombinants between subtypes B and F in two epidemiologically linked individuals from Brazil. J Virol. 1994;68:6340–6346. doi: 10.1128/jvi.68.10.6340-6346.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sharp P M, Robertson D L, Gao F, Hahn B. Origins and diversity of human immunodeficiency viruses. AIDS. 1994;8:S27–S42. [Google Scholar]
- 46.Sherefa K, Sâllberg M, Sönnerborg A. Evidence of no change in V3 loop antibody recognition pattern in HIV type 1-infected Ethiopians between 1988 and 1993. AIDS Res Hum Retroviruses. 1994;10:1551–1556. doi: 10.1089/aid.1994.10.1551. [DOI] [PubMed] [Google Scholar]
- 47.Sherefa K, Sönnerborg A, Steinbergs J, Sâllberg M. Rapid grouping of HIV-1 infection in subtypes A to E by V3 peptide serotyping and its relation to sequence analysis. Biochem Biophys Res Commun. 1994;205:1658–1664. doi: 10.1006/bbrc.1994.2858. [DOI] [PubMed] [Google Scholar]
- 48.Simon F, Loussert-Ajaka I, Danond F, Saragosti S, Barin F, Brun-Vezinet F. HIV-1 diversity in Northern Paris, France. AIDS Res Hum Retroviruses. 1996;12:1427–1433. doi: 10.1089/aid.1996.12.1427. [DOI] [PubMed] [Google Scholar]
- 49.Smith J D, Bruce C B, Featherstone A S R, Downing R G, Biryahawaho B, Clegg J C S, Carswell J W, Oram J D. Reactions of Ugandan antisera with peptides encoded by V3 loop epitopes of human immunodeficiency virus type 1. AIDS Res Hum Retroviruses. 1994;10:577–583. doi: 10.1089/aid.1994.10.577. [DOI] [PubMed] [Google Scholar]
- 50.Vanden Haeseveld M, Decourt J L, de Luys R J, Vanderborght B, van der Groen G, van Heuverswijn H, Saman E. Genomic cloning and complete sequence analysis of a highly divergent African human immunodeficiency virus isolate. J Virol. 1994;68:1586–1596. doi: 10.1128/jvi.68.3.1586-1596.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Zwart G, Wolf T F W, Bookelman R, Hartman S, Bakker M, Boucher C A B, Kuiken C, Goudsmit J. Greater diversity of the HIV-1 V3 neutralization domain in Tanzania compared with The Netherlands: serological and genetic analysis. AIDS. 1993;7:467–474. doi: 10.1097/00002030-199304000-00003. [DOI] [PubMed] [Google Scholar]