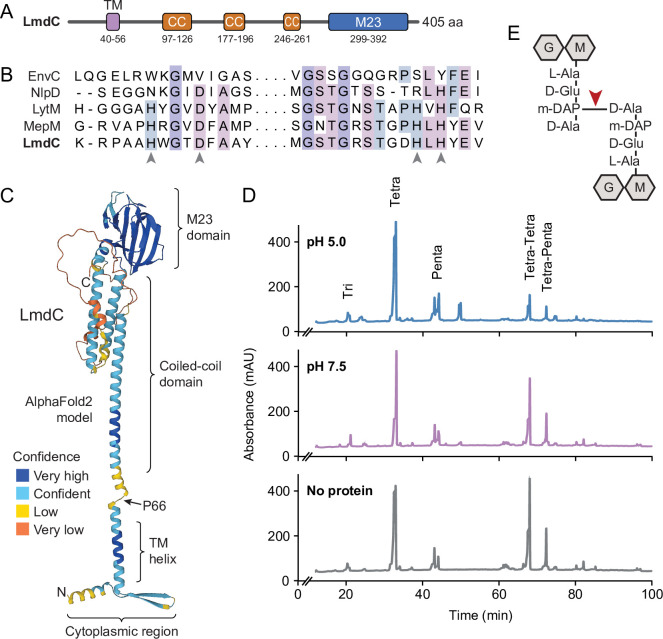
Figure 8. LmdC is a peptidoglycan hydrolase with DD-endopeptidase activity.
(A) Predicted domain architecture of H. neptunium LmdC. The predicted positions of the transmembrane helix (TM), the three coiled-coil regions (CC), and the M23 peptidase domain are indicated. (B) Alignment of the amino acid sequences of multiple M23 peptidases showing the conservation of the catalytic residues in LmdC. Residues required to coordinate the catalytic Zn2+ ion of LytM from S. aureus (Firczuk et al., 2005) are indicated by arrowheads. The proteins shown are EnvC from E. coli (UniProt: P37690), NlpD from E. coli (P0ADA3), LytM from S. aureus (O33599), MepM from E. coli (P0AFS9) and LmdC from H. neptunium (Q0BYX6). (C) Predicted molecular structure of H. neptunium LmdC, generated with Alphafold2 (Jumper et al., 2021). The different domains of the protein and the position of proline 66, which terminates the N-terminal fragment of LmdC used for the in vivo analyses in this study (LmdC1-66), are indicated. (D) Representative HPLC traces showing the muropeptide profile of peptidoglycan treated with LmdC. Cell walls were incubated with the isolated M23 peptidase domain of LmdC at pH 5.0 or pH 7.5. Subsequently, muropeptides were released with cellosyl, reduced, and separated by HPLC. A sample lacking LmdC (No protein) was analyzed as a reference. The nature of the major products is indicated above the peaks. Tri, Tetra, and Penta stand for N-acetylglucosamine-N-acetylmuramitol tripeptide, tetrapeptide, and pentapeptide, respectively. (E) Structure of a Tetra-Tetra muropeptide. Abbreviations: G (N-acetylglucosamine), M (N-acetylmuramic acid). The cleavage site of LmdC is indicated by a red arrowhead.