Abstract
Background
High-grade serous ovarian carcinoma (HGSOC) is a pathologic subtype of ovarian cancer (OC) with a more lethal prognosis. Extensive heterogeneity results in HGSOC being more susceptible to treatment resistance and adverse treatment effects. Revealing the heterogeneity involved is crucial.
Methods
We downloaded the single-cell RNA-seq (scRNA) data from GEO database and performed a scRNA analysis for cell landscape of HGSOC by using the Seurat package. The highly expressed genes were uploaded into the DAVID and KEGG database for enrichment analysis, and the AUCell package was used to calculate cancer-associated hallmark score. The SCENIC analysis was used for key regulons, the estrogen response enrichment scores in TCGA-OV RNA-seq dataset were calculated by using the GSVA package. Besides, the expression of STRA6 and IRF1 and the cell invasion and migration in si-STRA6 OC cells were detected by using the quantitative reverse transcription (qRT)-PCR method and Transwell assay respectively.
Results
We successfully constructed a single-cell atlas of HGSOC and delineated the heterogeneity of epithelial cells therein. There were five epithelial cell subpopulations, GLDC + Epithelial cells, PEG3+ leydig cells, STRA6+ granulosa cells, POLE2+ Epithelial cells, and AURKA + Epithelial cells. STRA6+ granulosa cells have the potential to promote tumor growth as well as the highest estrogen response early activity through the biological pathways analysis of highly expressed genes and estrogen response score of ssGSEA. We found that IRF1 and STRA6 expression was remarkably upregulated in the OC cancer cell line HEY. Silencing of STRA6 markedly decreased the invasion and migration ability of the OC cancer cell line HEY.
Conclusion
There is extreme heterogeneity of epithelial cells in HGSOC, and STRA6+ granulosa cells may be able to promote cancer progression. Our findings are benefit to the heterogeneity identification of HGSOC and develop targeted therapy strategy for HGSOC patients.
Keywords: High-grade serous ovarian carcinoma, Single-cell sequencing, STRA6+ granulosa cells, SCENIC, TFs, GRNs
1. Introduction
Ovarian cancer (OC) is characterized by high malignancy and easy metastasis, with high recurrence and mortality rates, and is the most common malignant tumor in the female genital organs [1,2].
With a morbidity rate of 3.4% and a mortality rate of 4.7%, OC is one of the malignant tumors that threaten women's health and lives [3]. According to the United States cancer statistics report, the estimated new cases of OC are 19,710, the estimated deaths are 13,270 in 2023 [4]. Despite the increasing level of treatment in recent years, the high heterogeneity results in a still poor prognosis for OC, with a survival rate of less than 30% [5]. Conventional treatments include surgery and platinum-based drug therapy, and a combination of the two is also a common treatment option [6,7]. In recent years, a number of novel targeted therapeutic agents have also been promising options, with anti-VEGF antibodies and PARP inhibitors available for patients who meet the criteria [8]. However, the high recurrence profile resulted in more than half of the patients having a recurrence within two years, with little to no improvement in overall survival [9]. Among OCs, high-grade serous ovarian carcinoma (HGSOC) originating from tubal epithelial tissue is the pathologic subtype with the least favorable prognosis, with a survival rate of less than 15% for advanced HGSOC [10]. The variety of HGSOC symptoms and lack of typical specific pathognomonic symptoms make clinical diagnosis a great challenge, and most HGSOC have progressed to advanced stages by the time they are diagnosed [11]. Specific biomarkers are important for diagnosis and improved prognosis.
HGSOC is a malignant tumor whose origin occurs in the ovarian epithelium and is extremely heterogeneous [10]. Therefore, exploring the heterogeneity among epithelial cells in HGSOC is necessary to gain insight into its pathogenesis. And single-cell sequencing (scRNA-seq) technology can reveal the heterogeneity of different cells at the transcriptional level by analyzing single-cell populations [12]. However, most of the existing studies have focused on the classification of specific molecular subpopulations or the study of tumor cell heterogeneity, such as Xu el. revealed that CCNE1-amplified tumors represent a subtype of HGSOC in clinical diagnosis [13], Yang el. demonstrated two immune “cold” patterns in HGSOC through the study of infiltrating T cells [14], and few studies have focused on revealing the heterogeneity of epithelial cells in HGSOC.
In addition, the HGSOC is a complex disease with huge heterogeneity, the patients with distinct histopathologies may exhibit variable responses to treatment [11]. Over the past 50 years, the next-generation sequencing and microarrays technologies have led to the discovery of many key molecular alterations and the cancer subtypes, and enable the further subclassification of these common subtypes, such as the HGSOC was classified into four transcriptional subtypes: “proliferative”, “mesenchymal”, “immunoreactive” and “proliferative” in The Cancer Genome Atlas (TCGA) project [15,16]. The mutational profiling provided valuable insight, such as the ubiquity TP53 mutations of HGSOC [17]. However, the technological advances of NGS also are help for the disease heterogeneity and underlying molecular mechanism exploring. Recently, a scRNA-seq study of primary HGSOC unraveled the immune cell subsets signatures in tumor microenvironment (TME), the identified IRF8+, CD274+ and NR1H2+ macrophage clusters were regarded as anti-tumor response [18]. In this study, we downloaded the scRNA-seq data of HGSOC tumor tissues from GEO database and marker genes from the CellMarker database, then performed a scRNA-seq analysis for the cell landscape of HGSOC. The FindAllMarkers function was used for the identification of significantly high expression genes, and the highly expressed genes were uploaded into the DAVID and KEGG database for enrichment analysis, and the AUCell package was used to calculate cancer-associated hallmark score. The SCENIC analysis was used for key regulons, the estrogen response enrichment scores in TCGA-OV RNA-seq dataset were calculated by using the GSVA package. Finally, we obtained a group of epithelial cells that promoting cancer progression.
2. Materials and methods
2.1. scRNA-seq sources and processing
In this study, two single-cell sequencing datasets (scRNA-seq) of primary HGSOC were downloaded from the Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/) database, including GSE192898 (HiSeq X Ten) and GSE211956 (Illumina NovaSeq 6000). GSE192898 contains 9 HGSOC tumor samples (ovCHA004, ovCHA017, ovCHA018, ovCHA034, ovCHA039, ovCHA066, ovCHA070, ovCHA107, ovCHA110). GSE211956 contains 5 HGSOC tumor samples (Y2, Y3, Y5, MJ10, MJ11). According to the processing program for scRNA-seq data, the Seurat package [19] was used to process the data in the dual dataset. The cell filtering followed by these criterions, expressing 200–9000 genes and the mitochondrial genes <25%. The SCTransform function was used to normalize the 14 samples. Principal component analysis (PCA) was then performed for the dimensionality reduction of the data [20]. Since the data were derived from different sequencing platforms, it was critical to remove batch effects from the samples. The Harmony package [21] was used for batch effect processing. The first 20 principal components in PCA were then selected for Uniform manifold approximation and projection (UMAP) dimensionality reduction analysis. Unsupervised cluster analysis of all cells was performed using the FindNeighbors and FindClusters functions built into the Seurat package to identify subpopulations of cells in HGSOC, with the parameter resolution = 0.12 set.
2.2. Cell annotation
To annotate identified cell subpopulations, we annotated cell subpopulations using the expression levels of marker genes. Downloading the list of marker genes for published cells in the CellMarker database (http://xteam.xbio.top/CellMarker/) [22], we performed cell annotation. For the identified epithelial cells, all epithelial cells were selected and further unsupervised cluster analysis was performed to identify molecular subgroups among them, with the parameter set to resolution = 0.2.
2.3. Differential expression analysis and Functional enrichment analysis between cell subpopulations
To explore the heterogeneity of gene expression patterns among cell subpopulations, the FindAllMarkers function built into the Seurat package performed differential expression analysis to identify highly expressed genes among them. Here we set the parameters: only.pos = T, min.pct = 0.25, logfc.threshold = 0.25. Functional enrichment analysis was performed in order to explore the biological pathways involved in different cell subpopulations. The highly expressed genes in each cell subpopulation were uploaded to the DAVID database (https://david.ncifcrf.gov/) [23], and we performed Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis. In addition, we downloaded the h.all.v2023.1.Hs.symbols.gmt file via the Molecular Signatures Database (MSigDB, https://www.gsea-msigdb.org/gsea/msigdb) to extract the cancer-related hallmark pathways. The AUCell package [24] was used to calculate AUCell enrichment scores for cancer-associated hallmark pathways in epithelial cell subpopulations.
2.4. Cell communication analysis
To probe the strength and number of ligand-receptor interaction pairs between epithelial cell subpopulations, we performed cell communication analysis. The CellChat package [25] was utilized to calculate the probability of ligand and receptor interactions between cell subpopulations, and bubble plots were used to demonstrate their interactions. The types of cell-cell interactions are categorized as cell-cell contact and secreted signaling.
2.5. Bulk RNA-seq data analysis
The RNA sequencing dataset of Ovarian serous cystadenocarcinoma (OV) was downloaded from the Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov/) database, and 378 tumor samples were included in TCGA-OV. Extracted HALLMARK_ESTROGEN_RESPONSE_EARLY gene set from h.all.v2023.1.Hs.symbols.gmt list. We calculated the estrogen response early enrichment scores for OC samples in TCGA-OV by the GSVA package [26] by means of single-sample gene set enrichment analysis (ssGSEA) [27]. The median values were used for grouping into higher ssGSEA score groups and lower ssGSEA score groups. Kaplan-Meier (K-M) survival analysis was performed in both groups to assess the prognostic status of patients in both groups. The estrogen response early enrichment score was then assessed separately in epithelial cell subpopulations. We also assessed the expression levels of marker genes in the estrogen pathway in subpopulations of epithelial cells. In addition, the pathway map of the estrogen pathway was downloaded from the KEGG official website (https://www.kegg.jp/), and we labeled the locations of the marker genes by entering their gene ids in the Mapper-Color module.
2.6. Single-cell regulatory network inference and clustering analysis
This study also mined transcription factors (TFs) in epithelial cell subpopulations in HGSOC with potential regulation of the estrogen pathway. We performed single-cell regulatory network inference and clustering (SCENIC) analysis [24,28,29]. According to the official description file of SCENIC (https://scenic.aertslab.org/), the GENIE3 package, the RcisTarget package, and the AUCell package were loaded in the R software and analyzed. The potential target gene of each TF were identified by using the GENIE3 method, the gene regulation networks (GRNs) were constructed by using the top10perTarget method. The cytoscape was used to visualize the GRNs. The TFs with the highest impact factor were determined based on the area under the curve (AUC) value of each regulon with the pearson correlation coefficient of the estrogen pathway. In addition, we explored the biological pathways involved in TFs-target genes.
2.7. Cell culture and transfection
Human ovarian epithelial cells IOSE-80 and epithelial ovarian cancer cell line Hey were purchased from Shanghai Cell Bank, Chinese Academy of Sciences. Hey and IOSE-80 cells were cultured in DMEM medium containing 10% fetal bovine serum at 37 °C with 5% CO2. Non-specific siRNA (si-NC) and si-STRA6 were purchased from Shanghai Jimma Genetics Co. Lipofectamine 2000 (Invitrogen, Thermo Fisher Scientific, Inc.) was used for transfection of si-STRA6 according to the instructions. The STRA6 siRNA sequence was 5′-GGCUCUGGAAGUGUGCUACAU-3′.
2.8. Quantitative reverse transcription (qRT)-PCR
The expression levels of IRF1 and STRA6 mRNA in cells were detected by qRT-PCR. Total RNA was extracted from IOSE-80, Hey cell lines using TRIzol reagent (Thermo Fisher, USA). Quantitative reverse transcription-polymerase chain reaction (qRT-PCR) was performed on RNA from per sample (2 μg) on a LightCycler 480 PCR system (Roche, USA) using FastStart Universal SYBR ®Green Master (Roche, USA). The cDNA was used as template and the reaction volume was 20 μl (2 μl cDNA template, 10 μl PCR mix, 0.5 μl forward and reverse primers and appropriate amount of water). The relative expression levels of IRF1 and STRA6 were calculated by the 2-ΔΔCt method using GAPDH as the internal reference gene. The primer sequences were shown in Table 1.
Table 1.
Primer sequences for IRF1 and STRA6.
| Gene | Forward primer sequence (5′-3′) | Reverse primer sequence (5′-3′) |
|---|---|---|
| IRF1 | AAGGGGTGTGGCCTTTTTAGA | TGTCCCTGTTCACCCCAAAG |
| STRA6 | CTATGGCAGCTGGTACATCG | TACAG GCCGGGTGGTATG |
| GAPDH | AATGGGCAGCCGTTAGGAAA | GCCCAATACGACCAAATCAGAG |
2.9. Transwell assay
Cell invasion and migration were detected by Transwell assay. Migration assay: 5 × 104 cells were added to the upper chamber of the Transwell migration plate, and 600 μL of complete medium was added to the lower chamber, and the cells were incubated at 5% CO2 and 37 °C for 24 h. The cells were fixed and stained with 0.1% crystal violet for 20 min, and quantified under a light microscope. Invasion assay: Using the pre-filled Matrigel gel chamber, 5 × 104 cells were added to the upper chamber and cultured at 5% CO2 and 37 °C for 48 h. The cells were fixed at the bottom of the membrane and stained with 0.1% crystal violet staining for 20 min, and the cells were quantified under a light microscope.
2.10. Statistical analysis
We used the wilcoxon rank sum test to calculate the difference between the two sets of continuous variables. For survival analysis, we divided the sample into two groups, high and low, based on the median of the continuous variables, and then used the log-rank test to calculate the difference in survival time between the two groups of patients. All statistical analyses were done by R language (version 3.6.0) and *p < 0.05 was considered statistically significant.
3. Results
3.1. Cell mapping in HGSOC
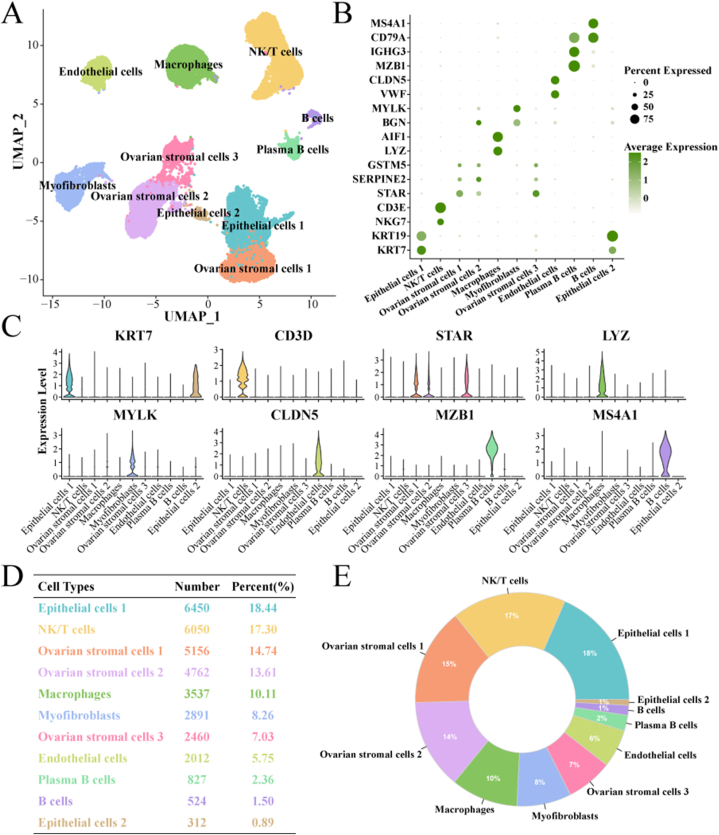
Two HGSOC single-cell datasets, GSE192898 (ovCHA004, ovCHA017, ovCHA018, ovCHA034, ovCHA039, ovCHA066, ovCHA070, ovCHA107, ovCHA110), GSE211956 (Y2, Y3, Y5, MJ10, MJ11). We first removed batch effects and performed data quality control in the double dataset, and data from 34,981 cells were retained (Supplementary figs. 1A–D). After single-cell data analysis with the Seurat package, 11 major cell subpopulations were identified, Endothelial cells, Macrophages, NK/T cells, B cells, Ovarian stromal cells 1, Ovarian stromal cells 2, Ovarian stromal cells 3, Plasma B cells, Myofibroblasts, Epithelial cells 1, Epithelial cells 2 (Fig. 1A). The marker genes specifically expressed in 11 cell subpopulations are shown in Fig. 1B–C. Ovarian stromal cells 1–3 all specifically express GSTM5, SERPINE2, STAR. GSTM5 is a prognostic biomarker for angiogenic features in Ovarian cancer [30]. SERPINE2 is likewise an angiogenic lymphangiogenic pro-oncogenic factor [31]. We counted the number of cells in 11 cell subpopulations, and three Ovarian stromal cells subpopulations including Ovarian stromal cells (1,2,3), had the highest number of cells (Fig. 1D–E).
Fig. 1.
Cell mapping in HGSOC.
A: Cell mapping of 11 cell subpopulations in HGSOC.
B–C: Expression levels of marker genes specifically expressed in 11 cell subpopulations.
D–E: cell numbers and proportions of 11 cell subpopulations.
3.2. Heterogeneity of epithelial cells in HGSOC
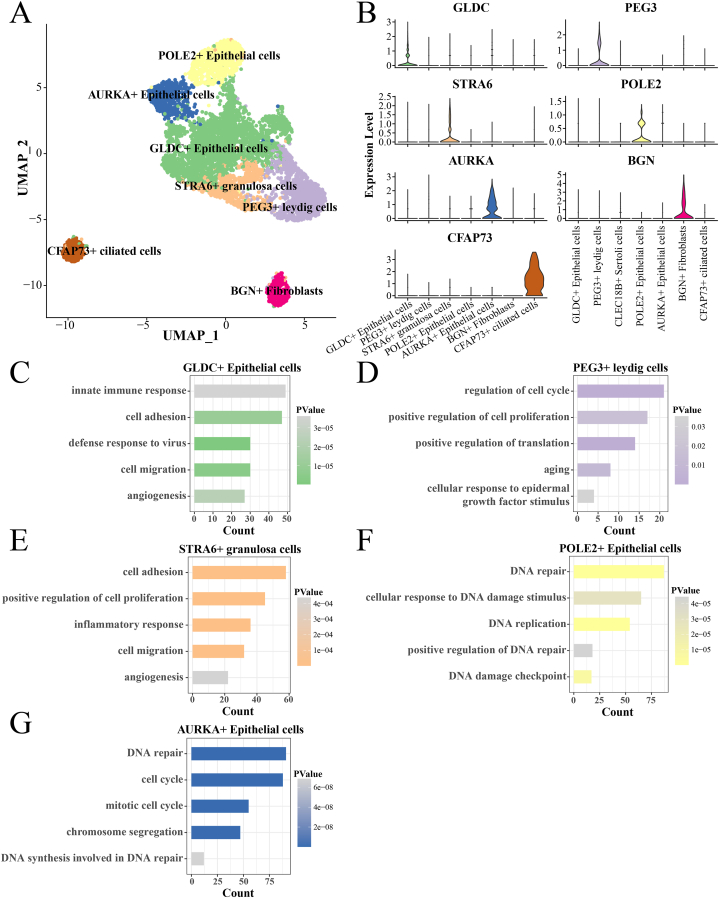
Since HGSOC is a malignant tumor that occurs in the ovarian epithelium, it is particularly important to portray the heterogeneity of HGSOC epithelial cells. We extracted cell data of Epithelial cells 1–2 for further cell typing. Seven cell subpopulations were present in Epithelial cells, POLE2+ Epithelial cells, AURKA + Epithelial cells, GLDC + Epithelial cells, STRA6+ granulosa cells, PEG3+ leydig cells, CFAP73+ ciliated cells, BGN + Fibroblasts (Fig. 2A–B). We found a small number of ciliated cells and fibroblasts with gene expression patterns similar to those of epithelial cells, which were mixed into the epithelial cell subpopulation, and therefore excluded BGN + Fibroblasts. The biological pathways involved in the remaining five cell subpopulations were analyzed. GLDC + Epithelial cells are involved in innate immune response, cell adhesion, defense response to virus, cell migration, and angiogenesis (Fig. 2C). PEG3+ leydig cells are involved in regulation of cell cycle, positive regulation of cell proliferation, positive regulation of translation, aging, and cell response to epidermal growth factor stimulus (Fig. 2D). STRA6+ granulosa cells are involved in cell adhesion, positive regulation of cell proliferation, inflammatory response, cell migration, angiogenesis (Fig. 2E)). POLE2+ Epithelial cells are involved in DNA repair, cell response to DNA damage stimulus, DNA replication, positive regulation of DNA repair, and DNA damage checkpoint (Fig. 2F). AURKA + Epithelial cells are involved in DNA repair, cell cycle, mitotic cell cycle, chromosome segregation, and DNA synthesis involved in DNA repair (Fig. 2G).
Fig. 2.
Five heterogeneous subpopulations of Epithelial cells ya in HGSOC and the biological pathways they are involved in.
A: Six heterogeneous subpopulations of Epithelial cells in HGSOC.
B: Marker genes specifically expressed in five heterogeneous subpopulations of Epithelial cells in HGSOC.
C: Biological pathways involved in GLDC + Epithelial cells.
D: Biological pathways involved in PEG3+ leydig cells.
E: Biological pathways involved in STRA6+ granulosa cells.
F: Biological pathways involved in POLE2+ Epithelial cells.
G: Biological pathways involved in AURKA + Epithelial cells.
3.3. STRA6+ granulosa cells promote HGSOC progression
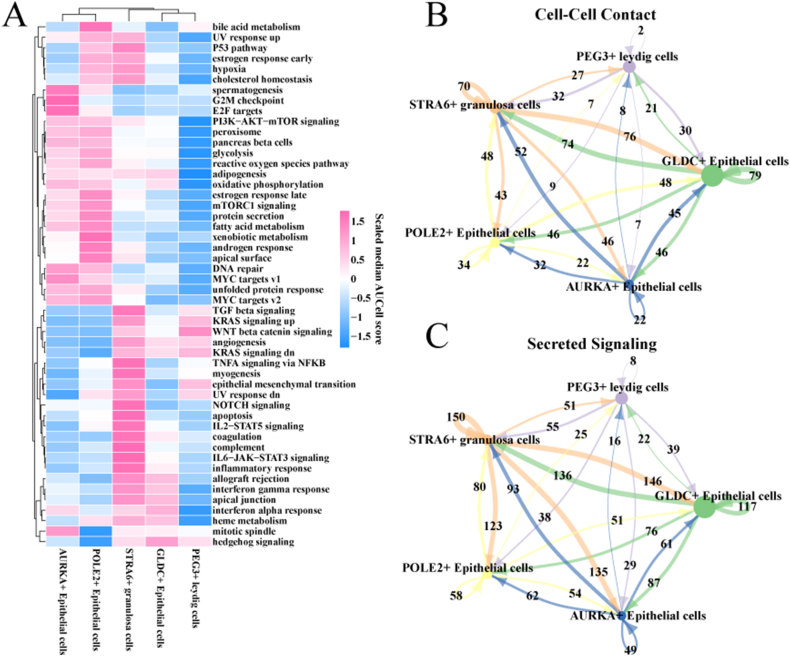
To characterize the degree of activation or inhibition of cancer-related pathways in different epithelial cells, we calculated enrichment scores for the hallmark gene set using the AUCell algorithm. We observed that PI3K-AKT-mTOR signaling, mTORC1 signaling, and DNA repair activities were activated in AURKA + Epithelial cells and POLE2+ Epithelial cells, which indicating that these two cell subpopulations had a greater proliferative capacity. We also observed that glycolysis, fatty acid metabolism, reactive oxygen pathway, oxidative phosphorylation in AURKA + Epithelial cells and POLE2+ Epithelial cells, adipogenesis all scored somewhat higher in activity. It suggested that they are more capable of generating energy, providing ample power for unlimited cell proliferation. Interestingly, we found TGF beta signaling, KRAS signaling, WNT beta catenin signaling, angiogenesis, epithelial mesenchymal transition in STRA6+ granulosa cells, inflammatory response had higher enrichment scores, and these pathways are closely related to tumor development, which suggested STRA6+ granulosa cells have the potential to promote the development of HGSOC. In contrast, most of the cancer-related pathways in GLDC + Epithelial cells and PEG3+ leydig cells were in an inhibited state (Fig. 3A). The results of the cell communication analysis showed a higher number and density of ligand-receptor interaction pairs between STRA6+ granulosa cells and epithelial cell subpopulations, both at the cell-cell contact level and between secreted signaling layers. In contrast, the intensity of communication exchanges between PEG3+ leydig cells and epithelial cell subpopulations was lower (Fig. 3B–C). These results suggested that STRA6+ granulosa cells have the potential to promote tumor growth.
Fig. 3.
Enrichment analysis of the hallmark pathway and cell communication analysis of epithelial cell subpopulations.
A: Heatmap of hallmark pathway enrichment scores for epithelial cell subpopulations.
B–C: Network diagram of the number of ligand-receptor pairs interacting between epithelial cell subpopulations. (B) Cell-cell contact type. (C) Secreted signaling type.
3.4. Exchange density between STRA6+ granulosa cells and epithelial cell subpopulations
To probe the binding status of ligand-receptor interaction pairs between STRA6+ granulosa cells and epithelial cell subpopulations, we performed further cell communication analyses. For cell-cell contact, there was a close cell interaction between STRA6+ granulosa cells and GLDC + Epithelial cells. Specifically, the proliferation of GLDC + Epithelial cells was promoted by JAG1-NOTCHs, HLAs- CD4, EFNBs-EPHBs, and EFNAs-EPHAs (Fig. 4A). For secreted signaling, HBEGF secreted in STRA6+ granulosa cells strongly interacted with ERBB2+ERBB4 and EGFR + ERBB2 in four epithelial cells. In addition, STRA6+ granulosa cells secreted WNT, VEGF, TGF-β, EGFR, FGFR-related ligand-receptors to promote epithelial cell proliferation (Fig. 4B).
Fig. 4.
Communication density between STRA6+ granulosa cells and epithelial cell subpopulations.
A: Cell-cell contact type.
B: Secreted signaling type.
3.5. STRA6+ granulosa cells activate estrogen response early activity
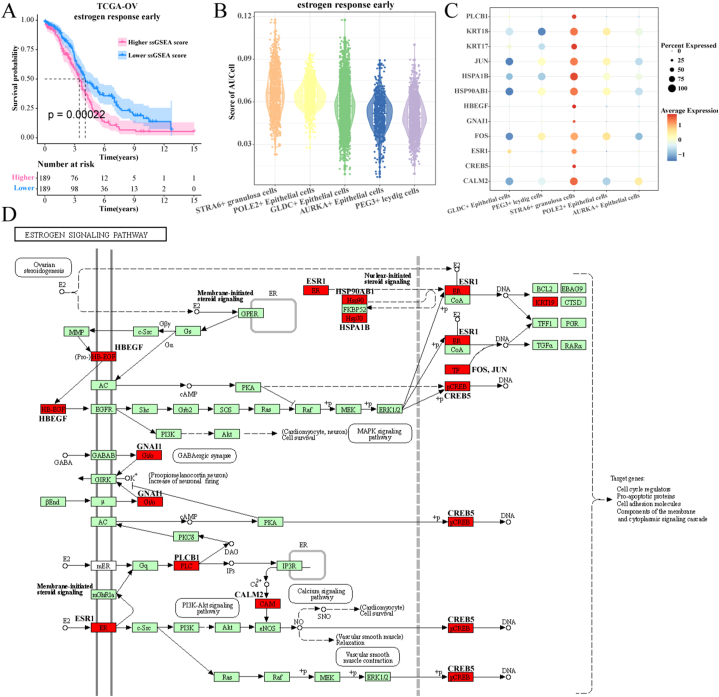
The ovaries compose of the central reproductive organ of female and produce multiple hormones including the estrogen and progesterone [32], thus estrogen levels are strongly correlated with OC [33]. Estrogens may stimulate the OC cell growth in vitro was observed [34]. Therefore, we evaluated the relationship between early estrogen response and TCGA-OV prognosis. First, the estrogen response score were calculated in the TCGA-OV cohort, the patients were divided by the median into high- and low-score groups with significantly different prognosis (p < 0.001), in which the patients with high estrogen score exhibited poor outcome (Fig. 5A). Subsequently, the high estrogen response score as the cancer progression indicators was calculated in five cell clusters, the results found that the STRA6+ granulosa cells exhibited the highest score in these cell clusters(Fig. 5B), supporting the STRA6+ granulosa cells as a crucial factor in HGSOC progression. Correspondingly, the highest expression levels of marker genes of the estrogen response early pathway were found in STRA6+ granulosa cells (Fig. 5C). Finally, the estrogen pathway map was downloaded from the KEGG official website to inscribe the location of marker genes of the estrogen pathway in STRA6+ granulosa cells (Fig. 5D).
Fig. 5.
STRA6+ granulosa cells activate estrogen response early activity.
A: K-M curves of patients in the estrogen response early subgroup.
B: estrogen response early enrichment score in a subpopulation of epithelial cells.
C: Expression levels of estrogen pathway-related genes in a subpopulation of epithelial cells. , and
D: Location of estrogen pathway-related genes highly expressed in STRA6+ granulosa cells within the KEGG pathway map.
3.6. STRA6+ granulosa cells regulate estrogen response early via IRF1 dominant GRN
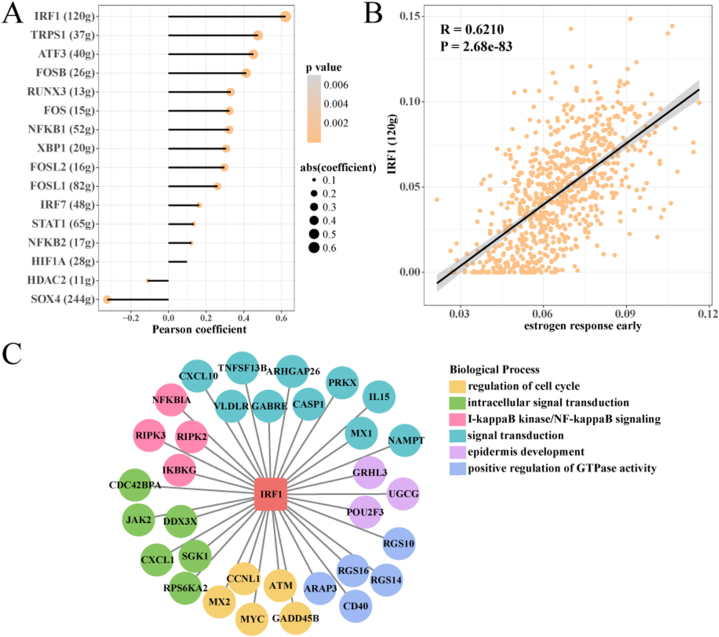
To further identify transcription factors that are closely associated with estrogen response early activity within STRA6+ granulosa cells, we performed SCENIC analysis. Sixteen significantly correlated TFs were identified by pearson correlation of the AUCell score of TFs with the estrogen response early score of STRA6+ granulosa cells analysis (Fig. 6A). IRF1 showed a significant positive correlation (R = 0.6210, P = 2.68e-83) with estrogen response early score in STRA6+ granulosa cells (Fig. 6B), implying that IRF1 is closely associated with estrogen response early activity. IRF1-dominant GRN is involved in regulation of cell cycle, intracell signal transduction, I-kappaB kinase/NF-kappaB signaling, signal transduction, epidermis development, and positive regulation of GTPase activity (Fig. 6C).
Fig. 6.
STRA6+ granulosa cells regulate estrogen response early via IRF1-dominant GRN.
A: pearson correlation analysis of AUCell score of TFs with estrogen response early score of STRA6+ granulosa cells.
B: Pearson correlation analysis of IRF1 with estrogen response early score in STRA6+ granulosa cells.
C: IRF1-dominated biological pathways involved in GRN. Boxes represent TFs, circles represent target genes, and different colors represent different biological processes.
3.7. Effects of IRF1 and STRA6 on OC invasion and migration
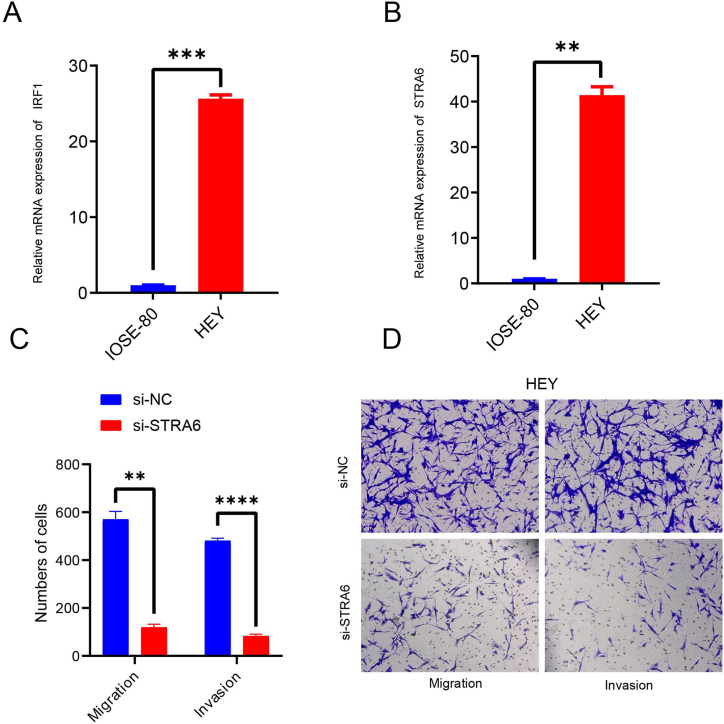
To verify the relationship between the expression of STRA6 and its transcription factors, we detected the expression level of IRF1 and STRA6 in normal (IOSE-80) and tumor (HEY) derived epithelial cell by using qRT-PCR. The results showed that IRF1 and STRA6 exhibited the consistent expression patterns, their expression levels were significantly upregulated in the OC cancer cell line HEY (p < 0.01, Fig. 7A–B). After silencing STRA6, the invasion and migration ability of OC cancer cell line HEY was markedly decreased (Fig. 7C–D), implying the high expression level of STRA6 is a crucial supporting factors of OC progression.
Fig. 7.
Effects of IRF1 and STRA6 on OC invasion and migration.
A: Expression levels of IRF1 in IOSE-80 and HEY.
B: Expression levels of STRA6 in IOSE-80 and HEY.
C–D: Effects of silencing STRA6 on OC cell invasion and migration.
(*p < 0.05, **p < 0.01, ***p < 0.001).
4. Discussion
HGSOC is a highly heterogeneous and complex cancer that lacks definitive and actionable targets and has a consistently poor prognosis [35]. The phenomenon of accumulation of epithelial cells that transform from benign to malignant is an important cause of the development of HGSOC [36]. In this study, we portrayed the cell landscape in HGSOC tumor tissues, in which the epithelial cells had the most numerous and proportionate. Further, we identified three subpopulations of epithelial cells, all of which specifically express GSTM5, SERPINE2, and STAR, but with differential expression levels. GSTM5, SERPINE2 were all shown to be pro-carcinogenic factors [30,31]. Thus, the three epithelial cell subpopulations may be malignant epithelial cells with pro-carcinogenic effects.
Epithelial cells are the largest cell clusters and composed of ∼31% in cells analysis of ovarian cancer (OC) [37]. However, the current single-cell studies of OC cannot precisely distinguish the cell cluster with specific clinical phenotypes [38]. Sun el. identified a groups Scissor + cells that associated with the poor prognosis through the Scissor algorithm [38], these genes including the JUN, GAS1 and MGP were significantly high expression in Scissor + cells as the signature of tumor progression [39]. Our scRNA analysis demonstrated the heterogeneity of epithelial cells in HGSOC tumors. Five cell subpopulations were present in epithelial cells, POLE2+ Epithelial cells, AURKA + Epithelial cells, GLDC + Epithelial cells, STRA6+ granulosa cells, PEG3+ leydig cells. They specifically express POLE2, AURKA, GLDC, STRA6, PEG3, respectively. Inhibition of POLE2 expression was found to inhibit epithelial-mesenchymal transition and migration capacity in tumor cells [40]. Reduced POLE2 promoter activity is associated with DNA damage and cell cycle mechanisms in tumor cells [41]. It has also been found that POLE2 regulates apoptotic and proliferative responses in tumor cells through the PI3K/AKT signaling pathway [42,43]. This is consistent with the trend of our findings that PI3K-AKT-mTOR signaling, DNA repair mutations were significantly activated in POLE2+ Epithelial cells, suggesting a strong cell proliferation capacity. AURKA can mediate the migration and adhesion capacity of epithelial ovarian cancer cells [44].
STRA6 has no relevant studies in OC for the time being. But in other cancer studies, the phenomenon we found was confirmed. In gastric cancer, inhibition of STRA6 expression suppresses the Wnt/β-linker protein signaling pathway, and this results in the ability of gastric cancer cell lines to migrate and invade [45]. It can be seen that STRA6 can be remembered to activate the Wnt/β-linker protein signaling pathway. This is consistent with our finding that Wnt/β-linker protein signaling pathway activity is significantly activated in STRA6+ granulosa cells. The dysregulated Wnt/β-collagen signaling pathway is also an important cause of cancer development [46]. The activities of TGF beta signaling [47], angiogenesis [48], and epithelial mesenchymal transition [49] were also significantly activated in STRA6+ granulosa cells. Dysregulation of these pathways is a major cause of cancer development, and clinical drugs against these targets have been shown to be promising potential agents for cancer. These identified unique subpopulations compose of an important heterogeneous characteristic of HGSOC and provide a new perspective on the TME of ovarian cancer. The STRA6+ granulosa cells may have the potential to promote the development of HGSOC, these specific genes could be the disease treatment targets and develop the novel treatment strategy for HGSOC intervention. During the HGSOCs progression, several key signaling pathways, such as oxidative phosphorylation, tumor cell proliferation and cell cycle, were markedly enhanced for energy supply, so the poly (ADP-ribose) polymerase (PARP) inhibition is a useful targeted strategy for HGSOC treatment [[50], [51], [52]]. Beyond that, the JAK/STAT inhibitor JSI-124 exhibited anti-tumor property in HGSOC cell lines and could be an effective anti-cancer in patients with HGSOC [53]. In our study, these cell cluster are associated with different cancer pathway activation for HGSOC progression, such as the activated PI3K/AKT and glycolysis, reactive oxygen pathway and fatty acid metabolism in AURKA+ and POLE2+ Epithelial cells, the activated TGF-β, KRAS and Wnt pathways in STRA6+ granulosa cells, thus the combined targeted therapy of drugs may benefit for the HGSOC patients. Meanwhile, these cell cluster also provide potential targeted genes for the development of HGSOC intervention strategy, but their actual function in these cancer pathways need further verification by using the gene knockout or silencing.
There is also a gene regulatory network of pro-carcinogenic features in STRA6+ granulosa cells that is dominated by IRF1. PD-L1 expressed on the surface of OC cells is the main reason for the occurrence of immune escape, in which IRF1 signaling plays an important role. Induction of IFNγ inhibits IRF1 expression, and inhibition of IRF1 affects PD-L1 expression in OC cells [54]. In a previous study of immunologic parameters, high expression of IRF1 in OC cells was indeed an important immunologic parameter predicting OC prognosis [55]. However, previous studies have not been able to identify the source of IRF1, and our study utilized single-cell sequencing data to identify STRA6+ granulosa cells-derived IRF1 as an important TF that promotes the development of HGSOC. Indeed, IRF1-dominated GRNs are mainly associated with the cell cycle and cell epidermal development. Alterations in these signals may provide a pro-carcinogenic environment. In addition, the strongest estrogen response early activity was found in STRA6+ granulosa cells. Inhibition of estrogen response is also one of the therapeutic options for OC [56]. Furthermore, cell assays confirmed that silencing STRA6 decreased the invasion and migration ability of OC cells.
Overall, our findings revealed that STRA6+ granulosa cells are an important cell cluster that supported OC progression, in which the STRA6 is a crucial TF for the activity of STRA6+ granulosa cells. However, there are also many limitations in this study, such as the conclusions of our study are mainly depended on the bioinformatics analyses of two retrospective cohorts, the sample sizes and retrospective nature may limit the conclusion interpretation. Besides, the function of STRA6 need to the further verification in vitro or in vivo.
5. Conclusion
Overall, we revealed the heterogeneity of epithelial cells in HGSOC by single-cell techniques, and STRA6+ granulosa cells have a potential pro-carcinogenic role. The STRA6+ granulosa cells may represent a specific subtype to explain the heterogeneity of HGSOC, meanwhile, our findings may provide new insight to understand the underly mechanisms of progress in HGSOC.
Funding
This research was supported by Hangzhou Biomedical and Health Industry Development Support Science and Technology Special Project Fund (No. 2022WJC254), Zhejiang Medical and Health Science and Technology Program Fund (No. 2023KY957) and Zhejiang Medical and Health Science and Technology Program Fund (No. 2023KY183).
Data availability statement
The datasets generated during and/or analyzed during the current study are available in the GSE repository [GSE192898] (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc= GSE192898) and [GSE211956] (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc= GSE211956).
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
CRediT authorship contribution statement
Xiaoting Liu: Writing – review & editing, Writing – original draft, Visualization, Software, Resources, Project administration, Investigation, Funding acquisition, Data curation. Zhaojun Chen: Writing – review & editing, Visualization, Supervision, Software, Methodology, Data curation, Conceptualization. Lahong Zhang: Writing – review & editing, Visualization, Supervision, Resources, Funding acquisition, Formal analysis.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgement
None.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.heliyon.2024.e27790.
Contributor Information
Zhaojun Chen, Email: hzczj2006@163.com.
Lahong Zhang, Email: zjhzzlh2007@163.com.
Abbreviations
- HGSOC
High-grade serous ovarian carcinoma
- OC
ovarian cancer
- SCENIC
single-cell regulatory network inference and clustering
- TFs
transcription factors
- GRNs
gene regulatory networks
- scRNA-seq
single-cell sequencing
- GEO
Gene Expression Omnibus
- PCA
Principal component analysis
- UMAP
Uniform manifold approximation and projection
- KEGG
Kyoto Encyclopedia of Genes and Genomes
- MSigDB
the Molecular Signatures Database
- OV
Ovarian serous cystadenocarcinoma
- TCGA
the Cancer Genome Atlas
- ssGSEA
single-sample gene set enrichment analysis
- K-M,
Kaplan-Meier
- AUC
area under the curve
Appendix A. Supplementary data
The following are the Supplementary data to this article.
References
- 1.Huang X., et al. Carrier-free multifunctional nanomedicine for intraperitoneal disseminated ovarian cancer therapy. J Nanobiotechnology. 2022;20(1):93. doi: 10.1186/s12951-022-01300-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Li S., et al. Long non-coding RNA SLC25A21-AS1 inhibits the development of epithelial ovarian cancer by specifically inducing PTBP3 degradation. Biomark. Res. 2023;11(1):12. doi: 10.1186/s40364-022-00432-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sung H., et al. Global cancer statistics 2020: GLOBOCAN Estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–249. doi: 10.3322/caac.21660. [DOI] [PubMed] [Google Scholar]
- 4.Siegel R.L., et al. Cancer statistics, 2023. CA Cancer J Clin. 2023;73(1):17–48. doi: 10.3322/caac.21763. [DOI] [PubMed] [Google Scholar]
- 5.Armstrong D.K., et al. NCCN guidelines(R) insights: ovarian cancer, version 3.2022. J Natl Compr Canc Netw. 2022;20(9):972–980. doi: 10.6004/jnccn.2022.0047. [DOI] [PubMed] [Google Scholar]
- 6.Kurnit K.C., Frumovitz M. Primary mucinous ovarian cancer: options for surgery and chemotherapy. Int J Gynecol Cancer. 2022 doi: 10.1136/ijgc-2022-003806. Oct 13:ijgc-2022-003806. Epub ahead of print. PMID: 36229081. [DOI] [PubMed] [Google Scholar]
- 7.Li G., et al. Synergetic delivery of artesunate and isosorbide 5-mononitrate with reduction-sensitive polymer nanoparticles for ovarian cancer chemotherapy. J Nanobiotechnology. 2022;20(1):471. doi: 10.1186/s12951-022-01676-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bartoletti M., et al. Emerging molecular alterations leading to histology-specific targeted therapies in ovarian cancer beyond PARP inhibitors. Cancer Treat Rev. 2021;101 doi: 10.1016/j.ctrv.2021.102298. [DOI] [PubMed] [Google Scholar]
- 9.Qiao L., et al. Correlation analysis and clinical significance of CA125, HE4, DDI, and FDP in type II epithelial ovarian cancer. Medicine (Baltim.) 2020;99(49) doi: 10.1097/MD.0000000000023329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Punzon-Jimenez P., et al. Molecular management of high-grade serous ovarian carcinoma. Int. J. Mol. Sci. 2022;23(22) doi: 10.3390/ijms232213777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Lisio M.A., et al. High-grade serous ovarian cancer: basic sciences, clinical and therapeutic standpoints. Int. J. Mol. Sci. 2019;20(4) doi: 10.3390/ijms20040952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Olbrecht S., et al. High-grade serous tubo-ovarian cancer refined with single-cell RNA sequencing: specific cell subtypes influence survival and determine molecular subtype classification. Genome Med. 2021;13(1):111. doi: 10.1186/s13073-021-00922-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Xu H., et al. CCNE1 copy number is a biomarker for response to combination WEE1-ATR inhibition in ovarian and endometrial cancer models. Cell Rep Med. 2021;2(9) doi: 10.1016/j.xcrm.2021.100394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Yang B., et al. Spatial heterogeneity of infiltrating T cells in high-grade serous ovarian cancer revealed by multi-omics analysis. Cell Rep Med. 2022;3(12) doi: 10.1016/j.xcrm.2022.100856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474(7353):609–615. doi: 10.1038/nature10166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chen G.M., et al. Consensus on molecular subtypes of high-grade serous ovarian carcinoma. Clin. Cancer Res. 2018;24(20):5037–5047. doi: 10.1158/1078-0432.CCR-18-0784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Jayson G.C., et al. Ovarian cancer. Lancet. 2014;384(9951):1376–1388. doi: 10.1016/S0140-6736(13)62146-7. [DOI] [PubMed] [Google Scholar]
- 18.Olalekan S., et al. Characterizing the tumor microenvironment of metastatic ovarian cancer by single-cell transcriptomics. Cell Rep. 2021;35(8) doi: 10.1016/j.celrep.2021.109165. [DOI] [PubMed] [Google Scholar]
- 19.Stuart T., et al. Comprehensive integration of single-cell data. Cell. 2019;177(7):1888–1902 e21. doi: 10.1016/j.cell.2019.05.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Zulibiya A., et al. Single-cell RNA sequencing reveals potential for endothelial-to-mesenchymal transition in tetralogy of fallot. Congenit. Heart Dis. 2023;18(6):611–625. [Google Scholar]
- 21.Korsunsky I., et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods. 2019;16(12):1289–1296. doi: 10.1038/s41592-019-0619-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Zhang X., et al. CellMarker: a manually curated resource of cell markers in human and mouse. Nucleic Acids Res. 2019;47(D1):D721–D728. doi: 10.1093/nar/gky900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Dennis G., Jr., et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 2003;4(5):P3. [PubMed] [Google Scholar]
- 24.Aibar S., et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods. 2017;14(11):1083–1086. doi: 10.1038/nmeth.4463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jin S., et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 2021;12(1):1088. doi: 10.1038/s41467-021-21246-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hanzelmann S., Castelo R., Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinf. 2013;14:7. doi: 10.1186/1471-2105-14-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Barbie D.A., et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature. 2009;462(7269):108–112. doi: 10.1038/nature08460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Van de Sande B., et al. A scalable SCENIC workflow for single-cell gene regulatory network analysis. Nat. Protoc. 2020;15(7):2247–2276. doi: 10.1038/s41596-020-0336-2. [DOI] [PubMed] [Google Scholar]
- 29.Bravo Gonzalez-Blas C., et al. SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks. Nat. Methods. 2023;20(9):1355–1367. doi: 10.1038/s41592-023-01938-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wang Y., et al. A comprehensive understanding of ovarian carcinoma survival prognosis by novel biomarkers. Eur. Rev. Med. Pharmacol. Sci. 2019;23(19):8257–8264. doi: 10.26355/eurrev_201910_19136. [DOI] [PubMed] [Google Scholar]
- 31.Sasahira T., et al. SERPINE2 is an oral cancer-promoting factor that induces angiogenesis and lymphangiogenesis. Int. J. Clin. Oncol. 2021;26(10):1831–1839. doi: 10.1007/s10147-021-01970-4. [DOI] [PubMed] [Google Scholar]
- 32.Richards J.S., Pangas S.A. The ovary: basic biology and clinical implications. J. Clin. Invest. 2010;120(4):963–972. doi: 10.1172/JCI41350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wang Z., et al. EVI1 overexpression promotes ovarian cancer progression by regulating estrogen signaling. Mol. Cell. Endocrinol. 2021;534 doi: 10.1016/j.mce.2021.111367. [DOI] [PubMed] [Google Scholar]
- 34.Cunat S., Hoffmann P., Pujol P. Estrogens and epithelial ovarian cancer. Gynecol. Oncol. 2004;94(1):25–32. doi: 10.1016/j.ygyno.2004.03.026. [DOI] [PubMed] [Google Scholar]
- 35.Shih A.J., et al. Identification of grade and origin specific cell populations in serous epithelial ovarian cancer by single cell RNA-seq. PLoS One. 2018;13(11) doi: 10.1371/journal.pone.0206785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Bulun S.E., Wan Y., Matei D. Epithelial mutations in endometriosis: link to ovarian cancer. Endocrinology. 2019;160(3):626–638. doi: 10.1210/en.2018-00794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kurman R.J., Shih Ie M. The origin and pathogenesis of epithelial ovarian cancer: a proposed unifying theory. Am. J. Surg. Pathol. 2010;34(3):433–443. doi: 10.1097/PAS.0b013e3181cf3d79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Sun D., et al. Identifying phenotype-associated subpopulations by integrating bulk and single-cell sequencing data. Nat. Biotechnol. 2022;40(4):527–538. doi: 10.1038/s41587-021-01091-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Bakir B., et al. EMT, MET, plasticity, and tumor metastasis. Trends Cell Biol. 2020;30(10):764–776. doi: 10.1016/j.tcb.2020.07.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zhang P., et al. POLE2 facilitates the malignant phenotypes of glioblastoma through promoting AURKA-mediated stabilization of FOXM1. Cell Death Dis. 2022;13(1):61. doi: 10.1038/s41419-021-04498-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Guo H., et al. Knockdown of HDAC10 inhibits POLE2-mediated DNA damage repair in NSCLC cells by increasing SP1 acetylation levels. Pulm. Pharmacol. Ther. 2023;83 doi: 10.1016/j.pupt.2023.102250. [DOI] [PubMed] [Google Scholar]
- 42.Ge S., et al. Silencing POLE2 promotes apoptosis and inhibits proliferation of oral squamous cell carcinomas by inhibiting PI3K/AKT signaling pathway. Med. Oncol. 2023;40(10):304. doi: 10.1007/s12032-023-02158-8. [DOI] [PubMed] [Google Scholar]
- 43.Zhu Y., et al. POLE2 knockdown reduce tumorigenesis in esophageal squamous cells. Cancer Cell Int. 2020;20:388. doi: 10.1186/s12935-020-01477-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Do T.V., et al. Aurora kinase A mediates epithelial ovarian cancer cell migration and adhesion. Oncogene. 2014;33(5):539–549. doi: 10.1038/onc.2012.632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Lin L., et al. STRA6 exerts oncogenic role in gastric tumorigenesis by acting as a crucial target of miR-873. J. Exp. Clin. Cancer Res. 2019;38(1):452. doi: 10.1186/s13046-019-1450-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Liu J., et al. Wnt/beta-catenin signalling: function, biological mechanisms, and therapeutic opportunities. Signal Transduct Target Ther. 2022;7(1):3. doi: 10.1038/s41392-021-00762-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Peng D., et al. Targeting TGF-beta signal transduction for fibrosis and cancer therapy. Mol. Cancer. 2022;21(1):104. doi: 10.1186/s12943-022-01569-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Ansari M.J., et al. Cancer combination therapies by angiogenesis inhibitors; a comprehensive review. Cell Commun. Signal. 2022;20(1):49. doi: 10.1186/s12964-022-00838-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Mortezaee K., Majidpoor J., Kharazinejad E. Epithelial-mesenchymal transition in cancer stemness and heterogeneity: updated. Med. Oncol. 2022;39(12):193. doi: 10.1007/s12032-022-01801-0. [DOI] [PubMed] [Google Scholar]
- 50.Pujade-Lauraine E., et al. Olaparib tablets as maintenance therapy in patients with platinum-sensitive, relapsed ovarian cancer and a BRCA1/2 mutation (SOLO2/ENGOT-Ov21): a double-blind, randomised, placebo-controlled, phase 3 trial. Lancet Oncol. 2017;18(9):1274–1284. doi: 10.1016/S1470-2045(17)30469-2. [DOI] [PubMed] [Google Scholar]
- 51.DiSilvestro P., et al. Overall survival with maintenance olaparib at a 7-year follow-up in patients with newly diagnosed advanced ovarian cancer and a BRCA mutation: the SOLO1/GOG 3004 trial. J. Clin. Oncol. 2023;41(3):609–617. doi: 10.1200/JCO.22.01549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kristeleit R., et al. A phase I-ii study of the oral PARP inhibitor rucaparib in patients with germline BRCA1/2-mutated ovarian carcinoma or other solid tumors. Clin. Cancer Res. 2017;23(15):4095–4106. doi: 10.1158/1078-0432.CCR-16-2796. [DOI] [PubMed] [Google Scholar]
- 53.Izar B., et al. A single-cell landscape of high-grade serous ovarian cancer. Nat Med. 2020;26(8):1271–1279. doi: 10.1038/s41591-020-0926-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Padmanabhan S., et al. IFNgamma-induced PD-L1 expression in ovarian cancer cells is regulated by JAK1, STAT1 and IRF1 signaling. Cell. Signal. 2022;97 doi: 10.1016/j.cellsig.2022.110400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Zeimet A.G., et al. Intratumoral interferon regulatory factor (IRF)-1 but not IRF-2 is of relevance in predicting patient outcome in ovarian cancer. Int. J. Cancer. 2009;124(10):2353–2360. doi: 10.1002/ijc.24214. [DOI] [PubMed] [Google Scholar]
- 56.Sarwar S., et al. Insights into the role of epigenetic factors determining the estrogen response in estrogen-positive ovarian cancer and prospects of combining epi-drugs with endocrine therapy. Front. Genet. 2022;13 doi: 10.3389/fgene.2022.812077. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets generated during and/or analyzed during the current study are available in the GSE repository [GSE192898] (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc= GSE192898) and [GSE211956] (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc= GSE211956).