Abstract
Background
Cardiorenal syndrome (CRS) type 4 is prevalent among the chronic kidney disease (CKD) population, with many patients dying from cardiovascular complications. However, limited data regarding cardiac transcriptional changes induced early by CKD is available.
Methods
We used a murine unilateral ureteral obstruction (UUO) model to evaluate renal damage, cardiac remodeling, and transcriptional regulation at 21 days post-surgery through histological analysis, RT-qPCR, RNA-seq, and bioinformatics.
Results
UUO leads to significant kidney injury, low uremia, and pathological cardiac remodeling, evidenced by increased collagen deposition and smooth muscle alpha-actin 2 expression. RNA-seq analysis identified 76 differentially expressed genes (DEGs) in UUO hearts. Upregulated DEGs were significantly enriched in cell cycle and cell division pathways, immune responses, cardiac repair, inflammation, proliferation, oxidative stress, and apoptosis. Gene Set Enrichment Analysis further revealed mitochondrial oxidative bioenergetic pathways, autophagy, and peroxisomal pathways are downregulated in UUO hearts. Vimentin was also identified as an UUO-upregulated transcript.
Conclusions
Our results emphasize the relevance of extensive transcriptional changes, mitochondrial dysfunction, homeostasis deregulation, fatty-acid metabolism alterations, and vimentin upregulation in CRS type 4 development.
Keywords: Cardiorenal, Transcriptomics, mRNA, Heart, Kidney, Uropathy, CKD
Graphical abstract

Highlights
-
•
UUO is a useful model to study pathological cardiac remodeling in early CKD.
-
•
Extensive cardiac transcriptional changes occur in response to UUO.
-
•
Upregulated genes are enriched in cell cycle and cardiac repair pathways.
-
•
Autophagic, mitochondrial, and peroxisomal pathways are downregulated in UUO hearts.
-
•
Mitochondrial dysfunction and vimentin upregulation participate in CRS type 4.
1. Introduction
Cardiorenal syndrome (CRS) encompasses acute and chronic pathophysiological interactions between the cardiovascular and renal systems [1]. In the case of CRS type 4, primary chronic kidney disease (CKD) results in chronic heart failure (HF) [2]. Hemodynamic changes, neurohumoral disturbances, anemia, mineral metabolism disorders, and uremic toxins are some of the mechanisms that converge in CRS, complicating its understanding [1,3].
CKD affects more than 10% of the general population worldwide and is a significant cause of mortality [4]. Cardiovascular disease (CVD) is more prevalent among the CKD population, with many patients dying from CVD complications [[5], [6], [7], [8]]. Hence, understanding the pathophysiological processes associated with CKD and its cardiac consequences is vital to improving the prevention, diagnosis, and treatment of CRS type 4.
Left-ventricular hypertrophy and fibrosis are frequently associated with uremia in end-stage renal disease (ESRD) patients heavily burdened by CVD [9]. However, there is evidence of pathological cardiac performance and remodeling occurring early in CKD when the renal dysfunction is mild [[10], [11], [12], [13]].
Transcriptome profiling is valuable for discovering biomarkers and therapeutic targets. Nevertheless, the number of transcriptomic studies in experimental models of CRS remains limited. Here, we explored the cardiac transcriptomic changes in an early-stage CKD murine model induced by unilateral ureteral obstruction (UUO) and identified novel pathways involved in the pathogenesis of CRS type 4. Our results suggest that early CKD can lead to the onset of widespread transcriptional changes implicated in CVD development.
2. Method and materials
2.1. Animals
8–12-week-old male C57BL/6 mice were housed in our local animal facility with free access to water and a standard rodent diet. All animal care and research methods followed institutional and federal regulations (NOM-062-200-1999), the ARRIVE guidelines, and the National Research Council's Guide for the Care and Use of Laboratory Animals. The Local Health Research Committee of CIBO, IMSS (CLIES R-2021-1305-013) approved all the procedures.
2.2. Murine model of UUO
The UUO surgery was previously described [14]. We performed a midline incision in mice (N = 5) under xylazine (10 mg/kg)/ketamine (100 mg/kg) i.m. anesthesia to expose, ligate, and cut the left ureter before closing the midline incision. A sham group (N = 5) underwent the same procedures except for ureter ligation. Cardiac blood, kidneys, and heart were collected under anesthesia on day 21 post-surgery. Tissues were fixed in 4% paraformaldehyde or immediately frozen and stored at −80 °C.
2.3. Renal function
Serum was obtained by centrifugation at 3500 × g for 15 min at 4 °C. We determined serum creatinine (sCR) and blood urea nitrogen (BUN) levels with kits (Valtek Diagnostics, Santiago, Chile) according to the manufacturer's protocols. The absorbances were recorded with an EPOCH spectrophotometer (BioTek, Winooski, VT, USA). The mRNA expression of the injury marker neutrophil gelatinase-associated lipocalin (Ngal) was determined by RT-PCR.
2.4. RT-qPCR
RNA was isolated from the whole kidney and heart with Trizol (Invitrogen, Carlsbad, CA). The RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher Scientific, Waltham, MA, USA) was used to synthesize cDNA. RT-qPCR was performed in a QuantStudio™ 5 instrument (Thermo Fisher Scientific, Waltham, MA, USA) with Green-2-Go-qPCR-Mastermix-ROX (BioBasic, Markham, ON, Canada) and the primers listed in Supplementary_Table_S1. We used the delta-CT method to determine gene expression differences normalizing with the glyceraldehyde 3-phosphate dehydrogenase (Gapdh) gene.
2.5. Histological analysis
Paraffin-embedded 5-μm slices were dewaxed, rehydrated, and stained with hematoxylin and eosin (H&E), periodic acid-Schiff (PAS), and Sirius red. For immunohistochemistry, we used citrate buffer (10 mM, pH6.0) for antigen retrieval and the peroxidase-based Novolink Polymer Detection System (Leica Biosystems, Wetzlar, Germany) with polyclonal primary antibodies against smooth muscle alpha-actin 2 (ACTA2) and vimentin (VIM) (CST, Danvers, MA, US). Images were recorded in a DMi1 light microscope at 400X (Leica, Wetzlar, Germany) and quantified in NIH ImageJ/Fiji software.
2.6. Library construction and RNA sequencing (RNA-seq)
We performed RNA-seq to investigate mRNA expression profiles of male murine hearts 21 days after UUO surgery (N = 3) vs. sham (N = 3). First, a poly(A) RNA-seq library was prepared following Illumina's TruSeq-stranded-mRNA sample preparation protocol (Illumina, San Diego, CA, USA) after passing RNA integrity checks (RIN>7) in an Agilent Technologies 2100 Bioanalyzer (Agilent, Santa Clara, CA, USA). Poly(A) tail-containing mRNAs were purified using oligo-(dT) magnetic beads. Poly(A) RNA was fragmented at high temperatures in a divalent cation buffer. Quality control and sequencing library quantification were performed using Agilent Technologies 2100 Bioanalyzer High Sensitivity DNA Chip. Paired-ended sequencing was performed on Illumina's NovaSeq 6000 system (LC Sciences, Houston, TX, USA). The raw sequence data have been submitted to NCBI Gene Expression Omnibus (GEO) repository under accession code GSE235751.
2.7. Bioinformatic analysis
2.7.1. Transcripts Assembly
Low-quality reads were removed with Cutadapt v1.10 and in-house perl scripts. We verified sequence quality with FastQC 0.10.1 (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and used HISAT2 2.0 to map reads to ftp://ftp.ensembl.org/pub/release-101/fasta/mus_musculus/dna/genome. The mapped reads of each sample were assembled using StringTie. All transcriptomes were merged to reconstruct a comprehensive transcriptome using perl scripts and gffcompare (https://github.com/gpertea/gffcompare).
2.7.2. Differential expression analysis of mRNAs
We used StringTie 1.3.4 and ballgown (http://www.bioconductor.org/packages/release/bioc/html/ballgown.html) to estimate expression levels by calculating Fragments Per Kilobase per Million (FPKM). mRNAs differential expression analysis was performed by R package DESeq2 between two different groups (and by R package edgeR between two samples). mRNAs with a false discovery rate (FDR) < 0.05 and absolute fold-change≥2 were considered differentially expressed genes (DEGs).
2.7.3. DEGs enrichment analysis
We used the Gene Ontology (GO) project (http://www.geneontology.org) and enrichment analysis for functional interpretation of the RNA-seq experimental data. The Kyoto Encyclopedia of Genes and Genomes (KEGG) (http://www.kegg.jp/) was used to identify the significant pathways (P < 0.05) associated with enriched DEGs. Gene Set Enrichment Analysis (GSEA) (https://www.gsea-msigdb.org/gsea/index.jsp) was used to identify pathways in which several genes change a small amount but in a coordinated way [15]. Gene sets with |NES|≥ 1, NOM p-val<0.05, and FDR q-val<0.25 are considered significant.
2.8. Statistical analysis
Data are presented as mean ± SEM. We used the Mann-Whitney U test and Spearman's correlation for comparisons between two groups of non-parametric data and correlation analysis, respectively. P ≤ 0.05 was considered statistically significant. Analyses were done in GraphPad Pro (GraphPad, San Diego, CA).
3. Results
3.1. UUO as an early CKD model
UUO resulted in significant hydronephrosis at day 21 post-surgery (Fig. 1A), and the left kidney weight/body weight ratio is higher compared to sham (Fig. 1B). Histological analysis of the obstructed kidneys indicated the presence of tubular dilation, cast formation, and extracellular matrix deposition (Fig. 1C). Despite the significant damage observed in UUO mice, sCR levels were undistinguishable from sham (Fig. 1D), whereas BUN presented a slight but significant (P < 0.05) increase (Fig. 1E). In contrast, Ngal expression in the obstructed kidneys had a >130-fold change increase (P < 0.005) (Fig. 1F). At 21 days post-UUO, mice experience severe damage in the obstructed kidney. Yet, the intact contralateral kidney allows this model to be mildly uremic, thus making it valuable for studying early CKD.
Fig. 1.
Kidney injury post-UUO. (A) Sham and UUO kidneys representative images. (B) Left kidney weight/Bw ratio. (C) Histological analysis of sham and UUO kidneys with PAS and Sirius red (400X, scale bar = 100 μm). Levels of (D) sCR (mg/dL) and (E) BUN (mg/dL). (F) Ngal/Gapdh fold-change by RT-qPCR. N = 5, **P < 0.005.
3.2. Pathological remodeling in UUO hearts
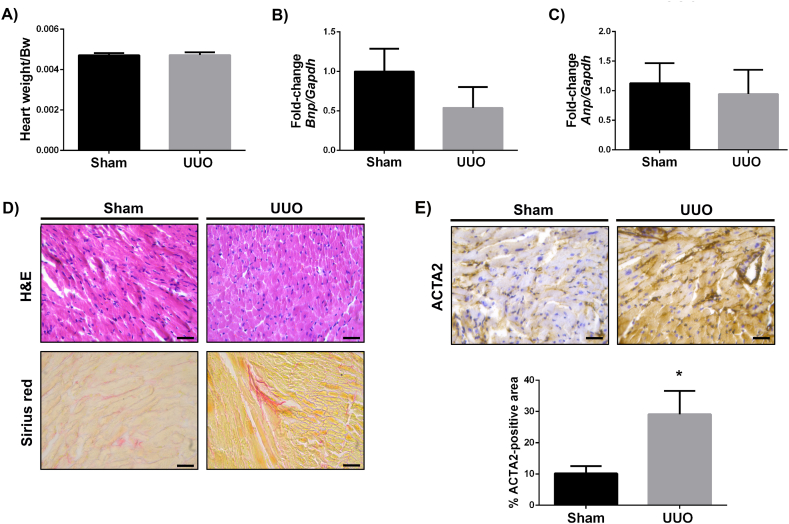
Previously, Ham O et al. reported cardiac hypertrophy and increased mRNA expression of the hypertrophy-associated genes in UUO mice at 21 days post-surgery [12]. However, we did not observe an increment in heart weight/BW ratio nor significant changes in gene expression for atrial natriuretic peptide (Anp) and brain natriuretic peptide (Bnp) in UUO animals vs. sham (Fig. 2A–C). Still, UUO induced pathological cardiac remodeling, as evidenced by increased collagen deposition and ACTA2 expression (Fig. 2D and E). These results correlate with previous reports of fibrosis development in the hearts of UUO animals that have been associated with early activation of the Transforming Growth Factor beta (TGF-β)/Smad signaling pathway [12,16].
Fig. 2.
Pathological remodeling in UUO hearts. (A) Heart weight/Bw ratio. (B) Bnp/Gapdh fold-change by RT-qPCR. (C) Anp/Gapdh fold-change by RT-qPCR. (D) Histological analysis of sham and UUO hearts with H&E and Sirius red, and (E) ACTA2 immunohistochemical detection and quantification (400X, scale bar = 100 μm). N = 5, *P < 0.05.
3.3. DEGs in UUO hearts
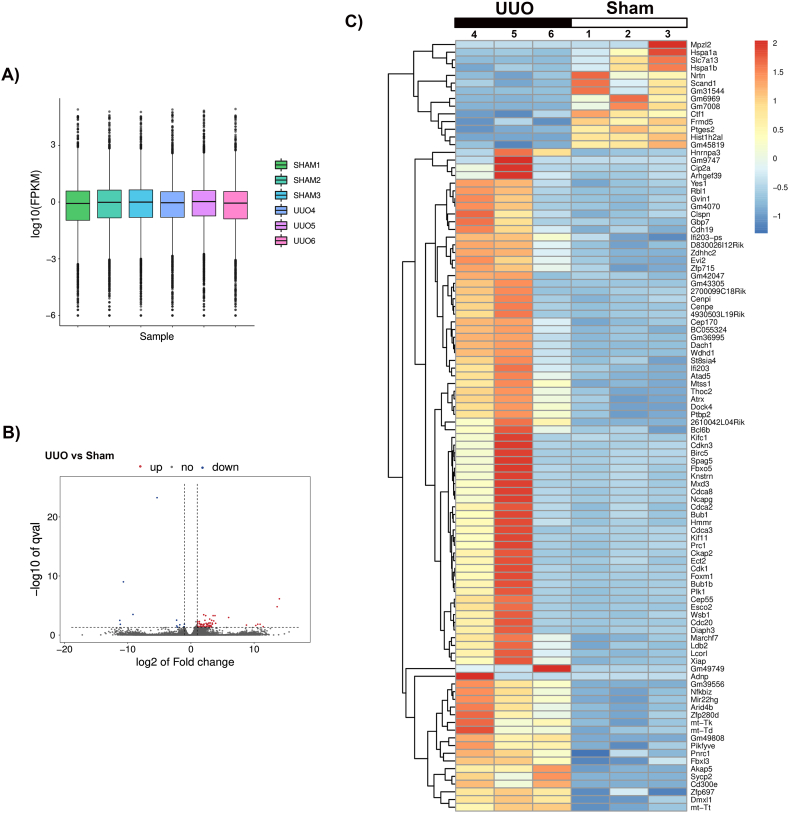
Next, we investigated mRNA expression profiles of male murine hearts 21 days after UUO surgery vs. sham. Fig. 3A shows a box plot of log2(FPKM) values across UUO and sham-expressed transcripts. RNA-seq identified 99 DEGs with a log2 (fold-change)>1. Among these, 76 DEGs (65 upregulated and 11 downregulated) were significantly and differentially expressed (P < 0.05) between UUO and sham hearts (Supplementary_file_1). The most relevant differentially expressed mRNAs are shown in Table 1. A volcano map reflects the distribution of DEGs (Fig. 3B), and a heatmap analysis illustrates DEG's expression pattern differences between experimental groups and replicates (Fig. 3C).
Fig. 3.
DEGs in UUO hearts. (A) Box plot of log2(FPKM) values. (B) Volcano map showing the distribution of DEGs in UUO vs. sham hearts. (C) Heatmap plot of DEGs. Red = upregulated and blue = downregulated. P < 0.05 and log2 (fold change) > 1. N = 3.
Table 1.
Differentially expressed cardiac mRNAs in UUO vs. sham.
| Gene | Description | Fold-change | P-value | Expression |
|---|---|---|---|---|
| Slc7a13 | solute carrier family 7, member 13 | 0.000 | 1.87E-06 | down |
| Hist1h2al | histone cluster 1, H2al | 0.001 | 7.53E-14 | down |
| Mpzl2 | myelin protein zero-like 2 | 0.002 | 6.22E-08 | down |
| Nrtn | neurturin | 0.209 | 1.37E-06 | down |
| Scand1 | SCAN domain-containing 1 | 0.211 | 5.61E-05 | down |
| Ctf1 | cardiotrophin 1 | 0.231 | 1.45E-04 | down |
| Frmd5 | FERM domain containing 5 | 0.463 | 1.02E-04 | down |
| Ptges2 | prostaglandin E synthase 2 | 0.490 | 1.67E-05 | down |
| Zfp715 | zinc finger protein 715 | 2.002 | 6.92E-05 | up |
| Ldb2 | LIM domain binding 2 | 2.084 | 1.22E-04 | up |
| Thoc2 | THO complex 2 | 2.102 | 7.26E-06 | up |
| Bcl6b | B cell CLL/lymphoma 6, member B | 2.121 | 9.53E-05 | up |
| Yes1 | YES proto-oncogene 1, Src family tyrosine kinase | 2.211 | 1.29E-04 | up |
| St8sia4 | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 4 | 2.252 | 1.35E-04 | up |
| Cdh19 | cadherin 19, type 2 | 2.293 | 1.58E-04 | up |
| Lcorl | ligand dependent nuclear receptor corepressor-like | 2.408 | 1.16E-04 | up |
| Nfkbiz | nuclear factor of kappa light polypeptide gene enhancer in B cells inhibitor, zeta | 2.544 | 3.27E-06 | up |
| Akap5 | A kinase (PRKA) anchor protein 5 | 2.551 | 1.06E-05 | up |
| Zdhhc2 | zinc finger, DHHC domain containing 2 | 2.785 | 6.22E-05 | up |
| Gbp 7 | guanylate binding protein 7 | 2.833 | 1.02E-04 | up |
| Wsb1 | WD repeat and SOCS box-containing 1 | 3.057 | 9.04E-05 | up |
| Ifi203-ps | interferon activated gene 203, pseudogene | 3.071 | 2.81E-05 | up |
| Gvin1 | GTPase, very large interferon inducible 1 | 3.271 | 2.12E-05 | up |
| Wdhd1 | WD repeat and HMG-box DNA binding protein 1 | 3.466 | 8.17E-05 | up |
| Zfp697 | zinc finger protein 697 | 3.988 | 6.59E-05 | up |
| Dach1 | dachshund family transcription factor 1 | 4.083 | 8.33E-08 | up |
| Cdc20 | cell division cycle 20 | 4.152 | 2.68E-05 | up |
| Plk1 | polo like kinase 1 | 4.567 | 1.09E-04 | up |
| Evi 2 | ecotropic viral integration site 2 | 4.666 | 1.73E-06 | up |
| Hnrnpa3 | heterogeneous nuclear ribonucleoprotein A3 | 5.077 | 1.59E-07 | up |
| Bub1b | BUB1B, mitotic checkpoint serine/threonine kinase | 5.302 | 1.04E-05 | up |
| Prc1 | protein regulator of cytokinesis 1 | 5.422 | 7.18E-05 | up |
| Clspn | claspin | 5.425 | 1.42E-04 | up |
| Knstrn | kinetochore-localized astrin/SPAG5 binding | 5.650 | 8.88E-05 | up |
| Foxm1 | forkhead box M1 | 5.708 | 2.34E-05 | up |
| Kif11 | kinesin family member 11 | 6.347 | 1.48E-05 | up |
| Fbxo5 | F-box protein 5 | 6.398 | 3.69E-05 | up |
| Cdca2 | cell division cycle associated 2 | 6.633 | 6.97E-05 | up |
| Hmmr | hyaluronan mediated motility receptor (RHAMM) | 6.938 | 8.62E-06 | up |
| Spag5 | sperm associated antigen 5 | 7.537 | 1.44E-04 | up |
| Ckap2 | cytoskeleton associated protein 2 | 7.671 | 4.25E-05 | up |
| Cdca3 | cell division cycle associated 3 | 7.726 | 1.04E-04 | up |
| Cdk1 | cyclin-dependent kinase 1 | 7.887 | 8.26E-07 | up |
| Cip2a | cell proliferation regulating inhibitor protein phosphatase 2A | 7.937 | 7.34E-06 | up |
| Birc5 | baculoviral IAP repeat-containing 5 | 8.057 | 1.42E-04 | up |
| Kifc1 | kinesin family member C1 | 8.315 | 6.67E-05 | up |
| Esco2 | establishment of sister chromatid cohesion N-acetyltransferase 2 | 8.408 | 7.60E-06 | up |
| Diaph3 | diaphanous related formin 3 | 9.062 | 4.34E-05 | up |
| Ect 2 | ect2 oncogene | 9.625 | 9.30E-05 | up |
| Cdca8 | cell division cycle associated 8 | 9.786 | 1.59E-06 | up |
| Bub1 | BUB1, mitotic checkpoint serine/threonine kinase | 10.483 | 8.43E-06 | up |
| Cdkn3 | cyclin-dependent kinase inhibitor 3 | 10.583 | 1.70E-05 | up |
| Cenpe | centromere protein E | 11.458 | 1.78E-07 | up |
| Ncapg | non-SMC condensin I complex, subunit G | 12.296 | 2.47E-06 | up |
| Cep 55 | centrosomal protein 55 | 14.075 | 1.52E-07 | up |
| Mxd3 | Max dimerization protein 3 | 15.098 | 9.97E-06 | up |
| Sycp2 | synaptonemal complex protein 2 | 440.922 | 3.66E-05 | up |
| Cd300e | CD300E molecule | 1227.930 | 4.57E-05 | up |
| Adnp | activity-dependent neuroprotective protein | 16364.127 | 8.33E-11 | up |
3.4. Upregulated DEGs are enriched in cell cycle and cell division pathways
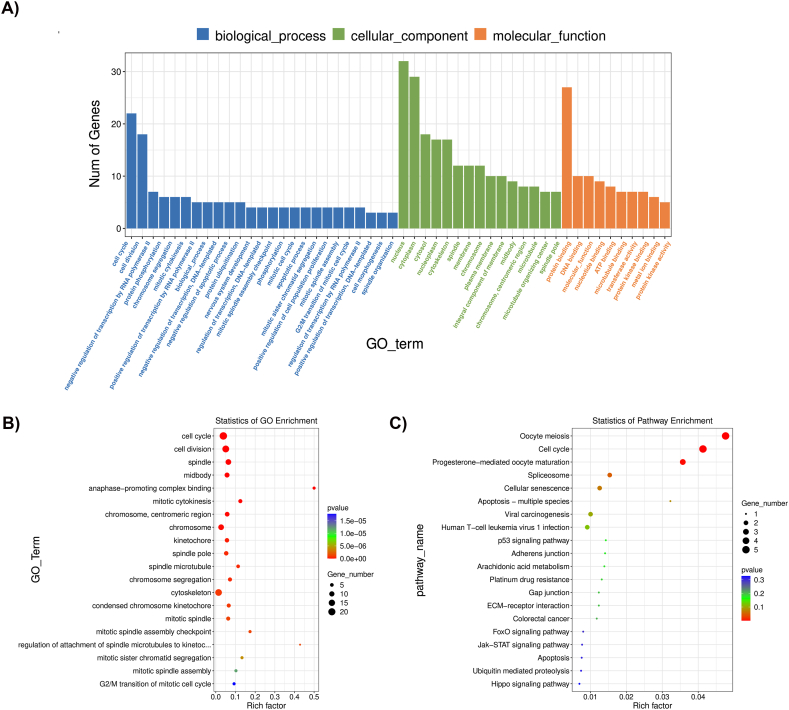
GO analysis of DEGs in UUO hearts revealed significant enrichment in 10 GO terms within three independent categories: biological process (46), cellular component (49), and molecular function (4) (Supplementary_file_2). The 50 most enriched GO terms (P < 0.05) are shown in Fig. 4A. Also, a scatterplot of the top enriched 20 GO terms indicates that several upregulated genes are enriched in cell cycle, cell division, spindle, midbody, anaphase-promoting complex binding, mitotic cytokinesis, chromosome-centromere, kinetochore, and spindle pole (Fig. 4B).
Fig. 4.
Enrichment analysis in UUO hearts. (A) Enriched GO terms. Scatter plot of the top enriched 20 (B) GO terms (C) KEGG pathways. Circle sizes, number of enriched genes per pathway, and color, P-value range. N = 3.
KEGG pathway analysis further revealed 29 different pathways in UUO hearts vs. sham, but only 4 were statistically significant (P < 0.05) (Supplementary_file_2). Consistently with the GO analysis, the cell cycle was a significantly enriched pathway along with oocyte meiosis, progesterone-mediated oocyte maturation, and spliceosome (Fig. 4C).
3.5. GSEA analysis reveals downregulated gene sets in UUO hearts
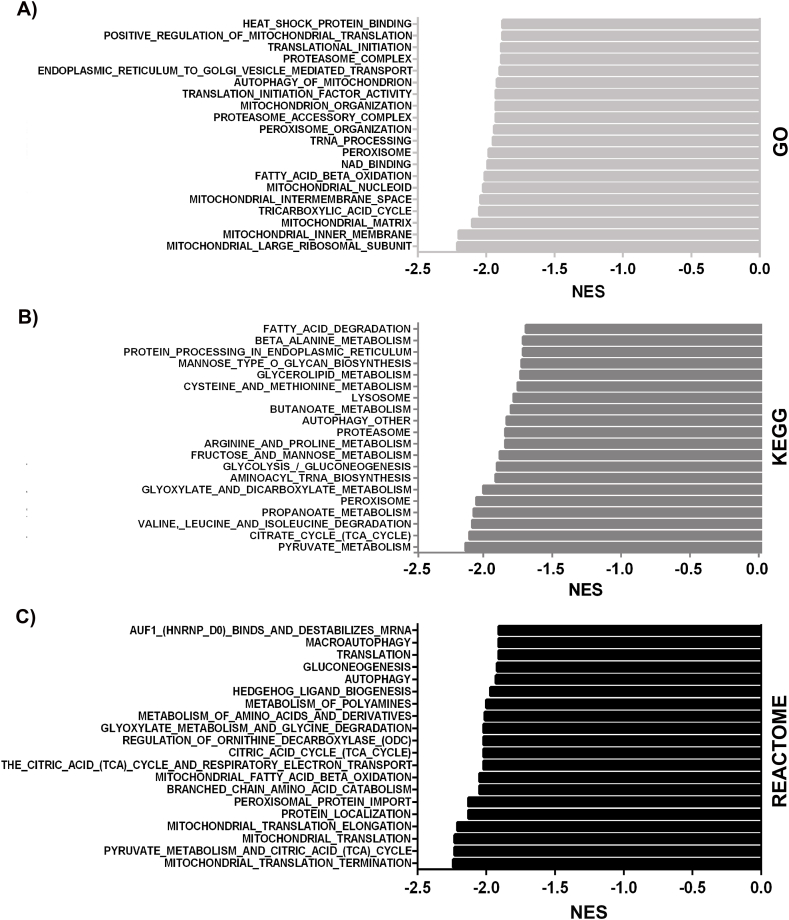
We performed GSEA analysis of DEGs in UUO hearts using three major pathway databases (GO, KEGG, and Reactome). The top 20 pathways, according to their NES (FDR q-val<0.25), are shown in Fig. 5A–C. GSEA analysis showed negative gene-set regulation related to mitochondrial oxidative bioenergetic pathways, peroxisomal processes, and autophagy. In UUO hearts, pyruvate metabolism and the tricarboxylic acid cycle, fatty-acid oxidation, glucose metabolism, respiratory electron transport, and branched-chain amino acid catabolism are highly negatively enriched mitochondrial pathways (Fig. 5A–C). Other negatively enriched signaling pathways in UUO hearts included mTOR/AMPK, retinoic acid, insulin receptors, hormone-associated pathways (prolactin, estrogen, thyroid hormone), cellular response to hypoxia (HIF1, VEGF), Hippo, WNT, and neddylation (Supplementary_file_3). Interestingly, KEGG analysis showed gene sets associated with the renin-angiotensin system as downregulated in UUO hearts (NES = −1.47, FDR q-val = 0.09). These data suggest that cardiovascular decline in CRS type 4 early in CKD progression might be associated with suppressing mitochondrial oxidative bioenergetic pathways, peroxisomal metabolic function, autophagy, and signaling pathways critical for cardiovascular function.
Fig. 5.
GSEA analysis in UUO hearts. Top 20 negatively enriched pathways (A) GO, (B) KEGG, and (C) REACTOME. |NES|≥ 1, NOM p-val<0.05, and FDR q-val<0.25. N = 3.
3.6. RT-qPCR validation of RNA-seq results
RT-qPCR was used to verify DEGs in UUO hearts. We selected 3 downregulated (Nrnt, Scand1, and Ctf1) and 6 upregulated DEGs (CD300E, Esco2, Spag5, Hmmr, Foxm1, and Plk1) (Table 1). In Fig. 6A, we compare log2 (fold-change) expression values of the selected DEGs from RNA‐seq data and RT-qPCR. Then, we performed a gene expression correlation between RT-qPCR and RNA-seq data based on the log2 (fold-change) value of DEGs (Spearman's correlation coefficient r = 0.7707 (P < 0.001) (Fig. 6B).
Fig. 6.
RT-qPCR validation. (A) Log2 (fold change) expression changes between RNA-seq (N = 3) and RT-qPCR (N = 5) for CD300E, Esco2, Spag5, Hmmr, Foxm1, Plk1, Ctf1, Scand1, and Nrnt. (B) Linear regression plot. The log2 (fold-change) values for RNA-seq and qRT-PCR are plotted along with the linear fit line. A significant Spearmanʼs correlation coefficient is also shown.
3.7. Cardiac vimentin expression is induced by UUO
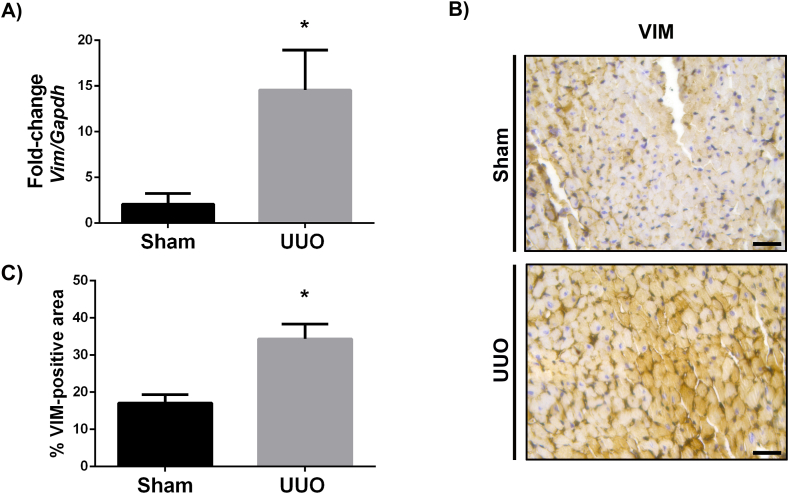
Additionally, we performed transcript analysis of RNA-seq data and found vimentin as a transcript highly expressed in UUO hearts (Supplementary_file_4). Given vimentin's role in myocardial remodeling, we verified its expression [17]. Through RT-qPCR, we found vimentin expression to be ∼15-fold higher in UUO hearts compared to sham (Fig. 7A), which correlated with increased vimentin expression in cardiac tissues (Fig. 7B and C).
Fig. 7.
Vimentin upregulation in UUO hearts. (A) Vim/Gapdh fold-change by RT-qPCR, *P < 0.05. (B) Vimentin immunohistochemical detection (400X, scale bar = 100 μm) and (C) quantification. N = 5.
4. Discussion
Efforts to understand CVD tend to focus on ESRD patients despite recent evidence pointing to cardiovascular risk increasing earlier within CKD progression, often when sCr is still within normal levels [18]. Here, we used UUO as a mildly uremic model akin to early CKD to study cardiac transcriptional changes involved in developing CRS type 4. UUO results in significant kidney injury, low uremia, pathological cardiac remodeling, and changes in cardiac gene expression.
This study used the same UUO model Ham O et al. described as an early-stage, non-uremic, CKD model in C57BL/6 mice that develops cardiac hypertrophy and fibrosis 21 days after UUO [12]. Ham O et al. showed a significant increase in sCr (∼1.7-fold) and the hypertrophy-associated markers Anp and Bnp three weeks post-obstruction, which in our hands was not reproducible despite the similarity in surgical procedure, time, and the use of C57BL/6 male mice. However, we did observe significant kidney injury, increased cardiac collagen deposition, and ACTA2 expression. Similar to our findings, Chen C et al. demonstrated the presence of myocardial fibrosis in four murine models of uremic cardiomyopathy (pole ligation, 5/6 nephrectomy, unilateral nephrectomy plus contralateral ischemia/reperfusion, and adenine) but were unable to detect increased expression of the hypertrophy biomarkers Anp and myosin heavy chain beta [19]. Due to the well-established link between uremia and cardiac hypertrophy markers, we hypothesize that our animals' modest increase in uremia after UUO resulted in lower Anp and Bnp expression than previously reported [20]. Though the lack of increase in the expression of natriuretic peptides represents a limitation, our results support the suitability of this model at 21 days to study early cardiac remodeling and molecular changes without the burden of severe kidney dysfunction and uremia consistent with the presence of an intact contralateral kidney. More in-depth studies are required to fully understand the time course for renal and cardiac marker expression after UUO.
A decline in renal function promotes angiotensin II and aldosterone production due to sodium and water accumulation, stimulates the calcium-parathyroid axis, and leads to arterial hypertension, all critical risk factors for developing CRS type 4 [[1], [2], [3]]. Even small parathyroid hormone (PTH) increases in CKD patients are associated with an elevated CVD risk independent of calcium, phosphorous, and vitamin D [21]. Hypertension raises cardiac workload, prompting structural and functional changes in the myocardium, including left ventricle hypertrophy, which can progress to HF [22]. Previous studies in rodents have demonstrated that UUO increases systolic and diastolic blood pressures as soon as two weeks after the intervention [12]. Yet, in several preclinical studies of unilateral UUO, creatinine, phosphorous, PTH, and fibroblast growth factor 23 levels are like control animals despite the confirmed presence of tubular injury [23,24]. Variations in bone mineral metabolism and serum biochemical markers of kidney injury and CVD risk in UUO models have been attributed mainly to the compensatory effect of the contralateral kidney on the excretory function, complicating their reliability at early times. In this study, we did not determine blood pressure, potassium, phosphate, or serum PTH levels. Therefore, we cannot exclude their role in the cardiac transcriptomic changes observed in mice after ureteral obstruction. Nonetheless, we found gene sets associated with the renin-angiotensin system downregulated in UUO hearts.
In UUO hearts, over 80% of DEGs were upregulated. Many were significantly enriched in cell cycle and cell division pathways related to chromatin remodeling (Adnp) and cohesin (Sycp2, Ceps5, Bub1, Esco2, Cip2a, Cdk1, Kif11, Clspn, Plk1, Cdc20, Wdhd1, Prc1), a ring-shaped protein complex involved in sister chromatid cohesion in mitosis, DNA replication and repair, genome organization, and gene expression [25,26]. Basal gene expression and myocardial stress-gene responses require a stable yet plastic chromatin architecture [27]. In cardiomyocytes, there is evidence of chromatin architecture reorganization in response to myocardial stress [28]. Moreover, cohesin depletion extensively affects gene expression, and slight decreases in cohesin levels can alter gene regulation without affecting cell division [29]. Although the direct relationship between chromatin architecture and transcriptional effects in this CRS model requires further clarification, our results suggest that early CKD can lead to the onset of extensive transcriptional changes, including the upregulation of multiple genes involved in epigenetic regulation that could drive the development of cardiovascular complications. Yet, whether these changes contribute directly to CRS pathogenesis or occur just as secondary phenomena remain to be addressed through gain- and loss-of-function in vitro and in vivo experiments of the up-and down-regulated genes.
In this transcriptomic study, we describe an abundance of changes occurring in the heart after UUO that comprise a broad spectrum of biological processes. Among them, we identified other UUO-upregulated cardiac genes (Birc5, Cd300E, Foxm1, Spag5, Hmmr, Dach1, Akap5, and Yes1) that could advance our understanding of the relationship between kidney disease and cardiac dysfunction due to their involvement in immune responses, cardiac repair, fibrosis, inflammation, proliferation, oxidative stress, and apoptosis [[30], [31], [32], [33]]. These genes could provide novel insights into CKD-related cardiac pathways contributing to CRS progression though their direct contribution needs to be validated experimentally. Importantly, we found differential expression of multiple genes and pathways in the hearts of UUO animals even without significant changes in sCr, further supporting the link between early kidney dysfunction and CVD risk [10,11,18]. Our data suggests that cardiovascular damage at the molecular level starts before traditional biomarkers of CRS can be unequivocally quantified.
The relationship between kidney function and CVD outcomes is less evident in low-risk populations compared to individuals who have some form of CVD or are at high risk for developing CVD [11]. CKD and CRS progression depend on several factors, including genetic background and sex. In animal models of UUO, the C57BL/6 mouse strain develops CKD soon after ureteral obstruction, which does not occur in other mouse strains resistant to CKD, such as BALB/c [34]. Thus, we cannot rule out that our strain choice is associated with a high prevalence of cardiovascular events. Likewise, female mice exhibit tolerance to renal damage in various animal models, and multiple studies have revealed the presence of sexual dimorphisms throughout the Cardiovascular System in health and disease [[35], [36], [37]]. A significant limitation of our study is that we only evaluated transcriptomic changes in male C57BL/6 mice, which introduces bias into our findings. Thus, an evaluation of the DEGs found in this study and their function will need to be further assessed in both sexes to determine their similarities and differences.
Initially, T cell responses promote myocardial healing through fibrosis [38]. Still, when T cell reactivity is sustained, these reparative efforts often result in adverse remodeling that, when chronic, can progress to HF. Birc5 encodes survivin, a known apoptosis inhibitor that regulates T cell responses identified as a critical gene in the progression of atrial fibrillation [39]. Knockdown or pharmacological inhibition of Birc5 in T cells attenuates acute allograft rejection survival of cardiac allografts after murine heterotopic heart transplantation by inducing apoptosis [30]. Additionally, targeting Birc5 through the lncRNA PART1/miR-503-5p pathway prevents apoptosis and improves mitochondrial function in myocardial ischemia-reperfusion injury [40]. Similarly, the myeloid cell surface receptor CD300e fine-tunes immune responses by triggering activation markers expression, releasing proinflammatory cytokines, and regulating T cell-mediated responses [41]. Importantly, focusing on T cells, including Tregs, can be a clue to reveal the reparative mechanism.
The Hippo pathway, a highly conserved signaling pathway that controls organ size, cell proliferation, apoptosis, and differentiation, has emerged as an essential regulator of cardiac development, homeostasis, and regeneration [42]. Moreover, the Yes1-associated transcriptional regulator (Yap1) leads to myofibroblast activation after myocardial infarction, and YAP1, accompanied by Foxm1, participates in various hypertrophic and fibrotic disorders [31]. YAP1 and Foxm1 upregulation reduces pathological injury in the myocardium [43,44]. In contrast, the cardiac expression of Yap1 and Foxm1 increases under hyperglycemic conditions leading to cardiomyocyte hypertrophy and fibrotic responses through increased AKT phosphorylation and Glycogen synthase kinase-3 beta inhibition [31]. FOXM1 also drives TGF-β-induced endothelial to mesenchymal transition and, in cardiac endothelial cells, cooperates with the Brahma-related gene-1 (Brg1) chromatin remodeler to trigger the angiotensin-converting enzymes pathological switch leading to angiotensin I-to-II conversion and cardiac hypertrophy [45,46]. Additionally, various studies report a cardioprotective role for A-kinase anchoring protein 5 (Akap5) by regulating calcineurin, calcium/calmodulin-dependent protein kinase II, and β-adrenergic receptors in the heart [[47], [48], [49]]. Targeting the Hippo pathway, Foxm1, and Akap5 has tremendous potential as a therapeutic strategy for cardiac fibrosis and hypertrophy, with implications for cardiovascular diseases like CRS.
Mitochondrial oxidative bioenergetic pathways, autophagy and peroxisomal pathways were also downregulated in UUO hearts, emphasizing the relevance of mitochondrial dysfunction, homeostasis deregulation, and fatty-acid metabolism alterations in CRS development. Moreover, critical signaling pathways for cardiovascular function like AMPK/mTOR, retinoic acid, HIF-1/VEGF, Hippo, WNT, and neddylation were also downregulated. These pathways have distinct molecular mechanisms for maintaining cardiac homeostasis and regulating metabolism, autophagy, endoplasmic reticulum stress, fibrosis, organ size, proliferation, regeneration, differentiation, angiogenesis, inflammation, and apoptosis [[50], [51], [52], [53], [54]]. Thus, preclinical studies considering them promising therapeutic targets for cardiovascular therapy could bring new perspectives for CRS treatments.
Furthermore, UUO induces cardiac vimentin expression as initially hinted by RNA-seq results (Supplementary_file_4), and vimentin has emerged as a myocardial remodeling marker in ischemic heart disease [17]. Although the muscle-specific desmin and laminin are the major intermediate filament proteins linked to cardiomyopathies and HF, we did not find them among the significant DEGs in UUO hearts by RNA-seq [55]. For this reason, we decided to evaluate only vimentin expression in our model. Although the role of vimentin in HF and cardiac hypertrophy is not entirely new, our results emphasize the importance of vimentin upregulation in CRS type 4.
CKD disrupts energy equilibrium, immunity, and neuroendocrine signaling and, in consequence, has systemic effects on multiple organs generating complex clinical phenotypes including inflammation, metabolic and nutritional abnormalities, autonomic and central nervous systems dysfunction, and cardiovascular, pulmonary, and bone diseases [56]. In this study, we only evaluated heart tissue, but transcriptomic changes induced by UUO probably occur in other organs. Though UUO is a widely used model of tubulointerstitial kidney fibrosis, multiple studies have reported changes in lung function, bone metabolism and structure, and cognitive and neuropathological damage following obstructive nephropathy [[57], [58], [59]]. Hence, the global changes occurring after UUO cannot be underestimated but instead studied further to understand and treat better the systemic complications of renal diseases.
In this study, we performed irreversible, unilateral UUO in which the left ureter was ligated and cut to induce hydronephrosis, severe inflammation, and interstitial scarring over 21 days as a model of early CKD. However, acute kidney injury secondary to obstructive nephropathy is frequent, impacting the morbidity and mortality of affected patients [60]. Thus, using UUO models in which the ureteric obstruction is surgically reversed (r-UUO) allows for kidney decompression and urinary flow restoration to the bladder while providing the opportunity to study the resolution of inflammation and tissue repair [34,61]. It is unclear whether removing the harmful stimulus at different times after ureteral obstruction could restore the DEGs in UUO hearts to Sham levels, mainly since there is evidence that relieving the urinary tract cannot fully recover kidney function and has residual impairments that can lead to ESRD [14]. Still, using r-UUO models in the future could be helpful to determine how the heart recovers after an acute kidney injury event.
Transcriptomic data from the whole heart is representative of changes occurring across all cell types, thus presenting a limitation of this study. More detailed, single-cell transcriptomic studies are needed to discover differences at the cellular level that the heterogeneity of the heart could be masking. Still, our results suggest that CRS type 4 development involves global transcriptional changes and highlight the importance of understanding the molecular mechanisms driving CVD risk at early points during renal disease progression.
5. Conclusions
This study identified novel pathways involved in the pathogenesis of CRS type 4 by investigating the cardiac transcriptomic changes in an early-stage CKD murine model induced by UUO. Our results suggest that early CKD can lead to the onset of widespread transcriptional changes implicated in CVD development. In UUO hearts, over 80% of DEGs were upregulated and enriched in cell cycle and cell division pathways related to chromatin remodeling, immune responses, cardiac repair, fibrosis, inflammation, proliferation, oxidative stress, and apoptosis. Interestingly, mitochondrial oxidative bioenergetic pathways, autophagy and peroxisomal pathways were downregulated in UUO hearts, emphasizing the role of altered mitochondrial function, homeostasis, and fatty-acid metabolism in the development of CRS type 4.
Institutional review board statement
The Local Health Research Committee of CIBO, IMSS (CLIES R-2021-1305-013) approved all the procedures.
Informed consent statement
Not applicable.
Data availability statement
The data that support the findings of this study are openly available in NCBI GEO repository under accession code GSE235751 and within the article's supplementary materials.
CRediT authorship contribution statement
Francisco Javier Munguia-Galaviz: Conceptualization, Formal analysis, Investigation. Yanet Karina Gutierrez-Mercado: Investigation, Resources. Alejandra Guillermina Miranda-Diaz: Conceptualization, Writing – review & editing. Eliseo Portilla de Buen: Writing – review & editing. Mario Eduardo Flores-Soto: Resources. Raquel Echavarria: Conceptualization, Formal analysis, Investigation, Writing – original draft.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgements
The graphical abstract was created with biorender.com.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.heliyon.2024.e27468.
Appendix A. Supplementary data
The following are the Supplementary data to this article.
References
- 1.Rangaswami J., Bhalla V., Blair J.E.A., Chang T.I., Costa S., Lentine K.L., Lerma E.V., Mezue K., Molitch M., Mullens W., Ronco C., Tang W.H.W., McCullough P.A., American Heart Association Council on the Kidney in Cardiovascular D, Council on Clinical C Cardiorenal syndrome: Classification, pathophysiology, diagnosis, and treatment Strategies: a scientific statement from the American heart association. Circulation. 2019;139(16):e840–e878. doi: 10.1161/CIR.0000000000000664. [DOI] [PubMed] [Google Scholar]
- 2.Clementi A., Virzi G.M., Goh C.Y., Cruz D.N., Granata A., Vescovo G., Ronco C. Cardiorenal syndrome type 4: a review. Cardiorenal Med. 2013;3(1):63–70. doi: 10.1159/000350397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hatamizadeh P., Fonarow G.C., Budoff M.J., Darabian S., Kovesdy C.P., Kalantar-Zadeh K. Cardiorenal syndrome: pathophysiology and potential targets for clinical management. Nat. Rev. Nephrol. 2013;9(2):99–111. doi: 10.1038/nrneph.2012.279. [DOI] [PubMed] [Google Scholar]
- 4.Kovesdy C.P. Epidemiology of chronic kidney disease: an update 2022. Kidney Int. Suppl. 2022;12(1):7–11. doi: 10.1016/j.kisu.2021.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jankowski J., Floege J., Fliser D., Bohm M., Marx N. Cardiovascular disease in chronic kidney disease: pathophysiological insights and therapeutic options. Circulation. 2021;143(11):1157–1172. doi: 10.1161/CIRCULATIONAHA.120.050686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Di Lullo L., Gorini A., Russo D., Santoboni A., Ronco C. Left ventricular hypertrophy in chronic kidney disease patients: from pathophysiology to treatment. Cardiorenal Med. 2015;5(4):254–266. doi: 10.1159/000435838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Tuegel C., Bansal N. Heart failure in patients with kidney disease. Heart. 2017;103(23):1848–1853. doi: 10.1136/heartjnl-2016-310794. [DOI] [PubMed] [Google Scholar]
- 8.Minciunescu A., Genovese L., deFilippi C. Cardiovascular alterations and structural changes in the setting of chronic kidney disease: a review of cardiorenal syndrome type 4. SN Compr Clin Med. 2023;5(1):15. doi: 10.1007/s42399-022-01347-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.de Albuquerque Suassuna P.G., Sanders-Pinheiro H., de Paula R.B. Uremic cardiomyopathy: a new piece in the chronic kidney disease-mineral and bone disorder puzzle. Front. Med. 2018;5:206. doi: 10.3389/fmed.2018.00206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lai S., Dimko M., Galani A., Coppola B., Innico G., Frassetti N., Mazzei E.D., Mariotti A. Early markers of cardiovascular risk in chronic kidney disease. Ren. Fail. 2015;37(2):254–261. doi: 10.3109/0886022X.2014.982489. [DOI] [PubMed] [Google Scholar]
- 11.Zhang L., Zuo L., Wang F., Wang M., Wang S., Lv J., Liu L., Wang H. Cardiovascular disease in early stages of chronic kidney disease in a Chinese population. J. Am. Soc. Nephrol. 2006;17(9):2617–2621. doi: 10.1681/ASN.2006040402. [DOI] [PubMed] [Google Scholar]
- 12.Ham O., Jin W., Lei L., Huang H.H., Tsuji K., Huang M., Roh J., Rosenzweig A., Lu H.A.J. Pathological cardiac remodeling occurs early in CKD mice from unilateral urinary obstruction, and is attenuated by Enalapril. Sci. Rep. 2018;8(1) doi: 10.1038/s41598-018-34216-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chinnappa S., White E., Lewis N., Baldo O., Tu Y.K., Glorieux G., Vanholder R., El Nahas M., Mooney A. Early and asymptomatic cardiac dysfunction in chronic kidney disease. Nephrol. Dial. Transplant. 2018;33(3):450–458. doi: 10.1093/ndt/gfx064. [DOI] [PubMed] [Google Scholar]
- 14.Hesketh E.E., Vernon M.A., Ding P., Clay S., Borthwick G., Conway B., Hughes J. A murine model of irreversible and reversible unilateral ureteric obstruction. J. Vis. Exp. 2014;94 doi: 10.3791/52559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Subramanian A., Tamayo P., Mootha V.K., Mukherjee S., Ebert B.L., Gillette M.A., Paulovich A., Pomeroy S.L., Golub T.R., Lander E.S., Mesirov J.P. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chang Y., Ben Y., Li H., Xiong Y., Chen G., Hao J., Ma X., Gao X., Qiang P., Shimosawa T., Wang X., Yang F., Xu Q. Eplerenone prevents cardiac fibrosis by inhibiting angiogenesis in unilateral urinary obstruction rats. J Renin Angiotensin Aldosterone Syst. 2022;2022 doi: 10.1155/2022/1283729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kondo T., Takahashi M., Yamasaki G., Sugimoto M., Kuse A., Morichika M., Nakagawa K., Sakurada M., Asano M., Ueno Y. Immunohistochemical analysis of vimentin expression in myocardial tissue from autopsy cases of ischemic heart disease. Leg. Med. 2022;54 doi: 10.1016/j.legalmed.2021.102003. [DOI] [PubMed] [Google Scholar]
- 18.Edwards N.C., Moody W.E., Chue C.D., Ferro C.J., Townend J.N., Steeds R.P. Defining the natural history of uremic cardiomyopathy in chronic kidney disease: the role of cardiovascular magnetic resonance. JACC Cardiovasc Imaging. 2014;7(7):703–714. doi: 10.1016/j.jcmg.2013.09.025. [DOI] [PubMed] [Google Scholar]
- 19.Chen C., Xie C., Wu H., Wu L., Zhu J., Mao H., Xing C. Uraemic cardiomyopathy in different mouse models. Front. Med. 2021;8 doi: 10.3389/fmed.2021.690517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Codognotto M., Piccoli A., Zaninotto M., Mion M., Plebani M., Vertolli U., Tona F., Ruzza L., Barchita A., Boffa G.M. Renal dysfunction is a confounder for plasma natriuretic peptides in detecting heart dysfunction in uremic and idiopathic dilated cardiomyopathies. Clin. Chem. 2007;53(12):2097–2104. doi: 10.1373/clinchem.2007.089656. [DOI] [PubMed] [Google Scholar]
- 21.Pontoriero G., Cozzolino M., Locatelli F., Brancaccio D. CKD patients: the dilemma of serum PTH levels. Nephron Clin. Pract. 2010;116(4):c263–c268. doi: 10.1159/000318787. [DOI] [PubMed] [Google Scholar]
- 22.Saheera S., Krishnamurthy P. Cardiovascular changes associated with hypertensive heart disease and aging. Cell Transplant. 2020;29 doi: 10.1177/0963689720920830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Zhang X., Li T., Wang L., Li Y., Ruan T., Guo X., Wang Q., Meng X. Relative comparison of chronic kidney disease-mineral and bone disorder rat models. Front. Physiol. 2023;14 doi: 10.3389/fphys.2023.1083725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Smith E.R., Tan S.J., Holt S.G., Hewitson T.D. FGF23 is synthesised locally by renal tubules and activates injury-primed fibroblasts. Sci. Rep. 2017;7(1):3345. doi: 10.1038/s41598-017-02709-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Horsfield J.A. Full circle: a brief history of cohesin and the regulation of gene expression. FEBS J. 2023;290(7):1670–1687. doi: 10.1111/febs.16362. [DOI] [PubMed] [Google Scholar]
- 26.Sun X., Yu W., Li L., Sun Y. ADNP controls gene expression through local chromatin architecture by association with BRG1 and CHD4. Front. Cell Dev. Biol. 2020;8:553. doi: 10.3389/fcell.2020.00553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lee D.P., Tan W.L.W., Anene-Nzelu C.G., Lee C.J.M., Li P.Y., Luu T.D.A., Chan C.X., Tiang Z., Ng S.L., Huang X., Efthymios M., Autio M.I., Jiang J., Fullwood M.J., Prabhakar S., Lieberman Aiden E., Foo R.S. Robust CTCF-based chromatin architecture underpins epigenetic changes in the heart failure stress-gene response. Circulation. 2019;139(16):1937–1956. doi: 10.1161/CIRCULATIONAHA.118.036726. [DOI] [PubMed] [Google Scholar]
- 28.Rosa-Garrido M., Chapski D.J., Schmitt A.D., Kimball T.H., Karbassi E., Monte E., Balderas E., Pellegrini M., Shih T.T., Soehalim E., Liem D., Ping P., Galjart N.J., Ren S., Wang Y., Ren B., Vondriska T.M. High-resolution mapping of chromatin conformation in cardiac myocytes reveals structural remodeling of the epigenome in heart failure. Circulation. 2017;136(17):1613–1625. doi: 10.1161/CIRCULATIONAHA.117.029430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Horsfield J.A., Anagnostou S.H., Hu J.K., Cho K.H., Geisler R., Lieschke G., Crosier K.E., Crosier P.S. Cohesin-dependent regulation of Runx genes. Development. 2007;134(14):2639–2649. doi: 10.1242/dev.002485. [DOI] [PubMed] [Google Scholar]
- 30.Xu H., Yu J., Cui J., Chen Z., Zhang X., Zou Y., Du Y., Li Y., Le S., Jiang L., Xia J., Wu J. Ablation of survivin in T cells attenuates acute allograft rejection after murine heterotopic heart transplantation by inducing apoptosis. Front. Immunol. 2021;12 doi: 10.3389/fimmu.2021.710904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Mondal A., Das S., Samanta J., Chakraborty S., Sengupta A. YAP1 induces hyperglycemic stress-mediated cardiac hypertrophy and fibrosis in an AKT-FOXM1 dependent signaling pathway. Arch. Biochem. Biophys. 2022;722 doi: 10.1016/j.abb.2022.109198. [DOI] [PubMed] [Google Scholar]
- 32.Chen Q., Tang Y., Deng H., Liang B., Li H., Li Z., Zhu H., Chen L. Curcumin improves keratinocyte proliferation, inflammation, and oxidative stress through mediating the SPAG5/FOXM1 Axis in an in vitro model of actinic dermatitis by ultraviolet. Dis. Markers. 2022;2022 doi: 10.1155/2022/5085183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wang Z., Zhang X., Zhu F., Zhou S., Wang Q., Wang H. A-kinase anchoring protein 5 anchors protein kinase A to mediate PLN/SERCA to reduce cardiomyocyte apoptosis induced by hypoxia and reoxygenation. Biochem. Cell. Biol. 2022;100(2):162–170. doi: 10.1139/bcb-2021-0466. [DOI] [PubMed] [Google Scholar]
- 34.Puri T.S., Shakaib M.I., Chang A., Mathew L., Olayinka O., Minto A.W., Sarav M., Hack B.K., Quigg R.J. Chronic kidney disease induced in mice by reversible unilateral ureteral obstruction is dependent on genetic background. Am J Physiol Renal Physiol. 2010;298(4):F1024–F1032. doi: 10.1152/ajprenal.00384.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Melo Z., Gutierrez-Mercado Y.K., Garcia-Martinez D., Portilla-de-Buen E., Canales-Aguirre A.A., Gonzalez-Gonzalez R., Franco-Acevedo A., Palomino J., Echavarria R. Sex-dependent mechanisms involved in renal tolerance to ischemia-reperfusion: role of inflammation and histone H3 citrullination. Transpl. Immunol. 2020;63 doi: 10.1016/j.trim.2020.101331. [DOI] [PubMed] [Google Scholar]
- 36.Lam C.S.P., Arnott C., Beale A.L., Chandramouli C., Hilfiker-Kleiner D., Kaye D.M., Ky B., Santema B.T., Sliwa K., Voors A.A. Sex differences in heart failure. Eur. Heart J. 2019;40(47):3859–3868c. doi: 10.1093/eurheartj/ehz835. [DOI] [PubMed] [Google Scholar]
- 37.Leinwand L.A. Sex is a potent modifier of the cardiovascular system. J. Clin. Invest. 2003;112(3):302–307. doi: 10.1172/JCI19429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kino T., Khan M., Mohsin S. The regulatory role of T cell responses in cardiac remodeling following myocardial infarction. Int. J. Mol. Sci. 2020;21(14) doi: 10.3390/ijms21145013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Zhao Z., Liu G., Zhang H., Ruan P., Ge J., Liu Q. BIRC5, GAJ5, and lncRNA NPHP3-AS1 are correlated with the development of atrial fibrillation-valvular heart disease. Int. Heart J. 2021;62(1):153–161. doi: 10.1536/ihj.20-238. [DOI] [PubMed] [Google Scholar]
- 40.Guo Z., Zhao M., Jia G., Ma R., Li M. LncRNA PART1 alleviated myocardial ischemia/reperfusion injury via suppressing miR-503-5p/BIRC5 mediated mitochondrial apoptosis. Int. J. Cardiol. 2021;338:176–184. doi: 10.1016/j.ijcard.2021.05.044. [DOI] [PubMed] [Google Scholar]
- 41.Coletta S., Salvi V., Della Bella C., Bertocco A., Lonardi S., Trevellin E., Fassan M., D'Elios M.M., Vermi W., Vettor R., Cagnin S., Sozzani S., Codolo G., de Bernard M. The immune receptor CD300e negatively regulates T cell activation by impairing the STAT1-dependent antigen presentation. Sci. Rep. 2020;10(1) doi: 10.1038/s41598-020-73552-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Meng F., Xie B., Martin J.F. Targeting the Hippo pathway in heart repair. Cardiovasc. Res. 2022;118(11):2402–2414. doi: 10.1093/cvr/cvab291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Song S., Wang Y., Wang H.Y., Guo L.L. Role of sevoflurane in myocardial ischemia-reperfusion injury via the ubiquitin-specific protease 22/lysine-specific demethylase 3A axis. Bioengineered. 2022;13(5):13366–13383. doi: 10.1080/21655979.2022.2062535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Zhang G., Yu K., Bao Z., Sun X., Zhang D. Upregulation of FoxM1 protects against ischemia/reperfusion-induced myocardial injury. Acta Biochim. Pol. 2021;68(4):653–658. doi: 10.18388/abp.2020_5536. [DOI] [PubMed] [Google Scholar]
- 45.Song S., Zhang R., Cao W., Fang G., Yu Y., Wan Y., Wang C., Li Y., Wang Q. Foxm1 is a critical driver of TGF-beta-induced EndMT in endothelial cells through Smad2/3 and binds to the Snail promoter. J. Cell. Physiol. 2019;234(6):9052–9064. doi: 10.1002/jcp.27583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yang J., Feng X., Zhou Q., Cheng W., Shang C., Han P., Lin C.H., Chen H.S., Quertermous T., Chang C.P. Pathological Ace2-to-Ace enzyme switch in the stressed heart is transcriptionally controlled by the endothelial Brg1-FoxM1 complex. Proc Natl Acad Sci U S A. 2016;113(38):E5628–E5635. doi: 10.1073/pnas.1525078113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Zhu F., Wang Q., Wang Z., Zhang X., Zhang B., Wang H. Metoprolol mitigates ischemic heart remodeling and fibrosis by increasing the expression of AKAP5 in ischemic heart. Oxid. Med. Cell. Longev. 2022;2022 doi: 10.1155/2022/5993459. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 48.Zhang X., Wang Q., Wang Z., Zhang H., Zhu F., Ma J., Wang W., Chen Z., Wang H. Interaction between A-kinase anchoring protein 5 and protein kinase A mediates CaMKII/HDAC signaling to inhibit cardiomyocyte hypertrophy after hypoxic reoxygenation. Cell. Signal. 2023;103 doi: 10.1016/j.cellsig.2022.110569. [DOI] [PubMed] [Google Scholar]
- 49.Li X., Matta S.M., Sullivan R.D., Bahouth S.W. Carvedilol reverses cardiac insufficiency in AKAP5 knockout mice by normalizing the activities of calcineurin and CaMKII. Cardiovasc. Res. 2014;104(2):270–279. doi: 10.1093/cvr/cvu209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Li X., Liu J., Lu Q., Ren D., Sun X., Rousselle T., Tan Y., Li J. AMPK: a therapeutic target of heart failure-not only metabolism regulation. Biosci. Rep. 2019;39(1) doi: 10.1042/BSR20181767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hemanthakumar K.A., Kivela R. Angiogenesis and angiocrines regulating heart growth. Vasc Biol. 2020;2(1):R93–R104. doi: 10.1530/VB-20-0006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Xie J., Wang Y., Ai D., Yao L., Jiang H. The role of the Hippo pathway in heart disease. FEBS J. 2022;289(19):5819–5833. doi: 10.1111/febs.16092. [DOI] [PubMed] [Google Scholar]
- 53.Zhao Y., Wang C., Hong X., Miao J., Liao Y., Hou F.F., Zhou L., Liu Y. Wnt/beta-catenin signaling mediates both heart and kidney injury in type 2 cardiorenal syndrome. Kidney Int. 2019;95(4):815–829. doi: 10.1016/j.kint.2018.11.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Li J., Zou J., Littlejohn R., Liu J., Su H. Neddylation, an emerging mechanism regulating cardiac development and function. Front. Physiol. 2020;11 doi: 10.3389/fphys.2020.612927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Tsikitis M., Galata Z., Mavroidis M., Psarras S., Capetanaki Y. Intermediate filaments in cardiomyopathy. Biophys Rev. 2018;10(4):1007–1031. doi: 10.1007/s12551-018-0443-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Zoccali C., Vanholder R., Massy Z.A., Ortiz A., Sarafidis P., Dekker F.W., Fliser D., Fouque D., Heine G.H., Jager K.J., Kanbay M., Mallamaci F., Parati G., Rossignol P., Wiecek A., London G., European R., Cardiovascular Medicine Working Group of the European Renal Association - European Dialysis Transplantation A The systemic nature of CKD. Nat. Rev. Nephrol. 2017;13(6):344–358. doi: 10.1038/nrneph.2017.52. [DOI] [PubMed] [Google Scholar]
- 57.Yang F., Chang Y., Zhang C., Xiong Y., Wang X., Ma X., Wang Z., Li H., Shimosawa T., Pei L., Xu Q. UUO induces lung fibrosis with macrophage-myofibroblast transition in rats. Int Immunopharmacol. 2021;93 doi: 10.1016/j.intimp.2021.107396. [DOI] [PubMed] [Google Scholar]
- 58.Gu S.S., Zhang Y., Chen X., Diao T.Y., Gebru Y., Wong M.S. Trabecular bone deterioration at the greater trochanter of mice with unilateral obstructive nephropathy. Asian J. Androl. 2013;15(4):564–566. doi: 10.1038/aja.2013.34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Ho Y.S., Lau C.F., Lee K., Tan J.Y., Lee J., Yung S., Chang R.C. Impact of unilateral ureteral obstruction on cognition and neurodegeneration. Brain Res. Bull. 2021;169:112–127. doi: 10.1016/j.brainresbull.2021.01.001. [DOI] [PubMed] [Google Scholar]
- 60.Chavez-Iniguez J.S., Navarro-Gallardo G.J., Medina-Gonzalez R., Alcantar-Vallin L., Garcia-Garcia G. Acute kidney injury caused by obstructive nephropathy. Int J Nephrol. 2020;2020 doi: 10.1155/2020/8846622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Narvaez Barros A., Guiteras R., Sola A., Manonelles A., Morote J., Cruzado J.M. Reversal unilateral ureteral obstruction: a mice experimental model. Nephron. 2019;142(2):125–134. doi: 10.1159/000497119. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data that support the findings of this study are openly available in NCBI GEO repository under accession code GSE235751 and within the article's supplementary materials.