Abstract
Emergency departments (ED) are complex, triage is a main task in the ED to prioritize patient with limited medical resources who need them most. Machine learning (ML) based ED triage tool, Score for Emergency Risk Prediction (SERP), was previously developed using an interpretable ML framework with single center. We aimed to develop SERP with 3 Korean multicenter cohorts based on common data model (CDM) without data sharing and compare performance with inter-hospital validation design. This retrospective cohort study included all adult emergency visit patients of 3 hospitals in Korea from 2016 to 2017. We adopted CDM for the standardized multicenter research. The outcome of interest was 2-day mortality after the patients’ ED visit. We developed each hospital SERP using interpretable ML framework and validated inter-hospital wisely. We accessed the performance of each hospital’s score based on some metrics considering data imbalance strategy. The study population for each hospital included 87,670, 83,363 and 54,423 ED visits from 2016 to 2017. The 2-day mortality rate were 0.51%, 0.56% and 0.65%. Validation results showed accurate for inter hospital validation which has at least AUROC of 0.899 (0.858–0.940). We developed multicenter based Interpretable ML model using CDM for 2-day mortality prediction and executed Inter-hospital external validation which showed enough high accuracy.
Subject terms: Computer science, Outcomes research
Introduction
Emergency department (ED) is complex and need urgent judgement for the better triage1,2. In order to determine the patient’s condition quickly, Korea Triage Acuity Scale (KTAS), New Early Warning Score and Modified Early Warning Score have been developed by expertise3,4. However, although most scores require complicated process to make, they are fixed score and have low reliability and poor outcome due to subjective assessment5. To solve this problem, data and machine learning (ML) based objective score has emerged6,7.
Those ML based models have problems of black box and external validation8,9. There has been some studies for interpretable triage in ED which utilized framework for interpretable scoring system called Autoscore10–12. However it was only conducted with limited population and specific for ER admission patients11. Each hospital have different population and characteristics, so we need to develop each hospital based unique score for the application.
Another tricky part for the external validation in ML research is data protection law and policy13,14. It is impossible to transfer the data into other hospital for preserving privacy. To solve this challenge, common data model (CDM) can be adopted for each hospital15. Through the CDM format, multicenter research could be done without data transfer. Standardized format of terminology and structure can be made for each hospital’s different electronic medical records format and policy. There has been some CDM based research regarding the ML16,17, there was no CDM based interpretable machine learning research in Korea.
The aim of the study is to develop, and inter-hospital external validate the interpretable ML score among the 3 big hospitals in Korea using novel framework using CDM.
Results
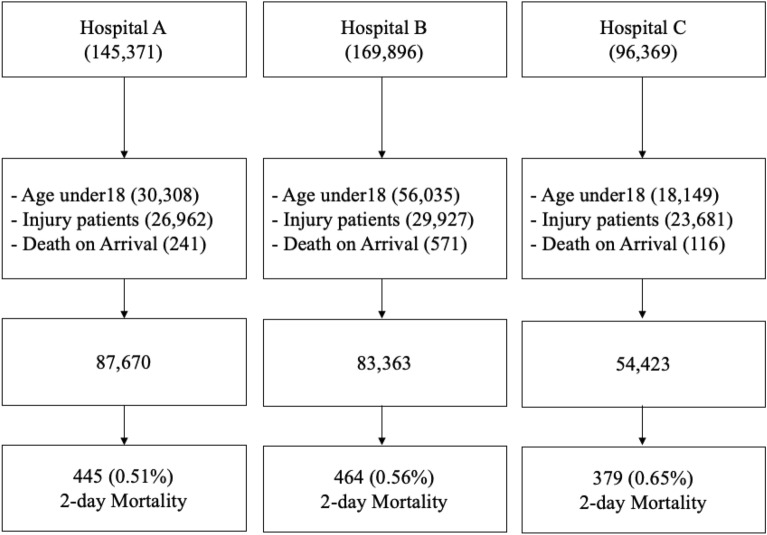
During the same study period for each hospital from 2016 to 2017 145,371, 169,896 and 96,369 patients visited ED in A, B and C respectively as shown in Fig. 1. Among them, totally 57,511, 86,533 and 41,946 patients were excluded due to age under 18, DOA, and trauma patient. Finally, 86,670, 83,363 and 54,423 patients were used for developing models. The mortality rate was from 0.51%, 0.55% and 0.65% for 2 days.
Figure 1.
Flow chart for each hospital from 2016 to 2017 emergency department visits. Age under 18, traumatic and death on arrival patient were excluded.
The distribution of ED patients’ demographics for each hospital is shown in Table 1. Each cohort included 445, 464 and 379 of events. (67.2 (14.3), 72.8 (14.4) and 72.5 (13.5) for age; 265 (59.6.%), 245 (52.8%) and 218 (57.5%) for male). Regarding the mortality patient, there were quite differences between hospitals, especially in patient conciseness of Alert at hospital A (70.8%) have higher than others (44.0 and 28.2%). Moreover, patient with severe (KTAS1 or KTAS2) at scene in hospital C (87.4%) was higher than other hospitals. (49.7% and 71.3%). Regarding the vital sign all hospital have different patterns, especially in SPO2 and BP. In terms of comorbidities history, Hospital A have much higher cancer related patients (73.9%) compared to B and C (9.5 and 5%). Whereas Hospital B and C have higher chronic disease including diabetes (28.2 and 28%). Synthetic minority over-sampling technique (SMOTE) based distribution and significance of difference for each variable were provided with standardized mean difference (SMD) were shown in Supplementary Tables 1–3.
Table 1.
Baseline Demographic for each hospital ED triage information from 2016 to 2017.
| Hospital A | Hospital B | Hospital C | |||||||
|---|---|---|---|---|---|---|---|---|---|
| No death (n = 87,225) | 2 d- mortality (n = 445) | p-value | No death (n = 82,899) | 2 d- mortality (n = 464) | p-value | No death (n = 54,044) | 2 d- mortality (n = 379) | p-value | |
| Sex | < 0.001 | 0.019 | < 0.001 | ||||||
| Male | 42,608 (48.8%) | 265 (59.6%) | 39,155 (47.2%) | 245 (52.8%) | 24,276 (44.9%) | 218 (57.5%) | |||
| Female | 44,617 (51.2%) | 180 (40.4%) | 43,744 (52.8%) | 219 (47.2%) | 29,768 (55.1%) | 161 (42.5%) | |||
| Age, mean (SD) | 55.3 ± 17.5 | 67.2 ± 14.3 | < 0.001 | 51.4 ± 19.1 | 72.8 ± 14.4 | < 0.001 | 51.7 ± 20.4 | 72.5 ± 13.6 | < 0.001 |
| Day of week | 0.202 | 0.909 | 0.288 | ||||||
| Midweek | 36,174 (41.5%) | 187 (42.0%) | 33,341 (40.2%) | 186 (40.1%) | 21,498 (39.8%) | 156 (41.2%) | |||
| Weekend | 24,699 (28.3%) | 137 (30.8%) | 25,376 (30.6%) | 143 (30.8%) | 17,053 (31.6%) | 105 (27.7%) | |||
| Friday | 12,147 (13.9%) | 47 (10.6%) | 11,331 (13.7%) | 59 (12.7%) | 7334 (13.6%) | 48 (12.7%) | |||
| Monday | 14,205 (16.3%) | 74 (16.6%) | 12,851 (15.5%) | 76 (16.4%) | 8159 (15.1%) | 70 (18.5%) | |||
| Shift time | 0.002 | 0.129 | 0.217 | ||||||
| 8 am to 4 pm | 40,217 (46.1%) | 218 (49.0%) | 35,176 (42.4%) | 200 (43.1%) | 20,596 (38.1%) | 163 (43.0%) | |||
| 4 pm to midnight | 31,533 (36.2%) | 128 (28.8%) | 30,223 (36.5%) | 183 (39.4%) | 20,974 (38.8%) | 128 (33.8%) | |||
| Midnight to 8 am | 15,475 (17.7%) | 99 (22.2%) | 17,500 (21.1%) | 81 (17.5%) | 12,474 (23.1%) | 88 (23.2%) | |||
| Triage categories | < 0.001 | < 0.001 | < 0.001 | ||||||
| 1 (most severe) | 564 (0.6%) | 73 (16.4%) | 506 (0.6%) | 84 (18.1%) | 665 (1.2%) | 197 (52.0%) | |||
| 2 | 7904 (9.1%) | 148 (33.3%) | 8809 (10.6%) | 247 (53.2%) | 7362 (13.6%) | 134 (35.4%) | |||
| 3 | 40,658 (46.6%) | 177 (39.8%) | 56,088 (67.7%) | 127 (27.4%) | 34,072 (63.0%) | 46 (12.1%) | |||
| 4 | 31,387 (36.0%) | 45 (10.1%) | 13,127 (15.8%) | 4 (0.9%) | 8986 (16.6%) | 2 (0.5%) | |||
| 5 (less severe) | 6712 (7.7%) | 2 (0.4%) | 4369 (5.3%) | 2 (0.4%) | 2959 (5.5%) | 0 (0.0%) | |||
| Consciousness | < 0.001 | < 0.001 | < 0.001 | ||||||
| Alert | 84,969 (97.4%) | 315 (70.8%) | 79,268 (95.6%) | 204 (44.0%) | 50,943 (94.3%) | 107 (28.2%) | |||
| Verbal | 1345 (1.5%) | 50 (11.2%) | 1980 (2.4%) | 88 (19.0%) | 2277 (4.2%) | 68 (17.9%) | |||
| Painful | 773 (0.9%) | 57 (12.8%) | 1485 (1.8%) | 124 (26.7%) | 723 (1.3%) | 66 (17.4%) | |||
| Unconsciousness | 138 (0.2%) | 23 (5.2%) | 166 (0.2%) | 48 (10.3%) | 101 (0.2%) | 138 (36.4%) | |||
| Route of arrival | < 0.001 | < 0.001 | < 0.001 | ||||||
| Direct | 69,632 (79.8%) | 282 (63.4%) | 65,011 (78.4%) | 216 (46.6%) | 46,039 (85.2%) | 289 (76.3%) | |||
| Other* | 17,593 (20.2%) | 163 (36.6%) | 17,888 (21.6%) | 248 (53.4%) | 8005 (14.8%) | 90 (23.7%) | |||
| Mode of transport | < 0.001 | < 0.001 | < 0.001 | ||||||
| Ambulance | 17,678 (20.3%) | 317 (71.2%) | 17,454 (21.1%) | 350 (75.4%) | 16,353 (30.3%) | 331 (87.3%) | |||
| Other* | 69,547 (79.7%) | 128 (28.8%) | 65,445 (78.9%) | 114 (24.6%) | 37,691 (69.7%) | 48 (12.7%) | |||
| Vital signs, mean (SD) | |||||||||
| Pulse, /min | 89.5 ± 20.1 | 108.0 ± 25.4 | < 0.001 | 87.9 ± 17.6 | 102.2 ± 28.1 | < 0.001 | 89.3 ± 19.0 | 100.8 ± 22.0 | < 0.001 |
| Blood pressure, mm Hg | |||||||||
| Systolic | 129.9 ± 25.0 | 118.0 ± 32.3 | < 0.001 | 131.7 ± 24.4 | 105.7 ± 30.5 | < 0.001 | 134.1 ± 24.4 | 112.6 ± 31.8 | < 0.001 |
| Diastolic | 77.1 ± 15.4 | 68.1 ± 20.1 | < 0.001 | 78.8 ± 15.4 | 63.0 ± 20.0 | < 0.001 | 81.9 ± 15.3 | 73.3 ± 21.2 | < 0.001 |
| Respiration, /min | 19.0 ± 2.4 | 22.7 ± 5.4 | < 0.001 | 16.2 ± 3.1 | 22.0 ± 6.3 | < 0.001 | 20.9 ± 2.9 | 22.0 ± 7.2 | 0.034 |
| SPo2, % | 97.3 ± 3.2 | 91.8 ± 9.8 | < 0.001 | 98.4 ± 2.2 | 93.3 ± 8.0 | < 0.001 | 97.5 ± 3.0 | 88.9 ± 12.2 | < 0.001 |
| Temperature, °C | 37.0 ± 0.8 | 36.8 ± 1.0 | < 0.001 | 36.8 ± 0.7 | 36.6 ± 1.2 | 0.008 | 36.8 ± 0.8 | 36.1 ± 1.1 | < 0.001 |
| Comorbidity | |||||||||
| Myocardial infarction | 1336 (1.5%) | 16 (3.6%) | 0.001 | 1240 (1.5%) | 22 (4.7%) | < 0.001 | 1075 (2%) | 23 (6.1%) | < 0.001 |
| Congestive heart failure | 4324 (5%) | 38 (8.5%) | 0.001 | 2042 (2.5%) | 47 (10.1%) | < 0.001 | 1349 (2.5%) | 21 (5.5%) | < 0.001 |
| Peripheral vascular disease | 2065 (2.4%) | 14 (3.1%) | 0.357 | 682 (0.8%) | 11 (2.4%) | 0.001 | 508 (0.9%) | 5 (1.3%) | 0.621 |
| Stroke | 7641 (8.8%) | 48 (10.8%) | 0.155 | 4835 (5.8%) | 48 (10.3%) | < 0.001 | 2723 (5%) | 24 (6.3%) | 0.304 |
| Dementia | 2860 (3.3%) | 26 (5.8%) | 0.004 | 905 (1.1%) | 15 (3.2%) | < 0.001 | 1235 (2.3%) | 25 (6.6%) | < 0.001 |
| Chronic pulmonary disease | 6121 (7%) | 55 (12.4%) | < 0.001 | 3820 (4.6%) | 39 (8.4%) | < 0.001 | 1800 (3.3%) | 20 (5.3%) | 0.050 |
| Rheumatoid disease | 1152 (1.3%) | 5 (1.1%) | 0.877 | 800 (1%) | 5 (1.1%) | 0.993 | 390 (0.7%) | 3 (0.8%) | 1.000 |
| Diabetes without complications | 3427 (3.9%) | 18 (4%) | 0.997 | 2281 (2.8%) | 27 (5.8%) | < 0.001 | 1460 (2.7%) | 14 (3.7%) | 0.304 |
| Diabetes with complication | 9756 (11.2%) | 75 (16.9%) | < 0.001 | 8723 (10.5%) | 131 (28.2%) | < 0.001 | 5777 (10.7%) | 106 (28%) | < 0.001 |
| Hemiplegia or paraplegia | 487 (0.6%) | 2 (0.4%) | 1.000 | 353 (0.4%) | 7 (1.5%) | 0.001 | 487 (0.9%) | 4 (1.1%) | 0.965 |
| Kidney disease | 5001 (5.7%) | 25 (5.6%) | 0.998 | 2869 (3.5%) | 41 (8.8%) | < 0.001 | 1519 (2.8%) | 17 (4.5%) | 0.071 |
| Local tumor, leukemia, and lymphoma | 31,269 (35.8%) | 329 (73.9%) | < 0.001 | 4304 (5.2%) | 44 (9.5%) | < 0.001 | 1594 (2.9%) | 19 (5%) | 0.027 |
| Metastatic solid tumor | 5516 (6.3%) | 87 (19.6%) | < 0.001 | 812 (1%) | 11 (2.4%) | 0.005 | 304 (0.6%) | 15 (4%) | < 0.001 |
| Mild liver disease | 7694 (8.8%) | 55 (12.4%) | 0.011 | 2513 (3%) | 32 (6.9%) | < 0.001 | 1,486 (2.7%) | 15 (4%) | 0.203 |
| Severe liver disease | 1321 (1.5%) | 11 (2.5%) | 0.146 | 823 (1%) | 24 (5.4%) | < 0.001 | 266 (0.5%) | 5 (1.3%) | 0.056 |
*P-value were calculated for t-test for numerical variable and chi-square test for categorical variable under 0.05 significance. SD Standard deviation,
Other Route of arrival contains transfer in, referral from outpatient, other and unknown. Other in Mode of transport contains walk-in,public transportation, Aeromedical transport, other cars, others and unknown.
Based on the variable importance from the Autoscore framework shown in Table 2 and parsimonious plot shown in Supplementary Fig. 1, we selected top 8 variables for score generations. Common feature for three hospitals were vital sign, age, patient consciousness. Vital sign such as systolic blood pressure (SBP) and heart rate (HR) were important in hospital A and B, whereas Consciousness was most important in hospital C. SBP, HR, Temperature were top 3 contributed variables in overall rank.
Table 2.
Top 14 contribution variables for each hospital.
| Top Variable | Hospital A | Hospital B | Hospital C |
|---|---|---|---|
| 1 | Systolic blood pressure | Heart rate | Consciousness |
| 2 | Heart rate | Systolic blood pressure | Systolic blood pressure |
| 3 | Temperature | Age | SpO2 |
| 4 | Diastolic blood pressure | Diastolic blood pressure | Temperature |
| 5 | Age | Temperature | Age |
| 6 | SpO2 | SpO2 | Diastolic blood pressure |
| 7 | Respiratory rate | Respiratory rate | Heart rate |
| 8 | Day of week | Day of week | Respiratory rate |
| 9 | KTAS | Consciousness | KTAS |
| 10 | Time of visit | Time of visit | Time of visit |
| 11 | Consciousness | KTAS | Day of week |
| 12 | Route of arrival | Route of arrival | Route of arrival |
| 13 | Gender | Gender | Gender |
| 14 | Ambulance use | Ambulance use | Ambulance use |
KTAS Korea Triage Acute Scale.
Scores for each hospital were presented in Table 3. The developed score for each hospital had different patterns. Among the included variables, Temperature and SpO2 were the highest effect in hospital A (17), patient consciousness for hospital B (27) and C (33). In hospital B, Age (13) was also high scored variables. Whereas Systolic blood pressure (14) was dominant at hospital C. Overall score was calculated with weighted score of number of patients and performance for each institutions. Score based on SMOTE was provided at Supplementary Table 4.
Table 3.
Score generated from each hospital.
| Score for 2-day mortality | ||||
|---|---|---|---|---|
| Variable | Hospital A | Hospital B | Hospital C | Overall |
| Age, year | ||||
| < 60 | 0 | 0 | 0 | 0 |
| 60–80 | 4 | 13 | 11 | 8 |
| ≥ 80 | 4 | 20 | 12 | 11 |
| Heart rate, /min | ||||
| < 50 | 4 | 7 | 2 | 5 |
| 50–100 | 0 | 0 | 0 | 0 |
| ≥ 100 | 9 | 7 | 2 | 7 |
| Respiration rate, /min | ||||
| < 24 | 0 | 0 | 0 | 0 |
| ≥ 24 | 13 | 7 | 6 | 10 |
| Temperature, °C | ||||
| < 24 | 17 | 7 | 10 | 12 |
| ≥ 24 | 0 | 0 | 0 | 0 |
| Blood pressure, mm Hg | ||||
| Systolic | ||||
| < 90 | 9 | 7 | 14 | 9 |
| ≥ 90 | 0 | 0 | 0 | 0 |
| Diastolic | ||||
| < 60 | 4 | 7 | 1 | 5 |
| ≥ 60 | 0 | 0 | 0 | 0 |
| SpO2, % | ||||
| < 90 | 17 | 13 | 14 | 15 |
| 90–95 | 4 | 7 | 5 | 5 |
| ≥ 95 | 0 | 0 | 0 | 0 |
| Patient consciousness | ||||
| Alert | 0 | 0 | 0 | 0 |
| Verbal | 9 | 13 | 12 | 11 |
| Painful | 13 | 20 | 19 | 17 |
| Unconsciousness | 13 | 33 | 40 | 24 |
Variables were selected from parsimonious plot shown in Supplementary Fig. 2. Overall score was calculated with weighted score for each institutions. weights are 0.472 for Hospital A, 0.410 for Hospital B and 0.116 for Hospital C.
We evaluated each score to the other hospital for the intra-institutional external validation. We used the testing cohort to evaluate the performance of each score. Table 4 depicts the AUROC with CI for the external validation which showed the best internal validation (0.913, 0.919 and 0.930) and dropped a little for the external results. Overall evaluation results show the quite good classification results from 0.904 to 0.933. Other metrics for original and SMOTE were shown in Supplementary Table 5.
Table 4.
Inter-hospital external validation result with AUROC (95% CI mortality) for each hospital.
| AUROC (Original) | Validation cohort | ||
|---|---|---|---|
| Development cohort | Hospital A | Hospital B | Hospital C |
| Hospital A | 0.913 (0.882–0.945) | 0.9124 (0.884–0.9407) | 0.928 (0.902–0.955) |
| Hospital B | 0.893 (0.854–0.931) | 0.919 (0.891–0.946) | 0.930 (0.902–0.958) |
| Hospital C | 0.885 (0.842–0.927) | 0.929 (0.9015–0.950) | 0.930 (0.899–0.960) |
| Overall | 0.904 (0.866–0.942) | 0.929 (0.9049–0.952) | 0.933 (0.904–0.961) |
AUROC area under the receiver operating characteristic.
Discussion
In this study, we developed interpretable score based on CDM Autoscore for ED and evaluated with 3 tertiary hospitals in Korea for inferring the 2-day mortality for ED visit patients. Although each hospitals have different characteristics, scores were accurate for their external validation results for other institutions which has at least of 0.885 (0.842–0.942) AUROC. Moreover, it was interpretable score, so it can be integrated easily into clinical practice. We found each scores from their own hospital, which is the internal validation results were accurate from 0.913 to 0.930 AUROC. We also identified the extent of lack of accuracy and acceptance when we apply the score to other institute.
To the best of our knowledge, this is the first study for interpretable machine learning using CDM framework in ED. Many policies or laws regarding the data protection or leak was published for the protection of private patient information18,19. For solving these problems, our framework can share the result without any transferring patient data. CDM is designed to standardize the structure and vocabulary of observational health data that can produce reliable evidence without sharing data. This approach creates a unique opportunity of implementing several existing data exploration and evidence generation tools and participating in world-wide distributed research network studies without raw data leakage20–22. Extensibility and generatability can be obtained based on our framework. More institutions can be added to analysis cohort for further development and validation because of the developed semi-automated ETL process enables CDM conversion for all institution’s NEDIS data in Korea.
Interpretable point-based score can be easily utilized for the real practice. A paper published from Netherlands in 2023 also developed international early warning score for predicting mortality in ED23. The score was consistent with our interpretable score in terms of having high impact on consciousness, systolic blood pressure and temperature and Spo2. Whereas old age was most impact factor in international score.
Another novelty for this study is it conducted the cross-external validation for identifying the generalizability. Patient distribution is different for each institution. In case of hospital C, almost mortality patients had severe KTAS level and consciousness was most important for predicting mortality. We need to develop each score for institution. Many previous study emphasized the importance of external validation for the generality of model14,24,25. Most of the studies conducted one model from one site to other sites26,27, but in this study all institutions made their one score and we can compare the results for each one.
There are some limitations for this study, first it was a retrospective, the score needs to be evaluated in prospectively for the checking the applicability. However, this score-based model development is easy to apply to EMR integration because of advantages of point-based score. Second, we need to consider the representative score for Korea. We can develop with national emergency department information system data which is data from 403 ED data for developing national level score for Korea.
In summary, we developed the K-SERP score for 3 hospitals in Korea using CDM Autoscore for ED and showed good cross-external validation results which were at least 0.899 of AUROC. We can expand the result with other emergency department site based on CDM framework. Each score could be interpreted and applied to clinical process easily.
Method
Study design and setting
This retrospective and validation study was executed across from 3 ED in Korea (A, B and C). A, B and C are tertiary hospitals located in a metropolitan city in Korea. Respectively, the hospital has approximately 2000, 1000, and 1000 inpatient beds. Approximately more than 80,000, 90,000 and 50,000 patients visit the ED annually. There are 16, 20 and 7 specialists working at each institution, respectively. All data were mapped to the Observational Medical Outcome Partnership Common Data Model (OMOP-CDM) for the multicenter study. This study was approved by the Samsung Medical Center Institutional Review Board (2023-02-036), and a waiver of informed consent was granted for EHR data collection and analysis because of the retrospective and de-identified nature of the data. All methods were performed in accordance with the relevant guidelines and regulations.
Selection of participants
Initially, ED patients from 2016 to 2017 were included for each hospital. Patient older than 18 with disease patients were included. We also excluded patient with left without being seen or death on arrival/cardiopulmonary resuscitation patients. We split into two cohort: development (70%) cohort for training the interpretable ML model and test (30%) for evaluation from each hospital.
Candidate predictors
We extracted data from each hospital’s electronic medical records system which all patient information was deidentified. Candidate input variables were considered with available features at the stage of ED triage including demographic characteristics such as age, gender, administrative variables including time of ED visit and clinical variables such as severity index, consciousness, and initial vital sign. Comorbidities were also obtained from hospital diagnosis records in the preceding 5 years before patients’ emergency visit and compared for each hospital. They were extracted from International Statistical Classification of Diseases and Related Health Problems, Tenth Revision (ICD-10). The list and description of candidate predictors and comorbidities are given in the supplementary Tables 6 and 7.
Outcomes
Emergency patients with semi-acute conditions typically undergo surgical procedure or are admitted to Intensive care unit (ICU) following emergency room treatment and given the imperative for patients to survive. Our primary outcome was 2-day mortality which was the target feature for analysis to build the interpretable ML model for each hospital.
Common data model (CDM)
For the multicenter study, we adopted OMOP CDM from the research network Observational Health Data Sciences and Informatics (OHDSI)28 for standardized structure and vocabularies to map emergency department data based on Systematized Nomenclature of Medicine–Clinical Terms (SNOMED-CT) and Logical Observation Identifiers Names and Codes (LOINC) as example shown Supplementary Fig. 1. Extract, Transformation and Load (ETL) process was performed with structured query language. Each ED care and diagnosis related information was mapped into proper CDM tables as shown in Fig. 2. For example, patient demographics and vital sign are mapped to Person and Measurement table, respectively. After transformation was completed into CDM format, all hospital can get the same structure and vocabularies, for executing same research query. All details of transformation and code are accessible on Gitgub29.
Figure 2.
Table mapping for converting clinical to common data model tables. CDM: common data model; ED: Emergency department.
CDM autoscore for ED framework
AutoScore Framework is a machine learning-based clinical score generator, consisting of six modules developed from Singapore12. Module 1 uses a random forest for ranking variables according to their importance. Module 2 transforms variables by categorizing continuous variables to improve interpretation with quantile information. Module 3 makes scores for each variable based on a logistic regression coefficient. Module 4 selects which variables could be included in the scoring model. In Module 5, clinical domain knowledge is incorporated to the score and cutoff points can be defined when categorizing continuous variables. Module 6 evaluates the performance of the score in a separate test dataset. The AutoScore framework provides a systematic and automated approach to develop score automatically, combining of advantage of machine learning for discriminating and the strength of logistic regression in its interpretability. For the overall score generation, We considered weighted average scores across all institutions. For each institutions i, a weight was formulated as = / × 100% where was the sample size, was the AUC value obtained based on the validation set, and M was the total number of institutions. Overall score was calculated with weighted score based on .
We defined our new novel framework “CDM Autoscore for ED”, combination of CDM based standardized format and autoscore based interpretable framework shown in Fig. 3. The analysis and preparation code using CDM format was also shared on GitHub29.
Figure 3.
Overall process of “CDM Autoscore for ED”. Each Institutions conducted Extract, Transformation and Load process for converting local data into CDM format. Algorithms from each of institution were derived using interpretable machine learning framework and validated inter-and intra- institutionally. EMR: Electronic medical records; ETL: Extract, transformation and Load; OMOP CDM: Observational Medical Outcome Partnership Common Data Model.
Statistical analysis
Categorical features were expressed as frequency and percentages and continuous features were expressed as means and standard deviations. Comparison tests for each hospital were performed with analysis of variance and chi-square tests at 5% significance levels. Standardized mean difference (SMD) was also calculated for comparing each hospital. Two types of validations for this study were conducted. First, we executed internal-institutional validation for each hospital’s score. We also performed intra-institutional validation pair-wisely for the external validation. Area under the curve in the receiver operating characteristic (AUROC) and 95% confidence interval (CI) with 1000 times of bootstrap was reported. Other metrics including accuracy, sensitivity, specificity, positive predictive value (PPV) and negative predictive value (NPV) were also reported. SMOTE was conducted for handling the imbalance problem. Twice of minority was oversampled and same number of majorities according to the number of minority was sampled with fixed seed number.
Supplementary Information
Abbreviations
- ED
Emergency department
- ML
Machine learning
- SERP
Score for emergency risk prediction
- CDM
Common data model
- AUROC
Area under receiver operating curve
- KTAS
Korea triage acuity scale
- SMOTE
Synthetic minority over-sampling technique
- SMD
Standardized mean difference
- OMOP-CDM
Observational medical outcome partnership common data model
- OHDSI
Observational health data sciences and informatics
- SNOMED-CT
Systematized nomenclature of medicine–clinical terms
- LOINC
Logical observation identifiers names and codes
- ETL
Extract transformation load
- EMR
Electronic medical records
- CI
Confidence interval
- SD
Standard deviation
Author contributions
Conceptualization: W.C.C.; data curation: J.Y.Y., S.Y.; formal analysis: J.Y.Y. D.Y.K; investigation: X.F.; methodology: X.F., M.E.H.O.; visualization: J.Y.Y.; writing—original Draft: J.Y.Y., writing—review and editing: J.Y.Y., H.J.J., K.W.J., R.W.P., J.M.G.,G.S.H., X.F., M.E.H.O., Y.Y.N., W.C.C.
Funding
This research was supported by a grant of Korea Health Technology R&D Project throught the Korea Health Industry Development Institute (KHIDI) and the Medical data-driven hospital support project through the Korea Health Information Service (KHIS), funded by the Ministry of Health & Welfare, Republic of Korea (Grant Number: HI19C1328).
Data availability
Data was available in study site clinical data warehouse. The datasets generated and analyzed during the current study are not publicly available due dataset includes although is de-identifed, part of patient information, but are available from the corresponding author on reasonable request.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Jae Yong Yu and Doyeop Kim.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-024-54364-7.
References
- 1.Hoot NR, Aronsky D. Systematic review of emergency department crowding: Causes, effects, and solutions. Ann. Emerg. Med. 2008;52:126–136. doi: 10.1016/j.annemergmed.2008.03.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Petrie DA, Comber S. Emergency department access and flow: Complex systems need complex approaches. J. Eval. Clin. Pract. 2020;26:1552–1558. doi: 10.1111/jep.13418. [DOI] [PubMed] [Google Scholar]
- 3.Mitsunaga T, Hasegawa I, Uzura M, et al. Comparison of the National Early Warning Score (NEWS) and the Modified Early Warning Score (MEWS) for predicting admission and in-hospital mortality in elderly patients in the pre-hospital setting and in the emergency department. PeerJ. 2019;7:e6947. doi: 10.7717/peerj.6947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kwon H, Kim YJ, Jo YH, et al. The Korean triage and acuity scale: Associations with admission, disposition, mortality and length of stay in the emergency department. Int. J. Qual. Health Care. 2019;31:449–455. doi: 10.1093/intqhc/mzy184. [DOI] [PubMed] [Google Scholar]
- 5.Choi H, Ok JS, An SY. Evaluation of validity of the Korean triage and acuity scale. J. Korean Acad. Nurs. 2019;49:26–35. doi: 10.4040/jkan.2019.49.1.26. [DOI] [PubMed] [Google Scholar]
- 6.Liu Y, Gao J, Liu J, et al. Development and validation of a practical machine-learning triage algorithm for the detection of patients in need of critical care in the emergency department. Sci. Rep. 2021;11:24044. doi: 10.1038/s41598-021-03104-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Yu JY, Jeong GY, Jeong OS, Chang DK, Cha WC. Machine learning and initial nursing assessment-based triage system for emergency department. Healthc. Inform. Res. 2020;26:13–19. doi: 10.4258/hir.2020.26.1.13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mueller B, Kinoshita T, Peebles A, Graber MA, et al. Artificial intelligence and machine learning in emergency medicine: A narrative review. Acute Med. Surg. 2022;9:e740. doi: 10.1002/ams2.740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Dugas AF, Kirsch TD, Toerper M, et al. An electronic emergency triage system to improve patient distribution by critical outcomes. J. Emerg. Med. 2016;50:910–918. doi: 10.1016/j.jemermed.2016.02.026. [DOI] [PubMed] [Google Scholar]
- 10.Yun H, Choi J, Park JH. Prediction of critical care outcome for adult patients presenting to emergency department using initial triage information: An XGBoost algorithm analysis. JMIR Med. Inform. 2021;9:e30770. doi: 10.2196/30770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Xie F, Ong MEH, Liew J, et al. Development and assessment of an interpretable machine learning triage tool for estimating mortality after emergency admissions. JAMA Netw. Open. 2021;4:e2118467. doi: 10.1001/jamanetworkopen.2021.18467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Xie F, Chakraborty B, Ong MEH, Goldstein BA, Liu N. AutoScore: A machine learning-based automatic clinical score generator and its application to mortality prediction using electronic health records. JMIR Med. Inform. 2020;8:e21798. doi: 10.2196/21798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ramspek CL, Jager KJ, Dekker FW, Zoccali C, van Diepen M. External validation of prognostic models: What, why, how, when and where? Clin. Kidney J. 2020;14:49–58. doi: 10.1093/ckj/sfaa188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Riley RD, Ensor J, Snell KIE, et al. External validation of clinical prediction models using big datasets from e-health records or IPD meta-analysis: Opportunities and challenges. BMJ. 2016;353:i3140. doi: 10.1136/bmj.i3140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Reps JM, Williams RD, You SC, et al. Feasibility and evaluation of a large-scale external validation approach for patient-level prediction in an international data network: Validation of models predicting stroke in female patients newly diagnosed with atrial fibrillation. BMC Med. Res. Methodol. 2020;20:102. doi: 10.1186/s12874-020-00991-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Choi YI, Park SJ, Chung JW, et al. Development of machine learning model to predict the 5-year risk of starting biologic agents in patients with inflammatory bowel disease (IBD): K-CDM network study. J. Clin. Med. 2020;9:3427. doi: 10.3390/jcm9113427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ryu B, Yoo S, Kim S, Choi J. Development of prediction models for unplanned hospital readmission within 30 days based on common data model: A feasibility study. Methods Inf. Med. 2021;60:e65–e75. doi: 10.1055/s-0041-1735166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kim Y. Uncertain future of privacy protection under the Korean public health emergency preparedness governance amid the COVID-19 pandemic. Cogent Soc. Sci. 2022;8:2006393. [Google Scholar]
- 19.Lee D, Park M, Chang S, Ko H. Protecting and utilizing health and medical big data: Policy perspectives from Korea. Healthc. Inform. Res. 2019;25:239–247. doi: 10.4258/hir.2019.25.4.239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.You SC, Rho Y, Bikdeli B, Kim J, Siapos A, et al. Association of ticagrelor vs clopidogrel with net adverse clinical events in patients with acute coronary syndrome undergoing percutaneous coronary intervention. JAMA. 2020;324(16):1640–1650. doi: 10.1001/jama.2020.16167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Schuemie MJ, Ryan PB, Pratt N, Chen R, You SC, et al. Principles of large-scale evidence generation and evaluation across a network of databases (LEGEND) J. Am. Med. Inform. Assoc. 2020;27(8):1331–1337. doi: 10.1093/jamia/ocaa103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Burn E, You SC, Sena AG, Kostka K, Abedtash H, Abrahão MTF, Alberga A, et al. Deep phenotyping of 34,128 adult patients hospitalised with COVID-19 in an international network study. Nat. Commun. 2020;11(1):5009. doi: 10.1038/s41467-020-18849-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Candel BGJ, Nissen SK, Nickel CH, et al. Development and external validation of the international early warning score for improved age- and sex-adjusted in-hospital mortality prediction in the emergency department. Crit. Care Med. 2023;51:881–891. doi: 10.1097/CCM.0000000000005842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Bleeker SE, Moll HA, Steyerberg EW, et al. External validation is necessary in prediction research: a clinical example. J. Clin. Epidemiol. 2003;56:826–832. doi: 10.1016/S0895-4356(03)00207-5. [DOI] [PubMed] [Google Scholar]
- 25.Collins GS, de Groot JA, Dutton S, et al. External validation of multivariable prediction models: A systematic review of methodological conduct and reporting. BMC Med. Res. Methodol. 2014;14:40. doi: 10.1186/1471-2288-14-40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lee YJ, Cho KJ, Kwon O, et al. A multicentre validation study of the deep learning-based early warning score for predicting in-hospital cardiac arrest in patients admitted to general wards. Resuscitation. 2021;163:78–85. doi: 10.1016/j.resuscitation.2021.04.013. [DOI] [PubMed] [Google Scholar]
- 27.Kwon JM, Kim KH, Jeon KH, et al. Development and validation of deep-learning algorithm for electrocardiography-based heart failure identification. Korean Circ. J. 2019;49:629–639. doi: 10.4070/kcj.2018.0446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hripcsak G, Duke JD, Shah NH, Reich CG, Huser V, Schuemie MJ, Suchard MA, Park RW, Wong IC, Rijnbeek PR, van der Lei J, Pratt N, Norén GN, Li YC, Stang PE, Madigan D, Ryan PB. Observational health data sciences and informatics (OHDSI): Opportunities for observational researchers. Stud. Health Technol. Inform. 2015;216:574–578. [PMC free article] [PubMed] [Google Scholar]
- 29.Kim DY. NEDIS CDM github GitHub. https://github.com/OHDSI/ETL---Korean-NEDIS.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data was available in study site clinical data warehouse. The datasets generated and analyzed during the current study are not publicly available due dataset includes although is de-identifed, part of patient information, but are available from the corresponding author on reasonable request.