Summary
Faster vegetative growth and early maturity/harvest reduce plant life cycle time and are important agricultural traits facilitating early crop rotation. GA is a key hormone governing developmental transitions that determine growth speed in plants. An EAR‐motif repressor, SlERF36 that regulates various growth transitions, partly through regulation of the GA pathway and GA levels, was identified in tomato. Suppression of SlERF36 delayed germination, slowed down organ growth and delayed the onset of flowering time, fruit harvest and whole‐plant senescence by 10–15 days. Its over‐expression promoted faster growth by accelerating all these transitions besides increasing organ expansion and plant height substantially. The plant life cycle and fruit harvest were completed 20–30 days earlier than control without affecting yield, in glasshouse as well as net‐house conditions, across seasons and generations. These changes in life cycle were associated with reciprocal changes in expression of GA pathway genes and basal GA levels between suppression and over‐expression lines. SlERF36 interacted with the promoters of two GA2 oxidase genes, SlGA2ox3 and SlGA2ox4, and the DELLA gene, SlDELLA, reducing their transcription and causing a 3–5‐fold increase in basal GA3/GA4 levels. Its suppression increased SlGA2ox3/4 transcript levels and reduced GA3/GA4 levels by 30%–50%. SlERF36 is conserved across families making it an important candidate in agricultural and horticultural crops for manipulation of plant growth and developmental transitions to reduce life cycles for faster harvest.
Keywords: early maturity, flowering time, GA2 oxidase, AP2‐ERF domain, DELLA, gibberellins
Introduction
Plant growth is marked by multiple growth cycles and transitions leading from seed germination to organ expansion, adult growth, onset of flowering, fruit formation and whole‐plant senescence. These growth transitions occur within a defined period primarily in response to developmental cues but are also strongly influenced by changes in the external environment. The duration of the growth cycles in plant life is important in agriculture since short durations can allow farmers to grow multiple crops in a year, reduce water/nutrient losses by early cropping and minimize exposure of the plant to vagaries of the external environment. The onset of growth transitions is regulated by a balance between different hormones and ensures completion of the life cycle in a given time frame. Any change in hormonal balance, brought about by a change in expression of regulatory genes, can alter the speed of transitions. GA is one of the key hormones that controls these transitions (Castro‐Camba et al., 2022; Weiss and Ori, 2007; Yamaguchi, 2008). Its interaction with other hormones governs the outcome of developmental changes leading to seed germination (Finch‐Savage and Leubner‐Metzger, 2006; Li et al., 2019b; Liu and Hou, 2018), bud break (Liu and Sherif, 2019; Wen et al., 2016), organ expansion (Achard et al., 2006; Fu and Harberd, 2003; Li et al., 2012), stem and tuber development (Chen et al., 2022), flowering (Bauerle, 2022; Blazquez et al., 1998; Izawa, 2020), fruit set (Serrani et al., 2008; Shinozaki et al., 2015) and adaptation (Dubois et al., 2013; Kuroha et al., 2018; Shohat et al., 2020, 2021a, 2021b). Changes in basal GA levels and/or responses change plant growth characteristics as well as stress responses and have had far‐reaching effects in agriculture including development of varieties that led to the Green Revolution in the sixties (Hedden, 2003; Khush, 1999).
GA levels are regulated in different tissues through fine control over GA biosynthesis by a family of GA20/GA3 oxidases while reduction in levels of active GAs is brought about by a family of GA2 oxidases. Multiple genes encoding these enzymes are differentially transcribed and collectively regulate stem elongation, leaf expansion, branching, flowering initiation, fruit set, trichome development, stresses etc. (Castro‐Camba et al., 2022; Chen et al., 2016; Illouz‐Eliaz et al., 2019, 2020; Kozaki and Aoyanagi, 2022; Li et al., 2012, 2019a; Magome et al., 2008; Martínez‐Bello et al., 2015; Matías‐Hernández et al., 2016; Monna et al., 2002; Shohat et al., 2020, 2021b; Yanai et al., 2011). A great deal of regulation is also exerted through GA signalling by GA receptors that facilitate GA action (Illouz‐Eliaz et al., 2019; Sun, 2010) and by DELLA proteins that suppress GA action, inhibit growth and thereby allow the plant to respond to environmental challenges (Achard et al., 2006; Colebrook et al., 2014; Sun, 2010). Despite considerable studies on how GA pathway components affect growth in plants like Arabidopsis, rice and tomato, the regulation of individual components of the pathway across development remain largely un‐deciphered.
The AP2 domain family of transcription factors is large family governing various aspects of development and adaptation in plants. It is characterized by the presence of a conserved 58–59 amino acid DNA‐binding AP2 domain with further functional sub‐groups based on distinctions within the domains and the presence of additional domains flanking these (Nakano et al., 2006; Pirrello et al., 2012). While most AP2 domain family members function as activators of transcription, a small sub‐group (with 8 members in Arabidopsis and 7 in tomato) is characterized by the presence of a short C‐terminal repressor motif (L/F)DLN(L/F)xP designated as the ERF‐associated amphiphilic repression (EAR) motif (Ohta et al., 2001). EAR‐motif proteins function as dominant repressors of transcription in combination with co‐repressors and histone deacetylases (HDACs) and are responsible for chromatin modification (Kagale and Rozwadowski, 2011). Detailed studies on some of these have demonstrated roles in governing sensitivity to hormones like ABA, ethylene, JA in processes like germination, fruit ripening, stomatal responses, abiotic stresses and herbivory (Deng et al., 2022; Licausi et al., 2013; Lu et al., 2011; McGrath et al., 2005; Plant et al., 2021; Song et al., 2005; Upadhyay et al., 2013; Yang et al., 2005). Nevertheless, detailed analysis of these genes remains a challenge in most plants.
In this paper, we show that SlERF36, an AP2 domain EAR‐motif suppressor in tomato, controls the timing of developmental transitions in tomato, partly by regulating GA levels and GA signalling by reducing the expression of the GA2 oxidase genes, GA2ox3 and GA2ox4, and the sole DELLA gene in tomato, SlDELLA. Changes brought about by SlERF36 alter plant growth cycle reproducibly by 30–40 days under glasshouse as well as nethouse conditions demonstrating the robustness of its regulation and usefulness of the gene under field conditions.
Results
SlERF36 is an EAR‐motif repressor and its levels regulate germination and organ expansion during juvenile/adult vegetative growth
SlERF36 was previously identified in our lab as an EAR‐motif ERF that accelerated flowering time when ectopically expressed in tobacco and Arabidopsis (Upadhyay et al., 2013, 2014). It is expressed ubiquitously across tissues at varying levels (Figure S1a). In order to elucidate its function in tomato, transgenic lines expressing SlERF36 in the sense and antisense orientations under the CaMV35S promoter were generated and monitored for various phenotypic changes ranging from seed germination to senescence. Three SlERF36 over‐expression lines, OEx1, OEx3 and OEx4 (with 3–5‐fold higher transcript levels in seedlings and 4–6‐fold higher transcript levels in leaves), and three suppression lines, Sup3, Sup6, and Sup7 (with transcript levels reduced to 36%–44% in seedlings and leaves), were chosen from ten lines generated for the study (Figure S1b). The expression of a closely related gene, SlERF37, was first monitored to ensure that its expression did not undergo a change in the transgenic lines (Figure S1c).
Detailed phenotypic analysis of transgenic SlERF36 lines revealed clear changes in the timing of growth transitions and the speed of plant growth, starting with germination. Compared to control seeds which germinated between days 2–5, seeds of all the over‐expression lines germinated from day 1 itself and completed germination by day 4. In contrast, the seeds of all suppression lines began germinating between days 3 (lines 6 and 7) and 6 (line 3) and reached completion only by days 9–11 (Figure 1a). Even at day 5, when control and OEx lines had completely germinated, suppression lines showed less than 20% germination. The most prominent differences were observed on day 2, with nearly 80% germination in over‐expression lines, 55% germination in control seeds, but no germination in suppression lines.
Figure 1.
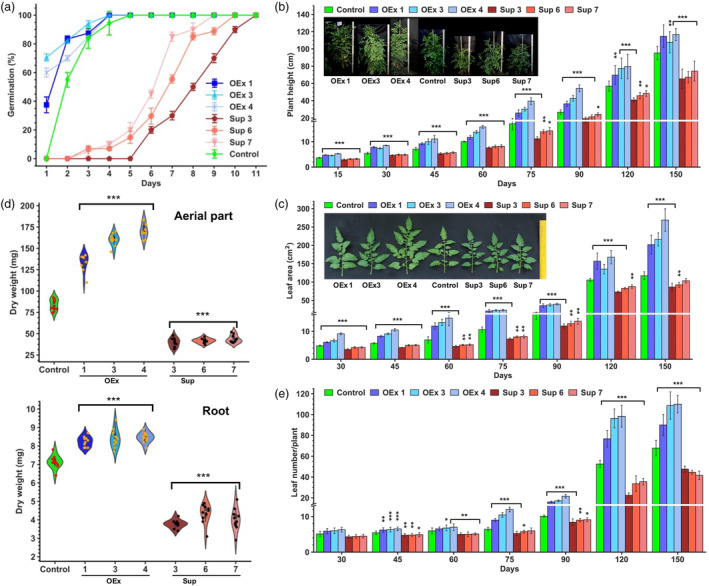
Manipulation of SlERF36 levels alters plant growth and the timing of developmental transitions in tomato. (a) Germination percentage of seeds of control and SlERF36 over‐expression and suppression lines on ½ MS medium. Germination was monitored based on radical emergence. The number of days excludes the two‐day stratification (n = 30). (b) Graphical representation of height of control and transgenic SlERF36 over‐expression and suppression lines during the course of their growth. Measurements were made in nethouse‐grown plants on days 15, 30, 45, 60, 75, 90 120 and 150. Values are averages (± SD) of ten plants per line. The inset shows height of 5‐month‐old plants of control and SlERF36 transgenic lines. (c) Graphical representation of changes in leaf area of control and SlERF36 transgenic lines. Measurements were made in nethouse‐grown plants on days 15, 30, 45, 60, 75, 90 120 and 150 using the 4th leaf from bottom. The values represent the mean (± SD) of leaves of five plants. The inset shows leaves of 5‐month‐old plants of control and SlERF36 transgenic lines. (d) Dry weight of the entire aerial part (inclusive of stem and leaves) and roots of 30‐day‐old plants. The values represent average ± SD of five replicates. (e) Graphical analysis of differences in the leaf number in control and transgenic SlERF36 over‐expression and suppression plants during the course of their development under nethouse conditions. The values are average ± SD of twenty plants per line.
Following seedling establishment, further growth differences were noted in subsequent stages of plant growth in tissues such as roots, leaves and stem between the SlERF36 OEx and suppression lines. All the OEx lines grew faster while the suppression lines grew slower as evident from the 34%–36% increase in root length of 1‐month‐old transgenic SlERF36 OEx plants and a reduction of 12%–16% in suppression lines over control roots (Figure S2a). The differences were even more apparent in aerial tissues with the OEx lines showing an increase in plant height ranging from 25% to 69% at 30 days and 50%–130% at 75–90 days, compared to control (Figure 1b). As the OEx plants attained their final height, the growth increase slowed down, although plants remained taller by 20%–24% than controls even at day 150. In comparison, suppression lines were slow‐growing and stayed shorter than controls by 10%–20% up to 30 days and by 15%–25% thereafter. At harvest, the plants were shorter by 20%–30% and never attained the height of the control. These large reciprocal differences between OEx and suppression lines were indicative of dose‐dependent changes in levels of SlERF36.
Leaf growth was also substantially affected in transgenic lines, with all three suppression lines showing a reduction in leaf area compared to control plants (Figure 1c). The reduction ranged from 12% to 27% up to 45 days and 25%–35% between 60–75 days. Even at harvest, leaves of suppression lines were smaller by 11%–25% compared to control. In contrast, OEx lines showed a much greater change in leaf area with an increase ranging from 45% to 85% by 45 days, 70%–112% by 60 days and 100–150% by 90 days. At 5 months, OEx plants had leaves that were ~two‐fold larger (70%–130% increase) than controls and >2.5‐fold larger than leaves of suppression lines.
A true indication of faster growth is an increase in dry weight of plants. To verify this, the fresh weight and dry weight of roots as well as aerial tissues was estimated in 30‐day‐old plants. All the suppression lines showed a 25%–40% reduction in the root fresh weight and 40%–50% reduction in root dry weight, compared to control (Figure 1d; Figure S2b). A similar change was observed for shoots with a 50%–55% reduction in fresh weight and a 47%–52% reduction in dry weight (Figure 1d; Figure S2b). Strikingly, OEx lines showed an increase of 50%–86% in root fresh weight and 15%–18% in dry weight while shoot fresh weight increased by 13%–21% and dry weight by 60%–100% (compared to control) suggesting a greater resource allocation to aerial parts upon SlERF36 over‐expression.
The effects of SlERF36 expression were not restricted to just organ expansion and growth but also affected leaf bud initiation with a 30%–38% reduction in leaf number in all suppression lines and a 32%–62% increase in OEx lines compared to controls at the final stage (Figure 1e). Thus, OEx lines not only had more leaves but also larger leaves while suppression lines had fewer and smaller leaves. Internodal distance was greater by 1.5–2‐fold in over‐expression lines but reduced by 20%–40% in suppression lines (Figure S2c).
Manipulation of SlERF36 expression alters the timing of flowering, senescence and fruit harvest without affecting yield
The changes in vegetative growth observed in SlERF36‐altered lines had a marked effect on the transition from the vegetative to reproductive phase as observed by the onset of flowering. The first flower buds were observed around 90.6 ± 1.1 days in control but were seen at least 6–7 days earlier at 84.6 ± 1.2 days, 84.4 ± 1 days, and 83.4 ± 0.9 days in the SlERF36 OEx lines OEx1, OEx3 and OEx4, respectively (Figure 2a). In contrast, suppression lines showed a delay of 6–8 days with initiation after 98.2 ± 1, 97.4 ± 0.96 and 96.4 ± 1.3 days in lines Sup3, Sup6 and Sup7, respectively. Thus, flowering time differed by 13–14 days between OEx and suppression lines.
Figure 2.
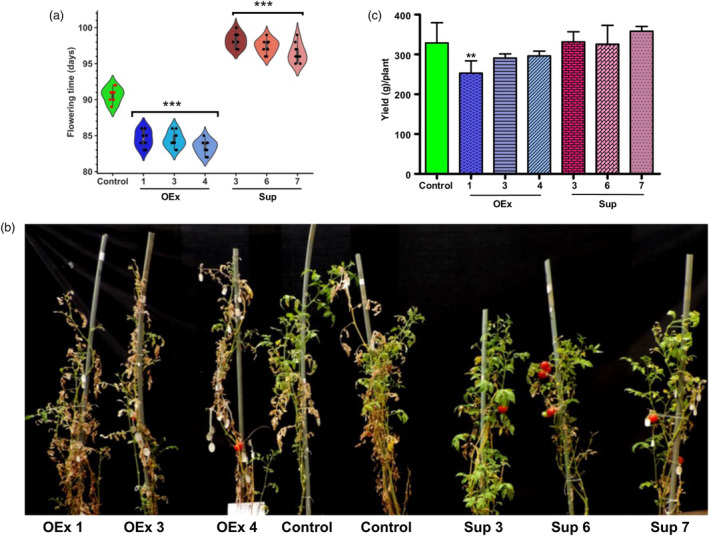
Manipulation of SlERF36 levels alters the timing of flowering, senescence and fruit harvest in tomato. (a) Graphical analysis of differences in the flowering time (time of appearance of the first bud on the plant) in control and transgenic SlERF36 over‐expression and suppression plants grown in the nethouse. The values are average ± SD of twenty plants per line. (b) Representative picture showing progression of whole‐plant senescence in nethouse‐grown plants of control and transgenic SlERF36 lines in the 6th month. (c) Graphical analysis of yield (total fruit weight per plant) in control and transgenic SlERF36 OEx and suppression line plants. The error bars represent SE, * indicates P < 0.05, ** indicates P < 0.01, *** indicates P < 0.001 (Student's t‐test).
Not unexpectedly, the onset of whole‐plant senescence post‐flowering and fruiting was also early in all the SlERF36 over‐expression lines, but delayed in the suppression lines. Senescence, examined as the time when plants had more than 50% senescent leaves (wilted and brown), was at least a week earlier in OEx lines but delayed by about 7–10 days in suppression lines compared to control (Figure 2b). The loss of chlorophyll, an indicator of senescence, was examined in detached leaves after every 12 h using a PAM fluorescence meter and occurred earlier within 1.5–2 days in OEx lines compared to 4 days in control and 5–9 days in suppression lines (Figure S2d).
Finally, manipulation of SlERF36 expression did not affect final yield (Figure 2c). However, the time taken to harvest at least 80% of the total fruits was considerably altered with OEx lines showing early harvest by 20–30 days compared to control while suppression lines showed delayed harvest by ~12–14 days compared to control (Table 1).
Table 1.
Time taken (days) for various transitions (including harvest) in control and SlERF36 transgenic lines under glasshouse/nethouse conditions in different seasons and generations
| Line | Germination (Days) | Flowering time initiation (Days) | Total time for >80% fruit harvest (days) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| GH1 | GH2 | NH1 | NH2 | GH1 | GH2 | NH1 | NH2 | ||
| Control | 5 | 89.8 ± 1.1 | 86.4 ± 1.1 | 90.6 ± 1.1 | 87.9 ± 1.7 | 168–182 | 165–180 | 170–183 | 166–180 |
| OEx 1 | 4 | 83.5 ± 1.2 | 78.7 ± 1.8 | 84.6 ± 1.2 | 80.1 ± 1.7 | 138–152 | 132–150 | 141–153 | 135–150 |
| OEx 2 | 4 | 82.8 ± 1 | 78.1 ± 1 | 84.4 ± 1.1 | 79.2 ± 1.1 | 138–152 | 132–150 | 141–153 | 135–150 |
| OEx 3 | 4 | 81.9 ± 1.1 | 77.5 ± 1.7 | 83.4 ± 1 | 78.9 ± 1.1 | 138–152 | 132–150 | 140–153 | 135–150 |
| Sup 1 | 11 | 97.7 ± 1 | 96.1 ± 1 | 98.2 ± 1.1 | 96.9 ± 1 | 182–194 | 180–192 | 183–195 | 180–192 |
| Sup 2 | 10 | 96.8 ± 1 | 95 ± 1.2 | 97.4 ± 1 | 95.9 ± 1.3 | 182–194 | 180–192 | 183–195 | 180–192 |
| Sup 3 | 9 | 95.9 ± 1.1 | 94.3 ± 1 | 96.4 ± 1.3 | 95 ± 1.1 | 182–194 | 180–192 | 183–195 | 180–192 |
GH1 = Glasshouse (Aug 2017‐Feb 2018), T3 generation, n = 25–30 plants (plants/line).
GH2 = Glasshouse (Dec 2018‐Jun 2019), T4 generation, n = 15–20 plants (plants/line).
NH1 = Net house (Aug 2019‐Feb 2020), T5 generation, n = 10–12 (plants/line).
NH2 = Net house (Oct 2020‐Apr 2021), T6 generation, n = 10–12 plants (plants/line).
These results showed that SlERF36 expression levels influenced whole‐plant growth considerably by altering the timings of different growth transitions and whole‐plant growth. The studies were repeated across different seasons in experiments carried out over a span of 4 years in T3 to T6 generation plants in glasshouse as well as nethouse (Table 1). These showed that the harvest of ~80% of fruits on the plants was completed in a range of 165–183 days in control, 132–153 days in OEx lines and 180–195 days in suppression lines in glasshouse as well as nethouse with slight differences. The studies indicated the robustness of regulation by SlERF36.
SlERF36 ‐mediated changes in growth in stem and leaves are dependent on GA levels as well as GA signalling.
Growth changes associated with developmental transitions are marked by changes in hormone homeostasis with GA being one of the key hormones responsible for many transitions (Castro‐Camba et al., 2022). Although most of the changes in traits observed in SlERF36‐altered lines (germination, height, leaf area, flowering time) are controlled by multiple factors, GA is a common regulatory factor in all these, making it a likely candidate for regulation by SlERF36. To test this, control and transgenic SlERF36 OEx and suppression line plants were grown with regular sprays of either water (as control), 100 μM GA3 or with 100 μM paclobutrazol (a GA biosynthesis inhibitor) at 3‐day intervals for 3 months to check whether differences in plant height and leaf area could be overcome by these. As shown in Figure 3a (Figure S3a), the OEx lines showed a significant increase in height that ranged from 20% to 40% over the control plants from 14 to 28 days under untreated conditions. This increased to 80%–130% from days 42 to 70 and reduced to 25%–30% by day 84. A similar quantum of increase was also observed in the leaf area of OEx lines over the controls at the corresponding stages (Figure 3b; Figure S3b). Suppression lines showed a 25–37% decrease in height compared to control at all stages from 14 to 70 days and a 15%–20% decrease by day 84 under untreated conditions with similar differences in leaf area at corresponding stages. Treatment with 100 μM GA increased the plant height in all the lines. However, the differences in height between control and SlERF36 OEx lines were largely reduced with significant differences of only 5%–10% from days 14 to 28. Thereafter, no differences were observed in heights between OEx and control plants. Although leaf area showed significant differences over the control at the corresponding stages, these too were only in the range of 4%–11%, except at day 14 where the difference was higher. Suppression lines also grew taller following GA treatment. Surprisingly, they continued to remain shorter than controls by 25%–35% from 14 to 56 days and by 10%–20% at days 70 and 84 with similar differences in leaf area at corresponding stages. When the plants were treated with paclobutrazol, there was marked reduction in the heights of all the lines at all stages. The differences between control and OEx lines were much reduced compared to untreated plants and were in the range of 8%–22% in different lines from days 14 to 70. There was nevertheless an increase of 25%–54% in heights in OEx lines at day 84 over the control. An increase of 10%–20% was also seen in leaf area of OEx lines from 14 to 42 days with a much higher increase of 25%–50% at days 70 and 84. In contrast, suppression lines showed a much greater reduction in height compared to controls with the decrease ranging from 21% to 33% from days 14 to 70 and little lower at 84 days. The leaf area of suppression lines also showed a much larger reduction in the range of 25%–37% compared to control at all stages. Thus, treatment with GA as well as paclobutrazol largely reduced differences in growth (height and leaf area) between OEx lines and controls but not as much between suppression lines and controls.
Figure 3.
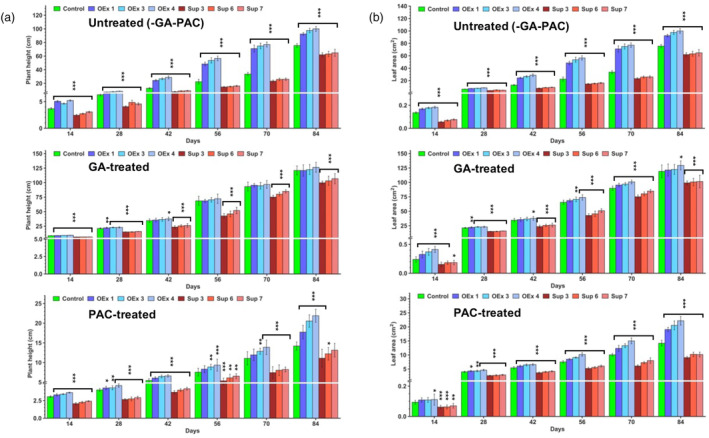
Sensitivity of control and transgenic SlERF36 altered lines to exogenous GA3 and paclobutrazol. Graphical representation of plant height (a) and leaf area (b) in control and transgenic SlERF36 plants upon exogenous application of GA3 and paclobutrazol. Control and transgenic SlERF36 plants were sprayed with either water (untreated) or with 100 μM GA3 or 100 μM paclobutrazol in three independent sets at three‐day intervals for 3 months and measurements for height and leaf area taken at 14, 28, 42, 56, 70 and 84 days post‐germination. Bars represent value (± SD) averaged over 10–15 measurements per line. The error bars represent SE, * indicates P < 0.05, ** indicates P < 0.01, *** indicates P < 0.001 (Student's t‐test).
The results indicated that GA levels as well as GA signalling (or GA‐independent processes) were, in part, responsible for the SlERF36‐mediated growth changes.
Changes in plant growth are associated with SlERF36 ‐mediated control of expression of GA biosynthesis/degradation and signalling
Since GA appeared to be a strong candidate that may influence the changes in SlERF36‐mediated plant growth, the GA pathway (biosynthesis, metabolism and signalling) was studied in greater detail. Real‐time PCR analysis of the GA pathway genes was carried out in SlERF36 over‐expression, suppression and control plants at two developmental stages namely, seed germination (at 3DPI) and in leaves of 2‐month‐old plants, when differences were most prominent between these lines. Clear reciprocal changes were observed in expression of several GA pathway genes between OEx and suppression lines. Transcript levels of the GA biosynthesis genes SlGA20ox1 and SlGA20ox3 increased by 2.5–3‐fold and 4–6‐fold respectively in all the OEx line seeds but were reduced to 50%–55% (SlGA20ox1) and just 20%–35% (SlGA20ox3) in all the suppression line seeds (Figure 4a). Expression of SlGA3ox1 increased 2–3‐fold in OEx lines but was reduced to only 20%–40% in suppression lines. In contrast, the expression profiles of the GA2ox genes (responsible for GA degradation) showed opposite effects with the most significant reduction in transcript levels of SlGA2ox3 and SlGA2ox4. These decreased by 30%–40% in OEx lines and increased 1.5‐fold in suppression lines compared to control. Other GA2oxs such as SlGA2ox1, SlGA2ox2 and SlGA2oxs 5–10 also showed a reduction in OEx lines and an increase in suppression lines although these were not as significant (Figure S4a).
Figure 4.
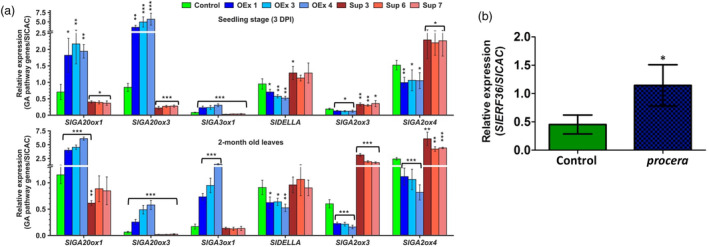
Manipulation of SlERF36 affects the expression of GA pathway genes in different tissues. (a) qRT‐PCR analysis showing relative changes in transcript levels of GA biosynthesis genes (SlGA20ox1, SlGA20ox3 and SlGA3ox1), GA catabolism genes (SlGA2ox3 and SlGA2ox4) and SlDELLA in control and transgenic SlERF36‐altered lines in germinating seeds (3DPI) (upper panel) and in 2‐month‐old leaves (lower panel). The values were normalized against SlCAC and represent means of three biological replicates. The error bars represent SE, * indicates P < 0.05, ** indicates P < 0.01, *** indicates P < 0.001 (Student's t‐test). (b) Real‐time PCR analysis of expression of SlERF36 in leaves of the procera mutant.
The differences in expression of GA pathway genes were more prominent in leaves of 2‐month‐old plants although SlERF36 transcript levels were in the range as seen in seedlings (4–5‐fold higher in OEx lines and 50%–70% reduced in suppression lines). As shown (Figure 4a), transcript levels of the GA biosynthesis genes were up‐regulated by 3.5–5‐fold (SlGA20ox1), 3–7.5‐fold (SlGA20ox3) and 4–8‐fold (SlGA3ox1) in all OEx lines but reduced by 20%–50% (SlGA20ox1), 50%–70% (SlGA20ox3) and ~20% (SlGA3ox1) in all the suppression lines compared to the control. In contrast, transcript levels of SlGA2ox3 and SlGA2ox4 (but not other GA2ox genes; Figure S4b) were reduced by 65%–70% and 55%–70%, respectively, in over‐expression lines and increased 3–5‐fold and 2–2.4‐fold in all the suppression lines.
Besides GA biosynthesis and degradation, transcript levels of the DELLA gene, SlDELLA, involved in GA signalling, also showed a significant 20%–40% reduction in seedlings and 35%–50% reduction in leaves of all the OEx lines but a slight increase in suppression lines. Interestingly, the expression of SlERF36 was about twofold higher in leaves of the SlDELLA mutant, procera (Figure 4b). No differences were observed in transcript levels of the three GID genes in the transgenic lines (data not shown).
Changes in SlERF36 ‐mediated plant growth are associated with changes in GA3 and GA4 levels in stem and leaves
In view of the correlation between the expression of the GA pathway genes and the large‐scale differences in height and leaf area of SlERF36‐altered lines, an estimation of endogenous levels of GA3 and GA4 was carried out in stem and leaves of 60‐day‐old plants of all the SlERF36 transgenic lines and control. A marked change in the basal GA content of 60‐day‐old leaf tissues was observed between controls and transgenic lines (Figure 5a). The levels of GA3 and GA4 in controls (5.5 ± 0.4 and 28.1 ± 0.7 ng g−1 FW, respectively) increased by 21, 93 and 135% for GA3 in OEx lines 1, 3 and 4, respectively and by 22%–35% for GA4 in leaves of all OEx lines. In contrast, leaves of 60‐day‐old suppression lines showed a significant decrease of 31%–47% for GA3 and 60%–66% for GA4.
Figure 5.
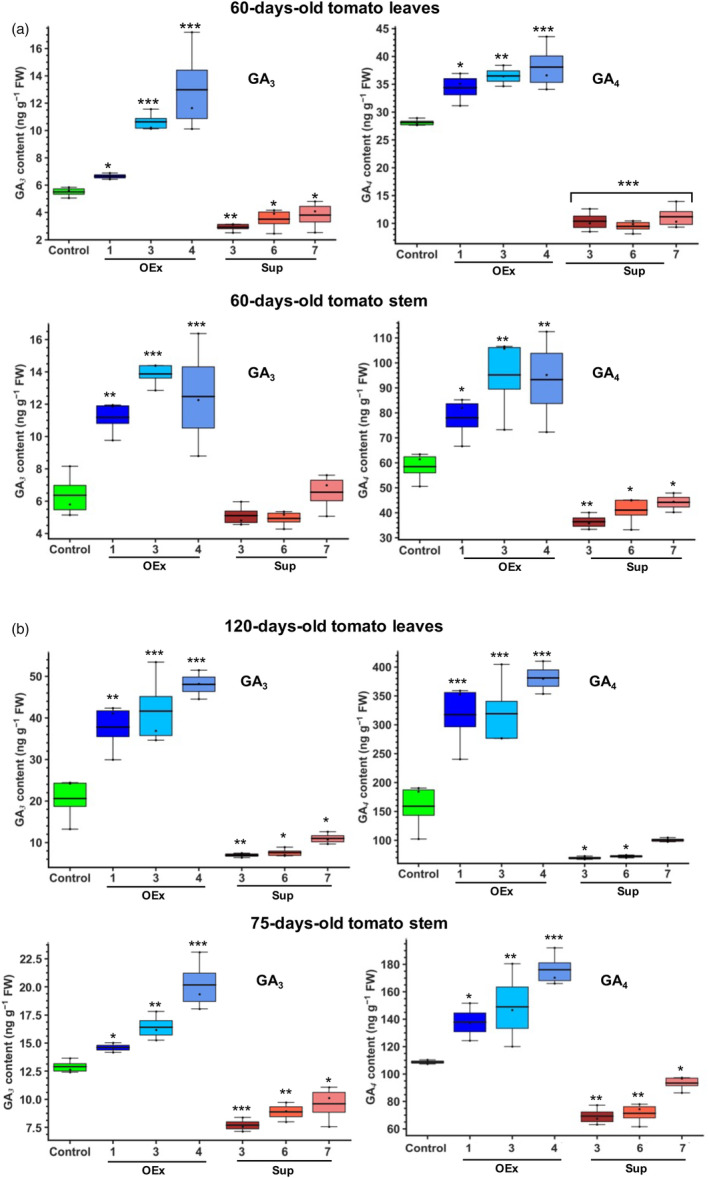
SlERF36 manipulation alters GA levels in stem and leaves. Box plots of liquid chromatography‐mass spectrometry estimation of GA3 and GA4 in (a) leaves and stems of 60‐day‐old plants of control and transgenic SlERF36 over‐expression and suppression lines and (b) leaves of 120‐day‐old plants and stem of 75‐day‐old plants. The values are average (± SD) of three replicates. The error bars represent SE, * indicates P < 0.05, ** indicates P < 0.01, *** indicates P < 0.001 (Student's t‐test).
The levels for GA3 and GA4 in corresponding stems of 60‐day‐old control plants were 6.3 ± 1.6 and 58 ± 7 ng g−1 FW, respectively. These increased by 75%–118% for GA3 and by 33%–62% for GA4 in stems of the OEx lines. In contrast, stems of 60‐day‐old suppression lines showed a slight decrease of 20%–22% for lines 3 and 6 for GA3 that was not significant but showed a significant decrease of 24%–38% for GA4 in all the lines.
An estimation of GA levels in leaves of 120‐day‐old plants and stems of 75‐day‐old plants (Figure 5b) revealed even more prominent differences. GA3 levels in control and transgenic OEx lines increased almost 3.5‐fold in leaves of 120‐day‐old plants over 60‐day‐old plants while GA4 levels shot up ~sixfold in control but 9–10‐fold in transgenic OEx lines. Compared to 120‐day‐old control plants, these were associated with ~twofold higher levels of GA3 and 2.5‐fold higher levels of GA4 in the OEx lines but a reduction by 50%–70% (GA3) and >70% (GA4) in suppression lines. GA levels also increased in 75‐day‐old stem by about 1.8‐fold for GA3 and GA4 over the 60‐day stem while they were significantly reduced in suppression lines. These were associated with an increase of 13%–56% in OEx lines (GA3) and 27%–62% (GA4) but a reduction by 26%–40% for GA3 and 14%–36% for GA4 for suppression lines, compared to controls. The results confirmed that GA levels were partly to substantially responsible for changes in the growth phenotypes of leaves and stem of SlERF36 transgenic lines.
SlERF36 functions as a repressor and binds the promoters of SlGA2ox3 , SlGA2ox4 as well as SlDELLA
EAR‐motif proteins are well known to function as repressors of transcription (Kagale and Rozwadowski, 2011; Ohta et al., 2001). In order to confirm whether SlERF36 functioned as a repressor, a protoplast transfection assay was performed with 35S‐GAL4‐DBD and 35S‐GAL4DBD‐SlERF36 as effector constructs, and GAL4(4X)‐CaMV35S‐GUS as reporter construct as described (Singh et al., 2019; Wang et al., 2007). The effector construct expressing GAL4DBD under the CaMV35S promoter could drive GUS activity from the reporter in absence of SlERF36. However, in presence of SlERF36 (under the control of the CaMV35S double promoter), there was 60% reduction in GUS activity (Figure 6a). Having established the ability of SlERF36 to function as a repressor, and given the observations of strong suppression of SlGA2ox3, SlGA2ox4 as well as SlDELLA in SlERF36 OEx lines by real‐time analysis (Figure 4a), yeast one‐hybrid assays were carried out to study whether SlERF36 could directly bind their promoters to regulate them. A region of the promoters comprising −1931 bp (SlGA2ox3), −2498 bp (SlGA2ox4) and −1983 bp (SlDELLA), upstream of the initiation codons, was chosen for study and showed clear interactions with SlERF36 (Figure 6b). These interactions were further confirmed in vivo using luciferase assays. As shown (Figure 6c), expression of luciferase driven by the promoters of SlGA2ox3, SlGA2ox4 and SlDELLA was observed as fluorescence in tobacco leaves agro‐infiltrated with constructs lacking the SlERF36 construct. However, co‐expression of these promoters with SlERF36 strongly reduced promoter‐driven luciferase activity and fluorescence to low/undetectable levels indicating the suppression of SlGA2ox3, SlGA2ox4 and SlDELLA promoters by SlERF36. Further confirmation of SlERF36 function was obtained at least for SlGA2ox3 through EMSA studies wherein binding of the SlERF36‐His tagged protein to the GCC‐box cis element at position −1343 in the SlGA2ox3 promoter led to a shift in the band that was competed out by 5x unlabelled probe (Figure 6d). No binding was observed when the cis element was mutated. Collectively, the studies highlighted how regulation of multiple growth transitions could be effectively controlled by SlERF36 at least partly through manipulation of GA levels and responses through suppression of SlGA2ox3, SlGA2ox4 and SlDELLA.
Figure 6.
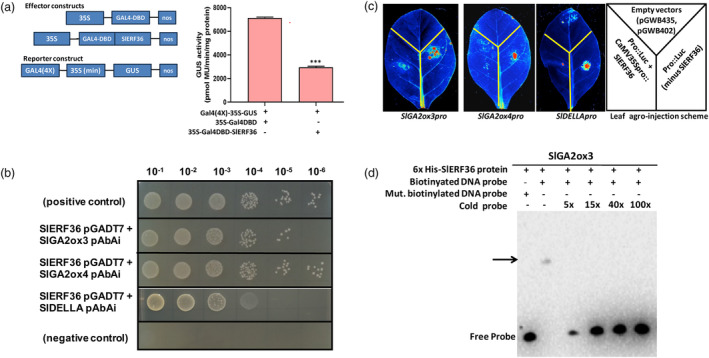
SlERF36 functions as a transcriptional repressor and physically interacts with the SlGA2ox3/SlGA2ox4/SlDELLA promoters. (a) SlERF36 functions as a repressor of GAL4 in protoplast transfection assays. The effectors GAL4DBD and GAL4DBD‐SlERF36 under the CaMV35 double promoter and the GAL4(4X)::GUS reporter constructs were co‐transfected into protoplasts extracted from 3 to 4 week‐old Arabidopsis Col‐0 plants. GUS activity was studied after 16 h of transfection in dark. Graph represents the mean ± standard error of three replicates. (b) Yeast one‐hybrid studies showing interactions between SlERF36 and the promoters of the GA catabolism genes P SlGA2ox3 (−1931 bp, second row), P SlGA2ox4 (−2498 bp, third row) and the GA signalling inhibitor P SlDELLA (−1983 bp, fourth row) in different serial dilutions. The positive and negative controls are present in the first and last rows, respectively. The minimal inhibitory concentration of the aureobasidin‐A (AbA) antibiotic for selection of positive bait‐prey interactions was optimized to 100 ng/mL and selection performed on SD/AbA/‐Ura agar plates. (c) Luciferase assays depicting suppression of the above promoters of P SlGA2ox3, P SlGA2ox4 and P SlDELLA by SlERF36 (driven by the CaMV35S promoter) in tobacco leaves. The scheme showing the agro‐injection of constructs in different regions of the leaves (empty vectors at top; promoter‐LUC constructs without SlERF36 on the right; and promoter‐LUC constructs with SlERF36 on the left) is provided in the extreme right panel. The fluorescence in the right part of the leaf indicates basal promoter activity in absence of SlERF36. (d) Electrophoretic mobility shift assays showing binding of SlERF36 to the biotin‐labelled primers containing the GCC box cis element at −1343 nt (upstream of the initiation codon) in the promoter of SlGA2ox3. Biotin‐labelled primer probes (double‐stranded) were electrophoresed with SlERF36 protein on a 6% native polyacrylamide gel as described in methods. Interactions were checked with biotinylated mutated probe and by competition with excess cold probe. The arrow shows the band of shifted probe bound to SlERF36.
Discussion
GA is one of the primary hormones associated with multiple growth cycles that mark transitions in a plant and can impact important agricultural traits such as maturity, harvest time and crop duration (Castro‐Camba et al., 2022; Hedden, 2003). The GA pathway needs to be tightly controlled as misregulation of GA levels/signalling can markedly affect plant growth and fitness. We have identified the tomato EAR‐motif repressor, SlERF36, as a global regulator of the GA pathway that alters GA levels and signalling across tissues, affecting multiple transitions and plant growth duration.
Manipulation of SlERF36 levels influences the timing of developmental events such that higher SlERF36 expression accelerates germination (Figure 1a), increases the speed of organ growth (root length, leaf area, internodal distance, height and dry weight) (Figure 1b–d), accelerates flowering time (in days) (Figure 2a) and whole‐plant senescence (Figure 2b). Its suppression has opposite effects that retard growth and delay transitions. These changes lead to a 6–8‐day difference in germination, 15‐day difference in flowering time and a difference of about 30–42 days in completion of the fruit harvest between SlERF36 OEx and suppression lines. The results are indicative of dose‐dependent effects of SlERF36 that alter the speed of plant growth.
Although each of these traits is influenced by a combination of hormones and different tissue‐specific regulators (Blazquez et al., 1998; Castro‐Camba et al., 2022; Finch‐Savage and Leubner‐Metzger, 2006; Fu and Harberd, 2003), GA plays a decisive role in determining how these changes progress and is strongly and reciprocally altered in SlERF36 OEx and suppression lines across tissues like germinating seeds, leaves and stem (Figures 4 and 5). SlERF36 indirectly activates transcription of the GA biosynthesis genes SlGA20ox1/3 and SlGA3ox1 (Figure 4a) and simultaneously reduces GA degradation by directly binding to the promoters of GA2 oxidase genes, SlGA2ox3 and SlGA2ox4, and the GA response inhibitor SlDELLA, reducing their transcription in seeds as well as leaves (Figures 4a and 6A–D). Its action increases basal GA3 and GA4 levels in stem and leaves throughout development with 3–5‐fold higher levels in SlERF36 OEx lines and 30%–70% reduction in suppression lines (Figure 5a,b). Although GA levels change across tissues, the changes are greater for GA4 compared to GA3 in leaves as well as stems indicating that the GA2ox3/4 enzymes may preferentially target GA4 than GA3. The greater differences in GA4 levels in stems suggest that it may contribute more to differences in height. The changes in GA levels and signalling seem to have a cascading effect on overall plant growth and transitions leading to marked changes in flowering time, fruit harvest and senescence. Tomato organ growth is known to be affected by changes in GA biosynthesis or degradation (Olimpieri et al., 2011; Schrager‐Lavelle et al., 2019; Shohat et al., 2021b; Yanai et al., 2011) and by GA signalling (Jasinski et al., 2008; Liu et al., 2016; Livne et al., 2015; Nir et al., 2017; Ramon et al., 2021). A reduction in GA levels by suppression of GA20ox1 reduces internodal distance and height in tomato plants and affects leaf morphology and floral tissues (Olimpieri et al., 2011; Shohat et al., 2021a) while over‐expression of CcGA20ox1 in tomato affects stem length and fruit set (García‐Hurtado et al., 2012). Alterations in GA levels are also caused by inactivation of SlGA2ox7 which affects GA2 oxidase activity leading to increased internode length in the tomato internode elongated‐1 mutant (Schrager‐Lavelle et al., 2019). The lanceolate mutation la in tomato and its semi‐dominant form La‐2 (which positively affects GA responses) affect leaf lamina growth by changing GA20ox1 and GA2ox4 expression in the margins, thereby affecting leaf complexity (Yanai et al., 2011). Mutations in the procera mutant, affecting the only DELLA gene in tomato, increase GA responses leading to elongated, thin stems and simpler leaves (Jasinski et al., 2008; Livne et al., 2015) while a gain‐of‐function mutant lacking DELLA domain is GA‐insensitive and severely reduces growth (Nir et al., 2017). SlDELLA expression also regulates stomatal responses and reduces stomatal opening in drought (Nir et al., 2017; Shohat et al., 2020, 2021a). SlERF36 seems to simultaneously control many of the above genes, and the GA pathway as a whole, through its action. It also affects growth independently of GA levels since treatment with exogenous GA or paclobutrazol (Figure 3a,b) does not completely overcome the growth phenotypes in transgenic lines. These effects, which are more prominent in SlERF36 suppression lines under GA‐limiting conditions (paclobutrazol treatment), could partly be mediated by regulating GA signalling through increased SlDELLA expression and its stability (due to high GA oxidase activity) but also through GA‐independent regulators (as evident from an ongoing transcriptomic study). The changes in SlDELLA stability in SlERF36‐manipulated lines could selectively affect its interactions with GID receptors in different tissues thereby regulating growth differentially, as previously shown (Illouz‐Eliaz et al., 2019). This aspect, along with GA pathway‐independent regulation, is currently under study. SlERF36 effects on germination and organ growth/expansion can be explained by SlERF36‐mediated changes in the GA pathway and an increase in basal GA levels and responses. Although flowering has also been shown to be promoted by GA in Arabidopsis (Blazquez et al., 1998; Yamaguchi et al., 2014) and some other plants, higher GA levels or higher GA responses (as in the procera mutant) have actually been shown to delay flowering in tomato (Carrera et al., 2012; García‐Hurtado et al., 2012). Interestingly, while SlERF36 over‐expression increases leaf number (an indication of delay in flowering, Figure 1e), flowering time is actually accelerated by ~7 days compared to control and delayed in suppression lines (Figure 2a). This is unlike the procera mutant, where leaf number is higher and the time to anthesis of the first flower is also delayed (Carrera et al., 2012). Early flowering was also observed upon ectopic expression of SlERF36 in tobacco and Arabidopsis (Upadhyay et al., 2013, 2014). The disparity between increased leaf number and reduced flowering time in SlERF36‐altered lines could stem from faster overall growth and morphogenetic development in OEx lines and growth retardation in suppression lines. The observations suggest that SlERF36 effects on flowering may be more complex and dependent on additional targets that differentially affect flowering time and plant growth, partly through GA‐independent effects. GA application is also known to delay senescence (Yu et al., 2009) although its effects on senescence are conflicting since DELLA function can also delay senescence (Zhang et al., 2018). It is likely that whole‐plant senescence that is activated in annuals post‐flowering, may be regulated differently by GA/DELLA than that of individual organs alone (although SlERF36 OEx leaf discs also show early senescence symptoms). Conversely, it may be regulated by SlERF36 differently through GA‐independent targets. A model explaining the possible mode of action of SlERF36 on whole‐plant development through GA‐dependent and independent pathways is shown (Figure 7). That SlERF36 action may also involve additional GA‐independent targets is implied through its growth phenotypes which unlike procera (which is tall and thin) cause plants to be sturdy because of an increase in stem girth (data not shown). The highly expanded leaves, that develop early on, increase the canopy area and allow greater capture of light and could aid in photo‐assimilation of carbon dioxide and would explain the faster increase in fresh and dry weight in SlERF36 OEx lines (Figure 1d; Figure S2b). A comprehensive analysis of SlERF36 targets using tagged versions would shed more light on the complexities of SlERF36 regulation on other aspects besides GA homeostasis. Regardless of the mechanismsgoverning flowering and senescence inSlERF36 altered lines, the overall effect of faster growth and earliness in flowering, harvest and senescence in OEx lines is an attribute of significance for farmers.
Figure 7.
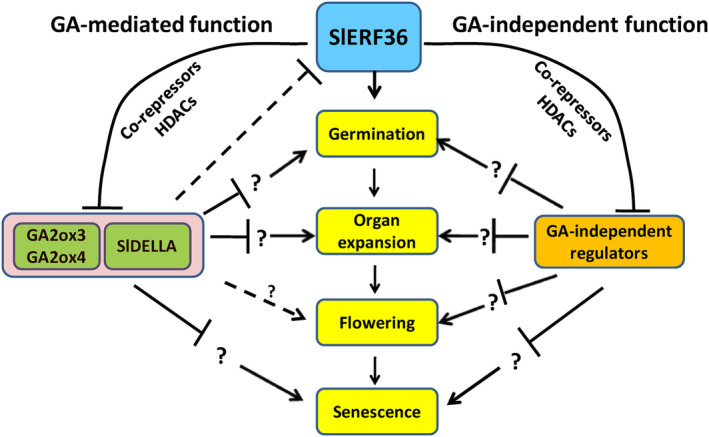
Model showing the function of SlERF36 in regulating developmental transitions through its action on the GA pathway and through putative GA‐independent regulators. SlERF36, through co‐repressors and HDACs, suppresses the GA2 oxidases, SlGA2ox3 and SlGA2ox4 and the GA action inhibitor, SlDELLA, to increase the GA pathway output which in turn activates intermediate components (some known, some yet to be identified) that activate various transitions. It also regulates these through suppression of GA‐independent regulators that suppress the transitions through other intermediaries.
As a member of the small EAR‐motif sub‐group of repressors within the large AP2/ERF family, SlERF36 is likely to function through recruitment of co‐repressors like SIN3 and TOPLESS, like other EAR‐motif repressors (Deng et al., 2022; Song et al., 2005). EAR‐motif repressors, although accounting for a small subgroup of the large AP2‐ERF domain family, seem to have major roles in modulating hormone responses during development and stress adaptation. AtERF7, a homologue of SlERF36, reduces ABA responses in Arabidopsis in stomatal closure leading to water loss under drought while its suppression increases ABA sensitivity in germination and stomatal closure (Song et al., 2005). AtERF4 negatively regulated JA responses to Fusarium oxysporum (McGrath et al., 2005) and also reduced sensitivity to ethylene and ABA (Yang et al., 2005) while OsERF3 increased JA responses upon wounding and ethylene responses in drought (Lu et al., 2011; Zhang et al., 2013). AtERF12 controls the transition from dormancy to germination by regulating ethylene responses (Li et al., 2019b) while SlERF12 in tomato negatively regulates the transition to fruit ripening through suppression of ethylene biosynthesis genes ACS2 and ACS4 (Deng et al., 2022). Although SlERF36 governs GA homeostasis and all the major transitions across development, a simultaneous effect on other hormones like ABA (akin to its Arabidopsis homologue AtERF7) cannot be ruled out considering that ABA and GA often function antagonistically in developmental processes (Liu and Hou, 2018).
Finally, controlling transitions and shortening plant duration in the field is important in agriculture as early maturity and early harvest can reduce water and nutrient requirement in fields, reduce exposure to adverse conditions (harsh temperatures, excess rains, biotic factors) and promote faster crop rotation. SlERF36 expression reduces plant duration substantially from 6 to 5 months by accelerating growth without reducing yield (Figures 1 and 2), making it an important candidate for smart agriculture. Further, studies in our lab show that it is functionally conserved in Arabidopsis, potentially extending its usefulness across families. Manipulation of this gene provides an avenue for tailoring plants in different families for faster growth and a shorter life cycle without yield loss.
Experimental procedures
Tomato transformation, growth conditions and phenotypic analysis of transgenic plants
All analyses were carried out in the background of the tomato variety Arka Vikas in homozygous progeny (T3‐T6 generations) of transgenic SlERF36 over‐expression and suppression lines (three independent lines each). Plants were grown in the glasshouse and in the field (nethouse) under natural conditions of light and temperature from August to March/April in four different years (2018–2021).
The SlERF36 ORF was cloned in pBI121 (Upadhyay et al., 2013, 2014) in sense and anti‐sense orientations under the CaMV35S promoter at BamHI and introduced in tomato through Agrobacterium‐mediated transformation (Madhulatha et al., 2007). Ten independent lines were generated of which three lines over‐expressing SlERF36, namely OEx1, OEx3 and OEx4 with threefold, fourfold and 4.5‐fold higher SlERF36 transcript levels, respectively, and three suppression lines, Sup3, Sup6 and Sup7 with SlERF36 transcript levels down to 30%, 35% and 40% of control, respectively, were chosen for study (Figure S1c). Homozygous plants of these lines in T3 generation were used for all subsequent studies on phenotyping, gene expression and hormone estimation as described below:
Germination: Seeds were grown on ½ MS and germination recorded as radical emergence after two days of darkness at 25°C (50 seeds/line). Experiments were replicated at least thrice. Differences due to seed germination were excluded when collecting data on subsequent plant growth parameters.
Plant height, leaf area and leaf number
These were measured at 15, 30, 45, 60, 75, 90, 120 and 150 days after seedling emergence. The height was measured from the base to the top of each plant, and the leaf area (4th leaf from bottom) was measured using a graph chart. For root length, 1‐month‐old plants were used.
Biomass
Roots and aerial parts (fresh/dry weight) of 15‐day and 30‐day‐old plants (10 plants each) were cleaned and separately measured. The weight of the entire aerial part was taken as the shoot weight. Dry weights were measured after drying in an oven (48 h, 70°C).
Flowering time
Flowering time (monitored on 20 plants) was measured as the time taken for the first bud to appear on the plant. The time for fruit harvest of at least 80% of the total fruits was measured for comparing earliness in harvest.
Whole plant and leaf senescence
The onset of whole‐plant senescence was considered when more than 50% of the leaves on the plant had wilted and turned yellow/brown after fruiting. The extent of leaf senescence was studied by chlorophyll fluorescence imaging of leaf discs of control and SlERF36 transgenic leaves using an Imaging‐PAM, M‐series Chlorophyll Fluorometer (Walz Effeltrich, Germany) over 4–5 days. The onset of senescence, indicated by damage to photosystem II, is seen as a change in fluorescence and decrease in Fv/Fm values with a colour change from blue to green‐yellow‐red and black (Singh et al., 2013).
GA and paclobutrazol (PAC) treatment
Control and transgenic seeds (30 seeds each) were treated separately with water, 10 μM GA3 and 10 μM paclobutrazol (Sigma‐Aldrich). After radical emergence, the seedlings were transferred to soilrite and sprayed separately with either water, 100 μM GA3 or 100 μM PAC every third day. Height and leaf areas for 10–15 plants from each set (watered and GA/paclobutrazol‐treated sets) were measured for 3 months at seven‐day intervals from the day of radical emergence.
RNA isolation and real‐time PCR analysis
Transcriptional analysis of various GA pathway genes was carried out in germinating tomato seedlings and in 2‐month‐old leaves. Seedlings were sampled on the third day post‐imbibition (DPI), excluding the first two days of dark period.
RNA was isolated as described (Asif et al., 2000) and cDNA prepared. Real‐time PCR was performed using SYBR Select (Applied Biosystems Inc, USA) on an ABI StepOnePlus PCR machine using gene‐specific primers (Table S1). Reactions were run in technical triplicates for three biological samples, and the data analysed were the mean of biological triplicates normalized against the validated reference control gene, SlCAC (Expósito‐Rodríguez et al., 2008) using the 2−∆CT method to obtain the fold change in expression (Livak and Schmittgen, 2001).
Protoplast isolation and transfection for SlERF36 repressor assay
Preparation of effector and reporter constructs
The effector (GAL4‐DBD and GAL4‐DBD‐SlERF36) and reporter constructs [GAL4(4X)::GUS] were prepared as described previously (Singh et al., 2019; Wang et al., 2007) in the pUC19 vector background under the control of the double CaMV35S promoter at Nde1 and Sac1 restriction sites with the full‐length SlERF36 ORF cloned in‐frame with an N‐terminal GAL4 DNA binding domain.
Protoplast Isolation
Arabidopsis protoplasts were isolated largely as described by Abel and Theologis (1994) but without vacuum infiltration. Following incubation (22–24°C, 6–8 h in dark with gentle swirling), the protoplasts were filtered using nylon filter (70 μm) and centrifuged at 100 × g for 5 min. The pellet was washed once with pre‐chilled W5 buffer and resuspended in 20 mL of the same (Abel and Theologis, 1994). Following a 30 min incubation on ice, the protoplasts were centrifuged at 300 × g for 5 min and resuspended in pre‐chilled MaMg solution (0.4 M mannitol, 15 mM MgCl2, and 4 mM MES, pH 5.7). Transfection was carried out as described (Abel and Theologis, 1994) using 100 μL protoplasts (2 × 104 cells) mixed with 15 μg plasmid (effector and reporter constructs) followed by addition of 110 μL of 40% PEG solution (containing 0.2 M mannitol and 0.1 M CaCl2). After incubation at room temperature for 30 min, the reaction mixture was diluted slowly with 0.44 mL of W5 solution, centrifuged at 300 × g for 3 min, and the protoplast pellet was resuspended in 1 mL W5 solution and incubated in dark for 16–18 h at room temperature. GUS activity was assayed as described by Jefferson et al. (1987) on a microtitre plate.
Arabidopsis protoplasts were isolated largely as described by Abel and Theologis (1994) but without vacuum infiltration. Following incubation (22–24°C, 6–8 h in dark with gentle swirling), the protoplasts were filtered using nylon filter (70 μm) and centrifuged at 100 × g for 5 min. The pellet was washed once with pre‐chilled W5 buffer and resuspended in 20 mL of the same (Abel and Theologis, 1994). Following a 30 min incubation on ice, the protoplasts were centrifuged at 300 × g for 5 min and resuspended in pre‐chilled MaMg solution (0.4 M mannitol, 15 mM MgCl2, and 4 mM MES, pH 5.7). Transfection was carried out as described (Abel and Theologis, 1994) using 100 μL protoplasts (2 × 104 cells) mixed with 15 μg plasmid (effector and reporter constructs) followed by addition of 110 μL of 40% PEG solution (containing 0.2 M mannitol and 0.1 M CaCl2). After incubation at room temperature for 30 min, the reaction mixture was diluted slowly with 0.44 mL of W5 solution, centrifuged at 300 × g for 3 min, and the protoplast pellet was resuspended in 1 mL W5 solution and incubated in dark for 16–18 h at room temperature. GUS activity was assayed as described by Jefferson et al. (1987) on a microtitre plate.
Yeast one‐hybrid and luciferase assays
The yeast one‐hybrid protein‐DNA interactions were performed in Y1H gold cells essentially as described in the manufacturer's protocol (Takara, Japan). Three bait sequences representing promoters of SlGA2ox3 (−1931 bp), SlGA2ox4 (−2498 bp) and SlDELLA (−1983 bp) were cloned in the pAbAi vector, integrated in Y1H gold yeast genome and selected on SD/‐uracil agar plates. A minimum inhibitory concentration of aureobasidin A required to inhibit growth of cells in absence of prey (100 ng/mL AbA) was established. For the prey construct, the open reading frame of SlERF36 (666 bp) driven by the constitutive ADH1 promoter was cloned in pGADT7‐AD vector at BamHI and selected in yeast on SD/−leu plates. Following mating, the yeast recombinants, where both bait and prey interacted, were selected by their ability to grow on SD/−leu and the inhibitory concentration of AbA (100 ng/mL) at 30°C for 3–5 days on different dilutions.
The above promoters were also used to drive luciferase expression in tobacco leaves by cloning in the Gateway cloning vector pGWB435 (Invitrogen). The coding sequence of SlERF36 was separately cloned in pGWB402 under the CaMV35S promoter to generate effectors. Empty vectors (pGWB435, pGWB402) and Pro::Luc reporters (with individual promoters) were used as negative and positive controls, respectively. The constructs were introduced in A. tumefaciens (strain GV3101), assays performed by agro‐infection in N. tabacum leaves and visualized by the ability of SlERF36 (co‐infiltrated with Pro::Luc vectors) to suppress fluorescence as described (Zhou et al., 2018). For each promoter, interactions were performed in a single leaf divided into three parts. Experiments were repeated at least thrice in different leaves.
Electrophoretic mobility shift assay (EMSA)
The full‐length coding region of SlERF36 was cloned into pET28a and expressed in Rosetta (DE‐3) strain of E. coli. The His‐tagged protein was induced at 37°C for 16 h. with 1 mM IPTG and purified on Ni‐NTA agarose.
For EMSA, the probes covering the GCC box in the promoters of SlGA2ox3 (−1343 bp), SlGA2ox4 (−1366 bp) and SlDELLA (−1166 bp) were labelled with biotin using the 3՛ end biotinylation kit (Thermo Fisher) and double‐stranded oligonucleotide prepared. Unlabelled probe was used for competitive binding. In the mutated version of the probe, ‘GCCGCC’ was changed to ‘AAAAA’ and biotinylated. The probe (15 μg) and purified fusion protein were incubated with 10x binding buffer containing 10 mM MgCl2, 1 μg/μl poly(dI.dC), 1% NP‐40, 50% glycerol for 30 min at room temperature. The reaction product was resolved on a 6% native polyacrylamide gel, transferred to positively charged nylon membrane (Bright star‐plus, Thermo Fisher Scientific) and the membrane visualized on a Chemidoc system (Bio‐Rad).
GA estimation
GA estimation was performed with fresh leaf tissue as described (Hou et al., 2008) with slight modifications. Crushed samples (1 g) were extracted by ultrasonication in 90% methanol (1 mL) on ice bath (with 30 s on/off cycles for 3 min). After centrifugation (10 000 × g , 15 min at 4°C), the supernatant was collected and pellet re‐extracted with 0.5 mL extraction solvent thrice. The supernatants were combined, dried under a nitrogen stream and re‐dissolved in 1 mL methanol followed by centrifugation (10 000 × g , 5 min at 4°C) and filtration through a 0.22 μm PTFE filter. Samples (5 μL) were analysed against standards of GA (Sigma‐Aldrich, Bengaluru, India) using liquid chromatography‐mass spectrometry (SCIEX Exion LC 2.0 system – SCIEX Triple‐TOF 5600 + system). LC separation was achieved using the SCIEX Exion LC 2.0 system (Framingham, MA, USA) and a Phenomenex Kinetex 1.8 μm, EVO C18 100 Å LC column 100 × 2.1 mm (Torrance, California, USA). A 4 min isocratic flow of water with 0.1% formic acid (5%) and methanol with 0.1% formic acid (95%) was used, with a flow rate of 0.5 mL/min. 5 μL of the sample was injected into the LC system. The SCIEX Triple‐TOF 5600 + system (Framingham, MA, USA) was operated with the Turbo V ion source using an electrospray ionization (ESI) probe. MRM‐HR transitions were monitored for GA3 and GA4 in negative mode by observing the product of 345.0 Da and 331.0 Da, respectively. The source parameter for mass analysis was GS1 (55.0 psi), GS2 (45.0 psi), CUR (35.0 psi), TEM (500°C), ISVF (4500 V), TOF Mass range (50–500 Da) and accumulation time (0.100 s). The compound parameter for GA3 and GA4 was −30 and − 40 declustering potential, −35 was the collision energy with 2.0 collision energy spread. Data acquisition was performed using Analyst TF 1.8.1 software.
Data processing of mass spectrometry acquired data was performed using MultiQuant 3.0.3 software in which calibration curves and peak integration statistics were generated.
Statistical analysis
Statistical analysis was carried out using data from different biological replicates. Significant differences between control and transgenic SlERF36 over‐expression and suppression lines were analysed using Student's t‐test with the GraphPad Prism software where * indicates P < 0.05, ** indicates P < 0.01, and *** indicates P < 0.001.
Author contributions
APS conceived the idea; RG performed the genetic manipulations and real‐time expression analysis; RG, HM performed the phenotypic analysis and GA/paclobutrazol treatments; RG/UC/HM performed the yeast‐one‐hybrid and luciferase assays while HM performed the repression studies; HM/PD performed the EMSA studies; RG/UC/HM prepared samples for GA estimation; RST/NGA performed the GA estimations, VAS analysed the data; RG, VAS and APS analysed all the results; RG, APS wrote the paper.
CSIR‐NBRI manuscript number: CSIR‐NBRI_MS/2023/04/12.
Competing interests
The authors have no competing interests.
Supporting information
Table S1 List of oligonucleotides used in the study.
Figure S1 SlERF36 expression in different tissues and in different OEx and suppression lines.
Figure S2 SlERF36 manipulation regulates various growth processes.
Figure S3 Plant phenotypes of control and transgenic SlERF36 lines upon exogenous treatment with GA3 and paclobutrazol.
Figure S4 qRT‐PCR analysis showing relative changes in transcript levels of GA2ox genes.
Acknowledgements
The authors thank Prof. David Weiss (Hebrew University of Jerusalem, Rehovot, Israel) for the kind gift of procera mutant seeds and Dr Shucai Wang (Northeast Normal University, Changchum China) for the effector and reporter constructs for the repressor assay. The authors thank Mr. Ram Awadh for maintaining plants in the glasshouse and nethouse. The authors are grateful to Mr. Shiv Narayan and Dr. PA Shirke (Dept of Plant Physiology, CSIR‐NBRI) for help with the PAM imaging system for leaf disc senescence studies. RG, HM and UC were supported by Senior Research fellowships from CSIR, India and from SERB, while PD was supported by CSIR funds. Financial support for the work was provided to APS by SERB (Science and Engineering Research Board), Department of Science and Technology, Govt of India, under the project EMR/2016/007736 (GAP3455).
References
- Abel, S. and Theologis, A. (1994) Transient transformation of Arabidopsis leaf protoplasts: a versatile experimental system to study gene expression. Plant J. 5, 421–427. [DOI] [PubMed] [Google Scholar]
- Achard, P. , Cheng, H. , De Grauwe, L. , Decat, J. , Schoutteten, H. , Moritz, T. , Van Der Straeten, D. et al. (2006) Integration of plant responses to environmentally activated phytohormonal signals. Science 311, 91–94. [DOI] [PubMed] [Google Scholar]
- Asif, M.H. , Dhawan, P. and Nath, P. (2000) A simple procedure for the isolation of high quality RNA from ripening banana fruit. Plant Mol. Biol. Rep. 18, 109–115. [Google Scholar]
- Bauerle, W.L. (2022) Gibberellin A3 induced flowering intensification in Humulus lupulus L.: Synchronizing vegetative phase change and photoperiod induction. Sci. Hortic., 302, 111183. [Google Scholar]
- Blazquez, M.A. , Green, R. , Nilsson, O. , Sussman, M.R. and Weigel, D. (1998) Gibberellins promote flowering of arabidopsis by activating the LEAFY promoter. Plant Cell 10, 791–800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carrera, E. , Ruiz‐Rivero, O. , Peres, L.E.P. , Atarés, A. and García‐Martínez, J.L. (2012) Characterization of the procera tomato mutant shows novel functions of the SlDELLA protein in the control of flower morphology, cell division and expansion, and the auxin‐signaling pathway during fruit‐set and development. Plant Physiol. 160, 1581–1596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castro‐Camba, R. , Sánchez, C. , Vidal, N. and Vielba, J.M. (2022) Plant development and crop yield: The role of gibberellins. Plan. Theory 11, 2650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen, S. , Wang, X. , Zhang, L. , Lin, S. , Liu, D. , Wang, Q. , Cai, S. et al. (2016) Identification and characterization of tomato gibberellin 2‐oxidases (GA2oxs) and effects of fruit‐specific SlGA2ox1 overexpression on fruit and seed growth and development. Hortic. Res. 3, 16059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen, P. , Yang, R. , Bartels, D. , Dong, T. and Duan, H. (2022) Roles of abscisic acid and gibberellins in stem/root tuber development. Int. J. Mol. Sci. 23, 4955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colebrook, E.H. , Thomas, S.G. , Phillips, A.L. and Hedden, P. (2014) The role of gibberellin signaling in plant responses to abiotic stress. J. Exp. Biol. 217, 67–75. [DOI] [PubMed] [Google Scholar]
- Deng, H. , Chen, Y. , Liu, Z. , Liu, Z. , Shu, P. , Wang, R. , Hao, Y. et al. (2022) SlERF.F12 modulates the transition to ripening in tomato fruit by recruiting the co‐repressor TOPLESS and histone deacetylases to repress key ripening genes. Plant Cell 34, 1250–1272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dubois, M. , Skirycz, A. , Claeys, H. , Maleux, K. , Dhondt, S. , De Bodt, S. , Vanden Bossche, R. et al. (2013) Ethylene Response Factor6 acts as a central regulator of leaf growth under water‐limiting conditions in Arabidopsis. Plant Physiol. 162, 319–332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Expósito‐Rodríguez, M. , Borges, A.A. , Borges‐Pérez, A. and Pérez, J.A. (2008) Selection of internal control genes for quantitative real‐time RT‐PCR studies during tomato development process. BMC Plant Biol. 8, 131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finch‐Savage, W.E. and Leubner‐Metzger, G. (2006) Seed dormancy and the control of germination. New Phytol. 171, 501–523. [DOI] [PubMed] [Google Scholar]
- Fu, X. and Harberd, N.P. (2003) Auxin promotes Arabidopsis root growth by modulating gibberellin response. Nature 421, 740–743. [DOI] [PubMed] [Google Scholar]
- García‐Hurtado, N. , Carrera, E. , Ruiz‐Rivero, O. , López‐Gresa, M.P. , Hedden, P. , Gong, F. and García‐Martínez, J.L. (2012) The characterization of transgenic tomato overexpressing gibberellin 20‐oxidase reveals induction of parthenocarpic fruit growth, higher yield, and alteration of the gibberellin biosynthetic pathway. J. Exp. Bot. 63, 5803–5813. [DOI] [PubMed] [Google Scholar]
- Hedden, P. (2003) The genes of the Green Revolution. Trends Genet. 19, 5–9. [DOI] [PubMed] [Google Scholar]
- Hou, S. , Zhu, J. , Ding, M. and Lv, G. (2008) Simultaneous determination of gibberellic acid, indole‐3‐acetic acid and abscisic acid in wheat extracts by solid‐phase extraction and liquid chromatography‐electrospray tandem mass spectrometry. Talanta 76, 798–802. [DOI] [PubMed] [Google Scholar]
- Illouz‐Eliaz, N. , Ramon, U. , Shohat, H. , Blum, S. , Livne, S. , Mendelson, D. and Weiss, D. (2019) Multiple gibberellin receptors contribute to phenotypic stability under changing environments. Plant Cell 31, 1506–1519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Illouz‐Eliaz, N. , Nissan, I. , Nir, I. , Ramon, U. , Shohat, H. and Weiss, D. (2020) Mutations in the tomato gibberellin receptors suppress xylem proliferation and reduce water loss under water‐deficit conditions. J. Exp. Bot. 71, 3603–3612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Izawa, T. (2020) What is going on with the hormonal control of flowering in plants? Plant J. 105, 431–445. [DOI] [PubMed] [Google Scholar]
- Jasinski, S. , Tattersall, A. , Piazza, P. , Hay, A. , Martinez‐Garcia, J.F. , Schmitz, G. , Theres, K. et al. (2008) PROCERA encodes a DELLA protein that mediates control of dissected leaf form in tomato. Plant J. 56, 603–612. [DOI] [PubMed] [Google Scholar]
- Jefferson, R.A. , Kavanagh, T.A. and Bevan, M.W. (1987) GUS fusions: beta‐glucuronidase as asensitive and versatile gene fusion marker in higher plants. EMBO J. 6, 3901–3907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kagale, S. and Rozwadowski, K. (2011) EAR motif‐mediated transcriptional repression in plants: an underlying mechanism for epigenetic regulation of gene expression. Epigenetics, 6, 141–146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khush, G.S. (1999) Green revolution: preparing for the 21st century. Genome 42, 646–655. [PubMed] [Google Scholar]
- Kozaki, A. and Aoyanagi, T. (2022) Molecular aspects of seed development controlled by gibberellins and abscisic acids. Int. J. Mol. Sci. 23, 1876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuroha, T. , Nagai, K. , Gamuyao, R. , Wang, D.R. , Furuta, T. , Nakamori, M. , Kitaoka, T. et al. (2018) Ethylene‐gibberellin signaling underlies adaptation of rice to periodic flooding. Science 361, 181–186. [DOI] [PubMed] [Google Scholar]
- Li, J. , Sima, W. , Ouyang, B. , Wang, T. , Ziaf, K. , Luo, Z. , Liu, L. et al. (2012) Tomato SlDREB gene restricts leaf expansion and internode elongation by downregulating key genes for gibberellin biosynthesis. J. Exp. Bot. 63, 6407–6420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li, C. , Zheng, L. , Wang, X. , Hu, Z. , Zheng, Y. , Chen, Q. et al. (2019a) Comprehensive expression analysis of Arabidopsis GA2‐oxidase genes and their functional insights. Plant Sci. 285, 1–13. [DOI] [PubMed] [Google Scholar]
- Li, X. , Chen, T. , Li, Y. , Wang, Z. , Cao, H. , Chen, F. and Liu, Y. (2019b) ETR1/RDO3 regulates seed dormancy by relieving the inhibitory effect of the ERF12‐TPL complex on DELAY OF GERMINATION1 expression. Plant Cell 31, 832–847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Licausi, F. , Ohme‐Takagi, M. and Perata, P. (2013) APETALA2/Ethylene Responsive Factor (AP2/ERF) transcription factors: mediators of stress responses and developmental programs. New Phytol. 199, 639–649. [DOI] [PubMed] [Google Scholar]
- Liu, X. and Hou, X. (2018) Antagonistic Regulation of ABA and GA in Metabolism and Signaling Pathways. Front. Plant Sci. 9, 251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu, J. and Sherif, S.M. (2019) Hormonal orchestration of bud dormancy cycle in deciduous woody perennials. Front. Plant Sci. 10, 1136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu, Q. , Guo, X. , Chen, G. , Zhu, Z. , Yin, W. and Hu, Z. (2016) Silencing SlGID2, a putative F‐box protein gene, generates a dwarf plant and dark‐green leaves in tomato. Plant Physiol. Biochem. 109, 491–501. [DOI] [PubMed] [Google Scholar]
- Livak, K.J. and Schmittgen, T.D. (2001) Analysis of relative gene expression data using real‐time quantitative PCR and the 2−ΔΔCt method. Methods 25, 402–408. [DOI] [PubMed] [Google Scholar]
- Livne, S. , Lor, V.S. , Nir, I. , Eliaz, N. , Aharoni, A. , Olszewski, N.E. , Eshed, Y. et al. (2015) Uncovering DELLA‐independent gibberellins responses by characterizing new tomato procera mutants. Plant Cell 27, 1579–1594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu, J. , Ju, H. , Zhou, G. , Zhu, C. , Erb, M. , Wang, X. , Wang, P. et al. (2011) An EAR‐motif‐containing ERF transcription factor affects herbivore‐induced signaling, defense and resistance in rice. Plant J. 68, 583–596. [DOI] [PubMed] [Google Scholar]
- Madhulatha, P. , Pandey, R. , Hazarika, P. and Rajam, M.V. (2007) High transformation frequency in Agrobacterium‐mediated genetic transformation of tomato by using polyamines and maltose in shoot regeneration medium. Physiol. Mol. Biol. Plants, 13, 191–198. [Google Scholar]
- Magome, H. , Yamaguchi, S. , Hanada, A. , Kamiya, Y. and Oda, K. (2008) The DDF1 transcriptional activator upregulates expression of a gibberellin‐deactivating gene, GA2ox7, under high‐salinity stress in Arabidopsis. Plant J. 56, 613–626. [DOI] [PubMed] [Google Scholar]
- Martínez‐Bello, L. , Moritz, T. and López‐Díaz, I. (2015) Silencing C19‐GA 2‐oxidases induces parthenocarpic development and inhibits lateral branching in tomato plants. J. Exp. Bot. 66, 5897–5910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matías‐Hernández, L. , Aguilar‐Jaramillo, A.E. , Cigliano, R.A. , Sanseverino, W. and Pelaz, S. (2016) Flowering and trichome development share hormonal and transcription factor regulation. J. Exp. Bot. 67, 1209–1219. [DOI] [PubMed] [Google Scholar]
- McGrath, K.C. , Dombrecht, B. , Manners, J.M. , Schenk, P.M. , Edgar, C.I. , Maclean, D.J. , Scheible, W.R. et al. (2005) Repressor‐ and activator‐type ethylene response factors functioning in jasmonate signaling and disease resistance identified via a genome‐wide screen of Arabidopsis transcription factor gene expression. Plant Physiol. 139, 949–959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Monna, L. , Kitazawa, N. , Yoshino, R. , Suzuki, J. , Masuda, H. , Maehara, Y. , Tanji, M. et al. (2002) Positional cloning of rice semi‐dwarfing gene, sd‐1: Rice “green revolution gene” encodes a mutant enzyme involved in gibberellin synthesis. DNA Res. 9, 11–17. [DOI] [PubMed] [Google Scholar]
- Nakano, T. , Suzuki, K. , Fujimura, T. and Shinshi, H. (2006) Genome‐wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol. 140, 411–432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nir, I. , Shohat, H. , Panizel, I. , Olszewski, N.E. , Aharoni, A. and Weiss, D. (2017) The tomato DELLA protein PROCERA acts in guard cells to promote stomatal closure. Plant Cell 29, 3186–3197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta, M. , Matsui, K. , Hiratsu, K. , Shinshi, H. and Ohme‐Takagi, M. (2001) Repression domains of class II ERF transcriptional repressors share an essential motif for active repression. Plant Cell 13, 1959–1968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olimpieri, I. , Caccia, R. , Picarella, M.E. , Pucci, A. , Santangelo, E. , Soressi, G.P. and Mazzucato, A. (2011) Constitutive co‐suppression of the GA 20‐oxidase1 gene in tomato leads to severe defects in vegetative and reproductive development. Plant Sci. 180, 496–503. [DOI] [PubMed] [Google Scholar]
- Pirrello, J. , Prasad, B.C.N. , Zhang, W. , Chen, K. , Mila, I. , Zouine, M. , Latché, A. et al. (2012) Functional analysis and binding affinity of tomato ethylene response factors provide insight on the molecular bases of plant differential responses to ethylene. BMC Plant Biol. 12, 190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plant, A.R. , Larrieu, A. and Causier, B. (2021) Repressor for hire! The vital roles of TOPLESS‐mediated transcriptional repression in plants. New Phytol. 231, 963–973. [DOI] [PubMed] [Google Scholar]
- Ramon, U. , Weiss, D. and Illouz‐Eliaz, N. (2021) Underground gibberellin activity: differential gibberellin response in tomato shoots and roots. New Phytol. 229, 1196–1200. [DOI] [PubMed] [Google Scholar]
- Schrager‐Lavelle, A. , Gath, N.N. , Devisetty, U.K. , Carrera, E. , López‐Díaz, I. , Blázquez, M.A. and Maloof, J.N. (2019) The role of a class III gibberellin 2‐oxidase in tomato internode elongation. Plant J. 97, 603–615. [DOI] [PubMed] [Google Scholar]
- Serrani, J.C. , Ruiz‐Rivero, O. , Fos, M. and Garcia‐Martinez, J.L. (2008) Auxin‐induced fruit‐set in tomato is mediated in part by gibberellins. Plant J. 56, 922–934. [DOI] [PubMed] [Google Scholar]
- Shinozaki, Y. , Hao, S. , Kojima, M. , Sakakibara, H. , Ozeki‐Iida, Y. , Zheng, Y. , Fei, Z. et al. (2015) Ethylene suppresses tomato (Solanum lycopersicum) fruit set through modification of gibberellin metabolism. Plant J. 83, 237–251. [DOI] [PubMed] [Google Scholar]
- Shohat, H. , Illouz Eliaz, N. , Kanno, Y. , Seo, M. and Weiss, D. (2020) The tomato DELLA protein PROCERA promotes abscisic acid responses in guard cells by upregulating an abscisic acid transporter. Plant Physiol. 184, 518–528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shohat, H. , Ceriker, H. , Vasuki, H. , Illouz‐Eliaz, N. , Blum, S. , Amsellem, Z. , Tarkowska, D. et al. (2021a) Inhibition of gibberellin accumulation by water deficiency promotes fast and long‐term 'drought avoidance' responses in tomato. New Phytol. 232, 1985–1998. [DOI] [PubMed] [Google Scholar]
- Shohat, H. , Eliaz, N.I. and Weiss, D. (2021b) Gibberellin in tomato: metabolism, signaling and role in drought responses. Mol. Hortic. 1, 15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singh, R. , Ranjan, S. , Nayaka, S. , Pathre, U.V. and Shirke, P.A. (2013) Functional characteristics of a fruticose type of lichen, Stereocaulon foliolosum Nyl. in response to light and water stress. Acta Physiol. Plant, 35, 1605–1615. [Google Scholar]
- Singh, P. , Singh, A.P. , Tripathi, S.K. , Kumar, V. and Sane, A.P. (2019) Petal abscission in roses is associated with the activation of a truncated version of the animal PDCD4 homologue, RbPCD1 . Plant Sci. 288, 110242. [DOI] [PubMed] [Google Scholar]
- Song, C.P. , Agarwal, M. , Ohta, M. , Guo, Y. , Halfter, U. , Wang, P. and Zhu, J.K. (2005) Role of an Arabidopsis AP2/EREBP‐type transcriptional repressor in abscisic acid and drought stress responses. Plant Cell 17, 2384–2396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun, T.P. (2010) Gibberellin‐GID1‐DELLA: a pivotal regulatory module for plant growth and development. Plant Physiol. 154, 567–570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Upadhyay, R.K. , Soni, D.K. , Singh, R. , Dwivedi, U.N. , Pathre, U.V. , Nath, P. and Sane, A.P. (2013) SlERF36,an EAR motif containing ERF gene from tomato, alters stomatal density and modulates photosynthesis and growth. J. Exp. Bot. 64, 3237–3247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Upadhyay, R.K. , Gupta, A. , Ranjan, S. , Singh, R. , Pathre, U.V. , Nath, P. and Sane, A.P. (2014) The EAR motif controls the early flowering and senescence phenotype mediated by over‐expression of SlERF36 and is partly responsible for changes in stomatal density and photosynthesis. PloS One, 9, e101995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang, S. , Chang, Y. , Guo, J. and Chen, J.G. (2007) Arabidopsis Ovate Family Protein 1 is a transcriptional repressor that suppresses cell elongation. Plant J. 50, 858–872. [DOI] [PubMed] [Google Scholar]
- Weiss, D. and Ori, N. (2007) Mechanisms of cross talk between gibberellin and other hormones. Plant Physiol. 144, 1240–1246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wen, L.H. , Zhong, W.J. , Huo, X.M. , Zhuang, W.B. , Ni, Z.J. and Gao, Z.H. (2016) Expression analysis of ABA‐and GA‐related genes during four stages of bud dormancy in Japanese apricot (Prunus mume Sieb. et Zucc). J. Hortic. Sci. Biotechnol. 91, 362–369. [Google Scholar]
- Yamaguchi, S. (2008) Gibberellin metabolism and its regulation. Annu. Rev. Plant Biol. 59, 225–251. [DOI] [PubMed] [Google Scholar]
- Yamaguchi, N. , Winter, C.M. , Wu, M.F. , Kanno, Y. , Yamaguchi, A. , Seo, M. and Wagner, D. (2014) Gibberellin acts positively then negatively to control onset of flower formation in Arabidopsis . Science 344, 638–641. [DOI] [PubMed] [Google Scholar]
- Yanai, O. , Shani, E. , Russ, D. and Ori, N. (2011) Gibberellin partly mediates LANCEOLATE activity in tomato. Plant J. 68, 571–582. [DOI] [PubMed] [Google Scholar]
- Yang, Z. , Tian, L.N. , Latoszek‐Green, M. , Brown, D. and Wu, K.Q. (2005) Arabidopsis ERF4 is a transcriptional repressor capable of modulating ethylene and abscisic acid responses. Plant Mol. Biol. 58, 585–596. [DOI] [PubMed] [Google Scholar]
- Yu, K. , Wei, J. , Ma, Q. , Yu, D. and Li, J. (2009) Senescence of aerial parts is impeded by exogenous gibberellic acid in herbaceous perennial Paris polyphylla. J. Plant Physiol. 166, 819–830. [DOI] [PubMed] [Google Scholar]
- Zhang, H. , Zhang, J. , Quan, R. , Xiaowu, P. , Wan, R. and Huang, R. (2013) EAR motif mutation of rice OsERF3 alters the regulation of ethylene biosynthesis and drought tolerance. Planta, 237, 1443–1451. [DOI] [PubMed] [Google Scholar]
- Zhang, Y. , Liu, Z. , Wang, X. , Wang, J. , Fan, K. , Li, Z. and Lin, W. (2018) DELLA proteins negatively regulate dark‐induced senescence and chlorophyll degradation in Arabidopsis through interaction with the transcription factor WRKY6. Plant Cell Rep. 37, 981–992. [DOI] [PubMed] [Google Scholar]
- Zhou, Z. , Bi, G. and Zhou, J.‐M. (2018) Luciferase complementation assay for protein‐protein interactions in plants. Curr. Protoc. Plant Biol. 3, 42–50. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1 List of oligonucleotides used in the study.
Figure S1 SlERF36 expression in different tissues and in different OEx and suppression lines.
Figure S2 SlERF36 manipulation regulates various growth processes.
Figure S3 Plant phenotypes of control and transgenic SlERF36 lines upon exogenous treatment with GA3 and paclobutrazol.
Figure S4 qRT‐PCR analysis showing relative changes in transcript levels of GA2ox genes.


