ABSTRACT
Single-strand RNA (ssRNA) and single-strand DNA phages elicit host lysis using a single gene, in each case designated as sgl. Of the 11 identified Sgls, three have been shown to be specific inhibitors of different steps in the pathway that supplies lipid II to the peptidoglycan (PG) biosynthesis machinery. These Sgls have been called “protein antibiotics” because the lytic event is a septal catastrophe indistinguishable from that caused by cell wall antibiotics. Here, we designate these as type I Sgls. In this formalism, the other eight Sgls are assigned to type II, the best-studied of which is protein L of the paradigm F-specific ssRNA phage MS2. Comparisons have suggested that type II Sgls have four sequence elements distinguished by hydrophobic and polar character. Environmental metatranscriptomics has revealed thousands of new ssRNA phage genomes, each of which presumably has an Sgl. Here, we describe methods to distinguish type I and type II Sgls. Using phase contrast microscopy, we show that both classes of Sgls cause the formation of blebs prior to lysis, but the location of the blebs differs significantly. In addition, we show that L and other type II Sgls do not inhibit the net synthesis of PG, as measured by radio-labeling of PG. Finally, we provide direct evidence that the Sgl from Pseudomonas phage PP7 is a type I Sgl, in support of a recent report based on a genetic selection. This shows that the putative four-element sequence structure suggested for L is not a reliable discriminator for the operational characterization of Sgls.
IMPORTANCE
The ssRNA phage world has recently undergone a metagenomic expansion upward of a thousandfold. Each genome likely carries at least one single-gene lysis (sgl) cistron encoding a protein that single-handedly induces host autolysis. Here, we initiate an approach to segregate the Sgls into operational types based on physiological analysis, as a first step toward the alluring goal of finding many new ways to induce bacterial death and the attendant expectations for new antibiotic development.
KEYWORDS: bacteriophage lysis, bacteriophages, bacteriophage evolution
INTRODUCTION
For host lysis, double-strand DNA phages encode multiple lysis proteins that actively attack each layer of the envelope according to a tightly regulated temporal schedule (1, 2). More than a dozen different functional types of proteins have been linked to this “multi-gene lysis” (MGL) process, with some phages encoding as many as six. By contrast, small lytic phages with single-strand RNA and DNA chromosomes lack the genomic space for an MGL system. Instead, these phages, the Microviridae and the ssRNA Leviviricetes, liberate their progeny by inducing lysis through the expression of a single gene (3, 4). These genes have been collectively named sgl (single-gene lysis; product Sgl). Importantly, Sgls do not have bacteriolytic enzyme activities, and thus, each is, formally, an inducer of host autolysis. As such, Sgls are of interest in terms of the development of new antibiotic strategies, each representing a potential modality for instigating autolysis of a bacterial cell (5). The position and character of the sgls are particularly striking in the ssRNA phages, which have three core genes that encode virion and replication proteins (Fig. 1A). In contrast to the core genes, which are always in the same order and approximately the same size, the location and size of the Sgl are highly variable, embedded out of frame in one of the three core genes or positioned in an intergenic space (3). This strongly indicates that there is an independent evolutionary path for each sgl. The classic case is the first Sgl, identified by genetic analysis in the canonical F-specific coliphage MS2 (6). In this case, SglMS2, given the gene name L for lysis, is a 75-codon reading frame overlapping the coat-rep junction (Fig. 1). Its Sgl character was unambiguously demonstrated by its ability to cause autolysis by induction from plasmid expression vectors (7–9). Soon after, the Sgls of two other paradigm coliphages were identified by the same functional criterion: E, the Sgl of the canonical single-strand DNA (ssDNA) Microvirus ϕX174, and A2, the dual-function maturation protein/Sgl of Qβ, another well-studied F-specific ssRNA phage (3). The homologs of L and A2 were identified in the genomes of many MS2 and Qβ-like F-specific coliphages (10). Outside of L, A2, and E, only seven other Sgls have been identified, encoded by ssRNA phages specific for the pili of conjugative plasmids of Escherichia coli (HgaI, C-1, and M) (10, 11) and Pseudomonas (PRR1) (12), as well as for the polar retractable pili of Pseudomonas (PP7) (13, 14), Acinetobacter (AP205) (15), and Caulobacter crescentus (phiCB5) (16, 17). Definitive mechanistic information is available for only E and A2. Biochemical and structural evidence indicates that each acts by binding and inhibiting an essential enzyme in the pathway for the biosynthesis of lipid II, the precursor of peptidoglycan biosynthesis: MraY and MurA, respectively (18–21). In addition, genetic and molecular evidence has indicated that the Sgls of coliphage M and the Pseudomonas phage PP7 inhibit MurJ, which flips lipid II to the periplasm as the last step of the pathway (22). The sobriquet “protein antibiotic,” originally coined for A2, alluding to the functional similarity to beta-lactams and other cell wall antibiotics (23), could reasonably be extended to all four of these Sgls.
Fig 1.
Peptidoglycan biosynthesis pathway and Sgl inhibition (A). Genomes of ssDNA (ϕX174) and ssRNA (Qβ, M, PP7, MS2, KU1, Hgal, and PRR1) phages. Adapted from reference (9). SsRNA phages have three core conserved genes in the same order: mat (orange), coat (black), and rep (green). These phages also encode a lysis gene (blue) that is embedded in an alternative reading frame in one of the three core genes or intergenic spaces. In Qβ, the maturation gene has a dual function, serving as both the maturation protein and the Sgl. (B). Sgls of ssRNA and ssDNA phages such as ϕx174, Qβ, and M inhibit different enzymatic steps in the PG biosynthesis pathway.
In contrast, mechanistic information for the other six Sgls is sparse. MS2 L has been the subject of multiple studies, beginning in the mid-1980s with a series of collaborative works from the groups of Holtje and van Duin. One monograph reported pulse labeling studies showing that L did not affect the rate of PG biosynthesis (24). In addition, immuno-electron microscopy data were cited indicating that L was enriched in zones of adhesion between the inner membrane (IM) and outer membrane (OM) (25). Finally, a synthetic peptide corresponding to the C-terminal 25 AA of L was reported to have a pore-forming character for liposomes, leading to the model that L-mediated autolysis involved membrane pore formation (26). More recently, Mezhyrova et al. (27) used in vitro liposome permeabilization experiments and in vivo cryo-electron microscopy to support this model. In addition, our group reported a genetic analysis of L (8). The results and comparison with L sequences from closely related F-specific MS2-like phages suggest that L has four elements, including an essential hydrophobic region preceding a conserved Leu-Ser dipeptide (Fig. S1). Moreover, the highly cationic N-terminal region, although previously shown to be non-essential for lytic function, was found to confer dependence on the host chaperone DnaJ (7, 9). Other than the three Sgls shown to be inhibitors of the lipid II biosynthesis pathway, all the other Sgls have similar elements, including the LS dipeptide, although none have significant sequence similarity (Fig. S1) (8). Taken together with the studies on L, these findings suggested that L and the other five Sgls might be functionally related (“L-like”). Recently, the significance of the LS dipeptide was brought into question when SglPP7, which has a similar multi-element structure, was implicated as being an inhibitor of MurJ by multi-copy suppression and genetic studies (22).
Recently, thousands of new ssRNA phage genomes have been discovered in environmental transcriptomes and viromes (28–31). The diversity of this new ssRNA genomic space is staggering but not surprising, considering the high mutagenesis rate that characterizes RNA-dependent RNA polymerases (32). Importantly, homologs of the known Sgls were very rare in these new genomes. Moreover, 33 new sgl genes were found by bioinformatic and functional analysis, mapping to at least 18 different positions in the genome (19). This unparalleled diversity raises the alluring possibility that Sgl lysis targets might titrate many of the known steps of PG biosynthesis and homeostasis, plus new ones, leading to multiple new antibiotic targets. Some success at identifying and characterizing the sgl genes from these collections was obtained by synthesizing more than 300 potential sgl genes, selected for having a potential transmembrane domain (TMD) as well as plausible reading frames and translational start (28). When these sgl candidates were synthesized and tested for inducible lysis in E. coli, about 10% were shown to have a lytic function, a reasonable success rate considering that the bacterial hosts for these hypothetical phages are not known. Given the number of Sgls to be tested, it would be very useful to be able to segregate the “protein antibiotic” Sgls from the L-like Sgls, based on the notion that each identifies a sensitive molecular interaction surface in a potential antibiotic target.
Given this new wide perspective and the likelihood that there will be many fundamentally different mechanistic systems, we propose to organize the Sgl database into a series of types (Table 1), as has been done profitably with the CRISPR system. We propose to reclassify the Sgls that inhibit PG biosynthesis as type I Sgls and the L-like Sgls as type II. For the purposes of the rest of this report, we will use this terminology. In addition, we will follow historical precedent in referring to the well-studied Sgls from MS2, Qβ, and ϕ174 as L, A2, and E, respectively, but all others as SglX, where X is the name of the phage.
TABLE 1.
Classification of Sgls of known plaque-forming ssDNA and ssRNA phages
| Phage | Host | Sgl | Reference |
|---|---|---|---|
| Type I Sgls | |||
| ϕX174 | E. coli | E | (33, 34) |
| Qβ | E. coli | A2 | (35, 36) |
| M | E. coli | SglM | (11) |
| Type II Sgls | |||
| MS2 | E. coli | L | (37, 38) |
| KU1 | E. coli | SglKU1 | (39) |
| Hgal | E. coli | SglHgal | (10) |
| C-1 | E. coli | SglC1 | (10) |
| PRR1 | P. aeruginosa | SglPRR1 | (12) |
| PP7 | P. aeruginosa | SglPP7 | (14) |
| Unknown classification | |||
| LeviOr01 | P. aeruginosa | Unknown | (40) |
| PhiCB5 | C. crescentus | SglCB5 | (16) |
| AP205 | Acinetobacter | SglAP205 | (15) |
RESULTS
Type I Sgls but not type II Sgls inhibit net PG synthesis
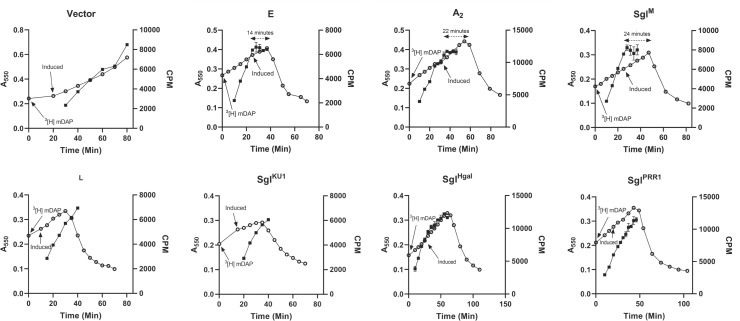
As noted above, it has been reported that, for lytic inductions of L, the rate of PG synthesis, as measured by pulse labeling with 3[H]mDAP, is unaffected up to the time of lysis (24). In contrast, for both E and A2, net PG synthesis, as measured by the continuous incorporation of the same label, is halted at least 20 min before the lytic event (23, 41). These results raised the possibility that type I Sgls can be distinguished from type II using PG labeling. The simplest method is to follow the incorporation of 3[H]mDAP into material that is insoluble in boiling SDS, i.e., the sacculi. However, 3[H]mDAP labeling requires a minimal medium, which is done with glucose as the carbon source. Moreover, in our hands, reproducible results with cloned sgl genes require the use of tightly regulated arabinose (Ara)-inducible vectors (see Fig. S2), because the appreciable basal level expression of these highly toxic genes leads to growth anomalies prior to induction. However, since Ara inductions are subject to catabolite repression, a different carbon source was required. Among the available carbon sources that do not cause catabolite repression, studies with E. coli B/r have shown that glycerol supports reasonable growth rates if the minimal medium is supplemented with all the amino acids (42–44). We tested glycerol as a carbon source for our standard induction system, using RY34348, a lysA derivative of the E. coli K-12 strain MG1655 (Fig. 2). Cultures exhibited reproducible growth rates of ~1 doubling/hr and, depending on the cloned sgl gene, showed robust, distinct lysis profiles at 10–30 min after the addition of arabinose. In the case of two well-established type I Sgls, E and SglM, the net incorporation of 3[H]mDAP stopped abruptly long before lysis was apparent (Fig. 3). In contrast, the incorporation continued unabated until lysis in the case of the type II paradigm, L, consistent with the previous pulse labeling results (15). Similar results were obtained with the Sgl from the MS2-like phage KU1, which is significantly diverged from L at only 28% identity (8), and the Pseudomonas phage PRR1, indicating that both are type II Sgls (Fig. 3). A modified labeling system was required for the sgl of phage Hgal, which for unknown reasons failed to support lysis when induced from the Ara-inducible platform (Fig. S2). In this case, the crystal violet-inducible vector pJexD was used (Fig. S3), and again, labeling continued until lysis (Fig. 3). Taken together, these data indicate all three are type II Sgls. Although none of these genes have been experimentally characterized, they all have marginal similarity to MS2 L, have a four-element structure, and are encoded in a reading frame similarly positioned to L overlapping the coat–rep boundary (8).
Fig 2.
Sgl lysis profiles in defined minimal media. Lysis profiles of Sgls in minimal media supplemented with 16 amino acids, 0.01% thiamine, and 0.2% glycerol as the carbon source. When the cultures reached an OD of ~0.2, they were induced with 0.4% L-arabinose. From top to bottom: pBAD24 vector with no gl (closed circles), E (closed downward triangle), A2 (open circles), SglM (closed diamonds), SglPP7 (open squares), L (closed squares), SglKU1 (closed upward triangles), and SglPRR1 (open triangles).
Fig 3.
3[H]mDAP labeling of peptidoglycan. Growth curve of E. coli is shown as open circle (O), and the amount of 3[H]mDAP counted at different time points is shown as closed squares (◼). The first arrow indicates when 3[H]mDAP was added to the bacterial culture, and the second arrow indicates the induction of the plasmid. The dual-sided arrows indicate the time between cessation of 3[H]mDAP incorporation and lysis. Each construct has a minimum of one repeat, and the representative one is shown in each case. For type I Sgls, after induction, each time point was filtered twice (see Materials and Methods). The average is plotted, and the error bars represent the variation.
Type I and type II Sgls cause different lytic morphology
Exposure to cell wall antibiotics causes a distinct morphological change in growing bacteria (45, 46). Most cells undergo catastrophic septal failures, with a large bleb emanating from the mid-cell. This presumably reflects the requirement for massive de novo PG biosynthesis localized to the developing septum, compared to the highly distributed nature of PG biosynthesis in the elongating cell walls (47). In principle, all type I Sgls should cause identical lytic morphology, given the common end-point (i.e., starvation for periplasmic lipid II). Indeed, E. coli cells infected with the ssDNA phage ϕX174, which uses the canonical type I Sgl E, exhibit mid-cell blebbing (48). This lysis morphology has been recapitulated with the cloned E gene (49–51), demonstrating that the expression of E is necessary and sufficient not only for lysis but also for the particular lysis morphology characterized by a septal catastrophe. We tested if this morphological phenotype could be used as a criterion for distinguishing type I and type II Sgls. We reproduced the E finding and found that the septal blebbing was also characteristic of other type I Sgls, A2 and SglM (Fig. 4 to 6). In contrast, L and the putative type II Sgls of KU1, PRR1, and Hgal gave rise to more distributed, numerous, and smaller blebs (Fig. 4 and 5). Moreover, the blebs exhibited by the type II Sgls were significantly less stable, as judged by the frequency of lysed cells in microscopy fields (Fig. S4) . Indeed, being able to visualize any significant number of intact blebs, as judged by phase contrast, required the addition of Mg++ ions and sucrose to stabilize the envelope. Stabilizing conditions were used to image both type I and type II Sgls for a direct comparison.
Fig 4.
Type I and type II lysis morphology. Shown are phase contrast images of E. coli after induction of different lysis genes. (A) Vector only. (B–D) Type I Sgls: E, A2, and SglM. (E–H) Type II Sgls: L, SglKU1, SglHgal, and SglPRR1. For panels (A–G), cultures were grown at 37°C to an optical density of ~0.2 and induced with 0.4% L-arabinose. For panel (H), cultures were induced with 0.075 mM crystal violet. Ten minutes prior to lysis, 1.5-μL samples were taken from culture flasks, put onto glass slides, covered with a cover slip, and imaged using the 100× objective lens. Collages were made using Inkscape, and each represents the population of cells seen in triplicate imaging experiments. Furthermore, to minimize bias in the selection of cells with blebs for the collages, all cells showing blebs on an image were incorporated into collages.
Fig 6.
SglPP7 portrays type I lysis morphology and isotope labeling. (A) Microscopy images of bleb formation after the induction of the SglPP7. (B) Isotope labeling of the peptidoglycan layer using 3[H]mDAP. The open circles represent the optical density of the culture, and the closed squares represent the counts per minute of 3[H]mDAP incorporated into insoluble material. The solid arrow on the left indicates the time at which 3[H]mDAP was added to the growing culture. The middle solid arrow shows when the culture was induced with 0.4% L-arabinose. The dual-sided arrow indicates the time between cessation of 3[H]mDAP incorporation and lysis.
Fig 5.
Analysis of lesions caused by Sgls. The y-axis represents the location of the bleb along the cell length with 0.0 representing the poles of the cell and 0.5 representing middle of the cell. The x-axis indicates the type of Sgl for each set of measurements, including type I (E, A2, SglM, and SgPP7) and type II (L, SglKU1, SglHgal, and SglPRR1) Sgls. Measurements of cells (n = 25 for each Sgl) were done using the measurement tool from ImageJ. The box plot was created using GraphPad Prism 9. To minimize bias in the selection of cells with blebs for the analysis, all cells showing blebs in an image were measured. A comparison between type I Sgls and type II Sgls was conducted using a Levene and Fligner–Killeen test, both of which yielded a P-value less than 0.001.
Type I character of the Sgl of Pseudomonas phage PP7
Among the Sgls originally assigned to the “L-like” or type II category by virtue of having a four-element structure that resembled that established for L is the Sgl of Pseudomonas phage PP7, which uses the polar pilus for its receptor (8, 52). However, during this project, Adler and colleagues reported the results of a screen for multi-copy suppressors for a number of Sgls; in this report, they identified murJ as a multi-copy suppressor for PP7 (22). In the same work, this unexpected finding was supported by genetic analysis, including the isolation of missense mutations in murJ conferring resistance to SglPP7. Interestingly, this Sgl had been classified as type II based on the presence of the four-element motif (8). Thus, we wondered if SglPP7 might have both type I and type II characters. However, applying the two methods developed here revealed solely type I phenotypes (Fig. 5 and 6). SglPP7 caused an abrupt halt in net PG biosynthesis, as judged by 3[H]mDAP incorporation, as well as a septal bleb lysis morphology.
DISCUSSION
Here, we show that the type I and type II Sgls can be distinguished by two methods, incorporation of 3[H]mDAP into PG and phase contrast microscopy of lysing cells. In addition, we provide physiological evidence that supports the recent finding that SglPP7 is a type I Sgl. The first criterion, incorporation of 3[H]mDAP into insoluble PG, is the most definitive and mechanistically relevant because the enzymes required for de novo biosynthesis and externalization of lipid II, the immediate precursor of PG polymer, are known and highly conserved. With the discovery that SglPP7 is a MurJ inhibitor, there are now four different small lytic phages that have been shown to use a type I Sgl for host lysis: ϕX174 and Qβ targeting MraY and MurA, respectively, and M and PP7 targeting MurJ. So far, none of the proteins in the multi-meric complexes [i.e., the elongasome and the divisome (47, 53)] that incorporate the disaccharide subunit of lipid II into the PG has shown up as an Sgl target. This could be interpreted as surprising, since the septal bleb morphological phenotype that characterizes type I Sgl function looks the same as that produced by beta-lactam antibiotics. However, efficient lysis in the context of a phage infection might require the inhibition of both the elongasome and divisome complexes, which would necessitate that the Sgl target two different proteins, one in each complex. It is also worth noting that two of the four known type I Sgls target the same protein, MurJ, the lipid II “flippase.” Moreover, all three of the type I Sgls that evolved within the boundaries of essential phage genes target membrane proteins (E targets MraY, and both SglPP7 and SglM target MurJ). It may be easier to evolve a functional Sgl that is localized to the membrane, where the lipidic environment imposes a helical structure on the polypeptide, than one required to fold in the aqueous phase. We have shown that, for candidate Sgls with TMDs, the ability to cause overt lysis in E. coli can be selected rapidly (28). In contrast, evolving de novo a polypeptide to target a soluble enzyme might be much easier with a pre-existing tertiary structural scaffold, i.e., A2 inhibiting MurA.
It should be noted that the original conclusion that L did not act by inhibiting cell wall synthesis used a pulse labeling protocol and thus did not rule out the possibility that L acted by inducing PG degradation. Our results show that net PG synthesis is unaffected by L, and therefore, the balance between overall PG synthesis and degradation is unchanged. This does not rule out localized imbalances. The simplest general concept is that L interferes with the proper distribution of PG biosynthesis, so that the multi-protein PG biosynthesis complexes are just as active but not positioned at the right places for homeostasis and growth. It is also important to note that at this point, type II Sgl function is mainly defined as the absence of type I characteristics. The notion that L was just one of many similarly functioning Sgls produced by convergent evolution was in part based on the attractive notion that the four-element structure, defined by genetic analysis of L, seemed to be present in other Sgls. This suggests that the presence of the four-element sequence motifs might not be a reliable criterion for distinguishing between type I and type II Sgls (22).
Finally, it must be noted that there is no evidence that these ssRNA genomes that are extracted from metatranscriptomes are from plaque-forming lytic phages. The ssRNA infection cycle does not directly interfere with the DNA-based life of the host cell, so it can be considered that many if not most RNA phages cause chronic infections in which virions accumulate indefinitely while allowing the host to continue growth and division cycles. Indeed, even with the known ssRNA phages, infections can lead to non-lytic intracellular accumulation of virions to levels of 104–105 per cell (13). Thus, many ssRNA phages may persist in an endemic carrier state; in this scenario, only occasional host lysis events occur every few generations, thus seeding the environment with virions that can take advantage of new host cells. The original isolate Q that gave rise to Qβ was just such a chronic phage, since it formed regions of retarded growth on plates, rather than sharply defined plaques, and gave rise to bacterial colonies that constitutively produced virions (54). Modern Qβ likely arose from mutations that increased the affinity of A2 for MurA. In fact, single base changes that increase the production rate of A2 and overcome host resistance due to missense changes in MurA arise at a high rate in the context of the phage (55).
MATERIALS AND METHODS
Bacterial strains, plasmids, and growth conditions
The bacterial strains used in this work, unless otherwise stated, were as follows: DH5α (ThermoFisher Scientific). The plasmids and primers that were used in this study can be found in (Table 2). Bacterial cultures were grown in lysogeny broth (LB) with 100 µg·mL−1 ampicillin at 37°C with constant aeration in a gyrotory water bath shaker (New Brunswick Scientific) for lysis curves and microscopy. Overnight cultures were grown in 18 × 150 mm glass culture tubes (Fisher Scientific 14-961-32). For overnight cultures of E. coli harboring the pBAD24 vector, glucose was added at a final concentration of 0.4% to prevent premature gene expression. For isotope labeling experiments, bacterial cultures were grown in defined M9 media supplemented with 0.2% glycerol, 0.01% thiamine, 100 µg·mL−1 ampicillin, and L-amino acids (20 μg·mL−1 His; 30 µg·mL−1 Gly, Ile, Phe, Pro, Ser, and Val; 50 µg·mL−1 Arg; 60 µg·mL−1 Ala; 70 µg·mL−1 Asn and Lys; 80 µg·mL−1 Glu; 90 µg·mL−1 Gln; 100 µg·mL−1 Met and The; and 110 µg·mL−1 Leu). High levels of Met and Thr were added to reduce the intracellular pool of mDAP (mDAP is the precursor for lysine) (45). Furthermore, the lysA strain RY3448 was used to prevent the conversion of mDAP to lysine, ensuring that our labeled mDAP was solely incorporated into the PG layer and not elsewhere. The induction of cultures was done with 0.4% (wt/vol) L-arabinose.
TABLE 2.
Strains, plasmids, and primers used in this study
| Strain, plasmid, or primer | Relevant genotype, features, or sequence (5′−3′) | Reference |
|---|---|---|
| Strains | ||
| DH5αΤ1Ρ | ϕ80 fhuA dΔ(lxc-argF) [ΔlacZM15] U169 phoA supE44 gyrA96 recA1 relA1 endA1 thi1 hsdR17(rK− mK+) | Thermo Scientific |
| RY34010 | MG1655 ΔlacIZYA <>frt | TB28 in (56) |
| RY34348 | RY34010 ΔlysA763 <>frt | (57) |
| Plasmids | ||
| pBAD24 | bla araC ParaBAD | (58) |
| pBAD24_L | bla araC ParaBAD::L | (8) |
| pBAD24_E | bla araC ParaBAD::E | Laboratory collection |
| pBAD24_A2 | bla araC ParaBAD::A2 | This work |
| pBAD24_PP7 | bla araC ParaBAD::SglPP7 | Laboratory collection |
| pBAD24_KU1 | bla araC ParaBAD::SglKU1 | Laboratory collection |
| pBAD24_M | bla araC ParaBAD:: SglM | Laboratory collection |
| pBAD24_PRR1 | bla araC ParaBAD:: SglPRR1 | Laboratory collection |
| pAJEXD_Hgal | eilR PJExD :: SglHgal | This work |
| Primers | ||
| KC30 | cggcagaaaagtccacattg | |
| KC31 | cacactaccatcggcgcta | |
| KC710 | ggcgatagtgcgaaccttgtcattattgactttctg | |
| KC711 | gcccgagatattttatagtctcgagcccttcgcc | |
| KC713 | ggtgaaagatttgagccggatgtcacgtctgtc | |
| FA_A2_F_Ecor1 | gagagtgaattccatatgcctaaattaccgcgtgg | |
| FA_A2_R_Hind3 | gtatataagcttatcaacgctttacgcgttggg | |
| Forward pAJEXD_FA | ggatccaaactcgagtaaggatctcc | |
| Reverse pAJEXD_FA | atgtatatctccttcttaaaagatcttttgaattcaaagttgg | |
| FW Hgal_pAJEXD_FA | actttgaattcaaaagatcttttaagaaggagatatacatatggccaacccgcgc | |
| RV Hgal_pAJEXD_FA | tttatttgatgcctggagatccttactcgagtttggatccttacgtcatgggaaccacataggcgt | |
Chemical reagents and enzymes
Tritiated diaminopimelic (250 µCi·mL−1) acid was purchased from VWR. All other chemicals were purchased from Sigma Aldrich unless otherwise stated.
Microscopy
Prior to microscopy, lysis curves of each plasmid construct in strain RY34010 were done by inoculating a 250-mL culture flask containing 25 mL of LB and an appropriate amount of antibiotic (100 µg·mL−1 ampicillin or 40 µg·mL−1 kanamycin) with a 1:200 dilution of bacteria. Cultures were grown with aeration in a water bath until they reached an A550 of ~0.2, at which they were then induced with 0.4% L-arabinose or 0.075 µM of crystal violet. For all microscopy images, samples were taken 10 min prior, 5 min prior, and at the time of expected lysis. At each time point, 1.5-µL samples were put onto a 25 mm × 75 mm × 1 mm glass slide (Globe Scientific) and covered with a 22 mm × 22 mm cover slip (Globe Scientific) and imaged in inverted orientation at 100× with oil immersion on a Zeiss microscope. Slides were imaged at an exposure of 100 ms for no longer than 20 min at a time to avoid artifacts.
Image analysis
The images were analyzed using ImageJ (59). The length from each pole to the center of the bleb was measured three times. The center of the bleb can be defined as the region at the base of the bleb that is equidistant from the left and right sides of the bleb (protruding membranes). These measurements were used to calculate the total length of the cell as well as calculate the location of the bleb.
3[H]mDAP incorporation and measurement of PG synthesis
For the labeling experiments, strains were grown in 25 mL of defined media in 250-mL culture flasks (mother culture) until an A550 ~0.1 was reached. At this point, 5 mL of the mother culture was placed into a pre-warmed 50-mL culture flask (daughter culture). Both cultures were then left to grow to an A550 ~0.2; the growth of cultures was measured via the mother flask. Once the culture reached the desired optical density (OD), 3[H]mDAP was added to the daughter flask to a final level of 1 µCi·mL−1 and given a 10-min pre-labeling period. After the pre-labeling period, 200-µL samples were taken at designated time points (Fig. 3) and put into a microcentrifuge tube with 800 µL boiling 5% SDS and vortexed vigorously. The samples were boiled for an hour and a half and then cooled to room temperature. After cooling to room temperature, the samples were vortexed vigorously again and briefly spun down. Filters were rinsed with 5 mL of lysine (70 µg·mL−1) before the addition of any sample. After the blocking step, 200 µL of each sample was filtered through a 0.22-µm cellulose nitrate membrane filter (Cytiva), washed with 30 mL of the lysine–water solution, and left to dry overnight. For type I Sgls, each time point had two samples filtered, such that an average for each time point could be taken. Next, the filters were put into plastic 20-mL vials (Fisher Scientific) with Ecoscint A scintillation fluid (National Diagnostics). The vials with the filters were then kept in the dark for a minimum of 1 hr prior to reading. The counts per minute of the samples were read in a Tri-Carb 2910 TR liquid scintillation counter (PerkinElmer) with a count time of 1 min.
ACKNOWLEDGMENTS
The authors want to thank Diobenhi Castellanos for her assistance with the lysis curve replicates. We also thank Jennifer Tran and Lorna Min for the assistance in cloning and preliminary experiments in cell morphology and Dr. Heath Blackmon for the assistance in conducting the statistical analysis.
This work was supported by R35GM136396 to R.Y. and the Center for Phage Technology at Texas A&M University, jointly sponsored by Texas A&M AgriLife. Additional funding for this project and support of S. Francesca Antillon were provided funding by the National Institute of General Medical Sciences, grant number T32GM135748. Funding for Diobenhi Castellanos was provided through the National Science Foundation REU Site in Biochemistry at Texas A&M University, award ID DBI-1949893.
S.F.A. and R.Y. were primarily responsible for the design of the studies and analysis of the results. T.G.B. designed and executed the original radioactive labeling experiments. S.F.A. performed most of the physiological experiments reported and created the figures. S.F.A. and R.Y. prepared and edited the manuscript. K.C. and T.G.B. provided edits to the manuscript.
Contributor Information
Ry Young, Email: ryland.young@ag.tamu.edu.
Joseph Bondy-Denomy, University of California San Francisco, San Francisco, California, USA.
DATA AVAILABILITY
Raw data generated during the course of this study are available from the corresponding author upon request. The code used in this study to analyze the data in Fig. 5 is available from a public repository at https://github.com/AntillonF/bleb_statistical_analysis.git.
SUPPLEMENTAL MATERIAL
The following material is available online at https://doi.org/10.1128/jb.00384-23.
Figures S1 to S4.
ASM does not own the copyrights to Supplemental Material that may be linked to, or accessed through, an article. The authors have granted ASM a non-exclusive, world-wide license to publish the Supplemental Material files. Please contact the corresponding author directly for reuse.
REFERENCES
- 1. Cahill J, Young R. 2019. Phage lysis: multiple genes for multiple barriers. Adv Virus Res 103:33–70. doi: 10.1016/bs.aivir.2018.09.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Young R. 2014. Phage lysis: three steps, three choices, one outcome. J Microbiol 52:243–258. doi: 10.1007/s12275-014-4087-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Chamakura K, Young R. 2019. Phage single-gene lysis: finding the weak spot in the bacterial cell wall. J Biol Chem 294:3350–3358. doi: 10.1074/jbc.TM118.001773 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Bernhardt TG, Wang IN, Struck DK, Young R. 2002. Breaking free: "protein antibiotics" and phage lysis. Res Microbiol 153:493–501. doi: 10.1016/s0923-2508(02)01330-x [DOI] [PubMed] [Google Scholar]
- 5. Fischetti VA. 2008. Bacteriophage lysins as effective antibacterials. Curr Opin Microbiol 11:393–400. doi: 10.1016/j.mib.2008.09.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Atkins JF, Steitz JA, Anderson CW, Model P. 1979. Binding of mammalian ribosomes to MS2 phage RNA reveals an overlapping gene encoding a lysis function. Cell 18:247–256. doi: 10.1016/0092-8674(79)90044-8 [DOI] [PubMed] [Google Scholar]
- 7. Berkhout B, de Smit MH, Spanjaard RA, Blom T, van Duin J. 1985. The amino terminal half of the MS2-coded lysis protein is dispensable for function: implications for our understanding of coding region overlaps. EMBO J 4:3315–3320. doi: 10.1002/j.1460-2075.1985.tb04082.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Chamakura KR, Edwards GB, Young R. 2017. Mutational analysis of the MS2 lysis protein L. Microbiology (Reading) 163:961–969. doi: 10.1099/mic.0.000485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Chamakura KR, Tran JS, Young R. 2017. MS2 lysis of Escherichia coli depends on host chaperone DnaJ. J Bacteriol 199:00058–17. doi: 10.1128/JB.00058-17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Kannoly S, Shao Y, Wang IN.. 2012. Rethinking the evolution of single-stranded RNA (ssRNA) bacteriophages based on genomic sequences and characterizations of two R-plasmid-dependent ssRNA phages, C-1 and Hgal1. J Bacteriol 194:5073-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Rumnieks J, Tars K. 2012. Diversity of pili-specific bacteriophages: genome sequence of IncM plasmid-dependent RNA phage M. BMC Microbiol 12:277. doi: 10.1186/1471-2180-12-277 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Ruokoranta TM, Grahn AM, Ravantti JJ, Poranen MM, Bamford DH. 2006. Complete genome sequence of the broad host range single-stranded RNA phage PRR1 places it in the Levivirus genus with characteristics shared with alloleviviruses. J Virol 80:9326–9330. doi: 10.1128/JVI.01005-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Bradley DE, Dewar CA, Robertson D. 1966. The structure and infective process of a Pseudomonas aeruginosa bacteriophage containing ribonucleic acid. J Gen Microbiol 45:83–96. doi: 10.1099/00221287-45-1-83 [DOI] [Google Scholar]
- 14. Olsthoorn RC, Garde G, Dayhuff T, Atkins JF, Van Duin J. 1995. Nucleotide sequence of a single-stranded RNA phage from Pseudomonas aeruginosa: kinship to coliphages and conservation of regulatory RNA structures. Virology 206:611–625. doi: 10.1016/s0042-6822(95)80078-6 [DOI] [PubMed] [Google Scholar]
- 15. Klovins J, Overbeek GP, van den Worm SHE, Ackermann H-W, van Duin J. 2002. Nucleotide sequence of a ssRNA phage from Acinetobacter: kinship to coliphages. J Gen Virol 83:1523–1533. doi: 10.1099/0022-1317-83-6-1523 [DOI] [PubMed] [Google Scholar]
- 16. Kazaks A, Voronkova T, Rumnieks J, Dishlers A, Tars K. 2011. Genome structure of Caulobacter phage phiCb5. J Virol 85:4628–4631. doi: 10.1128/JVI.02256-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. SCHMIDT JM, STANIER RY. 1965. Isolation and characterization of bacteriophages active against stalked bacteria. J Gen Microbiol 39:95–107. doi: 10.1099/00221287-39-1-95 [DOI] [PubMed] [Google Scholar]
- 18. Reed CA, Langlais C, Kuznetsov V, Young R. 2012. Inhibitory mechanism of the Qβ lysis protein A2. Mol Microbiol 86:836–844. doi: 10.1111/mmi.12021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Chamakura KR, Sham L-T, Davis RM, Min L, Cho H, Ruiz N, Bernhardt TG, Young R. 2017. A viral protein antibiotic inhibits lipid II flippase activity. Nat Microbiol 2:1480–1484. doi: 10.1038/s41564-017-0023-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Vollmer W, Blanot D, de Pedro MA. 2008. Peptidoglycan structure and architecture. FEMS Microbiol Rev 32:149–167. doi: 10.1111/j.1574-6976.2007.00094.x [DOI] [PubMed] [Google Scholar]
- 21. Silhavy TJ, Kahne D, Walker S. 2010. The bacterial cell envelope. Cold Spring Harb Perspect Biol 2:a000414. doi: 10.1101/cshperspect.a000414 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Adler BA, Chamakura K, Carion H, Krog J, Deutschbauer AM, Young RF, Mutalik VK, Arkin AP. 2023. Parallel multicopy-suppressor screens reveal convergent evolution of phage-encoded single gene lysis proteins. Nat Chem Biol 19:759–766. doi: 10.1038/s41589-023-01269-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Bernhardt TG, Wang IN, Struck DK, Young R. 2001. A protein antibiotic in the phage Qβ virion: diversity in lysis targets. Science 292:2326–2329. doi: 10.1126/science.1058289 [DOI] [PubMed] [Google Scholar]
- 24. Holtje J, van Duin J. 1984. MS2 phage induced lysis of E. coli depends upon the activity of the bacterial autolysins. Elsevier Science Publishers, New York, NY. [Google Scholar]
- 25. Walderich B, Höltje JV. 1989. Specific localization of the lysis protein of bacteriophage MS2 in membrane adhesion sites of Escherichia coli. J Bacteriol 171:3331–3336. doi: 10.1128/jb.171.6.3331-3336.1989 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Goessens WH, Driessen AJ, Wilschut J, van Duin J. 1988. A synthetic peptide corresponding to the C-terminal 25 residues of phage MS2 coded lysis protein dissipates the protonmotive force in Escherichia coli membrane vesicles by generating hydrophilic pores. EMBO J 7:867–873. doi: 10.1002/j.1460-2075.1988.tb02886.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Mezhyrova J, Martin J, Börnsen C, Dötsch V, Frangakis AS, Morgner N, Bernhard F. 2023. in vitro characterization of the phage lysis protein MS2-L. Microbiome Res Rep 2:28. doi: 10.20517/mrr.2023.28 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Chamakura KR, Tran JS, O’Leary C, Lisciandro HG, Antillon SF, Garza KD, Tran E, Min L, Young R. 2020. Rapid de novo evolution of lysis genes in single-stranded RNA phages. Nat Commun 11:6009. doi: 10.1038/s41467-020-19860-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Starr EP, Nuccio EE, Pett-Ridge J, Banfield JF, Firestone MK. 2019. Metatranscriptomic reconstruction reveals RNA viruses with the potential to shape carbon cycling in soil. Proc Natl Acad Sci USA 116:25900–25908. doi: 10.1073/pnas.1908291116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Callanan J, Stockdale SR, Shkoporov A, Draper LA, Ross RP, Hill C. 2020. Expansion of known ssRNA phage genomes: from tens to over a thousand. Sci Adv 6:eaay5981. doi: 10.1126/sciadv.aay5981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Neri U, Wolf YI, Roux S, Camargo AP, Lee B, Kazlauskas D, Chen IM, Ivanova N, Zeigler Allen L, Paez-Espino D, Bryant DA, Bhaya D, Krupovic M, Dolja VV, Kyrpides NC, Koonin EV, Gophna U, RNA Virus Discovery Consortium . 2022. Expansion of the global RNA virome reveals diverse clades of bacteriophages. Cell 185:4023–4037. doi: 10.1016/j.cell.2022.08.023 [DOI] [PubMed] [Google Scholar]
- 32. Domingo E, Holland JJ. 1997. RNA virus mutations and fitness for survival. Annu Rev Microbiol 51:151–178. doi: 10.1146/annurev.micro.51.1.151 [DOI] [PubMed] [Google Scholar]
- 33. Sanger F, Air GM, Barrell BG, Brown NL, Coulson AR, Fiddes CA, Hutchison CA, Slocombe PM, Smith M. 1977. Nucleotide sequence of bacteriophage Φx174 DNA. Nature 265:687–695. doi: 10.1038/265687a0 [DOI] [PubMed] [Google Scholar]
- 34. Roof WD, Horne SM, Young KD, Young R. 1994. Slyd, a host gene required for Φx174 lysis, is related to the FK506-binding protein family of peptidyl-prolyl cis-trans-isomerases. J Biol Chem 269:2902–2910. [PubMed] [Google Scholar]
- 35. Karnik S, Billeter M. 1983. The lysis function of RNA bacteriophage Qβ is mediated by the maturation (A2) protein. EMBO J 2:1521–1526. doi: 10.1002/j.1460-2075.1983.tb01617.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Winter RB, Gold L. 1983. Overproduction of bacteriophage Qβ maturation (A2) protein leads to cell lysis. Cell 33:877–885. doi: 10.1016/0092-8674(83)90030-2 [DOI] [PubMed] [Google Scholar]
- 37. Model P, Webster RE, Zinder ND. 1979. Characterization of Op3, a lysis-defective mutant of bacteriophage F2. Cell 18:235–246. doi: 10.1016/0092-8674(79)90043-6 [DOI] [PubMed] [Google Scholar]
- 38. Beremand MN, Blumenthal T. 1979. Overlapping genes in RNA phage: a new protein implicated in lysis. Cell 18:257–266. doi: 10.1016/0092-8674(79)90045-x [DOI] [PubMed] [Google Scholar]
- 39. Groeneveld H, Oudot F, van Duin JV. 1996. RNA phage KU1 has an insertion of 18 nucleotides in the start codon of its lysis gene. Virology 218:141–147. doi: 10.1006/viro.1996.0174 [DOI] [PubMed] [Google Scholar]
- 40. Pourcel C, Midoux C, Vergnaud G, Latino L. 2017. A carrier state is established in Pseudomonas aeruginosa by phage LeviOr01, a newly isolated ssRNA levivirus. J Gen Virol 98:2181–2189. doi: 10.1099/jgv.0.000883 [DOI] [PubMed] [Google Scholar]
- 41. Bernhardt TG, Struck DK, Young R. 2001. The lysis protein E of Φx174 is a specific inhibitor of the MraY-catalyzed step in peptidoglycan synthesis. J Biol Chem 276:6093–6097. doi: 10.1074/jbc.M007638200 [DOI] [PubMed] [Google Scholar]
- 42. Paliy O, Gunasekera TS. 2007. Growth of E. coli Bl21 in minimal media with different gluconeogenic carbon sources and salt contents. Appl Microbiol Biotechnol 73:1169–1172. doi: 10.1007/s00253-006-0554-8 [DOI] [PubMed] [Google Scholar]
- 43. Thakur CS, Brown ME, Sama JN, Jackson ME, Dayie TK. 2010. Growth of wildtype and mutant E. coli strains in minimal media for optimal production of nucleic acids for preparing labeled nucleotides. Appl Microbiol Biotechnol 88:771–779. doi: 10.1007/s00253-010-2813-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Bremer H, Churchward G, Young R. 1979. Relation between growth and replication in bacteria. J Theor Biol 81:533–545. doi: 10.1016/0022-5193(79)90051-1 [DOI] [PubMed] [Google Scholar]
- 45. Lederberg J, St Clair J. 1958. Protoplasts and L-type growth of Escherichia coli. J Bacteriol 75:143–160. doi: 10.1128/jb.75.2.143-160.1958 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. CIAK J, HAHN FE. 1957. Penicillin-induced lysis of Escherichia coli. Science 125:119–120. doi: 10.1126/science.125.3238.119 [DOI] [PubMed] [Google Scholar]
- 47. Rohs PDA, Bernhardt TG. 2021. Growth and division of the peptidoglycan matrix. Annu Rev Microbiol 75:315–336. doi: 10.1146/annurev-micro-020518-120056 [DOI] [PubMed] [Google Scholar]
- 48. Bradley DE, Dewar CA, Robertson D. 1969. Structural changes in Escherichia coli infected with a ΦX174 type bacteriophage. J Gen Virol 5:113–121. doi: 10.1099/0022-1317-5-1-113 [DOI] [PubMed] [Google Scholar]
- 49. Henrich B, Lubitz W, Plapp R. 1982. Lysis of Escherichia coli by induction of cloned ΦX174 genes. Mol Gen Genet 185:493–497. doi: 10.1007/BF00334146 [DOI] [PubMed] [Google Scholar]
- 50. Young KD, Young R. 1982. Lytic action of cloned ΦX174 gene E. J Virol 44:993–1002. doi: 10.1128/JVI.44.3.993-1002.1982 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Bläsi U, Henrich B, Lubitz W. 1985. Lysis of Escherichia coli by cloned ΦX174 gene E depends on its expression. J Gen Microbiol 131:1107–1114. doi: 10.1099/00221287-131-5-1107 [DOI] [PubMed] [Google Scholar]
- 52. Brockhurst MA, Buckling A, Rainey PB. 2005. The effect of a bacteriophage on diversification of the opportunistic bacterial pathogen, Pseudomonas aeruginosa. Proc Biol Sci 272:1385–1391. doi: 10.1098/rspb.2005.3086 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Egan AJF, Errington J, Vollmer W. 2020. Regulation of peptidoglycan synthesis and remodelling. Nat Rev Microbiol 18:446–460. doi: 10.1038/s41579-020-0366-3 [DOI] [PubMed] [Google Scholar]
- 54. Miyake T, Yanagisawa K, Watanabe I. 1966. Alteration of a host character by infection with RNA phage Qβ. Jpn J Microbiol 10:141–148. doi: 10.1111/j.1348-0421.1966.tb00302.x [DOI] [PubMed] [Google Scholar]
- 55. Reed CA, Langlais C, Wang IN, Young R. 2013. A2 expression and assembly regulates lysis in Qβ infections. Microbiology (Reading) 159:507–514. doi: 10.1099/mic.0.064790-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Bernhardt TG, de Boer PAJ. 2003. The Escherichia coli amidase AmiC is a periplasmic septal ring component exported via the twin-arginine transport pathway. Mol Microbiol 48:1171–1182. doi: 10.1046/j.1365-2958.2003.03511.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Cho H, Uehara T, Bernhardt TG. 2014. Beta-lactam antibiotics induce a lethal malfunctioning of the bacterial cell wall synthesis machinery. Cell 159:1300–1311. doi: 10.1016/j.cell.2014.11.017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Guzman LM, Belin D, Carson MJ, Beckwith J. 1995. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J Bacteriol 177:4121–4130. doi: 10.1128/jb.177.14.4121-4130.1995 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Schneider CA, Rasband WS, Eliceiri KW. 2012. NIH image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675. doi: 10.1038/nmeth.2089 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figures S1 to S4.
Data Availability Statement
Raw data generated during the course of this study are available from the corresponding author upon request. The code used in this study to analyze the data in Fig. 5 is available from a public repository at https://github.com/AntillonF/bleb_statistical_analysis.git.