Abstract
Entomological sampling and storage conditions often prioritise efficiency, practicality and conservation of morphological characteristics, and may therefore be suboptimal for DNA preservation. This practice can impact downstream molecular applications, such as the generation of high-throughput genomic libraries, which often requires substantial DNA input amounts. Here, we use a practical Tn5 transposase tagmentation-based library preparation method optimised for 96-well plates and low yield DNA extracts from insect legs that were stored under sub-optimal conditions for DNA preservation. The samples were kept in field vehicles for extended periods of time, before long-term storage in ethanol in the freezer, or dry at room temperature. By reducing DNA input to 6ng, more samples with sub-optimal DNA yields could be processed. We matched this low DNA input with a 6-fold dilution of a commercially available tagmentation enzyme, significantly reducing library preparation costs. Costs and workload were further suppressed by direct post-amplification pooling of individual libraries. We generated medium coverage (>3-fold) genomes for 88 out of 90 specimens, with an average of approximately 10-fold coverage. While samples stored in ethanol yielded significantly less DNA compared to those which were stored dry, these samples had superior sequencing statistics, with longer sequencing reads and higher rates of endogenous DNA. Furthermore, we find that the efficiency of tagmentation-based library preparation can be improved by a thorough post-amplification bead clean-up which selects against both short and large DNA fragments. By opening opportunities for the use of sub-optimally preserved, low yield DNA extracts, we broaden the scope of whole genome studies of insect specimens. We therefore expect these results and this protocol to be valuable for a range of applications in the field of entomology.
Introduction
In response to widespread declines in insect diversity (reviewed in [1]), and the subsequent severe implications for ecosystem functioning (reviewed in [2]), large-scale international insect survey and sampling schemes have been put in place (e.g. [3]). While the primary interest of such schemes is to monitor abundance and distribution changes, the application of genomic approaches to insect specimens can also be used to investigate spatial drivers of population trends, habitat connectivity and local adaptation (reviewed in [4]). These approaches rely on high-throughput sequencing (HTS) methods, which generally have considerably greater DNA quality requirements compared to more traditional PCR-based methods. However, entomological samples are often collected and stored using a variety of techniques which form a trade-off between practicality, morphological requirements and molecular demands. While samples should ideally be stored immediately following tissue death under optimal conditions [5], this is not always feasible during extended field periods. For instance, entomological specimens are often temporarily stored in warm fieldwork vehicles until sampling has concluded, and those captured by high-efficiency trapping methods, such as pitfall and flight interception traps, can remain in traps on site for several days, or even weeks, prior to collection and processing [6]. In addition, as entomological specimens are important biological resources with myriad applications in the fields of taxonomy and ecology, standard collection and storage methods are designed to prioritise morphological preservation. Given these considerations, alcohols such as ethanol are commonly used as both killing agents and preserving fluids (e.g. [3]). While this approach helps to prevent physical deterioration, it does not fully inhibit DNA degradation due to relatively high water content [7, 8]. The use of highly concentrated ethanol (>96%) can improve DNA preservation, although ethanol concentration may decrease due to evaporative loss. In addition, entomological specimens preserved in high concentration ethanol can become overly brittle, impacting morphological analyses [9]. Ethanol-based sampling and storage nevertheless remains the cheapest and most practical method compared to other options such as flash freezing with liquid nitrogen, and is therefore widespread (e.g. [10–12]). Library preparation methods which are compatible with material collected and stored using suboptimal preservation protocols are therefore required when working with entomological specimens.
An additional obstacle to the use of entomological specimens for HTS is the destructive nature of DNA extraction: certain protocols require the sample be physically ground or beaten with beads, while most call for tissue digestion through the use of corrosive chemicals and enzymes (e.g. [13]). While it is possible to extract from the entire insect in order to prioritise DNA quantity and quality (see [14], this destructive approach requires the sacrifice of biologically valuable entomological specimens. Some ‘non-destructive’ insect DNA extraction methods have been proposed, whereby the entire specimen is incubated during the digestion step and subsequently retained, allowing for the preservation of the whole insect [15–18]. However, this practice may result in some morphological damage such as pigment loss [16, 19], and its impacts on long-term preservation (i.e. >10 years) remain unclear. Importantly, while re-extractions of the same biological material may produce viable genomic DNA extracts, the first extraction is often the most effective in terms of DNA yield and endogenous DNA content [20]. Using the entire specimen therefore limits any future DNA applications, reduces the long-term biological value of entomological collections, and fails to incorporate ethical guidelines for the destructive sampling of biological specimens [21]. To most effectively preserve insect specimens, for both future entomological applications and future DNA extractions, a more conservative solution would be to sacrifice a small piece of tissue such as a leg [20]. This approach leaves the majority of the insect untouched, although the low sample volume and consequent low DNA yield may constrain specific downstream molecular applications.
While the cost of whole genome sequencing has decreased by several orders of magnitude since its inception [22], library preparation often remains expensive and now represents a substantial proportion of the overall cost of whole genome data generation. For instance, methods whereby DNA is physically fragmented are widely adopted, yet can be expensive, time consuming, and often require high DNA input [23]. Alternatively, tagmentation-based library preparation has streamlined the process by combining DNA fragmentation and adapter ligation into a single reaction facilitated by Tn5 transposase, and typically calls for lower DNA input [24, 25]. Commercial Tn5 transposase kits, however, remain relatively expensive, and their dependence on the use of undisclosed reagents inhibits methodological optimisation. Several adaptations to the tagmentation method have been developed, such as the in-house production of Tn5 transposase [26–28]. Despite significantly reducing costs, in-house enzyme production can be both time and resource intensive, may not be always possible and therefore lacks broad-scale practicality. Further modifications include reduction of reaction volumes, replacement of reagents with cheaper alternatives and the incorporation of magnetic bead-linked transposomes, highlighting the potential of matching low DNA input amounts with diluted transposase concentrations to increase both throughput and affordability [25, 29, 30]. Here, we explore the use of a commercially available hyperactive Tn5 transposase (Diagenode) to create high-throughput sequencing libraries from low-yield DNA extracts obtained from entomological specimens.
Specifically, we investigate the effectiveness of a practical and in-house developed Tn5 tagmentation-based library preparation method, using insect DNA extracted by applying minimally destructive methods to samples initially stored in ethanol for several weeks while kept in fieldwork vehicles at ambient summer temperatures. After fieldwork was completed, single legs were removed from the specimens and subsequently stored in 96% ethanol at -18°C, while the remaining specimens were dried, pinned and stored at room temperature. Our study utilises samples stored using both methods. We assessed the robustness of the protocol in response to varying tagmentation reaction time and Tn5 transposase enzyme concentration. We then rescaled our protocol to fit 96-well plates and a standardised DNA input of 6ng. Finally, we developed a double-sided post-amplification clean-up protocol in order to optimise library fragment size distribution and thus improve sequencing efficiency. We find that this economical, easily scalable library preparation protocol opens multiple opportunities for genomic studies, particularly in the cases of non-model organisms with limited funding, small organisms with low per-sample DNA yield or biological samples stored under non-optimal conditions for DNA preservation.
Materials and methods
Specimens, sampling and storage conditions
122 samples of two different bumblebee species (Bombus lapidarius and Bombus pascuorum) were stored following one of two protocols (see Table 1). All specimens were sampled in 2017 and 2022 by immediate submersion in 96% ethanol. They were then stored and transported in fieldwork vehicles for several weeks during the summer, before being dried, pinned and stored at room temperature with no preservative (Dry Protocol, n = 82). Samples stored using the Ethanol Protocol (n = 40) were comprised of single legs which were removed from the specimens collected in 2017 prior to drying, and subsequently stored in 96% ethanol at -18°C.
Table 1. Bumblebee specimen sampling and storage features.
Details of two bumblebee sample storage protocols, including year sampled, killing agent, storage preservative and storage temperature, and the number of legs used for DNA extraction. Also presented are the sample sizes from each group used in two different sets of analyses: DNA concentration comparison of DNA extracts, and library preparation and sequencing statistics.
| Protocol | Year sampled | Killing agent | Storage preservative | Temperature | Specimens (n) | One leg extracted (n) | Two legs extracted (n) | DNA concentration comparison (n) | Library preparation and sequencing (n) |
|---|---|---|---|---|---|---|---|---|---|
| Dry | 2017 / 2022 | Ethanol | None | Ambient | 82 | 20 | 62 | 20 | 82 |
| Ethanol | 2017 | Ethanol | Ethanol | -18°C | 40 | 40 | 0 | 40 | 8 |
DNA extraction
DNA was extracted from either 1 or 2 mid or hind legs, including the coxa, femur, tibia, and tarsomeres, using the Qiagen DNeasy Blood & Tissue Kit. Samples from the Ethanol Protocol were washed with molecular-grade water in order to remove any ethanol residue. All samples were incubated in 380μL digestion buffer (300μL ATL buffer, 50μL DTT, 30μL proteinase K) at 56° C and 350 RPM overnight (minimum 17 hours), as implemented by the Campos & Gilbert method (in [31]) for DNA extraction from chitin. Following overnight digestion, 7μL RNase A and 300μL AL buffer were added to the lysate, followed by a room temperature incubation (15 minutes) and the addition of 360 μL 96% ethanol. DNA was subsequently bound and purified following the manufacturer’s recommendations, then eluted in AE buffer (105μL, 90μL or 80μL) and stored at -18°C. DNA concentration was then estimated using the Qubit dsDNA Assays (Thermofisher) and standardised to 1ng/μL.
Initially, DNA extractions of dry-stored specimens were performed on single legs (n = 21). In order to consistently avoid very low DNA yields, two legs were used for the remaining 61 extractions, along with the addition of 5μL 1M CaCl2 solution to the digestion buffer to aid tissue digestion.
Library preparation
Two 8-sample Illumina compatible trial libraries were generated to evaluate the impact of two variables on DNA fragment size distribution: 1) tagmentation reaction time (Trial 1) and 2) Tn5 transposase enzyme concentration (Trial 2). For Trial 1, two B. pascuorum DNA extracts were incubated for five, six, seven or eight minutes with 1μL of undiluted Loaded Tn5 Tagmentase (Diagenode, C01070012). For trial two, the same DNA extracts were incubated with diluted Loaded Tn5 Tagmentase (either a 2X, 4X, 6X or 8X-fold dilution of the original Tn5 Tagmentase) for 7 minutes.
Following these trials, the protocol was adapted for a 96-well plate setup, and 96 libraries were generated using a 7 minute incubation and 6X Loaded Tn5 Tagmentase dilution. The libraries were built using extracts with DNA concentrations of at least 1ng/μL, consisting of 82 dry-stored samples, 8 ethanol-stored samples, and 6 additional samples which were either found to be misidentified bumblebee species, or stored using a different protocol. Following PCR amplification, the libraries were pooled and cleaned using two alternative clean-up protocols (Pool SP and Pool S4) in order to optimise fragment size distribution for sequencing (see also S1 Protocol for full Tn5 tagmentation library preparation, PCR and post-amplification clean-up protocol.).
Tn5 tagmentation library preparation and PCR
All DNA extracts were standardised to 1ng/μL. 6μL DNA extract were incubated (55°C) with 5μL of 2X Tagmentation Buffer (Diagenode, C01019043) and 1μL of Loaded Tn5 Tagmentase (Diagenode, C01070012, using various dilutions, see above). After 5, 6, 7 or 8 minutes (see above), 3μL 0.2% Sodium dodecyl sulfate (SDS) Solution (Molecular Biology/Electrophoresis, Fisher BioReagents™ MFCD00036175) was added to stop the tagmentation reaction, followed by a 5 minute room temperature incubation. Post-tagmentation PCR (12 cycles) was done in 40μL reactions using KAPA HiFi DNA Polymerase (HotStart and Ready Mix formulation, Roche, KK2601). The manufacturer’s recommendations were followed by adding 24μL Mastermix, 1μL N7 Primer (1μM) and 1μL N5 Primer (1μM) (IDT for Illumina UD Indexes Plate A/Set 1, Illumina) to each 15μL reaction. After amplification, 6μL of each of the 96 wells was pooled (without quantification) before proceeding to clean-up (AMPure XP, Beckman Coulter).
Post-amplification clean-up and sequencing
All bead clean-ups were performed using AMPure XP beads (Beckman Coulter). The Trial 1, Trial 2 and Pool SP libraries were cleaned once, removing small DNA fragments by using a 0.6 DNA/bead ratio, following the manufacturer’s recommendations. Pool S4 was built from the same PCR reaction as Pool SP, but cleaned twice, using different bead ratios to remove both large and small DNA fragments (as demonstrated in [32]), with a target median fragment size distribution of ~600bp. Firstly, a 0.5 DNA bead/ratio was used to remove large DNA fragments. Secondly, a 0.65 DNA/bead ratio was used to remove smaller DNA fragments and shift the fragment size distribution to peak between 500 and 1000 bp. This “double sided” clean-up process was performed twice, and the final library was eluted in 40μL molecular grade water. DNA fragment length for all amplified libraries (individual libraries for Trial 1 and Trial 2, Pool SP, and Pool S4) were analysed using a Fragment Analyzer™ (Advanced Analytical) with the DNF-474 High Sensitivity Fragment Analysis Kit after 10-fold dilution of library aliquots. Pool SP was sequenced using an Illumina Novaseq 6000 SP flowcell, generating 923,253,536 read pairs passing filters and Pool S4 was sequenced using an Illumina Novaseq 6000 S4 flowcell and generated 2,799,713,996 read pairs passing filters. Base calling and demultiplexing were performed with RTA v3.4.4 and bcl2fastq v2.20.0.422 in both cases.
Read data were aligned using PALEOMIX v1.2.14 [33] to BomLapEIv1 (https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_936014575.1/) and BomPasc1.1 [34] after using AdapterRemoval/2.3.1 [35] and the mem algorithm as implemented in BWA/0.7.17 [36]. Only reads with a minimum MapQ value of 15 were retained and considered endogenous.
Statistics
The DNA yield comparison between dry-stored and ethanol-stored specimens was carried out using a t-test. Mann-Whitney-Wilcoxon tests were used to compare sequencing statistics between a) dry-stored and ethanol-stored specimens and b) Pool SP and Pool S4, excluding the endogenous DNA comparison between Pool SP and Pool S4, for which a t-test was used. All statistical analyses were performed using base R 4.3.1.
Ethics statement
There are no legal limitations on sampling B. pascuorum and B. lapidarius in Scandinavia. Bees in Germany were collected under permit LLUR 515 from the Schleswig-Holstein Nature Conservation Bureau, issued to M. Kuhlmann.
Results
Sample storage impact on DNA yield
One leg was used to extract DNA from samples stored in ethanol at -18°C (n = 40) and either one leg (n = 20) or two legs (n = 62) were used from samples stored dried at ambient temperatures (Table 1). When comparing DNA yields of extracts generated from comparable tissue volumes (a single leg), we found that significantly more DNA is obtained from the dry-stored specimens than the ethanol-stored specimens (Fig 1). Given that DNA yields from ethanol-stored specimens were low, we chose to prioritise dry-stored specimens for library preparation.
Fig 1. DNA extract concentration measurements (ng) from single bumblebee legs (B. pascuorum and B. lapidarius) stored using 2 different methods.
Specimens were sampled in 2017 from various locations in Norway, Germany and Denmark and stored in ethanol in fieldwork vehicles at ambient summer temperatures for several weeks. Samples were then either stored dry (at room temperature) or in ethanol (at -18°C). See Materials and Methods for details regarding sample collection and DNA extraction. Following extraction, DNA was eluted in 105μL. DNA yields from dry-stored specimens (mean = 298.21±181.03) were significantly greater than from those stored in ethanol (mean = 72.98±51.79; t = 5.45, df = 20.57, P < 0.001).
Tn5 transposase library preparation optimisation
Changing tagmentation reaction time had minimal impact on fragment size distribution, with all libraries showing similar distribution profiles and fragment size peaking slightly above 300 bp (see Fig 2A). Fragmentation size distribution profiles of libraries built using 2X, 4X and 6X transposase enzyme dilutions were broader, and showed a modest increase in the proportion of long DNA fragments with each ascending dilution. This increase becomes more pronounced with the use of an 8X dilution (see Fig 2B). We therefore generated libraries in a 96-well plate using a 7 minute incubation time and 6X Loaded Tn5 Tagmentase dilution. These libraries were then amplified (12 cycles), after which 5μL from each library was pooled to perform bead clean-up on all libraries combined using two different approaches.
Fig 2. Fragment size distribution profiles of DNA libraries built from a standardised input of 6ng DNA from each of two B. pascuorum specimens, using different tagmentation reaction times and transposase dilutions.
(a) Libraries were built using four different tagmentation reaction times (5 minutes; 6 minutes; 7 minutes; 8 minutes) with a 1X transposase dilution. (b) Libraries were built using four different transposase dilutions (2X; 4X; 6X; 8X) with a 7 minute tagmentation reaction. RFU (Relative Fluorescence Units) signifies relative amount of DNA. A vertical line indicating a fragment size of 300 bp is included to assist visual comparison of the different treatments. See Materials and Methods for details regarding library preparation protocol.
Post-amplification clean-up optimisation
Following initial bead clean-up, we obtained a fragment size distribution situated between 400 bp and 1500 bp (Pool SP, Fig 3), a much broader range than the tight distribution around 600 bp recommended for Illumina DNA sequencing [37]. Pool SP was sequenced on an Illumina Novaseq SP flowcell with paired-end 150 bp reads, with a large number of reads collapsing into a single read (see further below). We therefore performed a more stringent clean-up on the original pooled libraries to tighten fragment size distribution, removing both very short and very long DNA fragments, resulting in a narrow distribution peaking around 600 bp (Pool S4, Fig 3). Pool S4 was subsequently sequenced on one quarter of an Illumina 6000 S4 flowcell, also with paired-end 150 bp reads.
Fig 3. Comparison of fragment size distribution of 96 pooled bumblebee DNA libraries which were cleaned up with different AMPure XP protocols: A single clean-up with a 0.6 DNA/bead ratio (Pool SP), and a double-sided clean-up using two ratios (0.5 DNA/bead ratio followed by 0.65 DNA/bead ratio) which was performed twice (Pool S4).
RFU (Relative Fluorescence Units) signifies relative amount of DNA.
Sequencing results
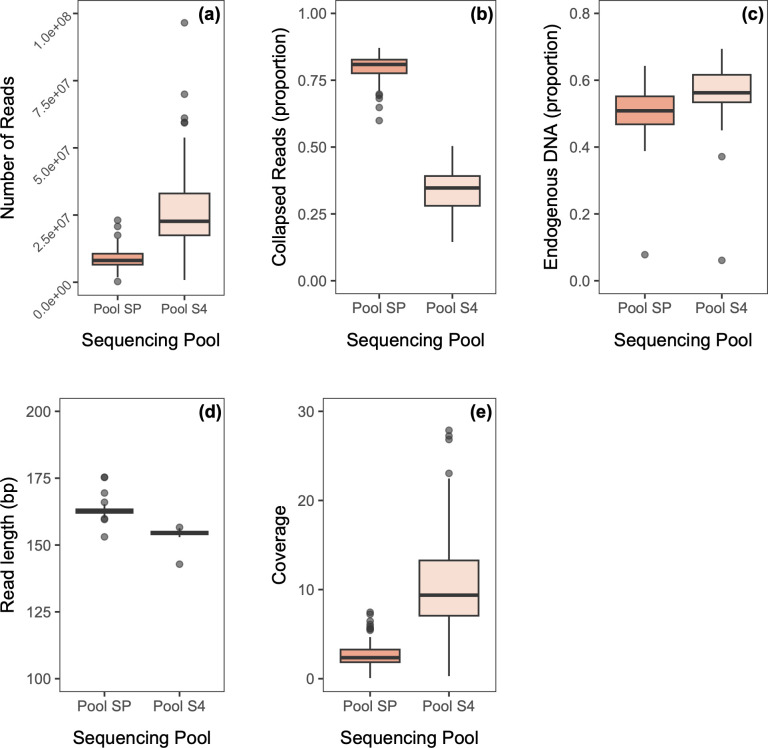
We obtained 795,694,049 reads ranging between 261,576 and 23,087,451 per specimen for Pool SP, while Pool S4 produced significantly more reads: 2,409,119,984, ranging between 835,001 and 96,527,020 per specimen (W = 429, P = <0.001; Table 2 and Fig 4A). Furthermore, Pool SP contained a significantly higher number of collapsed reads (W = 8100, P = <0.001; Fig 4B), slightly lower endogenous DNA proportion (t = -25.89, df = 89, P = <0.001; Fig 4C), and slightly higher overall read length (W = 8012, P = <0.001; Fig 4D) compared to Pool S4. Overall, Pool S4 resulted in higher coverage than Pool SP (W = 365, P = <0.001; Fig 4E), which was predominantly driven by the higher sequencing yield (Fig 4A). Nonetheless, when calculating the ratio of coverage against sequencing yield (calculated as coverage divided by number of paired reads), the coverage achieved by Pool S4 was on average greater than that of Pool SP by a factor of 1.45±0.12. This increase in efficiency for Pool S4 can be explained by both its reduced proportion of collapsed reads, leading to the mapping of more nucleotides per pair, and its higher proportion of endogenous DNA, leading to more unique mapped sequences per pair.
Table 2. a & b. Sequencing statistics for two sequencing pools (SP and S4) of 90 specimens of two bumblebee species: (a) Bombus lapidarius (n = 46) and (b) Bombus pascuorum (n = 44).
Bumblebee specimens were sampled between 2017 and 2022 from various locations in Norway, Germany, Denmark and Sweden. Libraries for both pools were generated simultaneously. The SP pool was cleaned up with a 0.6 DNA/bead ratio and was sequenced on an Illumina Novaseq SP flowcell. The S4 pool was cleaned up using a double-sided size selection protocol, with a 0.5 DNA/bead ratio clean-up followed by a 0.65 DNA/bead ratio clean-up (twice) and was sequenced on ¼ Illumina Novaseq S4 flowcell. Statistics presented for each sequencing pool are specimen ID, number of reads, collapsed reads (proportion), clonality (proportion), endogenous DNA (proportion), nuclear coverage (fold), and read length (bp). Also presented is combined per-specimen coverage of both sequencing runs.
| (a) | |||||||||||||
| Bombus lapidarius | |||||||||||||
| ID | SP | S4 |
Combined
coverage (fold) |
||||||||||
| Reads |
Collapsed reads
(proportion) |
Clonality
(proportion) |
Endogenous DNA
(proportion) |
Coverage
(fold) |
Read length
(bp) |
Reads |
Collapsed reads
(proportion) |
Clonality (proportion) |
Endogenous DNA
(proportion) |
Coverage
(fold) |
Read length
(bp) |
||
| BEE009 | 9974301 | 0,871 | 0,066 | 0,470 | 2,397 | 160,435 | 19387911 | 0,329 | 0,117 | 0,536 | 7,659 | 156,283 | 10,056 |
| BEE010 | 9973152 | 0,842 | 0,071 | 0,470 | 2,458 | 160,446 | 25403974 | 0,377 | 0,131 | 0,514 | 9,263 | 154,988 | 11,721 |
| BEE014 | 8076852 | 0,831 | 0,067 | 0,481 | 2,092 | 163,328 | 20241708 | 0,326 | 0,121 | 0,550 | 8,167 | 155,345 | 10,259 |
| BEE016 | 12203996 | 0,829 | 0,071 | 0,479 | 3,146 | 162,998 | 28735826 | 0,328 | 0,124 | 0,545 | 11,470 | 155,234 | 14,616 |
| BEE017 | 9640662 | 0,838 | 0,072 | 0,460 | 2,362 | 162,387 | 22589207 | 0,356 | 0,129 | 0,509 | 8,263 | 155,072 | 10,625 |
| BEE018 | 7609357 | 0,860 | 0,067 | 0,478 | 1,889 | 161,304 | 16140277 | 0,361 | 0,117 | 0,550 | 6,398 | 155,828 | 8,287 |
| BEE019 | 12168745 | 0,803 | 0,067 | 0,452 | 3,039 | 163,656 | 36439272 | 0,368 | 0,124 | 0,506 | 13,112 | 154,294 | 16,151 |
| BEE021 | 16261719 | 0,726 | 0,054 | 0,588 | 5,604 | 162,840 | 53827117 | 0,205 | 0,113 | 0,641 | 26,837 | 153,707 | 32,441 |
| BEE025 | 23087451 | 0,740 | 0,063 | 0,556 | 7,459 | 163,406 | 96527020 | 0,241 | 0,147 | 0,612 | 45,061 | 153,617 | 52,520 |
| BEE027 | 14102012 | 0,796 | 0,066 | 0,587 | 4,514 | 160,541 | 34124445 | 0,189 | 0,120 | 0,639 | 17,289 | 155,188 | 21,803 |
| BEE032 | 13389169 | 0,847 | 0,071 | 0,465 | 3,276 | 161,618 | 23670306 | 0,314 | 0,120 | 0,534 | 9,339 | 155,415 | 12,615 |
| BEE034 | 20745911 | 0,749 | 0,076 | 0,539 | 6,463 | 163,746 | 69948180 | 0,215 | 0,169 | 0,589 | 31,912 | 153,800 | 38,375 |
| BEE037 | 12311833 | 0,785 | 0,055 | 0,566 | 3,878 | 162,403 | 31316323 | 0,203 | 0,106 | 0,620 | 15,312 | 155,496 | 19,190 |
| BEE038 | 6629450 | 0,828 | 0,066 | 0,436 | 1,564 | 163,612 | 20515904 | 0,459 | 0,122 | 0,473 | 6,519 | 154,457 | 8,083 |
| BEE039 | 9868829 | 0,811 | 0,077 | 0,428 | 2,317 | 163,437 | 27932752 | 0,394 | 0,133 | 0,482 | 9,416 | 154,365 | 11,732 |
| BEE040 | 13433561 | 0,818 | 0,072 | 0,506 | 3,686 | 162,638 | 28159433 | 0,276 | 0,123 | 0,585 | 12,419 | 154,880 | 16,105 |
| BEE041 | 10106362 | 0,813 | 0,083 | 0,448 | 2,489 | 164,142 | 43363140 | 0,435 | 0,161 | 0,489 | 14,436 | 154,196 | 16,924 |
| BEE043 | 5924824 | 0,833 | 0,068 | 0,461 | 1,446 | 160,913 | 16846127 | 0,455 | 0,121 | 0,493 | 5,605 | 154,640 | 7,051 |
| BEE047 | 14470649 | 0,718 | 0,070 | 0,643 | 5,430 | 161,472 | 42097600 | 0,167 | 0,123 | 0,670 | 22,463 | 153,943 | 27,893 |
| BEE165 | 11932018 | 0,742 | 0,067 | 0,527 | 3,655 | 163,741 | 47903029 | 0,280 | 0,144 | 0,562 | 20,086 | 153,714 | 23,741 |
| BEE166 | 10186140 | 0,774 | 0,061 | 0,541 | 3,070 | 161,137 | 26039416 | 0,268 | 0,097 | 0,616 | 12,096 | 154,291 | 15,166 |
| BEE168 | 10210866 | 0,736 | 0,057 | 0,442 | 2,619 | 162,861 | 36326853 | 0,222 | 0,108 | 0,473 | 13,270 | 153,865 | 15,889 |
| BEE178 | 13903566 | 0,741 | 0,060 | 0,492 | 3,941 | 162,279 | 59516346 | 0,388 | 0,126 | 0,541 | 22,442 | 153,188 | 26,383 |
| BEE254 | 8588713 | 0,821 | 0,065 | 0,462 | 2,139 | 162,012 | 24208707 | 0,373 | 0,112 | 0,522 | 8,975 | 154,743 | 11,114 |
| BEE255 | 8851361 | 0,820 | 0,063 | 0,474 | 2,280 | 163,271 | 25350112 | 0,361 | 0,113 | 0,538 | 9,781 | 155,012 | 12,062 |
| BEE256 | 7091677 | 0,820 | 0,068 | 0,454 | 1,753 | 163,622 | 22236744 | 0,435 | 0,123 | 0,500 | 7,595 | 154,543 | 9,348 |
| BEE257 | 2427508 | 0,770 | 0,084 | 0,497 | 0,641 | 153,048 | 8916584 | 0,391 | 0,111 | 0,569 | 3,292 | 142,826 | 3,933 |
| BEE258 | 8132019 | 0,824 | 0,061 | 0,477 | 2,097 | 162,749 | 30014805 | 0,302 | 0,114 | 0,538 | 12,016 | 155,311 | 14,113 |
| BEE259 | 7600369 | 0,814 | 0,080 | 0,455 | 1,885 | 162,846 | 22660080 | 0,396 | 0,145 | 0,498 | 7,892 | 154,618 | 9,778 |
| BEE260 | 7354599 | 0,682 | 0,081 | 0,078 | 0,341 | 159,535 | 25988804 | 0,145 | 0,140 | 0,061 | 1,280 | 153,194 | 1,621 |
| BEE261 | 14375044 | 0,818 | 0,074 | 0,474 | 3,715 | 163,360 | 37035119 | 0,352 | 0,131 | 0,547 | 14,558 | 154,671 | 18,273 |
| BEE263 | 10276519 | 0,803 | 0,059 | 0,462 | 2,616 | 162,954 | 28463838 | 0,369 | 0,103 | 0,534 | 10,798 | 154,208 | 13,414 |
| BEE264 | 7120018 | 0,831 | 0,066 | 0,462 | 1,768 | 163,087 | 18442004 | 0,436 | 0,111 | 0,509 | 6,408 | 154,634 | 8,176 |
| BEE265 | 5453553 | 0,823 | 0,062 | 0,468 | 1,384 | 163,422 | 16730402 | 0,460 | 0,109 | 0,511 | 5,740 | 154,453 | 7,124 |
| BEE266 | 6616852 | 0,809 | 0,070 | 0,420 | 1,525 | 163,393 | 17800519 | 0,409 | 0,120 | 0,450 | 5,556 | 154,502 | 7,081 |
| BEE267 | 5917461 | 0,819 | 0,058 | 0,468 | 1,499 | 162,297 | 17694985 | 0,344 | 0,092 | 0,551 | 7,066 | 155,077 | 8,565 |
| BEE268 | 6664674 | 0,807 | 0,056 | 0,501 | 1,838 | 163,453 | 19275461 | 0,381 | 0,104 | 0,550 | 7,491 | 154,678 | 9,330 |
| BEE269 | 7465422 | 0,810 | 0,066 | 0,470 | 1,921 | 163,122 | 17086497 | 0,392 | 0,106 | 0,527 | 6,306 | 154,312 | 8,227 |
| BEE270 | 261576 | 0,798 | 0,056 | 0,455 | 0,067 | 165,995 | 835001 | 0,440 | 0,078 | 0,505 | 0,288 | 155,124 | 0,355 |
| BEE271 | 9085590 | 0,836 | 0,072 | 0,463 | 2,243 | 162,363 | 19003806 | 0,358 | 0,118 | 0,537 | 7,332 | 155,177 | 9,575 |
| BEE272 | 2456658 | 0,804 | 0,057 | 0,466 | 0,634 | 164,139 | 7233437 | 0,452 | 0,090 | 0,511 | 2,495 | 154,413 | 3,129 |
| BEE273 | 5807791 | 0,798 | 0,057 | 0,430 | 1,385 | 163,551 | 20537471 | 0,504 | 0,109 | 0,463 | 6,164 | 153,630 | 7,548 |
| BEE274 | 6422043 | 0,831 | 0,055 | 0,489 | 1,682 | 162,351 | 13546114 | 0,323 | 0,092 | 0,564 | 5,616 | 155,304 | 7,298 |
| BEE275 | 11070844 | 0,820 | 0,066 | 0,468 | 2,813 | 163,006 | 5837269 | 0,402 | 0,086 | 0,544 | 2,212 | 154,424 | 5,025 |
| BEE276 | 7392200 | 0,809 | 0,063 | 0,480 | 1,944 | 163,083 | 22690095 | 0,358 | 0,117 | 0,541 | 8,786 | 154,502 | 10,730 |
| BEE277 | 5956503 | 0,836 | 0,057 | 0,447 | 1,422 | 162,493 | 17784587 | 0,463 | 0,097 | 0,502 | 5,984 | 154,540 | 7,406 |
| (b) | |||||||||||||
| Bombus pascuorum | |||||||||||||
| ID | SP | S4 | Combined coverage (fold) | ||||||||||
| Reads | Collapsed reads (proportion) | Clonality (proportion) | Endogenous DNA (proportion) | Coverage (fold) | Read length (bp) | Reads | Collapsed reads (proportion) | Clonality (proportion) | Endogenous DNA (proportion) | Coverage (fold) | Read length (bp) | ||
| BEE008 | 6549854 | 0,862 | 0,056 | 0,574 | 2,230 | 160,364 | 14692031 | 0,289 | 0,108 | 0,657 | 8,376 | 156,090 | 10,605 |
| BEE011 | 9454360 | 0,811 | 0,066 | 0,547 | 3,251 | 162,735 | 28641027 | 0,338 | 0,116 | 0,625 | 14,955 | 154,549 | 18,207 |
| BEE012 | 1842509 | 0,648 | 0,039 | 0,560 | 0,796 | 175,369 | 12755700 | 0,316 | 0,106 | 0,537 | 5,768 | 153,834 | 6,564 |
| BEE013 | 9979839 | 0,855 | 0,072 | 0,604 | 3,597 | 160,225 | 19615704 | 0,256 | 0,116 | 0,694 | 12,048 | 156,115 | 15,645 |
| BEE015 | 6242105 | 0,847 | 0,056 | 0,620 | 2,348 | 161,792 | 15278897 | 0,255 | 0,109 | 0,693 | 9,411 | 156,607 | 11,759 |
| BEE020 | 17478850 | 0,731 | 0,064 | 0,620 | 7,235 | 161,673 | 59277548 | 0,195 | 0,127 | 0,665 | 35,583 | 153,820 | 42,818 |
| BEE022 | 13710915 | 0,698 | 0,056 | 0,595 | 5,601 | 162,110 | 61087559 | 0,185 | 0,126 | 0,634 | 35,015 | 153,281 | 40,615 |
| BEE023 | 6881537 | 0,796 | 0,080 | 0,532 | 2,338 | 163,275 | 20380076 | 0,338 | 0,130 | 0,598 | 10,186 | 154,551 | 12,524 |
| BEE024 | 6102715 | 0,774 | 0,050 | 0,600 | 2,476 | 169,461 | 18644941 | 0,230 | 0,101 | 0,670 | 11,108 | 154,536 | 13,583 |
| BEE028 | 6652996 | 0,862 | 0,071 | 0,528 | 2,076 | 159,861 | 14249671 | 0,359 | 0,116 | 0,604 | 7,161 | 155,907 | 9,238 |
| BEE031 | 10012093 | 0,793 | 0,063 | 0,508 | 3,255 | 162,919 | 29458652 | 0,294 | 0,120 | 0,567 | 14,298 | 154,369 | 17,553 |
| BEE033 | 6172151 | 0,850 | 0,063 | 0,560 | 2,084 | 161,214 | 13102527 | 0,304 | 0,103 | 0,650 | 7,324 | 155,882 | 9,408 |
| BEE035 | 4694766 | 0,809 | 0,066 | 0,504 | 1,504 | 164,292 | 17189720 | 0,422 | 0,127 | 0,542 | 7,365 | 154,092 | 8,870 |
| BEE036 | 7118608 | 0,796 | 0,064 | 0,502 | 2,285 | 163,293 | 26096664 | 0,347 | 0,117 | 0,565 | 12,219 | 154,174 | 14,504 |
| BEE042 | 7127099 | 0,794 | 0,062 | 0,527 | 2,391 | 162,470 | 24946535 | 0,388 | 0,111 | 0,591 | 11,895 | 154,038 | 14,286 |
| BEE044 | 10297225 | 0,802 | 0,062 | 0,574 | 3,711 | 161,244 | 24511141 | 0,287 | 0,103 | 0,653 | 13,801 | 154,889 | 17,512 |
| BEE045 | 6629392 | 0,842 | 0,055 | 0,534 | 2,169 | 162,890 | 18638735 | 0,348 | 0,111 | 0,603 | 9,375 | 155,248 | 11,544 |
| BEE046 | 9300192 | 0,771 | 0,066 | 0,515 | 3,112 | 162,764 | 36373177 | 0,412 | 0,121 | 0,563 | 16,234 | 153,618 | 19,346 |
| BEE048 | 11154948 | 0,782 | 0,059 | 0,553 | 3,962 | 162,237 | 34077198 | 0,282 | 0,107 | 0,628 | 18,432 | 154,256 | 22,394 |
| BEE049 | 8614332 | 0,790 | 0,064 | 0,530 | 2,936 | 163,321 | 26850114 | 0,340 | 0,114 | 0,590 | 13,188 | 154,188 | 16,124 |
| BEE152 | 14781975 | 0,726 | 0,060 | 0,609 | 6,077 | 162,831 | 53816848 | 0,209 | 0,127 | 0,658 | 31,612 | 153,303 | 37,689 |
| BEE153 | 9557459 | 0,744 | 0,059 | 0,543 | 3,439 | 162,238 | 38876394 | 0,327 | 0,111 | 0,602 | 19,513 | 153,380 | 22,952 |
| BEE163 | 11101633 | 0,724 | 0,062 | 0,585 | 4,380 | 162,459 | 48103020 | 0,238 | 0,128 | 0,644 | 27,243 | 153,556 | 31,623 |
| BEE172 | 7480370 | 0,693 | 0,047 | 0,619 | 3,253 | 165,221 | 34468096 | 0,182 | 0,109 | 0,663 | 20,711 | 153,299 | 23,964 |
| BEE174 | 15292573 | 0,708 | 0,069 | 0,562 | 5,832 | 161,538 | 50307717 | 0,205 | 0,121 | 0,619 | 27,876 | 153,383 | 33,708 |
| BEE176 | 11677843 | 0,729 | 0,055 | 0,598 | 4,674 | 162,071 | 39096839 | 0,237 | 0,104 | 0,670 | 23,040 | 153,484 | 27,714 |
| BEE177 | 10997949 | 0,766 | 0,062 | 0,551 | 3,918 | 161,049 | 34515569 | 0,269 | 0,116 | 0,605 | 18,085 | 153,794 | 22,003 |
| BEE190 | 10726503 | 0,796 | 0,064 | 0,537 | 3,692 | 163,885 | 33598043 | 0,378 | 0,120 | 0,598 | 16,318 | 153,998 | 20,011 |
| BEE191 | 6055388 | 0,833 | 0,064 | 0,521 | 1,948 | 162,587 | 14757467 | 0,377 | 0,107 | 0,594 | 7,179 | 155,071 | 9,127 |
| BEE192 | 6528502 | 0,806 | 0,062 | 0,504 | 2,085 | 163,119 | 21001357 | 0,411 | 0,113 | 0,549 | 9,183 | 154,200 | 11,268 |
| BEE193 | 6145356 | 0,824 | 0,067 | 0,510 | 1,954 | 162,975 | 17434203 | 0,430 | 0,117 | 0,551 | 7,577 | 154,462 | 9,530 |
| BEE194 | 7435272 | 0,818 | 0,061 | 0,560 | 2,604 | 162,739 | 19402838 | 0,328 | 0,107 | 0,638 | 10,410 | 154,608 | 13,014 |
| BEE195 | 6887512 | 0,811 | 0,059 | 0,542 | 2,355 | 163,167 | 20768851 | 0,359 | 0,099 | 0,616 | 10,555 | 154,517 | 12,910 |
| BEE243 | 2408720 | 0,599 | 0,036 | 0,388 | 0,746 | 175,276 | 15452830 | 0,162 | 0,100 | 0,371 | 5,245 | 152,942 | 5,990 |
| BEE244 | 4493003 | 0,791 | 0,061 | 0,536 | 1,554 | 164,153 | 16340055 | 0,359 | 0,122 | 0,581 | 7,801 | 154,089 | 9,356 |
| BEE245 | 9030446 | 0,807 | 0,063 | 0,552 | 3,155 | 163,165 | 23494047 | 0,281 | 0,114 | 0,625 | 12,692 | 154,744 | 15,847 |
| BEE246 | 6606412 | 0,843 | 0,068 | 0,527 | 2,116 | 161,448 | 17510158 | 0,348 | 0,118 | 0,594 | 8,684 | 155,479 | 10,800 |
| BEE247 | 5349295 | 0,844 | 0,074 | 0,509 | 1,643 | 160,640 | 14397938 | 0,400 | 0,130 | 0,557 | 6,471 | 155,148 | 8,114 |
| BEE248 | 4045813 | 0,833 | 0,065 | 0,513 | 1,277 | 162,198 | 10184154 | 0,418 | 0,110 | 0,550 | 4,462 | 154,950 | 5,739 |
| BEE249 | 4646831 | 0,810 | 0,063 | 0,457 | 1,344 | 163,601 | 14903320 | 0,420 | 0,115 | 0,494 | 5,840 | 154,397 | 7,183 |
| BEE250 | 5133027 | 0,812 | 0,063 | 0,500 | 1,612 | 162,613 | 15908544 | 0,444 | 0,109 | 0,541 | 6,711 | 154,227 | 8,323 |
| BEE251 | 8840707 | 0,815 | 0,063 | 0,509 | 2,817 | 162,530 | 23795123 | 0,381 | 0,114 | 0,570 | 11,023 | 154,297 | 13,839 |
| BEE252 | 8477852 | 0,789 | 0,060 | 0,554 | 3,013 | 163,030 | 27759739 | 0,333 | 0,110 | 0,618 | 14,342 | 154,220 | 17,355 |
| BEE253 | 7376703 | 0,813 | 0,064 | 0,541 | 2,493 | 161,961 | 20934610 | 0,338 | 0,109 | 0,608 | 10,643 | 154,794 | 13,136 |
Fig 4. Sequencing comparisons of two pools of 90 bumblebee libraries.
Bumblebee specimens were sampled between 2017 and 2022 from various locations in Norway, Germany, Denmark and Sweden. Both pools were generated from the same parallel library session. Pool SP was cleaned up with a 0.6 DNA/bead ratio and was sequenced on an Illumina Novaseq SP flowcell. Pool S4 was cleaned up using a double-sided size selection protocol, with a 0.5 DNA/bead ratio clean-up followed by a 0.65 DNA/bead ratio clean-up (twice) and was sequenced on ¼ Illumina Novaseq S4 flowcell. The boxplots compare (a) number of reads; (b) proportion of collapsed reads; (c) endogenous DNA proportion (unique reads only); (d) read length and (e) coverage (fold).
Ethanol-stored samples consistently produced preferable sequencing statistics, with significantly more paired reads (W = 553, P = <0.001; Fig 5A and see S1 Table), fewer collapsed reads (W = 1850, P = 0.007; Fig 5B), higher endogenous DNA proportions (W = 402, P = <0.001; Fig 5C), and similar read length (Fig 5D), leading to significantly higher overall coverage (W = 598, P = <0.001; Fig 5E).
Fig 5. Sequencing comparisons of two combined pools of 90 bumblebee libraries from the same specimens stored using 2 different methods.
Bumblebee specimens were sampled between 2017 and 2022 from various locations in Norway, Germany, Denmark and Sweden and were either stored dry at room temperature (n = 82) or in ethanol at -18°C (n = 8). The box plots compare (a) number of reads; (b) proportion of collapsed reads; (c) endogenous DNA proportion (unique reads only); (d) read length and (e) coverage (fold).
Finally, when combining the sequencing data of Pool SP and Pool S4, we obtain at least 3-fold overall coverage for 88 out of 90 samples, and at least 10-fold overall coverage for 58. (Table 2). Sequences exhibited minimal sequencing bias as a result of Tn5 library preparation (see S1 Fig).
Discussion
We here use an affordable and practical tagmentation-based genomic library preparation protocol to generate successful libraries from 6ng DNA extracted from entomological samples. We reduce costs by transposase enzyme dilution (similar to [25]), and pooling post-amplification libraries without quantification prior to clean-up. We also improve sequencing efficiency by applying a double-sided post-amplification clean-up protocol, selecting a narrow range of library fragments from the broad distribution of fragment sizes produced by the tagmentation reaction. We thus generated medium coverage (>3-fold) genomes for 88 out of 90 specimens, with an average of approximately 10-fold coverage. We estimate the overall economic cost for library preparation and PCR amplification of 96 reactions to be below 450 Euros. Below, we discuss a number of practical considerations.
First, initial extractions of bumblebee samples stored either dry or in ethanol revealed that dry-stored samples produced significantly higher overall DNA yields (up to 3-fold higher, see Fig 1); in fact, few extracts of ethanol-stored samples exhibited DNA concentrations beyond 1ng/μL. Assuming the dry-stored samples to be of higher quality, we chose to prioritise these samples for subsequent DNA extraction, library preparation and sequencing, resulting in the unbalanced sample sizes of the two groups (82 dry-stored samples, 8 ethanol-stored samples; see Table 1). Nonetheless, the eight extracts from ethanol-stored samples performed significantly better during sequencing, despite normalising the concentrations of all samples. Ethanol-stored samples obtained significantly more read pairs, which were also less often fully collapsed, indicating larger DNA fragments. They also yielded significantly greater proportions of endogenous DNA than the dry-stored sample sequences. We speculate that the combination of submerging samples in 96% ethanol with a storage temperature of -18°C reduces the presence of microbial or fungal contaminants, improves the proportion of endogenous DNA and decreases fragmentation of the little amount of DNA remaining. Given that our DNA library preparation protocol is of an adequate sensitivity to produce viable libraries from such low DNA amounts, we conclude that 96% ethanol-based storage of entomological specimens is preferable for high-throughput sequencing compared to dry storage, and that DNA yield is not a determining factor of library sequencing quality.
In order to increase affordability of library preparation, a 6X Tn5 transposase dilution was used, along with direct pooling of all amplified libraries prior to clean-up. As a result of reduced enzyme concentration, the tagmentation reaction produced a broad distribution of DNA library fragment sizes, with a large proportion of long fragments (see Figs 2B and 3, Pool SP). The ideal library fragment size distribution for Illumina DNA sequencing peaks tightly around approximately 600 bp [37]; in addition, long fragments have been found to produce significantly more error rates and reduced base qualities when sequenced on Illumina platforms [38]. In order to narrow library fragment size distribution around the recommended optimal length of 600 bp and thus increase sequencing efficiency, a more stringent double-sided post-amplification clean-up was performed on Pool S4, which removed both very short (<~400 bp) and very large (>~1000 bp) fragments. Consequently, Pool S4 produced significantly fewer collapsed reads, more endogenous reads and thus yielded approximately 1.5-fold more nucleotides per paired read compared to Pool SP. We therefore conclude that this stringent double-sided clean-up leads to a considerable improvement in sequencing efficiency.
To further reduce financial costs and workload, we pooled all libraries after PCR amplification and omitted individual quantification, which likely increased the variation and spread of coverage obtained by our sequences. Nonetheless, when combining the sequencing data of Pool SP and Pool S4, we find that 88 out of 90 samples have coverage of at least 3X, and 58 out of 90 samples with coverage over 10X. The vast majority of our samples are therefore suitable for a range of downstream analytical approaches, in spite of this pooling approach. While this 96-sample library preparation protocol (including PCR) runs in just a few hours, there is potential for increased efficiency through automation. The direct-post amplification pooling and lack of intermediate clean-up steps increases the ease with which this adaptation could be made.
Finally, we reiterate that 90 successful libraries were generated from a standardised DNA input of 6ng extracted from just one or two bumblebee legs. This low tissue volume minimises morphological damage to biologically valuable entomological specimens, helps to maintains their potential for future taxonomic, ecological and DNA-based research, and as a result reduces the requirement for additional entomological sampling efforts. In addition, library preparation protocols with low DNA input volumes are essential for the genomic sequencing of much smaller organisms than bumblebees, which may yield very little DNA even when entire specimens are used for DNA extraction. We therefore hope that this protocol can be of considerable value for the future of entomological genomics and monitoring.
Supporting information
(DOCX)
Sequencing statistics for two sequencing pools (SP and S4) of 90 specimens of two bumblebee species: (a) Bombus lapidarius (n = 46) and (b) Bombus pascuorum (n = 44). Bumblebee specimens were sampled between 2017 and 2022 from various locations in Norway, Germany, Denmark and Sweden, and samples were stored either dry at room temperature, or submerged in ethanol at -18°C. Libraries for both pools were generated simultaneously. The SP pool was cleaned up with a 0.6 DNA/bead ratio and was sequenced on an Illumina Novaseq SP flowcell. The S4 pool was cleaned up using a double-sided size selection protocol, with a 0.5 DNA/bead ratio clean-up followed by a 0.65 DNA/bead ratio clean-up (twice) and was sequenced on ¼ Illumina Novaseq S4 flowcell. Metadata presented for each sequencing pool are specimen ID, storage protocol, year sampled, number of reads (bp), collapsed reads (proportion), clonality (proportion), endogenous DNA (proportion), nuclear coverage (fold), mitochondrial clonality (proportion), mitochondrial coverage (fold) and read length (bp). Also presented is combined per-specimen coverage of both sequencing runs.
(XLSX)
While Tn5 transposase can cause sequence bias, this bias is associated with the first 10 bases of the reads only. We find that by using simple post-analysis filtering, preliminary data (not shown) indicate that these biases do not impact our ability to perform detailed population genetic analyses. Patterns were obtained using MapDamage v. 2.0.6 after down-sampling BAM files to 1,000,000 reads (Jónsson, H., Ginolhac, A., Schubert, M., Johnson, P. L. F., & Orlando, L. MapDamage2.0: Fast approximate Bayesian estimates of ancient DNA damage parameters. Bioinformatics. 2013;29(13): 1682–1684).
(TIF)
Acknowledgments
Sequencing was performed by the Norwegian Sequencing Centre (www.sequencing.uio.no), a national technology platform hosted by the University of Oslo and Oslo University Hospital, supported by the Research Council of Norway and South-Eastern Regional Health Authority.
Data Availability
All raw sequence read data are available under PRJEB65580 at the European Nucleotide Archive (ENA, https://www.ebi.ac.uk). All raw sequence statistics are in the Supplementary Table.
Funding Statement
This work was funded by a grant from the University of Oslo awarded to B. Star and M.A.K. Sydenham. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Wagner DL, Grames EM, Forister ML, Berenbaum MR, Stopak D. Insect decline in the Anthropocene: Death by a thousand cuts. Proc Natl Acad Sci U S A. 2021;118(2). doi: 10.1073/pnas.2023989118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.van der Sluijs JP. Insect decline, an emerging global environmental risk. Current Opinion in Environmental Sustainability. 2020;46:39–42. [Google Scholar]
- 3.Potts S, Dauber, J., Hochkirch, A., Oteman, B., Roy, D., Ahnre, K., et al. Proposal for an EU Pollinator Monitoring Scheme. Luxembourg; 2020. Report No.: ISBN 978-92-76-23859-1 Contract No.: JRC122225.
- 4.Saunders ME, Janes JK, O’Hanlon JC. Moving On from the Insect Apocalypse Narrative: Engaging with Evidence-Based Insect Conservation. Bioscience. 2020;70(1):80–9. [Google Scholar]
- 5.Graham CF, Glenn TC, McArthur AG, Boreham DR, Kieran T, Lance S, et al. Impacts of degraded DNA on restriction enzyme associated DNA sequencing (RADSeq). Mol Ecol Resour. 2015;15(6):1304–15. doi: 10.1111/1755-0998.12404 [DOI] [PubMed] [Google Scholar]
- 6.Knuff AK, Winiger N, Klein AM, Segelbacher G, Staab M, Kotze DJ. Optimizing sampling of flying insects using a modified window trap. Methods in Ecology and Evolution. 2019;10(10):1820–5. [Google Scholar]
- 7.Lindahl T. Instability and decay of the primary structure of DNA. Nature. 1993;362(6422):709–15. doi: 10.1038/362709a0 [DOI] [PubMed] [Google Scholar]
- 8.Soniat TJ, Sihaloho HF, Stevens RD, Little TD, Phillips CD, Bradley RD. Temporal-dependent effects of DNA degradation on frozen tissues archived at –80°C. Journal of Mammalogy. 2021;102(2):375–83. [Google Scholar]
- 9.King JR, Porter SD. Recommendations on the use of alcohols for preservation of ant specimens (Hymenoptera, Formicidae). Insectes Sociaux. 2004;51(2):197–202. [Google Scholar]
- 10.Moreau CS, Wray BD, Czekanski-Moir JE, Rubin BER. DNA preservation: a test of commonly used preservatives for insects. Invertebrate Systematics. 2013;27(1):81–6. [Google Scholar]
- 11.Pd Sousa, Henriques A Silva SE, Carvalheiro LG, Smagghe G, Michez D, et al. Genomic Patterns of Iberian Wild Bees Reveal Levels of Diversity, Differentiation and Population Structure, Supporting the “Refugia within Refugia” Hypothesis. Diversity (Basel). 2023;15(6):746. [Google Scholar]
- 12.Frampton M, Droege S, Conrad T, Prager S, Richards MH. Evaluation of Specimen Preservatives for DNA Analyses of Bees. Journal of Hymenoptera Research. 2008;17(2):195–200. [Google Scholar]
- 13.Qiagen. DNeasy® Blood & Tissue Handbook. 2020. [Google Scholar]
- 14.Beadle K, Singh KS, Troczka BJ, Randall E, Zaworra M, Zimmer CT, et al. Genomic insights into neonicotinoid sensitivity in the solitary bee Osmia bicornis. PLoS Genet. 2019;15(2):e1007903. doi: 10.1371/journal.pgen.1007903 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Santos D, Ribeiro GC, Cabral AD, Sperança MA. A non-destructive enzymatic method to extract DNA from arthropod specimens: Implications for morphological and molecular studies. PLoS One. 2018;13(2):e0192200. doi: 10.1371/journal.pone.0192200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tin MM-Y, Economo EP, Mikheyev AS. Sequencing degraded DNA from non-destructively sampled museum specimens for RAD-tagging and low-coverage shotgun phylogenetics. PLoS One. 2014;9(5):e96793-e. doi: 10.1371/journal.pone.0096793 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Castalanelli MA, Severtson DL, Brumley CJ, Szito A, Foottit RG, Grimm M, et al. A rapid non-destructive DNA extraction method for insects and other arthropods. Journal of Asia-Pacific Entomology. 2010;13(3):243–8. [Google Scholar]
- 18.Thomsen PF, Elias S, Gilbert MTP, Haile J, Munch K, Kuzmina S, et al. Non-Destructive Sampling of Ancient Insect DNA. PLoS One. 2009;4(4):e5048-e. doi: 10.1371/journal.pone.0005048 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Korlević P, McAlister E, Mayho M, Makunin A, Flicek P, Lawniczak MKN. A Minimally Morphologically Destructive Approach for DNA Retrieval and Whole-Genome Shotgun Sequencing of Pinned Historic Dipteran Vector Species. Genome Biol Evol. 2021;13(10). doi: 10.1093/gbe/evab226 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Cavill EL, Liu S, Zhou X, Gilbert MTP. To bee, or not to bee? One leg is the question. Mol Ecol Resour. 2022;22(5):1868–74. doi: 10.1111/1755-0998.13578 [DOI] [PubMed] [Google Scholar]
- 21.Pálsdóttir AH, Bläuer A, Rannamäe E, Boessenkool S, Hallsson JH. Not a limitless resource: ethics and guidelines for destructive sampling of archaeofaunal remains. R Soc Open Sci. 2019;6(10):191059-. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shendure J, Balasubramanian S, Church GM, Gilbert W, Rogers J, Schloss JA, et al. DNA sequencing at 40: past, present and future. Nature. 2017;550(7676):345–53. doi: 10.1038/nature24286 [DOI] [PubMed] [Google Scholar]
- 23.Tvedte ES, Michalski J, Cheng S, Patkus RS, Tallon LJ, Sadzewicz L, et al. Evaluation of a high-throughput, cost-effective Illumina library preparation kit. Scientific Reports. 2021;11(1):15925-. doi: 10.1038/s41598-021-94911-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Adey A, Morrison HG, Asan, Xun X, Kitzman JO, Turner EH, et al. Rapid, low-input, low-bias construction of shotgun fragment libraries by high-density in vitro transposition. Genome Biol. 2010;11(12):R119-R. doi: 10.1186/gb-2010-11-12-r119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jones A, Stanley D, Ferguson S, Schwessinger B, Borevitz J, Warthmann N. Cost-conscious generation of multiplexed short-read DNA libraries for whole-genome sequencing. PLoS One. 2023;18(1):e0280004-e. doi: 10.1371/journal.pone.0280004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Picelli S, Björklund AK, Reinius B, Sagasser S, Winberg G, Sandberg R. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 2014;24(12):2033–40. doi: 10.1101/gr.177881.114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Hennig BP, Velten L, Racke I, Tu CS, Thoms M, Rybin V, et al. Large-Scale Low-Cost NGS Library Preparation Using a Robust Tn5 Purification and Tagmentation Protocol. G3 (Bethesda). 2018;8(1):79–89. doi: 10.1534/g3.117.300257 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Vonesch SC, Li S, Tu CS, Hennig BP, Dobrev N, Steinmetz LM. Fast and inexpensive whole-genome sequencing library preparation from intact yeast cells. G3 (Bethesda). 2021;11(1). doi: 10.1093/g3journal/jkaa009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Baym M, Kryazhimskiy S, Lieberman TD, Chung H, Desai MM, Kishony R. Inexpensive multiplexed library preparation for megabase-sized genomes. PLoS One. 2015;10(5):e0128036-e. doi: 10.1371/journal.pone.0128036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Bruinsma S, Burgess J, Schlingman D, Czyz A, Morrell N, Ballenger C, et al. Bead-linked transposomes enable a normalization-free workflow for NGS library preparation. BMC Genomics. 2018;19(1):722-. doi: 10.1186/s12864-018-5096-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Shapiro B, Barlow A, Heintzman PD, Hofreiter M, Paijmans JLA, Soares AER. Ancient DNA: Methods and Protocols. New York, NY: Springer New York: Imprint: Humana; 2019. [Google Scholar]
- 32.Bronner IF, Quail MA. Best Practices for Illumina Library Preparation. Current Protocols in Human Genetics. 2019;102(1):n/a. doi: 10.1002/cphg.86 [DOI] [PubMed] [Google Scholar]
- 33.Schubert M, Ermini L, Der Sarkissian C, Jónsson H, Ginolhac A, Schaefer R, et al. Characterization of ancient and modern genomes by SNP detection and phylogenomic and metagenomic analysis using PALEOMIX. Nat Protoc. 2014;9(5):1056–82. doi: 10.1038/nprot.2014.063 [DOI] [PubMed] [Google Scholar]
- 34.Crowley LM, Sivell O, Sivell D. The genome sequence of the Common Carder Bee, Bombus pascuorum (Scopoli, 1763). Wellcome Open Research. 2023;8:142. doi: 10.12688/wellcomeopenres.19251.1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Schubert M, Lindgreen S, Orlando L. AdapterRemoval v2: rapid adapter trimming, identification, and read merging. BMC Res Notes. 2016;9(1):88. doi: 10.1186/s13104-016-1900-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Ithaca: Ithaca: Cornell University Library, arXiv.org; 2013. [Google Scholar]
- 37.Illumina DNA Prep Reference Guide. 2020. [Google Scholar]
- 38.Tan G, Opitz L, Schlapbach R, Rehrauer H. Long fragments achieve lower base quality in Illumina paired-end sequencing. Sci Rep. 2019;9(1):2856–. doi: 10.1038/s41598-019-39076-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
(DOCX)
Sequencing statistics for two sequencing pools (SP and S4) of 90 specimens of two bumblebee species: (a) Bombus lapidarius (n = 46) and (b) Bombus pascuorum (n = 44). Bumblebee specimens were sampled between 2017 and 2022 from various locations in Norway, Germany, Denmark and Sweden, and samples were stored either dry at room temperature, or submerged in ethanol at -18°C. Libraries for both pools were generated simultaneously. The SP pool was cleaned up with a 0.6 DNA/bead ratio and was sequenced on an Illumina Novaseq SP flowcell. The S4 pool was cleaned up using a double-sided size selection protocol, with a 0.5 DNA/bead ratio clean-up followed by a 0.65 DNA/bead ratio clean-up (twice) and was sequenced on ¼ Illumina Novaseq S4 flowcell. Metadata presented for each sequencing pool are specimen ID, storage protocol, year sampled, number of reads (bp), collapsed reads (proportion), clonality (proportion), endogenous DNA (proportion), nuclear coverage (fold), mitochondrial clonality (proportion), mitochondrial coverage (fold) and read length (bp). Also presented is combined per-specimen coverage of both sequencing runs.
(XLSX)
While Tn5 transposase can cause sequence bias, this bias is associated with the first 10 bases of the reads only. We find that by using simple post-analysis filtering, preliminary data (not shown) indicate that these biases do not impact our ability to perform detailed population genetic analyses. Patterns were obtained using MapDamage v. 2.0.6 after down-sampling BAM files to 1,000,000 reads (Jónsson, H., Ginolhac, A., Schubert, M., Johnson, P. L. F., & Orlando, L. MapDamage2.0: Fast approximate Bayesian estimates of ancient DNA damage parameters. Bioinformatics. 2013;29(13): 1682–1684).
(TIF)
Data Availability Statement
All raw sequence read data are available under PRJEB65580 at the European Nucleotide Archive (ENA, https://www.ebi.ac.uk). All raw sequence statistics are in the Supplementary Table.