Abstract
It has been predicted that the highly degenerate mammalian Y chromosome will be lost eventually. Indeed, Y was lost in the Ryukyu spiny rat Tokudaia osimensis, but the fate of the formerly Y-linked genes is not completely known. We looked for all 12 ancestrally Y-linked genes in a draft T. osimensis genome sequence. Zfy1, Zfy2, Kdm5d, Eif2s3y, Usp9y, Uty, and Ddx3y are putatively functional and are now located on the X chromosome, whereas Rbmy, Uba1y, Ssty1, Ssty2, and Sry are missing or pseudogenized. Tissue expressions of the mouse orthologs of the retained genes are significantly broader/higher than those of the lost genes, suggesting that the destinies of the formerly Y-linked genes are related to their original expressions. Interestingly, patterns of gene retention/loss are significantly more similar than by chance across four rodent lineages where Y has been independently lost, indicating a level of certainty in the fate of Y-linked genes even when the chromosome is gone.
Keywords: spiny rat, Y chromosome, evolution, gene loss, gene expression, Tokudaia osimensis
Significance.
Although sex is generally determined by the Y chromosome in therian mammals, Y is absent in the Ryukyu spiny rat Tokudaia osimensis, raising the question of the whereabouts of all formerly Y-linked genes. Our comparative genomic analysis shows that, of 12 ancestrally Y-linked genes, seven are now on the X chromosome whereas the rest are missing or pseudogenized. We find that the mouse orthologs of the T. osimensis retained genes tend to have broader/higher expressions than the lost genes, suggesting that the fate of the ancestrally Y-linked genes upon the Y loss is related to their original expressions, which may also explain our observation of similar gene retention/loss patterns across four rodent lineages that have independently lost the Y chromosome.
Loss of the Y Chromosome in Tokudaia osimensis
Therian mammals typically have the XX/XY sex determination system, where the X and Y sex chromosomes originated from a pair of autosomes. The male-determining Y chromosome, however, is highly degenerate, having lost most of its original genes during the evolution (Bachtrog 2013). This trend has led to the hypothesis that the entire Y chromosome will be lost eventually in the evolution of some mammals including humans (Graves 2004). Indeed, Y has been lost four times: in each of Ellobius talpinus, Ellobius lutescens, and Microtus oregoni and, according to a parsimony inference, in the common ancestor of Tokudaia osimensis and T. tokunoshimensis, all being rodents (Kuroiwa et al. 2010; Mulugeta et al. 2016; Couger et al. 2021). The fate of formerly Y-linked genes upon the loss of Y is of substantial interest, and a few scenarios are possible. First, these genes may be lost in company with the Y chromosome. Alternatively, they may be relocated to other chromosomes individually or en bloc. Several genomic studies have addressed the destinies of formerly Y-linked genes in E. talpinus, E. lutescens, and M. oregoni (Mulugeta et al. 2016; Couger et al. 2021). Here, we address this question in T. osimensis by analyzing its recently published draft genome sequence (Terao et al. 2022). We then compare patterns of gene retention/loss across the four rodent lineages that have independently lost the Y chromosome.
Tokudaia osimensis, or Ryukyu spiny rat, is an endangered species endemic to Amami Ōshima island in the Amami Islands of the Ryukyu archipelago of Japan. Tokudaia osimensis has lost its Y chromosome as well as the male sex determination gene Sry (Terao et al. 2022). A recent study discovered a male-specific duplication of an enhancer upstream of Sox9 on an autosome and proposed that the duplicated Sox9 enhancer has substituted for Sry in male sex determination in T. osimensis (Terao et al. 2022). Prior to genome sequencing, cDNA clones of five mouse Y-linked genes were used as probes to identify their homologs in T. osimensis using fluorescence in situ hybridization (FISH). It was reported that Kdm5d, Eif2s3y, Tspy, and Zfy were translocated to the distal end of the X chromosome by an unusual recombination between the nonpseudoautosomal regions of the X and Y chromosomes and that a pseudogenized copy of Rbmy exists on chromosome 2 (Arakawa et al. 2002; Kuroiwa et al. 2010). However, the fate of other formerly Y-linked genes remains elusive.
Retained and Lost Formerly Y-linked Genes in T. osimensis
We investigated the existence and location of the formerly Y-linked genes of T. osimensis by using the mouse (Mus musculus) orthologs of ancestrally Y-linked genes (Zfy1, Zfy2, Uba1y, Kdm5d, Eif2s3y, Uty, Ddx3y, Usp9y, Rbmy, Ssty1, Ssty2, and Sry) as queries to BLAST the T. osimensis genome sequence (see Materials and Methods). The so-called ancestrally Y-linked genes are those that presumably existed on the Y chromosome of the most recent common ancestor of T. osimensis and mouse (Suzuki et al. 2000), which existed ∼10 mya (Kumar et al. 2022). However, the currently available genome assembly of T. osimensis is at the scaffold level (Terao et al. 2022), making it difficult to chromosomally locate the formerly Y-linked genes. To this end, we utilized the available Hi-C data of T. osimensis (Terao et al. 2022) to improve the quality of the genome assembly (see Materials and Methods). After this improvement, the number of scaffolds with >50 million bases (Mb) increased from 1 to 13. The coding sequence (CDS) of each mouse ancestrally Y-linked gene was used as the query to BLAST against the newly obtained, assembled T. osimensis scaffolds. To ensure that the BLAST search identified the true formerly Y-linked genes in T. osimensis reference genome rather than the corresponding X homologs, the matched T. osimensis sequences were BLASTed back to the mouse genome to examine whether the best hits are the corresponding mouse Y orthologs. We found that the BLAST matches of mouse Zfy1, Zfy2, Kdm5d, Eif2s3y, Uty, and Ddx3y were on Scaffold 17 with high coverages and hit scores, indicating their existence in the T. osimensis genome (Table 1). Mouse Usp9y had a relatively poor BLAST match on the T. osimensis Scaffold 17, and nearly half of the sequence failed to align. Rbmy was detected on Scaffold 3. No BLAST matches were detected for Uba1y, Ssty1, Ssty2, and Sry, suggesting their absence from the T. osimensis genome (Table 1). To exclude the possibility that the negative BLAST result is due to a low genome assembly quality, we performed a BUSCO analysis (Manni et al. 2021) of the T. osimensis genome assembly. The BUSCO score revealed a completeness of 95.8% (complete: 95.8%, single copy: 94.7%, duplicated: 1.1%, fragmented: 0.8%, and missing: 3.4%), suggesting that the negative BLAST results most likely reflect the absence of Uba1y, Ssty1, Ssty2, and Sry in T. osimensis.
Table 1.
Status of ancestrally Y-linked genes in the Ryukyu spiny rat T. osimensis
| Gene name | Scaffold | Intact? | Number of introns | Number of introns in mouse | Genomic location |
|---|---|---|---|---|---|
| Zfy1 | 17 | Yes | 7 | 7 | Chromosome X |
| Zfy2 | 17 | Yes | 7 | 7 | Chromosome X |
| Uba1y | … | … | … | … | Lost |
| Kdm5d | 17 | Yes | 25 | 26 | Chromosome X |
| Eif2s3y | 17 | Yes | 12 | 12 | Chromosome X |
| Uty | 17 | Yes | 25 | 27 | Chromosome X |
| Ddx3y | 17 | Yes | 16 | 17 | Chromosome X |
| Usp9y | 17 | Yes | At least 24 | 43 | Chromosome X |
| Sry | … | … | … | … | Lost |
| Rbmy | 3 | No | 0 | 8 | Chromosome 2 (pseudogenized) |
| Ssty1 | … | … | … | … | Lost |
| Ssty2 | … | … | … | … | Lost |
For each retained formerly Y-linked T. osimensis gene, we used the BLAST result to compare its number of introns with the corresponding number in mouse, which informed the means of relocation of the T. osimensis gene. The intron numbers of T. osimensis Zfy1, Zfy2, Kdm5d, Eif2s3y, Uty, and Ddx3y are generally equal to those of the mouse Y orthologs (Table 1), whereas the number of introns detected in Usp9y of T. osimensis is approximately half that of the mouse due to the incomplete BLAST match. These results, along with the fact that the above seven genes are located on the same scaffold, suggest that Zfy1, Zfy2, Kdm5d, Eif2s3y, Uty, Ddx3y, and Usp9y were likely relocated by chromosomal translocation. By contrast, we found Rbmy to have lost all of its introns, suggesting that it was relocated via retroposition.
To further investigate the status of Usp9y in T. osimensis, we examined whether its CDS is intact. While no premature stop codon was found in Usp9y, a sequencing gap was detected. Specifically, compared to the CDS of mouse Usp9y, which is about 7.6 kb long, the CDS of T. osimensis Usp9y misses about 3.3 kb from the start codon. Consequently, the previous BLAST search of Usp9y resulted in a low query coverage but a high percent identity and hit score (query coverage: 57%; percent identity: 94.81%; bit score: 1177). These observations suggest that Usp9y is intact and putatively functional in T. osimensis. We also investigated whether other retained genes are putatively functional by examining premature stop codons in their CDSs obtained from the BLAST search. The CDS of each of these retained genes, except Rbmy, is complete and without premature stop codons. By contrast, premature stop codons are present in Rbmy, indicating that Rbmy has been pseudogenized in T. osimensis (Table 1).
To investigate the T. osimensis chromosomes to which the retained formerly Y-linked genes have been relocated, we mapped Scaffold 17, which contains all putatively functional, formerly Y-linked genes, and Scaffold 3, which contains pseudogenized Rbmy, to the mouse genome. Scaffold 3 was mapped to mouse chromosome 9, which is the counterpart of an arm of T. osimensis’ chromosome 2 (Nakamura et al. 2007), suggesting that Rbmy is relocated to an autosome in T. osimensis. Scaffold 17, however, was mapped to the mouse Y chromosome, which offers no additional insight because T. osimensis has lost the Y chromosome. Fortunately, an earlier FISH study suggested that T. osimensis Kdm5d, Eif2s3y, Tspy, and Zfy were translocated as a group to the distal end of the X chromosome (Kuroiwa et al. 2010). Given that the above four genes are all located on Scaffold 17, we can infer that Scaffold 17 and thereby all seven putatively functional, formerly Y-linked T. osimensis genes must be on the same distal end of the X chromosome (Table 1). Using FISH and other evidence, Kuroiwa et al. (2010) showed that Rbmy was retrotransposed to T. osimensis’ chromosome 2. The complete loss of all introns of Rbmy (Table 1) and the mapping of Scaffold 3 corroborate the above finding.
Mouse Tissue Expressions of Orthologs of T. osimensis Retained and Lost Genes
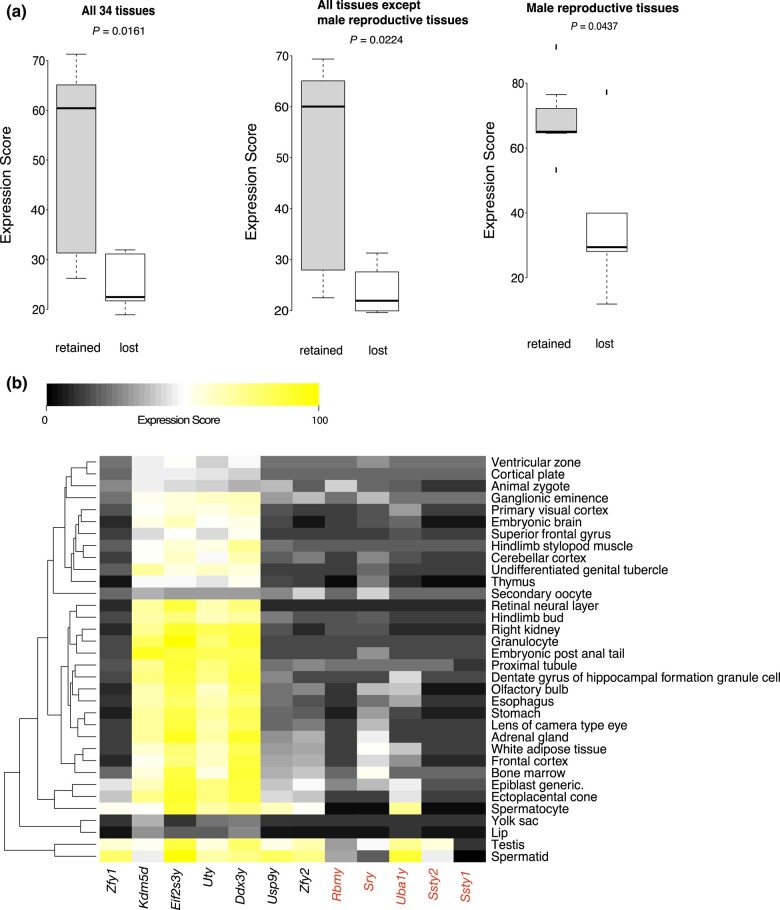
We then asked why the lost genes were lost whereas the retained genes are retained in T. osimensis. One potential reason is that the lost genes were functionally less important than the retained genes, for example, because the former were expressed in fewer tissues or at lower levels than the latter. From Bgee (Bastian et al. 2021), we obtained a dataset of mouse expression scores of each ancestrally Y-linked gene in 34 tissues. We calculated the average expression score for the mouse ortholog of each retained or lost gene across all 34 tissues. Considering the importance of Y chromosome in male sex determination and spermatogenesis (Subrini and Turner 2021), we also calculated the average expression score across male reproductive tissues (testis, spermatid, and spermatocyte), and across all tissues excluding the three male reproductive tissues. In all three analyses, the retained genes have significantly higher average expression scores than the lost genes (Fig. 1a). This said, the expression level varies substantially among tissues and genes such that some lost genes have higher expressions than some retained genes in certain tissues (Fig. 1b). Additionally, whether a gene will be lost probably also depends on whether it can be substituted. For example, the loss of Sry in T. osimensis was likely made possible by the male-specific duplication of a Sox9 enhancer that substituted for Sry's role in sex determination (Terao et al. 2022).
Fig. 1.
Mouse tissue expressions of 12 ancestrally Y-linked genes. a) Box plots of mean expression scores across tissues for the mouse orthologs of T. osimensis retained and lost formerly Y-linked genes, respectively. The lower and upper edges of a box represent the first and third quartiles, respectively, the horizontal line inside the box indicates the median, the whiskers extend to the most extreme values inside inner fences (median ± 1.5× interquartile range) and the dots show outliers. P-values are based on two-sample, two-tailed t-tests with unequal variances. b) Heatmap of expression scores in 34 tissues. Mouse orthologs of the genes that are lost or pseudogenized in T. osimensis are color-coded in red, whereas those of the retained genes are in black. The dendrogram produced by hierarchical clustering is shown on the leftmost of the heatmap.
Patterns of Gene Retention/Loss Across Lineages With Independent Y Losses
Finally, we compared the retained and lost (including pseudogenized) formerly Y-linked genes across four rodent species (T. osimensis, E. talpinus, E. lutescens, and M. oregoni) that have independently lost their Y chromosomes (Table 2). The genes retained/lost are virtually identical between E. talpinus and E. lutescens (i.e. E. talpinus lost one more gene relative to E. lutescens; Table 2), which is likely because the evolutionary events that led to gene retention occurred in the common ancestor of the two species despite their independent Y losses (Mulugeta et al. 2016). Because the gene retention/loss pattern of E. lutescens (but not that of E. talpinus) represents the pattern of the most recent common ancestor of the two Ellobius species, E. lutescens was chosen to represent the two species. While the number of genes retained varies substantially among T. osimensis (7), M. oregoni (11), and E. lutescens (6), the specific genes retained show some similarities (Table 2). Using a randomization test, we found that the patterns of gene retention/loss are significantly more similar across the evolutionary lineages of T. osimensis, M. oregoni, and Ellobius than by chance (P = 0.042; see Materials and Methods).
Table 2.
Status of ancestrally Y-linked genes in four rodents that independently lost the Y chromosome
| Gene name | Tokudaia osimensis (XO male) | Microtus oregoni (XX male) | Ellobius lutescens (XO male) | Ellobius talpinus (XX male) | ||||
|---|---|---|---|---|---|---|---|---|
| Presencea | Intactnessb | Presencea | Intactnessb | Presencea | Intactnessb | Presencea | Intactnessb | |
| Zfy1 | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ |
| Zfy2 | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ |
| Uba1y | × | × | ✓ | × | × | × | × | × |
| Kdm5d | ✓ | ✓ | ✓ | ✓ | × | × | × | × |
| Eif2s3y | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ |
| Uty | ✓ | ✓ | ✓ | ✓ | × | × | × | × |
| Ddx3y | ✓ | ✓ | ✓ | ✓ | × | × | × | × |
| Usp9y | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | × | × |
| Sry | × | × | ✓ | ✓ | × | × | × | × |
| Rbmy | ✓ | × | ✓ | ✓ | × | × | × | × |
| Ssty1 | × | × | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ |
| Ssty2 | × | × | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ |
a✓, present; ×, absent.
b✓, putatively functional; ×, pseudogenized.
Conclusions
In summary, of the 12 ancestrally Y-linked genes of T. osimensis, Zfy1, Zfy2, Kdm5d, Eif2s3y, Usp9y, Uty, and Ddx3y were translocated en bloc to the X chromosome and are putatively functional, Rbmy was retrotransposed to chromosome 2 but has been pseudogenized, and Uba1y, Sry, Ssty1, and Ssty2 have been lost from the genome. The mouse orthologs of the T. osimensis retained genes tend to have broader/higher expressions than the lost genes, suggesting that the fate of the ancestrally Y-linked genes upon the Y loss is influenced by their expressions, which may also explain why the gene retention/loss patterns are broadly similar in the four rodent lineages that have lost the Y chromosome independently.
Materials and Methods
Information about the mouse ancestrally Y-linked genes were obtained from Table 1 of a previous study (Soh et al. 2014). We define ancestrally Y-linked genes as those that existed in the most recent common ancestor of T. osimensis and mouse. Besides 10 long-lived, ancestrally Y-linked genes, Ssty1 and Ssty2, previously categorized as mouse acquired genes, likely existed on the Y chromosome of the common ancestor of T. osimensis and mouse (Soh et al. 2014; Mulugeta et al. 2016), so are included in our analysis. Additionally, Tspy likely existed on the Y-chromosome of the common ancestor of T. osimensis and mouse (Arakawa et al. 2002), but because it has been lost or pseudogenized in mouse and several other Y-carrying rodents (Kuroiwa et al. 2010; Couger et al. 2021), its presence/absence is unrelated to Y losses and hence is ignored in our analysis. The Hi-C reads of T. osimensis were obtained from the NCBI Sequence Read Archive (Terao et al. 2022). The mouse genome assembly GRCm39 was also retrieved from NCBI (Church et al. 2011). The Hi-C reads of T. osimensis were prompted into Arima Hi-C Mapping Pipeline (https://github.com/ArimaGenomics/mapping_pipeline), where the mouse genome was used as the reference, to produce a filtered and processed binary alignment map file to feed into YaHs (Zhou et al. 2023), to assemble the scaffold reads at the chromosomal level. The quality of the newly assembled genome was examined using BUSCO (Manni et al. 2021). The BLAST + executables (Camacho et al. 2009) were used to BLAST the coding sequences of mouse ancestrally Y-linked genes from NCBI CCDS database against the newly assembled T. osimensis genome. Visualization of the BLAST results was achieved through Kablammo (Wintersinger and Wasmuth 2015). We then mapped T. osimensis Scaffold 17 to the mouse genome and also visualized the result using nucmer in MUMmer (Kurtz et al. 2004) and Ribbon (Nattestad et al. 2021). To test the integrity of each formerly Y-linked gene in T. osimensis, its open reading frame was identified using Open Reading Frame Finder at NCBI (https://www.ncbi.nlm.nih.gov/orffinder/) and was inspected for the existence of premature stop codons.
The mouse gene expression data of ancestrally Y-linked genes were obtained from Bgee (Bastian et al. 2021). The expression scores in Bgee were calculated on a scale of 0 to 100, based on an integration of RNA-Seq, Affymetrix, in situ hybridization, and EST data produced by various studies. We extracted 34 tissues where all 12 ancestrally Y-linked genes have available data in the database and calculated the averages. The box and whisker plots showing the mean expression scores were generated by BoxPlotR (Spitzer et al. 2014). The heatmap summarizing expression scores of ancestrally Y-linked genes across all 34 tissues was generated by Heatmapper (Babicki et al. 2016).
Previous studies investigated the status of some ancestrally Y-linked genes of M. oregoni, E. lutescens, and E. talpinus. However, uncertainty of the presence or the intactness of some ancestrally Y-linked genes and incompleteness in functionality analysis remains (Mulugeta et al. 2016; Couger et al. 2021). To address these deficiencies, we performed the same BLAST search against the genome of M. oregoni, E. lutescens, and E. talpinus, respectively, to investigate the presence of the 12 ancestrally Y-linked genes. To examine the intactness of these genes, we again pasted the aligned sequences from BLAST searches to Open Reading Frame Finder (https://www.ncbi.nlm.nih.gov/orffinder/). To compare the pattern of gene retention/loss across T. osimensis, E. lutescens, and M. oregoni, we created an array representing the retention/loss status of the formerly Y-linked genes of each species, where putatively functional genes were denoted as 1 and nonfunctional or lost genes as 0. We then randomly shuffled the gene retention/loss array of each species independently, without altering the numbers of 1 and 0 s in each array. Upon the shuffling, we estimated Pearson's correlation coefficient (r) between every two arrays and computed the average of the three r values (denoted R). This process was repeated 10,000 times to obtain a null distribution of R. The R computed from the actual data (i.e. without array shuffling) was compared with the null distribution to acquire a one-tailed P-value, which is the fraction of the distribution that exceeds the R from the actual data.
Acknowledgements
This work was supported by the U.S. National Institutes of Health grant R35GM139484 to J.Z.
Contributor Information
Jiachen Li, Department of Ecology and Evolutionary Biology, University of Michigan, Ann Arbor, MI 48109, USA.
Siliang Song, Department of Ecology and Evolutionary Biology, University of Michigan, Ann Arbor, MI 48109, USA.
Jianzhi Zhang, Department of Ecology and Evolutionary Biology, University of Michigan, Ann Arbor, MI 48109, USA.
Data Availability
All data used are publicly available.
Literature Cited
- Arakawa Y, Nishida-Umehara C, Matsuda Y, Sutou S, Suzuki H. X-chromosomal localization of mammalian Y-linked genes in two XO species of the Ryukyu spiny rat. Cytogenet. Genome Res. 2002:99(1-4):303–309. 10.1159/000071608 [DOI] [PubMed] [Google Scholar]
- Babicki S, Arndt D, Marcu A, Liang Y, Grant JR, Maciejewski A, Wishart DS. Heatmapper: web-enabled heat mapping for all. Nucleic Acids Res. 2016:44(W1):W147–W153. 10.1093/nar/gkw419 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bachtrog D. Y-chromosome evolution: emerging insights into processes of Y-chromosome degeneration. Nat Rev Genet. 2013:14(2):113–124. 10.1038/nrg3366 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bastian FB, Roux J, Niknejad A, Comte A, Fonseca Costa SS, de Farias TM, Moretti S, Parmentier G, de Laval VR, Rosikiewicz M, et al. The Bgee suite: integrated curated expression atlas and comparative transcriptomics in animals. Nucleic Acids Res. 2021:49(D1):D831–D847. 10.1093/nar/gkaa793 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL. BLAST+: architecture and applications. BMC Bioinformatics. 2009:10(1):421. 10.1186/1471-2105-10-421 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Church DM, Schneider VA, Graves T, Auger K, Cunningham F, Bouk N, Chen HC, Agarwala R, McLaren WM, Ritchie GRS, et al. Modernizing reference genome assemblies. PLoS Biol. 2011:9(7):e1001091. 10.1371/journal.pbio.1001091 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Couger MB, Roy SW, Anderson N, Gozashti L, Pirro S, Millward LS, Kim M, Kilburn D, Liu KJ, Wilson TM, et al. Sex chromosome transformation and the origin of a male-specific X chromosome in the creeping vole. Science. 2021:372(6542):592–600. 10.1126/science.abg7019 [DOI] [PubMed] [Google Scholar]
- Graves JAM. The degenerate Y chromosome–can conversion save it? Reprod. Fertil. Dev. 2004:16(5):527–534. 10.1071/RD03096 [DOI] [PubMed] [Google Scholar]
- Kumar S, Suleski M, Craig JM, Kasprowicz AE, Sanderford M, Li M, Stecher G, Hedges SB. TimeTree 5: an expanded resource for species divergence times. Mol Biol Evol. 2022:39(8):msac174. 10.1093/molbev/msac174 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuroiwa A, Ishiguchi Y, Yamada F, Shintaro A, Matsuda Y. The process of a Y-loss event in an XO/XO mammal, the Ryukyu spiny rat. Chromosoma. 2010:119(5):519–526. 10.1007/s00412-010-0275-8 [DOI] [PubMed] [Google Scholar]
- Kurtz S, Shumway M, Antonescu C, Salzberg SL, Phillippy A, Smoot M, Delcher AL, Delcher AL. Versatile and open software for comparing large genomes. Genome Biol. 2004:5(2):12. 10.1186/gb-2004-5-2-r12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manni M, Berkeley MR, Seppey M, Simão FA, Zdobnov EM. BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol Biol Evol. 2021:38(10):4647–4654. 10.1093/molbev/msab199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mulugeta E, Wassenaar E, Sleddens-Linkels E, Van Ijcken WFJ, Heard E, Grootegoed JA, Just W, Gribnau J, Baarends WM. Genomes of Ellobius species provide insight into the evolutionary dynamics of mammalian sex chromosomes. Genome Res. 2016:26(9):1202–1210. 10.1101/gr.201665.115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakamura T, Kuroiwa A, Nishida-Umehara C, Matsubara K, Yamada F, Matsuda Y. Comparative chromosome painting map between two Ryukyu spiny rat species, Tokudaia osimensis and Tokudaia tokunoshimensis (Muridae, Rodentia). Chromosome Res. 2007:15(6):799–806. 10.1007/s10577-007-1163-9 [DOI] [PubMed] [Google Scholar]
- Nattestad M, Aboukhalil R, Chin CS, Schatz MC. Ribbon: intuitive visualization for complex genomic variation. Bioinformatics. 2021:37(3):413–415. 10.1093/bioinformatics/btaa680 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soh YQS, Alföldi J, Pyntikova T, Brown LG, Graves T, Minx PJ, Fulton RS, Kremitzki C, Koutseva N, Mueller JL, et al. Sequencing the mouse Y chromosome reveals convergent gene acquisition and amplification on both sex chromosomes. Cell. 2014:159(4):800–813. 10.1016/j.cell.2014.09.052 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spitzer M, Wildenhain J, Rappsilber J, Tyers M. BoxPlotR: a web tool for generation of box plots. Nat Methods. 2014:11(2):121. 10.1038/nmeth.2811 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subrini J, Turner J. Y chromosome functions in mammalian spermatogenesis. eLife. 2021:10:e67345. 10.7554/eLife.67345 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki H, Tsuchiya K, Takezaki N. A molecular phylogenetic framework for the Ryukyu endemic rodents Tokudaia osimensis and Diplothrix legata. Mol Phylogenet Evol. 2000:15(1):15–24. 10.1006/mpev.1999.0732 [DOI] [PubMed] [Google Scholar]
- Terao M, Ogawa Y, Takada S, Kajitani R, Okuno M, Mochimaru Y, Matsuoka K, Itoh T, Toyoda A, Kono T, et al. Turnover of mammal sex chromosomes in the Sry-deficient Amami spiny rat is due to male-specific upregulation of Sox9. Proc Natl Acad Sci U S A. 2022:119(49):e2211574119. 10.1073/pnas.2211574119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wintersinger JA, Wasmuth JD. Kablammo: an interactive, web-based BLAST results visualizer. Bioinformatics. 2015:31(8):1305–1306. 10.1093/bioinformatics/btu808 [DOI] [PubMed] [Google Scholar]
- Zhou C, McCarthy SA, Durbin R. YaHS: yet another Hi-C scaffolding tool. Bioinformatics. 2023:39(1):btac808. 10.1093/bioinformatics/btac808 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data used are publicly available.