Visual Abstract
Key Words: CRISPR-mediated adenine base editing, dilated cardiomyopathy, lamin A/ C, primate model
Highlights
-
•
For clinical findings, patients with LMD had an earlier stage of disease progression when receiving heart transplantation, and the degree of impairment to heart structure and function was relatively mild.
-
•
For the primate model, CRISPR-mediated adenine base editing constructed the first gene-edited primate model of DCM.
-
•
The primate model was highly similar to patients with LMD in terms of symptoms, disease progression, histopathology, and electrophysiological and transcriptional features.
-
•
Arrhythmia events in patients with LMD and the primate model predated impairment of heart structure and function.
Summary
To solve the clinical transformation dilemma of lamin A/C (LMNA)-mutated dilated cardiomyopathy (LMD), we developed an LMNA-mutated primate model based on the similarity between the phenotype of primates and humans. We screened out patients with LMD and compared the clinical data of LMD with TTN-mutated and mutation-free dilated cardiomyopathy to obtain the unique phenotype. After establishment of the LMNA c.357-2A>G primate model, primates were continuously observed for 48 months, and echocardiographic, electrophysiological, histologic, and transcriptional data were recorded. The LMD primate model was found to highly simulate the phenotype of clinical LMD. In addition, the LMD primate model shared a similar natural history with humans.
Dilated cardiomyopathy (DCM) is a type of cardiomyopathy characterized by left ventricular (LV) enlargement and reduced cardiac function, and it is an important cause of heart failure (HF) and sudden cardiac death worldwide.1 DCM has a heterogeneous etiology, with about 40% of patients having a genetic background, and pathogenic mutations mainly affecting the structural components of myocardial cells such as sarcomere, cytoskeleton, and nuclear envelope.2 Lamin A/C (LMNA) is a fibrous structure located in the inner nuclear membrane, playing a role in supporting nuclear membrane structure, maintaining nuclear function, and regulating transcription; mutations in its encoding gene account for 10% of genetic DCM cases.3 Patients with LMNA-mutated DCM (LMD) have a high incidence of adverse arrhythmic events (conduction block [73%], supraventricular arrhythmias [61%], and ventricular arrhythmias [50%]),4 and the progression rate of end-stage HF and the rate of requiring heart transplantation (HTx) are higher than with other types of genetic DCM, making it the worst prognosis type among the genetic DCMs.5
Implantable cardioverter-defibrillators (ICDs) and anti-HF therapy are the main treatment options for LMD, but their effectiveness is limited, and HTx is the only end-stage treatment for the disease.6 Targeted interventions against LMNA mutations and downstream signaling systems (eg, the mitogen-activated protein kinase signaling system) are important directions for precision treatment of LMD7,8; however, there are currently no targeted drugs with sufficient evidence-based medical support applied in actual clinical practice. The translational dilemma in LMD research partly arises from the disparity between LMD mice and humans. Although LMD mice successfully replicate alterations in cardiac structure and function, they do not exhibit the characteristic manifestations of conduction abnormalities and arrhythmias commonly observed in human patients with LMD.9
To bridge this gap in translational research, alternative models that can capture the complete spectrum of LMD-related cardiac phenotypes is necessary. Primates are highly similar to humans in terms of genome, transcriptome, proteome, and cardiovascular anatomy.10,11 The present study compared LMD vs TTN-mutated DCM (TMD) and a mutation-free DCM control group to extract unique clinical and pathologic features of LMD. Furthermore, we successfully generated the first LMD primate model using the adenine base editing (ABE) technique and conducted a comprehensive assessment of its cardiac structure, function, electrophysiology, and pathologic phenotypes compared with the clinical LMD cohort. RNA-sequencing was also performed to explore the transcriptional disturbance in LMD primates. This pioneering DCM primate model represents a significant advancement, providing a robust and consistent research platform for conducting preclinical experiments.
Methods
Study Population
The study population included patients with DCM (N = 302) who underwent HTx and gene testing from November 2004 to September 2017; this DCM cohort was of single-center origin (Department of Cardiac Surgery, Fuwai Hospital, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China). DCM was defined as the presence of LV dilatation or biventricular dilatation with systolic dysfunction, as measured by echocardiography and/or cardiac magnetic resonance imaging. LV dilatation was defined as LV end-diastolic diameter (LVEDD) >5.0 cm (female)/5.5 cm (male) or LVEDD >117% of the predicted value for body surface area and age. Systolic dysfunction was defined as an LV ejection fraction (LVEF) <45% and LV short-axis shortening rate <25%. Patients with secondary causative etiologies (including hypertension, valvular disease, congenital heart disease, and ischemic heart disease) were excluded.12
The normal control hearts (n = 9) in the study were derived from donor hearts originally considered for HTx but abandoned due to heart size or blood type mismatching factors. The donor hearts were derived from brain-dead donors whose hearts functioned normally in vivo and were transferred to the laboratory under refrigerated storage after cardioplegia infusion. Written informed consent for research use was obtained from families.13 The Ethics Committee of Fuwai Hospital approved and supervised this study.
Gene Capture and Sequencing
The genomic DNA was extracted from peripheral blood according to the manufacturer’s instructions (Blood DNA Extraction Kit, Enriching Biotechnology). Probands were screened with 189 cardiovascular disease–related genes (Supplemental Table 1), including 14 DCM-related genes (including DES, DMD, MYH7, MYBPC3, TNNT2, DSP, Titin [TTN], LMNA, MYH6, MYPN, sodium voltage-gated channel alpha subunit 5 [SCN5A], RBM20, ANKRD1, and RAF1) by captured next-generation sequencing using the HiSeq 2500 platform (Illumina). All variants were annotated by using several strategies. First, variants with a minor allele frequency <1% in 1,000 genomic data (1000g_all),14 esp6500siv2_all, and gnomAD data (gnomAD_ALL and gnomAD_EAS)15 were used. Second, only single nucleotide variants (SNVs) occurring in exons or splice sites (splicing junction 10 bp) were further analyzed because we were interested in amino acid changes. Third, synonymous SNVs that are not relevant to the amino acid alternation predicted by dbscSNV are then discarded, and the small fragment non-frameshift (<10 bp) indel in the repeat region defined by RepeatMasker are discarded. Fourth, variations are screened according to scores of SIFT,16 PolyPhen,17 MutationTaster,18 and CADD19 software. The potentially deleterious variations are reserved if the score of more than one-half of these four softwares support harmfulness of variations.20 Sites (>2 bp) that did not affect alternative splicing were removed. Fifth, for TTN variants, only truncated variants remained for pathogenesis analysis. Sanger sequencing was used to validate putatively pathogenic variants and screen family members.
To better predict the harmfulness of variation, the classification system of the American College of Medical Genetics and Genomics was used. The variations are classified into pathogenic (P), likely pathogenic (LP), variant of uncertain significance (VUS), likely benign, and benign.21 In addition, when selecting the DCM control group without gene mutation background, the criteria were non-P/LP/VUS carriers of cardiovascular diseases (n = 48) to reduce bias.
Clinical Characterization Collection
Clinical data, including HF symptoms (dyspnea, presyncope, syncope, abdominal distension, bilateral edema, and NYHA functional class) and palpitation, were collected from medical records. Data from electrocardiogram (ECG) monitoring, including atrial fibrillation, atrioventricular blocks, ventricular arrhythmias (premature ventricular complexes, ventricular premature contraction, and nonsustained ventricular tachycardia), were recorded; among these, major arrhythmic cardiovascular events were defined as arrhythmic syncope, sustained ventricular tachycardia, ventricular fibrillation, ICD implantation, cardiac resynchronization therapy (CRT), and CRT-defibrillator.22 LV and right ventricular structural remodeling and systolic function were assessed by using echocardiography.
Pathologic Examination and Quantitative Analysis
Full-thickness myocardial tissue was dissected. The tissue was fixed in 10% formaldehyde and then embedded in paraffin, and a 4-μm paraffin section was prepared by using a slicer (Leica). Hematoxylin and eosin staining and Masson staining were performed with standard protocols. The staining section images were obtained by a high-resolution digital scanner system (Aperio, Leica). Image-Pro Plus version 6.0 (Media Cybernetics) was used to quantify the areas of cardiomyocytes (red), adipose tissue (white), and fibrosis (blue). The myocyte disarray was semi-quantified by identifying areas of cellular interlacing, whorling, or herringbone patterns (Grade 0: 0% disarray area; Grade 1: 1%-25% disarray area; Grade 2: 25%-50% disarray area; Grade 3: >50% disarray area).23
Animals
All animals were housed at the Yunnan Key Laboratory of Primate Biomedical Research. All animal procedures were performed following the Association for Assessment and Accreditation of Laboratory Animal Care International for the ethical treatment of primates.
Preparation of Messenger RNA and Single Guide RNA
pCMV_ABEmax_P2A_GFP and pCMV_ABE7.10 plasmids were obtained from Addgene (#112101 and #102919). The plasmid was linearized with the restriction enzyme AgeI, and messenger RNA (mRNA) was synthesized and purified by using an in vitro RNA transcription kit (mMESSAGE mMACHINE T7 Ultra kit, Ambion). Single guide RNA (sgRNA) oligos were amplified and transcribed in vitro by using the GeneArt Precision gRNA Synthesis Kit (Thermo Fisher Scientific) and purified with the MEGAclear Kit (Thermo Fisher Scientific) according to the manufacturer’s instructions.
Oocyte Collection and In Vitro Fertilization
Oocyte collection and fertilization were performed as previously described.24 In brief, healthy female cynomolgus monkeys aged 5 to 8 years with regular menstrual cycles were selected as oocyte donors for superovulation, which was performed by intramuscular injection with recombinant human follitropin alfa (GONAL-f, Merck Serono) for 8 days, and then recombinant human chorionic gonadotropin alpha (Ovidrel, Merck Serono) administration on day 9. Oocytes were collected by laparoscopic follicular aspiration 32 to 35 hours after administration of recombinant human chorionic gonadotropin alpha. Follicular contents were placed in HEPES-buffered Tyrode’s albumin lactate pyruvate medium containing 0.3% body surface area at 37°C. Oocytes were stripped of cumulus cells by pipetting after a brief exposure (<1 minute) to hyaluronidase (0.5 mg/mL) in HEPES-buffered Tyrode’s albumin lactate pyruvate medium to allow visual selection of nuclear maturity metaphase II (first polar body present) oocytes. The maturity oocytes were immediately subjected to intracytoplasmic sperm injection and then cultured in CMRL-1066 containing 10% fetal bovine serum at 37°C in 5% carbon dioxide. Fertilization was confirmed by the presence of the second polar body and 2 pronuclei.
ABE mRNA/sgRNA Injection, Embryo Culture, and Transplantation
Six to eight hours after intracytoplasmic sperm injection, the zygotes were injected with a mixture of ABEmax or ABE7.10 mRNA (100 ng/μL) and sgRNA (50 ng/μL) with a total volume of 5 pL for each zygote. Microinjections were performed in the cytoplasm of oocytes using a microinjection system under standard conditions. Zygotes were then cultured in the chemically defined hamster embryo culture medium (HECM)-9 containing 10% fetal bovine serum (Gibco) at 37°C in 5% carbon dioxide to allow embryo development. The culture medium was replaced every other day until the blastocyst stage. The cleaved embryos with high quality at the two-cell to blastocyst stage were transferred into the oviduct of the matched recipient monkeys.
Fifteen monkeys were used as surrogate recipients. The earliest pregnancy diagnosis was performed by ultrasonography about 20 to 30 days after the embryo transfer. Both clinical pregnancy and the number of fetuses were confirmed by fetal cardiac activity and the presence of a yolk sac as detected by ultrasonography.
Fibroblast Isolation and Cell Culture
Ear skin was obtained and sterilized with 75% ethyl alcohol and washed with phosphate-buffered saline, then cut into pieces and adhered to the culture dish after removing the hair and fat tissues. The fibroblasts can be outgrown in about 1 week. All cells were cultured in Dulbecco's modified Eagle’s medium high-glucose (HyClone) medium containing 10% fetal bovine serum (Gibco), 1% GlutaMAX (Gibco), 1% penicillin/streptomycin (Gibco), 2.5 μg/mL Plasmocin (Invitrogen), and 1 mol/L tenofovir under 37°C, 5% carbon dioxide conditions.
Genomic DNA Extraction and Sequencing
After mRNA injection, embryos were cultured until the blastocyst stage, and the genomic DNA was extracted by using the REPLI-g Single Cell Kit (Qiagen). For fibroblast cells and heart tissues, the genomic DNA was extracted by using a Wizard Genomic DNA Purification Kit (#A1125, Promega) according to the manufacturer’s instructions. Sanger sequencing after polymerase chain reaction (PCR) was performed with primers as follows: F: 5′-GGCAAGCAGATGCAAACCAA-3′; R: 5′-CTCCAAGTCCTTCAGCCGAG-3′. The PCR products also were joined into a T vector, and 20 to 50 colonies were picked to TA-cloned sequencing every sample.
Reverse Transcription-PCR and Sequencing
Total RNA was extracted with TRIzol reagent (Invitrogen) and quantified by NanoDrop OneC (Thermo Fisher Scientific). Total RNA (1 μg) was reverse-transcribed to complementary DNA using the PrimeScript RT reagent Kit (Takara). PCR was performed with primers as follows: F: 5′-GCAGGAGCTCAATGATCGCT-3′; R: 5′-CTGTTCTCAGCATCCACCCG-3′. The reverse transcription-PCR products also were joined into a T vector, and 30 to 80 colonies were picked to TA-cloned sequencing every sample.
Western Blot and Semi-Quantitative Analysis
For cultured fibroblast cell protein analysis, cells were harvested and washed with cold phosphate-buffered saline and lysed with RIPA lysis buffer (Beyotime). For the monkey and human heart tissues, approximately 50 mg of heart tissue was washed by using cold phosphate-buffered saline solution and lysed with RIPA lysis buffer. The lysate was quantified by using the BCA Kit (Beyotime); approximately 20 μg proteins per lane were separated by sodium dodecyl sulphate-polyacrylamide gel electrophoresis and transferred to polyvinylidene difluoride membranes (Millipore). The membrane was blocked with 5% non-fat milk and probed with the indicated primary antibodies overnight at 37°C. Antibodies for Western blot were anti-LMNA (sc-7293, 1:1,000; Santa Cruz Biotechnology) and anti–glyceraldehyde 3-phosphate dehydrogenase (GAPDH) (60004-Ig, 1:1000; Proteintech). After incubation with the horseradish peroxidase–linked secondary antibodies (AP308P, 1:2000; Sigma), the signal was detected with the Pierce enhanced chemiluminescent Western Blotting Substrate kit (Thermo Fisher Scientific).
For the semi-quantitative analysis, Western blotting image data were processed with Image J software (National Institutes of Health), and the gray values of LMNA or GAPDH bands were calculated. LMNA was normalized with GAPDH.
Echocardiography in Primates
At baseline, 15 days, and 1, 2, 3, 6, 9, 12, 18, 24, 30, 36, 42, and 48 months, transthoracic echocardiography (Vivid iq, GE HealthCare) was performed in nonanesthetic primates in a supine position. The structural features of the heart (LVEF, fraction shortening, LVEDD, LV end-systolic diameter, LV posterior wall thickness, and left atrial diameter) were measured from two-dimensional images, and the observation sections included the LV long-axis section, the apical 4-chamber section, the apical 5-chamber section, and the apical 3-chamber section. In addition, spectral Doppler was used to measure the aortic peak systolic velocity.
ECG in Primates
For the evaluation of ECG parameters, the 12-lead ECG was evaluated by 2 independent senior electrophysiology experts of Fuwai Hospital who were blinded to the clinical, pathologic, and genetic diameters. Due to changes in the cardiac electrophysiological features under anesthesia,25 the 12-lead ECG was measured in nonanesthetic primates, given the inability of primates to remain stationary for an accurate baseline assessment. In addition, we continuously observed the cardiac electrophysiology of the primate model using the single-lead ECG results of the integrated monitoring device (Nicolet Monitor, Natus).
Circulatory Marker Testing in Primates
To assess HF and myocardial damage in primates, N-terminal pro–B-type natriuretic peptide, high-sensitivity troponin I, myoglobin, and creatine kinase-MB levels in plasma were measured at 42 months in all primates using a chemiluminescence analyzer (I2000, Abbott).
RNA-Sequencing
RNA-sequencing was performed to explore the transcriptional profile of the mutated type (MT) primates. LV samples from 2 sacrificed monkeys (192007 and 192011) were included, with 3 replicates per individual. Wild-type (WT) individuals (n = 6) served as a normal control group using data from the GSE219045 database. Detailed experiment and analysis information is described in the Supplemental Methods.
Statistical Analysis
All values are expressed as mean ± SD unless otherwise specified. Statistical analyses were performed by using Prism 8 (GraphPad Software). Unpaired Student’s t-test was performed when comparing data from 2 groups (normality and homogeneity of variance were tested by Shapiro-Wilk test and variance ratio test). Chi-square tests and Fisher exact tests were used to compare qualitative variables between 2 groups. For comparison of histologic features between 4 groups, based on normality (by Shapiro-Wilk test), the Kruskal-Wallis test (Holm-Bonferroni method was used to control for type I error for multiple comparisons), and Dunnett test were used. Kaplan-Meier method and log-rank test were used to compute survival curves. A 2-sided P value <0.05 was considered statistically significant. Statistical details of the RNA-sequencing analysis are described in the Supplemental Methods.
Results
Description of Genetic Mutations in the Clinical Study Population
Of the 302 patients who underwent HTx and genetic testing, 66 patients (21.9%) had P or LP genetic mutations associated with DCM (Supplemental Table 1). These patients were affected by mutations of TTN (n = 20 [6.6%]), MYBPC3 (n = 11 [3.6%]), MYPN (n = 10 [3.3%]), LMNA (n = 10 [3.3%]), MYH7 (n = 6 [2.0%]), TNNT2 (n = 5 [1.7%]), SCN5A (n = 4 [1.3%]), DSP (n = 3 [1.0%]), and RBM20 (n = 3 [1.0%]). In addition, 7 patients (2.3%) had a background of multiple mutated genes (Supplemental Figures 1A and 1B).
Among LMNA mutation types, 5 patients had missense mutations, 4 showed nonsense mutations, and 1 showed splice-site mutations. In addition, 1 patient (patient No. 2) carried two LMNA missense mutations (Figure 1A, Supplemental Table 2). Because TMD is the subtype with the highest proportion of hereditary DCM, its mild clinical characteristics (compared with the mutation-free DCM control group, the HTx rate and mortality were not affected by TTN mutation) have been relatively clear,26 and we included the TMD group as one of the controls. In the TMD group (n = 20), each patient carried one TTN mutation type, and the mutations were P or LP (Supplemental Table 2).
Figure 1.
Mapping of LMNA Variants and Histology Characteristics of Heart Tissue
(A) Diagrams representing the structure of lamin A/C (LMNA) and the distribution of LMNA variants. (B) Histologic characteristics. (B-i) Hematoxylin-eosin staining revealed nuclear malformation of cardiomyocytes in the group with LMNA-mutated dilated cardiomyopathy (LMD). (B-ii and B-iii) Histologic analysis suggested increased cell vacuolation and intercellular space in the LMD group. (B-iv) Heart tissue of end-stage LMD was characterized by fibrosis. (B-v) Heart tissue of LMD was characterized by myocyte disarray. DCM = dilated cardiomyopathy.
Through panel sequencing, 48 patients in the DCM control group without a P/LP and VUS cardiovascular disease gene mutation background were selected and compared with the LMD group. In addition to the gene/gene cluster of DCM, 191 cardiovascular disease genes (Supplemental Table 1) were included in the exclusion criteria of the DCM control group (Supplemental Figures 1C and 1D).
Clinical Phenotype of Patients With LMD
Compared with the TMD group and DCM control group, a relatively high incidence rate of arrhythmia events was a characteristic feature of patients with LMD. Arrhythmia events in LMD include atrial fibrillation (70.0%), third-degree atrioventricular block (60.0%), premature ventricular contractions (40.0%), nonsustained ventricular tachycardia (80.0%), and major arrhythmic cardiovascular events (70.0%).22 Moreover, the rate of ICD implantation, CRT, and CRT-defibrillator is 60.0% in patients with LMD. These arrhythmia-related events and treatments had a statistically significant higher incidence in the LMD group than in the DCM control and/or TMD groups.
In addition, in terms of holistic symptoms, the incidence of palpitation (70.0%), presyncope (70.0%), and syncope (40.0%) in patients with LMD was significantly higher than in patients of the DCM control and TMD groups. Because this cohort included patients who received HTx and were in end-stage disease, there were no significant differences in the symptoms of HF (including dyspnea, abdominal distension, bilateral edema, and NYHA functional score) between the LMD and the other 2 groups.
Patients with LMD received HTx at a time when the left ventricular structure (LVEDD) and function (LVEF) were less impaired, and the decision factors for relatively early surgery were refractory arrhythmia, quality of life impairment, and the disability-adjusted life years of high-risk patients as determined by a transplantation committee (Table 1). The natural history and cardiac events of each patient were recorded in the 3 groups (Supplemental Figure 2).
Table 1.
Baseline Characteristics of Patients With LMNA Variants and TTN Variants or DCM Control Subjects
| LMD (n = 10) | TMD (n = 20) | DCM Control (n = 48) |
P Value |
||
|---|---|---|---|---|---|
| LMD vs TMD | LMD vs DCM Control | ||||
| Male | 5 (50.0) | 18 (90.0) | 37 (77.1) | 0.026 | 0.12 |
| Age of onset, y | 37.5 ± 9.9 | 37.7 ± 13.2 | 41.2 ± 12.6 | 0.98 | 0.39 |
| Age of receiving HTx, y | 42.0 ± 10.8 | 43.4 ± 14.1 | 45.5 ± 13.3 | 0.79 | 0.45 |
| Dyspnea | 10 (100.0) | 17 (85.0) | 42 (87.5) | 0.53 | 0.58 |
| Palpitation | 7 (70.0) | 2 (10.0) | 11 (22.9) | 0.002 | 0.007 |
| Presyncope | 7 (70.0) | 1 (5.0) | 11 (22.9) | <0.001 | 0.007 |
| Syncope | 4 (40.0) | 0 (0.0) | 7 (14.6) | 0.008 | 0.083 |
| Abdominal distension | 5 (50.0) | 5 (20.0) | 16 (33.3) | 0.23 | 0.47 |
| Bilateral edema | 9 (90.0) | 11 (55.0) | 28 (58.3) | 0.10 | 0.077 |
| Arrhythmia | |||||
| Atrial fibrillation | 7 (70.0) | 6 (30.0) | 14 (29.2) | 0.056 | 0.027 |
| Third-degree ventricular block | 6 (60.0) | 0 (0.0) | 1 (2.1) | <0.001 | <0.001 |
| Ventricular premature contraction | 4 (40.0) | 2 (10.0) | 5 (10.4) | 0.14 | 0.039 |
| Premature ventricular complexes/24 h | 4 (40.0) | 5 (25.0) | 17 (35.4) | 0.43 | >0.99 |
| Nonsustained ventricular tachycardia | 8 (80.0) | 5 (25.0) | 11 (22.9) | 0.007 | 0.001 |
| MACE | 6 (60.0) | 1 (5.0) | 7 (14.6) | 0.002 | 0.005 |
| ICD/CRT/CRT-D | 5 (50.0) | 2 (10.0) | 9 (18.8) | 0.026 | 0.050 |
| NYHA functional class | 0.21 | 0.28 | |||
| I | 0 (0.0) | 0 (0.0) | 0 (0.0) | ||
| II | 0 (0.0) | 0 (0.0) | 1 (2.1) | ||
| III | 1 (10.0) | 7 (35.0) | 16 (33.3) | ||
| IV | 9 (90.0) | 13 (65.0) | 31 (64.6) | ||
| Cardiac ultrasound | (n = 9) | (n = 20) | (n = 33) | ||
| RV dilation | 5 (55.6) | 5 (25.0) | 16 (48.5) | 0.21 | >0.99 |
| RV wall motion abnormalities | 4 (44.4) | 3 (15.0) | 13 (39.4) | 0.16 | >0.99 |
| RVEDD, mm | 31.1 ± 9.1 | 26.0 ± 5.1 | 27.8 ± 5.5 | 0.078 | 0.18 |
| LV wall motion abnormalities | 9 (100.0) | 20 (100.0) | 33 (100.0) | >0.99 | >0.99 |
| Interventricular septal thickness, mm | 8.2 ± 1.0 | 8.0 ± 1.1 | 8.7 ± 1.5 | 0.60 | 0.42 |
| LVEDD, mm | 63.8 ± 5.2 | 71.2 ± 8.9 | 77.3 ± 12.8 | 0.029 | <0.001 |
| LVEF, % | 30.2 ± 5.2 | 24.7 ± 7.7 | 24.1 ± 5.9 | 0.066 | 0.007 |
Values are n (%) or mean ± SD. Bold values are <0.05, which are statistically significant.
CRT = cardiac resynchronization therapy; CRT-D = cardiac resynchronization therapy–defibrillator; DCM = dilated cardiomyopathy; HTx = heart transplantation; ICD = implantable cardioverter-defibrillator; LMD = lamin A/C–mutated dilated cardiomyopathy; LV = left ventricular; LVEDD = left ventricular end-diastolic diameter; LVEF = left ventricular ejection fraction; MACE = major arrhythmic cardiovascular events; RV = right ventricular; RVEDD = right ventricular end-diastolic diameter; TMD = TTN-mutated dilated cardiomyopathy.
Histologic Phenotype of Patients With LMD
Compared with analysis of the TMD and DCM control groups, histologic analysis of the LMD group revealed that cardiomyocytes of LMD had the typical characteristic of nuclear malformation, which was related to the decrease in the nuclear structure strength (Figure 1B-I, Supplemental Figure 3A).27 Cardiomyocyte vacuolation and increased intercellular space between cardiomyocytes were observed in the LMD group (Figures 1B-ii and 1B-iii, Supplemental Figures 3B and 3C).3,28 The patients with LMD had significant fibrosis characteristics (Figure 1B-iv); however, because this DCM cohort was in end-stage disease, there was no evidence that this pathologic feature was specific to LMD (Supplemental Figures 3D and 3E). In addition, significant myocyte disarray was observed in the LMD group (Figure 1B-v, Supplemental Table 3).23
ABEmax Was Used to Construct an LMNA Mutant Monkey Model
To better recapitulate human LMNA-related DCM phenotypes, ABE was used to generate the LMNA (c.357-2A>G, splice acceptor of Ex2) mutation in cynomolgus monkeys, which is a novel splice-site mutation in LMD human.29 An sgRNA Ex2-gA5 was designed to introduce the LMNA (c.357-2A>G) mutation and was co-injected into 61 monkey zygotes alone with ABEmax or ABE7.10 mRNA, and ABEmax displayed a higher editing efficiency (Supplemental Table 4). Thus, we chose to use ABEmax for constructing the non-human primate model carrying the LMNA (c.357-2A>G) mutation (Figure 2A).
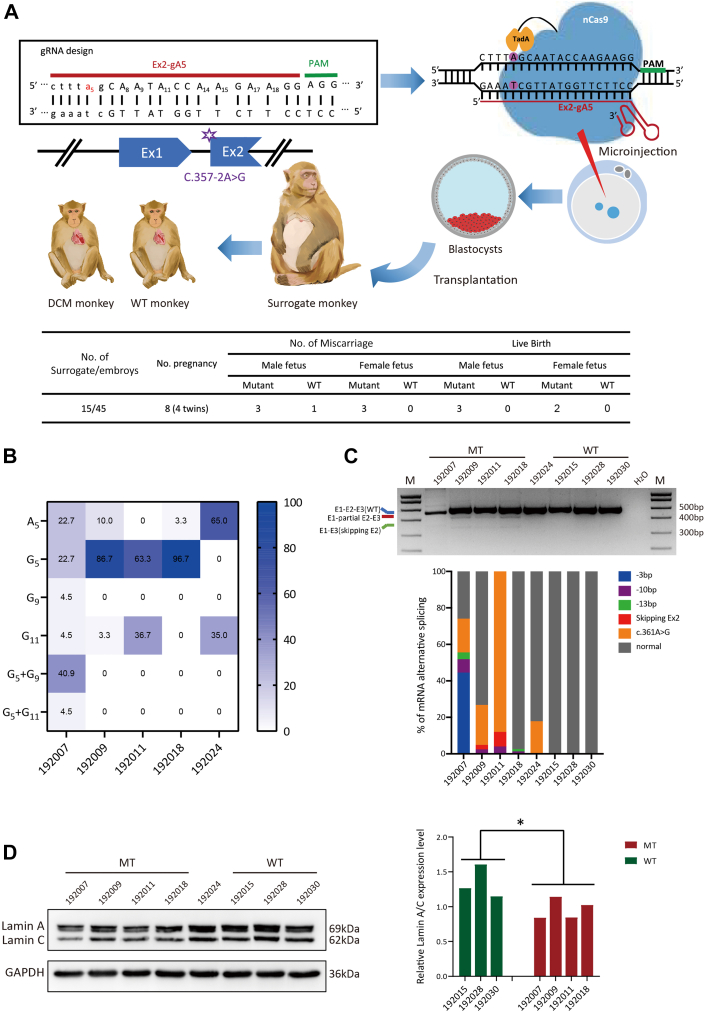
Figure 2.
Construction of LMNA c.357-2A>G Primate Model and Identification at the Molecular Level
(A) The schematic showed the process of generating DCM monkeys (upper) and general outcomes of LMNA c.357-2A>G mutations mediated by ABEmax in monkeys (below). (B) Heat map of the editing efficiency in mutated type (MT)-monkey fibroblast. (C) Reverse transcription–polymerase chain reaction to amplify LMNA gene Ex1 to Ex3 of MT and wild-type (WT) monkey’s fibroblast (upper) and percentage of messenger RNA (mRNA) alternative splicing–based TA-cloned sequencing (below). (D) Comparison of protein levels between the MT and WT monkeys via Western blot analysis. LMNA was normalized by glyceraldehyde 3-phosphate dehydrogenase (GAPDH). Compared with WT monkeys (n = 4), the MT monkeys (n = 3) exhibited lower LMNA protein expression using a 2-sided Student’s t-test (P = 0.047). ∗P < 0.05. ABE = adenine base editing; other abbreviations as in Figure 1.
After injection, 45 well-developed blastocysts were transplanted into 15 surrogate monkeys, and 8 surrogates were successfully impregnated. Finally, 5 full-term (approximately 150 days) monkey infants were born (3 male [192007, 192009, and 192011] and 2 female [192018 and 192024]). PCR and Sanger sequencing in fibroblast cells showed that the target site was edited with an efficiency of 66.3% to 96.7%. However, we observed bystander mutation A9>G9 and A11>G11, and monkey 192024 only displayed bystander mutation A11>G11 with an editing efficiency of 35.0% (Figure 2B, Supplemental Figure 4A). This mutation has not been reported to be associated with DCM; no DCM-like phenotypes were observed in the following study. We therefore did not include monkey 192024 in the MT or WT group for the subsequent statistical analysis.
To detect mutation at the mRNA level, we performed reverse transcription–PCR and Sanger sequencing. The results showed that there are abnormal mRNA types in the MT monkey; the types of alternative splicing included skipping exon2, 3 bp, 10 bp, or 13 bp deletion and single nucleotide alterations c.361A>G (Figure 2C). As a result of the mRNA alterations, we predicted the alterations of amino acid sequences (Supplemental Figure 4B). In addition, the Western blot analysis showed that the MT monkeys' LMNA was significantly reduced compared with WT monkeys in fibroblast (Figure 2D); this may be caused by aberrant mRNA splicing, which leads to premature termination of translation of the protein. Two MT monkeys were sacrificed, one at 3.5 years old (192011) and one at 4 years old (192007). We observed similar DNA editing efficiency, and the expression of LMNA was significantly reduced by approximately 35.0% in heart tissue (Supplemental Figures 4C to 4E). A similar result was observed in patients with LMD (LMNA-1,6,9,10) (Supplemental Table 2) which carried mutation of LMNA-truncated (Supplemental Figure 4F).
Clinical Phenotype of LMNA c.357-2A>G Primates
Echocardiography, circulatory markers associated with HF and myocardial injury, and cardiac electrophysiological characteristics were used to identify the clinical phenotypes of MT monkeys. It was found that except for monkey 192018 who did not develop symptoms of DCM for the time being, other MT monkeys showed cardiac remodeling, cardiac function reduction, myocardial damage, and arrhythmia phenotypes (Supplemental Figure 5).
Echocardiography revealed a significant decrease in cardiac function in the MT group, including LVEF (Figure 3A), fraction shortening (Figure 3B), and aortic peak systolic velocity (Figure 3C), except in the asymptomatic monkey (192018). In terms of cardiac remodeling, statistically significant LV dilatation (LV end-systolic diameter) was observed (Figure 3E). In addition, at the LVEDD, left atrial diameter, and LV posterior wall thickness levels, higher values were observed in the MT group, but these differences were not statistically different (Figures 3D, 3F, and 3G). There was no significant difference in body weight between the MT and WT groups (Figure 3H), which ruled out bias in the measurements of heart function and structure due to weight and growth differences.
Figure 3.
Echocardiographic and Weight Characteristics in Primates
The continuous change curve of echocardiographic and weight data between the WT group (192015, 192017, 192028, and 192030) and the MT group (192007, 192009, 192011, and 192018) (left). The month 42 echocardiographic and weight data between the WT group (192015, 192017, 192028, and 192030) and the symptomatic MT group (192007, 192009, and 192011) (right). (A) Left ventricular ejection fractions (LVEF). (B) Fraction shortening (FS). (C) Aortic peak systolic velocity (APSV). (D) Left ventricular end-diastolic diameter (LVEDD). (E) Left ventricular end-systolic diameter (LVESD). (F) Left ventricular posterior wall thickness (LVPW). (G) Left atrial diameter (LAD). (H) Weight. Values are presented as mean ± SD. Unpaired 2-sided Student’s t-test was performed when comparing data from 2 groups. ∗P < 0.05. ns = not significant; other abbreviations as in Figure 2.
By observing the symptoms in the LMNA c.357-2A>G primate model, we found that the arrhythmia phenotype in the symptomatic monkeys was earlier than the cardiac structural and functional abnormality (Figure 4A), which was similar to the natural history we observed in the LMD clinical cohort (Supplemental Figure 2). Fifty percent of the MT monkeys died by the fourth year (Figure 4B). Among them, monkey 192007 showed significant multi-organ edema (pericardial effusion: right ventricle lateral wall 0.4 cm, right ventricle anterior wall 0.2 cm, and left ventricle posterior wall 0.4 cm; bilateral pleural effusion [maximum depth was 2.2 cm]; and diffuse abdominal effusion), which was relieved after HF medication (spironolactone 2 mg/kg per day, furosemide 2 mg/kg per day, and valsartan-sacubitril 10 mg/kg per day) (Supplemental Figure 6).
Figure 4.
ECG and Histologic Characteristics in Primates
(1) Timeline of symptom onset in MT group primates. (B) Kaplan-Meier curves for the monkeys in the MT and WT groups. (C) Twelve-lead electrocardiogram (ECG) characteristics of WT (normal ECG phenotype) and MT (atrial fibrillation) monkeys. (D) Single-lead ECG characteristics of WT (normal ECG phenotype) and MT (192011, atrial flutter; 192007, atrial fibrillation; 192009, supraventricular premature contraction) individuals. (E) Histologic characteristics of LMNA c.357-2A>G primates (192007 and 192011). (Ei) Nuclear malformation of cardiomyocytes in LMNA c.357-2A>G primates. (Eii) Increased intercellular space in LMNA c.357-2A>G primates. (Eiii) Fibrosis of heart tissue in LMNA c.357-2A>G primates. (Eiv) Heart tissue of LMNA c.357-2A>G primates was characterized by myocyte disarray. HF = heart failure; other abbreviations as in Figures 1 and 2.
In terms of circulatory indicators, a noteworthy finding was the significant up-regulation of amino-terminal pro-brain natriuretic peptide, a recognized marker of HF, in monkeys 192007 and 192011 (Supplemental Figure 7A). In addition, high-sensitivity troponin I, a marker of myocardial injury, was observed to have higher expression in the MT group. High expression of myoglobin and creatine kinase-MB was observed only in monkey 192007 (Supplemental Figures 7B to 7D).
In terms of cardiac electrophysiological features, we captured atrial fibrillation in monkey 192007 using a 12-lead ECG (Figure 4C). A single-lead continuous monitoring ECG device was used, and atrial flutter was observed in monkey 192011, atrial fibrillation in monkey 192007, and supraventricular premature contraction in monkey 192009 (Figure 4D). The primates underwent biannual ECG monitoring; three (192007, 192011, and 192009) began to develop arrhythmias around age 3 years, and the arrhythmia events were continuously observed during subsequent monitoring.
Histologic Phenotype of LMNA c.357-2A>G Primates
Histologic analysis revealed that LMNA c.357-2A>G primates had similar histologic characteristics as humans. Compared with WT primates, the cardiomyocyte nucleus of LMNA c.357-2A>G primates (from 192007 and 192011) was malformed (Figure 4E-I, Supplemental Figure 8A). In addition, increased intercellular space (Figure 4E-ii, Supplemental Figure 8B), fibrosis (Figure 4E-iii, Supplemental Figures 8C and 8D), and the feature of myocyte disarray (Figure 4E-iv) were observed. However, no cardiomyocyte vacuolation was found in the LMNA c.357-2A>G primate model. Because vacuolar degeneration is associated with the accumulation of pathologic changes at the organelle level,30 the sacrificed monkeys might not have had a sufficient duration for pathologic changes to develop significant vacuolar degeneration in cardiomyocytes. Compared with clinical samples, the histologic findings of LMNA c.357-2A>G primates were highly similar to those of patients with LMD (Supplemental Figures 8E and 8F, Supplemental Table 5).
Transcriptional Profile of LMNA c.357-2A>G Primates
In terms of pathogenesis, in addition to the resulting structural defects and reduced mechanical strength of cardiomyocytes, LMNA mutation could affect transcription and gene localization through lamin–chromatin interactions.31 We compared transcriptional patterns in LV tissue of MT primates (n = 6, from 192007 and 192011) and WT primates (n = 6, from the GSE219045 database) using RNA-sequencing analysis. Results of the fragments per kilobase per million mapped fragments suggested that there was no batch effect between the data (Supplemental Figure 9A). We captured significantly differentially expressed genes in the MT group, with 714 genes down-regulated and 807 genes up-regulated (Figures 5A and 5B). It was found that there was a distinct clustering of samples from the MT and WT groups (Figure 5C, Supplemental Figure 9B).
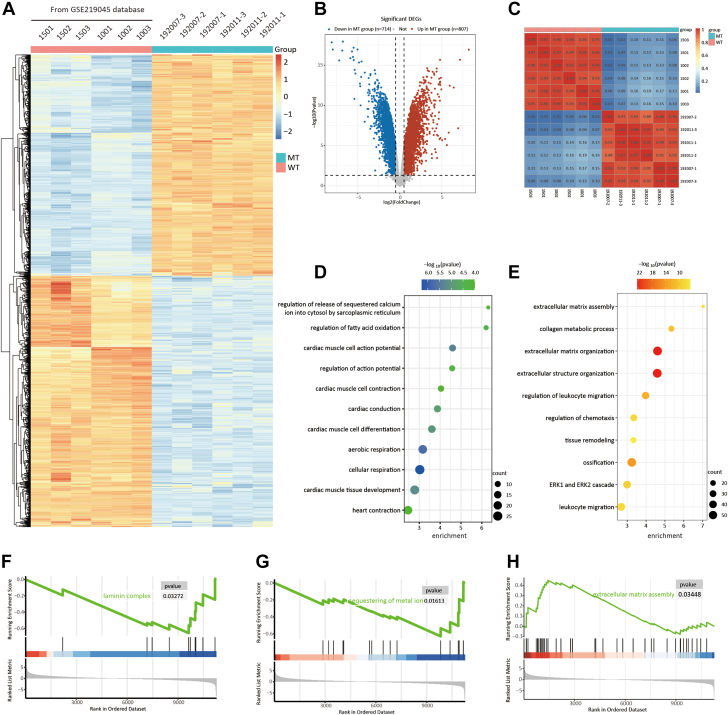
Figure 5.
RNA-Sequencing Analysis Revealed Transcriptional Alteration in LMNA c.357-2A>G Primates
(A) Heatmap showing the up-regulated and down-regulated differentially expressed genes (DEGs) in the left ventricular tissue of LMNA c.357-2A>G primates. (B) Volcano plot of DEGs. (C) Pearson correlation heatmap of RNA-sequencing showing distinct clustering of monkey samples from the WT and MT groups. (D) Gene ontology enrichment analysis of the down-regulated genes (n = 714) in the MT group. (E) Gene ontology enrichment analysis of the up-regulated genes (n = 807) in the MT group. (F) Gene set enrichment analysis (GSEA) plot showing the biological term of laminin complex (Gene Ontology [GO]: 0043256). (G) GSEA plot showing the biological term of sequestering of metal ion (GO: 0051238). (H) GSEA plot showing the biological term of extracellular matrix assembly (GO: 0085029). The Benjamini-Hochberg false discovery rate procedure was used for multiple testing. Abbreviations as in Figure 2.
Gene ontology analysis was performed on the up-regulated and down-regulated gene sets. The down-regulated differentially expressed genes were found to be enriched in cardiac action potential, cardiac conduction, cellular respiration, and fatty acid oxidation–related biological processes (Figure 5D, Supplemental Figure 9C); meanwhile, the up-regulated genes were mainly enriched in extracellular matrix and immune cell chemotaxis processes (Figure 5E, Supplemental Figure 9D). Gene set enrichment analysis suggested that the biological processes related to laminin complex (Figure 5F), ion exchange (Figure 5G), and extracellular matrix (Figure 5H) were disrupted in the myocardial tissue of MT monkeys. These results were consistent with the findings in the LMNA-related human induced pluripotent stem cells.32,33
Discussion
The rodent model is the most commonly used model to study LMD, among which the LMNA c.665 A>C mouse model developed by Arimura et al9 in 2005 is one of the representative models widely used. The pathogenesis of LMD, including alpha-tubulin acetylation,34 chromatin architecture disruption,35 and pathologic activation of the TP53 pathway,36 has been studied based on rodent models. However, due to the differences between the phenotypic simulation of LMD gene-edited rodents and actual clinical practice, there are issues with the translational efficiency of current LMD studies.
Large animal models have significant advantages in translational medicine compared with rodent models.37,38 In the field of DCM, large animal models, including gene-edited pig models with RBM20 editing by transcription activator–like effector nucleases, have been constructed, yielding successful induction of DCM phenotypes.39 Despite the similarities between pigs and humans in terms of anatomy, genome, and gene expression, primates have not yet been used for the construction of DCM models. With the mature application of gene-editing technology in primates,40 the development of an inherited DCM primate model becomes possible.
To provide a more intuitive phenotype for validation of the LMD monkey model, we observed the specificity of clinical and histologic features in patients with LMD using a DCM cohort. By comparing LMD with TMD and mutation-free DCM, it was found that LMD had a significant arrhythmia event rate, which was consistent with the characteristic clinical characteristics of LMD found in previous studies.41 These clinical data further confirmed that patients with TTN mutation types had milder clinical characteristics than patients with LMD.42 Moreover, in our cohort, we found that patients with LMD underwent HTx with a lower degree of cardiac remodeling (LVEDD) and function reduction (LVEF), and we found that it was related to the patient’s intolerance to arrhythmia and the priority consideration of the transplant committee. Histologically, the heart tissue of patients with LMD had characteristic pathologic changes (including nuclear malformation, cardiomyocyte vacuolation, increased intercellular space, and myocyte disarray). LMNA is the building block of the filamentous network lining the nucleoplasmic side of the inner nuclear membrane, and mutation of LMNA leads to impaired nuclear rigidity and nuclear malformation.43 The phenotype of cardiomyocyte vacuolation in patients with LMD was related to the presence of perinuclear autophagosomes and autolysosomes, which were used to dispose of nuclear wastes produced by nuclear damage.44 In addition, increased intercellular space between cardiomyocytes may be associated with cardiomyocyte atrophy caused by LMNA mutations.45 In this study, we found that patients with LMD, compared with TMD and DCM control patients, had a higher degree of myocyte disarray, which was related to cytoplasmic actin filament disturbance and structural remodeling in end-stage heart disease.45,46
To determine the characterization of clinical phenotypes in the LMNA c.357-2A>G primate model, we observed pathologic features, cardiac remodeling, reduced cardiac function, myocardial injury, and cardiac electrophysiological abnormalities in primates. In terms of pathologic features, the primate model exhibited typical features of nuclear malformation, fibrosis, enlargement of intercellular space, and myocytes disarray, but cardiomyocyte vacuolation was not found in the MT group, which might be related to the limited age of primates within 4 years. By comparison, the histologic phenotypes of human patients were not fully observed in the study of LMD mice, and the dominant histologic phenotypes of LMD mice were increased intercellular space and fibrosis28; however, fibrosis lacked LMD specificity. In terms of structural remodeling and cardiac function reduction, we found significant phenotypic changes as DCM by echocardiography, and circulatory markers were found to be significantly elevated in individuals with HF phenotypes. In addition, monkey 192011 (moribund in month 42) had high expression of circulatory markers of myocardial injury. In terms of cardiac electrophysiology, arrhythmic events were observed in MT individuals except for the asymptomatic monkey (192018).
Due to the differences in electrophysiology between the mouse and human,47 mouse models cannot faithfully simulate the characteristics of clinical arrhythmia.9 By comparing the natural history between the monkey model and the LMD clinical cohort, we found that arrhythmia events occurred earlier than the structural and functional abnormality of the heart, which, on the one hand, further supported the simulation of LMD in the primate model; on the other hand, it suggested that early intervention (including arrhythmia therapy, exercise intensity limitation, and avoiding medications related to heart damage) of LMD should take the occurrence of arrhythmia events as the initial time node and follow the cardiac structure and function closely after the occurrence of arrhythmia.48,49 It is difficult to observe a similar natural history in LMD mice due to premature death within few weeks,50 and because the treatment of LMD is a long-term process, LMD monkeys provide a platform for translational research with sufficient intervention time. At the transcriptional level, RNA-sequencing data of LMD monkeys confirmed the disturbance on action potential, ion transfer, fatty acid metabolism, and cardiomyocyte contraction. These results provided ideas for determining therapeutic targets, and the similarity in gene expression between monkeys and humans provides assurance for the translational value of these therapeutic targets.51
Study limitations
Our study focused on phenotypic analysis and lacked detailed exploration of the cellular biological mechanism of DCM caused by LMNA c.357-2A>G.
Conclusions
In this study, we constructed a primate model that is highly consistent with clinical LMD characteristics, which potentially has higher clinical translational value. In addition, because this primate model of LMD does not have the problems of fetal death and juvenile death, it can provide an experimental platform for the development of long-term therapeutic methods and continuous phenotype observation. We hope that this primate model can provide a bridge from basic research to clinical application for the precision intervention of LMD and gene therapy of hereditary diseases.
Perspectives.
COMPETENCY IN PATIENT CARE AND PROCEDURAL SKILLS: Based on the natural history of LMD, early cardiac risk management and diagnosis of carriers of LMNA mutations require a focus on the occurrence of arrhythmia events.
TRANSLATIONAL OUTLOOK: LMD primates exhibited ventricular dilatation and reduced cardiac function, consistent with the natural history in patients with LMD, and LMD primates showed high similarity to humans in electrophysiological, histologic, and transcriptional characteristics. These similarities supported the LMD primate model as a reliable platform for preclinical translational studies of LMNA-mutated DCM.
Funding Support and Author Disclosures
This work was supported by the National Natural Science Fund for Distinguished Young Scholars of China (82125004), the National Key Research and Development Program of China (2021YFA0805700), the National Natural Science Foundation of China (U2102204), and the Natural Science Foundation of Yunnan Province (grant numbers 202001BC070001 and 202102AA100053). The authors have reported that they have no relationships relevant to the contents of this paper to disclose.
Acknowledgments
The authors are grateful to Hong Wang, Ziyi Zhao, Baohong Tian, and all members of the animal facility of the State Key Laboratory of Primate Biomedical Research for excellent animal welfare and husbandry. In addition, the authors express their respect and gratitude to the experimental monkeys who were sacrificed for this study, and may they rest in peace. The animal data generated in the study are included in this published paper and its Supplemental Appendix. The human data are not publicly available due to privacy or ethical restrictions.
Footnotes
The authors attest they are in compliance with human studies committees and animal welfare regulations of the authors’ institutions and Food and Drug Administration guidelines, including patient consent where appropriate. For more information, visit the Author Center.
Appendix
For supplemental Methods, figures, and tables, please see the online version of this paper.
Contributor Information
Jiangping Song, Email: fwsongjiangping@126.com.
Yuyu Niu, Email: niuyy@lpbr.cn.
Appendix
References
- 1.Jefferies J.L., Towbin J.A. Dilated cardiomyopathy. Lancet. 2010;375:752–762. doi: 10.1016/S0140-6736(09)62023-7. [DOI] [PubMed] [Google Scholar]
- 2.Lakdawala N.K., Winterfield J.R., Funke B.H. Dilated cardiomyopathy. Circ Arrhythm Electrophysiol. 2013;6:228–237. doi: 10.1161/CIRCEP.111.962050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Captur G., Arbustini E., Bonne G., et al. Lamin and the heart. Heart. 2018;104:468–479. doi: 10.1136/heartjnl-2017-312338. [DOI] [PubMed] [Google Scholar]
- 4.Kayvanpour E., Sedaghat-Hamedani F., Amr A., et al. Genotype-phenotype associations in dilated cardiomyopathy: meta-analysis on more than 8000 individuals. Clin Res Cardiol. 2017;106:127–139. doi: 10.1007/s00392-016-1033-6. [DOI] [PubMed] [Google Scholar]
- 5.Paldino A., Dal Ferro M., Stolfo D., et al. Prognostic prediction of genotype vs phenotype in genetic cardiomyopathies. J Am Coll Cardiol. 2022;80:1981–1994. doi: 10.1016/j.jacc.2022.08.804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hasebe Y., Fukuda K., Nakano M., et al. Characteristics of ventricular tachycardia and long-term treatment outcome in patients with dilated cardiomyopathy complicated by lamin A/C gene mutations. J Cardiol. 2019;74:451–459. doi: 10.1016/j.jjcc.2019.03.019. [DOI] [PubMed] [Google Scholar]
- 7.Muchir A., Wu W., Choi J.C., et al. Abnormal p38α mitogen-activated protein kinase signaling in dilated cardiomyopathy caused by lamin A/C gene mutation. Hum Mol Genet. 2012;21:4325–4333. doi: 10.1093/hmg/dds265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Guénantin A.C., Jebeniani I., Leschik J., et al. Targeting the histone demethylase LSD1 prevents cardiomyopathy in a mouse model of laminopathy. J Clin Invest. 2021:131. doi: 10.1172/JCI136488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Arimura T., Helbling-Leclerc A., Massart C., et al. Mouse model carrying H222P-lmna mutation develops muscular dystrophy and dilated cardiomyopathy similar to human striated muscle laminopathies. Hum Mol Genet. 2005;14:155–169. doi: 10.1093/hmg/ddi017. [DOI] [PubMed] [Google Scholar]
- 10.Han L., Wei X., Liu C., et al. Cell transcriptomic atlas of the non-human primate macaca fascicularis. Nature. 2022;604:723–731. doi: 10.1038/s41586-022-04587-3. [DOI] [PubMed] [Google Scholar]
- 11.Liu Y.W., Chen B., Yang X., et al. Human embryonic stem cell-derived cardiomyocytes restore function in infarcted hearts of non-human primates. Nat Biotechnol. 2018;36:597–605. doi: 10.1038/nbt.4162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Pinto Y.M., Elliott P.M., Arbustini E., et al. Proposal for a revised definition of dilated cardiomyopathy, hypokinetic non-dilated cardiomyopathy, and its implications for clinical practice: a position statement of the esc working group on myocardial and pericardial diseases. Eur Heart J. 2016;37:1850–1858. doi: 10.1093/eurheartj/ehv727. [DOI] [PubMed] [Google Scholar]
- 13.Chen L., Hua K., Zhang N., et al. Multifaceted spatial and functional zonation of cardiac cells in adult human heart. Circulation. 2022;145:315–318. doi: 10.1161/CIRCULATIONAHA.121.055690. [DOI] [PubMed] [Google Scholar]
- 14.1000 Genomes Project Consortium. Auton A., Brooks L.D., et al. A global reference for human genetic variation. Nature. 2015;526:68–74. doi: 10.1038/nature15393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lek M., Karczewski K.J., Minikel E.V., Samocha K.E., Banks E., Fennell T., et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–291. doi: 10.1038/nature19057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kumar P., Henikoff S., Ng P.C. Predicting the effects of coding non-synonymous variants on protein function using the sift algorithm. Nat Protoc. 2009;4:1073–1081. doi: 10.1038/nprot.2009.86. [DOI] [PubMed] [Google Scholar]
- 17.Adzhubei I.A., Schmidt S., Peshkin L., et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–249. doi: 10.1038/nmeth0410-248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Schwarz J.M., Rodelsperger C., Schuelke M., Seelow D. Mutationtaster evaluates disease-causing potential of sequence alterations. Nat Methods. 2010;7:575–576. doi: 10.1038/nmeth0810-575. [DOI] [PubMed] [Google Scholar]
- 19.Kircher M., Witten D.M., Jain P., O’Roak B.J., Cooper G.M., Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet. 2014;46:310–315. doi: 10.1038/ng.2892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Muona M., Berkovic S.F., Dibbens L.M., et al. A recurrent de novo mutation in KCNC1 causes progressive myoclonus epilepsy. Nat Genet. 2015;47:39–46. doi: 10.1038/ng.3144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Richards S., Aziz N., Bale S., et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424. doi: 10.1038/gim.2015.30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Saguner A.M., Ganahl S., Baldinger S.H., et al. Usefulness of electrocardiographic parameters for risk prediction in arrhythmogenic right ventricular dysplasia. Am J Cardiol. 2014;113:1728–1734. doi: 10.1016/j.amjcard.2014.02.031. [DOI] [PubMed] [Google Scholar]
- 23.Cui H., Schaff H.V., Lentz Carvalho J., et al. Myocardial histopathology in patients with obstructive hypertrophic cardiomyopathy. J Am Coll Cardiol. 2021;77:2159–2170. doi: 10.1016/j.jacc.2021.03.008. [DOI] [PubMed] [Google Scholar]
- 24.Niu Y., Yu Y., Bernat A., et al. Transgenic rhesus monkeys produced by gene transfer into early-cleavage-stage embryos using a simian immunodeficiency virus-based vector. Proc Natl Acad Sci U S A. 2010;107:17663–17667. doi: 10.1073/pnas.1006563107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Owczuk R., Wujtewicz M.A., Zienciuk-Krajka A., Lasińska-Kowara M., Piankowski A., Wujtewicz M. The influence of anesthesia on cardiac repolarization. Minerva Anestesiol. 2012;78:483–495. [PubMed] [Google Scholar]
- 26.Herman D.S., Lam L., Taylor M.R., et al. Truncations of titin causing dilated cardiomyopathy. N Engl J Med. 2012;366:619–628. doi: 10.1056/NEJMoa1110186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Otomo J., Kure S., Shiba T., et al. Electrophysiological and histopathological characteristics of progressive atrioventricular block accompanied by familial dilated cardiomyopathy caused by a novel mutation of lamin A/C gene. J Cardiovasc Electrophysiol. 2005;16:137–145. doi: 10.1046/j.1540-8167.2004.40096.x. [DOI] [PubMed] [Google Scholar]
- 28.Chai R.J., Werner H., Li P.Y., et al. Disrupting the linc complex by AAV mediated gene transduction prevents progression of lamin induced cardiomyopathy. Nat Commun. 2021;12:4722. doi: 10.1038/s41467-021-24849-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zaragoza M.V., Fung L., Jensen E., et al. Exome sequencing identifies a novel lmna splice-site mutation and multigenic heterozygosity of potential modifiers in a family with sick sinus syndrome, dilated cardiomyopathy, and sudden cardiac death. PLoS One. 2016;11 doi: 10.1371/journal.pone.0155421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Takemura G., Kanamori H., Okada H., et al. Ultrastructural aspects of vacuolar degeneration of cardiomyocytes in human endomyocardial biopsies. Cardiovasc Pathol. 2017;30:64–71. doi: 10.1016/j.carpath.2017.06.012. [DOI] [PubMed] [Google Scholar]
- 31.Crasto S., My I., Di Pasquale E. The broad spectrum of LMNA cardiac diseases: from molecular mechanisms to clinical phenotype. Front Physiol. 2020;11:761. doi: 10.3389/fphys.2020.00761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Yang J., Argenziano M.A., Burgos Angulo M., Bertalovitz A., Beidokhti M.N., McDonald T.V. Phenotypic variability in iPSC-induced cardiomyocytes and cardiac fibroblasts carrying diverse lmna mutations. Front Physiol. 2021;12 doi: 10.3389/fphys.2021.778982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lee J., Termglinchan V., Diecke S., et al. Activation of PDGF pathway links LMNA mutation to dilated cardiomyopathy. Nature. 2019;572:335–340. doi: 10.1038/s41586-019-1406-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Le Dour C., Chatzifrangkeskou M., Macquart C., et al. Actin-microtubule cytoskeletal interplay mediated BY MRTF-A/SRF signaling promotes dilated cardiomyopathy caused by LMNA mutations. Nat Commun. 2022;13:7886. doi: 10.1038/s41467-022-35639-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Wang Y., Elsherbiny A., Kessler L., et al. Lamin A/C-dependent chromatin architecture safeguards naïve pluripotency to prevent aberrant cardiovascular cell fate and function. Nat Commun. 2022;13:6663. doi: 10.1038/s41467-022-34366-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Chen S.N., Lombardi R., Karmouch J., et al. DNA damage response/tp53 pathway is activated and contributes to the pathogenesis of dilated cardiomyopathy associated with lmna (lamin A/C) mutations. Circ Res. 2019;124:856–873. doi: 10.1161/CIRCRESAHA.118.314238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Pilz P.M., Ward J.E., Chang W.T., et al. Large and small animal models of heart failure with reduced ejection fraction. Circ Res. 2022;130:1888–1905. doi: 10.1161/CIRCRESAHA.122.320246. [DOI] [PubMed] [Google Scholar]
- 38.Blackwell D.J., Schmeckpeper J., Knollmann B.C. Animal models to study cardiac arrhythmias. Circ Res. 2022;130:1926–1964. doi: 10.1161/CIRCRESAHA.122.320258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Schneider J.W., Oommen S., Qureshi M.Y., et al. Dysregulated ribonucleoprotein granules promote cardiomyopathy in RBM20 gene-edited pigs. Nat Med. 2020;26:1788–1800. doi: 10.1038/s41591-020-1087-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kang Y., Dai S., Zeng Y., et al. Cloning and base editing of GFP transgenic rhesus monkey and off-target analysis. Sci Adv. 2022;8 doi: 10.1126/sciadv.abo3123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Lazarte J., Jurgens S.J., Choi S.H., et al. LMNA variants and risk of adult-onset cardiac disease. J Am Coll Cardiol. 2022;80:50–59. doi: 10.1016/j.jacc.2022.04.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ware J.S., Cook S.A. Role of titin in cardiomyopathy: from DNA variants to patient stratification. Nat Rev Cardiol. 2018;15:241–252. doi: 10.1038/nrcardio.2017.190. [DOI] [PubMed] [Google Scholar]
- 43.Broers J.L., Ramaekers F.C., Bonne G., Yaou R.B., Hutchison C.J. Nuclear lamins: Laminopathies and their role in premature ageing. Physiol Rev. 2006;86:967–1008. doi: 10.1152/physrev.00047.2005. [DOI] [PubMed] [Google Scholar]
- 44.Park Y.E., Hayashi Y.K., Bonne G., et al. Autophagic degradation of nuclear components in mammalian cells. Autophagy. 2009;5:795–804. doi: 10.4161/auto.8901. [DOI] [PubMed] [Google Scholar]
- 45.Dialynas G., Flannery K.M., Zirbel L.N., et al. LMNA variants cause cytoplasmic distribution of nuclear pore proteins in drosophila and human muscle. Hum Mol Genet. 2012;21:1544–1556. doi: 10.1093/hmg/ddr592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Perestrelo A.R., Silva A.C., Oliver-De La Cruz J., et al. Multiscale analysis of extracellular matrix remodeling in the failing heart. Circ Res. 2021;128:24–38. doi: 10.1161/CIRCRESAHA.120.317685. [DOI] [PubMed] [Google Scholar]
- 47.Elmadhun N.Y., Sabe A.A., Robich M.P., Chu L.M., Lassaletta A.D., Sellke F.W. The pig as a valuable model for testing the effect of resveratrol to prevent cardiovascular disease. Ann N Y Acad Sci. 2013;1290:130–135. doi: 10.1111/nyas.12216. [DOI] [PubMed] [Google Scholar]
- 48.Sidhu K., Castrini A.I., Parikh V., et al. The response to cardiac resynchronization therapy in LMNA cardiomyopathy. Eur J Heart Fail. 2022;24:685–693. doi: 10.1002/ejhf.2463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Guasch E., Mont L. Diagnosis, pathophysiology, and management of exercise-induced arrhythmias. Nat Rev Cardiol. 2017;14:88–101. doi: 10.1038/nrcardio.2016.173. [DOI] [PubMed] [Google Scholar]
- 50.Auguste G., Rouhi L., Matkovich S.J., et al. Bet bromodomain inhibition attenuates cardiac phenotype in myocyte-specific lamin A/C-deficient mice. J Clin Invest. 2020;130:4740–4758. doi: 10.1172/JCI135922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Qu J., Yang F., Zhu T., et al. A reference single-cell regulomic and transcriptomic map of cynomolgus monkeys. Nat Commun. 2022;13:4069. doi: 10.1038/s41467-022-31770-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.