Abstract
The gradual evolution of pharmacogenomics has shed light on the genetic basis for inter-individual drug response variations across diverse populations. This study aimed to identify pharmacogenomic variants that differ in Zhuang population compared with other populations and investigate their potential clinical relevance in gene-drug and genotypic-phenotypic associations. A total of 48 variants from 24 genes were genotyped in 200 Zhuang subjects using the Agena MassARRAY platform. The allele frequencies and genotype distribution data of 26 populations were obtained from the 1000 Genomes Project, followed by a comparison and statistical analysis. After Bonferroni correction, significant differences in genotype frequencies were observed of CYP3A5 (rs776746), ACE (rs4291), KCNH2 (rs1805123), and CYP2D6 (rs1065852) between the Zhuang population and the other 26 populations. It was also found that the Chinese Dai in Xishuangbanna, China, Han Chinese in Beijing, China, and Southern Han Chinese, China showed least deviation from the Zhuang population. The Esan in Nigeria, Gambian in Western Division, The Gambia, and Yoruba in Ibadan, Nigeria exhibited the largest differences. This was also proved by structural analysis, Fst analysis and phylogenetic tree. Furthermore, these differential variants may be associated with the pharmacological efficacy and toxicity of Captopril, Amlodipine, Lisinopril, metoclopramide, and alpha-hydroxymetoprolol in the Zhuang population. Our study has filled the gap of pharmacogenomic information in the Zhuang population and has provided a theoretical framework for the secure administration of drugs in the Zhuang population.
Keywords: Very important pharmacogene variant, Zhuang population, Single nucleotide variants, Potential clinical relevance, Personalized administration
Subject terms: Genetics, Risk factors
Introduction
Adverse drug reactions (ADRs) constitute a significant contributor to morbidity and mortality, ranking among the top 10 leading causes of death and disease in developed nations1,2. The characteristics of ADRs exhibit variability contingent upon factors such as genotype, age, gender, population, pathology, drug type, route of administration, and drug interaction3,4. According to Ingelman-Sundberg, genetic factors may account for approximately 10% to 20% of the occurrence of ADRs5. Genetic factors have been found to significantly influence pharmacokinetics, pharmacodynamics, and susceptibility to allergic reactions, resulting in changes in both local and systemic drug exposure and/or drug target functionality, ultimately impeding drug responses6. Recent investigations have elucidated the genetic underpinnings of ADRs7, thereby highlighting the close association between genetic factors and drug response.
Pharmacogenomics, as a key area of precision medicine, is the use of genomic and other “omic” information to personalize drugs selection and administration to avoid ADRs and maximize drugs therapeutic efficacy8,9. Pharmacogenomics accounted for 80% of the variations in drug treatment and safety. More than 400 genes were found to be involved in drug metabolism, and around 200 drug genes were linked to ADRs. It has been shown that substantial differences in distribution and frequencies of single nucleotide variants (SNVs) worldwide affect the key genes involved in drug absorption, distribution, metabolism, and elimination of abnormalities10. SNVs were a vast resource of genetic variation in humans, resulting in phenotypic differences among individuals11,12. The Pharmacogenetics and Pharmacogenomics Knowledge Base (PharmGKB; http://www.pharmgkb.org) that collects, organizes, and disseminates information on the impact of genetic variations in humans on drug responses. It provides free clinical-related information, including dosing guidelines, annotated drug labels, potentially viable gene-drug associations, and genotype–phenotype relationships13.
In recent years, numerous researchers have investigated very important pharmacogene (VIP) variants in ethnic minorities in China, such as the Tibetans14 and Lahu15. According to the 7th National Census, the Zhuang population totaled 15,721,956, ranking second only to the Han Chinese among the 56 populations. They are widespread in China’s Yunnan and Guizhou provinces, mainly in the Guangxi Zhuang Autonomous Region. Over a long period of time, they have developed customs and cultures with their own ethnic characteristics. However, we still have limited information on pharmacogenetic variants in the Zhuang population.
In this study, the VIP variants selected were derived from the PharmGKB, the SNP database of National Center for Biotechnology Information (NCBI, http://www.ncbi.nlm.nih.gov/SNP/), and the International HapMap Project (http://www.hapmap.org/), in addition to relevant pharmacogenomics literature. Then, the allele and genotype frequencies of the Zhuang population were compared with those of 26 other populations to obtain significant differences in SNVs after genotyping 200 unrelated Zhuang subjects from Yunnan province. The results of this study may complement current pharmacogenomics data of the Zhuang population, providing a theoretical basis for the safe use of drugs and predicting certain diseases in the Zhuang population.
Materials and methods
Study subjects
In total, 200 unrelated Zhuang subjects (110 females and 90 males) were recruited from Wen Shan in the Yunnan Province of China. The sample size and the proportion were determined using G*Power 3.1.9.2 software16. The participants were healthy based on their medical history and physical examination. Additionally, they had at least three generations of Zhuang ancestry, while none of the other populations had any known ancestral background. Subjects with chronic diseases, infectious diseases, drug or alcohol abuse, severe heart, liver or kidney dysfunction, immune disorders, pregnancy, and lactation were excluded. The informed consent forms have been signed by all subjects. According to the study protocol approved by the Clinical Research Ethics Committee of Northwest University, 5 mL of peripheral blood was collected from each subject and stored at 4 °C for 24 h.
Variants selection and genotyping
Through an extensive literature review on drug metabolism and toxicity, we identified 24 genes associated with these phenomena. By utilizing resources such as the PharmGKB database, the SNP database of NCBI, and the International HapMap Project, in addition to relevant pharmacogenomics literature, we selected variants linked to drug therapy responsiveness. A preliminary screening identified 59 variants. However, only homozygous genotypes were observed for 11 of these variants, making it impossible to compare the distribution and differences in genotype frequency. Consequently, these 11 variants were excluded from our analysis, leaving 48 variants for further investigation.
Genomic DNA was extracted from participants’ peripheral blood using GoldMag-Mini Whole Blood Genomic DNA Purification Kit (GoldMag Ltd., Xi’an, China). The concentration of genomic DNA was measured using NanoDrop 2000C spectrophotometer (Thermo Scientific, Waltham, MA, USA). Subsequently, multiplexed SNV MassEXTEND assays were designed using Agena MassARRAY Assay Design 4.0 software (San Diego, California, USA), which allowed for the design of PCR primers for the selected VIP variants. Agena MassARRAY RS1000 (San Diego, California, USA) was able to genotype the 48 VIP variants according to the manufacturer's instructions17. Finally, the data of SNV genotypes were collected and managed using Agena Typer 4.0 software18, as mentioned in previous studies.
Populations variation data
We downloaded the genotype data from the 1000 Genomes website (https://www.internationalgenome.org/). The 26 populations included: (1) Chinese Dai in Xishuangbanna, China (CDX); (2) Han Chinese in Beijing, China (CHB); (3) Southern Han Chinese, China (CHS); (4) Japanese in Tokyo, Japan (JPT); (5) Kinh in Ho Chi Minh City, Vietnam (KHV); (6) African Caribbeans in Barbados (ACB); (7) African Ancestry in Southwest USA (ASW); (8) Esan in Nigeria (ESN); (9) Gambian in Western Division, The Gambia (GWD); (10) Luhya in Webuye, Kenya (LWK); (11) Mende in Sierra Leone (MSL); (12) Yoruba in Ibadan, Nigeria (YRI); (13) Colombian in Medellin, Colombia (CLM); (14) Mexican Ancestry in Los Angeles, California (MXL); (15) Peruvian in Lima, Peru (PEL); (16) Puerto Rican in Puerto Rico (PUR); (17) Utah residents with Northern and Western European ancestry (CEU); (18) Finnish in Finland (FIN); (19) British in England and Scotland (GBR); (20) Iberian populations in Spain (IBS); (21) Toscani in Italy (TSI); (22) Bengali in Bangladesh (BEB); (23) Gujarati Indian in Houston, Texas (GIH); (24) Indian Telugu in the UK (ITU); (25) Punjabi in Lahore, Pakistan (PJL) and (26) Sri Lankan Tamil in the UK (STU).
Structure analysis and Fst analysis
The Structure 2.3.4 software was used to analyze the structure of 27 populations, and Arlequin3.1 software was used to evaluate pairwise Fst values for assessing the relationship between 27 population groups. In addition, MEGA11 software was utilized to plot phylogenetic tree.
Protein hazard prediction
We performed a functional analysis of missense variants using online tools such as Polyphen2 (http://genetics.bwh.harvard.edu/pph2/), SNAP2 (https://rostlab.org/services/snap/), Mutationassessor (http://mutationassessor.org/r3/), FATHMM (http://fathmm.biocompute.org.uk/index.html), and Mutationtaster (https://www.mutationtaster.org/) to assess the impact of SNVs mutations to predict protein function.
Mutant protein structure prediction
A single amino acid change has the potential to significantly affect protein activity and function. We downloaded the protein structures of CYP2D6 and KCNH2 from the PDB database (https://www.rcsb.org/) and utilized the Chimera v1.16 software to predict and visualize the mutant protein structures.
Statistical analysis
The data were compiled, ordered, and analyzed using Microsoft Excel 2019 (Microsoft, Redmond, WA, USA) and SPSS 26.0 (SPSS, Chicago, IL, USA). The χ2 test was utilized to estimate the Hardy–Weinberg equilibrium (HWE) and compare the divergences in genotype frequencies of 48 VIP variants between the Zhuang population and the other 26 populations. All statistical tests were two-tailed (p < 0.05). Bonferroni corrections were performed to determine the significance level. After the Bonferroni’s multiple tests, p < 4.01 × 10−5 was recognized as statistically significant.
Ethics approval
This study was conducted by the World Medical Association Declaration of Helsinki and was approved by the Northwestern University Clinical Research Ethics Committee (Approval number of Ethics Committee: 230,413,002). All subjects signed an informed consent form.
Results
Basic characteristics of candidate VIP variants
The 48 VIP variants on 24 genes that satisfied the HWE equation (p > 0.05) were collected in this study. Table 1 summarizes the fundamental characteristics of these variants, including gene name, SNVs ID, position, functional consequence, genotype frequency, and minor allele frequency (MAF) in the Zhuang population. Additionally, Table S1 shows the PCR primers for the gathered VIP variants.
Table 1.
Basic information of 48 selected VIP variants in the Zhuang population.
| Genes | SNVs ID | Chr | BP | Functional consequence | Zhuang Allele | Genotype frequencies | MAF | |||
|---|---|---|---|---|---|---|---|---|---|---|
| A | B | AA | AB | BB | ||||||
| CYP2J2 | rs11572325 | 1 | 59,896,030 | Intron Variant | T | A | 0 (0.000) | 21 (0.105) | 179 (0.895) | 0.053 |
| rs10889160 | 1 | 59,896,449 | Intron Variant | C | T | 1 (0.005) | 37 (0.185) | 162 (0.810) | 0.098 | |
| rs890293 | 1 | 59,926,822 | Upstream Transcript Variant | A | C | 0 (0.000) | 11 (0.055) | 189 (0.945) | 0.028 | |
| DPYD | rs1760217 | 1 | 97,137,438 | Genic Downstream Transcript Variant, Intron Variant | G | A | 11 (0.055) | 80 (0.400) | 109 (0.545) | 0.255 |
| rs1801159 | 1 | 97,515,839 | Coding Sequence Variant, Genic Downstream Transcript Variant, Intron Variant, Missense Variant | C | T | 26 (0.131) | 90 (0.455) | 82 (0.414) | 0.359 | |
| rs1801265 | 1 | 97,883,329 | Non-Coding Transcript Variant, Intron Variant, Coding Sequence Variant, 5 Prime UTR Variant, Missense Variant | G | A | 0 (0.000) | 40 (0.200) | 160 (0.800) | 0.100 | |
| PTGS2 | rs5275 | 1 | 186,673,926 | 3 Prime UTR Variant | G | A | 9 (0.046) | 67 (0.340 | 121 (0.614) | 0.216 |
| CACNA1S | rs12139527 | 1 | 201,040,054 | Missense Variant, Coding Sequence Variant, Intron Variant | G | A | 2 (0.010) | 36 (0.182) | 160 (0.808) | 0.101 |
| rs3850625 | 1 | 201,047,168 | Coding Sequence Variant, Missense Variant | A | G | 1 (0.005) | 7 (0.035) | 192 (0.960) | 0.023 | |
| RYR2 | rs2306238 | 1 | 237,550,803 | Intron Variant | A | G | 12 (0.060) | 62 (0.312) | 125 (0.628) | 0.216 |
| ABCG2 | rs2231142 | 4 | 88,131,171 | Coding Sequence Variant, Missense Variant | T | G | 8 (0.040) | 65 (0.327) | 126 (0.633) | 0.204 |
| rs2231137 | 4 | 88,139,962 | Coding Sequence Variant, Missense Variant | T | C | 27 (0.136) | 99 (0.497) | 73 (0.367) | 0.384 | |
| ADH1C | rs698 | 4 | 99,339,632 | Coding Sequence Variant, Non-Coding Transcript Variant, Missense Variant | C | T | 2 (0.010) | 49 (0.245) | 149 (0.745) | 0.133 |
| CYP3A5 | rs776746 | 7 | 99,672,916 | Intron Variant, splice acceptor variant, genic Downstream Transcript Variant, Downstream Transcript Variant | T | C | 19 (0.095) | 1 (0.005) | 180 (0.900) | 0.098 |
| CYP3A4 | rs2242480 | 7 | 99,763,843 | Intron Variant | T | C | 16 (0.08) | 83 (0.415) | 101 (0.505) | 0.288 |
| NAT2 | rs4646244 | 8 | 18,390,208 | Upstream Transcript Variant, Genic Upstream Transcript Variant, Intron Variant | A | T | 7 (0.035) | 66 (0.330) | 127 (0.635) | 0.200 |
| rs4271002 | 8 | 18,390,758 | Upstream Transcript Variant, Genic Upstream Transcript Variant, Intron Variant | C | G | 4 (0.020) | 50 (0.253) | 144 (0.727) | 0.146 | |
| rs1041983 | 8 | 18,400,285 | Coding Sequence Variant, Synonymous Variant | T | C | 24 (0.120) | 96 (0.480) | 80 (0.400) | 0.360 | |
| rs1801280 | 8 | 18,400,344 | Missense Variant, Coding Sequence Variant | C | T | 1 (0.005) | 6 (0.030) | 193 (0.965) | 0.020 | |
| rs1799929 | 8 | 18,400,484 | Coding Sequence Variant, Synonymous Variant | T | C | 1 (0.005) | 7 (0.035) | 192 (0.960) | 0.023 | |
| rs1799930 | 8 | 18,400,593 | Missense Variant, Coding Sequence Variant | A | G | 7 (0.035) | 69 (0.347) | 123 (0.618) | 0.209 | |
| rs1208 | 8 | 18,400,806 | Missense Variant, Coding Sequence Variant | G | A | 1 (0.005) | 7 (0.035) | 192 (0.960) | 0.023 | |
| rs1799931 | 8 | 18,400,860 | Missense Variant, Coding Sequence Variant | A | G | 4 (0.020) | 50 (0.250) | 146 (0.730) | 0.145 | |
| rs1495741 | 8 | 18,415,371 | None | A | G | 28 (0.146) | 90 (0.469) | 74 (0.385) | 0.380 | |
| ALOX5 | rs2115819 | 10 | 45,405,641 | Intron Variant | A | G | 8 (0.040) | 36 (0.181) | 155 (0.779) | 0.131 |
| CYP2C19 | rs12248560 | 10 | 94,761,900 | Upstream Transcript Variant | T | C | 0 (0.000) | 1 (0.005) | 199 (0.995) | 0.003 |
| rs4244285 | 10 | 94,781,859 | Coding Sequence Variant, Synonymous Variant | A | G | 17 (0.085) | 85 (0.425) | 98 (0.490) | 0.298 | |
| CYP2C8 | rs7909236 | 10 | 95,069,673 | Upstream Transcript Variant | T | G | 2 (0.010) | 44 (0.220) | 154 (0.770) | 0.120 |
| rs17110453 | 10 | 95,069,772 | Upstream Transcript Variant | C | A | 11 (0.055) | 82 (0.410) | 107 (0.535) | 0.260 | |
| CYP2E1 | rs3813867 | 10 | 133,526,101 | Non-Coding Transcript Variant, Upstream Transcript Variant | C | G | 4 (0.020) | 49 (0.245) | 147 (0.735) | 0.143 |
| rs6413432 | 10 | 133,535,040 | Intron Variant | A | T | 0 (0.000) | 44 (0.229) | 148 (0.771) | 0.115 | |
| rs2070676 | 10 | 133,537,633 | Intron Variant | G | C | 10 (0.050) | 57 (0.285) | 133 (0.665) | 0.193 | |
| KCNJ11 | rs5219 | 11 | 17,388,025 | Missense Variant, Stop Gained, 5 Prime UTR Variant, Intron Variant, Coding Sequence Variant | T | C | 12 (0.061) | 123 (0.628) | 61 (0.311) | 0.375 |
| SLCO1B1 | rs2306283 | 12 | 21,176,804 | Missense Variant, Coding Sequence Variant | A | G | 21 (0.106) | 71 (0.357) | 107 (0.538) | 0.284 |
| CYP1A2 | rs762551 | 15 | 74,749,576 | Intron Variant | C | A | 12 (0.060) | 90 (0.450) | 98 (0.490) | 0.285 |
| rs2472304 | 15 | 74,751,897 | Intron Variant | A | G | 2 (0.010) | 43 (0.216) | 154 (0.774) | 0.118 | |
| SULT1A1 | rs750155 | 16 | 28,609,251 | 5 Prime UTR Variant, Intron Variant, Genic Upstream Transcript Variant, Upstream Transcript Variant | C | T | 28 (0.144) | 118 (0.608) | 48 (0.247) | 0.448 |
| ACE | rs1800764 | 17 | 63,473,168 | None | C | T | 23 (0.116) | 102 (0.515) | 73 (0.369) | 0.374 |
| rs4291 | 17 | 63,476,833 | Upstream Transcript Variant | T | A | 0 (0.000) | 177 (0.898) | 20 (0.102) | 0.449 | |
| rs4267385 | 17 | 63,506,395 | None | T | C | 14 (0.070) | 71 (0.357) | 114 (0.573) | 0.249 | |
| CYP4F2 | rs2108622 | 19 | 15,879,621 | Missense Variant, Coding Sequence Variant | T | C | 3 (0.015) | 66 (0.332) | 130 (0.653) | 0.181 |
| rs3093105 | 19 | 15,897,578 | Missense Variant, Coding Sequence Variant | C | A | 0 (0.000) | 200 (1.000) | 0 (0.000) | 0.500 | |
| CYP2A6 | rs8192726 | 19 | 40,848,591 | Intron Variant | A | C | 7 (0.035) | 63 (0.315) | 130 (0.650) | 0.193 |
| SLC19A1 | rs1051298 | 21 | 45,514,912 | Intron Variant, 3 Prime UTR Variant | G | A | 34 (0.172) | 120 (0.606) | 44 (0.222) | 0.475 |
| rs1051296 | 21 | 45,514,947 | Intron Variant, 3 Prime UTR Variant | A | C | 24 (0.122) | 131 (0.668) | 41 (0.209) | 0.457 | |
| rs1131596 | 21 | 45,538,002 | Missense Variant, 5 Prime UTR Variant, Synonymous Variant, Genic Upstream Transcript Variant, Coding Sequence Variant | A | G | 32 (0.162) | 127 (0.644) | 38 (0.193) | 0.485 | |
| CYP2D6 | rs1065852 | 22 | 42,130,692 | Intron Variant, Missense Variant, Coding Sequence Variant | A | G | 45 (0.238) | 117 (0.619) | 27 (0.143) | 0.452 |
| KCNH2 | rs1805123 | 7 | 150,948,446 | Missense Variant, Coding Sequence Variant, Genic Downstream Transcript Variant | G | T | 151 (0.774) | 44 (0.226) | 0 (0.000) | 0.113 |
SNVs: single nucleotide variants, Chr: chromosome, BP: base pairs, ID: identity documents, MAF: minor allele frequency.
SNVs with significant differences in genotype frequencies between the Zhuang population and the other 26 populations
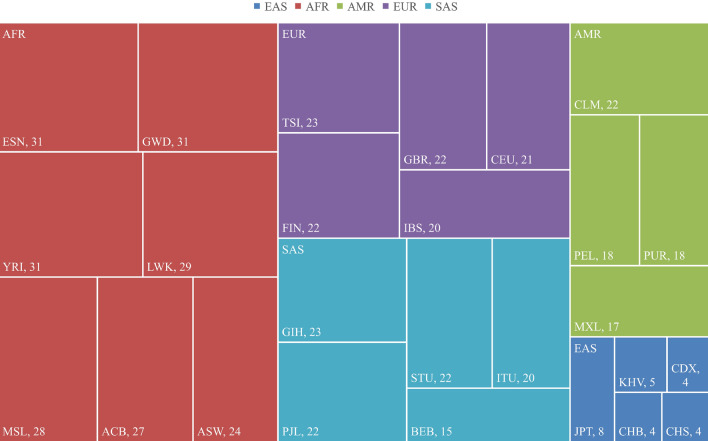
We compared the discrepancies in the genotype frequency distribution of the selected VIP variants between the Zhuang population and 26 other populations based on Chi-square tests. After the Bonferroni correction, the results were considered significant when p < 4.01 × 10–5. The number of SNVs with significant differences in genotype frequencies between the Zhuang population and 26 populations is shown in Fig. 1. The investigation demonstrated that the Zhuang population exhibited significant differences in four SNVs when compared to CDX, CHS, and KHV, and 31 SNVs when compared to ESN, GWD, and YRI. The Zhuang population exhibited differences in a number of SNVs when compared to other populations, including JPT (8), KHV (5), ACB (27), ASN (24), LWK (29), MSL (28), CLM (22), MXL (17), PEL (18), PUR (18), CEU (21), FIN (22), GBR (22), IBS (20), TSI (23), BEB (15), GIH (23), ITH (20), PJL (22), and STU (22). Furthermore, the Zhuang population showed significant differences in rs776746 (CYP3A5), rs4291 (ACE), and rs1805123 (KCNH2) compared to 26 other populations. Moreover, the Zhuang population exhibited significant differences in rs1065852 (CYP2D6) compared to 21 other populations (refer to Table 2 and Table S2).
Figure 1.
The amount of difference variants between the Zhuang population and 26 populations. The size of the rectangle indicates the number of different variants between the Zhuang population and the other 26 populations from five regions.
Table 2.
Genotype frequency distribution differences of 26 populations compared with the Zhuang population after Bonferroni’s multiple adjustment.
| SNVs ID | Genes | EAS | AFR | AMR | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CDX | CHB | CHS | JPT | KHV | ACB | ASW | ESN | GWD | LWK | MSL | YRI | CLM | ||
| rs11572325 | CYP2J2 | – | 0.376 | – | – | – | 8.15E-06 | 3.82E-05 | 1.96E-05 | 5.08E-05 | 1.51E-04 | 0.001 | 4.14E-05 | 0.024 |
| rs10889160 | CYP2J2 | 0.602 | 0.014 | 0.048 | 2.54E-04 | 0.201 | 1.47E-19 | 9.71E-11 | 2.94E-22 | 8.02E-16 | 8.45E-17 | 1.82E-19 | 9.11E-22 | 0.304 |
| rs890293 | CYP2J2 | – | 0.338 | – | – | – | 0.001 | 1.17E-06 | 8.44E-10 | 1.99E-07 | 1.56E-06 | 2.57E-11 | 3.43E-07 | - |
| rs1760217 | DPYD | 0.054 | 0.391 | 0.306 | 0.002 | 0.978 | 0.091 | 0.697 | 0.612 | 0.007 | 0.093 | 0.436 | 0.020 | 0.037 |
| rs1801159 | DPYD | 0.147 | 0.066 | 0.001 | 0.038 | 0.191 | 2.47E-07 | 6.90E-05 | 8.52E-05 | 7.27E-13 | 0.360 | 2.49E-11 | 4.31E-07 | 1.49E-04 |
| rs1801265 | DPYD | 0.012 | – | – | 0.028 | – | 1.64E-14 | 7.54E-19 | 1.56E-18 | 1.82E-21 | 3.06E-22 | 1.85E-14 | 1.34E-19 | 2.30E-05 |
| rs5275 | PTGS2 | 0.947 | 0.088 | 0.452 | 0.546 | 0.780 | 9.28E-19 | 2.50E-12 | 6.19E-23 | 5.67E-18 | 6.59E-19 | 1.92E-21 | 3.04E-23 | 5.31E-05 |
| rs12139527 | CACNA1S | 0.998 | 0.416 | 0.928 | 0.723 | 0.509 | 1.10E-23 | 1.42E-19 | 1.26E-31 | 3.10E-34 | 1.04E-26 | 1.08E-30 | 7.57E-31 | 0.451 |
| rs3850625 | CACNA1S | 0.417 | 0.004 | 0.353 | 0.762 | 0.103 | 0.774 | 0.503 | 0.131 | 0.098 | 0.131 | 0.174 | 0.109 | 1.52E-05 |
| rs2306238 | RYR2 | 0.160 | 0.253 | 0.212 | 0.128 | 0.634 | 1.70E-05 | 0.004 | 6.93E-08 | 1.02E-06 | 4.01E-06 | 4.26E-07 | 2.66E-06 | 0.133 |
| rs2231142 | ABCG2 | 0.784 | 0.014 | 0.284 | 0.003 | 0.001 | 1.30E-09 | 0.004 | 3.59E-11 | 1.42E-10 | 3.59E-11 | 5.40E-08 | 5.17E-12 | 0.063 |
| rs2231137 | ABCG2 | 0.093 | 0.090 | 0.387 | 2.13E-07 | 0.907 | 3.27E-17 | 1.40E-14 | 2.72E-17 | 2.11E-20 | 7.43E-11 | 6.58E-12 | 1.04E-17 | 2.06E-08 |
| rs698 | ADH1C | 0.577 | 0.005 | 0.043 | 0.048 | 0.146 | 0.336 | 0.409 | 0.044 | 0.201 | 0.946 | 0.361 | 0.048 | 0.001 |
| rs776746 | CYP3A5 | 8.15E-22 | 5.32E-20 | 1.62E-19 | 5.81E-22 | 6.45E-22 | 2.28E-43 | 5.81E-38 | 9.74E-50 | 1.68E-46 | 7.50E-47 | 2.72E-45 | 2.35E-51 | 1.03E-13 |
| rs2242480 | CYP3A4 | 0.715 | 0.415 | 0.420 | 0.540 | 0.660 | 9.64E-26 | 1.65E-17 | 4.74E-38 | 7.33E-31 | 2.10E-38 | 1.13E-34 | 9.72E-33 | 0.978 |
| rs4646244 | NAT2 | 0.456 | 0.936 | 0.162 | 0.078 | 2.04E-04 | 0.214 | 0.243 | 0.159 | 0.909 | 0.480 | 0.215 | 0.991 | 0.279 |
| rs4271002 | NAT2 | 2.87E-04 | 0.134 | 0.116 | 0.153 | 0.351 | 0.253 | 0.649 | 9.08E-05 | 2.06E-04 | 0.091 | 0.316 | 0.072 | 0.185 |
| rs1041983 | NAT2 | 0.002 | 0.988 | 0.068 | 0.248 | 7.78E-05 | 0.005 | 0.148 | 1.35E-04 | 0.613 | 0.206 | 0.001 | 0.003 | 0.220 |
| rs1801280 | NAT2 | 0.044 | 0.382 | 0.146 | 0.715 | 0.065 | 4.85E-19 | 4.86E-19 | 1.62E-19 | 2.96E-24 | 1.05E-27 | 5.47E-16 | 5.09E-16 | 9.78E-27 |
| rs1799929 | NAT2 | 0.077 | 0.498 | 0.224 | 0.762 | 0.201 | 6.55E-15 | 6.25E-15 | 6.18E-13 | 3.38E-20 | 1.00E-23 | 9.37E-12 | 3.67E-09 | 1.12E-24 |
| rs1799930 | NAT2 | 0.756 | 0.655 | 0.441 | 0.163 | 3.97E-04 | 0.339 | 0.141 | 0.200 | 0.499 | 0.197 | 0.381 | 0.923 | 0.338 |
| rs1208 | NAT2 | 0.077 | 0.498 | 0.132 | 0.762 | 0.058 | 2.31E-25 | 7.37E-22 | 4.83E-27 | 9.05E-33 | 1.32E-32 | 4.64E-24 | 2.36E-27 | 1.05E-26 |
| rs1799931 | NAT2 | 0.001 | 0.137 | 0.326 | 0.098 | 0.411 | 3.98E-04 | 0.018 | 4.30E-06 | 1.84E-06 | 1.26E-06 | 0.004 | 1.98E-04 | 0.024 |
| rs1495741 | NAT2 | 2.03E-04 | 0.908 | 0.014 | 0.509 | 7.39E-06 | 1.42E-05 | 1.05E-07 | 2.29E-04 | 5.63E-05 | 3.17E-09 | 0.002 | 0.007 | 3.46E-12 |
| rs2115819 | ALOX5 | 0.002 | 1.09E-04 | 0.040 | 0.003 | 0.051 | 1.49E-38 | 8.51E-26 | 4.34E-38 | 4.74E-42 | 7.49E-34 | 9.70E-33 | 6.76E-41 | 4.00E-17 |
| rs12248560 | CYP2C19 | – | – | – | – | – | 4.50E-25 | 1.48E-17 | 7.51E-22 | 1.18E-21 | 1.46E-15 | 7.69E-23 | 4.81E-21 | 2.28E-11 |
| rs4244285 | CYP2C19 | 0.606 | 0.197 | 0.100 | 0.800 | 0.905 | 0.001 | 0.002 | 0.054 | 1.47E-05 | 0.077 | 0.010 | 0.001 | 2.61E-06 |
| rs7909236 | CYP2C8 | 0.620 | 0.769 | 0.102 | 0.158 | 0.011 | 0.004 | 0.454 | 1.43E-06 | 2.42E-07 | 6.42E-05 | 8.66E-06 | 4.56E-07 | 1.26E-06 |
| rs17110453 | CYP2C8 | 0.927 | 0.100 | 0.026 | 0.005 | 0.575 | 1.75E-13 | 1.36E-09 | 7.43E-14 | 5.84E-17 | 3.10E-15 | 1.82E-13 | 1.21E-15 | 8.33E-06 |
| rs3813867 | CYP2E1 | 0.779 | 0.003 | 0.140 | 0.282 | 0.067 | 0.007 | 0.131 | 0.081 | 0.056 | 5.46E-05 | 0.023 | 0.097 | 0.820 |
| rs6413432 | CYP2E1 | 1.81E-06 | 3.90E-07 | 7.62E-06 | 7.17E-06 | 2.88E-07 | – | – | 0.119 | 0.031 | – | 0.020 | 0.237 | 0.039 |
| rs2070676 | CYP2E1 | 0.021 | 0.266 | 0.677 | 0.423 | 0.058 | 1.78E-25 | 1.45E-13 | 1.04E-24 | 3.05E-26 | 8.00E-31 | 2.96E-26 | 3.50E-24 | 0.073 |
| rs5219 | KCNJ11 | 1.13E-04 | 0.001 | 0.230 | 0.013 | 0.034 | 2.98E-19 | 6.49E-09 | 5.01E-28 | 3.94E-29 | 3.34E-27 | 3.40E-25 | 8.78E-30 | 8.52E-07 |
| rs2306283 | SLCO1B1 | 0.082 | 0.287 | 0.086 | 0.207 | 0.153 | 0.086 | 0.650 | 6.16E-05 | 0.036 | 0.004 | 0.082 | 0.023 | 1.48E-06 |
| rs762551 | CYP1A2 | 0.302 | 0.041 | 0.136 | 0.006 | 0.832 | 0.013 | 0.219 | 3.48E-06 | 0.001 | 4.53E-08 | 0.002 | 2.86E-05 | 0.321 |
| rs2472304 | CYP1A2 | 0.380 | 0.969 | 0.170 | 0.008 | 0.187 | 0.205 | 0.568 | 7.35E-06 | 4.86E-05 | 8.28E-05 | 4.17E-05 | 8.77E-06 | 3.48E-12 |
| rs750155 | SULT1A1 | 0.385 | 5.03E-05 | 0.596 | 3.47E-06 | 0.075 | 0.002 | 1.01E-04 | 8.71E-10 | 9.53E-13 | 8.18E-06 | 1.42E-06 | 6.52E-08 | 0.072 |
| rs1800764 | ACE | 0.587 | 0.085 | 0.573 | 0.175 | 0.068 | 1.07E-26 | 4.95E-17 | 4.17E-34 | 6.98E-40 | 4.78E-29 | 5.98E-37 | 2.54E-39 | 0.080 |
| rs4291 | ACE | 2.44E-17 | 9.38E-22 | 7.87E-18 | 4.20E-15 | 1.50E-21 | 5.93E-16 | 4.76E-15 | 4.06E-17 | 1.65E-14 | 3.91E-21 | 1.36E-13 | 4.08E-22 | 2.53E-16 |
| rs4267385 | ACE | 0.548 | 0.987 | 0.413 | 0.580 | 0.412 | 5.38E-26 | 1.54E-16 | 1.35E-27 | 1.57E-33 | 7.19E-35 | 1.15E-31 | 1.41E-31 | 9.42E-07 |
| rs2108622 | CYP4F2 | 0.623 | 0.352 | 0.696 | 0.003 | 0.042 | 0.010 | 0.041 | 4.11E-06 | 1.84E-04 | 0.058 | 0.047 | 1.65E-05 | 0.004 |
| rs3093105 | CYP4F2 | – | – | – | 2.56E-59 | – | 1.15E-37 | 1.86E-30 | 5.69E-33 | 2.44E-41 | 7.01E-38 | 2.46E-36 | 3.69E-35 | 5.66E-44 |
| rs8192726 | CYP2A6 | 0.464 | 0.225 | 0.044 | 0.115 | 0.061 | 1.18E-04 | 0.043 | 0.010 | 3.55E-05 | 0.006 | 0.001 | 0.005 | 3.04E-06 |
| rs1051298 | SLC19A1 | 0.075 | 0.073 | 0.398 | 0.065 | 0.035 | 0.004 | 0.187 | 0.573 | 1.26E-04 | 0.046 | 0.054 | 0.149 | 0.097 |
| rs1051296 | SLC19A1 | 0.011 | 0.001 | 0.275 | 0.004 | 0.011 | 0.050 | 0.003 | 0.001 | 0.002 | 1.82E-04 | 0.034 | 0.011 | 0.003 |
| rs1131596 | SLC19A1 | 0.049 | 0.017 | 0.756 | 0.013 | 0.289 | 3.32E-09 | 0.024 | 0.002 | 3.17E-14 | 1.37E-10 | 3.85E-09 | 2.32E-08 | 0.022 |
| rs1065852 | CYP2D6 | 0.001 | 0.030 | 0.001 | 5.47E-07 | 0.005 | 7.31E-23 | 4.59E-18 | 8.94E-30 | 1.34E-28 | 8.17E-37 | 3.59E-20 | 9.51E-29 | 1.66E-19 |
| rs1805123 | KCNH2 | 1.30E-51 | 3.59E-60 | 4.17E-60 | 3.57E-59 | 3.69E-53 | 4.89E-62 | 6.05E-51 | 1.44E-64 | 1.08E-66 | 1.29E-63 | 1.58E-61 | 1.60E-66 | 1.41E-42 |
| SNVs ID | Genes | AMR | EUR | SAS | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MXL | PEL | PUR | CEU | FIN | GBR | IBS | TSI | BEB | GIH | ITU | PJL | STU | ||
| rs11572325 | CYP2J2 | – | – | 7.89E-05 | 0.039 | 4.74E-04 | 0.158 | 0.044 | 0.049 | – | – | 0.372 | – | – |
| rs10889160 | CYP2J2 | 0.458 | 0.407 | 1.48E-04 | 0.382 | 1.34E-05 | 0.341 | 0.016 | 0.179 | 0.785 | 0.157 | 0.203 | 0.859 | 0.221 |
| rs890293 | CYP2J2 | – | – | 0.099 | – | 0.045 | 0.252 | – | – | – | – | 0.138 | – | – |
| rs1760217 | DPYD | 0.819 | 0.206 | 0.002 | 0.862 | 0.008 | 0.013 | 0.026 | 0.066 | 0.184 | 0.671 | 1.07E-05 | 0.004 | 4.81E-05 |
| rs1801159 | DPYD | 0.020 | 0.044 | 2.76E-04 | 3.12E-06 | 2.63E-07 | 3.21E-05 | 0.001 | 0.009 | 2.43E-09 | 4.13E-10 | 4.57E-15 | 1.80E-10 | 4.52E-13 |
| rs1801265 | DPYD | 9.52E-06 | 0.013 | 1.00E-06 | 0.024 | 2.85E-09 | 0.008 | 1.50E-04 | 6.20E-05 | 0.003 | 1.16E-11 | 4.66E-10 | 6.11E-09 | 0.001 |
| rs5275 | PTGS2 | 0.005 | 3.84E-05 | 0.017 | 8.01E-05 | 0.457 | 0.099 | 0.007 | 0.053 | 7.24E-05 | 1.07E-04 | 0.001 | 5.02E-09 | 1.52E-05 |
| rs12139527 | CACNA1S | 0.021 | 0.170 | 0.017 | 0.752 | 0.916 | 0.626 | 0.879 | 0.877 | 0.030 | 0.006 | 0.575 | 0.337 | 0.355 |
| rs3850625 | CACNA1S | 1.91E-04 | 6.07E-05 | 0.127 | 0.001 | 3.04E-11 | 1.07E-07 | 0.001 | 3.77E-05 | 2.24E-08 | 1.17E-19 | 2.68E-12 | 4.27E-11 | 7.10E-09 |
| rs2306238 | RYR2 | 0.004 | 0.001 | 0.021 | 0.531 | 0.301 | 0.794 | 0.827 | 0.499 | 0.154 | 0.358 | 0.012 | 0.006 | 0.003 |
| rs2231142 | ABCG2 | 0.958 | 0.280 | 0.010 | 0.030 | 0.002 | 0.155 | 9.27E-05 | 1.28E-05 | 0.066 | 8.87E-05 | 0.010 | 0.009 | 0.001 |
| rs2231137 | ABCG2 | 0.001 | 0.237 | 1.38E-08 | 3.39E-19 | 4.61E-13 | 2.17E-18 | 4.02E-18 | 5.28E-15 | 0.002 | 8.91E-09 | 1.42E-12 | 1.07E-11 | 1.12E-05 |
| rs698 | ADH1C | 2.91E-04 | 0.354 | 2.54E-10 | 1.08E-17 | 5.94E-19 | 6.67E-14 | 9.12E-07 | 9.77E-07 | 0.302 | 2.55E-05 | 0.002 | 1.37E-07 | 2.15E-08 |
| rs776746 | CYP3A5 | 1.45E-17 | 1.19E-09 | 1.15E-15 | 1.61E-05 | 7.01E-06 | 1.47E-06 | 2.30E-07 | 1.23E-06 | 5.72E-24 | 5.31E-21 | 2.07E-22 | 1.10E-21 | 2.70E-22 |
| rs2242480 | CYP3A4 | 0.068 | 1.40E-09 | 0.137 | 5.05E-10 | 1.86E-08 | 2.29E-08 | 4.78E-06 | 5.34E-08 | 0.088 | 0.479 | 0.060 | 0.010 | 0.130 |
| rs4646244 | NAT2 | 0.122 | 0.004 | 0.801 | 0.003 | 0.067 | 0.018 | 0.042 | 0.072 | 0.210 | 2.12E-05 | 4.70E-04 | 4.15E-04 | 9.95E-07 |
| rs4271002 | NAT2 | 0.535 | 0.015 | 0.200 | 0.027 | 0.584 | 0.779 | 0.775 | 0.647 | 0.241 | 0.930 | 0.415 | 0.909 | 0.788 |
| rs1041983 | NAT2 | 0.185 | 0.005 | 0.184 | 0.072 | 0.224 | 0.098 | 0.480 | 0.269 | 0.870 | 0.112 | 0.289 | 0.201 | 0.003 |
| rs1801280 | NAT2 | 4.50E-24 | 1.34E-20 | 1.78E-27 | 3.56E-31 | 2.53E-33 | 5.89E-33 | 1.73E-36 | 2.44E-32 | 1.69E-24 | 5.68E-24 | 1.80E-25 | 7.16E-31 | 1.41E-20 |
| rs1799929 | NAT2 | 1.84E-22 | 3.15E-19 | 1.08E-23 | 2.04E-30 | 3.87E-31 | 1.07E-30 | 1.03E-35 | 1.30E-31 | 2.11E-22 | 7.73E-21 | 7.06E-23 | 1.13E-26 | 9.82E-19 |
| rs1799930 | NAT2 | 0.148 | 0.001 | 0.782 | 0.007 | 0.233 | 0.019 | 0.031 | 0.097 | 0.263 | 1.11E-05 | 0.001 | 2.15E-04 | 2.54E-07 |
| rs1208 | NAT2 | 6.69E-28 | 7.16E-20 | 9.13E-27 | 4.93E-29 | 1.85E-30 | 1.02E-30 | 9.31E-36 | 2.80E-32 | 1.17E-26 | 2.75E-23 | 3.12E-24 | 7.53E-31 | 2.92E-22 |
| rs1799931 | NAT2 | 0.479 | 0.030 | 0.073 | 3.37E-07 | 2.68E-04 | 4.28E-05 | 9.27E-05 | 4.32E-06 | 0.242 | 0.004 | 0.007 | 0.008 | 0.055 |
| rs1495741 | NAT2 | 1.12E-07 | 0.004 | 7.38E-11 | 1.93E-13 | 1.66E-15 | 8.63E-16 | 6.88E-20 | 6.96E-14 | 1.18E-12 | 1.13E-17 | 3.28E-15 | 6.09E-21 | 2.66E-19 |
| rs2115819 | ALOX5 | 1.53E-11 | 2.59E-07 | 1.26E-15 | 4.32E-25 | 3.56E-20 | 3.24E-19 | 6.08E-21 | 7.28E-22 | 9.82E-17 | 4.99E-24 | 1.42E-21 | 1.00E-16 | 2.89E-15 |
| rs12248560 | CYP2C19 | 5.71E-09 | - | 7.65E-16 | 1.15E-20 | 1.68E-19 | 2.79E-21 | 1.33E-19 | 1.33E-19 | - | 7.28E-12 | 5.54E-12 | 3.36E-12 | 1.76E-11 |
| rs4244285 | CYP2C19 | 4.23E-04 | 2.99E-09 | 2.39E-05 | 4.15E-05 | 0.058 | 3.32E-04 | 9.91E-05 | 6.12E-08 | 0.702 | 0.235 | 0.157 | 0.095 | 0.018 |
| rs7909236 | CYP2C8 | 1.27E-05 | 8.08E-10 | 0.070 | 4.53E-05 | 3.35E-05 | 0.006 | 0.071 | 0.132 | 0.010 | 0.001 | 0.010 | 0.016 | 0.227 |
| rs17110453 | CYP2C8 | 0.001 | 1.36E-08 | 6.18E-05 | 2.48E-06 | 0.273 | 3.68E-05 | 0.189 | 9.12E-06 | 0.105 | 0.094 | 0.158 | 0.369 | 0.017 |
| rs3813867 | CYP2E1 | 0.832 | 0.780 | 0.013 | 0.013 | 3.79E-04 | 1.63E-04 | 1.83E-05 | 0.003 | 1.20E-05 | 1.00E-06 | 1.16E-06 | 2.78E-06 | 3.11E-07 |
| rs6413432 | CYP2E1 | 0.114 | 0.008 | 0.015 | 0.361 | 0.015 | 0.060 | 0.078 | 0.335 | 0.002 | 3.46E-06 | 0.003 | 0.106 | 0.002 |
| rs2070676 | CYP2E1 | 0.299 | 0.113 | 0.020 | 0.074 | 0.005 | 0.024 | 0.253 | 0.051 | 0.578 | 0.322 | 0.705 | 0.492 | 0.886 |
| rs5219 | KCNJ11 | 0.103 | 0.106 | 3.07E-04 | 0.384 | 0.037 | 0.004 | 0.018 | 0.001 | 0.002 | 0.001 | 0.013 | 2.17E-06 | 0.005 |
| rs2306283 | SLCO1B1 | 7.78E-10 | 3.53E-07 | 1.09E-04 | 8.99E-11 | 5.18E-09 | 5.74E-13 | 2.70E-11 | 1.20E-13 | 0.004 | 0.001 | 0.022 | 9.27E-08 | 0.001 |
| rs762551 | CYP1A2 | 0.029 | 2.17E-04 | 0.842 | 0.503 | 0.255 | 0.321 | 0.054 | 0.022 | 0.001 | 5.13E-06 | 2.96E-05 | 5.70E-05 | 1.26E-07 |
| rs2472304 | CYP1A2 | 1.01E-04 | 0.989 | 2.22E-19 | 1.27E-31 | 3.26E-24 | 3.18E-30 | 2.90E-28 | 7.29E-22 | 0.084 | 0.234 | 0.790 | 0.001 | 0.921 |
| rs750155 | SULT1A1 | 0.218 | 1.17E-12 | 6.26E-05 | 0.004 | 0.330 | 0.005 | 0.002 | 3.76E-04 | 3.83E-12 | 6.89E-08 | 6.20E-11 | 2.10E-08 | 1.25E-17 |
| rs1800764 | ACE | 0.129 | 0.001 | 0.079 | 0.011 | 0.172 | 0.114 | 0.384 | 0.001 | 0.685 | 0.147 | 0.210 | 0.580 | 0.949 |
| rs4291 | ACE | 3.27E-19 | 2.23E-26 | 9.16E-16 | 1.28E-17 | 1.51E-12 | 2.50E-18 | 1.41E-15 | 8.52E-19 | 3.55E-16 | 6.16E-16 | 5.81E-22 | 5.34E-17 | 6.88E-16 |
| rs4267385 | ACE | 3.89E-04 | 0.875 | 4.81E-10 | 3.57E-11 | 2.23E-10 | 1.49E-12 | 1.72E-13 | 3.89E-22 | 0.074 | 4.20E-04 | 0.131 | 2.91E-04 | 0.004 |
| rs2108622 | CYP4F2 | 0.188 | 0.123 | 0.002 | 0.016 | 0.345 | 0.001 | 3.00E-06 | 3.05E-06 | 8.50E-09 | 7.07E-11 | 6.54E-09 | 5.88E-08 | 5.84E-10 |
| rs3093105 | CYP4F2 | 1.39E-42 | 1.35E-55 | 1.54E-44 | 1.21E-44 | 1.84E-49 | – | 7.51E-35 | 1.34E-39 | 6.52E-45 | 1.47E-46 | 1.85E-42 | 3.34E-42 | 2.80E-46 |
| rs8192726 | CYP2A6 | 3.72E-04 | 4.46E-05 | 3.48E-06 | 2.15E-05 | 0.090 | 6.09E-06 | 1.27E-05 | 1.38E-04 | 0.096 | 0.213 | 0.019 | 0.356 | 0.019 |
| rs1051298 | SLC19A1 | 2.48E-05 | 1.15E-04 | 0.092 | 0.005 | 0.083 | 3.25E-04 | 0.008 | 0.023 | 0.196 | 0.029 | 0.205 | 0.342 | 0.021 |
| rs1051296 | SLC19A1 | 6.24E-08 | 9.47E-08 | 0.001 | 7.75E-05 | 0.002 | 1.09E-06 | 1.71E-04 | 1.69E-04 | 0.092 | 0.025 | 0.002 | 0.003 | 0.003 |
| rs1131596 | SLC19A1 | 4.37E-07 | 1.21E-05 | 0.006 | 0.045 | 0.022 | 2.49E-06 | 0.001 | 0.003 | 0.002 | 0.001 | 4.88E-04 | 2.09E-04 | 0.001 |
| rs1065852 | CYP2D6 | 6.20E-18 | 8.28E-30 | 1.30E-21 | 8.70E-14 | 4.41E-23 | 5.54E-14 | 4.52E-21 | 8.40E-18 | 1.55E-13 | 1.24E-22 | 1.76E-20 | 4.59E-27 | 3.85E-23 |
| rs1805123 | KCNH2 | 6.73E-41 | 3.21E-51 | 3.55E-41 | 3.82E-40 | 1.03E-44 | 4.00E-36 | 8.87E-38 | 9.24E-37 | 4.67E-34 | 3.55E-39 | 1.83E-37 | 3.46E-42 | 7.87E-42 |
Bolded font indicates significant results.
EAS, East Asian; SAS, South Asian; EUR, European; AFR, African; AMR, American; CDX, Chinese Dai in Xishuangbanna, China; CHB, Han Chinese in Beijing, China; CHS, Southern Han Chinese, China; JPT, Japanese in Tokyo, Japan; KHV, Kinh in Ho Chi Minh City; Vietnam; BEB, Bengali in Bangladesh; GIH, Gujarati Indian in Houston, Texas; ITU, Indian Telugu in the UK; PJL, Punjabi in Lahore, Pakistan; STU, Sri Lankan Tamil in the UK; CEU, Western European ancestry; FIN, Finnish in Finland; GBR, British in England and Scotland; IBS, Iberian populations in Spain; TSI, Toscani in Italy; ACB, African Caribbeans in Barbados; ASW, African Ancestry in Southwest USA; ESN, Esan in Nigeria; GWD, Gambian in Western Divisions, The Gambia; LWK, Luhya in Webuye, Kenya; MSL, Mende in Sierra Leone; YRI, Yoruba in Ibadan, Nigeria; CLM, Colombian in Medellin, Colombia; MXL, Mexican Ancestry in Los Angeles, Colombia; PEL, Peruvian in Lima, Peru; PUR, Puerto Rican in Puerto Rico.
Genetic structure analysis of 27 populations
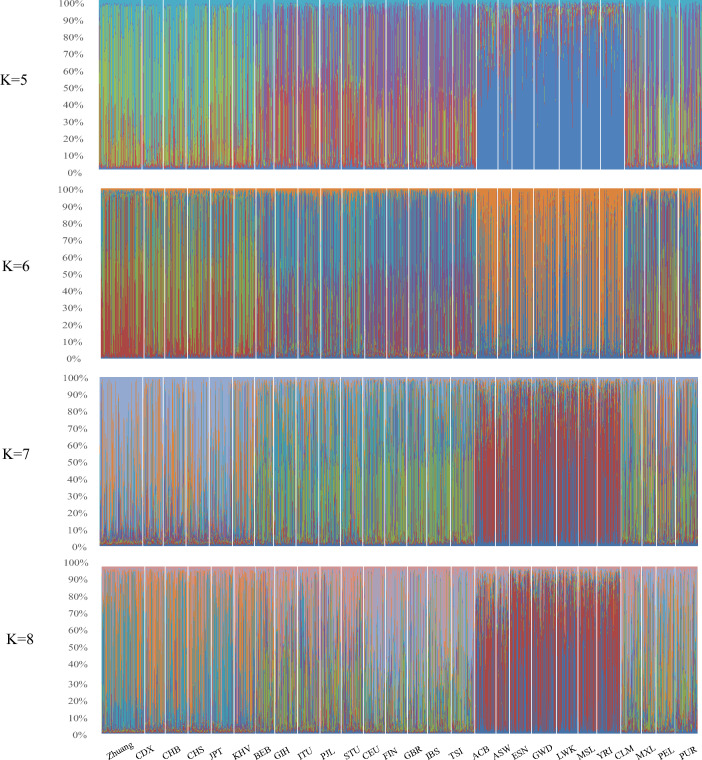
A model-based clustering approach was used to analyze the genetic structure of the 27 populations distributed in Africa, America, East Asia, Europe and South Asia to further analyze their relationship. Based on the Structure 2.3.1 Software, different K values ranging from 5 to 8 were hypothetically considered in structure analysis. When K = 5, the groups were divided into 5 subgroups based on the relative majority probability of assigning individuals to subgroups (subgroup 1: GWD and LWK; Subgroup 2: BEB, CEU, FIN, GBR, IBS, TSI, CLM, MXL and PUR; Subgroup 3: Zhuang, CDX, CHB, CHS, JPT, KHV and PUR; Subgroup 4: GIH, ITU, PJL and STU; Subgroup 5: ACB, ASW, ESN, MSL and YRI). It can be observed from Fig. 2 that Zhuang population have a stronger affinity with CDX, CHB, CHS, JPT, KHV and PUR. This is consistent with the results in Table 2.
Figure 2.
Structure analysis of the genetic relationship between the Zhuang population and the other 26 populations. K denotes the possible numbers of parental population clusters. Each vertical bar represents a sample, dividing into color sections. K = 5 were utilized to evaluate the relationship between Zhuang and 26 populations.
The pairwise Fst values were used to assess relationships among 27 populations, as shown in Table 3 and Fig. 3A. The Fst values between the Zhuang population and the East Asian population (CDX, CHB, CHS, JPT and KHV) were small, which were 0.065, 0.068, 0.066, 0.073 and 0.067, respectively (Table 3). Smaller Fst values indicate closer relationships between the two groups and suggest that they share similar genetic backgrounds. The result is confirmed by the phylogenetic trees of 27 populations shown in Fig. 3B.
Table 3.
Pairwise Fst values among the Zhuang and 26 populations.
| Zhuang | CDX | CHB | CHS | JPT | KHV | BEB | GIH | ITU | PJL | STU | CEU | FIN | GBR | IBS | TSI | ACB | ASW | ESN | GWD | LWK | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Zhuang | 0.000 | ||||||||||||||||||||
| CDX | 0.065 | 0.000 | |||||||||||||||||||
| CHB | 0.068 | 0.013 | 0.000 | ||||||||||||||||||
| CHS | 0.066 | 0.009 | 0.005 | 0.000 | |||||||||||||||||
| JPT | 0.073 | 0.026 | 0.012 | 0.015 | 0.000 | ||||||||||||||||
| KHV | 0.069 | 0.007 | 0.013 | 0.006 | 0.024 | 0.000 | |||||||||||||||
| BEB | 0.092 | 0.067 | 0.064 | 0.064 | 0.060 | 0.066 | 0.000 | ||||||||||||||
| GIH | 0.108 | 0.082 | 0.081 | 0.079 | 0.072 | 0.078 | 0.010 | 0.000 | |||||||||||||
| ITU | 0.103 | 0.082 | 0.076 | 0.075 | 0.069 | 0.078 | 0.006 | 0.005 | 0.000 | ||||||||||||
| PJL | 0.122 | 0.095 | 0.094 | 0.091 | 0.082 | 0.091 | 0.010 | 0.007 | 0.006 | 0.000 | |||||||||||
| STU | 0.108 | 0.077 | 0.075 | 0.070 | 0.065 | 0.071 | 0.012 | 0.011 | 0.007 | 0.011 | 0.000 | ||||||||||
| CEU | 0.140 | 0.123 | 0.124 | 0.122 | 0.111 | 0.122 | 0.057 | 0.054 | 0.058 | 0.047 | 0.069 | 0.000 | |||||||||
| FIN | 0.138 | 0.115 | 0.115 | 0.110 | 0.104 | 0.115 | 0.049 | 0.042 | 0.047 | 0.037 | 0.057 | 0.010 | 0.000 | ||||||||
| GBR | 0.146 | 0.128 | 0.131 | 0.128 | 0.119 | 0.128 | 0.057 | 0.056 | 0.059 | 0.047 | 0.069 | 0.005 | 0.010 | 0.000 | |||||||
| IBS | 0.129 | 0.116 | 0.119 | 0.114 | 0.105 | 0.115 | 0.042 | 0.040 | 0.044 | 0.033 | 0.053 | 0.011 | 0.012 | 0.009 | 0.000 | ||||||
| TSI | 0.125 | 0.114 | 0.117 | 0.116 | 0.105 | 0.115 | 0.048 | 0.045 | 0.050 | 0.041 | 0.059 | 0.011 | 0.015 | 0.008 | 0.006 | 0.000 | |||||
| ACB | 0.237 | 0.182 | 0.195 | 0.196 | 0.180 | 0.188 | 0.146 | 0.145 | 0.141 | 0.136 | 0.143 | 0.179 | 0.175 | 0.186 | 0.162 | 0.158 | 0.000 | ||||
| ASW | 0.198 | 0.147 | 0.158 | 0.161 | 0.146 | 0.154 | 0.107 | 0.107 | 0.102 | 0.097 | 0.108 | 0.138 | 0.134 | 0.144 | 0.125 | 0.121 | 0.009 | 0.000 | |||
| ESN | 0.283 | 0.228 | 0.236 | 0.240 | 0.221 | 0.232 | 0.188 | 0.187 | 0.179 | 0.176 | 0.180 | 0.230 | 0.225 | 0.237 | 0.213 | 0.206 | 0.013 | 0.021 | 0.000 | ||
| GWD | 0.280 | 0.226 | 0.235 | 0.239 | 0.219 | 0.231 | 0.179 | 0.178 | 0.170 | 0.168 | 0.176 | 0.213 | 0.208 | 0.219 | 0.194 | 0.187 | 0.011 | 0.019 | 0.013 | 0.000 | |
| LWK | 0.283 | 0.227 | 0.241 | 0.242 | 0.226 | 0.233 | 0.181 | 0.179 | 0.174 | 0.165 | 0.174 | 0.222 | 0.213 | 0.225 | 0.199 | 0.194 | 0.015 | 0.023 | 0.012 | 0.016 | 0.000 |
| MSL | 0.277 | 0.222 | 0.233 | 0.236 | 0.218 | 0.227 | 0.188 | 0.188 | 0.181 | 0.177 | 0.181 | 0.229 | 0.225 | 0.235 | 0.210 | 0.204 | 0.009 | 0.022 | 0.006 | 0.010 | 0.016 |
| YRI | 0.272 | 0.220 | 0.228 | 0.233 | 0.213 | 0.226 | 0.184 | 0.183 | 0.176 | 0.174 | 0.178 | 0.222 | 0.218 | 0.229 | 0.204 | 0.197 | 0.008 | 0.020 | 0.004 | 0.008 | 0.015 |
| CLM | 0.107 | 0.077 | 0.084 | 0.082 | 0.075 | 0.080 | 0.029 | 0.031 | 0.033 | 0.028 | 0.042 | 0.021 | 0.021 | 0.020 | 0.018 | 0.017 | 0.128 | 0.093 | 0.172 | 0.157 | 0.164 |
| MXL | 0.112 | 0.084 | 0.082 | 0.087 | 0.080 | 0.090 | 0.032 | 0.042 | 0.040 | 0.032 | 0.052 | 0.040 | 0.036 | 0.039 | 0.038 | 0.037 | 0.159 | 0.116 | 0.200 | 0.190 | 0.190 |
| PEL | 0.125 | 0.091 | 0.097 | 0.098 | 0.101 | 0.102 | 0.088 | 0.100 | 0.098 | 0.096 | 0.108 | 0.110 | 0.100 | 0.110 | 0.107 | 0.104 | 0.201 | 0.165 | 0.242 | 0.237 | 0.231 |
| PUR | 0.103 | 0.078 | 0.084 | 0.082 | 0.074 | 0.084 | 0.028 | 0.035 | 0.032 | 0.026 | 0.039 | 0.023 | 0.021 | 0.021 | 0.018 | 0.017 | 0.112 | 0.079 | 0.153 | 0.139 | 0.143 |
| LWK | MSL | YRI | CLM | MXL | PEL | PUR | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Zhuang | |||||||||||||||||||||
| CDX | |||||||||||||||||||||
| CHB | |||||||||||||||||||||
| CHS | |||||||||||||||||||||
| JPT | |||||||||||||||||||||
| KHV | |||||||||||||||||||||
| BEB | |||||||||||||||||||||
| GIH | |||||||||||||||||||||
| ITU | |||||||||||||||||||||
| PJL | |||||||||||||||||||||
| STU | |||||||||||||||||||||
| CEU | |||||||||||||||||||||
| FIN | |||||||||||||||||||||
| GBR | |||||||||||||||||||||
| IBS | |||||||||||||||||||||
| TSI | |||||||||||||||||||||
| ACB | |||||||||||||||||||||
| ASW | |||||||||||||||||||||
| ESN | |||||||||||||||||||||
| GWD | |||||||||||||||||||||
| LWK | 0.000 | ||||||||||||||||||||
| MSL | 0.016 | 0.000 | |||||||||||||||||||
| YRI | 0.015 | 0.004 | 0.000 | ||||||||||||||||||
| CLM | 0.164 | 0.171 | 0.165 | 0.000 | |||||||||||||||||
| MXL | 0.190 | 0.201 | 0.194 | 0.017 | 0.000 | ||||||||||||||||
| PEL | 0.231 | 0.244 | 0.235 | 0.058 | 0.035 | 0.000 | |||||||||||||||
| PUR | 0.143 | 0.151 | 0.146 | 0.009 | 0.019 | 0.067 | 0.000 | ||||||||||||||
Figure 3.
Fst value heamap and phylogenetic tree among 27 populations. (A) Heatmap based on the pairwise Fst values between 27 populations. (B) The phylogenetic tree was constructed by the neighboring-joining method among 27 populations.
Genotype frequencies of four significantly different SNVs
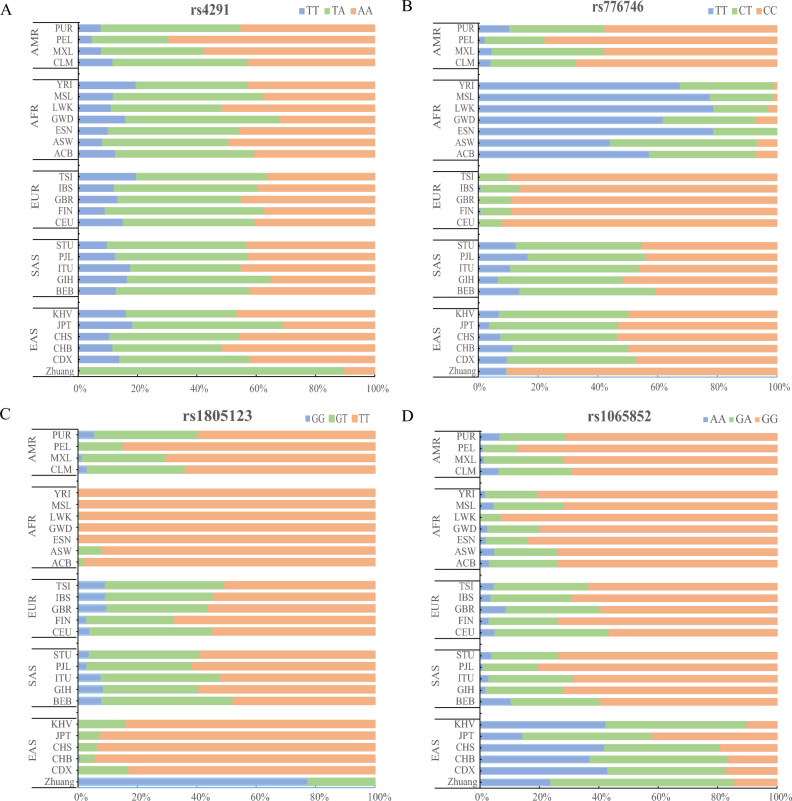
Moreover, the genotype frequency distribution of rs776746 (CYP3A5), rs4291 (ACE), rs1805123 (KCNH2), and rs1065852 (CYP2D6) in 26 populations are shown in Fig. 4. The genotype frequency of rs4291-AT in the Zhuang population is remarkably higher than that of the other 26 populations. The CC genotype frequency of rs776746 is similar to that of EUR and significantly higher than that of AFR. In the Zhuang population, the frequency of the rs1805123-GG genotype is notably higher compared to that observed in the other 26 populations. The frequency of rs1065852-GG is similar to that of KHV, CHS, CHB, and CDX, and lower than that of other populations.
Figure 4.
The distribution of genotype frequencies for significantly different SNVs in 27 populations at the rs776746, rs4291, rs1805123 and rs1065852.
MAF distribution of four significantly different SNVs
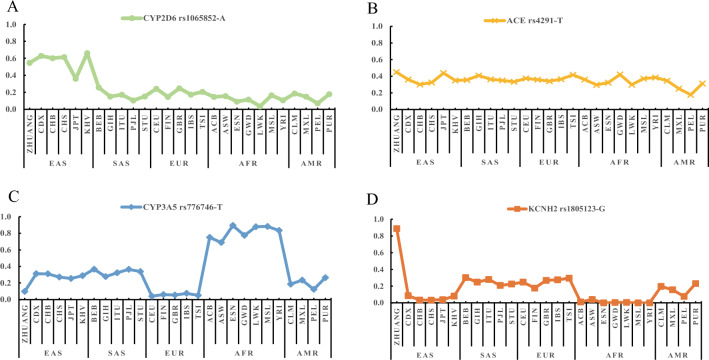
Based on the allele frequencies calculated in this study, we plotted a map of the MAF distribution of VIP variants that substantially differed from the other 26 populations. According to Fig. 5, the allele frequency of rs776746 at CYP3A5 in the Zhuang population was similar to that in the European population, despite their close genetic affinity with East Asians. The G allele of rs1805123 at KCNH2 was nearly fixed in the Zhuang population and had a low frequency in other global populations. The MAF of rs1065852 (CYP2D6) was similar to that of East Asians and higher than other populations. However, there were no significant differences in T allele frequency for rs4291 at ACE among different populations.
Figure 5.
The map of the allele frequency distribution for significantly different SNVs in 27 populations at the rs1065852, rs4291, rs776746 and rs1805123.
Clinical relevance of significant variants
The Table 4 presents the clinical annotation information of the VIP variants in PharmGKB. The genotype frequency of rs776746 (CYP3A5) has been shown to have an impact on the dose, toxicity and metabolism of tacrolimus19–21. Specific mutations in rs4291 (ACE) have been implicated in the metabolism of anti-hypertensive drugs such as amlodipine, sodium chlorthalidone and lisinopril22. Furthermore, they also influenced the risk of aspirin intolerance in asthmatics exposed to aspirin23. The efficacy of metoclopramide in patients with gastric disease was found to be associated with rs1805123 (KCNH2)24. Rs1065852 played an indispensable role in the regulation of the α-hydroxymetoprolol metabolism in patients with non-small cell lung cancer25.
Table 4.
Clinical annotation of very important pharmacogenomic variants with significant differences.
| Gene | Variant | PMID | Molecules | Association | P-value | Type | Phenotype |
|---|---|---|---|---|---|---|---|
| CYP3A5 | rs776746 | 23,073,468 | Tacrolimus | Genotype CC is associated with decreased dose of tacrolimus in people with Kidney Transplantation as compared to genotypes CT + TT | 0.016 | Dosage | Kidney Transplant |
| CYP3A5 | rs776746 | 21,677,300 | Tacrolimus | Allele T is associated with increased risk of tacrolimus nephrotoxicity when treated with tacrolimus in people with Kidney Transplantation as compared to allele C | 0.025 | Toxicity | Kidney Transplant |
| CYP3A5 | rs776746 | 24,120,259 | Tacrolimus | Genotype CT is associated with increased dose of tacrolimus in people with Kidney Transplantation as compared to genotype CC | < 0.001 | Metabolism/PK | Kidney Transplant |
| ACE | rs4291 | 27,546,928 | Captopril | Genotype AA is associated with decreased severity of Kidney Failure when treated with captopril in people with Alzheimer Disease as compared to genotypes AT + TT | 0.029 | Efficacy | Alzheimer disease |
| ACE | rs4291 | 18,727,619 | Aspirin | Genotypes AT + TT are associated with increased risk of aspirin intolerance when exposed to aspirin in people with Asthma as compared to genotype AA | 0.015 | Toxicity/ADR | Asthma |
| ACE | rs4291 | 20,577,119 | Amlodipine/lisinopril/chlorthalidone | Genotypes AA + AT are associated with decreased fasting glucose when treated with amlodipine, chlorthalidone or lisinopril in people with Hypertension as compared to genotype TT | 0.001 | Efficacy | Anti-Hypertension |
| CYP2D6 | rs1065852 | 10,223,777 | Alpha-hydroxymetoprolol | Allele A is associated with decreased clearance of alpha-hydroxymetoprolol in healthy individuals as compared to allele G | < 0.050 | Metabolism/PK | Carcinoma, Non-Small-Cell LungMesothelioma |
| CYP2D6 | rs1065852 | 24,528,284 | Citalopramescitalopram | Allele A is associated with plasma concentration of S-didesmethyl-citalopram when treated with citalopram or escitalopram in people with Depressive Disorder, Major as compared to allele G | 2E-16 | Other | Depressive Disorder |
| CYP2D6 | rs1065852 | 23,277,250 | Iloperidone | Genotype GG is associated with increased QTc interval when treated with iloperidone in people with Schizophrenia as compared to genotypes AA + AG | 0.028 | Other | Schizophrenia |
| KCNH2 | rs1805123 | 22,688,145 | Metoclopramide | The efficacy of metoclopramide in patients with gastric disease was correlated with the polymorphism of KCNH2 (rs1805123, P = 0.020) gene | 0.020 | Dose effect | Gastric disease |
ADR: Adverse drug reactions.
Prediction of functional damage in proteins
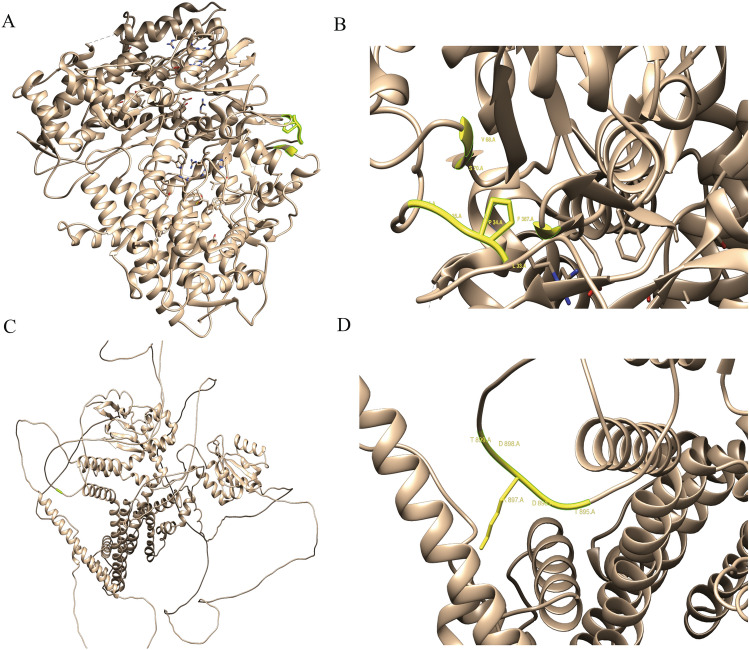
Subsequently, we used the PolyPhen-2, SNAP2, FATHMM, Mutationtaster, and Mutationassessor online databases to predicte whether the four SNVs would affect protein structure and function (Table 5). The results indicated that rs1805123 would cause a mutation from K to T at position 897 of KCNH2, however, this mutation was considered benign and less harmful to the protein in most databases. In contrast, rs1065852 caused a mutation from P to A at the 34th position of CYP2D6. The database predicted that this change would severely impair the protein's function and potentially contribute to certain diseases. Additionally, Chimera v1.16 was utilized for predicting the structure of point mutations of CYP2D6 and KCNH2, as shown in Fig. 6.
Table 5.
The functional analysis of missense variants using PolyPhen-2, SNAP2, Mutationassessor, FATHMM, and Mutationtaster.
| SNVs ID | Gene | AA change | PolyPhen-2 | SNAP2 | FATHMM | Mutation taster | Mutationassessor | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Score | Predicted effect | Score | Predicted effect | Coding Score | Predicted effect | Prob | Predicted | Func.Impact | FI score | |||
| rs1805123 | KCNH2 | K897T | 0 | Benign | -53 | Neutral | 0.760 | pathogenic | 0.205 | polymorphism | low | 1.735 |
| rs1065852 | CYP2D6 | P34A | 0.953 | Deleterious | 60 | Effect | 0.820 | pathogenic | 0.999 | disease causing | high | 4.080 |
Figure 6.
Structural prediction of point mutated proteins. (A) 3D structure of the CYP2D6 protein, with the yellow part being a SNV. (B) rs1065852 mutated local structure. (C) 3D structure of the KCNH2 protein, with the yellow part being a SNV mutation. (D) rs1805123 mutated local structure.
Discussion
During the development of biological sciences, it has gradually been realized that genetic differences between populations have an essential influence on drug metabolism, dosages and ADRs. This can potentially affect the efficacy of certain medications in specific populations. Pharmacogenomics research is gradually illuminating the genetic factors responsible for variations in drug utilization among diverse populations. For instance, an important study conducted by Wen et al. demonstrated that there were significant differences in allele frequencies of key genetic variants affecting drug selection and dosing between Hmong and East Asian populations26. Furthermore, pharmacogenomics studies have been reported on Mongolian27, Tibetan28, and Blang29, among others. However, there have been few pharmacogenomics studies conducted on the Zhuang population.
In this study, 200 Zhuang subjects in Yunnan Province were recruited and genotyped for 48 VIP variants on 24 candidate genes. The genotypic distribution was compared to that of 26 populations from the 1000G dataset. The results revealed significant differences in CYP3A5 (rs776746), ACE (rs4291), KCNH2 (rs1805123), and CYP2D6 (rs1065852) between Zhuang population and the other 26 populations. We also used the PharmGKB database to annotate significantly different SNVs. Our study on VIP polymorphism in the Zhuang population may provide tailored therapy for the Zhuang population.
The cytochrome P450 (CYP) superfamily is an ancient enzyme family found in hundreds of eukaryotic and prokaryotic organisms30. The human genome encodes 57 putative functional CYP genes, as well as 58 pseudogenes. Among these 57 functional human CYPs, 12 are involved in the metabolism of 70–80% commonly used drugs, including CYP2D6 and CYP3A531. The human CYP2D6 gene is relatively short, spanning only about 4.3 Kbps on the long arm of chromosome 22 (22q13.2). The CYP2D6 gene is composed of 9 exons and encodes the CYP2D6 protein, which is localized in the endoplasmic reticulum. This protein exhibited highly expressed levels in the liver, brain, intestinal tissues, and lymphocytes32. There were large population differences in the distribution of CYP2D6 alleles, which could lead to variations in drug utilization among different populations33. rs1065852 has been reported to result in reduced protein stability and a poor response to drugs such as iloperidone, atorvastatin, antidepressants, and antipsychotics34–37. Moreover, allele A was associated with decreased clearance of alpha-hydroxymetoprolol in healthy individuals and a higher plasma concentration of S-didesmethyl-citalopram when treated with citalopram or escitalopram in people with Depressive Disorder compared to allele G25,38. In this study, the MAF of rs1065852-A (54.80%) was higher than that in SAS (16.70%), EUR (20.30%), AFR (11.60%), and AMR (14.60%). In addition, the frequency of rs1065852-GG was lower than that in other populations except for EAS. Therefore, the differences in drug efficacy and safety caused by CYP2D6 rs1065852 should be taken into consideration in the Zhuang population.
CYP3A5, which is located in chromosome 7q21.1, is involved in the metabolism of many drugs. Tacrolimus, one of the substrates of CYP3A5, is widely used as an immunosuppressive agent for organ transplantation39. The expression of CYP3A5 varied among different populations, which may have an impact on drug metabolism in those populations21. One study identified that genotype CT was associated with a higher tacrolimus dose in renal transplant patients compared to genotype CC21. The results of Flores-Pérez et al. revealed that critically ill Mexican pediatric patients with the CYP3A5*3 allele variant (rs776746) had increased plasma levels of midazolam and higher drug clearance 3 h after the end of the infusion compared to carriers with the normal allele40. The study by Liang et al. pointed out that individuals with the rs776746-CC had an increased risk of amlodipine-induced peripheral edema in a dominant model among Chinese Han hypertensive patients41. In our study, the frequencies of CT, TT and CC of rs776746 were 0.5%, 9.5% and 90.0%, separately. The frequency of CT genotype in the Zhuang was lower than that in the other 26 populations, highlighting the importance of considering metabolism and absorption of specific drugs in the Zhuang population.
KCNH2 is a gene that encodes a component of voltage-activated potassium channel found in cardiac muscle, neuronal cells, and microglia. Four copies of this protein interact with a copy of the KCNE2 protein to form a functional potassium channel. Mutations in this gene can lead to long QT syndrome type 2 (LQT2)42. A recent study has identified KCNH2 p.Gly262AlafsTer98 as a novel pathogenic variant associated with long QT syndrome in a Spanish population43. In a separate study, it was found that KCNH2 mutations cause fetal biventricular densified cardiomyopathy with pulmonary stenosis and bradycardia44. The efficacy of metoclopramide in patients with gastric disease was found to be correlated with the polymorphism of KCNH2 gene (rs1805123, p = 0.020)24. A Marjamaa et al. found that allele G of rs1805123 was associated with a shorter QT interval in a Finnish population compared to the TT genotype45. In our study, the frequency of rs1805123-G was significantly higher in the Zhuang population than in the other 26 populations (88.70%). The rs1805123 causes a K-T mutation at site 897 of KCNH2. Although most databases predict this mutation to be benign, attention should be paid to the shorter/long QT interval and the dose of metoclopramide in the Zhuang population.
ACE, which encodes an enzyme, is known to participate in the regulation of blood pressure and electrolyte balance. Numerous studies have shown that ACE is closely associated with nervous system diseases46,47, cardiovascular diseases48, and hypertension49,50. In a previously published study, we found that the rs4291 genotype influenced drug dosing in the treatment of the disease. De Oliveira et al. found a correlation between the use of brain-penetrating angiotensin converting enzyme inhibitors (ACEIs) (such as captopril or perindopril) in antihypertensive therapy and rs429146. Another study has confirmed that the AA genotype of rs4291, compared to the genotype AT + TT, is associated with a reduced severity of renal failure in patients with Alzheimer's disease treated with captopril51. Furthermore, rs1800764 and rs4291 also formed haplotypes. A study discovered that ACEIs decelerated cognitive decline in individuals carrying the ACE haplotype with rs1800764-T and rs4291-A, as well as those carrying the APOE4 haplotype with either rs1800764-T or rs4291-T, regardless of changes in blood pressure52. Our study demonstrated that the allele frequency of rs4291 (ACE) differed significantly between the Zhuang population and the other 26 populations, which has been found to be associated with drug metabolism for various diseases, such as captopril, aspirin, and amlodipine. It may provide guidance for precision drug administration in the Zhuang population.
Our results are likely to complement the pharmacogenomic information of the Zhuang population and refine the study on the differences between the Zhuang population and the other 26 populations. More importantly, this study may provide certain theoretical support for drug use in the Zhuang population. Nonetheless, there are some limitations to our study. Our sample size was relatively small in this study. To design a comprehensive, systematic, disease-specific treatment protocol for the Zhuang population, we need to further expand the sample size for more in-depth studies. In addition, only Agena MassARRAY was used for genotyping in this study, and no other orthogonal method was employed to validate the sequencing, which will be utilized in subsequent studies.
Conclusion
In short, the genotype frequencies of CYP3A5 (rs776746), ACE (rs4291), CYP2D6 (rs1065852), and KCNH2 (rs1805123) showed significant disparities between the Zhuang population and 26 other populations. Our study can not only enrich the pharmacogenomics database of the Zhuang population but also provide a theoretical basis for tailored therapy in this population and ensure safe drug use for patients.
Supplementary Information
Acknowledgements
We would like to thank not only the subjects for participating in this study but also the clinicians and hospital staff who obtained the blood samples and collected the data.
Author contributions
T.B.J.: assisted in the conception and study design; Y.J.L.: writed and revised the manuscript and data analysis; Y.T.C. and Y.Y: reviewed and revised the article; X.Y.M., W.Q.Z., and H.Z.: participated in manuscript editing and statistical analysis; J.P.G. and J.W.: participated in the investigation and genotyping;. All authors have read and approved the manuscript.
Funding
No financial assistance was received to support the study.
Data availability
The datasets used or analyzed during the current study are available from the corresponding author upon reasonable request.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-024-58092-w.
References
- 1.Cacabelos R, Naidoo V, Corzo L, Cacabelos N, Carril JC. Genophenotypic factors and pharmacogenomics in adverse drug reactions. Int. J. Mol. Sci. 2018;22:155. doi: 10.3390/ijms222413302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Osanlou O, Pirmohamed M, Daly AK. Pharmacogenetics of adverse drug reactions. Adv. Pharmacol. 2018;83:155–190. doi: 10.1016/bs.apha.2018.03.002. [DOI] [PubMed] [Google Scholar]
- 3.Elzagallaai AA, Greff M, Rieder MJ. Adverse drug reactions in children: The double-edged sword of therapeutics. Clin. Pharmacol. Therapeut. 2017;101:725–735. doi: 10.1002/cpt.677. [DOI] [PubMed] [Google Scholar]
- 4.Malki MA, Pearson ER. Drug-drug-gene interactions and adverse drug reactions. Pharmacogenom. J. 2018;20:355–366. doi: 10.1038/s41397-019-0122-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ingelman-Sundberg M. Pharmacogenetics of cytochrome P450 and its applications in drug therapy: The past, present and future. Trends Pharmacol. Sci. 2004;25:193–200. doi: 10.1016/j.tips.2004.02.007. [DOI] [PubMed] [Google Scholar]
- 6.Pirmohamed M. Personalized pharmacogenomics: predicting efficacy and adverse drug reactions. Annual Rev. Genom. Human Genet. 2014;15:349–370. doi: 10.1146/annurev-genom-090413-025419. [DOI] [PubMed] [Google Scholar]
- 7.Swen JJ, et al. A 12-gene pharmacogenetic panel to prevent adverse drug reactions: An open-label, multicentre, controlled, cluster-randomised crossover implementation study. Lancet. 2020;401:347–356. doi: 10.1016/S0140-6736(22)01841-4. [DOI] [PubMed] [Google Scholar]
- 8.Cecchin E, Stocco G. Pharmacogenomics and personalized medicine. Genes. 2020;11:679. doi: 10.3390/genes11060679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Weinshilboum RM, Wang L. Pharmacogenomics: Precision medicine and drug response. Mayo Clin. Proc. 2017;92:1711–1722. doi: 10.1016/j.mayocp.2017.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cacabelos R. Pharmacogenomics of Alzheimer’s and Parkinson’s diseases. Neurosci. Lett. 2020;726:133807. doi: 10.1016/j.neulet.2018.09.018. [DOI] [PubMed] [Google Scholar]
- 11.Abecasis GR, et al. A map of human genome variation from population-scale sequencing. Nature. 2010;467:1061–1073. doi: 10.1038/nature09534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lek M, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–291. doi: 10.1038/nature19057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Whirl-Carrillo M, et al. An evidence-based framework for evaluating pharmacogenomics knowledge for personalized medicine. Clin. Pharmacol. Therapeut. 2021;110:563–572. doi: 10.1002/cpt.2350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.He C, et al. Population genetic difference of pharmacogenomic VIP variants in the tibetan population. Pharmacogenom. Personal. Med. 2021;14:1027–1040. doi: 10.2147/PGPM.S316711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Cheng Y, et al. Analysis of very important pharmacogenomics variants in the chinese lahu population. Pharmacogenom. Personal. Med. 2021;14:1275–1289. doi: 10.2147/PGPM.S324410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Faul F, Erdfelder E, Lang A, Buchner AJBRM. G*Power 3: A flexible statistical power analysis program for the social. Behav. Biomed. Sci. 2021;39:175–191. doi: 10.3758/bf03193146. [DOI] [PubMed] [Google Scholar]
- 17.Trembizki E, et al. High-throughput informative single nucleotide polymorphism-based typing of Neisseria gonorrhoeae using the Sequenom MassARRAY iPLEX platform. J. Antimicrob. Chemother. 2014;69:1526–1532. doi: 10.1093/jac/dkt544. [DOI] [PubMed] [Google Scholar]
- 18.Thomas RK, et al. High-throughput oncogene mutation profiling in human cancer. Nat. Genet. 2014;39:347–351. doi: 10.1038/ng1975. [DOI] [PubMed] [Google Scholar]
- 19.Glowacki F, et al. CYP3A5 and ABCB1 polymorphisms in donor and recipient: impact on Tacrolimus dose requirements and clinical outcome after renal transplantation. Nephrol. Dial. Transplant. Off. Pub. Eur. Dial. Transplant Assoc.–Eur. Renal. Assoc. 2011;26:3046–3050. doi: 10.1093/ndt/gfr253. [DOI] [PubMed] [Google Scholar]
- 20.Niioka T, et al. Comparison of pharmacokinetics and pharmacogenetics of once- and twice-daily tacrolimus in the early stage after renal transplantation. Transplantation. 2012;94:1013–1019. doi: 10.1097/TP.0b013e31826bc400. [DOI] [PubMed] [Google Scholar]
- 21.Vannaprasaht S, et al. Personalized tacrolimus doses determined by CYP3A5 genotype for induction and maintenance phases of kidney transplantation. Clin. Therapeut. 2013;35:1762–1769. doi: 10.1016/j.clinthera.2013.08.019. [DOI] [PubMed] [Google Scholar]
- 22.Irvin MR, et al. Pharmacogenetic association of hypertension candidate genes with fasting glucose in the GenHAT Study. J. Hypertens. 2010;28:2076–2083. doi: 10.1097/HJH.0b013e32833c7a4d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kim TH, et al. Association of angiotensin I-converting enzyme gene polymorphisms with aspirin intolerance in asthmatics. Clin. Exp. Allergy J. Br. Soc. Allergy Clin. Immunol. 2008;38:1727–1737. doi: 10.1111/j.1365-2222.2008.03082.x. [DOI] [PubMed] [Google Scholar]
- 24.Parkman HP, et al. Clinical response and side effects of metoclopramide: Associations with clinical, demographic, and pharmacogenetic parameters. J. Clin. Gastroenterol. 2012;46:494–503. doi: 10.1097/MCG.0b013e3182522624. [DOI] [PubMed] [Google Scholar]
- 25.Huang J, Chuang SK, Cheng CL, Lai ML. Pharmacokinetics of metoprolol enantiomers in Chinese subjects of major CYP2D6 genotypes. Clin. Pharmacol. Therapeut. 1999;65:402–407. doi: 10.1016/S0009-9236(99)70134-7. [DOI] [PubMed] [Google Scholar]
- 26.Wen YF, et al. Potential clinical relevance of differences in allele frequencies found within very important pharmacogenes between hmong and east asian populations. Pharmacotherapy. 2020;40:142–152. doi: 10.1002/phar.2360. [DOI] [PubMed] [Google Scholar]
- 27.Guo L, et al. Very important pharmacogenes polymorphism landscape and potential clinical relevance in the Chinese Mongolian. Gene. 2023;850:146960. doi: 10.1016/j.gene.2022.146960. [DOI] [PubMed] [Google Scholar]
- 28.Rong H, et al. Analysis of very important pharmacogene variants in the Tibetan population from China. Clin. Exp. Pharmacol. Physiol. 2021;48:668–678. doi: 10.1111/1440-1681.13327. [DOI] [PubMed] [Google Scholar]
- 29.Wang Y, et al. Genetic polymorphisms of very important pharmacogene variants in the blang population from Yunnan province in China. Pharmacogen. Personal. Med. 2021;14:1647–1660. doi: 10.2147/PGPM.S327313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kawashima A, Satta Y. Substrate-dependent evolution of cytochrome P450: Rapid turnover of the detoxification-type and conservation of the biosynthesis-type. PloS one. 2014;9:e100059. doi: 10.1371/journal.pone.0100059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zanger UM, Schwab M. Cytochrome P450 enzymes in drug metabolism: Regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacol. Therapeut. 2013;138:103–141. doi: 10.1016/j.pharmthera.2012.12.007. [DOI] [PubMed] [Google Scholar]
- 32.Taylor C, et al. A Review of the important role of CYP2D6 in pharmacogenomics. Genes. 2020;11:1295. doi: 10.3390/genes11111295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ingelman-Sundberg M, Sim SC, Gomez A, Rodriguez-Antona C. Influence of cytochrome P450 polymorphisms on drug therapies: Pharmacogenetic, pharmacoepigenetic and clinical aspects. Pharmacol. Therapeut. 2007;116:496–526. doi: 10.1016/j.pharmthera.2007.09.004. [DOI] [PubMed] [Google Scholar]
- 34.Pei Q, et al. Influences of CYP2D6(*)10 polymorphisms on the pharmacokinetics of iloperidone and its metabolites in Chinese patients with schizophrenia: A population pharmacokinetic analysis. Acta Pharmacologica Sinica. 2016;37:1499–1508. doi: 10.1038/aps.2016.96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Peng C, et al. Polymorphisms in CYP450 genes and the therapeutic effect of atorvastatin on ischemic stroke: A retrospective cohort study in chinese population. Clin. Therapeut. 2018;40:469–477. e462. doi: 10.1016/j.clinthera.2018.02.002. [DOI] [PubMed] [Google Scholar]
- 36.Sun CJ, Li L, Li XY, Zhang WY, Liu XW. Associations of polymorphisms of CYP2D6 and CYP2C9 with early onset severe pre-eclampsia and response to labetalol therapy. Arch. Gynecol. Obstetr. 2018;298:125–132. doi: 10.1007/s00404-018-4791-8. [DOI] [PubMed] [Google Scholar]
- 37.Xin J, Yuan M, Peng Y, Wang J. Analysis of the deleterious single-nucleotide polymorphisms associated with antidepressant efficacy in major depressive disorder. Front. Psychiatry. 2020;11:151. doi: 10.3389/fpsyt.2020.00151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ji Y, et al. Citalopram and escitalopram plasma drug and metabolite concentrations: Genome-wide associations. Br. J. Clin. Pharmacol. 2014;78:373–383. doi: 10.1111/bcp.12348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rodriguez-Antona C, et al. PharmVar GeneFocus: CYP3A5. Clin. Pharmacol. Therapeut. 2022;112:1159–1171. doi: 10.1002/cpt.2563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Flores-Pérez C, et al. The rs776746 variant of CYP3A5 is associated with intravenous midazolam plasma levels and higher clearance in critically ill Mexican paediatric patients. J. Clin. Pharm. Therapeut. 2021;46:633–639. doi: 10.1111/jcpt.13388. [DOI] [PubMed] [Google Scholar]
- 41.Liang H, et al. Association of CYP3A5 gene polymorphisms and amlodipine-induced peripheral edema in chinese han patients with essential hypertension. Pharmacogenom. Personal. Med. 2021;14:189–197. doi: 10.2147/PGPM.S291277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ng CA, et al. A massively parallel assay accurately discriminates between functionally normal and abnormal variants in a hotspot domain of KCNH2. Am. J. Human Genet. 2022;109:1208–1216. doi: 10.1016/j.ajhg.2022.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Lorca R, et al. KCNH2 p.Gly262AlafsTer98: A new threatening variant associated with long QT syndrome in a spanish cohort. Life. 2022;12:556. doi: 10.3390/life12040556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sun H, et al. Case report: Biventricular noncompaction cardiomyopathy with pulmonary stenosis and bradycardia in a fetus With KCNH2 mutation. Front. Genet. 2022;13:821226. doi: 10.3389/fgene.2022.821226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Marjamaa A, et al. Common candidate gene variants are associated with QT interval duration in the general population. J. Int. Med. 2009;265:448–458. doi: 10.1111/j.1365-2796.2008.02026.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.de Oliveira FF, Bertolucci PH, Chen ES, Smith MC. Brain-penetrating angiotensin-converting enzyme inhibitors and cognitive change in patients with dementia due to Alzheimer's disease. J. Alzheimer's Dis. JAD. 2014;42(Suppl 3):s321–324. doi: 10.3233/JAD-132189. [DOI] [PubMed] [Google Scholar]
- 47.Wang XB, et al. Angiotensin-converting enzyme gene polymorphisms and risk for sporadic Alzheimer's disease: a meta-analysis. J. Neural Transm. 2014;122:211–224. doi: 10.1007/s00702-014-1235-x. [DOI] [PubMed] [Google Scholar]
- 48.Baghai TC, et al. Polymorphisms in the angiotensin-converting enzyme gene are associated with unipolar depression, ACE activity and hypercortisolism. Mol. Psychiatry. 2006;11:1003–1015. doi: 10.1038/sj.mp.4001884. [DOI] [PubMed] [Google Scholar]
- 49.Kehoe PG, et al. Common variants of ACE contribute to variable age-at-onset of Alzheimer's disease. Human Genet. 2004;114:478–483. doi: 10.1007/s00439-004-1093-y. [DOI] [PubMed] [Google Scholar]
- 50.Xin XY, Lai ZH, Ding KQ, Zeng LL, Ma JF. Angiotensin-converting enzyme polymorphisms AND Alzheimer's disease susceptibility: An updated meta-analysis. PloS one. 2021;16:e0260498. doi: 10.1371/journal.pone.0260498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.de Ferreira Oliveira F, Berretta JM, Suchi Chen E, Cardoso Smith M, Ferreira Bertolucci PH. Pharmacogenetic effects of angiotensin-converting enzyme inhibitors over age-related urea and creatinine variations in patients with dementia due to Alzheimer disease. Colombia Medica. 2016;47:76–80. doi: 10.25100/cm.v47i2.2188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.de Oliveira FF, Chen ES, Smith MC, Bertolucci PHF. Pharmacogenetics of angiotensin-converting enzyme inhibitors in patients with alzheimer's disease dementia. Current Alzheimer Res. 2018;15:386–398. doi: 10.2174/1567205014666171016101816. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets used or analyzed during the current study are available from the corresponding author upon reasonable request.