Abstract
Prostate cancer harbors several genetic alterations, the most prevalent of which is TMPRSS2:ERG gene fusion, affecting nearly half of all cases. Capitalizing on the increasing availability of whole-slide images (WSI), this study introduces a deep learning (DL) model designed to detect TMPRSS2:ERG fusion from H&E-stained WSIs of radical prostatectomy specimens. Leveraging the TCGA prostate adenocarcinoma cohort, which comprises 436 WSIs from 393 patients, we developed a robust DL model, trained across 10 different splits, each consisting of distinct training, validation, and testing sets. The model's best performance achieved an AUC of 0.84 during training, and 0.72 on the TCGA test set. This model was subsequently validated on an independent cohort comprising 314 WSIs from a different institution, in which it has a robust performance at predicting TMPRSS2:ERG fusion with an AUC of 0.73. Importantly, the model identifies highly-attended tissue regions associated with TMPRSS2:ERG fusion, characterized by higher neoplastic cell content and altered immune and stromal profiles compared with fusion-negative cases. Multivariate survival analysis revealed that these morphologic features correlate with poorer survival outcomes, independent of Gleason grade and tumor stage. This study underscores the potential of DL in deducing genetic alterations from routine slides and identifying their underlying morphologic features, which might harbor prognostic information.
Implications:
Our study illuminates the potential of deep learning in effectively inferring key prostate cancer genetic alterations from the tissue morphology depicted in routinely available histology slides, offering a cost-effective method that could revolutionize diagnostic strategies in oncology.
Introduction
Prostate cancer is the second most common solid organ malignancy among men worldwide, with more than 1 million annually diagnosed new cases (1, 2). The prostate cancer genome is characterized by several gene rearrangements involving E26 transformation-specific (ETS) transcription factors (3, 4). Of these, fusion between ERG and the androgen-regulated transmembrane protease, serine 2 (TMPRSS2) is the most frequent, being present in nearly 40% to 50% of prostate cancer cases (3). The TMPRSS2:ERG fusion was found to be instrumental during the transformation of prostate intraepithelial neoplastic lesions into invasive adenocarcinoma as well as during prostate cancer progression and metastasis (5).
The diagnosis of prostate cancer is based on the light microscopic examination of hematoxylin and eosin (H&E)-stained tissue sections obtained from the prostate gland (6–8). However, H&E staining alone cannot differentiate between ERG fusion-positive (those with TMPRSS2:ERG fusion) and negative tumors. Instead, ERG fusion status is usually detected using FISH or reverse transcription-PCR (RT-PCR) whereas IHC staining for the ERG protein can be used to infer the TMPRSS2:ERG gene fusion status with optimal sensitivity and specificity (9). Although RT-PCR and IHC have been routinely used and are considered cost-effective in well-equipped clinical settings, their accessibility can be limited in under-resourced or remote areas. The continuous advancements in AI-driven methods, coupled with the increasing availability of low-cost slide scanners, present an opportunity to introduce alternative methods for ERG fusion detection in regions where traditional methods may be less accessible. Specifically, a tool that can utilize routine H&E-stained slides for such task would potentially achieve the aforementioned goals given the practicality and widespread availability of these slides.
Since 2000, whole-slide imaging (WSI) started to become common, as digital slide scanners became commercially available (10). Currently, modern pathology practice is moving toward a digital workflow through the deployment of artificial intelligence (AI) and computer vision systems for image preprocessing, segmentation, feature detection, and quantification on H&E-stained WSIs (10, 11). Deep learning (DL) systems in particular have been employed extensively over the past few years in several tasks involving the use of H&E-stained WSIs for phenotype prediction, classification, and subtyping especially in cancer research (12, 13). For instance, DL models have been developed to automatically detect tumor regions in several cancer types including breast, lung, and prostate cancers (14–19). In addition, DL has been used in more advanced tasks including classification of tumor subtypes (20–22), tumor grading (19, 23), and prediction of therapeutic response (24). Notably, the utilization of DL systems in molecular pathology has gained momentum over the past years with more models being deployed to predict genetic alterations from histopathology images (12). For instance, Coudray and colleagues have developed a model for predicting several key mutations in lung adenocarcinoma (25), whereas Bilal and colleagues developed a DL system for predicting several key genetic alterations in colorectal cancer (26). Similar studies have aimed at identifying other molecular alterations and phenotypes including ER status in breast cancer (27), BRAF mutations in melanoma (28), and SPOP mutations in prostate cancer (29).
Here, we introduce a semi-supervised DL model capable of inferring the TMPRSS2:ERG gene fusion status solely from H&E-stained WSIs. This model has been trained and validated using two large imaging cohorts with 750 WSIs derived from radical prostatectomy specimens of patients with prostate cancer with known ERG fusion status. In addition, we deciphered the cellular composition of the top patches contributing to the prediction of each ERG phenotype and examined the association of this composition with overall, progression-free survival (PFS), and metastasis-free survivals.
Materials and Methods
Patients and slides selection
We queried the Genomic Data Commons (GDC) portal and The Cancer Genome Atlas (TCGA) for whole slide images (WSI) that match our inclusion criteria for this analysis which included formalin-fixed paraffin-embedded (FFPE) slides of radical prostatectomy specimens from men with primary prostate cancer and known TMPRSS2:ERG gene fusion status. The following terms were used: Project ID: TCGA-PRAD; Data Type: Slide Image; and Experimental Strategy: Diagnostic Slide. The prostate cancer TCGA dataset included 436 FFPE H&E-stained WSIs from 393 patients with prostate cancer with known TMPRSS2:ERG fusion status (positive vs. negative) determined by FusionSeq (30, 31). Subsequently, a dataset of 314 FFPE-derived WSIs from 314 unique patients with prostate cancer has been provided by the Johns Hopkins University (thereafter termed the natural history cohort) and was used as an additional testing set. The ERG status in this cohort has been determined using IHC staining of FFPE tumor tissue from radical prostatectomy specimens to detect the ERG/TMPRSS2 fusion protein (9). Slides from the TCGA and natural history cohorts were digitized using Aperio (Leica Biosystems) and Hamamatsu (Hamamatsu Photonics K.K.) slide scanners, respectively. The WSIs were fully de-identified before we accessed them, and the use of these data for research purposes was approved by the relevant Institutional Review Boards.
Image preprocessing
Preprocessing and tissue segmentation
During the preprocessing of WSIs, we converted RGB images to Hue, Saturation, Value (HSV) space. This conversion separates the luminance (value) from the color information (hue and saturation), allowing our DL model to emphasize texture and structure, which are less affected by staining variability (32). To further enhance the model's focus on morphologic features, we applied a median blur filter with a kernel size of 7. This step helps in smoothing the image and reducing noise that could obscure histologic detail. Subsequently, we applied morphological closing with a 4 × 4 structuring element to bridge small gaps in tissue, facilitating the model's recognition of contiguous structures. Tissue segmentation was then performed on the preprocessed images using Otsu's method to differentiate between tissue and background, further refining the input for our model (33).
Tiling and feature extraction
Following segmentation, tissue masks were tiled into 2048 × 2048 patches at 40× magnification. We have used patches encompassing both cancerous and noncancerous regions since by training the model on all tiles, we ensure that it learns from diverse tissue morphologies. Regions such as the stroma, even if noncancerous, can hold significant contextual information that might be pertinent to determining ERG status. This approach potentially allows for more nuanced understanding and prediction of ERG status, as it captures the broader tissue context. Subsequently, we followed by down sampling the patches by a factor of 4 resulting into 512 × 512 patches at 10× magnification on which feature extraction was performed using a pre-trained ResNet50 model (34). Notably, tiles derived from slides corresponding to the same patient were grouped together to avoid data leakage between the training and testing sets which could affect the model's generalizability.
Training the TMPRSS2:ERG fusion status prediction model
The DL framework
To build a model capable of inferring the ERG status from H&E WSIs, we used Clustering-constrained Attention Multiple Instance Learning (CLAM; ref. 34). CLAM is a modified multiple instance learning (MIL) framework, which aggregates patch-level into slide-level representations using an attention-based pooling function instead of max pooling (35). In the CLAM network, the first layer is a fully connected linear layer that takes the 1024-dimensional vector representing the extracted patch features and returns a 512-dimensional vector, which is subsequently inputted into the attention network. The attention network is based on a gated attention mechanism that assigns different weights to instances (patches) within a bag (WSI) based on their contributions to the slide-level prediction (35). This network then splits into two separate branches, one for each class (ERG-positive and negative). Notably, slide-level representations are scored by two class-specific separate classifiers and a softmax function is then used to convert these into class-specific probability scores for each WSI.
WSIs datasets
For developing the model, we divided the TCGA PRAD cohort into training, validation, and testing sets, using 10 different splits. For each split, slides were assigned to training, validation, or testing sets using a stratified sampling process to ensure equal ratios of ERG-positive and negative cases in each subset. Notably, in case of multiple slides derived from the same patient, these slides were always assigned to the same set to minimize the bias arising from training and testing on slides from the same patient which could inflate the model's performance. For each split, the training set comprised 70% of all slides (n = 318 WSIs), whereas the validation and testing sets each comprised 15% of all slides (n = 59 WSIs). Model training was performed on the training set while the validation set was used to optimize its hyperparameters.
Training parameters
The model's learning was guided by the cross-entropy loss function, which was used to compare the slide-level predictions to the true labels and the model's internal weights were adjusted using Adam Optimizer with an α of 0.0001 and weight decay of 0.00001. Finally, a maximum of 150 epochs was used for training the model and we used early stopping to stop training and save the model if the error in the validation set did not decrease for over 20 epochs. This was done to minimize overfitting and to ensure an optimal balance between learning and generalization capabilities.
Independent evaluation of performance
In addition to the testing set derived from the TCGA cohort (n = 59), we tested the model on the entire natural history cohort (n = 314) to provide a completely independent assessment of performance. The model was utilized to infer class-specific predicted probabilities for each WSI in the natural history cohort. Utilizing the optimal threshold from the ROC curve of the training data, we translated these probability scores into binary class predictions (ERG positive vs. negative) and compared these predictions with the true class labels to calculate the accuracy and Matthews Correlation Coefficient (MCC; ref. 36).
Nuclei segmentation and classification
To unravel the cellular composition of the highly attended regions by our model, we used the HoVer-Net model trained on the PanNuke dataset (37, 38) to segment and classify the nuclei in these regions. Specifically, we pooled the patches with highest attention scores (15 patches per WSI) from all WSIs predicted as either positive or negative. These patches, obtained after basic preprocessing and tissue segmentation, maintained a size of 512 × 512 at 10× magnification. HoVer-Net was used on patches from each class separately to segment and classify the nuclei into 5 different types: benign epithelial, neoplastic, inflammatory/immune, necrotic, and stromal (37). Following this process, we then compared the prevalence of distinct nuclear types in the highly targeted regions of cases predicted as ERG-positive to those predicted as ERG-negative in both the TCGA and natural history cohorts.
Survival analyses
To examine the association between the nuclear/cellular content in the highly attended patches and the survival probability, we calculated the number and ratio of different nuclear types in each WSI in the TCGA and natural history cohorts. The ratio of each nuclear type was computed by dividing the absolute number of each nuclear type by the number of all nuclei in the highly attended patches of each slide (top 15 patches). Thereafter, this ratio was binary classified into high and low content, making use of maximally selected log-rank statistics to ascertain the best cutoff that presented the most significant association with the survival outcome (39, 40). We subsequently computed the association between the binarized nuclear content and the overall survival (OS) (TCGA and natural history cohorts), PFS (TCGA cohort), and metastases-free survival (natural history cohort) using Kaplan–Meier (KM) survival curves (41). Notably, only the correctly classified slides from each cohort were included in this analysis.
Statistical analyses and software
The performance of our model was assessed based on the AUC, accuracy, balanced accuracy, sensitivity, specificity, and Matthews Correlation Coefficient (MCC; ref. 36). ROC curves were plotted using the predicted probability scores together with the ground truth labels. For all cohorts, the probability scores were binarized into predicted classes using the best threshold from the ROC curves of the training data. Slide preprocessing and training the DL model were performed using python (v3.7.5), openslide (v3.4.1), and PyTorch (v1.3.1). Regarding the segmentation and classification of nuclei, we employed the PyTorch (v1.6) implementation of the HoVer-Net model, previously trained on the PanNuke dataset (37, 38). Survival analyses were performed using the survival (v3.3–1; refs. 42, 43) and survminer (v0.4.9; ref. 44) packages. The code used to perform this analysis is publicly available on GitHub and can be accessed using the following link: https://github.com/MohamedOmar2020/pca_ERG
Data availability
Digitized whole slide images from the TCGA prostate adenocarcinoma cohort are publicly available through the GDC data portal and can be accessed using the following link: https://portal.gdc.cancer.gov/.
Results
Patient selection
The prostate cancer TCGA cohort includes 436 slides from 393 unique patients with available slide-level information about the ERG fusion status. The slides in this cohort were split into a training set consisting of 318 WSIs (188 ERG-negative and 130 ERG-positive) together with a validation and testing sets each consisting of 59 WSIs (33 ERG-negative and 26 ERG-positive). In addition, we used the entire natural history cohort as an independent testing set to further validate the model's performance. This cohort includes 314 WSIs (185 ERG-negative and 129 ERG-positive) from patients who underwent radical prostatectomy between 1992 and 2010 and received no treatment prior to the procedure (Fig. 1).
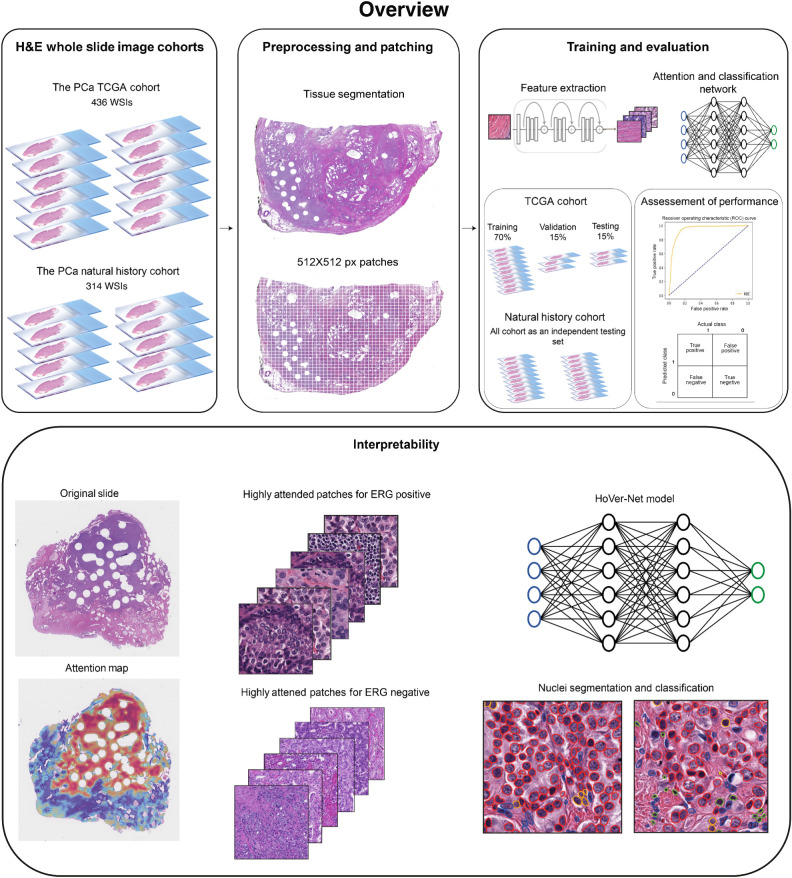
Figure 1.
Detecting TMPRSS2:ERG fusion status using H&E-stained radical prostatectomy specimens from patients with prostate cancer. Whole slide images from the prostate cancer TCGA (n = 436) and natural history (n = 314) cohorts were used in this study. Following tissue segmentation, each WSI was tiled into 2048 × 2048 patches at 40× magnification and were further down sampled by a factor of 4 to extract features from 512 × 512 patches at 10× magnification. WSIs from the prostate cancer TCGA cohort were split into training (70%), validation (15%), and testing (15%) sets whereas the natural history cohort was used as an additional testing set. HoVer-Net model was used for nuclei segmentation and classification in the top patches of WSIs predicted as ERG fusion positive or negative.
Predicting TMPRSS2:ERG fusion status from H&E-stained whole slide images
The ERG fusion prediction model was trained on H&E-stained WSIs to distinguish slides derived from patients with ERG-positive from those with ERG-negative using tissue morphologic and spatial features only. Because ERG fusion is known to induce changes in the tumor microenvironment, we hypothesized that this can also be associated with morphologic changes that are not limited only to the tumor but also involve the stroma and other regions surrounding the tumor. With this in mind, we used the whole tissue sections (after preprocessing) for making predictions instead of using only tumor regions. We trained 10 distinct models using 10 different splits of the TCGA dataset, each split consisting of training (n = 318), validation (n = 59), and testing (n = 59) sets (Figs. 1 and 2A). The average AUC for the 10 models in the training data is 0.79 (SD = 0.03) whereas the average accuracy is 0.71 (SD = 0.03). The best performing model has an AUC of 0.84 and accuracy of 0.77 in the training data (Fig. 2A). We further evaluated this model on the TCGA testing set (n = 59), in which it has an AUC of 0.72, accuracy of 0.70, and MCC of 0.38 using the best threshold derived from the ROC curve of the training data (Fig. 2B). The performance metrics of the model using different thresholds are shown in Supplementary Table S1.
Figure 2.
Performance of models predicting TMPRSS2:ERG fusion status using H&E-stained whole slide images. A, Performance of the models in the 10 different training folds. The prostate cancer TCGA cohort was divided 10 times into training, validation, and testing sets with each fold having different slides in each set. In each fold, models were trained on the training set while the validation set was used for tuning the model hyperparameters. The best performing model (fold 3) was used for downstream evaluation on the TCGA testing set as well as the natural history cohort. B and C, Performance of the model in the TCGA testing set (n = 59) and the natural history cohort (n = 314).
Independent evaluation of performance on the natural history cohort
In addition to testing the model on a separate testing set derived from the TCGA cohort, we also assessed its performance in a totally separate patient cohort from a different institution and different scanner. The model was deployed on the natural history cohort which includes 314 WSIs of tissue sections from radical prostatectomy specimens, in which the ERG status has been inferred using IHC. In this cohort, the ERG model can detect the ERG status with an AUC of 0.73, accuracy of 0.69, and MCC of 0.35 using the best threshold from the ROC curve of the training data (Fig. 2C) and also using different thresholds (see Supplementary Table S2). This performance is similar to the one seen in the TCGA testing set, which highlights our model robustness to technical variability. This variability is often observed in tasks involving slides from multiple institutions, prepared by different personnel, and scanned with different slide scanners.
Highly attended patches show distinct morphologic features associated with ERG fusion
To further decipher the cellular architecture contributing to the model's prediction, we extracted the top 15 highly attended patches (highest attention scores) from each slide predicted as either ERG-positive or negative. We then used HoVer-Net model to perform nuclear segmentation and classification into five categories: benign epithelial, tumor, stroma, inflammatory/immune, and necrotic cells and compared the frequency of these nuclear types between the two predicted classes. Subsequently, we compared the cellular composition in these top patches between the ERG-positive and negative cases. On average, the highly attended patches from the TCGA WSIs predicted as ERG-positive tend to have more neoplastic content and a lower stromal-to-neoplastic cell ratio (Fig. 3A) compared with those predicted as negative which are characterized by a higher stromal to neoplastic cell ratio combined with a more prominent inflammatory infiltration (Fig. 3B). The same pattern is observed in the natural history cohort in which the ERG-positive highly attended patches have a higher neoplastic and less inflammatory content (Fig. 4A) compared with the ERG-negative patches (Fig. 4B).
Figure 3.
Distinct morphological features corresponding to TMPRSS2:ERG fusion in the TCGA cohort. A and B, Example of a slide predicted as ERG positive (A) and negative (B) with the corresponding top 15 tiles with highest attention scores. HoVer-Net model was used to segment and classify the nuclei in these patches into five types: neoplastic, inflammatory, stroma, necrotic, and benign epithelial.
Figure 4.
Distinct morphologic features corresponding to TMPRSS2:ERG fusion in the natural history cohort. A and B, Example of a slide predicted as ERG positive (A) and negative (B) with the corresponding top 15 tiles with highest attention scores. HoVer-Net model was used to segment and classify the nuclei in these patches into five types: neoplastic, inflammatory, stroma, necrotic, and benign epithelial.
To further elucidate the relationship between TMPRSS2:ERG fusion status and cellular composition in relation to Gleason score, we conducted a stratified analysis of the cellular architecture within the most informative patches, as highlighted by our DL model. This analysis revealed a distinct cellular distribution pattern correlating with TMPRSS2:ERG fusion status that varies with tumor grade. On average, we observe a higher neoplastic ratio in ERG-positive cases, which is more pronounced in those with higher Gleason scores. Specifically, ERG-positive cases with a Gleason score of ≥8 exhibited an average neoplastic ratio of 0.72 compared with 0.68 in cases with a Gleason score of <8. These suggest a greater tumor cell density associated with higher-grade and potentially more aggressive disease (Supplementary Fig. S1A). The immune ratio, while slightly increased across both fusion statuses in higher Gleason scores, was notably lower in ERG-positive cases, indicating a potential dampening of the immune microenvironment in these tumors (Supplementary Fig. S1B). Importantly, the stroma to neoplastic cell ratio decreased in ERG-positive cases with higher Gleason scores (0.23 for ≥8 vs. 0.71 for <8), mirroring the characteristics of aggressive tumor phenotypes such as Gleason pattern 4. Conversely, ERG-negative cases presented an inverse relationship, with a higher stroma to neoplastic cell ratio (0.74 for ≥8 vs. 0.43 for <8), aligning with features typically observed in Gleason pattern 3 (Supplementary Fig. S1C). These findings enrich our understanding of the morphologic spectrum associated with TMPRSS2:ERG fusion status and provide further validation for the use of DL models to capture prognostically significant histologic features from routine H&E-stained slides.
The cellular composition in the highly attended patches captures clinical outcome
We further examined whether the cellular composition in the highly attended patches is associated with the survival probability in both cohorts. For each WSI correctly predicted as positive or negative, we computed the number and ratio of each nuclear type predicted by the HoVer-Net model in the highly attended patches (top 15 patches with the highest attention scores for each WSI). We subsequently computed the association between the cellular composition and the PFS in the TCGA cohort together with OS and metastases-free survival in the natural history cohort. In the TCGA, a high fraction of neoplastic cells in the highly attended patches is significantly associated with shorter PFS (P-value = 0.01) whereas high fractions of necrotic, and stromal cells are significantly associated with longer PFS (P-values = 0.031 and 0.002, respectively; Fig. 5). In addition, we report a significant association between the ratio of stromal to neoplastic cells and PFS (P-value = 0.006; Fig. 5). With the cellular composition of the highly-attended tissue regions providing accurate prognostic estimates, we sought to determine whether this prognostic value is comparable with the cellular composition of the entire slide. Therefore, we deployed the HoVer-Net model on all WSIs to construct slide-level cellular composition features, and examined their association with survival. Notably, the slide-level cellular composition did not provide additional prognostic value when compared with the composition derived from the highly-attended tissue regions identified by our ERG prediction model (Supplementary Fig. S2).
Figure 5.
Cellular composition in the highly attended patches is associated with PFS in the TCGA cohort. KM curves showing the association between the fraction of each nuclear type and PFS. The fraction of each cell type was calculated by dividing the absolute number of that cell type over the number of all cells in the highly attended patches of each slide. The stromal to neoplastic ratio was calculated by dividing the number of stromal cells by that of neoplastic cells in the highly attended patches of each slide. These fractions were then then binarized into high versus low using maximally selected log-rank statistics.
Next, we performed multivariate Cox proportional analyses examining the association of each cell type ratio with PFS after adjusting for Gleason grade and pathologic T and N stages. Notably, we found that a high neoplastic content within the highly attended patches is significantly associated with worse PFS (HR, 2.1; 95% CI, 1.1–3.9) in the multivariate models, whereas the opposite is true for high stromal (HR, 0.41; 95% CI, 0.21–0.78) and stromal to neoplastic (HR, 0.41; 95% CI, 0.21–0.81) fractions (Table 1).
Table 1.
Multivariate Cox proportional hazards model examining the association of cellular composition of the highly attended patches with PFS in the TCGA cohort.
| Variable | HR | Lower 95% CI | Upper 95% CI | P value | |
|---|---|---|---|---|---|
| Cellular composition (high vs. low)a | Neoplastic | 2.1 | 1.1 | 3.9 | 0.02 |
| Inflammatory | 0.54 | 0.26 | 1.1 | 0.096 | |
| Stromal | 0.41 | 0.21 | 0.78 | 0.006 | |
| Necrotic | 0.34 | 0.09 | 1.2 | 0.083 | |
| Benign epithelial | 1.2 | 0.64 | 2.3 | 0.562 | |
| Stromal-to-neoplastic ratio | 0.41 | 0.21 | 0.81 | 0.011 | |
| Gleason score (Gleason <8 as reference) | Gleason ≥8 | 3.85 | 1.74 | 8.5 | <0.001 |
| Pathologic T-stage (T2a as reference) | T2b | 0.11 | 0.01 | 2.4 | 0.16 |
| T2c | 0.13 | 0.01 | 1.3 | 0.08 | |
| T3a | 0.38 | 0.04 | 3.3 | 0.38 | |
| T3b | 0.62 | 0.07 | 5.8 | 0.68 | |
| T4 | 0.34 | 0.03 | 4.5 | 0.42 | |
| Pathologic N-stage (N0 as reference) | N1 | 0.57 | 0.25 | 1.3 | 0.17 |
Note: The multivariate model was fit using the fraction of each cell type separately (high vs. low) together with the overall Gleason score, pathologic T and N stages. The P values are derived from Wald test.
aThe fraction of each cell type was categorized into high and low based on the optimal cutpoint identified using the maximally selected rank statistics.
In the natural history cohort, a high fraction of neoplastic cells in the highly attended patches is significantly associated with shorter OS (P-value = 0.02) whereas high ratios of immune (P-value = 0.004), stromal (P-value = 0.002), and stromal to neoplastic cells (P-value = 0.01) are significantly associated with longer OS (Fig. 6A). Similarly, high ratios of immune (P-value = 0.01), necrotic (P-value = 0.01), benign epithelial (P-value < 0.001), stromal (P-value = 0.002), and stromal to neoplastic ratio (P-value = 0.001) are each associated with significantly longer metastasis-free survival (Fig. 6B). Multivariate survival analysis confirmed these findings after adjusting for important clinico-pathologic variables including pathological T-stage, preoperative prostate-specific antigen (PSA) levels, age at the time of diagnosis, and receiving androgen-deprivation therapy (ADT; Table 2). Altogether, these results show that the ERG status prediction model is also capable of deciphering large gigapixel WSIs to capture biologically informative small tissue regions whose cellular composition is associated with survival in patients with prostate cancer.
Figure 6.
Cellular composition in the highly attended patches is associated with overall and metastasis-free survival in the natural history cohort. KM curves showing the association between the fraction of each nuclear type and overall (A) and metastasis-free survivals (B). The fraction of each cell type was calculated by dividing the absolute number of that cell type over the number of all cells in the highly attended patches of each slide. The stromal-to-neoplastic ratio was calculated by dividing the number of stromal cells by that of neoplastic cells in the highly attended patches of each slide. These fractions were then then binarized into high versus low using maximally selected log-rank statistics.
Table 2.
Multivariate Cox proportional hazards model examining the association of cellular composition of the highly attended patches with OS in the natural history cohort.
| Variable | HR | Lower 95% CI | Upper 95% CI | P value | |
|---|---|---|---|---|---|
| Cellular composition (high vs. low)a | Neoplastic | 2.5 | 1.05 | 6.0 | 0.04 |
| Inflammatory | 0.39 | 0.16 | 0.94 | 0.04 | |
| Stromal | 0.31 | 0.11 | 0.83 | 0.02 | |
| Necrotic | 3.58 | 1.03 | 12.4 | 0.04 | |
| Benign epithelial | 0.50 | 0.17 | 1.5 | 0.21 | |
| Stromal-to-neoplastic ratio | 0.33 | 0.13 | 0.86 | 0.02 | |
| Gleason score (Gleason <8 as reference) | Gleason > = 8 | 0.92 | 0.27 | 3.1 | 0.9 |
| Pathologic T-stage (T2 as reference) | T3a | 0.50 | 0.13 | 2.0 | 0.32 |
| T3b | 1.4 | 0.4 | 5.0 | 0.6 | |
| Preoperative PSA | 0.97 | 0.93 | 1.0 | 0.18 | |
| Age | 0.94 | 0.86 | 1.0 | 0.18 | |
| ADT (no therapy as reference) | yes | 9.8 | 2.4 | 39.1 | 0.001 |
Note: The multivariate model was fit using the fraction of each cell type separately (high versus low) together with the overall Gleason score, pathologic T stage, preoperative PSA levels, age at diagnosis, and whether the patient received ADT. The P values are derived from Wald test.
aThe fraction of each cell type was categorized into high and low based on the optimal cutpoint identified using the maximally selected rank statistics.
Discussion
Over the past few years, there has been a tremendous growth in the research and clinical applications of AI and DL in the fields of computational pathology and cancer research. These applications have allowed for automated or semi-automated inspection of large numbers of histopathology images to extract informative spatially resolved features that can be associated with phenotypes of interest. Although DL systems have the potential of reducing the workload of pathologists by automatically detecting known morphological features, they can also help detect previously uncharacterized features. Collectively, these advances have facilitated a broad spectrum of diagnostic and prediction tasks, enabling detection of various clinical and molecular phenotypes, improving disease subtyping, and enhancing our understanding of different cancer types (12, 45). Particularly, semi-supervised learning has been utilized extensively to implement prediction tasks on WSIs using only slide-level labels instead of pixel-level annotations. This has covered a wide array of research and clinical interests with variable complexity (46). For instance, DL systems have been deployed on H&E-stained histopathology slides to detect tumor tissue (16, 17, 27) and for tumor subtyping (20, 22), grading (19), and prognostication (47–49). In addition, DL models have been employed to predict several molecular alterations including for example ER status in breast cancer, BRAF (28) and TP53 (50) mutations, and microsatellite instability (51).
In this study, we introduce a semi-supervised DL model capable of inferring the TMPRSS2:ERG gene fusion status from digitized H&E-stained WSIs The best performing model in our training cohort, which comprised 318 WSIs from the TCGA dataset, achieved an AUC and accuracy of 0.84 and 0.77, respectively. When tested on the TCGA testing set, which included 59 WSIs, this model maintained an AUC of 0.72 and accuracy of 0.70. In addition, we deployed the model on an internal testing cohort (the natural history cohort) with 314 WSIs in which our model maintained its good performance with an AUC of 0.73 and accuracy of 0.69. These results show that our model could maintain its predictive performance on slide cohorts from different institutions and scanned by different technologies.
Although many studies in this field have focused on reporting the predictive performance with little regard for biological interpretation, in our study, we thoroughly addressed the interpretability of our model to understand what distinct morphologic features are associated with its predictions. Specifically, the use of attention-based DL in our study allowed us to assign attention scores for patches contributing to slide-level representation (34) with high scores suggesting the importance of these patches in predicting either ERG-positive or negative cases. With this in mind, we computed the attention scores for all the slide patches and examined the highly attended patches from each slide predicted as either positive or negative to see if there are pathomorphologic or cellular composition features specific to ERG status. To characterize the cellular composition in these highly attended patches, we used HoVer-Net model (38) trained on the PanNuke dataset (37) to segment and classify the nuclei into one of five main nuclear types; neoplastic, immune, stromal (connective tissue), necrotic, and benign epithelium. Notably, highly attended patches for the positive class were enriched in more neoplastic content than their ERG-negative counterparts and were more enriched in necrotic, immune, and stromal cells. We hypothesized that the unique cellular content in these regions might capture prognostic information. For this reason, we examined whether the ratio of each nuclear type is associated with PFS in the TCGA cohort as well as OS and metastasis-free survival in the natural history cohort. Notably, the ratio of neoplastic cells in the highly attended patches was significantly associated with shorter PFS and OS in the TCGA and natural history cohorts, respectively. In contrast, the ratio of immune cells was associated with longer PFS (TCGA cohort), OS, and metastasis-free survival (natural history cohort). These results show that the cellular composition in the relevant tissue regions reflects known biology and can serve as additional validation of our results.
In this study, our model could predict the TMPRSS2:ERG gene fusion status using routine histopathologic images and only slide-level labels without expert pixel-level annotation. This together with other studies with similar scope (28, 50, 51) highlight the rising importance and relevance of AI and computer vision in the field of pathology by assisting pathologists and improving the cost and efficiency of the diagnostic process. It is important to place our findings in the context of the existing diagnostic landscape. Although RT-PCR, and IHC are effective and affordable in many settings, the potential for our model shines in settings where these resources might be sparse. As low-cost scanners gain traction, especially in remote areas, our DL approach to detect ERG fusion from H&E-stained WSIs could become a vital diagnostic tool. Although the accuracy of most existing DL models, including ours, may currently fall short of the threshold for clinical deployment, the potential therapeutic implications of ERG fusion make it an area of continued research interest. The detection of ERG fusion not only has diagnostic potential but can also inform therapeutic decisions, especially in light of recent studies exploring ERG as a therapeutic target. As DL models, like ours, continue to evolve and improve in accuracy, they have the potential to provide a valuable tool in the diagnostic and therapeutic landscape of prostate cancer.
This study has some inherent limitations. First, although RT-PCR and IHC serve as established methods for determining the ERG fusion status, it is important to note that they do not have perfect agreement. For our training cohort, RT-PCR was used, and for the testing cohort, IHC was employed. Although this distinction introduced a layer of complexity, our model's ability to maintain performance despite these differences further underscores its robustness. Second, although the performance of our model was optimal and stable when tested on slides from a different institution and scanned by a different slide scanner (Hamamatsu vs. Aperio), this performance can still improve with further training and re-evaluation on future multi-institutional cohorts. In addition, our model has been trained and tested on radical prostatectomy specimens and would need additional evaluation on prostate biopsy specimens which would further enhance its clinical utility as a diagnostic or screening test for ERG status. Moreover, it is important to note that several DL models have demonstrated performance on par with or even surpassing that of human practitioners. We believe that with further training and validation on larger scale datasets, we would be able to get higher quality diagnostic networks that can be implemented seamlessly into clinical practice. With further advancements, automated ERG fusion prediction from H&E-stained histopathologic slides could soon evolve into a robust, affordable diagnostic tool, significantly reducing the time and effort currently required for costly molecular investigations (52). Finally, although our findings do suggest a higher prevalence of cancer cells in patches associated with TMPRSS2:ERG fusion, and further associate high cancer cell prevalence with worse survival, it is essential to consider this in the context of the broader literature. The relationship between TMPRSS2:ERG fusion and survival outcomes has indeed been a subject of much debate in various studies. Most of these studies present a mixed picture and a strong direct link between fusion and poor outcomes is not consistently supported (53, 54).
Here, we presented a DL system that can predict the status of TMPRSS2:ERG fusion using digitized H&E-stained WSIs from prostate cancer radical prostatectomy specimens. Such tool can potentially be used by clinicians to infer ERG fusion status quickly and accurately. We thoroughly examined the cellular composition of the highly attended patched for cases predicted as either ERG-positive or negative and found a significant association between this composition and OS, PFS, and metastasis-free survivals. Altogether, these findings show the utility of semi-supervised DL models in predicting complex molecular and clinical phenotypes from routine histopathologic slides even without known morphological features or pixel-level annotation.
Supplementary Material
Supplementary Table 1. Performance metrics for the ERG prediction model in the TCGA testing cohort. The table shows the model’s performance in the TCGA testing cohort consisting of 59 whole slide images using different thresholds derived from the training data. PPV: positive predictive value; NPV: negative predictive value; MCC: Matthews Correlation Coefficient. Supplementary Table 2. Performance metrics for the ERG prediction model in the natural history cohort. The table shows the model’s performance in the natural history cohort consisting of 314 whole slide images using different thresholds derived from the training data. PPV: positive predictive value; NPV: negative predictive value; MCC: Matthews Correlation Coefficient.
Supplementary Figure 1. Progression-free survival in the TCGA cohort using the cellular composition of the whole slide images. Kaplan-Meier curves showing the association between the fraction of each nuclear type and progression-free survival. The fraction of each cell type was calculated by dividing the absolute number of that cell type over the number of all cells in the whole slide. The stromal to neoplastic ratio was calculated by dividing the number of stromal cells by that of neoplastic cells in the whole slide. These fractions were then then binarized into high versus low using maximally selected log-rank statistics.
Supplementary Figure 2. Comparison of Cellular Composition by TMPRSS2:ERG Fusion Status and Gleason Score in the TCGA Cohort. The average immune, neoplastic, and stroma to neoplastic ratios across two categories of TMPRSS2:ERG fusion status (Negative and Positive) stratified by Gleason score (<8 and >=8). The immune ratio panel shows the proportion of immune cells, the neoplastic ratio panel displays the proportion of neoplastic cells, and the stroma to neoplastic ratio panel depicts the relative abundance of stromal to neoplastic cells within the analyzed tissue sections. Red bars represent cases with a Gleason score of less than 8, while blue bars represent cases with a Gleason score of 8 or higher.
Acknowledgments
L. Marchionni and M. Omar are supported by the NIH grants U54CA273956 and R01CA200859. M.K. Alexanderani is a fellow supported by the NIH grant T32CA260293.
Footnotes
Note: Supplementary data for this article are available at Molecular Cancer Research Online (http://mcr.aacrjournals.org/).
Authors' Disclosures
M.K. Alexanderani reports support from NIH-NCI T32 CA260293-01A1 during the conduct of the study. T.L. Lotan reports grants from AIRA Matrix, DeepBio, Roche, and nonfinancial support from outside the submitted work. L. Marchionni reports grants from NIH-NCI during the conduct of the study. No disclosures were reported by the other authors. The Editor-in-Chief of Molecular Cancer Research is an author on this article. In keeping with AACR editorial policy, a senior member of the Molecular Cancer Research editorial team managed the consideration process for this submission and independently rendered the final decision concerning acceptability.
Authors' Contributions
M. Omar: Conceptualization, data curation, formal analysis, supervision, visualization, methodology, writing–original draft, project administration, writing–review and editing. Z. Xu: Resources, software, formal analysis, visualization, methodology. S.B. Rand: Software, visualization, methodology. M.K. Alexanderani: Writing–review and editing. D.C. Salles: Resources, data curation. I. Valencia: Writing–review and editing. E.M. Schaeffer: Resources, data curation. B.D. Robinson: Resources, data curation. T.L. Lotan: Conceptualization, resources, data curation. M. Loda: Conceptualization, resources, supervision, funding acquisition, project administration, writing–review and editing. L. Marchionni: Conceptualization, resources, data curation, supervision, funding acquisition, project administration, writing–review and editing.
References
- 1. Rawla P. Epidemiology of prostate cancer. World J Oncol 2019;10:63–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2021;71:209–49. [DOI] [PubMed] [Google Scholar]
- 3. Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun X-W, et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 2005;310:644–8. [DOI] [PubMed] [Google Scholar]
- 4. Perner S, Mosquera J-M, Demichelis F, Hofer MD, Paris PL, Simko J, et al. TMPRSS2-ERG fusion prostate cancer: an early molecular event associated with invasion. Am J Surg Pathol 2007;31:882. [DOI] [PubMed] [Google Scholar]
- 5. Adamo P, Ladomery MR. The oncogene ERG: a key factor in prostate cancer. Oncogene 2016;35:403–14. [DOI] [PubMed] [Google Scholar]
- 6. Gonzalez RS, Messing S, Tu X, McMahon LA, Whitney-Miller CL. Immunohistochemistry as a surrogate for molecular subtyping of gastric adenocarcinoma. Hum Pathol 2016;56:16–21. [DOI] [PubMed] [Google Scholar]
- 7. Mandel P, Wenzel M, Hoeh B, Welte MN, Preisser F, Inam T, et al. Immunohistochemistry for prostate biopsy: impact on histological prostate cancer diagnoses and clinical decision making. Curr Oncol 2021;28:2123–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Shao Y, Nir G, Fazli L, Goldenberg L, Gleave M, Black P, et al. Improving prostate cancer classification in H&E tissue micro arrays using Ki67 and P63 histopathology. Comput Biol Med 2020;127:104053. [DOI] [PubMed] [Google Scholar]
- 9. Chaux A, Albadine R, Toubaji A, Hicks J, Meeker A, Platz EA, et al. Immunohistochemistry for ERG expression as a surrogate for TMPRSS2-ERG fusion detection in prostatic adenocarcinomas. Am J Surg Pathol 2011;35:1014–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Aeffner F, Zarella MD, Buchbinder N, Bui MM, Goodman MR, Hartman DJ, et al. Introduction to digital image analysis in whole-slide imaging: a white paper from the digital pathology association. J Pathol Inform 2019;10:9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Huang W, Randhawa R, Jain P, Iczkowski KA, Hu R, Hubbard S, et al. Development and validation of an artificial intelligence–powered platform for prostate cancer grading and quantification. JAMA Netw Open 2021;4:e2132554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Shmatko A, Ghaffari Laleh N, Gerstung M, Kather JN. Artificial intelligence in histopathology: enhancing cancer research and clinical oncology. Nat Cancer 2022;3:1026–38. [DOI] [PubMed] [Google Scholar]
- 13. Tran KA, Kondrashova O, Bradley A, Williams ED, Pearson JV, Waddell N. Deep learning in cancer diagnosis, prognosis and treatment selection. Genome Medicine 2021;13:152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Yang Z, Ran L, Zhang S, Xia Y, Zhang Y. EMS-net: ensemble of multiscale convolutional neural networks for classification of breast cancer histology images. Neurocomputing 2019;366:46–53. [Google Scholar]
- 15. Araújo T, Aresta G, Castro E, Rouco J, Aguiar P, Eloy C, et al. Classification of breast cancer histology images using Convolutional Neural Networks. PLoS One 2017;12:e0177544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Chuang W-Y, Chang S-H, Yu W-H, Yang C-K, Yeh C-J, Ueng S-H, et al. Successful identification of nasopharyngeal carcinoma in nasopharyngeal biopsies using deep learning. Cancers 2020;12:507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Cruz-Roa A, Gilmore H, Basavanhally A, Feldman M, Ganesan S, Shih NNC, et al. Accurate and reproducible invasive breast cancer detection in whole-slide images: a deep learning approach for quantifying tumor extent. Sci Rep 2017;7:46450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Campanella G, Hanna MG, Geneslaw L, Miraflor A, Werneck Krauss Silva V, Busam KJ, et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat Med 2019;25:1301–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Ström P, Kartasalo K, Olsson H, Solorzano L, Delahunt B, Berney DM, et al. Artificial intelligence for diagnosis and grading of prostate cancer in biopsies: a population-based, diagnostic study. Lancet Oncol 2020;21:222–32. [DOI] [PubMed] [Google Scholar]
- 20. Sirinukunwattana K, Domingo E, Richman SD, Redmond KL, Blake A, Verrill C, et al. Image-based consensus molecular subtype (imCMS) classification of colorectal cancer using deep learning. Gut 2021;70:544–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Wang X, Chen H, Gan C, Lin H, Dou Q, Tsougenis E, et al. Weakly supervised deep learning for whole slide lung cancer image analysis. IEEE Transactions on Cybernetics 2020;50:3950–62. [DOI] [PubMed] [Google Scholar]
- 22. Kiani A, Uyumazturk B, Rajpurkar P, Wang A, Gao R, Jones E, et al. Impact of a deep learning assistant on the histopathologic classification of liver cancer. NPJ Digit Med 2020;3:23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Bulten W, Pinckaers H, van Boven H, Vink R, de Bel T, van Ginneken B, et al. Automated deep-learning system for Gleason grading of prostate cancer using biopsies: a diagnostic study. Lancet Oncol 2020;21:233–41. [DOI] [PubMed] [Google Scholar]
- 24. Harder N, Schönmeyer R, Nekolla K, Meier A, Brieu N, Vanegas C, et al. Automatic discovery of image-based signatures for ipilimumab response prediction in malignant melanoma. Sci Rep 2019;9:7449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Coudray N, Ocampo PS, Sakellaropoulos T, Narula N, Snuderl M, Fenyö D, et al. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat Med 2018;24:1559–67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Bilal M, Raza SEA, Azam A, Graham S, Ilyas M, Cree IA, et al. Development and validation of a weakly supervised deep learning framework to predict the status of molecular pathways and key mutations in colorectal cancer from routine histology images: a retrospective study. The Lancet. Digital Health 2021;3:e763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Couture HD, Williams LA, Geradts J, Nyante SJ, Butler EN, Marron JS, et al. Image analysis with deep learning to predict breast cancer grade, ER status, histologic subtype, and intrinsic subtype. NPJ Breast Cancer 2018;4:30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Kim RH, Nomikou S, Coudray N, Jour G, Dawood Z, Hong R, et al. A deep learning approach for rapid mutational screening in melanoma. 610311 ( 2020).
- 29. Schaumberg AJ, Rubin MA, Fuchs TJ. H&E-stained whole slide image deep learning predicts spop mutation state in prostate cancer. 064279 ( 2018).
- 30. Abeshouse A, Ahn J, Akbani R, Ally A, Amin S, Andry CD, et al. The molecular taxonomy of primary prostate cancer. Cell 2015;163:1011–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Sboner A, Habegger L, Pflueger D, Terry S, Chen DZ, Rozowsky JS, et al. FusionSeq: a modular framework for finding gene fusions by analyzing paired-end RNA-sequencing data. Genome Biol 2010;11:R104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Cheng HD, Jiang XH, Sun Y, Wang J. Color image segmentation: advances and prospects. Pattern Recognit 2001;34:2259–81. [Google Scholar]
- 33. Otsu N. A threshold selection method from gray-level histograms. IEEE Trans Syst Man Cybern 1979;9:62–66. [Google Scholar]
- 34. Lu MY, Williamson DFK, Chen TY, Chen RJ, Barbieri M, Mahmood F. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat Biomed Eng 2021;5:555–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Ilse M, Tomczak JM, Welling M. Attention-based deep multiple instance learning. Proceedings of the International Conference on Machine Learning; 2018.
- 36. Matthews BW. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochim Biophys Acta 1975;405:442–51. [DOI] [PubMed] [Google Scholar]
- 37. Gamper J, Koohbanani NA, Benes K, Graham S, Jahanifar M, Khurram SA, et al. Pannuke dataset extension, insights and baselines. arXiv preprint arXiv:2003.10778; 2020.
- 38. Graham S, Vu QD, Raza SEA, Azam A, Tsang YW, Kwak JT, et al. Hover-Net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med Image Anal 2019;58:101563. [DOI] [PubMed] [Google Scholar]
- 39. Lausen B, Schumacher M. Maximally selected rank statistics. Biometrics 1992;48:73–85. [Google Scholar]
- 40. Lausen B, Hothorn T, Bretz F, Schumacher M. Assessment of optimal selected prognostic factors. Biometr J 2004;46:364–74. [Google Scholar]
- 41. Kaplan EL, Meier P. Nonparametric estimation from incomplete observations. J Am Statist Assoc 1958;53:457–81. [Google Scholar]
- 42. Therneau TM., until 2009), T. L. (original S.→R port and maintainer R, Elizabeth A, Cynthia C. survival: Survival Analysis 2022.
- 43. survival. https://cran.r-project.org/web/packages/survival/citation.html.
- 44. Kassambara A, Kosinski M, Biecek P, Fabian S. survminer: Drawing survival curves using “ggplot2. 2021.
- 45. Baxi V, Edwards R, Montalto M, Saha S. Digital pathology and artificial intelligence in translational medicine and clinical practice. Mod Pathol 2022;35:23–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Echle A, Rindtorff NT, Brinker TJ, Luedde T, Pearson AT, Kather JN. Deep learning in cancer pathology: a new generation of clinical biomarkers. Br J Cancer 2021;124:686–96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Kulkarni PM, Robinson EJ, Sarin Pradhan J, Gartrell-Corrado RD, Rohr BR, Trager MH, et al. Deep learning based on standard h&e images of primary melanoma tumors identifies patients at risk for visceral recurrence and death. Clin Cancer Res 2020;26:1126–34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Wulczyn E, Steiner DF, Xu Z, Sadhwani A, Wang H, Flament-Auvigne I, et al. Deep learning-based survival prediction for multiple cancer types using histopathology images. PLoS One 2020;15:e0233678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Bychkov D, Linder N, Turkki R, Nordling S, Kovanen PE, Verrill C, et al. Deep learning based tissue analysis predicts outcome in colorectal cancer. Sci Rep 2018;8:3395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Noorbakhsh J, Farahmand S, Foroughi pour A, Namburi S, Caruana D, Rimm D, et al. Deep learning-based cross-classifications reveal conserved spatial behaviors within tumor histological images. Nat Commun 2020;11:6367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Kather JN, Pearson AT, Halama N, Jäger D, Krause J, Loosen SH, et al. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat Med 2019;25:1054–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Hameed BMZ, Shah M, Naik N, Ibrahim S, Somani B, Rice P, et al. Contemporary application of artificial intelligence in prostate cancer: an i-TRUE study. Therapeutic Advances in Urology 2021;13:1756287220986640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Graff RE, Pettersson A, Lis RT, DuPre N, Jordahl KM, Nuttall E, et al. The TMPRSS2:ERG fusion and response to androgen deprivation therapy for prostate cancer. Prostate 2015;75:897–906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Hägglöf C, Hammarsten P, Strömvall K, Egevad L, Josefsson A, Stattin P, et al. TMPRSS2-ERG expression predicts prostate cancer survival and associates with stromal biomarkers. PLoS One 2014;9:e86824. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Table 1. Performance metrics for the ERG prediction model in the TCGA testing cohort. The table shows the model’s performance in the TCGA testing cohort consisting of 59 whole slide images using different thresholds derived from the training data. PPV: positive predictive value; NPV: negative predictive value; MCC: Matthews Correlation Coefficient. Supplementary Table 2. Performance metrics for the ERG prediction model in the natural history cohort. The table shows the model’s performance in the natural history cohort consisting of 314 whole slide images using different thresholds derived from the training data. PPV: positive predictive value; NPV: negative predictive value; MCC: Matthews Correlation Coefficient.
Supplementary Figure 1. Progression-free survival in the TCGA cohort using the cellular composition of the whole slide images. Kaplan-Meier curves showing the association between the fraction of each nuclear type and progression-free survival. The fraction of each cell type was calculated by dividing the absolute number of that cell type over the number of all cells in the whole slide. The stromal to neoplastic ratio was calculated by dividing the number of stromal cells by that of neoplastic cells in the whole slide. These fractions were then then binarized into high versus low using maximally selected log-rank statistics.
Supplementary Figure 2. Comparison of Cellular Composition by TMPRSS2:ERG Fusion Status and Gleason Score in the TCGA Cohort. The average immune, neoplastic, and stroma to neoplastic ratios across two categories of TMPRSS2:ERG fusion status (Negative and Positive) stratified by Gleason score (<8 and >=8). The immune ratio panel shows the proportion of immune cells, the neoplastic ratio panel displays the proportion of neoplastic cells, and the stroma to neoplastic ratio panel depicts the relative abundance of stromal to neoplastic cells within the analyzed tissue sections. Red bars represent cases with a Gleason score of less than 8, while blue bars represent cases with a Gleason score of 8 or higher.
Data Availability Statement
Digitized whole slide images from the TCGA prostate adenocarcinoma cohort are publicly available through the GDC data portal and can be accessed using the following link: https://portal.gdc.cancer.gov/.