Abstract
Anaplastic large cell lymphoma (ALCL) is a mature T-cell lymphoma that accounts for for 10–15% of childhood lymphomas. Despite the observation that more than 90% of pediatric cases harbor the anaplastic lymphoma kinase (ALK) rearrangement resulting in aberrant ALK kinase expression, there is significant clinical, morphologic, and biological heterogeneity. To gain insights into the genomic aberrations and molecular heterogeneity within ALK-positive ALCL(ALK+ ALCL), we analyzed 46 pediatric ALK+ ALCLs by whole-exome sequencing, RNA-sequencing, and DNA methylation profiling. Whole-exome sequencing found on average 25 SNV/Indel events per sample with recurring genetic events in regulators of DNA damage (TP53, MDM4), transcription (JUNB), and epigenetic regulators (TET1, KMT2B, KMT2A, KMT2C, KMT2E). Gene expression and methylation profiling consistently subclassified ALK+ ALCLs into two groups characterized by diferential ALK expression levels. The ALK-low group showed enrichment of pathways associated with immune response, cytokine signaling, and a hypermethylated predominant pattern compared to the ALK− high group, which had more frequent copy number changes, and was enriched with pathways associated with cell growth, proliferation, metabolic pathways, and. Taken together, these findings suggest that there is molecular heterogeneity within pediatric ALK+ALCL, predicting distinct biological mechanisms that may provide novel insights into disease pathogenesis and represent prognostic markers.
Introduction
Anaplastic large cell lymphoma (ALCL) is a peripheral T-cell lymphoma that accounts for 10–15% of all childhood lymphomas 1, 2. ALCLs exhibit a broad morphologic spectrum characterized by infiltrates of pleomorphic cells and expression of CD30 antigen 3, 4. ALCL is divided into ALK-positive (ALK+) and ALK-negative (ALK−) based on the presence of the translocation involving ALK 3, 5. More than 90% of ALCL in children carry a translocation of the ALK gene at the 2p23 locus. In ALK+ ALCL, nucleophosmin 1 (NPM1) is the major fusion partner, which is described by the translocation t(2;5)(p23;q35)6. The translocation results in the expression of a constitutively active oncogenic ALK kinase, which activates multiple downstream signaling pathways, driving malignant transformation and growth of the tumor cells 7–9.
ALCL is an aggressive lymphoma, and most pediatric patients with ALCL present with advanced-stage disease 10. Even though the standar treatment of newly diagnosed patients with ALK+ALCL is anthracycline-based combination chemotherapy, the most effective and safe treatment for children with advanced ALCL remains to be established11 12. While several therapeutic strategies have been explored in clinical trials for pediatric ALCL, they have not led to significant improvement in event-free survival (EFS), with approximately 30% of patients continuing to experience disease progression or recurrence within two years of diagnosis 10, 13–19. Notably, clinical, morphologic, and biological factors are associated with prognosis and risk of treatment failure, such as morphological patterns and the assessment of minimal disseminated disease (MDD) at diagnosis 18, 20–22.
Integrative strategies using genomic, transcriptomic, and epigenetic profiling offer complementary opportunities to elucidate novel pathogenic insights, functional modules, and deregulated networks in lymphoma pathogenesis. Although the gene expression, genomic and methylation profiles of peripheral T-cell lymphomas 23, 24, including ALCL 25–27, have been previously characterized, an integrative approach, specifically focused on pediatric ALK+ ALCL has not been performed. Herein, we describe the genomic landscape of pediatric ALK+ ALCL and define the spectrum of somatic mutations, copy number alterations, and gene expression and methylation patterns within ALK+ALCL to identify molecular mechanisms that contribute to its biological heterogeneity.
Material and Methods
Subject cohorts and sample details
ALCL cases were collected with informed consent prospectively under a protocol approved by the St. Jude Children’s Research Hospital Institutional Review Board (IRB). The study cohort comprised of 46 fresh frozen tumor samples obtained from the St. Jude tissue resources core facility (N=10), Children’s Cancer and Leukaemia Group (CCLG) (N=18), and Children’s Oncology Group (COG) (N=18). For cases with available material for reviewed, the diagnosis was confirmed by a hematopathologist (V.L.) according to the revised 4th edition of the WHO classification of hematolymphpid tumors 28. Information about the % fraction of neoplastic cells in the specimens was available for 45 out of 46 specimens, and estimated to exceed 40% (ranging from 60% to 100%) as determined based on morphologic and/or immunohistochemical studies (applying ALK1 immunostain) for 27 samples with available tissue sections or provided by the tissue biorepository for the COG samples. Matched germline DNA was available for 12 patients. For the St. Jude samples (n=8), germline DNA was obtained from morphologically and immunohistochemically negative bone marrow biopsies obtained at the time of diagnosis; for the CCLG samples (n=4) matched germline DNA was provided by the tissue bank. The clinical and histological characteristics of the pediatric patients with ALCL are summarized in Table 1, and detailed information of the study with available data per patient is given in Supplementary Table S1.
Table 1.
Clinical and pathological characteristics of the pediatric ALCL cohort.
| Clinical and biological characteristics | N=46 ¶ | |
|---|---|---|
|
| ||
| Gender | Female | 12 |
| Male | 34 | |
| Median age at diagnosis (yrs) | ≤ 8.5 | 15 |
| >8.5 | 31 | |
| Stage at diagnosis | I-III | 26 |
| IV | 7 | |
| Unavailable | 13 | |
| Mediastinal involvement | Yes | 17 |
| No | 8 | |
| Unavailable | 21 | |
| Skin involvement | Yes | 5 |
| No | 26 | |
| Unavailable | 15 | |
| Histological patterns | Common | 27 |
| Other* | 7 | |
| Unavailable | 12 | |
Other: other than common patterns, including the small cell variant (SCV) and lymphohistiocytic (LH) variant.
One paired sample (diagnostic-relapsed) is included.
Whole exome sequencing analysis
Whole exome sequence (WES) analysis was performed as previously described 29. Briefly, putative SNVs and indel variants were detected using SNPdetector 30. The reference human genome assembly NCBI Build 37 was used to map all samples. Further evaluation of SNVs and indels was performed by manual review of the BAM files using Bambino 31. Non-silent coding variations present exclusively in tumors were considered somatic mutations. For variants derived from non-matching germline samples, we applied the unpaired pipeline as has been described previously 32. Non-silent mutations were compared to the Exome Variant Server (EVS) (National Heart, Lung, and Blood Institute Exome Sequencing Project; https://evs.gs.washington.edu/EVS/) and to a database of germline variations identified in the Pediatric Cancer Genome Project 33. Novel variants that passed this germline filter were annotated with dbSNP v141 and manually reviewed. Known somatic sequence mutations and genes were annotated by the Catalogue of Somatic Mutations in Cancer (COSMIC) database 34 and grouped as putative somatic mutations while others were considered variants of unknown origin. The mapping statistics and coverage for each sample are summarized in Supplementary Table S2.
Validation of Somatic Variants with targeted captured sequencing (TCS)
Mutations were validated using TCS (n =22 samples; AZENTA Life Sciences) designed to cover SNV and indel variants called by WES data (information about the lise of variant evaluated by TCS is included in supplements table S5 and S4). Raw BCL files generated by the sequencer were converted to FASTQ files for each sample using bcf2fastq v2.19. Sequencing adapters and low quality bases in raw reads were trimmed using Trimmomatic 0.39. Cleaned reads were then aligned to the GRCh37 reference genome using Sentieon 202112.01. Each variant was visualized and inspected manually in the IGV viewer35.
Among the 46 patients with WES, TCS was performed on 22 patients with available fresh frozen material for analysis (AZENTA Life Sciences, Indianapolis, IN, USA). A TargetGxOne custome gene panel was developed based on the mutated genes detected by WES in the 22 cases submitted for TCS (Supplemental Table S13). Illumins MiSeq, 2×150bp configuration was used for the sequencing (AZENTA Life Siences). 196 somatic SNV/Indel calls by the WES variant-calling pipeline from 46 patients were covered by the custom capture panel, and 180 out of 196 variants (90.9%) were validated (Supplementary Table S5, S6). For the 36 cases (32 diagnostic and 4 relapsed samples) with RNAseq data, 126 of 132 (95.4%) somatic SNV calls from WES data were validated by RNAseq (Supplementary Table S5). In three cases with the recurrent JUNB A282V mutation was also validate by Sanger sequencing (Suppl. Fig3.B).
Copy number variant analysis using WES data
Copy number analysis was performed using CNVkit version 0.9.1 on the exome sequencing data 36. Target and anti-target regions were defined based on Nextera rapid capture region and “access-5k-mappable.hg19.bed”. The copy number reference control was pooled from germline exome sequencing. Due to the inherent noise of the exome-based copy number analysis, for each copy number altered region, we manually visualized the B-allele frequency shift to validate the copy-number alterations (Supplementary Figure S4, B-C).
RNA sequencing, mapping, fusion detection, and relapse signature analysis
Total RNA was prepared from cryopreserved tumor samples by a single-step RNA isolation method (TRIzol Reagent, Life Technologies), mechanically homogenized with the TissueRuptor (Qiagen), and quantified using the Agilent 2100 Bioanalyzer (Agilent Technologies). Indexed RNA sequencing (RNA-seq) libraries were constructed from 250 ng of total RNA by using the TruSeqTotal Stranded RNA Library Prep Kit (Illumina). Each library was sequenced in a paired-end mode, using 1 lane of the HiSeq 2000 (Illumina) flow cell, which generated 2–3 × 100 bp reads. RNA-seq reads were aligned by using the BWA37 (0.5.10) aligner to the following database files: (i) the humanGRCh37-lite reference sequence; (ii) RefSeq; (iii) a sequence file representing all possible combinations of non-sequential pairs in RefSeq exons; and (iv) the AceView database flat file downloaded from the UCSC Genome Browser, which represents transcripts constructed from human ESTs. The mapping results from databases (ii)–(iv) were aligned to human reference genome coordinates, and a BAM file was constructed by selecting the best alignment from the 4 databases. The mapping statistics and coverage for each sample are summarized in Supplementary Table S3. Structural variations were detected by using CICERO, a novel algorithm that uses de novo assembly to identify structural variation in RNA-seq data 38. The ALK fusion transcript was manually reviewed by using the Bambino viewer 31 and the Integrated Genomics Viewer 39. Mapped reads were counted with HTseq 40 and normalized to FPKM (fragments per kilobase of transcript per million mapped reads) (Supplementary Table S4).
To generate the gene expression signature of relapsed ALCL, differential gene expression analysis was performed comparing relapse (n = 4) vs. diagnosis (n = 32) samples. Genes significantly up-regulated in relapsed samples (P < 0.01, Log2FC > 1, and adjusted p-value < 0.25) were used as the gene set for single-sample gene set enrichment analysis (ssGSEA). Gene expression pathway analysis was performed using GSEA 41. The Hallmark, KEGG and Oncogene databases were used for GSEA. Genes were ranked using signal2noise with 1000 permutations performed on the gene set. xCell was applied to define the immune infiltration 42.
DNA methylation analysis
Genomic DNA (1 μg) from 37 ALCL was bisulfite converted by using the EZ DNA Methylation kit (Zymo Research Corp, Irvine, CA, USA). Converted samples were processed and hybridized to the Infinium MethylationEPIC BeadChip (850K) system (Illumina, San Diego, CA, USA). The methylation score of each CpG site in the array is represented as a beta (β) value and was computed using the methylation module of the GenomeStudio software (version 1.9.0; Illumina). Beta-values were converted to M-values 43 prior to differential methylation analysis and clustering. Each probe was correlated to the ALK gene expression by the Spearman rank correlation. An absolute rho value of 0.75 was used as the cutoff to identify methylated genes for downstream pathway analysis by ENRICHR. The differentially methylated regions were characterized by the Wilcoxon Test with Benjamini Hochberg (BH) Adjusted Pval < 0.05. Probes residing in chromosome X and Y and with SNP VAF > 0.01 were removed from the analysis.
Cluster analysis of RNAseq and DNA methylation data
Clustering for both RNAseq and methylation array data was performed by calculating the top variable genes/probes using MAD, Variance, and DIP. Bootstrap hierarchical clustering was performed using a combination of “complete” and “average” hierarchical clustering algorithms. To determine the number of features that produce a stable hierarchical clustering assignment, we calculate the bootstrap probability while steadily increasing the optimal number of features. The cluster assignment was then compared to the top 1000 variable gene/probe’s cluster assignment. Finally, we took the consensus across the different methods for the final group assignment. Differential gene expression analysis was performed by LIMMA 44 using log2 fold chang (log2FC) > 1.0 and FDR adjusted p-value < 0.05 as cutoffs.
Statistical methods
The student’s t-test, two-tailed, assuming equal variances, and Wilcox signed-rank test were used to compare two experimental groups. The Fisher’s exact test was used to test for the consistency of the subgroup assignment by RNAseq and methylation. Overall survival and progression-free survival were defined as the time difference between the date of diagnosis and the date of death or relapse. Patients who were alive at the time of the last follow up were considered censored. Survival curves were compared via log-rank tests.
Data availability and accession codes
Genomic data have been deposited in the European Genome-phenome Archive (EGA), which is hosted by the European Bioinformatics Institute (EBI), under accession EGAS00001004189. The accession number for the Illumina methylation reported is GEO: GSE186487. All other remaining data are available within the Article and Supplementary Files.
Results
Sequencing of pediatric ALK+ ALCL
We performed next-generation sequencing on a cohort of 46 pediatric ALK+ ALCLs, including 42 tumor samples obtained at diagnosis and four relapsed samples (1 patient with paired diagnostic-relapsed samples) (Supplementary Figure S1.). The histological pattern, defined as common, lymphohistiocytic (LH) or small cell variant (SCV), was available for a subset of patients (34 out of 46) in our cohort. For the remaining cases the subtype was not included in the pathology report or a tissue section available for morphologic assessment was not sufficient to define the histological pattern with confidence. RNA sequencing was performed on 44 samples with available material; however, 10 samples failed quality control and were excluded from the analysis. For all the cases, the presence of the ALK fusion was confirmed by the RNA sequencing data, and for cases with available tissue material, the expression was also verified by ALK1 immunohistochemistry. RNASeq analysis confirmed that the NPM1-ALK was the most common fusion identified in 40 out of 46 patient samples (Figure 1; Supplementary Table S8). Remaining ALK+ patients were characterized by ATIC::ALK (n = 2), TPM4::ALK (n = 2), and EEF1G::ALK (n = 2). The two patients with a novel EEF1G::ALK fusion were previously published by our group 45.
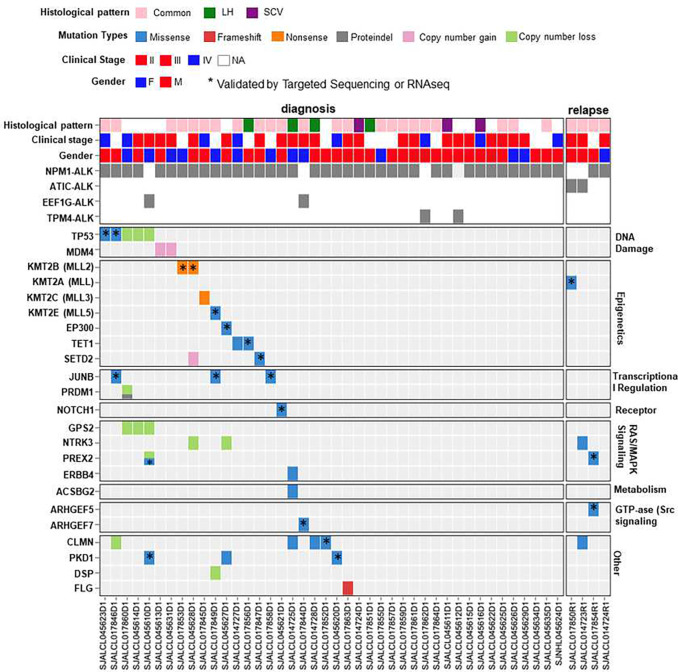
Figure 1. Genomic alterations in pediatric ALCL.
Heatmap demonstrates the somatic mutation profile and copy number (CN) alterations identified in the cohort of pediatric ALK+ALCL samples by WES, separated by gene functional groups. Only genetic alterations with presumed functional consequences are shown. Split cell for the PRDM1 gene indicates more than one mutation. (*) indicates mutations validated by targeted sequencing or RNAseq.
Whole exome sequencing (WES) was performed on all cases with a median coverage of 92X in the tumor (Supplementary Table S4). Matched germline DNA was available for 12 cases. Sample read coverage varied from 36 to 198 million reads with an average of approximately 120 million reads per sample, and ≥96% reads were mapped to the reference human genome (hg19). For the 46 cases, the WES for the 12 samples with paired germline DNA identified a total of 238 SNVs and 18 indels. The somatic mutation rate was estimated to be 25.6 events per sample in paired germline specimens. For the 34 samples with tumor-only material, mutations were filtered to exclude silent and UTR mutations, resulting in a total of 300 amino acid variants (192 SNVs and 108 indels; Supplementary Table S5, S6, S7). Next, we compared the mutation burden of the ALCL samples to other pediatric malignancies analyzing WES data from the St Jude Pediatric Genome Project 46, 47. The mutational burden in pediatric ALCL samples was higher than the mutational burden in pediatric acute leukemia but lower compared to most common pediatric solid tumors, such as adrenocortical carcinoma, osteosarcoma, rhabdomyosarcoma, and high-grade glioma (Supplementary Figure 2B).
In Figure 1, we highlighted genes with genomic alterations that (i) were identified in more than 2 cases, (ii) validated by RNA seq, (iii) are known to have biological significance in cancer, and (iv) have been previously shown to be altered in lymphoma 48, 49. 25 out of 46 patient samples observed somatic events (SNV, CNV) in genes associated with DNA damage, epigenetic and transcriptional regulators, metabolism, Rho-family GTPases, and RAS-MAPK signaling pathways. Mutations in epigenetic regulators (EP300, KMT2A, KMT2D, KMT2C, KMT2E, TET1, SETD2) were observed in 9 out of 46 cases. Genomic alterations of components of the JAK/STAT pathway, which frequently occurs in ALK− ALCL 25 were absent in our cohort. Two cases contained single nucleotide mutation in TP53, affecting the DNA binding (R280K) and tetramerization domains (Q331H). Three out of 46 cases demonstrated a recurrent JUNB mutation (A282V) (Supplementary Figure 3A). This mutation affects the conserved DNA binding domain of JUNB and has been recently described in three cases in a cohort of ALK− ALCL50. JUNB is a member of the AP-1 transcription factors, and has been shown in previous studies to regulate tumor growth and proliferation in ALK+ALCL 50–53. 50. The mutation was validated by RNAseq in all three cases but with a higher VAF possibly indicating that there is a preference for the expression of the mutated allele (Supplementary Figure 3B). Of note, all the cases with the JUNB mutation showed similarly high tumor content.
Copy number analysis detected 45 genetic alterations in 13 out of 46 ALCL tumors (Supplementary Table S9; Supplementary Figure S4A) and validated by B-allele frequency (Supplementary Figure S4B, S4C). CN alterations included losses of 17p13.1 (3/46 samples), which encompasses TP53 and GPS2, 6q21/PRDM1 (one sample with both indel and copy number loss), and 10p4/GATA3 (2 samples), and gains of different regions of chromosome 1q, including the region of 1q32.1/MDM4 (2 samples) and 1q44/AKT3 (2 samples). In agreement with previous studies 54, 55, TP53 alterations were infrequent in our pediatric cohort. Collectively, there were five cases (5/46; 11%) with genomic alterations affecting TP53 and these did not overlap with the two cases that showed gain of 1q23.1/MDM4.
Gene expression signatures and methylation profiling identify two subgroups of pediatric ALCL
To identify distinct molecular signatures and to assess potential molecular heterogeneity within ALK+ALCL, we performed an unsupervised clustering analysis of RNAseq data from 32 diagnostic samples, excluding the four relapsed samples (Supplementary Table S1 and Figure S1). Two distinct subgroups of patients were identified with greater than 70% bootstrapping support across clustering methods (Figure 2A; Supplementary Table S4). The two groups could be defined based on differential ALK expression levels into ALK-high (n=14) and ALK-low group (n=18) (Figure 2B) 56, 57.
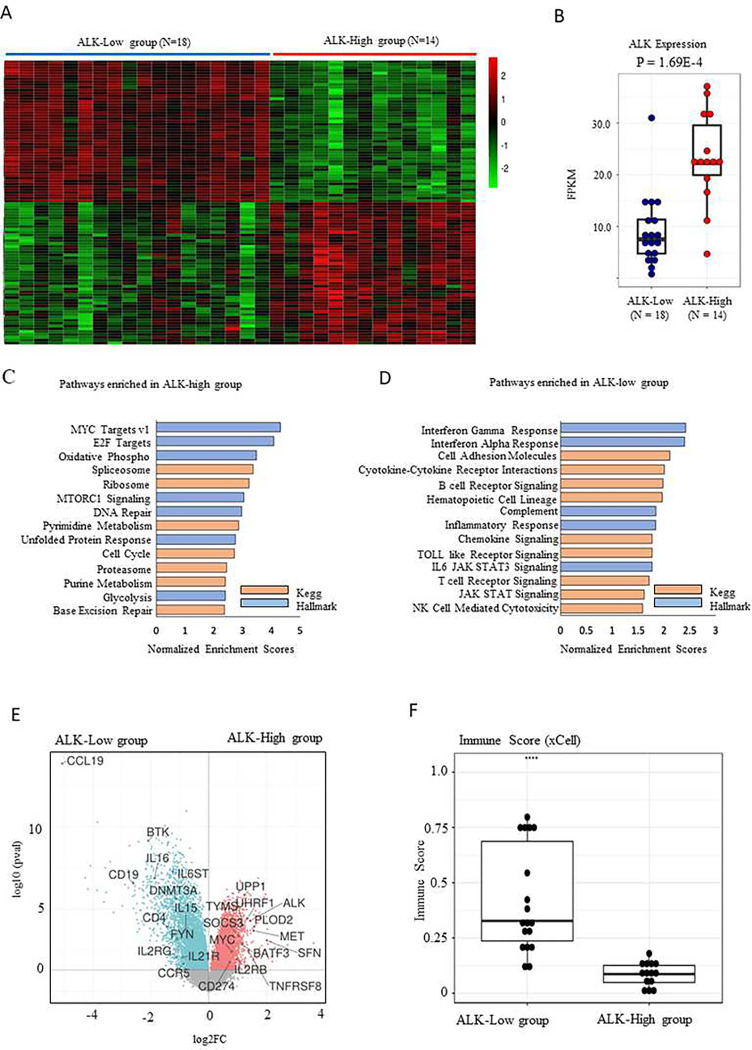
Figure 2. Gene expression profile classifies pediatric ALCL into two groups.
(A) Unsupervised hierarchical clustering reveals two distinct groups. The top 100 differentially expressed genes are shown as a heatmap. (B) Boxplot of differential ALK expression levels in ALCL samples detected by RNA seq analysis (P=0.026). FPKM, fragments per kilobase of exon per million mapped fragments. (C and D) Pathway analysis in ALK-low and ALK-high groups (KEGG and MSigDB), p-value <0.05 and FDR < 0.05. (E) Differential expression represented as a volcano plot. ALK-low group shows up-regulation of cytokine and immune related markers. ALK-high group shows up-regulation of ALK, MYC, and other proliferative markers. (F) xCell analysis shows differential Immune Scores in ALCL samples.
Among the differentially expressed genes, 2878 were up-regulated, and 3008 were downregulated in the ALK-high group when compared with the ALK-low group. Gene set enrichment analysis using the MSigDB, and KEGG databases found enriched pathways of cell proliferation, growth (MYC, E2F, and MTORC1), pyrimidine/purine metabolism, and glycolysis in the ALK-high group (Figure 2C). Upregulated genes in the ALK high group included UHRF1, a critical regulator for maintaining DNA methylation 58, 59, and the immunoinhibitory molecule, CD274 (PD-L1) 60(Figure 2E). Genes up-regulated in the ALK-low group included the epigenetic regulator, DNMT3A, and genes involved in cytokine-cytokine interaction, including CCL19, ligand of CCR7 61, and interleukin signaling, such as IL16, IL21R, and IL2RG 62 (Figure 2E). Overexpression of the components of the IL2R has been previously shown in ALK+ALCL cell lines and patient samples as we and others observed in the ALK-low group56, 63. Consistently, pathways enriched among ALK-low samples related to immunologic functions, including interferon response, B-cell receptor signaling, cytokine-cytokine receptor interactions, and chemokine signaling pathways (Figure 2D). Using the xCell algorithm, the ALK-low patient samples were characterized by an overall higher immune score compared to the ALK-high group (Figure 2F; Supplementary Figure S5A; T-Test p-value < 0.001; Supplementary Table S11), and higher numbers of immune cells such as B-cells, activated dendritic cells, and T-cells (Supplementary Figure S5B-D).
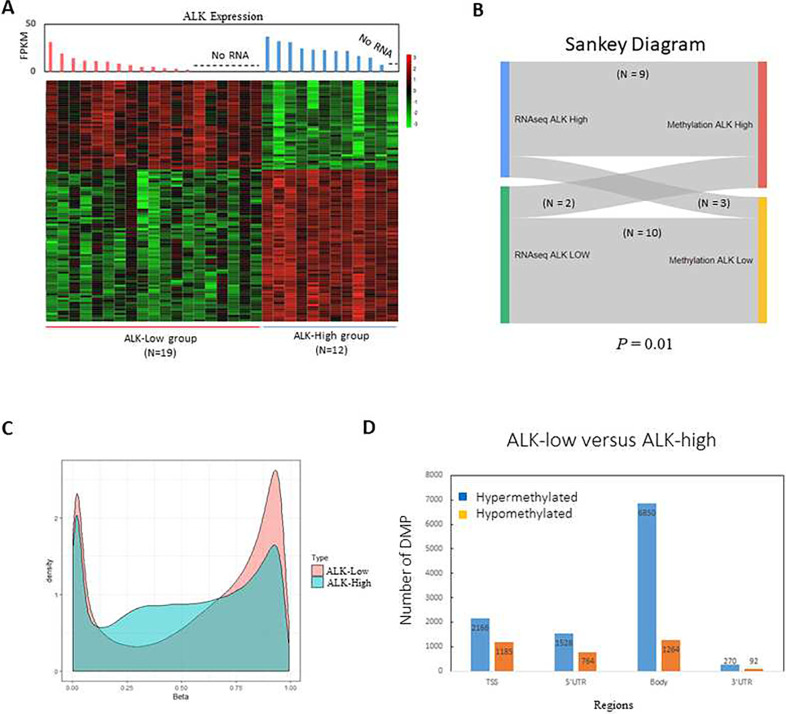
In addition to gene expression analysis, we identified distinct groups of pediatric ALCL using DNA methylation array profiling, an approach that has been extensively used to define molecular subgroups in tumors of the central nervous system 64–66. After excluding cases with <40% tumor content, DNA methylation analysis was performed on 31 diagnostic samples (Supplemental Figure S1). Unsupervised clustering revealed two distinct clusters with greater than 70% bootstrapping support (Figure 3A). For the twenty-four diagnostic samples analyzed by both DNA methylation and RNAseq, 19/24 samples (79%) had a consistent ALK-high or ALK-low group assignment (p-value < 1.96E-3; Figure 3B). For seven samples analyzed by DNA methylation only, six samples were grouped with the ALK-low group, and one sample was grouped with the ALK-high group. The DNA methylation pattern in ALK-low samples resembled a bimodal distribution. ALK-high samples contained moderate levels of methylated probes with a beta value ranging from 0.33 to 0.66 (Figure 3C). More hypermethylated regions were found in the ALK-low samples, particularly in the gene body (6850 hypermethylated, 1264 hypomethylated) and transcription start site (TSS) region (2166 hypermethylated, 1385 hypomethylated) (Figure 3D). This finding is in line with the observation of DNTM3a overexpression in the ALK-low group that may contribute to its hypermethylation profile. The association between DNTM3a and highly methylated genome affecting mostly CpG islands has been described in T-lymphoblastic leukemia67. Interstingly, Hassler et al described a differential methylation pattern between ALK− and ALK+ ALCL tumors mainly based on hypomethylating MVPs in the ALK+ tumors27 that supports our observation of a more hypomethylated profile in the ALK-high group compared to the ALK-low group.
Figure 3. Genome-wide DNA methylation classifies pediatric ALCL in two groups.
(A) Unsupervised clustering with bootstrapping identified two separate clusters. Each probe was then summarized to a single value for each associated genomic region. Each methylation value was annotated as Hyper-methylation or Hypo-methylation. Differential methylation was performed using Wilcoxon rank-sum test. FPKM, fragments per kilobase of exon per million mapped fragments. (B) Sankey diagram representation of the unsupervised hierarchical clustering result. The diagram depicts the group clustering based on RNA expression and methylation profiling. (C) Histogram of the methylation probe intensity (Beta value) in the ALK-high and ALK-low group (D) Bar plot representation of the differentially methylated gene regions. Hyper-methylated regions were more frequently differentially methylated than hypo-methylated regions. DMP, differentially methylated positions; TSS, transcription start site.
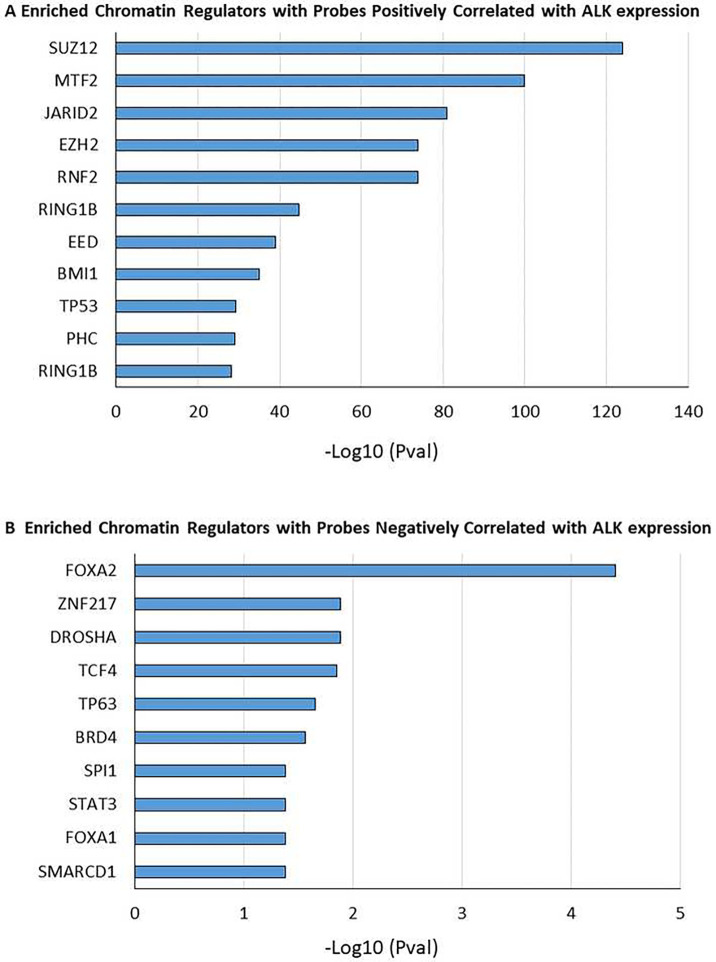
To identify biologically relevant functions and pathways affected by DNA methylation that could be associated with ALK expression, each probe methylation was correlated with the ALK expression levels in the 24 samples with available RNA seq and DNA methylation array data, excluding relapsed samples and samples with tumor content <40%. Methylated regions correlated (positive and negative)with ALK expression were selected for pathway analysis using the ChEA transcription factor database 68. Hypermethylated genes that track with high ALK levels (positively correlated), were enriched for the Polycomb-repressor complex 2 (SUZ12, EZH2, and JARID2 complex; Figure 4A). Hypermethylted genes that track with low ALK expression level (negatively correlated) were enriched for the BRD4 and TCF4 chromatin regulators (Figure 4B), which have been shown to induce MYC expression 69, 70. This finding is consistent with the expression profile of the ALK-high group characterized by increased MYC expression. Prior studies have also described MYC amplification as a mechanism of its overexpression that we did not observe in our cohort based on the analysis of WES data for CNA71, 72. Altogether, the findings suggest that ALK may contribute to differential DNA methylation patterns between the two groups that may subsequently contribute to reprogramming of transcription and distinct expression profiles.
Figure 4. Genes that show correlation between probe methylation status and ALK expression.
Correlation analysis is based on Spearman Rank with positive correlation shown in (A) and negative correlation shown in (B). Both are filtered based on an absolute rho value cutoff of 0.75. Bar plots show enriched pathways from EnrichR’s ChEA_2016 sorted by their log transformed p-value score.
Prognostic significance of genetic alterations and group assignment in pediatric ALCL
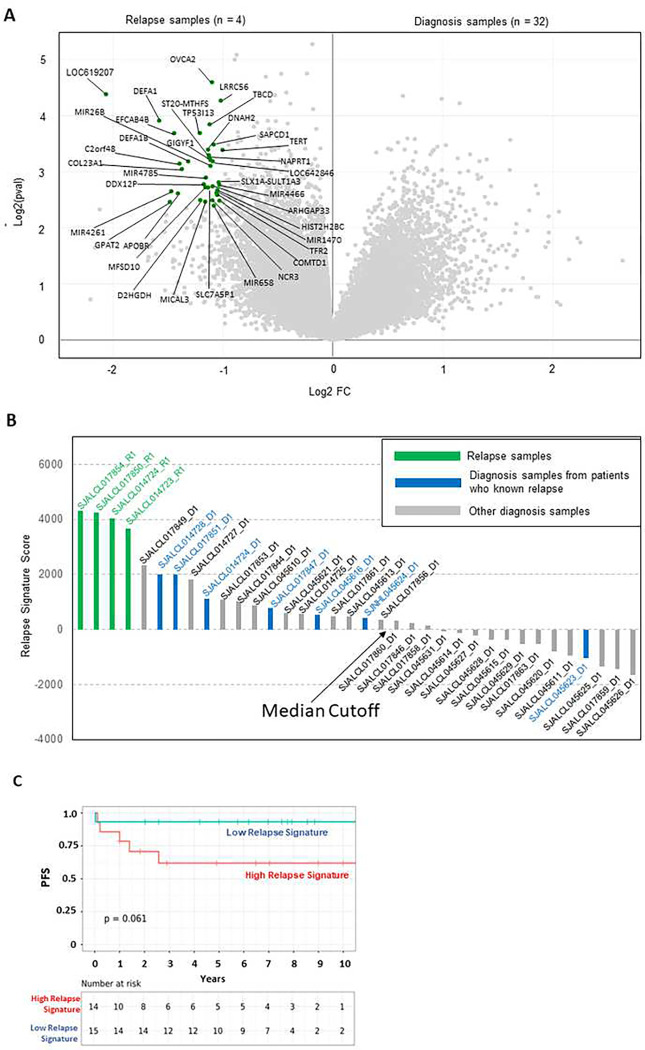
To identify molecular events associated with ALCL relapse, differential gene expression analysis was performed comparing four relapsed samples to 32 diagnostic samples. Differential expression analysis revealed 36 genes up-regulated in relapse compared to diagnostic samples (Figure 5A). Gene set enrichment analysis (GSEA) demonstrated that MYC targets and negative regulation of genes involved in the inflammatory response pathways are enriched in the relapsed samples (Supplementary Figure 6A-B). While the relapse-associated genes from Camille et al. 73 were not consistently upregulated in our cohort of relapsed RNAseq samples (Supplementary Figure 6C-D), diagnosis-associated genes were consistently up-regulated in our cohort of diagnosis ALK+ ALCL, which were enriched for genes involved in the extracellular matrix region (Supplementary Figure 6E-F). To evaluate the relationship between relapse gene signature and clinical outcome, we developed a relapse signature score by single-sample GSEA and analyzed the RNA sequencing data from our pediatric ALCL cohort (Supplementary Table S12). Interestingly, we found an elevated relapse signature present in the diagnosis sample of patients who eventually relapsed (Figure 5B). Of note, only for one patient (SJALCL014724) in our cohort there were available both a diagnostic and a relapse specimen, showing the relapsed signature. Notably, the diagnostic sample for this patient showed the small cell variant that has been consistently associated with worse prognosis in the pediatric ALK+ALCL. Of clinical relevance, the diagnosis samples with high relapse score (above median) trended toward a worse prognosis (OS, p = 0.056; PFS, p = 0.061; Figure 5C). Overall, these results suggest that the gene expression signature at diagnosis could be predictive of relapse associated with an inferior prognosis in a subset of pediatric ALCL patients.
Figure 5. Relapsed signature score in pediatric ALK+ALCL patients.
(A) Differential gene expression analysis between relapse and diagnosis samples from ALCL patients. Genes differentially up-regulated in relapsed samples are highlighted in green. (B) The relapse signature score is calculated based on single-sample GSEA. Samples were ordered based on the calculated relapse signature score. Relapse samples are colored in red. Diagnosis samples of patients who eventually relapse are colored in blue. The remaining diagnosis samples are colored in gray. The median cutoff for the diagnosis samples is indicated by the arrow. (C) Kaplan-Meier graph showing progression free survival (PFS) according to the relapse signature score. The relapse signature score was categorized into HIGH and LOW based on the median level. A statistically significant difference of the Kaplan–Meier survival curves between the high and low malignancy-risk groups was determined by the two-sided log-rank test. The number of patients at risk is listed below the survival curves.
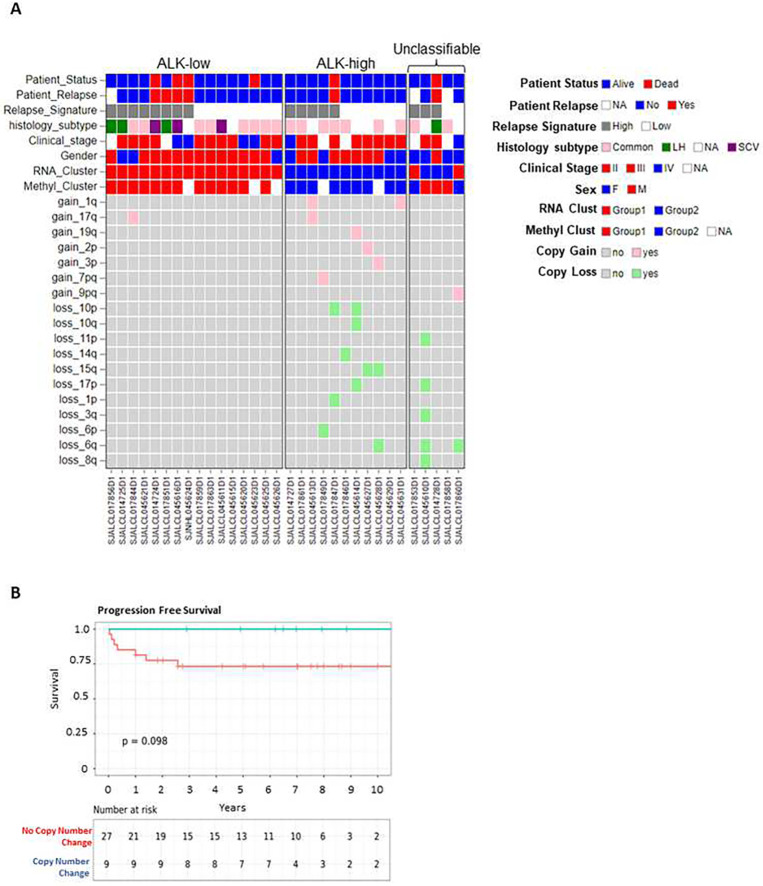
Next, to investigate the potential prognostic role of group assignment and genetic alterations in our cohort, we grouped the diagnostic only ALCL cases (N=32) based on their expression and methylation classification into ALK-low and ALK-high group. Cases with disagreement in group assignment between expression and methylation profile were designated as unclassifiable. (Figure 6A). CN alterations were enriched in the ALK-high group (72.7%; 8/11) compared to the ALK-low group (6.25%;1/16) (p-value < 0.0198; Figure 6A). Six out of seven cases with the non-common histology (3 cases of LH and 3 cases of SCV) were assigned to the ALK-low group by both expression and methylation analysis. Of clinical relevance, the group of patients without CN alterations trended toward an inferior progression-free survival (P=0.098, Figure 6B), which was previously observed in a cohort of systemic ALK+ALCL not strictly pediatric 26. However, group assignment in ALK-low or ALK-high based on differential expression and methylation profile did not show an association with clinical outcome (Supplemental Figure 7A-B). Also, due to the sample size, we could not investigate in detail whether specific genomic lesions could stratify patients further regarding prognosis and clinical outcome.
Figure 6. Overview of clinical findings and copy number changes associated with the two ALCL groups and impact on outcome.
(A) Heatmap with the distribution of clinical findings and copy number alterations in the diagnostic ALCL samples; cases are assigned to ALK-low and ALK-high group based on the expression and methylation profile. Five cases were classified differently based on RNA expression and DNA methylation analysis (designated as unclassifiable) (B) Kaplan-Meier curve showing progression free survival (PFS) in ALCL according to the presence of copy number changes.
Discussion
Previous studies on ALCL have focused on identifying genetic alterations in ALK negative ALCL (ALK-ALCL) and gene expression signatures that differ between ALK+ and ALK− ALCL 57, supporting their distinction into separate entities. In this study, we sought to define molecular heterogeneity in pediatric ALK+ ALCL (diagnostic and relapsed samples) using an integrated analysis including WES, RNA-sequencing, and DNA methylation array. Unsupervised cluster analysis of the expression data (RNAseq) identified two distinct subgroups of pediatric ALK+ ALCL defined by defferential ALK expression levels, the ALK-high and ALK-low groups, consistent with previous array expression data 56, 57. Through our integrative approach, we identified distinct molecular features associated with each of the two subtypes. The ALK-high group was enriched with somatic mutations and copy number changes, including genes associated with chromatin modification, cell cycle, and receptor tyrosine kinase signaling. The ALK-low group was significantly enriched in genes associated with cytokines and immune-related gene-sets. Differentially expression of methylation regulators was also notable between the two groups especially in association with differences in methylation patterns (discussed below), possibly related to DNMT3a overexpression in the ALK-low group, and UFHR1, a regulator of DNMT1, in the ALK-high group. A pattern of hypermethylation especially in CpG islands has been described in a T-ALL that was also correlated with DNMT3B 67. Overall, our findings support a molecular heterogeneity within pediatric ALK+ ALCL, while it is uniformly characterized by the transforming activity of the aberrant ALK expression. We found mutations affecting epigenetic regulators that have been described previously in different subtypes of mature T-cell lymphomas 74, 75. The functional role of genetic alterations of epigenetic regulators in ALK+ ALCLs is unknown and worth future investigation.
Mutations affecting the JAK/STAT3 pathway, observed in ALK− ALCL cases 25, were absent in our cohort of pediatric ALK+ ALCL. This corroborates the existing literature that supports the role of ALK fusion as a strong driver oncogene in ALK+ ALCLs. In our cohort of pediatric ALK+ALCL, one case showed PRDM1 alterations affecting both alleles (indel and CN loss) with concurrent loss of TP53 loci (17p). Previous studies have shown that loss of TP53 and PRDM1, more frequently observed in ALK− ALCL than in ALK+ALCL cell lines, play a role in the pathogenesis and are associated with a less favorable outcome 26. The CN alterations identified in our cohort using the WES data included gains of 1q, and losses of 6q and 17p. These findings are similar to the CNA reported by Boi et al using SNP array analysis in a cohort of ALCL (ALK+ and ALK−) patients26, while differing from the genomic imbalances described in earlier studies that used GCH arrays that included gains of 2p, 7p, 17p and 17q, and losses of 4q, 11q and 13q76. The different findings between older studies focused on copy number alterations could be most likely related to applying different technical approaches. For our copy number analysis approach using the WES data the main limitation was the lack of paired tumor-germ line DNA for a substantial number of the cases.
Arm level aneuploidy appears to be highly prevalent in ALK-high group, likely resembling aneuploidy and chromosome instability 77 but largely absent in the ALK-low group. The ALK-high group revealed up-regulated pathways involved in cell cycle and cancer proliferation, which recapitulates an accelerated tumorigenesis observed in a chromosome instability induced aneuploidy lymphoma model 78. Previous studies have also reported similar observations in adult solid tumors, associating somatic copy number change with immune evasion and reduced immunotherapy response 79. Our data suggest that this observation is not unique to solid tumors and may be relevant for lymphoma pathogenesis.
We further demonstrated that the molecular heterogeneity within the pediatric ALK+ALCL is supported by differential methylation patterns between the two groups identified by the expression cluster analaysis. ALK expression was associated with methylation of regions/probes harboring repressive histone marks as indicated by the enrichment of chromatin regulators of the PRC2 complex (EZH2, SUZ12, and JARID2), which has been described as “epigenetic switching” in cancer 80. Negative correlation between ALK expression and methylation was observed for the chromatin regulators, BRD4 and TCF4, suggesting a role in the regulation of MYC expression that was found to be upregulated in the ALK-high group. These findings provide a rationale for the evaluation of novel targeted therapies such as MYC/bromodomain inhibitors 81 and PRC2 inhibitors in ALK+ALCL 82. Additionally, this observation suggests that ALK-expression levels may be associated with DNA methylation and gene expression levels.
In our series, there was no statistically significant difference in outcome between the ALK-high and ALK-low groups by differential expression and DNA methylation. However, a tendency for worse outcomes was observed in cases without copy number changes that predominantly included ALK-low samples.
A transcriptional relapse signature was derived from four samples in our cohort that predicted relapse in the diagnosis samples. By comparing our expression profile to Camille et al. 73, the down-regulated relapse signature was found consistent between the two datasets but inconsistent in the up-regulated relapse signature. The discrepancy might be due to the relatively limited number of evaluated cases in our cohort, the high frequency (54%) of variant histology in the study by Camille et al. versus 15% in this study, and the heterogeneous nature of different therapies. Confirmation of these findings requires validation on an independent cohort of homogeneously treated pediatric ALK+ ALCL patients.
Our comprehensive, integragrative analysis of WES, RNA-seq, and DNA methylation data provide important features and further insights into the molecular heterogeneity in a strictly pediatric cohort of ALK+ALCL. In addition to supporting the previous observations of two distinct groups in ALK+ALCL based on the expression profile and ALK expression level (ALK-low and ALK-high), we have incorporated genomic (copy number alterations) and DNA methylation data to further characterize the molecular heterogeneity in this entity. Our observation of differentially expressed patterns between the two groups highlighting the role of immune response and proliferation markers, along with differences in the overall methylation profile, may help to focus on further investigation of novel molecular mechanisms that contribute to lymphogenesis, progression and as targeted therapies for children with newly-diagnosed and/or relapsed ALK+ALCL.
Acknowledgments
The authors thank the Children’s Oncology Group and the Children’s Cancer and Leukaemia Group (CCLG) Tissue Bank for access to specimens, and the Genomic Sequencing Facility of the Hartwell Center for Bioinformatics and Biotechnology of St. Jude Children’s Research Hospital. The CCLG Tissue Bank is funded by Cancer Research UK and CCLG. This work has been supported in part by the Biostatistics and Bioinformatics Shared Resource at the Moffitt Cancer Center (NCI P30 CA076292), the Moffitt Cancer Center Department of Biostatistics and Bioinformatics Pilot Project (T.I.S.) and Florida Department of Health Live Like Bella Pediatric Research Initiative (T.I.S.).
Footnotes
Conflict of Interest
T.I. Shaw reports a patent for EBD CAR pending. C.G. Mullighan reports personal fees from Illumina during the conduct of the study, as well as grants from Pfizer and AbbVie, and other support from Amgen outside the submitted work. No disclosures were reported by the other authors.
References
- 1.Cairo MS, Pinkerton R. Childhood, adolescent and young adult non-Hodgkin lymphoma: state of the science. Br J Haematol 2016. May 2. [DOI] [PubMed] [Google Scholar]
- 2.Stein H, Mason DY, Gerdes J, O’Connor N, Wainscoat J, Pallesen G, et al. The expression of the Hodgkin’s disease associated antigen Ki-1 in reactive and neoplastic lymphoid tissue: evidence that Reed-Sternberg cells and histiocytic malignancies are derived from activated lymphoid cells. Blood 1985. Oct; 66(4): 848–858. [PubMed] [Google Scholar]
- 3.Stein H, Foss HD, Durkop H, Marafioti T, Delsol G, Pulford K, et al. CD30(+) anaplastic large cell lymphoma: a review of its histopathologic, genetic, and clinical features. Blood 2000. Dec 1; 96(12): 3681–3695. [PubMed] [Google Scholar]
- 4.Jaffe ES. The 2008. WHO classification of lymphomas: implications for clinical practice and translational research. Hematology Am Soc Hematol Educ Program 2009: 523–531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jiang M, Bennani NN, Feldman AL. Lymphoma classification update: T-cell lymphomas, Hodgkin lymphomas, and histiocytic/dendritic cell neoplasms. Expert Rev Hematol 2017. Mar; 10(3): 239–249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Morris SW, Kirstein MN, Valentine MB, Dittmer KG, Shapiro DN, Saltman DL, et al. Fusion of a kinase gene, ALK, to a nucleolar protein gene, NPM, in non-Hodgkin’s lymphoma. Science 1994. Mar 4; 263(5151): 1281–1284. [DOI] [PubMed] [Google Scholar]
- 7.Turner SD, Tooze R, Maclennan K, Alexander DR. Vav-promoter regulated oncogenic fusion protein NPM-ALK in transgenic mice causes B-cell lymphomas with hyperactive Jun kinase. Oncogene 2003. Oct 30; 22(49): 7750–7761. [DOI] [PubMed] [Google Scholar]
- 8.Bischof D, Pulford K, Mason DY, Morris SW. Role of the nucleophosmin (NPM) portion of the non-Hodgkin’s lymphoma-associated NPM-anaplastic lymphoma kinase fusion protein in oncogenesis. Mol Cell Biol 1997. Apr; 17(4): 2312–2325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Chiarle R, Gong JZ, Guasparri I, Pesci A, Cai J, Liu J, et al. NPM-ALK transgenic mice spontaneously develop T-cell lymphomas and plasma cell tumors. Blood 2003. Mar 1; 101(5): 1919–1927. [DOI] [PubMed] [Google Scholar]
- 10.Brugieres L, Pacquement H, Le Deley MC, Leverger G, Lutz P, Paillard C, et al. Single-drug vinblastine as salvage treatment for refractory or relapsed anaplastic large-cell lymphoma: a report from the French Society of Pediatric Oncology. J Clin Oncol 2009. Oct 20; 27(30): 5056–5061. [DOI] [PubMed] [Google Scholar]
- 11.Noguchi K, Ikawa Y. Strategy for Pediatric Patients with Relapsed or Refractory Anaplastic Lymphoma Kinase-Positive Anaplastic Large Cell Lymphoma: A Review. Cancers (Basel) 2023. Dec 7; 15(24). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wrobel G, Mauguen A, Rosolen A, Reiter A, Williams D, Horibe K, et al. Safety assessment of intensive induction therapy in childhood anaplastic large cell lymphoma: report of the ALCL99 randomised trial. Pediatr Blood Cancer 2011. Jul 1; 56(7): 1071–1077. [DOI] [PubMed] [Google Scholar]
- 13.Seidemann K, Tiemann M, Schrappe M, Yakisan E, Simonitsch I, Janka-Schaub G, et al. Short-pulse B-non-Hodgkin lymphoma-type chemotherapy is efficacious treatment for pediatric anaplastic large cell lymphoma: a report of the Berlin-Frankfurt-Munster Group Trial NHL-BFM 90. Blood 2001. Jun 15; 97(12): 3699–3706. [DOI] [PubMed] [Google Scholar]
- 14.Laver JH, Kraveka JM, Hutchison RE, Chang M, Kepner J, Schwenn M, et al. Advanced-stage large-cell lymphoma in children and adolescents: results of a randomized trial incorporating intermediate-dose methotrexate and high-dose cytarabine in the maintenance phase of the APO regimen: a Pediatric Oncology Group phase III trial. J Clin Oncol 2005. Jan 20; 23(3): 541–547. [DOI] [PubMed] [Google Scholar]
- 15.Le Deley MC, Rosolen A, Williams DM, Horibe K, Wrobel G, Attarbaschi A, et al. Vinblastine in children and adolescents with high-risk anaplastic large-cell lymphoma: results of the randomized ALCL99-vinblastine trial. J Clin Oncol 2010. Sep 1; 28(25): 3987–3993. [DOI] [PubMed] [Google Scholar]
- 16.Lowe EJ, Sposto R, Perkins SL, Gross TG, Finlay J, Zwick D, et al. Intensive chemotherapy for systemic anaplastic large cell lymphoma in children and adolescents: final results of Children’s Cancer Group Study 5941. Pediatric blood & cancer 2009. Mar; 52(3): 335–339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Pillon M, Gregucci F, Lombardi A, Santoro N, Piglione M, Sala A, et al. Results of AIEOP LNH-97 protocol for the treatment of anaplastic large cell lymphoma of childhood. Pediatr Blood Cancer 2012. Nov; 59(5): 828–833. [DOI] [PubMed] [Google Scholar]
- 18.Williams DM, Hobson R, Imeson J, Gerrard M, McCarthy K, Pinkerton CR, et al. Anaplastic large cell lymphoma in childhood: analysis of 72 patients treated on The United Kingdom Children’s Cancer Study Group chemotherapy regimens. Br J Haematol 2002. Jun; 117(4): 812–820. [DOI] [PubMed] [Google Scholar]
- 19.Alexander S, Kraveka JM, Weitzman S, Lowe E, Smith L, Lynch JC, et al. Advanced stage anaplastic large cell lymphoma in children and adolescents: Results of ANHL0131, a randomized phase III trial of APO versus a modified regimen with vinblastine: A report from the children’s oncology group. Pediatr Blood Cancer 2014. Aug 23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Damm-Welk C, Mussolin L, Zimmermann M, Pillon M, Klapper W, Oschlies I, et al. Early assessment of minimal residual disease identifies patients at very high relapse risk in NPM-ALK-positive anaplastic large-cell lymphoma. Blood 2014. Jan 16; 123(3): 334–337. [DOI] [PubMed] [Google Scholar]
- 21.Mussolin L, Damm-Welk C, Pillon M, Zimmermann M, Franceschetto G, Pulford K, et al. Use of minimal disseminated disease and immunity to NPM-ALK antigen to stratify ALK-positive ALCL patients with different prognosis. Leukemia 2013. Feb; 27(2): 416–422. [DOI] [PubMed] [Google Scholar]
- 22.Damm-Welk C, Lovisa F, Contarini G, Ludersen J, Carraro E, Knorr F, et al. Quantification of Minimal Disease by Digital PCR in ALK-Positive Anaplastic Large Cell Lymphoma: A Step towards Risk Stratification in International Trials? Cancers (Basel) 2022. Mar 27; 14(7). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Elenitoba-Johnson KSJ, Lim MS. New Insights into Lymphoma Pathogenesis. Annu Rev Pathol 2018. Jan 24; 13: 193–217. [DOI] [PubMed] [Google Scholar]
- 24.Sandell RF, Boddicker RL, Feldman AL. Genetic Landscape and Classification of Peripheral T Cell Lymphomas. Curr Oncol Rep 2017. Apr; 19(4): 28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Crescenzo R, Abate F, Lasorsa E, Tabbo F, Gaudiano M, Chiesa N, et al. Convergent mutations and kinase fusions lead to oncogenic STAT3 activation in anaplastic large cell lymphoma. Cancer Cell 2015. Apr 13; 27(4): 516–532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Boi M, Rinaldi A, Kwee I, Bonetti P, Todaro M, Tabbo F, et al. PRDM1/BLIMP1 is commonly inactivated in anaplastic large T-cell lymphoma. Blood 2013. Oct 10; 122(15): 2683–2693. [DOI] [PubMed] [Google Scholar]
- 27.Hassler MR, Pulverer W, Lakshminarasimhan R, Redl E, Hacker J, Garland GD, et al. Insights into the Pathogenesis of Anaplastic Large-Cell Lymphoma through Genome-wide DNA Methylation Profiling. Cell Rep 2016. Oct 4; 17(2): 596–608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.swerdlow sh ce, harris nl. WHO Classification of haematopoietic and lymphoid tussues. iarc; 2008. [Google Scholar]
- 29.Roberts KG, Li Y, Payne-Turner D, Harvey RC, Yang YL, Pei D, et al. Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia. N Engl J Med 2014. Sep 11; 371(11): 1005–1015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Zhang J, Wheeler DA, Yakub I, Wei S, Sood R, Rowe W, et al. SNPdetector: a software tool for sensitive and accurate SNP detection. PLoS Comput Biol 2005. Oct; 1(5): e53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Edmonson MN, Zhang J, Yan C, Finney RP, Meerzaman DM, Buetow KH. Bambino: a variant detector and alignment viewer for next-generation sequencing data in the SAM/BAM format. Bioinformatics 2011. Mar 15; 27(6): 865–866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Iacobucci I, Wen J, Meggendorfer M, Choi JK, Shi L, Pounds SB, et al. Genomic subtyping and therapeutic targeting of acute erythroleukemia. Nat Genet 2019. Apr; 51(4): 694–704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Downing JR, Wilson RK, Zhang J, Mardis ER, Pui CH, Ding L, et al. The pediatric cancer genome project. Nat Genet 2012; 44(6): 619–622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Forbes SA, Bhamra G, Bamford S, Dawson E, Kok C, Clements J, et al. The Catalogue of Somatic Mutations in Cancer (COSMIC). Curr Protoc Hum Genet 2008. Apr; Chapter 10: Unit 10 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, et al. Integrative genomics viewer. Nat Biotechnol 2011. Jan; 29(1): 24–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Talevich E, Shain AH, Botton T, Bastian BC. CNVkit: Genome-Wide Copy Number Detection and Visualization from Targeted DNA Sequencing. PLoS Comput Biol 2016. Apr; 12(4): e1004873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 2010. Mar 1; 26(5): 589–595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Tian L, Li Y, Edmonson MN, Zhou X, Newman S, McLeod C, et al. CICERO: a versatile method for detecting complex and diverse driver fusions using cancer RNA sequencing data. Genome Biol 2020. May 28; 21(1): 126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Thorvaldsdottir H, Robinson JT, Mesirov JP. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 2013. Mar; 14(2): 178–192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Anders S, Pyl PT, Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics 2015. Jan 15; 31(2): 166–169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 2005. Oct 25; 102(43): 15545–15550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Aran D, Hu Z, Butte AJ. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol 2017. Nov 15; 18(1): 220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Du P, Zhang X, Huang CC, Jafari N, Kibbe WA, Hou L, et al. Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinformatics 2010. Nov 30; 11: 587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 2015. Apr 20; 43(7): e47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Palacios G, Shaw TI, Li Y, Singh RK, Valentine M, Sandlund JT, et al. Novel ALK fusion in anaplastic large cell lymphoma involving EEF1G, a subunit of the eukaryotic elongation factor-1 complex. Leukemia 2017. Mar; 31(3): 743–747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Zhou X, Wang J, Patel J, Valentine M, Shao Y, Newman S, et al. Exploration of Coding and Non-coding Variants in Cancer Using GenomePaint. Cancer Cell 2021. Jan 11; 39(1): 83–95 e84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.McLeod C, Gout AM, Zhou X, Thrasher A, Rahbarinia D, Brady SW, et al. St. Jude Cloud: A Pediatric Cancer Genomic Data-Sharing Ecosystem. Cancer Discov 2021. May; 11(5): 1082–1099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Roy B, Haupt LM, Griffiths LR. Review: Alternative Splicing (AS) of Genes As An Approach for Generating Protein Complexity. Curr Genomics 2013. May; 14(3): 182–194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zhu L, Xie S, Yang C, Hua N, Wu Y, Wang L, et al. Current Progress in Investigating Mature T- and NK-Cell Lymphoma Gene Aberrations by Next-Generation Sequencing (NGS). Cancer Manag Res 2021; 13: 5275–5286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Luchtel RA, Zimmermann MT, Hu G, Dasari S, Jiang M, Oishi N, et al. Recurrent MSC (E116K) mutations in ALK-negative anaplastic large cell lymphoma. Blood 2019. Jun 27; 133(26): 2776–2789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Atsaves V, Lekakis L, Drakos E, Leventaki V, Ghaderi M, Baltatzis GE, et al. The oncogenic JUNB/CD30 axis contributes to cell cycle deregulation in ALK+ anaplastic large cell lymphoma. Br J Haematol 2014. Nov; 167(4): 514–523. [DOI] [PubMed] [Google Scholar]
- 52.Atsaves V, Zhang R, Ruder D, Pan Y, Leventaki V, Rassidakis GZ, et al. Constitutive control of AKT1 gene expression by JUNB/CJUN in ALK+ anaplastic large-cell lymphoma: a novel crosstalk mechanism. Leukemia 2015. Nov; 29(11): 2162–2172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Schiefer AI, Vesely P, Hassler MR, Egger G, Kenner L. The role of AP-1 and epigenetics in ALCL. Front Biosci (Schol Ed) 2015. Jun 1; 7: 226–235. [DOI] [PubMed] [Google Scholar]
- 54.Rassidakis GZ, Thomaides A, Atwell C, Ford R, Jones D, Claret FX, et al. JunB expression is a common feature of CD30+ lymphomas and lymphomatoid papulosis. Mod Pathol 2005. Oct; 18(10): 1365–1370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Drakos E, Atsaves V, Schlette E, Li J, Papanastasi I, Rassidakis GZ, et al. The therapeutic potential of p53 reactivation by nutlin-3a in ALK+ anaplastic large cell lymphoma with wild-type or mutated p53. Leukemia 2009. Dec; 23(12): 2290–2299. [DOI] [PubMed] [Google Scholar]
- 56.Pomari E, Basso G, Bresolin S, Pillon M, Carraro E, d’Amore ES, et al. NPM-ALK expression levels identify two distinct subtypes of paediatric anaplastic large cell lymphoma. Leukemia 2017. Feb; 31(2): 498–501. [DOI] [PubMed] [Google Scholar]
- 57.Lamant L, de Reynies A, Duplantier MM, Rickman DS, Sabourdy F, Giuriato S, et al. Gene-expression profiling of systemic anaplastic large-cell lymphoma reveals differences based on ALK status and two distinct morphologic ALK+ subtypes. Blood 2007. Mar 1; 109(5): 2156–2164. [DOI] [PubMed] [Google Scholar]
- 58.Hahm JY, Park JW, Kang JY, Park J, Kim CH, Kim JY, et al. Acetylation of UHRF1 Regulates Hemi-methylated DNA Binding and Maintenance of Genome-wide DNA Methylation. Cell Rep 2020. Jul 28; 32(4): 107958. [DOI] [PubMed] [Google Scholar]
- 59.Du J, Johnson LM, Jacobsen SE, Patel DJ. DNA methylation pathways and their crosstalk with histone methylation. Nat Rev Mol Cell Biol 2015. Sep; 16(9): 519–532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Zhang JP, Song Z, Wang HB, Lang L, Yang YZ, Xiao W, et al. A novel model of controlling PD-L1 expression in ALK(+) anaplastic large cell lymphoma revealed by CRISPR screening. Blood 2019. Jul 11; 134(2): 171–185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Comerford I, Harata-Lee Y, Bunting MD, Gregor C, Kara EE, McColl SR. A myriad of functions and complex regulation of the CCR7/CCL19/CCL21 chemokine axis in the adaptive immune system. Cytokine Growth Factor Rev 2013. Jun; 24(3): 269–283. [DOI] [PubMed] [Google Scholar]
- 62.Mathy NL, Scheuer W, Lanzendorfer M, Honold K, Ambrosius D, Norley S, et al. Interleukin-16 stimulates the expression and production of pro-inflammatory cytokines by human monocytes. Immunology 2000. May; 100(1): 63–69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Liang HC, Costanza M, Prutsch N, Zimmerman MW, Gurnhofer E, Montes-Mojarro IA, et al. Super-enhancer-based identification of a BATF3/IL-2R-module reveals vulnerabilities in anaplastic large cell lymphoma. Nat Commun 2021. Sep 22; 12(1): 5577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Broniscer A, Chamdine O, Hwang S, Lin T, Pounds S, Onar-Thomas A, et al. Gliomatosis cerebri in children shares molecular characteristics with other pediatric gliomas. Acta Neuropathol 2016. Feb; 131(2): 299–307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Capper D, Jones DTW, Sill M, Hovestadt V, Schrimpf D, Sturm D, et al. DNA methylation-based classification of central nervous system tumours. Nature 2018. Mar 22; 555(7697): 469–474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Sturm D, Orr BA, Toprak UH, Hovestadt V, Jones DTW, Capper D, et al. New Brain Tumor Entities Emerge from Molecular Classification of CNS-PNETs. Cell 2016. Feb 25; 164(5): 1060–1072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Hetzel S, Mattei AL, Kretzmer H, Qu C, Chen X, Fan Y, et al. Acute lymphoblastic leukemia displays a distinct highly methylated genome. Nat Cancer 2022. Jun; 3(6): 768–782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res 2016. Jul 8; 44(W1): W90–97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Spriano F, Stathis A, Bertoni F. Targeting BET bromodomain proteins in cancer: The example of lymphomas. Pharmacol Ther 2020. Nov; 215: 107631. [DOI] [PubMed] [Google Scholar]
- 70.Weilemann A, Grau M, Erdmann T, Merkel O, Sobhiafshar U, Anagnostopoulos I, et al. Essential role of IRF4 and MYC signaling for survival of anaplastic large cell lymphoma. Blood 2015. Jan 1; 125(1): 124–132. [DOI] [PubMed] [Google Scholar]
- 71.Abate F, Todaro M, van der Krogt JA, Boi M, Landra I, Machiorlatti R, et al. A novel patient-derived tumorgraft model with TRAF1-ALK anaplastic large-cell lymphoma translocation. Leukemia 2015. Jun; 29(6): 1390–1401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Lyapichev KA, Tang G, Li S, You MJ, Cheng TJ, Miranda RN, et al. MYC expression is associated with older age, common morphology, increased MYC copy number, and poorer prognosis in patients with ALK+ anaplastic large cell lymphoma. Hum Pathol 2021. Feb; 108: 22–31. [DOI] [PubMed] [Google Scholar]
- 73.Camille D, Chloe B, Sebastien D, Veronique AL, Therese C, Pyronnet S, et al. Gene Expression Signature Associated with Clinical Outcome in ALK-Positive Anaplastic Large Cell Lymphoma. Cancers (Basel) 2021. Nov 3; 13(21). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Zhang P, Zhang M. Epigenetic alterations and advancement of treatment in peripheral T-cell lymphoma. Clin Epigenetics 2020. Nov 7; 12(1): 169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Watatani Y, Sato Y, Miyoshi H, Sakamoto K, Nishida K, Gion Y, et al. Molecular heterogeneity in peripheral T-cell lymphoma, not otherwise specified revealed by comprehensive genetic profiling. Leukemia 2019. Dec; 33(12): 2867–2883. [DOI] [PubMed] [Google Scholar]
- 76.Salaverria I, Bea S, Lopez-Guillermo A, Lespinet V, Pinyol M, Burkhardt B, et al. Genomic profiling reveals different genetic aberrations in systemic ALK-positive and ALK-negative anaplastic large cell lymphomas. Br J Haematol 2008. Mar; 140(5): 516–526. [DOI] [PubMed] [Google Scholar]
- 77.Holland AJ, Cleveland DW. Boveri revisited: chromosomal instability, aneuploidy and tumorigenesis. Nat Rev Mol Cell Biol 2009. Jul; 10(7): 478–487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Foijer F, Xie SZ, Simon JE, Bakker PL, Conte N, Davis SH, et al. Chromosome instability induced by Mps1 and p53 mutation generates aggressive lymphomas exhibiting aneuploidy-induced stress. Proc Natl Acad Sci U S A 2014. Sep 16; 111(37): 13427–13432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Davoli T, Uno H, Wooten EC, Elledge SJ. Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science 2017. Jan 20; 355(6322). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Shen H, Laird PW. Interplay between the cancer genome and epigenome. Cell 2013. Mar 28; 153(1): 38–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Boi M, Todaro M, Vurchio V, Yang SN, Moon J, Kwee I, et al. Therapeutic efficacy of the bromodomain inhibitor OTX015/MK-8628 in ALK-positive anaplastic large cell lymphoma: an alternative modality to overcome resistant phenotypes. Oncotarget 2016. Nov 29; 7(48): 79637–79653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Li B, Chng WJ. EZH2 abnormalities in lymphoid malignancies: underlying mechanisms and therapeutic implications. J Hematol Oncol 2019. Nov 21; 12(1): 118. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Genomic data have been deposited in the European Genome-phenome Archive (EGA), which is hosted by the European Bioinformatics Institute (EBI), under accession EGAS00001004189. The accession number for the Illumina methylation reported is GEO: GSE186487. All other remaining data are available within the Article and Supplementary Files.