Abstract
BZLF1 is a member of the extended AP-1 family of transcription factors which binds to specific BZLF1 sequence motifs within early Epstein-Barr virus (EBV) promoters and to closely related AP-1 motifs. BZLF1’s activity is regulated at the transcriptional level as well as through protein interactions and posttranslational modifications. Phorbol esters or immunoglobulin cross-linking both reactivate EBV from latently infected B cells via transactivation of BZLF1. We report here that the phorbol ester 12-O-tetradecanoylphorbol-13-acetate (TPA) is capable of inducing BZLF1’s activity even further. The induction occurs at the posttranscriptional level and depends on a single serine residue located in the DNA binding domain of BZLF1. This serine residue (S186) is phosphorylated by protein kinase C in vitro and in vivo after stimulation with TPA. Phosphorylation of S186 per se interferes with the DNA binding affinity of BZLF1 in vitro but is mandatory for TPA-induced increase in DNA binding of BZLF1, as shown in gel retardation assays and reconstruction experiments with cellular extracts. In transcriptional reporter assays, S186 is essential for the activation of BZLF1 by TPA. Presumably, a yet-to-be-identified cellular factor restores the DNA binding affinity and enhances the transcriptional activity of S186-phosphorylated BZLF1, which is required to induce the lytic phase of EBV’s life cycle.
A critical aspect in the life cycle of all herpesviruses is the decision between maintenance and disruption of viral latency. Epstein-Barr virus (EBV), for example, is a human herpesvirus which infects B cells in vivo and in vitro to establish a latent infection (for a review, see reference 38). In these latently infected cells, the induction of the lytic phase of EBV’s life cycle can be achieved by compounds such as 12-O-tetradecanoylphorbol-13-acetate (TPA) or cross-linking of surface immunoglobulin (62, 69). These agents activate expression of the viral immediate-early protein BZLF1, also termed EB1, Zta, Z, or ZEBRA (10, 38 [and references therein], 45, 61, 63). Conversly, BZLF1 is repressed in EBV’s latent phase to prevent the onset of the lytic phase and the eventual death of the host cell.
BZLF1 is a sequence-specific DNA binding protein related to the bZip family of transcription factors which transactivates several early, lytic-phase viral promoters via cis-acting ZRE (for Zta responsive element) DNA binding motifs (66). BZLF1 contains a carboxy-terminal domain (see Fig. 1A, upper part) that mediates homodimerization through a coiled-coil interaction (9, 42) and a basic region which shares sequence homology with the DNA binding domain of members of the AP-1 family of transcription factors (18, 66). Hence, BZLF1 binds to TRE (for TPA responsive element) or AP-1 sequence motifs with high affinity (18, 46). The amino-terminal domain of BZLF1 plays a role in transcriptional activation (11, 12, 22) and is also directly involved in the activation of the EBV lytic origin of DNA replication, oriLyt (2, 56–58).
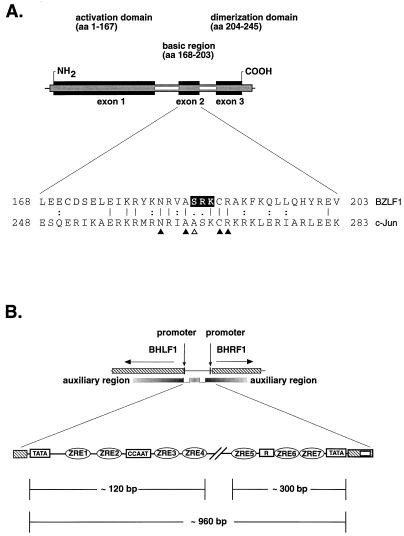
FIG. 1.
Trans- and cis-acting elements of the lytic cycle of EBV. (A) (Upper part) The schematic modular structure of the functional domains in BZLF1 is similar to that of other members of the AP-1 family. BZLF1 consists of a transcriptional activation domain involved in DNA replication as well as in transcriptional activation, a basic DNA binding domain, and a dimerization domain which is responsible for homodimerization of BZLF1. (Lower part) Alignment of the amino acid sequences of the basic DNA binding domains of human c-Jun and BZLF1. Identical amino acids are indicated by vertical lines; double and single points mark similar and less similar residues, respectively. Arrowheads indicate amino acids in c-Jun that were shown to contact DNA bases (26). The serine at aa 186 in BZLF1 differs from the highly conserved alanine (open arrowhead) found at the equivalent position in the majority of bZip proteins. The amino acids serine 186, arginine 187, and lysine 188 in BZLF1, which form a conserved PKC motif, are highlighted. (B) Fine structure of oriLyt, with its cis-acting elements. oriLyt is located in a divergently transcribed promoter region which is shown schematically at the top. The flanking genes BHLF1 and BHRF1 are illustrated on the left and right (hatched boxes) together with their promoters (black bars). The open rectangles delineate the essential sequence elements which encompass the minimal oriLyt (59). The shaded regions represent poorly defined regions which function as auxiliary elements of oriLyt. An enlarged view of oriLyt is given at the bottom of the figure, including known functional sequence elements which are indicated by icons representing seven binding sites for BZLF1 (ZRE1 to ZRE7), one cluster of binding sites for the viral enhancer factor R, and the TATA and CCAAT boxes.
Suppression of BZLF1 is critical in order to sustain the latent phase of EBV’s life cycle, but specific stimuli relieve suppression under certain conditions to allow the transition to EBV’s lytic phase. Therefore, BZLF1 expression is not detectable in latently infected B cells, but the BZLF1 promoter region contains several TPA-responsive binding motifs for transcription factors like ATFα, ATF1, ATF2, and c-Jun (67), as well as ZRE sites which trigger BZLF1’s expression via a positive feedback mechanism (14, 20). As a consequence, stimuli which are capable of inducing AP-1 activity are potent inducers of BZLF1 expression (6). Besides transcriptional control, BZLF1’s activity is regulated by protein-protein interactions (24, 32, 36, 37, 60, 68) and phosphorylation (13, 41, 55).
Phosphorylation is a fundamental regulatory modification of cellular and viral transcription factors, altering their affinity for DNA binding or transcriptional activities (for a review, see reference 30). For instance, the regulation of c-Jun, the paradigmatic member of the AP-1 family, has been found to be mediated by two distinct phosphorylation events. Phosphorylation of two serine residues located in the N-terminal transcription activation domain by Jun N-terminal kinase (JNK) in response to various stimuli, such as TPA, UV radiation, or intracellular signal transducers, leads to enhanced transcriptional activity of c-Jun (5, 16, 53). On the other hand, phosphorylation of c-Jun at sites next to its DNA binding domain decreases DNA binding affinity, while dephosphorylation of one of these sites results in increased DNA binding (7). These phosphorylations regulate each other, since phosphorylation of the c-Jun N-terminal transactivation domain by JNK/stress-activated protein kinase seems to alter the accessibility of the carboxy-terminal sites to phosphatases and kinases (50). Similarly, BZLF1 is phosphorylated close to its basic DNA binding domain by casein kinase II in vitro, which affects the affinity of BZLF1 binding (41). BZLF1 is also found phosphorylated in vivo, but the relevance of this finding is not clear (13). In addition, a consensus phosphorylation motif for protein kinase C (PKC) has been proposed within the DNA binding domain of BZLF1 at serine amino acid residue 186, which was found to be critical for the full biological activity of BZLF1 (23).
Besides regulation of DNA binding, phosphorylation can affect complex formation of proteins to direct their activation potential. The interaction of the cyclic AMP responsive factor CREB with the coactivator CREB-binding protein (CBP), for example, requires the phosphorylation of CREB at a serine residue located in the CBP-binding domain (51). This recruitment is critical for targeted gene induction in response to cyclic AMP. Therefore, like the src homology 2 domains which bind phosphotyrosine-containing peptides, signal transduction through serine/threonine kinase pathways may also require protein interaction motifs which are capable of recognizing phosphorylated amino acids.
PKC is a serine/threonine protein kinase which is directly activated by phorbol esters like TPA (for a review, see reference 49). PKC isozymes are crucially implicated in signal transduction cascades following a variety of different stimuli altering cellular functions and proliferation. In vitro, a large number of proteins have been shown to be phosphorylated by the different isoforms of PKC. Only a few proteins have been demonstrated to be physiological substrates for PKC by in vivo studies (for a review, see reference 27), e.g., MARCKS (for myristoylated alanine-rich C kinase substrate) (for a review, see reference 1), cytoskeletal protein receptors (31), and intracellular transducers such as Raf-1 (39) and p53 (64).
Being interested in the functional characterization of BZLF1, we noticed that its transcriptional activity can be augmented by TPA. We therefore addressed the hypothesis of whether BZLF1 could be a substrate for a TPA-activated kinase and whether this putative relationship has any functional significance. In this paper, we provide genetic, biochemical, and biological evidence that the complete activation of BZLF1 is dependent on phosphorylation of S186, which might be influential for the recruitment of a cooperating cellular factor.
MATERIALS AND METHODS
Plasmids for protein expression in Escherichia coli.
All proteins were expressed in E. coli DH5α (29) as glutathione S-transferase (GST) fusions in pGEX-1λT (Pharmacia). The BZLF1-encoding cDNA sequence was derived from EBV strain B95.8 (4, 10). Full-length BZLF1 cDNA (amino acids [aa] 1 to 245) was inserted into the BamHI/EcoRI-cut pGEX-1λT plasmid to provide a thrombin cleavage site between the two protein domains. This recombinant plasmid was termed pGST:BZLF1-wt. pGST:BZLF1-T159A, pGST:BZLF1-S186A, pGST:BZLF1-T159A/S186A, and pGST:BZLF1-S186D, expressing different substitution mutants of BZLF1, were generated by site-directed mutagenesis in the pBluescript II SK(−) vector (Stratagene). The coding parts of all plasmids which were used to express BZLF1 and its mutants were subjected to sequence analysis.
Recombinant eukaryotic plasmids.
The reporter plasmids pBHRF1-Luc and pBHLF1-Luc were constructed as described previously (57, 59). p4xZRE5tk-Luc is a reporter plasmid containing four copies of the ZRE5 binding site, which was multimerized by ligation of synthetic oligonucleotides (described below). The multimeric binding site of ZRE5 was inserted upstream and in close proximity to a minimal thymidine kinase promoter from herpes simplex virus type 1 fused to the luciferase reporter gene. Plasmid pCMV-BZLF1-wt is an expression vector which efficiently induces the lytic phase of EBV’s life cycle. The BZLF1 gene is driven in this retroviral vector construct by the promoter of the immediate-early genes of the human cytomegalovirus, as described previously (28). The BZLF1 chimeric transcription factor plasmid pCMV-BZLF1(trans):E2 was cloned as reported in detail previously (57). pCMV-BZLF1(trans):E2 consists of the transcriptional activation domain of BZLF1 (aa 1 to 169) fused to the DNA binding and dimerization domain (aa 281 to 410) of the bovine papillomavirus E2 transcription and replication factor. Other recombinant or native transcription factors expressed from various vector plasmids [pCMV-c-jun(trans):E2 and pCMV-E2-TR] were generous gifts from Paul Lambert (44). Plasmid pLPV-BZLF1-wt expresses a cDNA of the BZLF1 gene from the promoter of the early genes of the lymphotropic papovavirus (52). BZLF1 substitution mutants were generated by site-directed mutagenesis, as described above, and the corresponding plasmids were designated pCMV-BZLF1-S186A, pLPV-BZLF1-S186A, pLPV-BZLF1-S186D, and pLPV-BZLF1-S186E.
Bacterial protein synthesis and purification.
E. coli DH5α strains transformed with expression plasmids for GST-BZLF1 variants were grown in 400 ml of Luria-Bertani medium to an optical density at 600 nm of 0.5 and induced for 1 h with 2 mM IPTG (isopropyl-β-d-thiogalactopyranoside) at 37°C. Cells were harvested and the bacterial pellet was resuspended in 5 ml of cold lysis buffer (1× Tris-EDTA, 1% Triton X-100, and the following protease inhibitors: 0.5 μg of leupeptin per ml, 1 μg of pepstatin per ml, 1 mM phenylmethylsulfonyl fluoride, 50 mM benzamidin, 1 mM pefabloc, and 0.5 μg of aprotinin per ml) and sonified for 10 min. After ultracentrifugation at 45,000 rpm for 1 h, the cleared extracts were bound to Sepharose beads coupled with glutathione (Pharmacia). BZLF1 was released by enzymatic cleavage with 1 U of thrombin (Sigma) for 30 min from its fusion with GST. Proteins were analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and Coomassie staining. Protein content was determined by standard procedures.
Transient transcription assays.
Human embryonic kidney 293 cells and the EBV-negative Burkitt lymphoma cell line BL41 were grown in RPMI 1640 medium supplemented with 10% fetal calf serum. BL41 cells (5 × 106 per DNA transfection) were transfected by electroporation with 5 μg of reporter plasmid (pBHRF1-Luc, pBHLF1-Luc, p7xE2-BHRF1-Luc, or p4xZRE5tk-Luc) together with 2 μg of activator plasmid [pCMV-BZLF1-wt or pCMV-BZLF1-S186A; pLPV-BZLF1-wt, pLPV-BZLF1-S186A, pLPV-BZLF1-S186D, or pLPV-BZLF1-S186E; or pCMV-c-Jun(trans):E2, pCMV-BZLF1(trans):E2, or pCMV-E2-TR]. The same amounts of plasmid DNAs were used to transfect 5 × 106 293 cells per DNA transfection with Lipofectamine (Gibco BRL Life Technologies). Eight hours after transfection, cells were treated with 100 ng of TPA per ml and/or 1 μmol of PKC inhibitor GF109203X (Biomol) per liter for 12 h. The luciferase activity was measured in cell extracts, as described in detail previously (59). All reactions were performed at least in quadruplicate.
Transient replication assay.
The cell line D98HR1 was derived from a somatic cell hybrid between the EBV genome-positive Burkitt’s lymphoma cell line P3HR1 and the human epithelial cell line D98 (25). This adherent cell line contains ∼20 copies of the EBV genome (data not shown) and was maintained in Dulbecco’s Eagle’s medium containing 5% fetal and 5% newborn calf serum. Ten micrograms of the oriLyt plasmid p526 were cotransfected with 5 μg of either pCMV-BZLF1-wt or pCMV-BZLF1-S186A by electroporation. Plasmid p526 has been described previously (28). The concomitant introduction of pCMV-BZLF1 efficiently induces the lytic cycle of EBV. Two days after cotransfection, DNA was prepared, input plasmid DNA was digested with BamHI and DpnI to eliminate methylated prokaryotic DNA, and the efficiency with which p526 replicated was quantitated according to specific signals detected after Southern blot hybridization and autoradiography by standard procedures.
Nuclear extraction.
293 cells were transfected with 2 μg of plasmid pCMV-BZLF1-wt or pCMV-BZLF1-S186A as described above or mock transfected and then treated with TPA (100 ng/ml) for various periods up to 1 h. The cells were harvested immediately and resuspended in 300 μl of hypotonic buffer A (10 mM HEPES [pH 7.9], 10 mM KCl, 0.1 mM EDTA, 1.5 mM MgCl2, 0.5 mM dithiothreitol [DTT], protease inhibitors [see above], and the phosphatase inhibitors β-glycerolphosphate, NaF, pyrophosphate, molybdate, orthovanadate, and ocadaic acid at 1 mM each) and incubated on ice for 30 min. Permeabilized nuclei were recovered by brief centrifugation, and the crude nuclear pellet was resuspended in 200 μl of hypertonic buffer B (20 mM HEPES [pH 7.9], 0.4 M NaCl, 0.1 mM EDTA, 1.5 mM MgCl2, 0.5 mM DTT, 25% glycerol, and protease and phosphatase inhibitors as described above) followed by incubation on ice for 30 min. After centrifugation, the supernatant was used for gel retardation assays and for immunoblot analysis with the anti-BZLF1 monoclonal antibody Z130 (48) (see below).
Gel retardation assays.
Gel shift experiments were performed with bacterially expressed BZLF1 as well as with nuclear extracts from cells transfected with BZLF1 expression plasmids. The different protein preparations were mixed with 32P-labeled oligonucleotides (20,000 dpm per sample) containing an AP-1 site or different ZRE sites in a 25-μl solution of 10 mM HEPES (pH 7.9), 10% (vol/vol) glycerol, 60 mM KCl, 0.1 mM EDTA, 0.25 mM DTT, and 2 μg of poly(dI-dC). The DNA probes were labeled with T4 polynucleotide kinase. Binding reactions were carried out at room temperature for 30 min in the presence or absence of the anti-BZLF1 monoclonal antibody Z125 (48) and separated on a 7.5% polyacrylamide gel (20:1 acrylamide-bisacrylamide) in 0.2× TBE (1× TBE is 90 mM Tris, 64.6 mM boric acid, and 2.5 mM EDTA [pH 8.3]). The gels were dried and exposed to Kodak XAR-5 film. The following oligonucleotides were used for gel shift experiments (binding sites are underlined): AP-1, 5′-TCGAAGCTATGACTCATCCGGTCGA-3′ (54); ZRE2/7, 5′-CTAGCTCACCTTGAGCAATTTGGTCTAGAA-3′ (9); ZRE3A, 5′-CTAGCTATGCATGAGCCACAGATC-3′ (65); and ZRE5, 5′-CTAGATGTCACCTTTGCACATTTGGTCAG-3′ (46).
Western blot analysis.
Aliquots of the various protein samples were boiled in Laemmli buffer, subjected to SDS-PAGE on 15% polyacrylamide gels, and electrotransferred onto nitrocellulose membranes. BZLF1 was detected by Western blotting with monoclonal antibodies which recognize an epitope between aa 59 and 93 (Z125) or an epitope between aa 177 and 196 (Z130) of BZLF1 (48), a horseradish peroxidase-conjugated goat anti-mouse immunoglobulin G (Promega), and ECL detection reagent (Amersham).
In vitro phosphorylation.
Purified BZLF1 was phosphorylated in a reaction mixture (15 μl) containing kinase buffer (25 mM Tris-HCl [pH 7.5], 1.32 mM CaCl2, 5 mM MgCl2, 1 mM EDTA, 1.25 mM EGTA, and 1 mM DTT) supplemented with 10 nM TPA, 5 μg of phosphatidylserine per ml, 5 mM ATP, and 0.5 μCi of [γ-32P]ATP. Reaction mixtures were incubated at room temperature with purified PKC for 15 min. Products were subsequently separated by electrophoresis in SDS–15% polyacrylamide gels and visualized by autoradiography.
In vivo labeling.
293 cells were transfected with 10 μg of pCMV-BZLF1-wt or pCMV-BZLF1-S186A, as described above. Twelve hours after transfection, 293 cell cultures (in 90-mm dishes) were rinsed twice with phosphate-buffered saline and incubated in 5 ml of phosphate-free Dulbecco’s modified Eagle’s medium (Sigma) for 3 h. [32P]orthophosphate (2.5 mCi) (Amersham) was then added. After a 4-h labeling period (the last hour in the presence or absence of TPA [100 ng/ml]), the cells were washed twice with ice-cold phosphate-buffered saline and labeled proteins were immunoprecipitated from cell lysates with the Z130 monoclonal antibody directed against BZLF1 coupled to protein G-Sepharose (Pharmacia). Bound proteins were eluted by boiling in Laemmli buffer and were analyzed by SDS-PAGE. The SDS-polyacrylamide gel was blotted onto a nitrocellulose membrane, and the labeled proteins were detected by autoradiography (data not shown). The amount of immunoprecipitated BZLF1 was evaluated by immunodetection with biotinylated Z125 (data not shown).
Mass spectrometry.
Mass spectrometry analysis was performed on an ABI Micropurification System 173 connected to a Perkin-Elmer API 100 quadrupole mass spectrometer and a Berthold radioactivity detector. BZLF1 was phosphorylated and digested as described below. Tryptic peptides from 5 μg of BZLF1 were loaded onto an ABI 100- by 0.3-mm reversed-phase (C18) column. A gradient of 0 to 35% acetonitrile in 100 min, and subsequently of 35 to 70% in 20 min, was run at 5 μl/min. Masses were determined in 0.1-atomic mass unit steps over an m/z range of 500 to 1,500, with an orifice voltage of 20 V.
Phosphopeptide mapping.
BZLF1 was radiolabeled in vivo, immunoprecipitated, and subjected to SDS-PAGE, as described above. After autoradiography, the BZLF1 band was excised from the corresponding area of the gel, washed extensively with water, and digested to completion with trypsin in 50 mM NH4HCO3 at 37°C overnight. After trypsin treatment, the released peptides were subjected to lyophilization, resuspended in water, and applied to high-performance liquid chromatography or thin-layer cellulose sheets for two-dimensional peptide mapping. Separation of the labeled phosphopeptides was performed in the horizontal dimension by electrophoresis at pH 1.9 for 20 min at 1,000 V and in the vertical dimension by ascending chromatography. The same peptide mapping procedures were utilized for recombinant BZLF1 phosphorylated in vitro by PKCα, as described above.
Induction of GFP-tagged EBV and titer determination.
A fully recombinant EBV genome has recently been established; it carries the gene for green fluorescent protein (GFP) under control of the human cytomegalovirus immediate-early promoter/enhancer. Virus production was induced by transfecting up to 1 μg of pCMV-BZLF1-wt into 293 cells which carry the recombinant EBV genome in a latent fashion. Transfected 293 cells were kept in the presence of 1 μmol of PKC inhibitor GF109203X per liter (or dimethyl sulfoxide [DMSO] only, as a negative control) for 4 days. Supernatants were harvested from these 293 cells and used to superinfect 5 × 104 Raji cells in a dose-dependent manner. Raji cells, an EBV-positive Burkitt lymphoma cell line, were grown in RPMI 1640 medium supplemented with 10% fetal calf serum. The virus titers were determined by analyzing the number of green Raji cells by UV light microscopy or by fluorescence-activated cell sorter (FACS) analysis 4 days after infection, as described in detail elsewhere (15).
RESULTS
Expression of BZLF1-responsive genes is stimulated by TPA.
The expression of BZLF1 in latently infected EBV-positive B cells activates as many early promoters as does treatment of these cells with TPA (10). TPA induction of EBV early promoters is generally explained by TPA-mediated upregulation of the BZLF1 promoter Zp via its AP-1 responsive sites (6, 14, 21, 40). Subsequently, the BZLF1 gene product transactivates its own promoter, as well as the promoters of a number of lytic genes, thereby initiating a cascade of viral gene expression (6, 14, 19). Our initial observations indicated that TPA might also be capable of modulating BZLF1’s transcriptional activity independently of the induction of the BZLF1 gene.
The upstream regions of the divergently transcribed EBV early genes BHRF1 and BHLF1 constitute BZLF1-responsive promoters which colocalize with oriLyt, the lytic origin of DNA replication of EBV (Fig. 1B) (28). In order to measure the TPA inducibility of these BZLF1-responsive promoters, we replaced the open reading frames of the BHRF1 and BHLF1 genes with luciferase to measure the relative transcriptional activities enzymatically (59). To exclude effects due to induction of endogenous BZLF1, we chose the EBV-negative Burkitt lymphoma cell line BL41 for these and most of the following experiments.
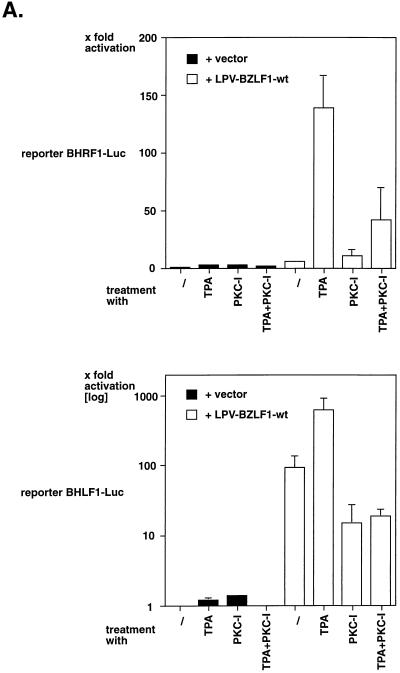
As shown in Fig. 2A, cotransfection of a BZLF1 expression plasmid (pLPV-BZLF1-wt) with the reporter plasmid BHRF1-Luc led to a 5- to 10-fold upregulation of the BHRF1 promoter, which was dramatically stimulated by the addition of TPA. TPA treatment in conjunction with the PKC inhibitor GF109203X resulted in repression of the TPA effect, whereas treatment of the cells with TPA and/or PKC inhibitor in the absence of BZLF1 did not induce the BHRF1 reporter. Very similar observations were made with the BHLF1 promoter (Fig. 2A, lower panel) as well as in different cell lines, including 293 cells (see below).
FIG. 2.
TPA induces BZLF1-responsive gene expression. (A) BL41 cells were transiently transfected with the luciferase reporter plasmid pBHRF1-Luc (upper panel) or pBHLF1-Luc (lower panel). Where indicated, the BZLF1 expression plasmid pLPV-BZLF1-wt was cotransfected. The cells were treated with either TPA, the PKC inhibitor GF109203X, or a combination of both. Relative transcriptional activation was calculated on the basis of the luciferase activity in cells transfected with the reporter plasmid and vector (left column), which was set to one. (B) Neither TPA nor PKC inhibitor affect steady-state protein levels of BZLF1. Aliquots of the cell lysates which were analyzed in parallel for luciferase activity, as shown in panel A of this figure, were analyzed for the amount of BZLF1 by Western blotting. (C) Transcriptional activation of the mutant reporter construct p7xE2-BHRF1-Luc. This plasmid is identical to the reporter plasmid pBHRF-1-Luc shown in panel A of this figure, with the exception of the seven ZRE motifs which were replaced by seven E2 binding sites. Chimeric transcription factors [pCMV-c-Jun(trans):E2 and pCMV-BZLF1(trans):E2] which consist of the transactivation domains of c-Jun and BZLF1, respectively, fused to the DNA binding domain E2 of bovine papillomavirus were cotransfected with the reporter plasmid in comparison with pCMV-BZLF1-wt (which is unable to bind to the reporter plasmid) and a version of E2 itself (pCMV-E2-TR). Cells were treated subsequently with TPA or were left untreated, as indicated. Relative transcriptional activation is based on the measured luciferase activity in cells transfected only with the reporter plasmid (left column). TPA induction in cells transfected with the reporter plasmid alone led to an about twofold increase in luciferase activity.
To confirm that treatment with TPA or PKC inhibitor or the combination of both had no effect on the levels of BZLF1 expressed from the transfected plasmid pLPV-BZLF1-wt, we analyzed the cell lysates for BZLF1 protein by Western blotting. No difference in the steady-state levels of BZLF1 was detected (Fig. 2B and data not shown).
TPA-mediated activation of c-Jun is partly due to phosphorylation of its activation domain by c-Jun N-terminal kinase (16, 35). Since BZLF1 shares some characteristics with c-Jun (18), we asked whether TPA induction of BZLF1-responsive promoters relied on a similar mechanism. For this purpose, chimeric transcription factors which consist of the DNA binding domain of the bovine papillomavirus E2 protein fused to the activation domain of c-Jun [c-Jun(trans):E2] (44) or BZLF1 [BZLF1(trans):E2] (57) were used. The TPA responsiveness of these chimeras was analyzed with a BHRF1 luciferase reporter construct in which all seven ZRE sites had been replaced by E2 binding sites (57) (Fig. 2C). Transcriptional activation of this reporter plasmid depends on transcription factors which contain the DNA binding domains of E2 (57).
As expected, the chimeric transcription factor c-Jun(trans):E2 was induced by TPA about eightfold (Fig. 2C). In contrast, TPA induction with BZLF1(trans):E2 was only about twofold. This level of induction was also observed with the reporter plasmid alone, with wild-type BZLF1 (which cannot bind to the reporter construct, which lacks all binding sites for wild-type BZLF1), or with the transcriptional repressor E2-TR.
Together, these data suggest that the induction of the BHRF1 promoter depends on (i) DNA binding of BZLF1 to cis-acting ZRE sites and (ii) BZLF1’s carboxy-terminal basic and/or dimerization domains but not on its N-terminal transactivation domain.
EBV titer is influenced by PKC levels.
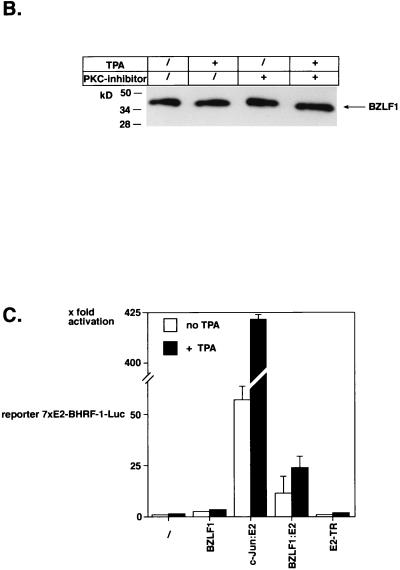
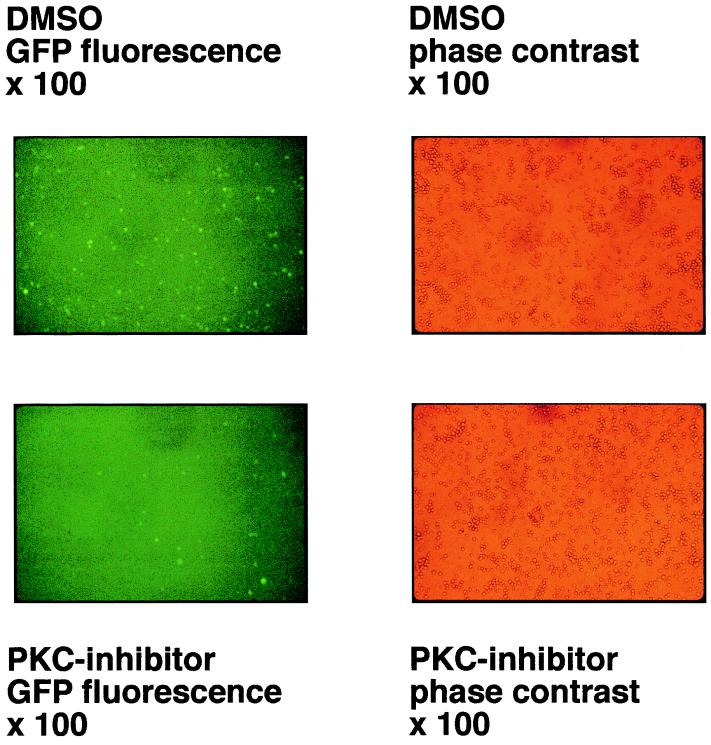
Recently, we cloned the complete 172-kbp genome of EBV strain B95.8 in E. coli onto an F-factor-derived plasmid which also includes the genes for hygromycin resistance and GFP as genetic and phenotypic markers, respectively. After transfection into 293 cells, the B95.8–F-factor DNA molecules are maintained extrachromosomally under selection with hygromycin. Virus production can be achieved by transfecting an expression vector encoding BZLF1, and virus titers can be easily assessed by superinfection of Raji cells and visualization of GFP-positive cells (15). Since growth factors present in the cell culture media provide a low constitutive level of endogenous PKC activity, we speculated that PKC inhibitors might influence the efficiency of lytic-phase induction in 293 cells carrying B95.8–F-factor genomes. The pCMV-BZLF1-wt expression plasmid was transfected into these cells by Lipofectamine, and the cells were kept either in the presence of the PKC inhibitor GF109203X or in equivalent concentrations of its solvent DMSO only, for 4 days. Cell-free supernatants were used to infect Raji cells as described in Materials and Methods, and GFP-positive cells were counted in corresponding sets or analyzed by FACS 4 days following superinfection. As exemplified in Fig. 3, EBV titers were reduced by a factor of five to six in supernatants obtained from cells treated with the PKC inhibitor compared to the negative control. The same tendency could be seen with different multiplicities of infection of Raji cells (data not shown), indicating that induction of EBV’s lytic phase and/or virus maturation depend to some extent on endogenous PKC levels within the cell which support virus production. Since the protein levels of constitutively overexpressed BZLF1 were not affected by PKC activation or inhibition (Fig. 2B), GF109203X could influence the posttranscriptional modification of BZLF1, mediated directly or indirectly by PKC.
FIG. 3.
A PKC inhibitor reduces the titer of EBV released from 293 cells. A fully recombinant EBV genome which carries the gene for GFP under control of the human cytomegalovirus immediate-early promoter/enhancer was established in a latent fashion in the epitheloid cell line 293, as described previously (15). Virus production was induced by transfecting pCMV-BZLF1-wt. Half of the transfected 293 cells were kept in the presence of the PKC inhibitor GF109203X or DMSO only, as a negative control, for 4 days. Cell-free supernatants were harvested from these 293 cells and 1/10 each of virus supernatants was used to infect Raji cells. The virus titers were determined by analyzing the number of green Raji cells by UV light microscopy, as shown in comparison to phase contrast micrographs. Supernatants which were obtained from 293 cells treated with PKC inhibitor show lower EBV titers (factor of five to six) than cells treated with DMSO only, as a negative control.
Phosphorylation of BZLF1 by PKC in vitro.
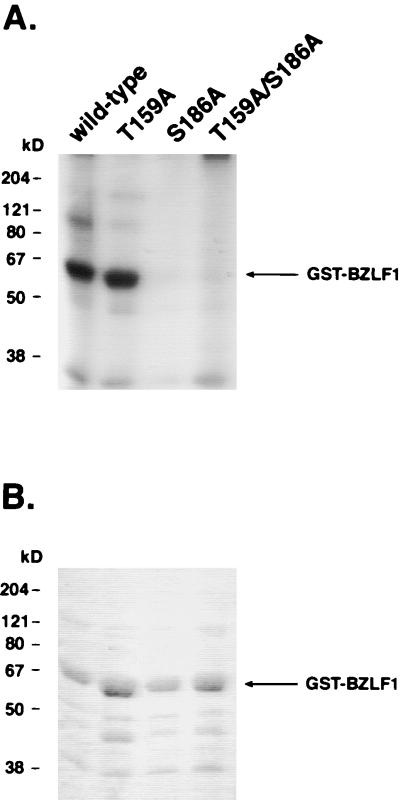
As TPA and the PKC inhibitor GF109203X directly affect the enzymatic functions of PKC, we asked whether BZLF1 could be a substrate for PKC. As shown in Fig. 4A, bacterially expressed BZLF1 protein was readily phosphorylated by PKCα in vitro. In order to map BZLF1’s phosphorylation sites, bacterially expressed BZLF1 was incubated with [γ-32P]ATP in the presence of PKCα and analyzed by mass spectrometry after trypsin cleavage. We were able to identify two possible candidate sites for PKC phosphorylation of BZLF1: a threonine residue at position 159 (T159) and a serine residue at position 186 (S186), both of which are part of a perfect PKC consensus motif (S/T-R-K) (see Fig. 1A, lower part, for the S186 motif) (3). BZLF1 mutants with single-amino-acid substitutions were generated. Both T159 and S186 were replaced by alanine (A), changing the PKC consensus motif to the sequence A-R-K. Mutation of the motif S186 (but not T159) led to a dramatic reduction of phosphorylation (Fig. 4), indicating that S186 is the main target for PKC-dependent phosphorylation of BZLF1 in vitro. PKCη, PKCδ, and PKCζ, which belong to the different categories of novel and atypical PKCs (34), each phosphorylated BZLF1 to a similar extent (data not shown).
FIG. 4.
BZLF1 is phosphorylated by PKCα at serine 186 in vitro. (A) Autoradiography of in vitro-labeled BZLF1. BZLF1-wt and point mutants BZLF1-T159A, BZLF1-S186A, and BZLF1-T159A/S186A were expressed as GST fusion proteins in E. coli and phosphorylated in vitro by PKCα. Following separation on an SDS gel, labeled proteins were analyzed by autoradiography. Only BZLF1-wt and the BZLF1 mutant T159A were efficiently phosphorylated by PKCα in vitro. (B) Coomassie stain of SDS gel from panel A of this figure. The amounts of the different GST-BZLF1 fusion proteins were comparable, as verified by Coomassie staining.
Phosphorylation of BZLF1 in vivo.
Together with the transactivation assays (Fig. 2), our data suggested that BZLF1 might be a target for phosphorylation by TPA-activated kinases, e.g., PKCs, in vivo. To address the phosphorylation status of BZLF1 in vivo, we transfected 293 cells with either pCMV-BZLF1-wt or pCMV-BZLF1-S186A. The cells were incubated with [32P]orthophosphate and treated with TPA for 1 h. BZLF1 was immunoprecipitated from the cell lysate, and precipitated proteins were analyzed by SDS-PAGE prior to autoradiography in parallel with immunodetection of BZLF1 (data not shown). Following tryptic digestion and two-dimensional gel electrophoresis, phosphopeptide maps of immunoprecipitated BZLF1-wt and BZLF1-S186A were compared with bacterially expressed and in vitro-phosphorylated BZLF1-wt (Fig. 5).
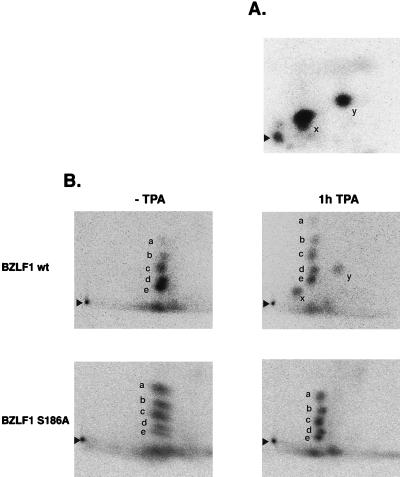
FIG. 5.
Phosphopeptide mapping of BZLF1. (A) Bacterially expressed GST-BZLF1-wt was phosphorylated by PKCα in vitro prior to SDS-PAGE. The radiolabeled GST-BZLF1 band was identified by autoradiography, excised, and subjected to tryptic digestion. The released peptides were separated by two-dimensional electrophoresis and visualized by autoradiography. Two signals (x and y), which correspond to two partially digested peptides which carry S186, were detected. BZLF1-S186A failed to be phosphorylated by PKCα in vitro (Fig. 4) and consequently did not produce peptide spots (data not shown). (B) Two-dimensional tryptic phosphopeptide mapping of BZLF1-wt and BZLF1-S186A proteins from transiently transfected 293 cells which were metabolically labeled with [32P]orthophosphate in vivo. The cells were left untreated or were incubated with TPA for 1 h prior to harvest. BZLF1 was immunoprecipitated and run on SDS-PAGE. The band was excised and subjected to tryptic digestion and two-dimensional phosphopeptide chromatography. BZLF1 is constitutively phosphorylated, as indicated by five spots labeled a through e which can be seen in all four panels. In addition, after TPA stimulation, BZLF1-wt but not BZLF1-S186A revealed two additional signals, labeled x and y. These signals correspond to the same spots seen with in vitro PKC-phosphorylated BZLF1, as shown in panel A of this figure. Arrowheads indicate the positions where the peptide samples were applied.
In vitro, two spots (x and y [Fig. 5A]) were detected, although the tryptic digestion of BZLF1 should theoretically yield only one peptide from aa 184 to 187. The detection of two spots is most likely due to partial trypsin cleavage (8), since trypsin usually cleaves C-terminally of an arginine residue or a lysine residue and S186 is followed immediately by both amino acids (Fig. 1A). As mentioned above, BZLF1-S186A was not detectably phosphorylated in vitro, and therefore, no phosphopeptides were detected (data not shown). As shown in Fig. 5B, BZLF1-wt and BZLF1-S186A are constitutively phosphorylated in nonstimulated 293 cells (spots a to e), as has been reported previously (13). Treatment with TPA induced two additional phosphorylated peptides, designated x and y, in BZLF1-wt, which revealed a migration pattern identical to the phosphopeptides derived from BZLF1-wt phosphorylated in vitro. Phosphopeptide mapping of BZLF1-S186A yielded no additional spots after TPA treatment (Fig. 5B), indicating that S186 is the target for phosphorylation in vivo, which might involve TPA-activated PKC.
S186A abrogates TPA activation of BZLF1-responsive promoters and TPA responsiveness.
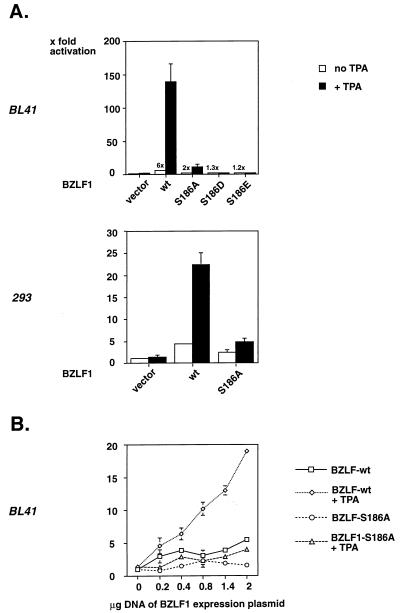
S186 is located in the DNA binding domain of BZLF1, which raised some concerns regarding whether its phosphorylation could be responsible for the enhancement of BZLF1 functions by TPA. To scrutinize this hypothesis, wild-type BZLF1 and the BZLF1-S186A mutant carrying an alanine substitution at position 186 were analyzed for the capacity to transactivate the BHRF1 luciferase reporter construct pBHRF1-Luc in BL41 cells in the presence or absence of TPA. As shown in Fig. 6A (upper panel). BZLF1-S186A expressed from the LPV promoter was slightly impaired in transactivating the BHRF1 promoter, but its TPA inducibility was dramatically reduced. Similar results at a somewhat lower level were obtained in the embryonic kidney cell line 293, indicating that no B-cell-specific factor is necessarily involved in this phenomenon (Fig. 6A, lower panel). In addition, two BZLF1 point mutants carrying the acidic amino acid aspartic acid (D) or glutamic acid (E) at position 186, BZLF1-S186D and BZLF1-S186E, were tested. Neither of them caused measurable transactivation (Fig. 6A, upper panel), indicating the requirement for phosphorylation at this position, which cannot be replaced by a negative charge. The BZLF1 mutant carrying a threonine substitution at position 159 (BZLF1-T159A) exhibited a phenotype comparable to that of BZLF1-wt, showing that S186 is crucial in mediating the TPA response (data not shown).
FIG. 6.
Mutation of serine 186 reduces the activation of BZLF1-dependent promoters and their TPA inducibility. (A) (Upper panel) BL41 cells were transiently transfected with the luciferase reporter plasmid BHRF1-Luc and a constitutive expression vector for either BZLF1-wt, BZLF1-S186A, BZLF1-S186D, or BZLF1-S186E. After treatment with TPA for 12 h, the cells were harvested and lysed, and luciferase activity was determined and compared with extracts from untreated cells. The reporter activity is expressed as fold stimulation based on relative light units in cells transfected with reporter plasmid and vector (left column). BZLF1 mutants were expressed at levels similar to wild-type (wt) BZLF1, as confirmed by Western blotting (data not shown). (Lower panel) The same assay as described in the upper panel was repeated with 293 cells, yielding comparable results. (B) Transient transfection of BL41 cells with the minimal reporter construct 4xZRE5tk-Luc, containing four ZRE5 sites upstream of the basal thymidine kinase promoter. The cells were cotransfected with increasing amounts of pLPV-BZLF1-wt or pLPV-BZLF1-S186A with or without TPA treatment, and the luciferase activities of the different cellular extracts were determined as in panel A.
Since the BHRF1 promoter is structurally complex, we also tested a minimal luciferase reporter plasmid containing four multimerized ZRE5 sites upstream of the basal thymidine kinase promoter (p4xZRE5tk-Luc). Up to fivefold transactivation and TPA inducibility were demonstrated with increasing amounts of BZLF1-wt but not with BZLF1-S186A (Fig. 6B).
The expression of BZLF1 is sufficient to trigger activation of the lytic cycle of EBV to induce DNA amplification from the lytic origin of DNA replication, oriLyt (for a review, see reference 38). We therefore asked whether the mutation of serine 186 has any significance on BZLF1’s capacity to induce DNA replication. oriLyt activation was measured in transient replication assays with the oriLyt plasmid p526, as described previously (28, 57, 59). Amplification of this plasmid is a direct consequence of DNA replication during the lytic phase. Substitution of serine 186 by alanine in BZLF1 completely abolished p526 amplification (data not shown), as has been concluded recently (23).
Taken together, these results emphasize the importance of serine 186 as a functional residue contributing to both transcription and DNA replication. Yet these data raised the question of whether phosphorylation of serine 186 primarily modulates the DNA binding affinity of BZLF1 for its different DNA binding motifs.
Serine 186 is critical for TPA-induced enhanced DNA binding of BZLF1.
Since S186 is located in the DNA binding domain of BZLF1, it was expected that mutation of this amino acid would affect BZLF1’s DNA binding affinity. Interestingly, S186 lies in a stretch of amino acids which is homologous to other transcription factors of the AP-1-family, such as c-Jun, c-Fos, and GCN4 (21). The crystal structure of c-Fos–c-Jun heterodimers bound to DNA demonstrates that five amino acids in the DNA binding domain contact DNA (26). Four of these five amino acids are conserved between c-Jun and BZLF1; they correspond to asparagine (N) 182, alanine (A) 185, cysteine (C) 189, and arginine (R) 190 in the DNA binding domain of BZLF1 (Fig. 1A, lower part). The fifth amino acid, alanine in c-Fos–c-Jun, corresponds to S186 in BZLF1. BZLF1-S186A, therefore, is identical to the core DNA binding domain of c-Jun, as has been noticed previously (40). In gel retardation assays, we observed that bacterially expressed BZLF1-S186A showed a slightly reduced relative binding affinity to some but not all BZLF1 binding sites. The formation of DNA-protein complexes with BZLF1-S186A was weaker with regard to ZRE3A and ZRE5 but as strong as BZLF1-wt with regard to ZRE2/7 and AP-1 sites, which are both better targets than ZRE3A and ZRE5 (Fig. 7A and data not shown) (46).
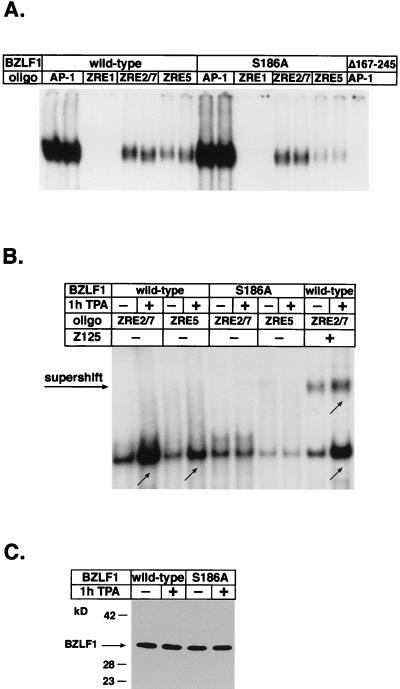
FIG. 7.
Role of S186 mutation and TPA treatment in DNA binding affinity of BZLF1. (A) Gel retardation assay with bacterially expressed and purified BZLF1-wt and BZLF1-S186A. Equal concentrations of BZLF1 protein were incubated with labeled oligonucleotides with different ZRE or AP-1 motifs prior to electrophoresis. Each reaction sample was performed in duplicate with extracts from different preparations. For unclear reasons, complexes with the ZRE1 oligonucleotide were not detected. As a control, a GST fusion protein which lacks the DNA binding domain of BZLF1 (GST-BZLF1Δ167-245) was used. (B) Gel retardation assay with nuclear extracts of 293 cells transfected with BZLF1-wt and BZLF1-S186A and treated or not with TPA for 1 h. Nuclear extracts were incubated with labeled ZRE2/7 or ZRE5 oligonucleotides in the absence or presence of the BZLF1-specific antibody Z125. (C) Immunodetection of BZLF1 in nuclear extracts of the transfected 293 cells used in panel B demonstrates equal amounts of BZLF1 protein in the differently treated extracts.
To determine the influence of TPA on BZLF1’s DNA binding affinity in vivo, we performed gel retardation assays with nuclear extracts of 293 cells transiently transfected with pCMV-BZLF1-wt and pCMV-BZLF1-S186A with or without TPA treatment. In parallel, nuclear extracts were subjected to Western blot analysis to ensure equal amounts of BZLF1 (Fig. 7C). As shown in Fig. 7B, TPA treatment reproducibly enhanced the DNA binding affinity of BZLF1-wt but not of BZLF1-S186A, which exhibited a reduced DNA affinity similar to bacterially expressed BZLF1-S186A (Fig. 7A). These observations suggested that TPA treatment improves the DNA binding affinity of BZLF1 and underscored the significance of residue S186, which again proved to be critical for the TPA effect.
To confirm that the DNA-protein complexes actually contained BZLF1, supershift assays with a BZLF1-specific antibody (Z125) (48) were performed. As expected, supershift complexes revealed a higher DNA binding affinity for ZRE2/7 (Fig. 7B) as well as AP-1 sites (data not shown) with BZLF1-wt after TPA treatment. Supershift complexes with BZLF1-S186A exhibited no TPA response (data not shown). In competitive gel shift analyses, DNA-protein complexes could be readily competed with the respective unlabeled oligonucleotides in a concentration-dependent manner, indicating the specificity of DNA recognition by BZLF1 (data not shown).
Putative recruitment of a cell factor upon BZLF1 phosphorylation.
Phosphorylation of amino acids which are integral parts of DNA binding domains is inhibitory of DNA binding affinity in general (30). Consequently, incorporation of a negatively charged phosphate at S186 which, in analogy to c-Jun, might contact DNA directly is likely to cause repulsion of DNA molecules. To address this problem, we analyzed DNA binding of recombinant purified BZLF1 which was phosphorylated in vitro by PKCα up to 50% (data not shown). As expected, in vitro phosphorylation decreased the efficiency of DNA-protein complex formation considerably (Fig. 8; compare lanes 5 and 6 to lanes 1 and 2 and lanes e and f to lanes a and b).
FIG. 8.
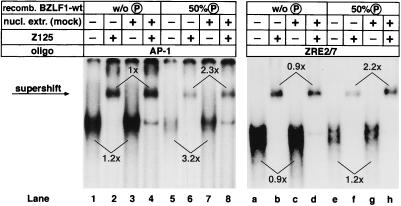
A putative cellular factor restores the DNA binding activity of BZLF1 phosphorylated in vitro. The relative binding affinity of bacterially expressed BZLF1 for different oligonucleotides with either AP-1 (left panel) or ZRE2/7 (right panel) motifs was analyzed after in vitro phosphorylation of BZLF1 by PKCα. The percentage of phosphate incorporation was about 50%, as determined with [32P]ATP (data not shown). Comparable amounts of nonphosphorylated and phosphorylated BZLF1-wt were used in each case, as confirmed by Western blotting (data not shown). Nuclear extracts prepared from mock-transfected 293 cells were added prior to incubation with the appropriate oligonucleotide in the indicated samples. To demonstrate the presence of BZLF1 in the protein-DNA complexes, a BZLF1-specific antibody (Z125) was added to yield supershifted complexes. Numbers above and below the autoradiography signals mark the relative enhancement of signal intensities in corresponding sample pairs following incubation with mock-transfected nuclear extracts. Signal intensities were quantified by a phosphorimager.
In order to establish experimental conditions similar to those in vivo, in vitro-phosphorylated BZLF1 was incubated with nuclear extracts from 293 cells prior to incubation with the labeled oligonucleotides in the presence of phosphatase inhibitors. This reconstruction experiment led to a reproducible two- to threefold increase of the DNA binding affinity of only phosphorylated BZLF1 (Fig. 8; compare lanes 5 and 6 to lanes 7 and 8 and lanes e and f to lanes g and h), which is about the same magnitude seen with BZLF1-wt extracted from TPA-treated 293 cells (Fig. 7B). No such effect was observed with nonphosphorylated BZLF1 (Fig. 8; compare lanes 5 and 7 to lanes 1 and 3 and lanes f and h to lanes b and d), suggesting the feasible involvement of a putative cell factor in a phosphorylation-dependent manner. However, no evidence was obtained in these experiments for any additional factor in the shift complex which is recruited upon phosphorylation of S186.
DISCUSSION
Our findings suggest that phorbol esters induce the posttranslational phosphorylation of BZLF1 to further activate native and artificial BZLF1-responsive promoters. Transcriptional activation assays with mutant and chimeric BZLF1 point to the serine residue at position 186 in BZLF1’s basic domain. Our data argue for the phosphorylation of S186 in response to TPA treatment, leading to enhanced DNA binding and transcriptional activity of the BZLF1 protein, while other mechanisms could be excluded (e.g., increased nuclear translocation of BZLF1 after TPA treatment [data not shown]).
The identification of S186 as the target for PKC in vitro (Fig. 4) and in vivo after TPA stimulation (Fig. 5) suggests that PKC might be the kinase phosphorylating BZLF1, since S186 is located within a conserved PKC phosphorylation motif (Fig. 1A). The phenotype of the BZLF1-S186A mutant is characterized by a marked reduction of the TPA effect on BZLF1-responsive promoters (Fig. 6), an effect which is comparable to that of a PKC inhibitor (Fig. 2A). Therefore, the most straightforward explanation for this finding involves PKC phosphorylating S186 in order to activate BZLF1 in a rapid response to TPA or related stimuli. The finding that virus titers depend on endogenous PKC levels is in line with this assumption.
Our data contradict those by Daibata and coworkers (13). They suggest a TPA-induced dephosphorylation and, after treatment with a PKC inhibitor, an enhanced phosphorylation of BZLF1. Although these discrepancies could be the result of cell-specific differences or experimental setup variations, we also noticed intensity changes in phosphopeptide spots a to e between mutant and wild-type BZLF1 (Fig. 5). Thus, we cannot rule out the possibility that phosphorylation of S186 has consequences for other phosphoacceptor residues in BZLF1 as well. It will be interesting to determine if S186 or S173 (41) is also a target for phosphatases, as has been shown for c-Jun (7). Furthermore, Jun family members are phosphorylated by a variety of cellular kinases (16, 43, 47), some of which might also be responsible for constitutive phosphorylation of BZLF1 (Fig. 5) (41). Future studies should reveal the nature of the PKC isoform and the subcellular location where BZLF1 phosphorylation takes place.
S186 phosphorylation seems to have opposite consequences for the DNA binding potential of BZLF1 in vitro and in vivo. The addition of nuclear fractions in gel shift experiments partially restored and improved binding of purified and phosphorylated BZLF1, which may indicate that BZLF1 cooperates with another protein in a phosphorylation-dependent manner (Fig. 8). Alternatively, phosphatase activity present in nuclear extracts or allosteric conformational changes might also modulate the DNA binding affinity of S186-phosphorylated BZLF1. Although we lack physical evidence for a cellular factor, this is an intriguing hypothesis, as several protein-protein interactions involve the basic DNA binding domain of transcription factors to stabilize DNA binding (17, 33). This finding is also supported by a report of Lieberman and coworkers, who observed the induction of a cellular binding activity together with BZLF1 in Raji cells after treatment with TPA (46). If this hypothesis is true, mutation of S186 to alanine would interfere with the recruitment of the BZLF1 interacting factor(s), which results in the inability to respond to TPA or similar signals.
A BZLF1-S186A mutant has been analyzed independently by Francis and coworkers (23) and was found to be unaffected in its ability to bind to ZRE sites or to activate transcription. BZLF1-S186A, but not a BZLF1-S186T mutant, failed to disrupt viral latency in EBV-infected Raji cells (22a), which implies that the threonine residue at position 186 in BZLF1-S186T could well serve as a PKC substrate like S186. However, no attempt was made to analyze these mutants in the presence of PKC activators, such as phorbol esters.
We propose that the mere binding of BZLF1 is not sufficient to fulfill its function as a transcription and replication factor. For full activation, BZLF1 needs to become phosphorylated at S186, presumably by PKC or a related kinase. By providing a phosphoserine-specific docking site, S186 might recruit an interacting cellular factor which stabilizes DNA binding and enhances transcriptional activation. Thus, future work will concentrate on the identification of this putative factor.
ACKNOWLEDGMENTS
This work was supported by a grant from the “Dr. Mildred Scheel Stiftung für Krebsforschung” (10-1016-Ze 1) to M.B. and by grant Mi 489/1-2 from the Deutsche Forschungsgemeinschaft to H.M. We were also supported by grant Wi319-/11-3 from the Deutsche Forschungsgemeinschaft, by grant CA70723-1 from the National Institutes of Health, and by institutional funds to W.H.
We thank Edith Pfitzner and Paul Lambert for gifts of recombinant plasmids and Emmanuel Drouet for providing the BZLF1-specific antibodies Z125 and Z130. We also thank George Miller for valuable data prior to publication (23).
REFERENCES
- 1.Aderem A. The MARCKS brothers: a family of protein kinase C substrates. Cell. 1992;71:713–716. doi: 10.1016/0092-8674(92)90546-o. [DOI] [PubMed] [Google Scholar]
- 2.Ašković S, Baumann R. Activation domain requirements for disruption of Epstein-Barr virus latency by ZEBRA. J Virol. 1997;71:6547–6554. doi: 10.1128/jvi.71.9.6547-6554.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Azzi A, Boscoboinik D, Hensey C. The protein kinase C family. Eur J Biochem. 1992;208:547–557. doi: 10.1111/j.1432-1033.1992.tb17219.x. [DOI] [PubMed] [Google Scholar]
- 4.Baer R, Bankier A T, Biggin M D, Deininger P L, Farrell P J, Gibson T J, Hatfull G, Hudson G S, Satchwell S C, Seguin C, Tufnell P S, Barell B G. DNA sequence and expression of the B95-8 Epstein-Barr virus genome. Nature (London) 1984;310:207–211. doi: 10.1038/310207a0. [DOI] [PubMed] [Google Scholar]
- 5.Binetruy B, Smeal T, Karin M. Ha-Ras augments c-Jun activity and stimulates phosphorylation of its activation domain. Nature (London) 1991;351:122–127. doi: 10.1038/351122a0. [DOI] [PubMed] [Google Scholar]
- 6.Borras A M, Strominger J L, Speck S H. Characterization of the ZI domains in the Epstein-Barr virus BZLF1 gene promoter: role in phorbol ester induction. J Virol. 1996;70:3894–3901. doi: 10.1128/jvi.70.6.3894-3901.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Boyle W J, Smeal T, Defize L H, Angel P, Woodgett J R, Karin M, Hunter T. Activation of protein kinase C decreases phosphorylation of c-Jun at sites that negatively regulate its DNA-binding activity. Cell. 1991;64:573–584. doi: 10.1016/0092-8674(91)90241-p. [DOI] [PubMed] [Google Scholar]
- 8.Boyle W J, van der Geer P, Hunter T. Phosphopeptide mapping and phosphoamino acid analysis by two-dimensional separation on thin-layer cellulose plates. Methods Enzymol. 1991;201:110–152. doi: 10.1016/0076-6879(91)01013-r. [DOI] [PubMed] [Google Scholar]
- 9.Chang Y N, Dong D L, Hayward G S, Hayward S D. The Epstein-Barr virus Zta transactivator: a member of the bZIP family with unique DNA-binding specificity and a dimerization domain that lacks the characteristic heptad leucine zipper motif. J Virol. 1990;64:3358–3369. doi: 10.1128/jvi.64.7.3358-3369.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chevallier-Greco A, Manet E, Chavrier P, Mosnier C, Daillie J, Sergeant A. Both Epstein-Barr virus (EBV)-encoded trans-acting factors, EB1 and EB2, are required to activate transcription from an EBV early promoter. EMBO J. 1986;5:3243–3249. doi: 10.1002/j.1460-2075.1986.tb04635.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chi T, Carey M. The ZEBRA activation domain: modular organization and mechanism of action. Mol Cell Biol. 1993;13:7045–7055. doi: 10.1128/mcb.13.11.7045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chi T, Lieberman P, Ellwood K, Carey M. A general mechanism for transcriptional synergy by eukaryotic activators. Nature (London) 1995;377:254–257. doi: 10.1038/377254a0. [DOI] [PubMed] [Google Scholar]
- 13.Daibata M, Humphreys R E, Sairenji T. Phosphorylation of the Epstein-Barr virus BZLF1 immediate-early gene product ZEBRA. Virology. 1992;188:916–920. doi: 10.1016/0042-6822(92)90553-2. [DOI] [PubMed] [Google Scholar]
- 14.Daibata M, Speck S H, Mulder C, Sairenji T. Regulation of the BZLF1 promoter of Epstein-Barr virus by second messengers in anti-immunoglobulin-treated B cells. Virology. 1994;198:446–454. doi: 10.1006/viro.1994.1056. [DOI] [PubMed] [Google Scholar]
- 15.Delecluse H-J, Hilsendegen T, Pich D, Zeidler R, Hammerschmidt W. Propagation and recovery of intact, infectious Epstein-Barr Virus from prokaryotic to human cells. Proc Natl Acad Sci USA. 1998;95:8245–8250. doi: 10.1073/pnas.95.14.8245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Derijard B, Hibi M, Wu I H, Barrett T, Su B, Deng T, Karin M, Davis R J. JNK1: a protein kinase stimulated by UV light and Ha-Ras that binds and phosphorylates the c-Jun activation domain. Cell. 1994;76:1025–1037. doi: 10.1016/0092-8674(94)90380-8. [DOI] [PubMed] [Google Scholar]
- 17.Du W, Maniatis T. The high mobility group protein HMG I(Y) can stimulate or inhibit DNA binding of distinct transcription factor ATF-2 isoforms. Proc Natl Acad Sci USA. 1994;91:11318–11322. doi: 10.1073/pnas.91.24.11318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Farrell P J, Rowe D T, Rooney C M, Kouzarides T. Epstein-Barr virus BZLF1 trans-activator specifically binds to a consensus AP-1 site and is related to c-fos. EMBO J. 1989;8:127–132. doi: 10.1002/j.1460-2075.1989.tb03356.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Flemington E, Speck S H. Autoregulation of Epstein-Barr virus putative lytic switch gene BZLF1. J Virol. 1990;64:1227–1232. doi: 10.1128/jvi.64.3.1227-1232.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Flemington E, Speck S H. Evidence for coiled-coil dimer formation by an Epstein-Barr virus transactivator that lacks a heptad repeat of leucine residues. Proc Natl Acad Sci USA. 1990;87:9459–9463. doi: 10.1073/pnas.87.23.9459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Flemington E, Speck S H. Identification of phorbol ester response elements in the promoter of Epstein-Barr virus putative lytic switch gene BZLF1. J Virol. 1990;64:1217–1226. doi: 10.1128/jvi.64.3.1217-1226.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Flemington E K, Borras A M, Lytle J P, Speck S H. Characterization of the Epstein-Barr virus BZLF1 protein transactivation domain. J Virol. 1992;66:922–929. doi: 10.1128/jvi.66.2.922-929.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22a.Francis, A. L., et al. Personal communication.
- 23.Francis A L, Gradoville L, Miller G. Alteration of a single serine in the basic domain of the Epstein-Barr virus ZEBRA protein separates its functions of transcriptional activation and disruption of latency. J Virol. 1997;71:3054–3061. doi: 10.1128/jvi.71.4.3054-3061.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Furnari F B, Zacny V, Quinlivan E B, Kenney S, Pagano J S. RAZ, an Epstein-Barr virus transdominant repressor that modulates the viral reactivation mechanism. J Virol. 1994;68:1827–1836. doi: 10.1128/jvi.68.3.1827-1836.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Glaser R, Nonoyama M. Host cell regulation of induction of Epstein-Barr virus. J Virol. 1974;14:174–176. doi: 10.1128/jvi.14.1.174-176.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Glover J N, Harrison S C. Crystal structure of the heterodimeric bZIP transcription factor c-Fos–c-Jun bound to DNA. Nature (London) 1995;373:257–261. doi: 10.1038/373257a0. [DOI] [PubMed] [Google Scholar]
- 27.Goodnight J, Mischak H, Mushinski J F. Selective involvement of protein kinase C isozymes in differentiation and neoplastic transformation. Adv Cancer Res. 1994;64:159–209. doi: 10.1016/s0065-230x(08)60838-6. [DOI] [PubMed] [Google Scholar]
- 28.Hammerschmidt W, Sugden B. Identification and characterization of oriLyt, a lytic origin of DNA replication of Epstein-Barr virus. Cell. 1988;55:427–433. doi: 10.1016/0092-8674(88)90028-1. [DOI] [PubMed] [Google Scholar]
- 29.Hanahan D. Techniques for transformation of E. coli. In: Glover D, editor. DNA cloning. A practical approach. Vol. 1. Oxford, England: IRL Press; 1985. pp. 109–135. [Google Scholar]
- 30.Hill C S, Treisman R. Transcriptional regulation by extracellular signals: mechanisms and specificity. Cell. 1995;80:199–211. doi: 10.1016/0092-8674(95)90403-4. [DOI] [PubMed] [Google Scholar]
- 31.Hocevar B A, Burns D J, Fields A P. Identification of protein kinase C (PKC) phosphorylation sites on human lamin B. Potential role of PKC in nuclear lamina structural dynamics. J Biol Chem. 1993;268:7545–7552. [PubMed] [Google Scholar]
- 32.Hong Y, Holley-Guthrie E, Kenney S. The bZip dimerization domain of the Epstein-Barr virus BZLF1 (Z) protein mediates lymphoid-specific negative regulation. Virology. 1997;229:36–48. doi: 10.1006/viro.1996.8413. [DOI] [PubMed] [Google Scholar]
- 33.Jain J, McCaffrey P G, Miner Z, Kerppola T K, Lambert J N, Verdine G L, Curran T, Rao A. The T-cell transcription factor NFATp is a substrate for calcineurin and interacts with Fos and Jun. Nature (London) 1993;365:352–355. doi: 10.1038/365352a0. [DOI] [PubMed] [Google Scholar]
- 34.Jaken S. Protein kinase C isoenzymes and substrates. Curr Opin Cell Biol. 1996;8:168–173. doi: 10.1016/s0955-0674(96)80062-7. [DOI] [PubMed] [Google Scholar]
- 35.Karin M. The regulation of AP-1 activity by mitogen-activated protein kinases. J Biol Chem. 1995;270:16483–16486. doi: 10.1074/jbc.270.28.16483. [DOI] [PubMed] [Google Scholar]
- 36.Katz D A, Baumann R P, Sun R, Kolman J L, Taylor N, Miller G. Viral proteins associated with the Epstein-Barr virus transactivator, ZEBRA. Proc Natl Acad Sci USA. 1992;89:378–382. doi: 10.1073/pnas.89.1.378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kenney S C, Holley-Guthrie E, Quinlivan E B, Gutsch D, Zhang Q, Bender T, Giot J-F, Sergeant A. The cellular oncogene c-myb can interact synergistically with the Epstein-Barr virus BZLF1 transactivator in lymphoid cells. Mol Cell Biol. 1992;12:136–146. doi: 10.1128/mcb.12.1.136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kieff E. Epstein-Barr virus and its replication. In: Fields B N, Knipe D M, Howley P M, Chanock R M, Monath T P, Melnick J L, Roizman B, Straus S E, editors. Virology. 3rd ed. Philadelphia, Pa: Lippincott-Raven Publishers; 1996. pp. 2343–2396. [Google Scholar]
- 39.Kolch W, Heidecker G, Kochs G, Hummel R, Vahidi H, Mischak H, Finkenzeller G, Marme D, Rapp U R. Protein kinase C alpha activates RAF-1 by direct phosphorylation. Nature (London) 1993;364:249–252. doi: 10.1038/364249a0. [DOI] [PubMed] [Google Scholar]
- 40.Kolman J L, Taylor N, Gradoville L, Countryman J, Miller G. Comparing transcriptional activation and autostimulation by ZEBRA and ZEBRA/c-Fos chimeras. J Virol. 1996;70:1493–1504. doi: 10.1128/jvi.70.3.1493-1504.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kolman J L, Taylor N, Marshak D R, Miller G. Serine-173 of the Epstein-Barr virus ZEBRA protein is required for DNA binding and is a target for casein kinase II phosphorylation. Proc Natl Acad Sci USA. 1993;90:10115–10119. doi: 10.1073/pnas.90.21.10115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kouzarides T, Packham G, Cook A, Farrell P J. The BZLF1 protein has a coiled coil structure dimerisation domain without a heptad leucine repeat but with homology to the C/EBP leucine zipper. Oncogene. 1991;6:195–204. [PubMed] [Google Scholar]
- 43.Kyriakis J M, Banerjee P, Nikolakaki E, Dai T, Rubie E A, Ahmad M F, Avruch J, Woodgett J R. The stress-activated protein kinase subfamily of c-Jun kinases. Nature (London) 1994;369:156–160. doi: 10.1038/369156a0. [DOI] [PubMed] [Google Scholar]
- 44.Lambert P F, Spalholz B A, Howley P M. A transcriptional repressor encoded by BPV-1 shares a common carboxy-terminal domain with the E2 transactivator. Cell. 1987;50:69–78. doi: 10.1016/0092-8674(87)90663-5. [DOI] [PubMed] [Google Scholar]
- 45.Laux G, Freese U K, Fischer R, Polack A, Kofler E, Bornkamm G W. TPA-inducible Epstein-Barr virus genes in Raji cells and their regulation. Virology. 1988;162:503–507. doi: 10.1016/0042-6822(88)90496-5. [DOI] [PubMed] [Google Scholar]
- 46.Lieberman P M, Hardwick J M, Sample J, Hayward G S, Hayward S D. The Zta transactivator involved in induction of lytic cycle gene expression in Epstein-Barr virus-infected lymphocytes binds to both AP-1 and ZRE sites in target promoter and enhancer regions. J Virol. 1990;64:1143–1155. doi: 10.1128/jvi.64.3.1143-1155.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lin A, Frost J, Deng T, Smeal T, al-Alawi N, Kikkawa U, Hunter T, Brenner D, Karin M. Casein kinase II is a negative regulator of c-Jun DNA binding and AP-1 activity. Cell. 1992;70:777–789. doi: 10.1016/0092-8674(92)90311-y. [DOI] [PubMed] [Google Scholar]
- 48.Mikaelian I, Drouet E, Marechal V, Denoyel G, Nicolas J C, Sergeant A. The DNA-binding domain of two bZIP transcription factors, the Epstein-Barr virus switch gene product EB1 and Jun, is a bipartite nuclear targeting sequence. J Virol. 1993;67:734–742. doi: 10.1128/jvi.67.2.734-742.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Newton A C. Regulation of protein kinase C. Curr Opin Cell Biol. 1997;9:161–167. doi: 10.1016/s0955-0674(97)80058-0. [DOI] [PubMed] [Google Scholar]
- 50.Papavassiliou A G, Treier M, Bohmann D. Intramolecular signal transduction in c-Jun. EMBO J. 1995;14:2014–2019. doi: 10.1002/j.1460-2075.1995.tb07193.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Parker D, Ferreri K, Nakajima T, LaMorte V J, Evans R, Koerber S, Hoeger C, Montminy M R. Phosphorylation of CREB at Ser-133 induces complex formation with CREB-binding protein via a direct mechanism. Mol Cell Biol. 1996;16:694–703. doi: 10.1128/mcb.16.2.694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Pawlita M, Clad A, zur Hausen H. Complete DNA sequence of lymphotropic papovavirus: prototype of a new species of the polyomavirus genus. Virology. 1985;143:196–211. doi: 10.1016/0042-6822(85)90108-4. [DOI] [PubMed] [Google Scholar]
- 53.Pulverer B J, Kyriakis J M, Avruch J, Nikolakaki E, Woodgett J R. Phosphorylation of c-jun mediated by MAP kinases. Nature (London) 1991;353:670–674. doi: 10.1038/353670a0. [DOI] [PubMed] [Google Scholar]
- 54.Ryseck R-P, Bravo R. c-Jun, Jun B, and Jun D differ in their binding affinities to AP-1 and CRE consensus sequences: effect of Fos proteins. Oncogene. 1991;6:533–542. [PubMed] [Google Scholar]
- 55.Sairenji T. The mechanisms of Epstein-Barr virus activation in B cells. Nippon Rinsho. 1997;55:316–320. [PubMed] [Google Scholar]
- 56.Sarisky R T, Gao Z, Lieberman P M, Fixman E D, Hayward G S, Hayward S D. A replication function associated with the activation domain of the Epstein-Barr virus Zta transactivator. J Virol. 1996;70:8340–8347. doi: 10.1128/jvi.70.12.8340-8347.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Schepers A, Pich D, Hammerschmidt W. A transcription factor with homology to the AP-1 family links RNA transcription and DNA replication in the lytic cycle of Epstein-Barr virus. EMBO J. 1993;12:3921–3929. doi: 10.1002/j.1460-2075.1993.tb06070.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Schepers A, Pich D, Hammerschmidt W. Activation of oriLyt, the lytic origin of DNA replication of Epstein-Barr virus, by BZLF1. Virology. 1996;220:367–376. doi: 10.1006/viro.1996.0325. [DOI] [PubMed] [Google Scholar]
- 59.Schepers A, Pich D, Mankertz J, Hammerschmidt W. cis-acting elements in the lytic origin of DNA replication of Epstein-Barr virus. J Virol. 1993;67:4237–4245. doi: 10.1128/jvi.67.7.4237-4245.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Sista N D, Barry C, Sampson K, Pagano J. Physical and functional interaction of the Epstein-Barr virus BZLF1 transactivator with the retinoic acid receptors RAR alpha and RXR alpha. Nucleic Acids Res. 1995;23:1729–1736. doi: 10.1093/nar/23.10.1729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Speck S H, Chatila T, Flemington E. Reactivation of Epstein-Barr virus: regulation and function of the BZLF1 gene. Trends Microbiol. 1997;5:399–405. doi: 10.1016/S0966-842X(97)01129-3. [DOI] [PubMed] [Google Scholar]
- 62.Takada K. Cross-linking of cell surface immunoglobulin induces Epstein-Barr virus in Burkitt lymphoma lines. Int J Cancer. 1984;33:27–32. doi: 10.1002/ijc.2910330106. [DOI] [PubMed] [Google Scholar]
- 63.Takada K, Ono Y. Synchronous and sequential activation of latently infected Epstein-Barr virus genomes. J Virol. 1989;63:445–449. doi: 10.1128/jvi.63.1.445-449.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Takenaka I, Morin F, Seizinger B R, Kley N. Regulation of the sequence-specific DNA binding function of p53 by protein kinase C and protein phosphatases. J Biol Chem. 1995;270:5405–5411. doi: 10.1074/jbc.270.10.5405. [DOI] [PubMed] [Google Scholar]
- 65.Taylor N, Flemington E, Kolman J L, Baumann R P, Speck S H, Miller G. ZEBRA and a Fos-GCN4 chimeric protein differ in their DNA-binding specificities for sites in the Epstein-Barr virus BZLF1 promoter. J Virol. 1991;65:4033–4041. doi: 10.1128/jvi.65.8.4033-4041.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Urier G, Buisson M, Chambard P, Sergeant A. The Epstein-Barr virus early protein EB1 activates transcription from different responsive elements including AP-1 binding sites. EMBO J. 1989;8:1447–1453. doi: 10.1002/j.1460-2075.1989.tb03527.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Wang Y C, Huang J M, Montalvo E A. Characterization of proteins binding to the ZII element in the Epstein-Barr virus BZLF1 promoter: transactivation by ATF1. Virology. 1997;227:323–330. doi: 10.1006/viro.1996.8326. [DOI] [PubMed] [Google Scholar]
- 68.Zhang Q, Gutsch D, Kenney S. Functional and physical interaction between p53 and BZLF1: implications for Epstein-Barr virus latency. Mol Cell Biol. 1994;14:1929–1938. doi: 10.1128/mcb.14.3.1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.zur Hausen H, O’Neill F J, Freese U K. Persisting oncogenic herpesvirus induced by tumour promoter TPA. Nature (London) 1978;272:373–375. doi: 10.1038/272373a0. [DOI] [PubMed] [Google Scholar]