Abstract
Peptide vaccine based on tumor-associated antigen (TAA), which usually belongs to self-antigen with poor immunogenicity, has been considered as an attractive option for treatment of malignant tumors. The ideal TAA epitopes should have stable affinity to major histocompatibility complex (MHC) molecules and elicit strong anti-tumor immune response. Although point-mutation technology of TAA peptide may increase the binding capability to MHC molecules, some previous studies have revealed that part of the variant peptides results in lymphocyte not to effectively cross-recognize and kill the target tumor expressed wild-type TAA. Here, we designed a novel HLA-A2-restricted mutated TAA Survivin epitope nonapeptide Sur79L2 (KLSSGCAFL) that showed higher binding ability compared to wild-type peptide Sur79 (KHSSGCAFL) in T2-binding assays. To investigate whether Sur79L2 can induce Survivin-specific anti-hepatocellular carcinoma (HCC) response, we stimulated tumor-associated lymphocytes from a HCC patient with Sur79L2 in vitro. IFN-γ release and cytotoxicity assays showed Sur79L2 could effectively cross-recognize and lysis T2 cell plus peptide Sur79 and HCC cell lines (expression of wild-type Survivin antigen) in an HLA-A2-restricted manner. In contrast, peptide Sur95 (ELTLGEFLKL) that has been reported as a very promising anti-tumor epitope in a variety of tumors except HCC were not able to generate detectable cytotoxic immune responses against HCC in this study. Our results suggest that point-mutated peptide Sur79L2 is a new HLA-A2-restricted CTL epitope and may be useful for the immunotherapy for patients with HCC.
Electronic supplementary material
The online version of this article (doi:10.1007/s00262-012-1323-4) contains supplementary material, which is available to authorized users.
Keywords: Cross-recognition, HCC, Survivin, Point-mutation peptide, CTL
Introduction
Hepatocellular carcinoma (HCC) is one of the most prevalent cancers in the world, which represents the third leading cause of cancer death world-wide. The Far East is the high incidence of HCC, and most deaths occur in this region. Resection, chemo-embolization, local ablation and radiotherapy are the most commonly curative treatments in the clinic. However, these treatments have not remarkably shown a survival advantage for HCC patients [1].
Immunotherapy has been demonstrated as an attractive therapeutic option for malignant tumors in the past few years, which has the advantages of strong anti-tumor immunogenicity and weak side effects. One of the tactics for tumor immunotherapy is tumor-associated antigen (TAA) vaccine that has been shown repeatedly to induce a specific and sometimes long-lasting response by the immune system in some studies. An ideal universal TAA for tumor vaccine should have the following characteristics: (1) present on the majority of human cancers but rarely on normal tissues, (2) include peptide sequences that bind to major histocompatibility complex (MHC) molecules effectively and (3) be recognized by the T cell repertoire of tumor patients in an MHC-restricted fashion and can elicit specific T cell response [2, 3]. A number of TAA epitope peptides, including α-fetoprotein (AFP), NY-ESO1 and MAGE-A, have been the productions of anti-HCC vaccine to conduct clinical trials in the last decade. A randomized phase II trial based on AFP has shown that there were increased frequencies of circulating AFP-specific T cells after vaccination, but no clinical responses were detected in the patients [4–6]. Therefore, the identification of appropriate CTL epitope from TAA has become the key step in the development of peptide-based HCC vaccine.
Survivin belongs to the inhibitory apoptotic protein family (IAP), which has been implicated in blocking mitochondrial-dependent apoptosis by targeting caspase 9 and promoting tumor progression. Survivin is aberrantly expressed in virtually every human cancer but is undetectable in normal differentiated adult tissues. Among them, approximately 30–90 % of patients with hepatocellular carcinoma were confirmed overexpression of Survivin [7–9]. Some clinical trials based on Survivin-peptide vaccine have performed on advanced melanoma, breast, colon and urothelial cancer patients. Published partial results have shown that in several cases, there was induction of antigen-specific T cell responses to the Survivin-peptide vaccine [10–12]. However, currently, there are not yet reported on the Survivin-peptide vaccine for treatment of primary HCC up to date.
Most of the identified Survivin CTL-reactive epitope peptides were HLA-A2- and HLA-A24-restricted, which were generally wild-type and shown to have a good binding stability with class I HLA molecule. Survivin 95–104 (ELTLGEFLKL, Sur95) was a much-studied HLA-A2-restricted epitope decapeptide with an intermediate affinity for the HLA-A2 molecules, which has been shown previously to be a T cell epitope eliciting specific CTLs in melanoma, breast and colorectal cancer patients [13, 14]. However, few data are available concerning a T cell-mediated response directed against Sur95 in HCC patients. Some recent studies have reported that T lymphocytes of tumor patients lacked the high affinity TCR molecule for TAA-specific reorganization because of thymic negative selection. Although TAA peptide-induced CTL reactivity could be observed in ex vitro experiments, the efficacy of in vivo experiments was not ideal. Possible explanation is that wild-type TAA peptide could not effectively activate and expand T lymphocytes with lower affinity T cell receptor LATCR in vivo. Technology of point mutation of TAA peptide has been validated to be able to increase the affinity to MHC molecules and the combination with the TCR molecule, which would lead to the activation of LATCR T lymphocytes [15–17]. However, Jordan et al. [18] recently have revealed that some variant peptides that strongly stimulated a specific T cell clone in vitro, but elicited fewer tumor-specific T cells in vivo, indicating that appropriate mutated peptide-specific T cells should have the capability of cross-reorganization to wild-type TAA.
In the present study, we filtered one novel HLA-A2-restricted mutated Survivin CTL epitope nonapeptide Sur79L2 (KLSSGCAFL) against HCC based on our previous research [26], the residue at position 2 of which was replaced with Leucine (Leu). A HLA-A2-positive HCC patient has been selected and his tumor-associated lymphocytes (TALs) from ascites were stimulated with Survivin peptides to induce Survivin-specific CD8+ T cell immunity in vitro. To investigate the cross-recognition to wild-type Survivin antigen with Sur79L2-specific T cell, we then detected the Survivin-specific anti-HCC immune response using HCC cell lines that all express wild-type Survivin. Our results showed that Sur79L2 may become an appropriate candidate epitope for anti-HCC peptide vaccine. In addition, Sur95 proved to not be able to elicit a peptide-specific CTL response on the HCC patient TALs in our experiments, which was distinguished with previous studies related to other carcinomas.
Materials and methods
Epitope prediction and synthesizing
Two matrix-based prediction algorithms, BIMAS (http://bimas.dcrt.nih.gov/molbio/hla_bind/) and SYFPEITHI (http://www.syfpeithi.de/Scripts/MHCServer.dll/EpitopePrediction.htm), were used for the prediction of peptides binding to HLA-A2 and HLA-A24. Two nonapeptides and one decapeptide derived from the human Survivin amino acid sequence, and one nonapeptide from HIV virus [HIVpol (476–484) (ILLEPVHGV)], which served as a positive control in MHC peptide-binding assay and a negative control in inducing Survivin-specific CTL reactions. All peptides were purchased from Sinoasis Pharmaceuticals, INC (Guangzhou, China) and provided at 90 % purity, as verified by HPLC and MS analysis. All peptides used are listed in Table 1.
Table 1.
Survivin candidate epitope predicted by computational methods
| Name | Position of peptide | Sequence | BIMAS score | SYFPEITHI score | FIb for HLA-A2-binding | ||
|---|---|---|---|---|---|---|---|
| HLA-A2 | HLA-A24 | HLA-A2 | HLA-A24 | ||||
| HIVpol476 | HIV-1pol (476–484) | ILKEPVHGV | 39.025 | 0.12 | 30 | 2 | 5.92 |
| Sur95 | Survivin (95–104) | ELTLGEFLKL | 3.044 | 5.28 | 19 | 13 | 1.63 |
| Sur79 | Survivin (79–87) | KHSSGCAFL | 0.286 | 0.8 | 15 | 11 | 0.46 |
| Sur79L2 | SurvivinM (79–87)a | KLSSGCAFL | 411.226 | 8 | 25 | 11 | 6.02 |
aThe Histidine (H) residue at position 80 is mutated to Leucine (L)
bFI = (MFI sample − MFI background)/MFI background, where MFI background represents the value without peptide, according to the results of Fig S1
MHC stabilization assay
The ability of peptides to bind to HLA-A2 molecules was evaluated using a major histocompatibility complex (MHC) class I stabilization assay. Briefly, T2 cells were grown overnight at 37 °C in Iscove’s modified Dulbecco’s medium (Gibco/Invitrogen, Carlsbad, CA) supplemented with 20 % fetal bovine serum (FBS, Gibco/Invitrogen). Cells were harvested and washed twice with serum-free AIM-V medium (Gibco/Invitrogen) and then incubated (5 × 105/250 μl) with 50 μM of individual peptides at 37 °C for 18 h in AIM-V medium. Following incubation, cells were washed twice with staining buffer (PBS plus 3 % FBS) and stained for 30 min at 4 °C with the fluorescein isothiocyanate (FITC)-conjugated anti-HLA-A2.1 antibody (Santa Cruz Biotechnology, Inc., USA, Clone BB7.2) at a concentration of 500 ng per reaction in 100 μl of staining buffer. Stained cells were washed twice with staining buffer and resuspended in PBS. HLA-A2 expression was measured flow cytometrically using an Epics-XL flow cytometer (Beckman Coulter, Inc. Miami, FL), and the data were analyzed using the EXPO32 v1.2 software. The fluorescence index (FI) was calculated using the following formula: FI = [mean fluorescence intensity (MFI) sample − MFI background]/MFI back ground, where MFI background represents the value without peptide. A test peptide capable of causing a ≥200 % increase in MFI values (FI ≥ 2) over the no-peptide control was considered a “high binder.”
Patient TALs and HCC cell lines
The patient (C367, HLA-A2+, HLA-A24+) was a 63-year-old man who had been diagnosed with hepatocellular carcinoma (HCC) for 6 months. CT examination is recognized as the central type of liver cancer with peritoneal metastasis (Fig. S4). Bloody ascites is the typical symptom in this patient, and he had received chemotherapy with dacarbazine alone at the PANYU Central Hospital of Guangzhou. TALs were isolated by Ficoll-Paque Plus (GE Healthcare, Uppsala, Sweden) density gradient centrifugation from the ascites samples. Erythrocytes were lysed using ACK hypotonic solution, and then, mononuclear cells were washed twice with PBS, resuspended in RPMI 1640 supplemented with 30 % heat-inactivated FBS (Gibco/Invitrogen) and 10 % DMSO (Sigma-Aldrich, Madrid, Spain) and cryopreserved in liquid nitrogen. In addition, one HLA-A2-positive healthy volunteer (Donor 1, aged 45 years old) and one HLA-A24-positive healthy volunteer (Donor 2, aged 48 years old) were enrolled as control group, and their PBLs from 50 ml peripheral blood were separated using density gradient centrifugation in this study. Before our experiments, informed consent was obtained from all the patient and donors (Fig. S5).
The human TAP-deficient T2 cell line and the human hepatocellular carcinoma cell line HepG2 (Survivin+, HLA-A2+, HLA-A24+) were purchased from the American Type Culture Collection (Rockville, MD). Other hepatocellular carcinoma cell lines, Huh-1 (Survivin+, HLA-A2+, HLA-A11+) and Bel-7402 (Survivin+, HLA-A68+, HLA-A2−) were maintained in our laboratory. The cell lines were cultured in RPMI (Gibco/Invitrogen) or DMEM (Gibco/Invitrogen), respectively, supplemented with 10 % FBS (Gibco/Invitrogen) and antibiotics (penicillin G 100 IU/ml and Streptomycin 50 μg/ml).
Prior to our study, HLA-A allele phenotype of the HCC patient, healthy donors and HCC cell lines were verified by a flow cytometer (Beckman Coulter) analysis using the PE-conjugated HLA-A2-specific antibody (MBL Inc, Japan, Clone BB7.2) and PE-conjugated HLA-A24-specific antibody (MBL Inc, Japan, Clone 22E1).
Detection of Survivin expression
The ascites supernatant were used to detect the Survivin levels with the PathScan Survivin Sandwich ELISA Kit(Cell Signaling Technology, Danvers, MA)according to the manufacturer’s directions.
RT-PCR was carried out to detect the expression of Survivin mRNA in HCC cell lines, HepG2, Huh-1 and Bel-7402. Briefly, total RNA was extracted from HCC cell lines and reverse-transcribed with avian myeloblastosis virus (AMV) reverse transcriptase and oli-godT (Clontech, Palo Alto, CA, USA). Quality of the cDNA was confirmed by PCR for GAPDH. The sequences of primers used to amplify Survivin were as follows: forward primer: 5′-AGCGGATGGCCGAGGCTGGCTTCATC-3′; reverse primer: 5′-CTTCTTATTGTTGGTTTCCTTTGC-3′. The cycling conditions were as follows: initial denaturation at 94 °C for 5 min, followed by 30 cycles at 94 °C for 30 s, 55 °C for 40 s and 72 °C for 40 s. Products were analyzed on 1 % agarose gels containing 0.01 mg/ml ethidium bromide. The PCR products of Survivin gene fragments were gel extracted and sequenced to confirm no gene mutation at Sur79 and Sur95.
Survivin expression of HepG2, Huh-1 and Bel-7402 were assessed by Western blot analysis. Briefly, cells were lysed in buffer containing 1 % Triton X-100, 20 mM Tris–HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 1 mM EGTA, 10 % glycerin, 1 mM PMSF (phenylmethylsulfonyl fluoride) and 1 mg/mL Leupeptin. Protein concentration was determined by using protein assay kit (Pierce, Rockford, IL). Total protein (50 μg) was separated on 15 % polyacrylamide gel, blotted on polyvinylidene difluoride (PVDF) membrane (Millipore, Bedford, MA), and incubated with a Survivin-specific antibody (Cell Signalling, Danvers, MA, Clone 6E4). Bands of proteins were detected by an enhanced chemiluminescence detection kit (ECL, Amersham Pharmacia Biotech, NJ, USA).
Antigen stimulation of TALs
To induce the generation of CD8+ Survivin-specific T cells, TALs from patients were first sensitized for 2 week with 10 μg/ml of the Survivin-derived peptide. At day 0, TALs were plated in 2 ml/well at a concentration of 2 × 106 cells in 24-well plates (Nunc, Roskilde, Denmark) in AIM-V medium (Gibco/Invitrogen), 5 % heat-inactivated fetal bovine serum and 2 mM of l-glutamine in the presence of 10 mM of peptide. In each experiment, a well without peptide was also included. Two days later, 300 IU/ml recombinant interleukin 2 (R&D InC, Minneapolis, USA) were added to the cultures; then, cells were maintained under the same conditions. Restimulation was performed every 6 days, using autologous TALs pulsed with 10 μg/ml of Survivin peptides for 2 h and then γ-irradiated (3,000 rad). Free peptide was washed out and APCs added at a stimulator: responder ratio of 5:1. The cultured cells were tested for reactivity in the ELISPOT on day 12.
ELISPOT assay
The ELISPOT assay was used to quantify peptide epitope-specific IFN-γ releasing effector cells and has been described previously [19]. Briefly, PVDF membrane-bottomed 96-well plates (Millipore) were coated with an anti-IFN-γ antibody (U-CyTech, Amsterdam, the Netherlands, Clone MD-1), and nonspecific binding was blocked using AIM-V (Life Technologies, Inc., Gaithersburg, MD). Lymphocytes were added at different cell concentrations together with the specific peptides and were incubated overnight at 37 °C. A positive control (5 μg/ml concanavalin [ConA]) and a negative control (negative peptide HIVpol476 [NP]) were included in all assays. After two washes, the biotinylated detector antibody (U-Cytech) was added. Specific binding was visualized using alkaline phosphatase-avidin together with the respective substrate (Life Technologies, Inc.). The reaction was terminated on the appearance of dark purple spots, which were quantitated using the AlphaImager System (Alpha Innotech, San Leandro, CA). The assays were all performed in duplicates for each peptide antigen. PBLs from healthy donors were used as control.
Isolation of CD8+ Survivin-specific T cells
To obtain CD8+ Survivin-specific T cells, TALs stimulated with peptides for 14 days were isolated by negative selection using magnetic beads, according to the protocol provided by the manufacturer (Miltenyi Biotec, Bergisch Gladbach, Germany). The purity (≥95 % CD8+ T cells) was confirmed by flow cytometry using an FITC-conjugated rat anti-human CD8 antibody. The CD8+ T lymphocytes were then cultured in RPMI1640 medium containing 10 % human serum and 100 IU/ml recombinant interleukins (IL-2, R&D) for 3 days for CTL assays.
CTL assay
CTL assays were performed using a nonradioactive CTL assay kit, CytoTox96 (Promega, Madison, WI), that measures lactate dehydrogenase (LDH) released by target cells in accordance with manufacturer’s instructions. T2 cells were used as target cells and were pulsed overnight with individual test peptides (10 μM). Peptide-stimulated CD8+ cells were incubated with these peptide-loaded T2 target cells at different effector/target (E/T) cell ratios ranging from 40:1 to 10:1 in a 96-well U-bottom plate for 4 h at 37 °C. Fifty microliters of culture supernatants was then harvested from each well and tested for LDH activity using a chromogenic substrate. Absorbance was measured at 495 nm using an enzyme-linked immunosorbent assay (ELISA) reader (Bio-Rad, Hercules, CA). The percent specific cell lysis was calculated using the following formula: 100 × [(experimental-effector spontaneous) − target spontaneous]/[target maximum − target spontaneous].
CTL assays were also used to evaluate the CTL activity for hepatoma cell line HepG2 and Huh-1 cells at various E/T ratios. For antibody blocking experiments, hepatoma cells were incubated for 1 h at 4 °C with 10 μg/ml anti-HLA-A24 mAb (MBL, Clone 22E1) and anti-HLA-A2 mAb (MBL, Clone BB7.2) before seeding in 96-well plates.
Morphological analysis of HCC incubated with CTL
Hepatoma cell lines and Survivin peptide-induced TALs were co-cultured as effector cell and target cell, respectively. Morphological changes of HCC cells were monitored daily by an inverted phase-contrast microscope (IX51; Olympus, Tokyo, Japan) attached a QImage Micropublisher 5.0 RTV camera, and images were acquired at 200× magnification for incubation of 48 h.
Intracellular Granzyme B staining
Intracellular Granzyme B (GzmB) staining of effecter CD8+ cells was performed as described previously. Briefly, the effectors and targets were incubated together at 37 °C, 5 % CO2 for 4 h; at the end of the incubation lymphocytes were collected, fixed, permeabilized (for intracellular GzmB staining) and washed with PBS three times. Cells labeled with PE-conjugated anti-Granzyme B antibody (eBioscience Inc, Headquarters, Clone GB11) and FITC-conjugated anti-CD8 (eBioscience Inc, Clone HIT8a) antibody in separate tubes. Cells are incubated in dark on ice for 30 min. After washing and fixing, cells were analyzed by flow cytometry (Beckman Coulter). Appropriate isotype control (IgG1) is included in all experiments.
Statistical analysis
The results were expressed as the mean ± standard deviation (SD). The values were compared by one-way analysis of variance with 0.05 as the minimum level of significance.
Results
Mutated Survivin peptide Sur79L2 has the higher HLA-A2-binding scores with computational prediction
Two HLA-peptide-binding prediction computational methods BIMAS and SYFPEITHI were performed to analyze the HLA-A2- and HLA-A24-restricted CTL epitopes. High binding scores of BIMAS and SYFPEITHI mean high affinities to HLA-A molecules. The sequences and binding scores of four peptides, including three peptides derived from Survivin protein and one nonapeptide as positive control from HIV-1 pol protein, were listed in Table 1. The mutated peptide Sur79L2, in which leucine replaced the natural histidine at position 2, had the higher HLA-A2-binding scores vs. Sur79. The peptide Sur95 with a BIMAS HLA-A2-binding score of 19 and a SYFPEITHI score of 5.28 was the intermediate-scoring peptide, which had been shown could stimulate HLA-A2-restricted CTL response in some previous studies [13, 14]. As shown in Table 1, these four peptides all had lower HLA-A24-binding scores using predicting software BIMAS and SYFPEITHI.
MHC stabilization assay showed that Sur79L2 had higher binding capacity to HLA-A2 molecule
The binding affinity of the four synthetic peptides to HLA-A2 was determined using the HLA-A2 TAP-deficient T2 lymphoma cell line, which could enhance HLA-A2 expression when exposed to exogenous HLA-A2-binding peptides. As shown in Fig. S1 and Table 1, the peptide Sur79L2 was capable of up-regulating HLA-A2 molecular expression and showed high affinity to HLA-A2 (FI = 6.02), whereas the wild-type peptide Sur79 had low affinities to HLA-A2 (FI = 0.46). The peptide Sur95 bound in comparison with intermediate affinity (FI = 1.63).
The expression levels of Survivin and HLA-A in target samples were detected by ELISA, RT-PCR and Western blot analysis
The expression of Survivin protein in ascites was detected by ELISA. The concentration of Sur protein reached to 53.7 ng/ml in ascites (data not shown), indicating high expression of this antigen in this patient. The expression of Sur mRNA and protein in all cell lines in this study was analyzed by RT-PCR and Western blot. The results demonstrated that the expression of Sur mRNA and protein could both be detected in HepG2, BEL-7402 and HuH-1 cancer cells (Fig. S2A, S2B). For the identification of the expression of HLA-A2 molecule in tumor cell line and TALs from the patient ascites, we used flow cytometry assay. The results showed that the expression of patient TALs, HepG2 and HuH-1 was 83.4, 82.4 or 81.3 %. Meanwhile, HLA-A2 expression in BEL-7402 was only 1.0 % (Fig. S3A). The same analysis was carried out to test the expression of HLA-A24 on the cell surface using HLA-A24 mAb. The results indicated that HepG2 and patient TALs were HLA-A24-positive cells rather than HuH-1 and BEL-7402 (Fig. S3B).
High-, intermediate- and low-frequency IFN-γ-producing T lymphocytes are detectable by ELISPOT assay when Survivin-derived epitope peptides are used
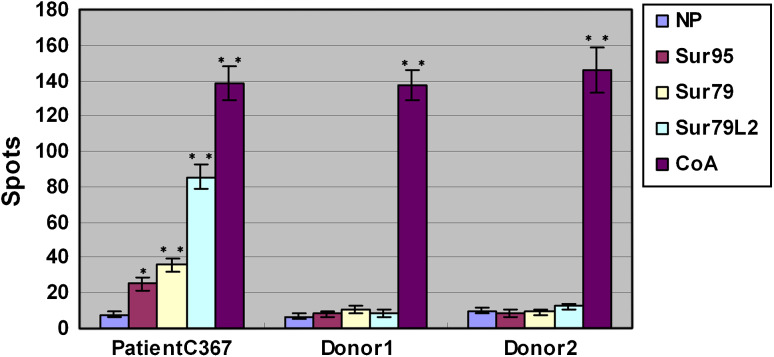
TALs from the ascites of patient C367 were stimulated with peptides in vitro for CTL priming and expansion before examination in the ELISPOT. This procedure would extend the sensitivity of the ELISPOT. Responses against the following Survivin peptides were examined: Sur95, Sur79 and Sur79L2. As shown in Fig. 1, the patient TALs responded strongly against the strong HLA-A2-binding peptide analog Sur79L2 (85.4 ± 6.88 peptide-specific spots/105cells) and elicit a response of lower intensity against Sur95 (25.2 ± 3.84 spots/105cells). Interestingly, the weak HLA-A2-binding peptide Sur79 also could generate a peptide-response of moderate intensity (35.8 ± 3.67 spots/105cells) in the patient. No response was detected against the negative peptide. This may imply that the peptide-binding capacity to HLA is not the only factor to determine its immunogenicity. PBLs from two healthy donors were analyzed to investigate whether a response against Survivin could be detected in healthy individuals. No response was observed in any of the controls against any of the Survivin deduced peptides.
Fig. 1.
IFN-γ-producing cells were enumerated by ELISPOT assay against peptide Sur95, peptide Sur79 and peptide Sur79L2. Each experiment was performed with 105 cells/well, and the average number of spots was calculated. Negative peptide (NP), HIVpol476, served as a negative control. Concanavalin (CoA) served as a positive control. Bar graphs indicates mean ± SD. *P < 0.05, **P < 0.01 compared to the NP group
Survivin-reactive CTLs can lyse peptide-pulsed T2 target cells
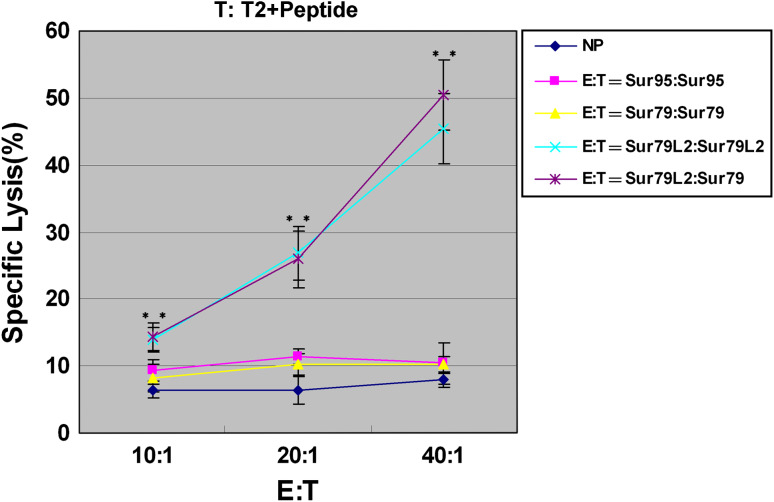
To characterize the functional capacity of Survivin-reactive CTLs, these peptides-stimulated TALs were isolated by means of magnetic beads coated with anti-CD8 antibody. Three days after isolation, their cytolytic activity by measuring their ability to serve as effector cells that kill peptide-pulsed T2 target cells expressing the HLA-A2 molecule was tested by the nonradioactive CTL assays (CytoTox96, Promega, Madison, WI). As shown in Fig. 2, Sur79L2-specific CD8+ T cell could efficiently lysed both the Sur79L2-pulsed and the Sur95-pulsed target T cells at an effector/target cell (E:T) ratio from 20:1 to 40:1. However, the CTLs induced by peptides Sur95 and Sur79 could not lyse the mentioned target cells. In some previous studies [13, 14], the Sur95-reactive CTL were often shown significant cytotoxicity in other tumor patient, which was distinguished from our study.
Fig. 2.
Specific lysis of CTLs generated from different Survivin-derived peptides against peptide-pulsed T2 cells. Immune effector cells were incubated with target cells at the effector/target ratios shown. CTL assays were performed following a 4-h incubation period. E represents peptide-induced CTLs effectors and T represents peptide-pulsed T2 target cells. Cytotoxic T lymphocytes generated from HIVpol476, served as a NP. Data points represent the mean ± SD as measured by quantitation of LDH release. **P < 0.01 compared to the NP group
Sur79L2-induced CTLs can produce Granzyme B and kill hepatoma cell lines
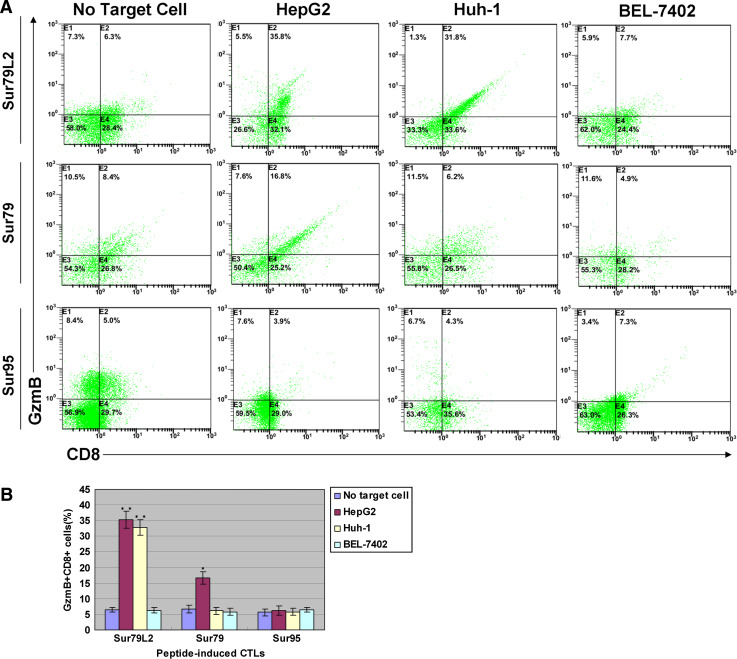
To further confirm whether mutated peptide Sur79L2-induced CTLs recognized Survivin-positive HLA-A2 target HCC cell lines, we tested the intracellular Granzyme B (GzmB) production in CD8+ T cell after incubation with different target cells for 4 h. As shown in Fig. 3, after incubated with HepG2 and HuH-1, the frequency of intracellular GzmB containing Sur79L2-specific CD8+ T cells increased to 35.4 ± 2.78 and 32.9 ± 2.49 % compared with the BEL-7402 incubation group (6.3 ± 0.91 %) and blank control group (6.4 ± 0.78 %). Analysis of Sur79-specific CTLs revealed that 16.6 ± 2.02 % CD8+ T cells expressed GzmB reacted with HepG2 and minimal numbers of T cells in HuH-1 and BEL-7402 incubation groups expressed GzmB. In contrast, the percentage of GzmB-positive CD8+ T cells were not observed significantly improved in Sur95-induced CTLs/tumor cell reaction groups.
Fig. 3.
FACS analysis for intracellular Granzyme B expression in peptide-induced CD8+ CTLs incubated with different target HCC cell lines for 4 h. The flow cytometric plots from the analysis of the response measured in a representative experiment are shown in a. The percentage value in upper right corner of each subpanel indicated the proportion of double positive cells (CD8+, GzmB+). The responses are measured in triplicate. The percentages of GzmB+ CD8+ cells for three peptide-induced CTLs against different HCC cell lines are shown in b. Bar graphs indicates mean ± SD. *P < 0.05, **P < 0.01 compared to the No target cell group
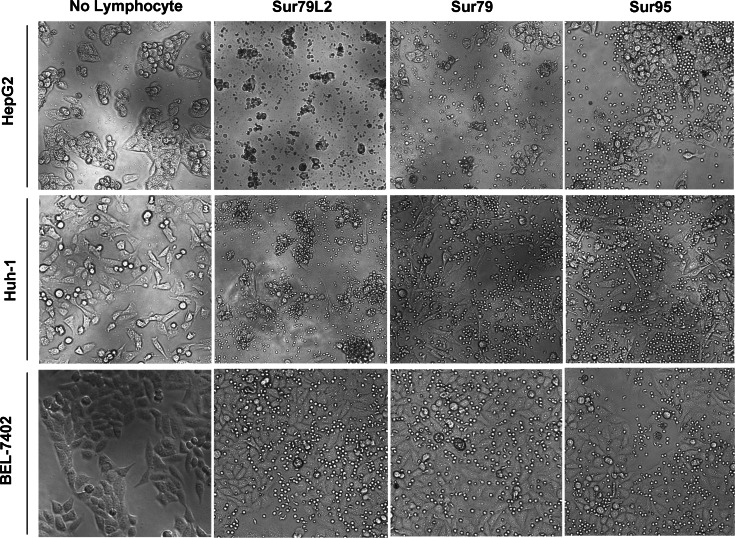
Similar results were observed in tumor cell morphological analysis after incubation. As shown in Fig. 4, Sur79L2-specific CTLs could effectively induce significant morphological change such as membrane blebbing, cell shrinkage and shedding in HepG2 and HuH-1 instead of BEL-7402. Similarly, the occurrence of morphological changes in part of the HepG2 could be also found in Sur79-specific CTLs action group. However, Sur95-specific CTLs failed to induce apparent morphological changes against tumor target cells.
Fig. 4.
Morphological analysis of the HCC cell lines after incubation with peptide-induced CTLs at 48 h. Cells were visualized under ×200 magnification
Antibody inhibition assay showed that Sur79-induced CTLs lyse hepatoma cell lines depend on HLA-A24-restricted manner
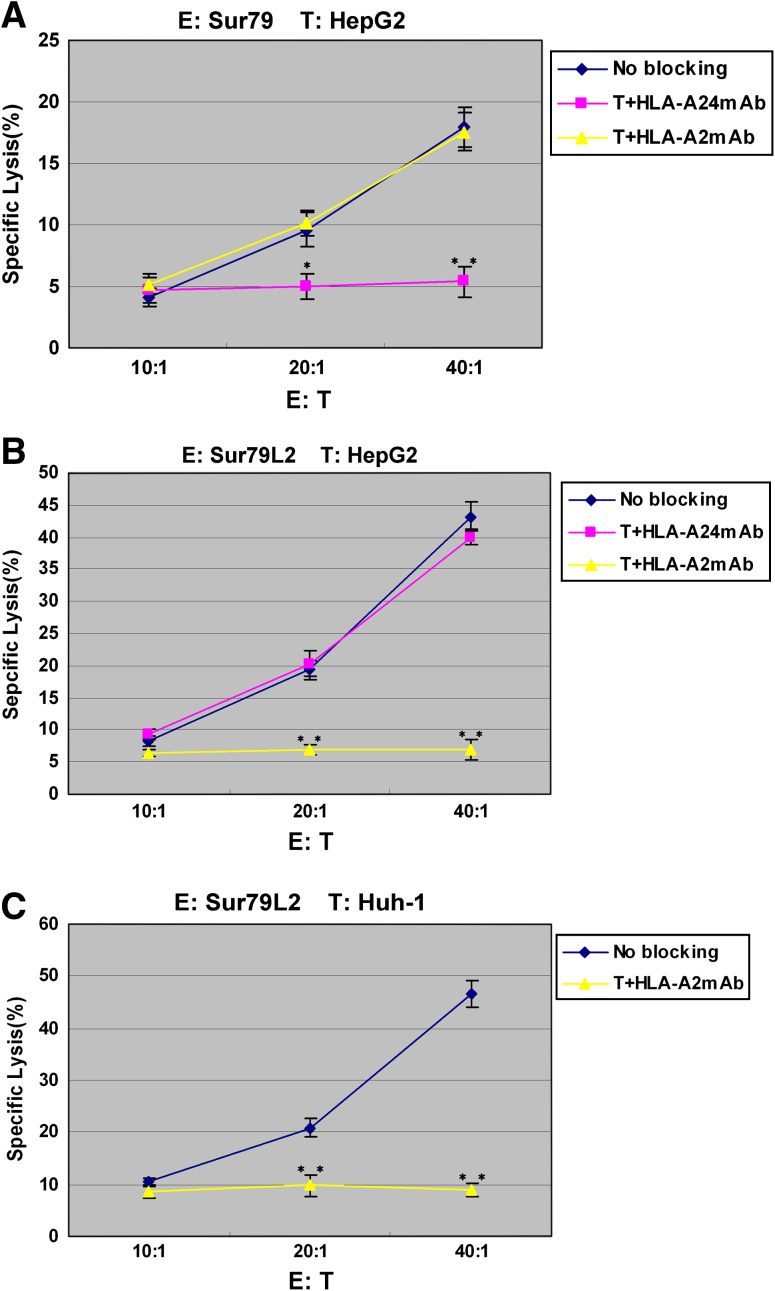
To investigate whether Sur79-specific CTLs killing HepG2 depend on HLA-A24-restricted manner, we performed antibody blocking experiments. In the HLA-A24 blocking experiment, anti-HLA-A24 mAb 22E1 markedly inhibited cytotoxic activity against HepG2 cells in CTL assay of CTLs generated from patient ascites by stimulation with the peptide Sur79 (Fig. 5a), but anti-HLA-A24 mAb did not inhibit the response of Sur79L2-specific CTLs (Fig. 5b). In the HLA-A2 blocking experiment, the results showed that after blocking the HLA-A2 sites on the surface of HepG2 and Huh-1 with HLA-A2 mAb, the specific killing effects of Sur79L2-specific CTLs could be significantly eliminated (Fig. 5b, c). However, HLA-A2 mAb could not eliminate the killing effect of Sur79-specific CTLs (Fig. 5a). These results clearly indicate that these Sur79-specific CTLs recognized HepG2 in a HLA-A24-restricted manner.
Fig. 5.
Anti-HLA-A mAb blocked the peptide-specific CTL reaction on target HCC cell lines. HCC target cells were incubated with or without anti-HLA-A2 and anti-HLA-A24 antibody for 1 h at 4 °C. Immune effector cells were incubated with target cells at the effector/target ratios shown. CTL assays were performed following a 4-h incubation period. Data points represent the mean ± SD as measured by quantitation of LDH release. a E: Sur79-specific CTLs, T: HepG2 cells. b E: Sur79L2-specific CTLs, T: HepG2 cells. c E: Sur79L2-specific CTLs, T: Huh-1 cells. *P < 0.05, **P < 0.01 compared to the No blocking group
Discussion
Vaccination for the prevention of infectious diseases has achieved great successes in medicine. Inspired by these achievements, a considerable number of researchers have been employed in an attempt to develop therapeutic cancer vaccines. Although the first cancer vaccine, Sipuleucel-T (Provenge) that is specific against prostate cancer, has received the FDA approval in 2010, in most of clinical trials to date, TAA peptide-based vaccines rarely eliminated tumors [2]. There are many hurdles we need to overcome, among which the selection of the appropriate target Ags appears critical. An on-going eight years of clinical trial involving HCC vaccine based on AFP has shown that although the AFP epitope peptide could induce antigen-specific T cell response in vitro, no potent anti-tumor immune response was observed in HCC volunteers. The possible explanation is that autologous AFP peptide only elicited relatively weak T cell responses in vivo due to expression of lower affinity TCR molecules on these T cells [20].
One current approach currently being investigated is the optimization of the MHC class I anchor residues in tumor epitopes using mutated TAA to enhance binding of the peptide to the MHC class I molecule. Compared with wild-type TAA, mutated TAA may has potential advantages, such as the following: (1) their T cell repertoire should not be deleted, and they should be recognized as nonself by the immune system eliciting stronger immune response, as is the case with viral Ags; and (2) their potential resistance to negative selection in case the mutated protein is essential for cell survival [3]. However, some of the studies revealed that not all the mutated peptides with higher binding capacity to MHC molecule are able to induce an effective anti-tumor response in vivo; the possible reason was that part of the mutated peptide-specific T cell lacks the ability to cross-recognize wild-type antigens [21, 22]. Therefore, the key questions remain as to how alterations of mutated anchor residues, while enhancing the peptide–MHC (pMHC) class I interaction, simultaneously increasing TCR cross-recognition on wild-type TAA.
Survivin has been considered to be a very attractive candidate for cancer immunotherapy in several previous studies because of its expression in a vast majority of human tumors rather than in differentiated normal cells. There have been some clinical trial results showed that partial cancer vaccines based on Survivin peptides could generate anti-tumor CTL response in vitro and lead to effective eradication of tumor cells in animal models, which were implicated with chronic lymphatic leukemia (CLL) [19], melanoma [13], breast [13], gastric [23], pancreatic [24], colon [14] and prostate [25] cancer, but not with primary HCC. This study was aimed to identify an appropriate mutated Survivin epitope that would be for vaccine design to boost specific CTL responses against HCC.
Based on our previous findings [26], three HLA-A2-restricted Survivin antigen epitopes, Sur79, Sur79L2 and Sur95, were selected in our study. Sur79L2 is a point-mutation nonapeptide derived from wild-type peptide Sur79 that the residue of histidine at positions 2 is replaced with Leucine. As a positive control for our experiment, Sur95 has been shown previously to be a T cell epitope eliciting specific CTLs in a variety of cancer patients. At first, we predicted the binding scores between the peptides and MHC class I molecules based on two classic computer algorithms, BIMAS and SYFPEITHI. The results showed that point-mutated peptide Sur79L2 owned the significantly higher MHC class I-binding ability than the other two wild-type peptides, which was consistent with the evaluation of HLA-A2-binding affinity using flow cytometric assay based on the use of the HLA-A-0201 TAP-deficient T2 lymphoma cell line. In fact, Sur95 peptide did only show an intermediate affinity for the HLA-A2 molecules.
To investigate whether Sur79L2 could induce Survivin-specific response in T lymphocytes of HCC patient, we measured the ability of peptides to induce IFN-γ-secreting cells separated from ascites samples of a HLA-A2-positive patient with advanced primary hepatic carcinoma. As shown in results of ELISPOT assay, Sur79L2 peptide was found to generate stronger peptide-specific T cell responses by virtue of their ability to induce increased frequencies of IFN-γ-producing T cells than the other peptides tested. To evaluate the CTL activity of the Sur79L2 peptide-specific T cells against HLA-A2-restricted target cells, which including T2 cells loaded with Sur79L2 and Sur79, in vitro cross-reorganization and cytotoxicity experiments were conducted. We found that both Sur79L2 and Sur79 could induce Survivin-specific CTL immune responses. Furthermore, this study demonstrated that the anti-HCC immunity to three HCC cell lines, HepG2, Huh-1 and BEL-7402 which are all the expression of wild-type Survivin antigen, induced by Sur79L2 peptide was Survivin-specific and HLA-A2-restricted.
Interestingly, although wild-type peptide Sur79 showed a lower affinity to the HLA-A2 molecules, certain IFN-γ-secreting T cells could be detected in TALs of HCC patient stimulated with Sur79. Moreover, after incubation with HepG2 liver cancer cells, Sur79-activated T cells could promote part of the target cells undergoing apoptosis or necrosis, which may be associated with the antigen-presenting in an HLA-A24 manner as the result of the positive expression of HLA-A24 molecules in HCC patient and HepG2 cell line. The software prediction score of Sur79-binding HLA-A24 was not high in our study, which was different from one previous research involved the actual binding capacity assay between Sur79 and HLA-A24 [27]. In order to understand the mechanism, antibody inhibition assay were carried out and the results confirmed our expectation.
Notably, Sur95 peptide has been considered to be an attractive epitope in eliciting Survivin-specific CTL response in a variety of malignant tumors, such as CLL [19], melanoma, breast cancer [13] and CRC [14]. However, no significant anti-tumor reactions were observed in this study using the Sur95-induced TALs derived from the patient with hepatic carcinoma, even though part of which could secret IFN-γ after sensation of Sur95. We assume that the inconsistent results may be caused by the difference of tumor types and the diversity of individual patient’s immune status. Preta et al. [28] had reported that some of colon-cancer cell lines were poorly recognized by Sur95-specific CTLs, which were implicated in the function of Serine-peptidase, tripeptidyl peptidase II (TPPII).
In summary, we show here that one novel Survivin deduced point-mutated epitope Sur79L2 is capable of inducing HLA-A2-restricted CD8+ CTLs in a HCC patient, which can efficiently cross-recognize and kill the target HCC cell lines expressing wild-type Survivin antigens. Our results suggest that Sur79L2 would serve as a good candidate to design the peptide-based anti-tumor vaccine for the treatment of Survivin-bearing HCC patients. Of course, we need to expand the number of HCC case and further validate the anti-HCC effects of Sur79L2. Meanwhile, a recent study have revealed that maintaining the amino acids at positions 2, 4, 6 and 9 of TAA nonapeptide and altering the remaining five amino acids may help create the stronger ligand for TAA-specific T cell repertoire [18]. Therefore, further optimization of the design Sur antigen epitope is also necessary.
It is important to note, however, that in clinical trials to date, the overwhelming majority of patients who received wild-type and variant TAA-peptide vaccines therapy for their underlying cancer did not exhibit objective evidence of cancer regression [2]. The most plausible reason is that the already existing immune dysfunction of cancer patients may not lead to enough TAA-specific T cells activated by the TAA peptide vaccines. In recent years, some applications of the antigen-specific TCR transgenic technology have been attempted for tumor therapy, which may bring a hope to solve this dilemma [29, 30]. Therefore, we are planning to identify the Sur79L2-specific TCR gene and evaluate the possibility for anti-HCC with the TCR gene-transferred PBLs.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Acknowledgments
The authors thank Dr. Yu Liu of Wuhan University for her helpful reviews and suggestions. This work was supported by the National Major Projects of Science and Technology of China (No. 2009ZX09103-708), National Natural Science Foundation of China (No. 31100664) and Natural Science Foundation of Guangdong Province (No. 10151022401000024).
Conflict of interest
The authors declare that they have no conflict of interest.
References
- 1.Finn RS. Development of molecularly targeted therapies in hepatocellular carcinoma: where do we go now? Clin Cancer Res. 2010;16:390–397. doi: 10.1158/1078-0432.CCR-09-2084. [DOI] [PubMed] [Google Scholar]
- 2.Klebanoff CA, Acquavella N, Yu Z, Restifo NP. Therapeutic cancer vaccines: are we there yet? Immunol Rev. 2011;239:27–44. doi: 10.1111/j.1600-065X.2010.00979.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Palucka K, Ueno H, Banchereau J. Recent developments in cancer vaccines. J Immunol. 2011;186:1325–1331. doi: 10.4049/jimmunol.0902539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Zerbini A, Pilli M, Soliani P, Ziegler S, Pelosi G, Orlandini A, Cavallo C, Uggeri J, Scandroglio R, Crafa P, Spagnoli GC, Ferrari C, Missale G. Ex vivo characterization of tumor-derived melanoma antigen encoding gene-specific CD8 + cells in patients with hepatocellular carcinoma. J Hepatol. 2004;40:102–109. doi: 10.1016/S0168-8278(03)00484-7. [DOI] [PubMed] [Google Scholar]
- 5.Shang XY, Chen HS, Zhang HG, Pang XW, Qiao H, Peng JR, Qin LL, Fei R, Mei MH, Leng XS, Gnjatic S, Ritter G, Simpson AJ, Old LJ, Chen WF. The spontaneous CD8 + T-cell response to HLA-A2-restricted NY-ESO-1b peptide in hepatocellular carcinoma patients. Clin Cancer Res. 2004;10:6946–6955. doi: 10.1158/1078-0432.CCR-04-0502. [DOI] [PubMed] [Google Scholar]
- 6.Butterfield LH, Ribas A, Dissette VB, Lee Y, Yang JQ, De la Rocha P, Duran SD, Hernandez J, Seja E, Potter DM, McBride WH, Finn R, Glaspy JA, Economou JS. A phase I/II trial testing immunization of hepatocellular carcinoma patients with dendritic cells pulsed with four alpha-fetoprotein peptides. Clin Cancer Res. 2006;12:2817–2825. doi: 10.1158/1078-0432.CCR-05-2856. [DOI] [PubMed] [Google Scholar]
- 7.Sah NK, Khan Z, Khan GJ, Bisen PS. Structural, functional and therapeutic biology of survivin. Cancer Lett. 2006;244:164–171. doi: 10.1016/j.canlet.2006.03.007. [DOI] [PubMed] [Google Scholar]
- 8.Yamamoto H, Ngan CY, Monden M. Cancer cells survive with surviving. Cancer Sci. 2008;99:1709–1714. doi: 10.1111/j.1349-7006.2008.00870.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mamori S, Asakura T, Ohkawa K, Tajiri H. Survivin expression in early hepatocellular carcinoma and post-treatment with anti-cancer drug under hypoxic culture condition. World J Gastroenterol. 2007;13:5306–5311. doi: 10.3748/wjg.v13.i40.5306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tsuruma T, Iwayama Y, Ohmura T, Katsuramaki T, Hata F, Furuhata T, Yamaguchi K, Kimura Y, Torigoe T, Toyota N, Yagihashi A, Hirohashi Y, Asanuma H, Shimozawa K, Okazaki M, Mizushima Y, Nomura N, Sato N, Hirata K. Clinical and immunological evaluation of anti-apoptosis protein, survivin-derived peptide vaccine in phase I clinical study for patients with advanced or recurrent breast cancer. J Transl Med. 2008;6:24. doi: 10.1186/1479-5876-6-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kameshima H, Tsuruma T, Torigoe T, Takahashi A, Hirohashi Y, Tamura Y, Tsukahara T, Ichimiya S, Kanaseki T, Iwayama Y, Sato N, Hirata K. Immunogenic enhancement and clinical effect by type-I interferon of anti-apoptotic protein, survivin-derived peptide vaccine, in advanced colorectal cancer patients. Cancer Sci. 2011;102:1181–1187. doi: 10.1111/j.1349-7006.2011.01918.x. [DOI] [PubMed] [Google Scholar]
- 12.Honma I, Kitamura H, Torigoe T, Takahashi A, Tanaka T, Sato E, Hirohashi Y, Masumori N, Tsukamoto T, Sato N. Phase I clinical study of anti-apoptosis protein survivin-derived peptide vaccination for patients with advanced or recurrent urothelial cancer. Cancer Immunol Immunother. 2009;58:1801–1807. doi: 10.1007/s00262-009-0691-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Andersen MH, Pedersen LO, Capeller B, Bröcker EB, Becker JC, thor Straten P. Spontaneous cytotoxic T-cell responses against survivin-derived MHC class I-restricted T-cell epitopes in situ as well as ex vivo in cancer patients. Cancer Res. 2001;61:5964–5968. [PubMed] [Google Scholar]
- 14.Casati C, Dalerba P, Rivoltini L, Gallino G, Deho P, Rini F, Belli F, Mezzanzanica D, Costa A, Andreola S, Leo E, Parmiani G, Castelli C. The apoptosis inhibitor protein survivin induces tumor-specific CD8 + and CD4 + T cells in colorectal cancer patients. Cancer Res. 2003;63:4507–4515. [PubMed] [Google Scholar]
- 15.Yu Z, Theoret MR, Touloukian CE, Surman DR, Garman SC, Feigenbaum L, Baxter TK, Baker BM, Restifo NP. Poor immunogenicity of a self/tumor antigen derives from peptide-MHC-I instability and is independent of tolerance. J Clin Invest. 2004;114:551–559. doi: 10.1172/JCI21695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.McMahan RH, McWilliams JA, Jordan KR, Dow SW, Wilson DB, Slansky JE. Relating TCR-peptide-MHC affinity to immunogenicity for the design of tumor vaccines. J Clin Invest. 2006;116:2543–2551. doi: 10.1172/JCI26936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Rizzuto GA, Merghoub T, Hirschhorn-Cymerman D, Liu C, Lesokhin AM, Sahawneh D, Zhong H, Panageas KS, Perales MA, Altan-Bonnet G, Wolchok JD, Houghton AN. Self-antigen-specific CD8 + T cell precursor frequency determines the quality of the antitumor immune response. J Exp Med. 2009;206:849–866. doi: 10.1084/jem.20081382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jordan KR, McMahan RH, Kemmler CB, Kappler JW, Slansky JE. Peptide vaccines prevent tumor growth by activating T cells that respond to native tumor antigens. Proc Natl Acad Sci USA. 2010;107:4652–4657. doi: 10.1073/pnas.0914879107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Andersen MH, Pedersen LO, Becker JC, Straten PT. Identification of a cytotoxic T lymphocyte response to the apoptosis inhibitor protein survivin in cancer patients. Cancer Res. 2001;61:869–872. [PubMed] [Google Scholar]
- 20.Butterfield LH. Recent advances in immunotherapy for hepatocellular cancer. Swiss Med Wkly. 2007;137:83–90. doi: 10.4414/smw.2006.11077. [DOI] [PubMed] [Google Scholar]
- 21.Colella TA, Bullock TN, Russell LB, Mullins DW, Overwijk WW, Luckey CJ, Pierce RA, Restifo NP, Engelhard VH. Self-tolerance to the murine homologue of a tyrosinase-derived melanoma antigen: implications for tumor immunotherapy. J Exp Med. 2000;91:1221–1232. doi: 10.1084/jem.191.7.1221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.McWilliams JA, Sullivan RT, Jordan KR, McMahan RH, Kemmler CB, McDuffie M, Slansky JE. Age-dependent tolerance to an endogenous tumor-associated antigen. Vaccine. 2008;26:1863–1873. doi: 10.1016/j.vaccine.2008.01.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Idenoue S, Hirohashi Y, Torigoe T, Sato Y, Tamura Y, Hariu H, Yamamoto M, Kurotaki T, Tsuruma T, Asanuma H, Kanaseki T, Ikeda H, Kashiwagi K, Okazaki M, Sasaki K, Sato T, Ohmura T, Hata F, Yamaguchi K, Hirata K, Sato N. A potent immunogenic general cancer vaccine that targets survivin, an inhibitor of apoptosis proteins. Clin Cancer Res. 2005;11:1474–1482. doi: 10.1158/1078-0432.CCR-03-0817. [DOI] [PubMed] [Google Scholar]
- 24.Wobser M, Keikavoussi P, Kunzmann V, Weininger M, Andersen MH, Becker JC. Complete remission of liver metastasis of pancreatic cancer under vaccination with a HLA-A2 restricted peptide derived from the universal tumor antigen surviving. Cancer Immunol Immunother. 2006;55:1294–1298. doi: 10.1007/s00262-005-0102-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Fuessel S, Meye A, Schmitz M, Zastrow S, Linné C, Richter K, Löbel B, Hakenberg OW, Hoelig K, Rieber EP, Wirth MP. Vaccination of hormone-refractory prostate cancer patients with peptide cocktail-loaded dendritic cells: results of a phase I clinical trial. Prostate. 2006;66:811–821. doi: 10.1002/pros.20404. [DOI] [PubMed] [Google Scholar]
- 26.Sun YC, Shen J, Shen H, Shao HW, Huang SL. Study of Survivin gene cloning and HLA-A2-restricted CTL epitope prediction in Hela cell line. Chin J Immunol. 2007;23:1044–1051. [Google Scholar]
- 27.Bachinsky MM, Guillen DE, Patel SR, Singleton J, Chen C, Soltis DA, Tussey LG. Mapping and binding analysis of peptides derived from the tumor-associated antigen survivin for eight HLA alleles. Cancer Immun. 2005;5:6. [PubMed] [Google Scholar]
- 28.Preta G, Marescotti D, Fortini C, Carcoforo P, Castelli C, Masucci M, Gavioli R. Inhibition of serine-peptidase activity enhances the generation of a survivin-derived HLA-A2-presented CTL epitope in colon-carcinoma cells. Scand J Immunol. 2008;68:579–588. doi: 10.1111/j.1365-3083.2008.02175.x. [DOI] [PubMed] [Google Scholar]
- 29.Morgan RA, Dudley ME, Wunderlich JR, Hughes MS, Yang JC, Sherry RM, Royal RE, Topalian SL, Kammula US, Restifo NP, Zheng Z, Nahvi A, de Vries CR, Rogers-Freezer LJ, Mavroukakis SA, Rosenberg SA. Cancer regression in patients after transfer of genetically engineered lymphocytes. Science. 2006;314:126–129. doi: 10.1126/science.1129003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Coccoris M, Straetemans T, Govers C, Lamers C, Sleijfer S, Debets R. T cell receptor (TCR) gene therapy to treat melanoma: lessons from clinical and preclinical studies. Expert Opin Biol Ther. 2010;10:547–562. doi: 10.1517/14712591003614756. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.