Abstract
Progression of chronic liver disease is precipitated by hepatocyte loss, inflammation and fibrosis. This process results in the loss of critical hepatic functions, increasing morbidity and the risk of infection. Medical interventions that treat complications of hepatic failure, including antibiotic administration for systemic infections and lactulose treatment for hepatic encephalopathy, can impact gut microbiome composition and metabolite production. Using shotgun metagenomic sequencing and targeted metabolomic analyses on 847 faecal samples from 262 patients with acute or chronic liver disease, we demonstrate that patients hospitalized for liver disease have reduced microbiome diversity and a paucity of bioactive metabolites, including short chain fatty acids and bile acid derivatives, that impact immune defenses and epithelial barrier integrity. We find that patients treated with the orally administered but non-absorbable disaccharide lactulose have increased densities of intestinal Bifidobacteria and reduced incidence of systemic infections and mortality. Bifidobacteria metabolize lactulose, produce high concentrations of acetate and acidify the gut lumen in humans and mice, which, in combination, can reduce the growth of antibiotic-resistant bacteria such as Vancomycin-resistant Enterococcus faecium in vitro. Our studies suggest that lactulose and Bifidobacteria serve as a synbiotic to reduce rates of infection in patients with severe liver disease.
INTRODUCTION
Chronic liver disease (CLD) is increasingly common due to high rates of metabolic syndrome and alcohol use1-4. Untreated, all etiologies of CLD converge on the final common endpoint of cirrhosis with similar complications. CLD is typically clinically silent until liver decompensation leads to development of ascites, variceal bleeding or hepatic encephalopathy (HE). Decompensating events are increasingly frequent as CLD progresses, resulting in death or the need for liver transplant5,6. Treatment of advanced CLD is largely supportive with limited ability to modify the overall clinical course. Gut microbial metabolism, including bacterial ammonia production, exacerbates HE in patients with decompensated cirrhosis, often leading to repeated antibiotic treatment. A current first line treatment for HE is the non-absorbable disaccharide lactulose7. The mechanism by which lactulose reduces serum ammonia levels is incompletely defined, and its potential role as a prebiotic that modifies microbiome compositions and metabolic activities remains controversial 8-13. Moreover, there are varying reports regarding lactulose’s impact on development of complications of CLD, including systemic infections14,15.
The gut microbiome impacts human health and a wide range of diseases. Given the bidirectional communication between the liver and the gut via the portal vein and biliary tree, it is postulated that the microbiome also plays a role in liver disease pathogenesis. Preclinical studies implicate the microbiome as a potential driver of non-alcoholic fatty liver disease (NAFLD) and alcoholic liver disease16-18. While robust clinical evidence linking the microbiome to progression of liver disease is lacking, multiple studies have reported gut microbiome “signatures” of advanced fibrosis and cirrhosis9,19-22. Observational studies have also associated gut microbiome compositions with complications of advanced CLD9,17,23,24. Different taxa are implicated in these studies, and, consistent with studies in non-hepatic diseases, patients with higher burdens of potentially pathogenic taxa (e.g. Proteobacteria or Enterococcus) and lower prevalence of obligate anaerobic commensals (e.g. Lachnospiraceae and Oscillospiraceae) generally have poor prognoses25-27.
The consequences of microbiome compositional differences on production of microbiome-derived metabolites remain incompletely defined. Microbe-derived metabolites contribute to intestinal epithelial cell differentiation and barrier formation28-30 and regulate mucosal innate and adaptive immune defenses27,31,32. In the case of liver disease, fecal bile acid (BA) profiles have been correlated with progression of NAFLD to non-alcoholic steatohepatitis (NASH) and subsequently advanced fibrosis33-35. Additionally, both total serum BA and specific immunomodulatory circulating BA have been implicated in the progression and prognosis of liver disease36-38. While recent studies have identified bacterial species that generate health promoting metabolites, whether these reduce or enhance progression of CLD remains largely unexplored.
To better understand the role of the gut microbiome in the development of CLD complications, including infections, we performed a single center observational cross-sectional cohort study of hospitalized patients with liver disease. Metagenomic and metabolomic profiles of fecal samples were correlated with liver disease outcomes. We demonstrate that the commonly prescribed disaccharide lactulose drives expansion of Bifidobacteria and, in the absence of systemic antibiotic administration, results in a protective fecal metabolome. Bifidobacteria expansion associates with decreased abundance of antibiotic-resistant pathobionts and improved patient outcomes, including reduced incidence of systemic infections and prolonged survival. Our study provides insights into the mechanism by which lactulose impacts outcomes of CLD patients and provides rationale for optimizing gut microbiome compositions and functions to minimize complications of liver disease.
RESULTS
Patient recruitment and disease characteristics
We enrolled 356 hospitalized patients with liver disease. Of these 356 patients, 262 (73.6%) produced 847 stool samples that were analyzed by shotgun metagenomic sequencing and paired targeted metabolomics. Demographics and admission disease characteristics of patients who produced samples are shown (Table 1). Most patients enrolled were hospitalized with decompensated cirrhosis (n = 196, 74.8%), and of those, alcohol use was the most common liver disease etiology (n = 124, 63.3%). The cohort had a median model of end stage liver disease MELD-sodium (MELD-Na) score of 18.69, which was higher in patients with decompensated cirrhosis compared to other disease states. Of the 262 patients with samples, 183 (69.8%) had clinically significant portal hypertension, which was more common in patients with decompensated cirrhosis. Over half of enrolled patients had end organ dysfunction on admission, which was defined using NACSELD criteria39. Acute on chronic liver failure (ACLF) (i.e. 2 or more organ failures) was present in 14.7% of patients with decompensated cirrhosis upon enrollment. End organ dysfunction was more prevalent in patients with decompensated cirrhosis, but only severe HE was statistically significant. Consistent with the poor prognosis suggested by enrollment disease characteristics, patients with decompensated cirrhosis had a 20.9% 90-day mortality rate.
TABLE 1:
Patient demographics and baseline disease characteristics of all patients that produced at least one stool sample. Patients are stratified by liver disease chronicity at the time of consent. Clinically significant portal hypertension was defined by hepatic venous pressure gradient (HVPG) ≥ 10mmHg, characteristic imaging (enlarged portal vein, intraabdominal varices, splenomegaly, or ascites), or clinical features (ascites, varices, or hepatic encephalopathy).
| Acute | Chronic | Cirrhosis (Compensated) | Cirrhosis (Decompensated) | p | ||
|---|---|---|---|---|---|---|
| n | 35 | 18 | 13 | 196 | ||
| Basic Demographics | Age (median [IQR]) | 57.30 [34.00, 67.25] | 60.20 [48.57, 64.60] | 63.10 [54.90, 69.70] | 58.05 [47.62, 65.93] | 0.413 |
| Sex = Male (%) | 12 ( 34.3) | 10 ( 55.6) | 9 ( 69.2) | 123 ( 62.8) | 0.014 | |
| Race (%) | 0.894 | |||||
| African American | 10 ( 28.6) | 8 ( 44.4) | 4 ( 30.8) | 64 ( 32.7) | ||
| Asian / Pacific Islander | 3 ( 8.6) | 1 ( 5.6) | 1 ( 7.7) | 7 ( 3.6) | ||
| Caucasian | 17 ( 48.6) | 6 ( 33.3) | 7 ( 53.8) | 95 ( 48.5) | ||
| Hispanic | 5 ( 14.3) | 2 ( 11.1) | 1 ( 7.7) | 26 ( 13.3) | ||
| Other | 0 ( 0.0) | 1 ( 5.6) | 0 ( 0.0) | 4 ( 2.0) | ||
| BMI (median [IQR]) | 25.28 [21.61, 28.50] | 26.61 [21.84, 32.33] | 30.67 [26.75, 34.17] | 27.99 [23.62, 33.04] | 0.059 | |
| Disease Etiology | Liver Disease Etiology (%) | <0.001 | ||||
| AIH | 5 ( 14.3) | 0 ( 0.0) | 1 ( 7.7) | 6 ( 3.1) | ||
| Alcohol | 5 ( 14.3) | 1 ( 5.6) | 3 ( 23.1) | 124 ( 63.3) | ||
| Checkpoint Blockade Hepatitis | 4 ( 11.4) | 0 ( 0.0) | 0 ( 0.0) | 1 ( 0.5) | ||
| DILI | 6 ( 17.1) | 0 ( 0.0) | 0 ( 0.0) | 0 ( 0.0) | ||
| HBV | 0 ( 0.0) | 0 ( 0.0) | 0 ( 0.0) | 3 ( 1.5) | ||
| HCV | 0 ( 0.0) | 0 ( 0.0) | 1 ( 7.7) | 15 ( 7.7) | ||
| Metabolic | 0 ( 0.0) | 0 ( 0.0) | 0 ( 0.0) | 4 ( 2.0) | ||
| NASH | 0 ( 0.0) | 1 ( 5.6) | 3 ( 23.1) | 23 ( 11.7) | ||
| Other | 7 ( 20.0) | 3 ( 16.7) | 2 ( 15.4) | 6 ( 3.1) | ||
| PBC | 0 ( 0.0) | 0 ( 0.0) | 0 ( 0.0) | 3 ( 1.5) | ||
| PSC | 0 ( 0.0) | 1 ( 5.6) | 0 ( 0.0) | 4 ( 2.0) | ||
| Vascular | 8 ( 22.9) | 12 ( 66.7) | 3 ( 23.1) | 7 ( 3.6) | ||
| Disease Severity | MELD-Na at consent (median [IQR]) | 10.93 [5.30, 15.47] | 10.40 [2.56, 20.19] | 13.06 [4.74, 19.98] | 21.30 [13.46, 31.92] | <0.001 |
| Clinically Significant Portal HTN (%) | 2 ( 5.7) | 2 ( 11.1) | 5 ( 38.5) | 174 ( 88.8) | <0.001 | |
| HCC (%) | 1 ( 2.9) | 1 ( 5.6) | 3 ( 23.1) | 22 ( 11.2) | 0.171 | |
| NACSELD-ACLF at Consent (%) | 0 (0.0) | 0 (0.0) | 0 (0.0) | 29 ( 14.8) | 0.012 | |
| Hepatic Enceaphalopathy (%) | 2 ( 5.7) | 1 ( 5.6) | 0 (0.0) | 56 ( 28.6) | 0.001 | |
| Shock (%) | 2 ( 5.7) | 3 ( 16.7) | 1 ( 7.7) | 26 ( 13.3) | 0.54 | |
| Respiratory Failure (%) | 1 ( 2.9) | 1 ( 5.6) | 1 ( 7.7) | 25 ( 12.8) | 0.29 | |
| Renal Failure (%) | 1 ( 2.9) | 0 ( 0.0) | 0 ( 0.0) | 24 ( 12.2) | 0.081 | |
| Medications and Diet | Lactulose | |||||
| PTA (%) | 1 ( 2.9) | 2 ( 11.1) | 0 ( 0.0) | 85 ( 43.6) | <0.001 | |
| Inpatient (%) | 7 ( 20.0) | 5 ( 27.8) | 5 ( 38.5) | 148 ( 75.9) | <0.001 | |
| Rifaximin | ||||||
| PTA (%) | 0 ( 0.0) | 1 ( 5.6) | 0 ( 0.0) | 60 ( 30.8) | <0.001 | |
| Inpatient (%) | 5 ( 14.3) | 1 ( 5.6) | 2 ( 15.4) | 103 ( 52.8) | <0.001 | |
| Broad Antibiotics (non-rifaximin) | ||||||
| PTA (%) | 5 ( 14.3) | 3 ( 16.7) | 1 ( 7.7) | 52 ( 26.7) | 0.169 | |
| Inpatient (%) | 10 ( 28.6) | 4 ( 22.2) | 5 ( 38.5) | 80 ( 41.0) | 0.263 | |
| Diet at time of consent (%) | 0.52 | |||||
| Diet Order | 31 ( 88.6) | 15 ( 83.3) | 13 (100.0) | 155 ( 79.5) | ||
| NPO | 4 ( 11.4) | 3 ( 16.7) | 0 ( 0.0) | 37 ( 19.0) | ||
| Tube Feeds | 0 ( 0.0) | 0 ( 0.0) | 0 ( 0.0) | 3 ( 1.5) | ||
| Disease Complications | SBP (%) | 0 ( 0.0) | 1 ( 5.6) | 0 ( 0.0) | 20 ( 10.2) | 0.133 |
| Bacteremia (%) | 2 ( 5.7) | 2 ( 11.1) | 0 (0.0) | 15 ( 7.7) | 0.663 | |
| 90-day Survival (%) | 33 ( 94.3) | 18 ( 100.0) | 12 ( 92.3) | 155 ( 79.1) | 0.02 |
Microbiome and metabolite profiles of liver disease patients
To determine the fecal microbiome compositions in patients with liver disease, we performed shotgun sequencing on DNA from 847 fecal samples collected from 262 liver disease patients and 22 healthy donors (median age 35.00 years-old, IQR 25.75 - 42.50). We used MetaPhlan4 to assign taxonomic compositions and inverse Simpson to assess microbial diversity (Figure 1A). Most fecal samples have reduced taxonomic alpha-diversity (Figure 1A, inverse Simpson range: 1.00 – 37.55, mean = 6.35, median = 3.85) with expansion of bacterial taxa belonging to the Enterococcus genus and Enterobacteriaceae family, common hallmarks of dysbiosis associated with antibiotic exposure25,40. Pathobiont expansion was detected in fecal samples across all alpha-diversity tertiles in patients with liver disease compared to healthy controls (Figure 1A, B). Marked expansion of the genus Bifidobacterium (Figure 1A, 1B) was also detected in a subset of samples across alpha-diversity groups. While Bifidobacterium expansion is common in breast fed infants, it is rarely detected in adults, suggesting that this may be liver disease-specific.
Figure 1: Fecal samples from hospitalized patients with liver disease display a wide range of microbiome and metabolomic profiles.
A total of 847 fecal samples from hospitalized patients with liver disease and 22 healthy donor fecal samples were analyzed by shotgun metagenomics and targeted metabolomics. (A) Relative taxa abundance by shotgun metagenomics is shown for each sample. Metagenomic alpha-diversity was quantified using the inverse Simpson metric, and samples are arranged from left to right in order of increasing metagenomic alpha-diversity. Samples from patients with liver disease were categorized as either low, medium, or high alpha-diversity based on tertiles of inverse Simpson levels. Quantitative targeted metabolite concentrations for each corresponding sample are shown below the metagenomic data. (B) Relative abundance of indicated taxa and (C) select metabolite concentrations were plotted for each tertile of alpha-diversity. Units for SCFA are mM, and units for BA derivatives are in μg/mL. For panels B and C, each point represents a single value where n = 283 (low diversity), 282 (medium diversity), 282 (high diversity), and 22 (healthy donor). Median and interquartile range are indicated by the horizontal line and box, respectively. The lower vertical line depicts Q1 – 1.5*IQR and the upper vertical line depicts Q3 + 1.5*IQR. Statistical comparisons between individual groups were analyzed using a two-tailed Wilcoxon rank sum test. Individual groups were compared to the healthy donor group. using the Benjamini-Hochberg procedure and represented as follows: *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001 *.
While microbiome-derived or modified metabolites mediate many beneficial impacts on mucosal immune defenses and epithelial barrier functions, little is known about their production in liver disease patients harboring vastly different microbial populations. We used targeted GC- and LC-MS to determine relative amounts of 82 metabolites in our dataset (Figure E1). Ranking fecal samples by alpha-diversity demonstrates a correlation between microbiome diversity and relative amounts of short chain fatty acids (SCFAs), branched chain FAs, aminated FAs, secondary BA, and various indole compounds and tryptophan metabolites. As expected, low diversity fecal samples have markedly reduced secondary BA levels but also reduced concentrations of SCFAs and a subset of indole compounds.
We next quantified concentrations of SCFAs and primary and secondary BA in fecal samples from patients with liver disease. Higher proportions of commensal anaerobes and alpha-diversity coincided with higher concentrations of SCFA, secondary and modified secondary BA and reduced concentrations of conjugated primary BA (Figure 1A-C). While high diversity fecal samples from patients with liver disease had microbial taxonomic and metabolomic profiles that approached those seen in healthy donors, there was reduced representation of Lachnospiraceae and Oscillospiraceae taxa, SCFA production, and secondary BA synthesis (Figure 1B, C). Thus, while the majority of patients hospitalized with liver disease have reduced microbiome diversity and microbiome-derived metabolite concentrations, our findings demonstrate a steep gradient extending from relatively minor to absolute loss of microbiome compositions and health-associated metabolites.
Fecal metabolites correlate with distinct microbial taxa
To associate fecal metabolite concentrations with microbiome compositions, fecal samples were plotted on a taxonomic Uniform Manifold Approximation and Projection (taxUMAP) and assigned to one of twelve taxonomic groups based on the most prevalent taxons (Fig E2A). Samples dominated by Enterococci, Proteobacteria, Actinobacteria (predominantly Bifidobacteria) and Lactobacilli form distinct clusters on the taxUMAP. Overlaying fecal metabolite concentrations demonstrates that Enterococcus, Proteobacteria and Lactobacillus clusters have lower SCFA concentrations compared to clusters dominated by Bifidobacteria, Lachnospiraceae or Bacteroidetes (Fig E2A-D, H-J). Bifidobacteria-expanded samples had higher acetate concentrations (Fig E2B,H) but similar concentrations of butyrate (Fig E2C,I) and propionate (Fig E2D, J) compared to samples with high Bacteroidetes or Lachnospiraceae abundance.
While all taxonomic clusters had detectable concentrations of cholic acid, a primary BA, the highest concentrations were detected in Enterococcus dominated samples, likely reflecting the ability of bile salt hydrolase (BSH) expressing Enterococci to deconjugate glycocholic and taurocholic acid (Fig E2M). Conjugated primary BA concentrations were lower (Fig E2E,K, L) and secondary (Fig E2F,N,O) and modified secondary BA (Fig E2G,P,R) concentrations were higher in samples with high levels of obligate anaerobes. Compared to Bacteroidetes- and Lachnospiraceae-dominated samples, Bifidobacteria-expanded samples had higher unconjugated primary BA concentrations, likely reflecting Bifidobacteria-associated BSH activity, and modestly reduced secondary BA concentrations. Conversion of primary BA to secondary BA and subsequent modifications are mediated by a subset of bacterial species in the Lachnospiraceae family and Bacteroidales class, which are present in Bifidobacteria-dominated samples but largely absent in pathobiont-dominated samples (i.e. those dominated by Enterococcus or Proteobacteria). Compared to pathobiont dominated samples, Bifidobacteria-dominated samples mediate the full range of BA metabolism, albeit producing slightly lower concentrations of secondary BAs. Our results indicate that a subset of patients with liver disease have expanded populations of Bifidobacteria in their GI tract, resulting in increased acetate concentrations and enhanced bile salt deconjugation while retaining only a modest ability to generate secondary BA.
Lactulose enhances Bifidobacterium expansion
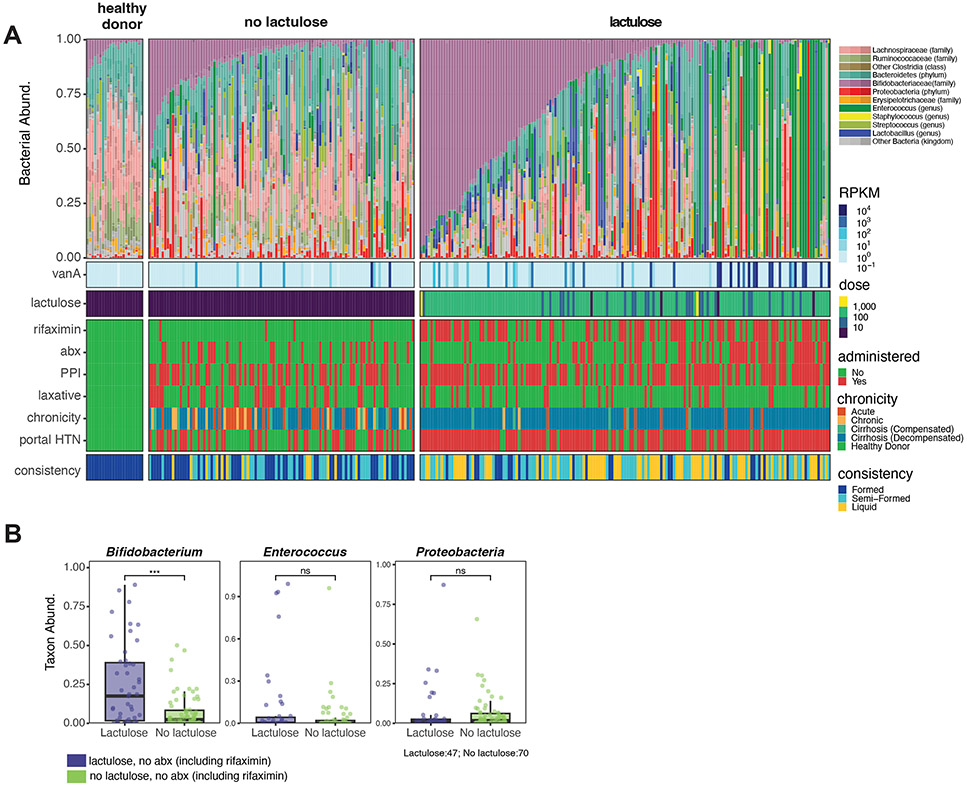
Lactulose is reported to have prebiotic activity that may impact microbiome compositions, but its impact on specific bacterial species and their production of metabolites remains poorly defined. To test whether lactulose is a driver of Bifidobacterium expansion in liver disease patients, we ranked one fecal sample from each of the 262 patients by Bifidobacterium abundance and stratified patients by whether or not they had received lactulose within 7 days prior to sample collection. Fecal samples from lactulose treated patients were taxonomically diverse, with 45.2% having ≥10% Bifidobacterium abundance (Fig 2A; range: 0.0 – 94.7%, mean: 22.2%, stdev: 28.4%). In contrast, fecal samples from patients not treated with lactulose had significantly lower Bifidobacterium abundance (Fig 2A; range: 0.0 – 50.1%, mean: 6.8%, stdev: 10.4%, padj = 0.009). High abundances (≥10%) of potential pathobionts including Enterobacteriaceae and Enterococcus were detected in lactulose treated and untreated patients (Fig 2A). Enterococcus expansion was restricted to fecal samples with low Bifidobacterium abundance and was associated with the presence of the vancomycin-resistance gene vanA, most strikingly in samples from lactulose treated patients (Fig 2A). These results suggest that lactulose-mediated Bifidobacterium expansion may limit intestinal Vancomycin-resistant Enterococcus (VRE) expansion. Treatment with broad-spectrum antibiotics likely reduces the density of Bifidobacterium species, potentially enabling VRE to benefit from lactulose treatment (Fig 2A)25,41. In support of this hypothesis, logistic regression demonstrates that lactulose administration positively associates with both Bifidobacterium and Enterococcus abundance (Table S1). Conversely, broad spectrum antibiotic administration negatively associates with Bifidobacterium abundance but positively associates with Enterococcus abundance (Table S1). Although the osmotic laxative effect of lactulose has been suggested to impact fecal microbiome compositions, we did not detect an association between stool consistency and Bifidobacteria abundance (Figure 2A, Table S1).
Figure 2: Lactulose use is associated with increased Bifidobacteria species abundance and reduced VRE abundance in the absence of systemic antibiotic use.
(A) Relative abundance by shotgun taxonomy of one sample per subject is shown (n = 22 healthy donors, 262 patients with liver disease). The sample with highest Bifidobacteria abundance for each subject was analyzed. Samples were arranged first by whether they were obtained without lactulose (left) or within 7 days after lactulose administration (right) and then by decreasing Bifidobacterium relative abundance. Among the Bifidobacteriaceae family, a small number of non-Bifidobacteria members are part of the Actinobacteria phylum and are also shown in shades of purple. The antibiotic resistance (vanA) gene was queried and colored based on expression level normalized to gene length (RPKM). Cumulative oral lactulose dose (grams) for 7 days prior to sample collection is shown. Rifaximin, broad-spectrum antibiotics, proton pump inhibitors (PPI), and alternative laxatives exposure prior to sample collection is shown. Red = treatment was administered within 7 days prior to the sample being collected; green = treatment not given in this period. The chronicity of liver disease and whether a patient had clinically significant portal hypertension (green = no, red = yes) at time of consent is shown. Stool consistency was categorized as either liquid (yellow), semi-formed (turquoise), or formed (royal blue) for each sample. (B) Relative abundances of Bifidobacteria, Enterococcus, and Proteobacteria were quantified for samples from patients that were not exposed to antibiotics (including rifaximin) in the 7 preceding days. Samples were stratified based on lactulose exposure. One sample per patient was analyzed, and each dot represents a single value (n=47 (lactulose exposure), n=70 (no lactulose exposure). Median and interquartile range are indicated by the horizontal line and box, respectively. The lower vertical line depicts Q1 – 1.5*IQR and the upper vertical line depicts Q3 + 1.5*IQR. Statistical comparisons between groups were analyzed using a two-tailed Wilcoxon rank sum test. P-values are adjusted for multiple comparisons using the Benjamini-Hochberg procedure and represented as: *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001.
To exclude the potentially dramatic impact of antibiotics on lactulose-associated Bifidobacterium expansion, we assessed the relative abundance of select taxa in samples obtained from patients who had not received antibiotics in the preceding 7 days (Figure 2B). In this antibiotic-naïve cohort, Bifidobacteria density was significantly increased in fecal samples obtained from lactulose-treated patients while Enterococcus and Proteobacteria densities were similar between groups (Figure 2B).
Lactulose promotes bioactive metabolite production
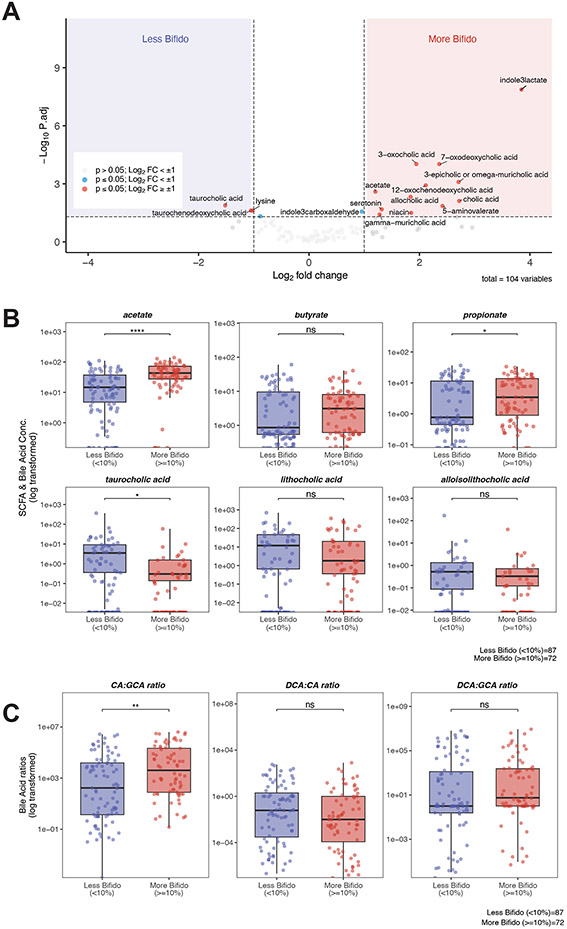
Bifidobacterium expansion, defined as relative abundance ≥10%, was associated with marked changes in fecal metabolite profiles in fecal samples from patients receiving lactulose (Fig 3, E3). In patients with Bifidobacterium expansion, the fecal metabolome was characterized by increased fatty acid concentrations (including acetate), reduced conjugated primary BA concentrations (taurocholic and taurochenodeoxycholic acids), increased secondary and modified secondary BA (3-oxo-cholic acid, 7-oxo-deoxycholic), and indole metabolites (indole-3-lactic acid and to a lesser extent, indole-3-carboxaldehyde) (Fig 3A,B). Consistent with a known property of Bifidobacterium to hydrolyze conjugated primary BA, ratios of unconjugated to conjugated primary BA were substantially higher in fecal samples with Bifidobacterium expansion (Fig 3C). In patients not receiving lactulose, fecal samples with ≥10% Bifidobacteria had only modestly increased indole-3-lactic acid levels (Figure E3). These results suggest that lactulose impacts the metabolic activity of the gut microbiota in liver disease patients (Fig 3 and E3).
Figure 3: Lactulose-mediated Bifidobacteria expansion is associated with significant changes in bioactive fecal metabolites.
(A) Volcano plot (log2fold change vs. log10p-value) of qualitative metabolites comparing samples with low (< 10%) vs. high (≥10%) Bifidobacteria abundance after lactulose exposure. P-values were calculated using a two-tailed Wilcoxon rank sum test and are corrected for multiple comparisons using the Benjamini-Hochberg procedure. Values with log2 fold-change > 1 (corresponding to a 2-fold change with a p-value < 0.05) were considered significant. (B) select SCFA and BA were quantified. Units for SCFA are mM, and units for BA derivatives are in μg/mL. (C) BA conversion from conjugated-primary BA to primary BA and then to secondary BAs was tested for each sample. Each point represents a molar ratio for an individual sample. For all comparisons (A-C), there is one sample per patient that was chosen based on the highest relative abundance of Bifidobacteria, and sample size was n=87 (lactulose exposure <10% Bifidobacteria) and n=72 (lactulose exposure with ≥10% Bifidobacteria). For panels B and C, each point represents a single sample. Median and interquartile range are indicated by the horizontal line and box, respectively. The lower vertical line depicts Q1 – 1.5*IQR and the upper vertical line depicts Q3 + 1.5*IQR. Statistical comparisons between individual groups were analyzed using a two-tailed Wilcoxon rank sum test. P-values are adjusted for multiple comparisons using the Benjamini-Hochberg procedure and represented as follows: *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001. For both panels B and C, Samples were grouped by whether they had expanded Bifidobacteria in the presence of lactulose. Median and interquartile range are indicated by the line and box, respectively. The lower vertical line depicts Q1 – 1.5*IQR and the upper vertical line depicts Q3 + 1.5*IQR. CA: cholic acid; GCA: glycocholic acid; DCA: deoxycholic acid.
Lactulose enhances B. longum-mediated VRE inhibition
Many fecal samples obtained from lactulose-treated patients had high abundances of pathobionts belonging to Enterobacteriaceae family (range: 0.0 – 98.8%, mean: 7.9%, median: 0.9%, stdev: 17.7%) and the Enterococcus genus (range: 0.0 – 100%, mean: 19.4%, median: 0.9%, stdev: 34.4%), with higher densities than those seen in fecal samples taken from patients not receiving lactulose (Proteobacteria mean: 5.3%, median: 1.3%; Enterococcus mean: 5.0%, median: 0.05%). While high abundances of Proteobacteria species were detected across the range of Bifidobacterium abundances, Enterococcus domination was detected more commonly in fecal samples with reduced to absent Bifidobacterium abundance (Fig 2A, 4A).
Figure 4: Lactulose-mediated Bifidobacteria expansion is associated with exclusion of antibiotic-resistant Enterococcus species.
(A) Relative abundance of potentially pathogenic taxa Enterococcus and Proteobacteria were plotted based on lactulose exposure (no lactulose, left; lactulose, right) and relative abundance of Bifidobacteria (< 10%, blue; ≥10%, red). Each point represents a single sample with the following sample sizes: total n = 262; no lactulose, < 10%, n=79; no lactulose, ≥10%, n=24; lactulose, < 10%, n=87; lactulose, ≥10%, n=72. Median and interquartile range are indicated by the horizontal line and box, respectively. The lower vertical line depicts Q1 – 1.5*IQR and the upper vertical line depicts Q3 + 1.5*IQR. Statistical comparisons between individual groups were analyzed using a two-tailed Wilcoxon rank sum test. P-values are adjusted for multiple comparisons using the Benjamini-Hochberg procedure and represented as follows: *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001. (B) A human-derived B. longum species was grown in BHIS media (blue) or BHIS supplemented with 50mM lactulose (red) or 50mM sucrose (green), and growth curves (OD600 over time) are shown. The growth curve shows mean +/− standard deviation for an experiment done in triplicate. This graph is representative of three independent experiments, also done in triplicate, all with consistent results. (C) Schematic of experimental design for B. longum (purple bacilli) and VRE (green cocci) co-culture. B. longum and VRE were grown to steady state and diluted to low density (OD600 = 0.05) prior to inoculation in either BHIS or BHIS supplemented with 50mM lactulose. Cultures were inoculated with both bacteria simultaneously (top) or B. longum was given either a 24-hour or 48-hour lead time prior to VRE inoculation (bottom). (D) After 24-hours of co-culture, serial dilutions of three replicates were plated for VRE c.f.u. counts. Each replicate is plotted as an individual point, and the median is represented by the horizontal line. *, p < 0.05 two-tailed student’s t-test. The plot is from one experiment that is representative of three independent experiments done in triplicate all with consistent results.
To investigate the inverse relationship between Bifidobacterium and VRE abundance, we cultured and whole genome sequenced a healthy donor-derived Bifidobacterium longum strain that encodes genes required for lactulose metabolism and acetate production42,43. B. longum culture growth was augmented by lactulose and to a lesser extent, sucrose (Fig 4B). Lactulose-induced growth was accompanied by media acidification and acetate production (Fig E4A,B). B. longum monocultures efficiently deconjugated taurocholic and glycocholic acid but, as expected, did not convert cholic acid to deoxycholic acid (Fig E4C,D).
We next colonized germ-free (GF) mice with B. longum and treated mice with or without lactulose (Fig E4E). Lactulose was detectable in the stool of GF mice and led to increased stool water content (Fig E4F,G). B. longum efficiently colonized and persisted in the intestines of gnotobiotic mice independent of lactulose (Fig E4H). B. longum-colonized mice had measurable fecal acetate levels but did not generate butyrate or propionate (Fig E4I-K). Lactulose treatment for 10 days resulted in elevated acetate and unconjugated primary BA concentrations and reduced concentrations of conjugated BA concentrations in B. longum colonized mice (Fig E4I,L,M). We next colonized GF mice with more complex, 8-11 strain consortia that contain B. longum (Fig E5 and Table S2). With all consortia, lactulose administration increased the relative abundance of B. longum (Fig E5B,C). Lactulose reduced fecal taurocholic acid concentrations in mice colonized with all three consortia, potentially a result of enhanced B. longum-mediated BA deconjugation (Fig E5B,C). Lactulose administration only modestly increased fecal acetate concentrations, possibly resulting from acetate consumption by other members of the administered consortia (Fig. E5B,C).
To test whether B. longum inhibits VRE growth and to determine the impact of lactulose administration, we performed co-culture assays. Simultaneous inoculation of B. longum and VRE into media did not reduce VRE growth, but pre-culture of B. longum for 24h or 48h fully inhibited VRE growth in the presence of lactulose and only partially inhibited VRE growth in the absence of lactulose (Fig 4C-D). Serial dilutions of B. longum culture supernatant with fresh media sequentially reduced the inhibitory effect on VRE growth (Fig E6A, B). Alkalization of B. longum supernatant pH from 5.8 to 7.2 restored VRE’s initial log phase growth but did not enable VRE to reach the maximal OD600 obtained in fresh, neutral pH media (Fig E6C). While lactulose modestly augments VRE growth, VRE growth is inhibited by addition of SCFA to the media, and inhibition is augmented at lower media pH (Fig E6D-F). These results suggest that B. longum inhibits VRE growth through parallel mechanisms that include media acidification, SCFA production, and, to a lesser extent, nutrient deprivation.
Bifidobacterium expansion reduces infection risk
To determine whether lactulose-mediated Bifidobacterium expansion and associated metabolite changes are associated with clinical benefits, we correlated microbiome compositional and metabolomic data with development of common infections in cirrhosis (Figs 5, E7, and E8).
Figure 5: Bifidobacteria expansion and associated metabolite production are associated with decreased incidence of systemic infection and prolonged survival.
(A) A Linear discriminant analysis effect size (LEfSe) showing the significant (Wilcoxon rank-sum, two-tailed, p ≤ 0.05) effect sizes of taxa between groups. (B) Acetate (mM) and taurocholic acid (μg/mL) were quantified, and (C) conversion from conjugated-primary BA to primary BA and then to secondary BAs was tested for each sample. Each point represents a molar ratio for an individual sample. (A-C) Sample size: n = 101 (no SBP), n = 21 (SBP)). (D) LEfSe showing the significant (Wilcoxon rank- sum, two-tailed, p ≤ 0.05) effect sizes of taxa between groups. (E) Acetate (mM) and taurocholic acid (μg/mL) were quantified, and (F) conversion from conjugated-primary BA to primary BA and then to secondary BAs was tested for each sample. Each point represents a molar ratio for an individual sample. (D-F) Sample size: n = 227 (no bacteremia), n = 19 (bacteremia)). (B,C,E, and F) Each point represents a single sample. Median and interquartile range are indicated by the horizontal line and box, respectively. The lower vertical line depicts Q1 – 1.5*IQR and the upper vertical line depicts Q3 + 1.5*IQR. Statistical comparisons between individual groups were analyzed using a two-tailed Wilcoxon rank sum test. P-values are adjusted for multiple comparisons using the Benjamini-Hochberg procedure. Survival curves were stratified based on (G) initial stool sample alpha-diversity or (H) lactulose administration and Bifidobacteria expansion of the initial stool sample. The number at risk for each condition at each 10-day interval is shown below the survival curve. Survival analysis was performed using Kaplan-Meier curves, and the p-value was obtained from a log-rank test. CA: cholic acid, TCA: taurocholic acid, DCA: deoxycholic acid.
LEfSe analysis demonstrated that Bifidobacterium is negatively associated with development of SBP (Fig 5A). We identified 122 ascites samples with near concurrent fecal samples, and 21 patients were diagnosed with SBP (Fig 5A, E7A,B, Table S3). We found that 1 of 21 patients (4.8%) with ≥10% Bifidobacterium frequency was diagnosed with SBP while 20 of 81 patients (24.7%) with <10% Bifidobacterium developed SBP. Upon correcting for MELD-Na at the time of ascitic fluid sampling and concurrent PPI use, Bifidobacterium expansion remained an independent predictor of remaining SBP-free (OR 0.09, 95% CI 0.00 – 0.54, p value 0.03, Tables S4, S5). Qualitative metabolomic analyses demonstrated that glycolithocholic acid, serotonin, and niacin were significantly associated with being SBP-free (Fig E7C). The Bifidobacteria-generated metabolite indole-3-lactic acid was also increased in fecal samples from SBP-free patients, albeit to a lesser extent (p < 0.05, log2 fold-change 0.95, Fig E7C). Quantitative metabolomic analysis demonstrated that acetate was significantly higher in SBP-free fecal samples and taurocholic acid was higher, but not statistically significantly, in fecal samples from patients with SBP (Fig 5B). Consistent with the BA-modifying properties of Bifidobacteria, samples from patients without SBP efficiently convert conjugated to unconjugated primary BA (CA:TCA ratio) but have similar capacity to convert primary to secondary BA (Fig 5C).
We next paired 246 blood cultures with adjacent fecal samples for analysis and identified 19 diagnoses of bacteremia (Fig 5D, E8A,B, Table S6). LEfSe analysis demonstrated that Bifidobacteria were associated with remaining bacteremia-free (Fig 5D). There was 1 diagnosis of bacteremia associated with the 59 (1.7%) samples with ≥10% Bifidobacteria and 18 of 187 samples (9.6%) with <10% Bifidobacteria (OR: 0.16, 95% CI: 0.015 to 0.96, p = 0.05, Table S7). When correcting for MELD-Na at the time of blood culture and concurrent PPI, the odds ratio remained 0.16 with CI not including 1. However, likely due to the small number of events (only 19 bacteremia diagnoses), the p-value was 0.07 (Table S8). Qualitative metabolomics demonstrated that conjugated primary BA and primary BA were all significantly increased in bacteremia-associated samples whereas higher levels of the Bifidobacteria-derived indole-3-lactic acid were associated with sterile cultures (Fig E8C). Targeted quantitative metabolomics confirmed that samples from bacteremic patients had increased taurocholic acid concentrations (Fig 5E) but only modestly reduced ratio of unconjugated to conjugated BA ratio (Fig 5F). These data demonstrate an association between Bifidobacteria expansion and protection from SBP and bacteremia and support the hypothesis that acetate production and BA deconjugation contribute to protection against some of the most common infectious complications of advanced liver disease.
Bifidobacteria expansion associates with improved survival
We next assessed 90-day survival in patients stratified by either initial sample alpha-diversity (Fig 5G) or lactulose exposure and Bifidobacterium expansion (Fig 5H). Patients with low initial sample alpha-diversity had significantly reduced 90-day survival compared to those with medium or high initial sample alpha-diversity (Figure 5G). Compared to patients who received lactulose but did not expand Bifidobacterium, patients with lactulose-associated Bifidobacterium expansion had significantly improved overall 90-day survival as assessed by Kaplan-Meier analysis (Fig 5H). While many patient characteristics between the groups were similar, patients with low alpha-diversity and those who received lactulose but did not expand Bifidobacterium had other clinical parameters that portend a poor prognosis (Tables E1 and E2). While the unadjusted Kaplan-Meir analyses revealed significant differences between groups, when adjusting for multiple clinical parameters that could affect mortality with Cox proportional hazard models, the p-values of 0.26 (alpha-diversity) and 0.12 (Bifidobacterium expansion) fall short of the traditional 0.05 threshold despite hazard ratios of 1.52 (low alpha-diversity) and 0.42 (Bifidobacterium expansion) (Tables S9, S10). While microbiome diversity and the degree of Bifidobacterium expansion inversely correlate the risk of mortality, the lack of independence from clinical parameters that predict mortality is likely attributable to the intensity of interventions, including antibiotic treatment, that patients receive as the severity of CLD increases (Tables E1 and E2).
DISCUSSION
Lactulose has been used to treat HE for over 50 years44. Despite widespread use, lactulose’s mechanism of action has remained incompletely defined7,15. While lactulose is postulated to decrease ammonia absorption by decreasing bowel transit time and gut lumen acidification, its role in altering the microbiome has been largely unexplored. We demonstrate that in patients with liver disease, lactulose leads to marked expansion of Bifidobacteria species in the absence of broad-spectrum antibiotics. This produces a distinct gut microbiome taxonomic and metabolic profile that is associated with exclusion of antibiotic-resistant pathobionts. Bifidobacteria inhibit in vitro growth of antibiotic-resistant Enterococcal species, which is augmented by lactulose. Our findings suggest a protective role of lactulose-mediated Bifidobacteria expansion in patients with liver disease. Consistent with this, Bifidobacteria expansion and associated metabolites associate with reduced incidence of infection and improved 90-day survival.
In addition to treating precipitating factors (e.g. infection), lactulose has replaced dietary modification, antibiotics, and laxatives as first line treatment for HE. All HE therapies are aimed at reducing gut ammonia production and absorption, and the initial reports of lactulose use for HE concluded that this is due to luminal acidification44,45. It has also been assumed that, similar to healthy subjects and experimental models12,46,47, lactulose exhibits a prebiotic effect, but this has not been consistently shown in liver disease10,13,44,48,49. The initial trial of lactulose was unable to detect changes in fecal bacteria44; however, this study was limited by culture-based techniques. More recent studies characterizing the microbiomes of cirrhotic patients did not reveal Bifidobacteria expansion in patients treated with lactulose9,13, likely because 16S rRNA sequencing platforms often do not amplify Bifidobacterium genes50. Similarly, lactulose withdrawal demonstrated only slight decreases in Faecalibacterium species using sequencing techniques10. We avoided 16S rRNA amplification bias by metagenomically sequencing fecal samples and demonstrate marked expansion of Bifidobacteria in a large cohort of cirrhotic patients receiving lactulose.
Bifidobacterium species are dominant fecal microbes in breast fed infants and are considered a prototypical health-promoting bacterium51. Bifidobacterium species imprint the human immune system52, are associated with decreased atopy and autoimmune diseases53, and administration decreases rates of necrotizing enterocolitis in preterm infants54. In adults, fecal Bifidobacteria modulate anti-tumor immunity and enhance immunotherapy efficacy in humans with melanoma and synergize with immunotherapies to reduce melanoma growth in mouse models55,56. Bifidobacteria also produce acetate and lactic acid, which antagonize pathogens to reduce the incidence of enteric infections in infants. While Bifidobacteria do not provide colonization resistance to pathogenic E. coli in a murine model, specific acetate-producing strains limit systemic disease via acetate-mediated promotion of epithelial integrity and immune surveillance42. Gut acidification, mucosal barrier enhancement, and immunomodulatory effects of Bifidobacteria may all benefit patients with liver disease, who are immunocompromised and at high risk of enteric pathogen colonization, growth and dissemination.
While Bifidobacteria attenuate liver disease progression in rodents57,58, our study suggests that Bifidobacteria may reduce complications of advanced liver disease and enhance approaches for supportive care. We report an association between Bifidobacteria expansion and reduced incidence of infection and prolonged survival. This beneficial association has multiple plausible explanations that may be driven by distinct metabolite profiles, including increased acetate levels and enhanced hydrolysis of conjugated primary BA. Similar metabolite profiles appear to be protective against both SBP and bacteremia. SCFA production has been linked to gut colonization resistance of common gram-negative enteric pathogens59. Similarly, Bifidobacteria expansion was associated with reduced abundance of the common gram-positive pathogen, VRE in our patients, and inhibited VRE growth in vitro, likely via both media acidification and SCFA production. Bifidobacteria-mediated VRE growth inhibition was significantly enhanced by lactulose, which also significantly increased acetate production and acidification. While colonization and expansion are important initial steps in infection development, SCFAs also limit pathogen translocation by enhancing both epithelial integrity and mucosal immune function29,30,42,60. Moreover, Bifidobacteria are associated with decreased intestinal permeability in patients with alcohol dependence, which is common in our cohort61. In our study, Bifidobacteria-expanded samples also had higher levels of indole-3-carboxaldehyde, which activates the aryl hydrocarbon receptor to increase mucosal IL-22 production and maintain reactivity towards pathogens.
Patients with liver disease are commonly treated with broad spectrum antibiotics with resultant dysbiosis. Infections precipitate hepatic decompensation and contribute to morbidity and mortality62,63. The gut microbiome has been implicated in the development of infections in many disease states25,64, including cirrhosis65. A recent study used untargeted serum metabolites and fecal 16S data to predict infections in cirrhosis66; however, many identified metabolites were not microbially derived and did not substantially improve the ability to predict infection beyond standard clinical metrics66. Antibiotics, while highly effective for treatment and prophylaxis against common infections 67,68, are associated with increasing antibiotic resistance genes in the gut microbiomes and subsequent poor outcomes69-72. Our study will help identify patients who may benefit from targeted microbiome therapy in future interventional trials.
Small studies of probiotics and fecal microbiota transfer (FMT) have been performed primarily to control HE with overall positive results73-77, but reports of FMT-mediated transmission of toxin-producing and drug-resistant pathogens to patients78,79 are among concerns that have limited widespread adoption of FMT in CLD80. In outpatients with liver disease, synbiotic approaches with lactic acid bacteria and fermentable fiber sources reduce blood ammonia levels, endotoxemia, and incidence of HE81,82. These studies do not assess durability or metabolic function of the probiotic or correlate use with other clinical outcomes. In subjects without liver disease, both prebiotic and probiotic approaches expand Bifidobacteria, and a synbiotic approach prolongs functional Bifidobacteria engraftment12,43,83,84. Another important consideration that has not been well-studied is the metabolomic change that marked Bifidobacteria expansion has on the gut microbiome.
While associated with improved outcomes in our population, a Bifidobacteria-expanded microbiome is not reflective of healthy adult microbiome. Acetate prevents gut pathogen dissemination in a murine model, but butyrate is better studied with regard to epithelial barrier function and mucosal immune function29,30,32,42. Secondary BA are considered markers of a “healthy” gut microbiome but have also been implicated in prevention of enteric infections64 and reduced intestinal inflammation85. Optimal design of probiotic therapies to reconstitute microbiome functions will likely require a spectrum of bacterial species that cooperatively produce beneficial metabolites. Bifidobacteria will likely be an important component of therapeutic consortia and are known to contribute to the development of the human gut microbiome. Lactulose-mediated Bifidobacteria expansion may therefore allow for expansion and engraftment of additional important commensal organisms. Recent studies have demonstrated that alcohol consumption increases acetate production and expands Bacteroidetes86, supporting the plausibility that Bifidobacteria-derived metabolites may support additional microbiota components.
Our study demonstrating associations between microbiome compositions and functions with clinical outcomes in patients with severe liver disease has important limitations. First, our observational study design cannot demonstrate causation. While in vitro and animal model experiments support the hypotheses generated by our clinical observations, a more important role for our findings is to provide a rationale for future interventional studies. Second, we include patients with a wide range of disease etiologies and severities. The majority of patients in our study had decompensated cirrhosis, the final endpoint of CLD. While microbiome parameters may differ between primary disease etiology early in disease pathogenesis, patients with decompensated cirrhosis have more in common with each other than patients with early stage liver disease of the same etiology. Of those with decompensated cirrhosis, alcohol was the most common etiology, which is representative of patients hospitalized with liver disease at our university and increasingly common worldwide. Finally, despite recruiting a large cohort, there remained relatively few clinical events that could be paired with collected fecal samples. While this limits our power to detect statistically significant associations, we still observed strong associations with Bifidobacteria expansion and associated metabolite profiles and important clinical outcomes.
In conclusion, we report that lactulose-mediated gut Bifidobacteria expansion is associated with a distinct fecal microbiome compositional and metabolic profile, reduced antibiotic-resistant pathobiont burden, and improved clinical outcomes in hospitalized patients with liver disease, including reduced incidence of infection and prolonged survival. Our study provides insight into the mechanism of action of a commonly prescribed medication and the rationale for future studies designed to precisely target the gut microbiome to prevent complications of advanced liver disease.
METHODS
Study Design
This was a prospective cohort study of consecutive hospitalized adult hepatology patients at a single institution from April 2021 to April 2022. Inclusion criteria were age ≥18 years, ability to provide informed consent (either themselves or by proxy if unable to provide consent), and being treated on the hepatology consult service. Subjects who were younger than 18 years, unable to provide consent, had prior solid organ transplant, or a prior colectomy were excluded. Patients were enrolled as soon as possible upon hospital admission, most within 48 hours. Samples were obtained under a protocol approved by the Institutional Review Board at the University of Chicago (IRB21-0327). Written informed consent was obtained from all participants or their surrogate decision makers. Participants were not compensated to take part in this observational study. No statistical methods were used to pre-determine sample sizes but our sample sizes are similar to (or larger than) those reported in previous observational human fecal microbiome studies9,27. Subjects were not randomized, and there was no formal blinding in this observational study. All subjects who produced at least one sample were included in the analyses. Specific samples were chosen for analysis as indicated in each figure.
Specimen Collection and Storage
After enrollment, an order for stool collection was placed in the electronic medical record. Stool samples were collected by the clinical nursing teams on inpatient wards and intensive care units. After collection, samples were immediately sent to the microbiology lab through the pneumatic tubing system and stored at +4°C until collection by the research team. The research team collected freshly obtained refrigerated samples twice daily, and all samples were all aliquoted and stored at -80°C within 24 hours of sample production. If able to provide additional samples, fecal samples were collected approximately every 2 days during hospitalization. Samples were collected in a similar manner on re-hospitalization up until 1 year post-enrollment or until death or transplant. Samples remained stored at -80°C until they were processed for metagenomics and metabolomics.
Clinical data collection:
Upon enrollment, all patients were given a unique patient ID that was linked to their unique medical record numbers (MRNs). The unique ID was stored in a RedCap database along with prior to admission medications and disease characteristics, which were obtained by a combination of patient/family recollection and verification with medical records when available. After enrollment, clinical data (including inpatient medication information and laboratory values) was all gathered from the Center for Research Informatics (CRI: https://cri.uchicago.edu/) at the University of Chicago. This is a clinical data warehouse that contains all medication administration records (MAR), laboratory values, and additional clinical parameters linked to each MRN. The CRI database and RedCap data were merged through MRNs, and select records were verified by chart review to ensure accuracy. All survival data was verified with chart review. There were no discrepancies identified between CRI database and the electronic medical record.
Infectious data:
Infectious data was obtained through the CRI data warehouse and verified with manual chart review. Infections were defined using standard clinical criteria. That is, ascitic fluid infections without an evident intraabdominal source (e.g. bowel perforation) were all considered spontaneous bacterial peritonitis (SBP) and were diagnosed with the standard clinical definition of either polymorphonuclear cells (PMN) of 250cells/mm3 or greater or a positive ascites culture67,68. Cultures growing common contaminants (i.e. components of skin flora) were considered negative if PMN were < 250cells/mm3 and the clinical team did not treat for SBP. Blood stream infection (i.e. bacteremia) was defined as having a blood culture with bacterial or fungal growth. Skin contaminants were again excluded (i.e. considered “negative” cultures) if the clinical team did not treat for bacteremia. Clinical samples were paired with the closest stool sample that was within 14 days prior to the ascitic sample or 3 days after the ascitic sample. If a clinical sample indicated infection, all subsequent samples for the subsequent 28 days were excluded from analysis to minimize observing the effects of directed antibiotic therapy. If an initial ascitic fluid sample or blood culture was negative, subsequent clinical samples were excluded for 14 days. Flow diagrams for this approach are shown in Figure S11.
Metagenomic Analyses
Fecal samples underwent shotgun DNA sequencing. After undergoing mechanical disruptions with a bead beater (BioSpec Product), samples were further purified with QIAamp mini spin columns (Qiagen). Purified DNA was quantified with a Qubit 2.0 fluorometer and sequenced on the Illumina HiSeq platform, producing around 7 to 8 million PE reads per sample with read length of 150 bp. Adapters were trimmed off from the raw reads, and their quality were assessed and controlled using Trimmomatic (v.0.39)87, then human genome were removed by kneaddata (v0.7.10, https://github.com/biobakery/kneaddata). Taxonomy was profiled using metaphlan4 using the resultant high-quality reads88. Microbial reads then were assembled using MEGAHIT (v1.2.9)89, genes are called by prodigal (https://github.com/hyattpd/Prodigal). In addition, high-quality reads are queried against genes of interest, such as virulence factors, cazymes, and antibiotic resistance genes, using DIAMOND (v2.0.4)90, and hits are filtered with threshold > 80% identity, > 80% protein coverage, then abundance is tabulated into counts per million or reads per million mapped reads (RPKM).
Alpha-diversity of fecal samples was estimated using the Inverse Simpson Index, while beta diversity was assessed by using taxumap (https://github.com/jsevo/taxumap). We applied Uniform Manifold Approximation and Project (UMAP) on taxonomy profiles on the 847 Liver Disease samples using a slightly modified approach, taxUMAP91. Number of neighbors was 375, while no custom weighting of the aggregations of taxon abundances was applied. Each sample is represented by a single point and colored by most abundant/dominant taxon as indicated. Samples with no taxon > 5% relative abundance were considered to have no most abundant taxon and were labeled as “other.” Metagenomic information is publicly available on NCBI under BioProject ID PRJNA912122 (liver disease cohort) and BioProject ID PRJNA838648 (healthy donor cohort).
Mouse fecal DNA isolation
DNA was extracted using the QIAamp PowerFecal Pro DNA kit (Qiagen). Prior to extraction, samples were subjected to mechanical disruption using a bead beating method. Briefly, samples were suspended in a bead tube (Qiagen) along with lysis buffer and loaded on a bead mill homogenizer (Fisherbrand). Samples were then centrifuged, and supernatant was resuspended in a reagent that effectively removed inhibitors. DNA was then purified routinely using a spin column filter membrane and quantified using Qubit.
16S sequencing
16S sequencing was performed for murine studies in which known bacterial strains that were previously whole genome sequenced were given to ex-germ-free mice. V4-V5 region within 16S rRNA gene was amplified using universal bacterial primers – 563F (5’-nnnnnnnn-NNNNNNNNNNNN-AYTGGGYDTAAA- GNG-3’) and 926R (5’-nnnnnnnn-NNNNNNNNNNNN-CCGTCAATTYHT-TTRAGT-3’), where ‘N’ represents the barcodes, ‘n’ are additional nucleotides added to offset primer sequencing. Approximately ~412bp region amplicons were then purified using a spin column-based method (Minelute,Qiagen), quantified, and pooled at equimolar concentrations. Illumina sequencing-compatible Unique Dual Index (UDI) adapters were ligated onto the pools using the QIAseq 1-step amplicon library kit (Qiagen). Library QC was performed using Qubit and Tapestation and sequenced on Illumina MiSeq platform to generate 2x250bp reads.
16S qPCR
16S qPCR was performed for murine studies in which ex-germ-free mice were monocolonized with Bifidobacteria given that all mice had 100% relative abundance of Bifidobacteria. Extracted DNA was diluted to 20ng/ul to ensure concentrations fell within measurable range. Degenerate primers were diluted to 5.5mM concentration. Primer sequences are as follows: 563F (5’- AYTGGGYDTAAAGNG-3’) and 926Rb (5’-CCGTCAATTYHTTTRAGT-3’). Standard curves were generated using linearized TOPO pcr2.1TA vector (containing V4-V5 region of the 16S rRNA gene) transformed into DH5α competent bacterial cells. Five-fold serial dilution was performed on the purified plasmid from 108 to 103 copies/μl per tube. PCR products were detected using PowerTrack SYBR Green Master Mix (A46109). qPCR was run on QuantStudio 6 Pro (Applied Biosystems) with the following cycling conditions: 95°C for 10 min, followed by 40 cycles of 95°C for 30 s, 52°C for 30s, and 72°C for 1min. Copy numbers for samples were calculated using the Design and Analysis v2 software
Metabolomic Analyses
Short chain fatty acids (SCFA, i.e. butyrate, acetate, propionate, and succinate) were derivatized with pentafluorobenzyl bromide (PFBBr) and analyzed via negative ion collision induced-gas chromatography-mass spectrometry ([−]CI-GC-MS, Agilent 8890)92. Eight bile acids (BA, i.e. primary: cholic acid; conjugated primary: glycocholic acid, taurocholic acid; secondary: deoxycholic acid, lithocholic acid [LCA], isodeoxycholic acid; modified secondary: alloisolithocholic acid [alloisoLCA] and 3-oxolithocholic acid [3-oxoLCA]) were quantified (μg/mL) by negative mode liquid chromatography-electrospray ionization-quadrupole time-of-flight-MS ([−]LC-ESI-QTOF-MS, Agilent 6546). Eleven indole metabolites were quantified by UPLC-QqQ LC-MS. Eighty-five additional compounds were relatively quantified using normalized peak areas relative to internal standards. Data analysis was performed using MassHunter Quantitative Analysis software (version B.10, Agilent Technologies) and confirmed by comparison to authentic standards. Quantitative fecal metabolomic information paired to fecal metagenomic information is publicly available on NCBI under BioProject ID PRJNA912122 (liver disease cohort) and BioProject ID PRJNA838648 (healthy donor cohort). Raw data files are publicly available on MassIVE repository under Dataset IDs MSV000092750 and MSV000092751(both liver disease and healthy donor cohorts).
Bacterial culture
The Bifidobacteria longum strains MSK.11.12 and DFI.2.45 were previously derived from two distinct healthy donor stool samples and whole genome sequenced (BioSample ID: SAMN19731851 and SAMN22167409). The vancomycin resistant Enterococcus faecium (VRE) strain used in this study was obtained from ATCC (strain 700221). Both bacterial strains were grown in anaerobic conditions in Brain-heart infusion broth (BHI broth, BD 237500). The pH was adjusted to 7.0 with NaOH. Media was supplemented with lactulose (Thermo Scientific, J60160-22) or sucrose (Fisher BP220-1) where indicated. For growth in media supplemented with short chain fatty acids, media was supplemented with either sodium butyrate (Sigma 303410), sodium succinate (Sigma S2378) or sodium acetate (Sigma S5636). For studies of bile acid metabolism, 10μg/mL of conjugated primary bile acid, either glycocholic acid (Sigma T4009) or taurocholic acid (EMD Millipore, 360512), was added to the media. All growth curves were obtained in anaerobic culture conditions at 37°C on a BioTek EPOCH2 microplate reader with BioTek Gen 5 3.11 software. Growth curves were analyzed in GraphPad Prism Version 9.4.0. Lactulose concentrations in were measured using the EnzyChrom™ Lactulose Assay Kit (ELTL-100). No data was excluded from analysis.
Mouse studies
All mouse studies were approved by The University of Chicago Institutional Animal Care and Use Committee (IACUC, Protocol 72599). For germ free studies, 6-18 week-old wild type (WT) male and female C57BL/6 mice were used for all studies. Mice were initially obtained from The Jackson Laboratory and subsequently bred and raised in a germ-free isolator. After removal from the germ-free isolator, mice were handled in a sterile manner and individually housed in sealed negative pressure (BCU) isolators. Throughout breeding and experimental conditions, mice were housed within the University of Chicago Gnotobiotic Research Animal Facility (GRAF). The GRAF is operated with adherence to the Guide for Care and Use of Laboratory Animals, 8th edition, 2011, National Academy of Sciences. This includes standardized 12-hour light/dark cycles (light from 6AM - 6PM), centrally monitored temperature (maintained between 68°F and 79°F) and centrally monitored humidity (maintained between 30-70%). Gnotobiotic mice were fed an ad libitum diet of autoclaved Teklad Global 18% Protein Rodent Diet (Sterilizable) (2018S/2018SC). For experiments, mice were treated with either regular sterile water or sterile water supplemented with filter sterilized lactulose at a final concentration of 20g lactulose/L of water in the timing indicated in Figures E4 and E5. For monocolonization, B. longum was grown to steady state in BHIS, pelleted, and resuspended in an equal volume of PBS, and previously germ-free mice were gavaged with 200μL of a freshly prepared suspension on 3 consecutive days. For experiments with consortia, the consortia strains are as follows. CON.1 and CON.2: A. hadrus, B. longum, B. ovatus, C. comes, C. scindens, C. symbiosum, E. lenta, L. gasseri, P. distasonis, P. merdae, and R. gnavus. CON.3: A. caccae, B. longum, B. ovatus, C. scindens, C. symbiosum, E. lenta, P. distasonis, and P. vugatus. Details regarding strain ID and Biosample ID for each strain are given in Table S2. Each consortia strain was grown to early steady state and normalized to OD600 = 0.3. Consortia strains were grown in BHIS, Wilkins-Chalgren (Fisher), de Man, Regosa, and Sharpe (MRS) broth (Fisher), or modified Yeast Casitone Fatty Acids (YCFA) medium termed YTFA medium (recipe shown in Table S11). Stocks of the consortia were stored at -80°C in 20% glycerol, 0.1% cysteine until ready for use, and previously germ-free mice were gavaged with 100μL (CON.1 and CON.2) or 200μl (CON.3) of the consortia for 3 consecutive days. Fecal pellets were collected at the indicated timepoints for 16S rRNA metagenomic analysis and targeted metabolomic analysis. No statistical methods were used to pre-determine sample sizes but our sample sizes are similar to those reported in previous studies of commensal consortia administered to gnotobiotic mice93. Mice were grouped to equalize the number of male and female mice in each group, but no formal randomization was done due to scarcity of gnotobiotic animals. While there was no formal blinding, microbiome data was collected and analyzed by core facilities with staff that were blinded to the experimental design (commensal exposure and treatments) that microbiome data was linked to. All mice that produced fecal pellets were included in the analyses, and no collected data was excluded from the analyses.
Statistical analyses
All statistical analyses were conducted using the R programming language (version 4.2.2). Data was not assumed to be normal, but this was not formally tested. Adjusted p-values of the tests were considered to be statistically significant for all analyses conducted if p ≤ 0.05. Continuous variables were compared between the groups using Wilcoxon rank-sum test (rstatix::wilcox_test) and multiple test correction were adjusted following the Benjamini-Hochberg method (stats:: p.adjust). Categorial variables were compared using Fisher’s Exact test (stats::fisher.test). Linear regression (stats::lm) was used to estimate the response of an outcome (Figure 2: Bifidobacteria and Enterococcus abundance) to multiple factors (Figure 2: lactulose exposure, PPI use, antibiotics, and stool consistency). Results of linear regression are shown in Table S1. Logistic regression (stats::glm) was used to estimate the odds of multiple factors (Figure 5: Bifidobacteria abundance, PPI use, MELD-Na score) affecting outcomes (Figure 5: SBP and bacteremia). Results of logistic regression are shown in Tables S4 and S7. Kaplan–Meier curves for survival were generated after stratifying for selected microbiome parameters including alpha-diversity (high, medium, and low, Figure 5G) and Bifidobacteria abundance after lactulose exposure (Figure 5H) (survival::Surv, survfit, ggsurvplot). A Cox proportional hazards regression model for mortality was used to estimate the effects of microbiome parameters (alpha-diversity and Bifidobacterium abundance after lactulose exposure) adjusting for known risk factors for mortality in liver disease (survival::coxph). Results of Cox proportional hazard testing are shown in Tables S9 and S10.
Supplementary Material
ACKNOWLEDGEMENTS
The authors would like to acknowledge the staff in the Duchossois Family Institute (DFI) metagenomics and metabolomics core facilities for technical assistance. We are grateful to all members of the DFI for enthusiastic discussions that guided this study. We would also like to acknowledge the University of Chicago Center for Research Informatics (CRI) for assistance with medical data extraction and the University of Chicago Gnotobiotic Research Animal Facility (GRAF) for their care of animals used in this study and guidance with performing gnotobiotic experiments.
FUNDING
T32DK007074 (M.A.O.)
U01AA026975 (T.G.C.)
R35GM146969 (S.H.L.)
Searle Scholars Program (S.H.L)
Duchossois Family Institute
Footnotes
COMPETING INTERESTS STATEMENT
The authors have no relevant competing interests to disclose.
DATA AVAILABILITY
In keeping with the Health Insurance Portability and Accountability Act (HIPAA) and the University of Chicago IRB policy, clinical data that we obtained through the Center for Research Informatics is stored on secure, encrypted and password protected servers and is not all publicly available. Select, non-identifiable clinical data is available on the data repository: https://github.com/DFI-Bioinformatics/DFILiverDiseaseMicrobiome. Metagenomic information is publicly available on NCBI under BioProject ID PRJNA912122 (liver disease patient cohort and murine 16S data) and BioProject ID PRJNA838648 (healthy donor cohort). Quantitative fecal metabolomic information paired to fecal metagenomic information is publicly available on NCBI under BioProject ID PRJNA912122 (liver disease cohort) and BioProject ID PRJNA838648 (healthy donor cohort). Raw data files are publicly available on MassIVE repository under Dataset IDs MSV000092750 and MSV000092751(both liver disease and healthy donor cohorts). In addition to the repositories specified above, all raw data included in the manuscript is publicly available at: https://github.com/DFI-Bioinformatics/DFILiverDiseaseMicrobiome.
CODE AVAILABILITY
All code used for data analysis is publicly available at: https://github.com/DFI-Bioinformatics/DFILiverDiseaseMicrobiome.
REFERENCES
- 1.Moon AM, Singal AG & Tapper EB Contemporary Epidemiology of Chronic Liver Disease and Cirrhosis. Clin Gastroenterol H 18, 2650–2666 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Younossi ZM et al. Epidemiology of chronic liver diseases in the USA in the past three decades. Gut 69, 564 (2020). [DOI] [PubMed] [Google Scholar]
- 3.Termeie O. et al. Alarming Trends: Mortality from Alcoholic Cirrhosis in the United States. Am J Medicine (2022) doi: 10.1016/j.amjmed.2022.05.015. [DOI] [PubMed] [Google Scholar]
- 4.Collaborators, G. 2017 C. et al. The global, regional, and national burden of cirrhosis by cause in 195 countries and territories, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet Gastroenterology Hepatology 5, 245–266 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Franchis R. de et al. BAVENO VII - RENEWING CONSENSUS IN PORTAL HYPERTENSION Report of the Baveno VII Consensus Workshop: personalized care in portal hypertension. J Hepatol 76, 959–974 (2021).35120736 [Google Scholar]
- 6.Ge PS & Runyon BA Treatment of Patients with Cirrhosis. New Engl J Medicine 375, 767–777 (2016). [DOI] [PubMed] [Google Scholar]
- 7.Vilstrup H. et al. Hepatic encephalopathy in chronic liver disease: 2014 Practice Guideline by the American Association for the Study Of Liver Diseases and the European Association for the Study of the Liver. Hepatology 60, 715–735 (2014). [DOI] [PubMed] [Google Scholar]
- 8.Riggio O. et al. Effect of Lactitol and Lactulose Administration on the Fecal Flora in Cirrhotic Patients. J Clin Gastroenterol 12, 433–436 (1990). [DOI] [PubMed] [Google Scholar]
- 9.Bajaj JS et al. Altered profile of human gut microbiome is associated with cirrhosis and its complications. J Hepatol 60, 940–947 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Bajaj JS et al. A longitudinal systems biology analysis of lactulose withdrawal in hepatic encephalopathy. Metab Brain Dis 27, 205–215 (2012). [DOI] [PubMed] [Google Scholar]
- 11.Haemmerli Peter, U. & Bircher J and Wrong Idea, Good Results (The Lactulose Story). New England Journal of Medcine (1969). [DOI] [PubMed] [Google Scholar]
- 12.Ruszkowski J & Witkowski JM Lactulose: Patient- and dose-dependent prebiotic properties in humans. Anaerobe 59, 100–106 (2019). [DOI] [PubMed] [Google Scholar]
- 13.Wang JY et al. Lactulose Improves Cognition, Quality of Life and Gut Microbiota in Minimal Hepatic Encephalopathy: A Multi-Center, Randomized Controlled Trial. J. Dig. Dis 20, 547–556 (2019). [DOI] [PubMed] [Google Scholar]
- 14.Bajaj JS et al. Nosocomial Infections Are Frequent and Negatively Impact Outcomes in Hospitalized Patients With Cirrhosis. Am J Gastroenterol 114, 1091–1100 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gluud LL, Vilstrup H & Morgan MY Nonabsorbable disaccharides for hepatic encephalopathy: A systematic review and meta-analysis. Hepatology 64, 908–922 (2016). [DOI] [PubMed] [Google Scholar]
- 16.Llopis M. et al. Intestinal microbiota contributes to individual susceptibility to alcoholic liver disease. Gut 65, 830 (2016). [DOI] [PubMed] [Google Scholar]
- 17.Duan Y. et al. Bacteriophage targeting of gut bacterium attenuates alcoholic liver disease. Nature 575, 505–511 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hermanson JB et al. Dietary Cholesterol-Induced Gut Microbes Drive Nonalcoholic Fatty Liver Disease Pathogenesis in a Murine Model. FASEB (2022). [Google Scholar]
- 19.Qin N. et al. Alterations of the human gut microbiome in liver cirrhosis. Nature 513, 59–64 (2014). [DOI] [PubMed] [Google Scholar]
- 20.Dubinkina VB et al. Links of gut microbiota composition with alcohol dependence syndrome and alcoholic liver disease. Microbiome 5, 141 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Caussy C. et al. A gut microbiome signature for cirrhosis due to nonalcoholic fatty liver disease. Nat Commun 10, 1406 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Loomba R. et al. Gut Microbiome-Based Metagenomic Signature for Non-invasive Detection of Advanced Fibrosis in Human Nonalcoholic Fatty Liver Disease. Cell Metab 25, 1054–1062.e5 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Bajaj JS et al. Gut microbial RNA and DNA analysis predicts hospitalizations in cirrhosis. Jci Insight 3, e98019 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ahluwalia V. et al. Impaired Gut-Liver-Brain Axis in Patients with Cirrhosis. Sci Rep-uk 6, 26800 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Taur Y. et al. Intestinal Domination and the Risk of Bacteremia in Patients Undergoing Allogeneic Hematopoietic Stem Cell Transplantation. Clin Infect Dis 55, 905–914 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Peled JU et al. Microbiota as Predictor of Mortality in Allogeneic Hematopoietic-Cell Transplantation. New Engl J Med 382, 822–834 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Stutz MR et al. Immunomodulatory fecal metabolites are associated with mortality in COVID-19 patients with respiratory failure. Nat Commun 13, 6615 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Roediger WEW Utilization of Nutrients by Isolated Epithelial Cells of the Rat Colon. Gastroenterology 83, 424–429 (1982). [PubMed] [Google Scholar]
- 29.Augeron C & Laboisse CL Emergence of permanently differentiated cell clones in a human colonic cancer cell line in culture after treatment with sodium butyrate. Cancer Res 44, 3961–9 (1984). [PubMed] [Google Scholar]
- 30.Peng L, Li Z-R, Green RS, Holzman IR & Lin J Butyrate Enhances the Intestinal Barrier by Facilitating Tight Junction Assembly via Activation of AMP-Activated Protein Kinase in Caco-2 Cell Monolayers. J Nutrition 139, 1619–1625 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hang S. et al. Bile acid metabolites control Th17 and Treg cell differentiation. Nature 576, 143–148 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Arpaia N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T cell generation. Nature 504, 451–455 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Mouzaki M. et al. Bile Acids and Dysbiosis in Non-Alcoholic Fatty Liver Disease. Plos One 11, e0151829 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kasai Y. et al. Association of Serum and Fecal Bile Acid Patterns With Liver Fibrosis in Biopsy-Proven Nonalcoholic Fatty Liver Disease: An Observational Study. Clin Transl Gastroen 13, e00503 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Adams LA et al. Bile acids associate with specific gut microbiota, low-level alcohol consumption and liver fibrosis in patients with non-alcoholic fatty liver disease. Liver Int 40, 1356–1365 (2020). [DOI] [PubMed] [Google Scholar]
- 36.Leonhardt J. et al. Circulating Bile Acids in Liver Failure Activate TGR5 and Induce Monocyte Dysfunction. Cell Mol Gastroenterology Hepatology 12, 25–40 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Alm R, Carlson J & Eriksson S Fasting Serum Bile Acids in Liver Disease. Scand J Gastroentero 17, 213–218 (2010). [DOI] [PubMed] [Google Scholar]
- 38.Ferslew BC et al. Altered Bile Acid Metabolome in Patients with Nonalcoholic Steatohepatitis. Digest Dis Sci 60, 3318–3328 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.O’Leary JG et al. NACSELD acute-on-chronic liver failure (NACSELD-ACLF) score predicts 30-day survival in hospitalized patients with cirrhosis. Hepatology 67, 2367–2374 (2018). [DOI] [PubMed] [Google Scholar]
- 40.Stoma I. et al. Compositional Flux Within the Intestinal Microbiota and Risk for Bloodstream Infection With Gram-negative Bacteria. Clin Infect Dis 73, ciaa068 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ubeda C. et al. Vancomycin-resistant Enterococcus domination of intestinal microbiota is enabled by antibiotic treatment in mice and precedes bloodstream invasion in humans. J Clin Invest 120, 4332–4341 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Fukuda S. et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 469, 543–547 (2011). [DOI] [PubMed] [Google Scholar]
- 43.Yoshida K. et al. Bifidobacterium response to lactulose ingestion in the gut relies on a solute-binding protein-dependent ABC transporter. Commun Biology 4, 541 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Bircher J, Müller J, Guggenheim P & Haemmerli UP TREATMENT OF CHRONIC PORTAL-SYSTEMIC ENCEPHALOPATHY WITH LACTULOSE. Lancet 287, 890–893 (1966). [DOI] [PubMed] [Google Scholar]
- 45.Elkington SG, Floch MH & Conn HO Lactulose in the Treatment of Chronic Portal-Systemic Encephalopathy — A Double-Blind Clinical Trial. New Engl J Medicine 281, 408–412 (1969). [DOI] [PubMed] [Google Scholar]
- 46.Mao B. et al. Lactulose Differently Modulates the Composition of Luminal and Mucosal Microbiota in C57BL/6J Mice. J Agr Food Chem 64, 6240–6247 (2016). [DOI] [PubMed] [Google Scholar]
- 47.Karakan T, Tuohy KM & Solingen GJ Low-Dose Lactulose as a Prebiotic for Improved Gut Health and Enhanced Mineral Absorption. Frontiers Nutrition 8, 672925 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Bajaj JS et al. Colonic mucosal microbiome differs from stool microbiome in cirrhosis and hepatic encephalopathy and is linked to cognition and inflammation. Am J Physiol-gastr L 303, G675–G685 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Riggio O. et al. Effect of Lactitol and Lactulose Administration on the Fecal Flora in Cirrhotic Patients. J Clin Gastroenterol 12, 433–436 (1990). [DOI] [PubMed] [Google Scholar]
- 50.Sim K. et al. Improved Detection of Bifidobacteria with Optimised 16S rRNA-Gene Based Pyrosequencing. Plos One 7, e32543 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Yoshioka H, Iseki K & Fujita K Development and differences of intestinal flora in the neonatal period in breast-fed and bottle-fed infants. Pediatrics 72, 317–21 (1983). [PubMed] [Google Scholar]
- 52.Henrick BM et al. Bifidobacteria-mediated immune system imprinting early in life. Cell 184, 3884–3898.e11 (2021). [DOI] [PubMed] [Google Scholar]
- 53.Vatanen T. et al. Variation in Microbiome LPS Immunogenicity Contributes to Autoimmunity in Humans. Cell 165, 842–853 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Patole SK et al. Benefits of Bifidobacterium breve M-16V Supplementation in Preterm Neonates - A Retrospective Cohort Study. Plos One 11, e0150775 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Sivan A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti–PD-L1 efficacy. Science 350, 1084–1089 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Matson V. et al. The commensal microbiome is associated with anti–PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Xue L. et al. Probiotics may delay the progression of nonalcoholic fatty liver disease by restoring the gut microbiota structure and improving intestinal endotoxemia. Sci Rep-uk 7, 45176 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Xu R, Wan Y, Fang Q, Lu W & Cai W Supplementation with probiotics modifies gut flora and attenuates liver fat accumulation in rat nonalcoholic fatty liver disease model. J Clin Biochem Nutr 50, 72–77 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Sorbara MT et al. Inhibiting antibiotic-resistant Enterobacteriaceae by microbiota-mediated intracellular acidification. J Exp Medicine 216, 84–98 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Yang W. et al. Intestinal microbiota-derived short-chain fatty acids regulation of immune cell IL-22 production and gut immunity. Nat Commun 11, 4457 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Leclercq S. et al. Intestinal permeability, gut-bacterial dysbiosis, and behavioral markers of alcohol-dependence severity. Proc National Acad Sci 111, E4485–E4493 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Bajaj JS, Kamath PS & Reddy KR The Evolving Challenge of Infections in Cirrhosis. New Engl J Med 384, 2317–2330 (2021). [DOI] [PubMed] [Google Scholar]
- 63.Fernández J. et al. Bacterial infections in cirrhosis: Epidemiological changes with invasive procedures and norfloxacin prophylaxis. Hepatology 35, 140–148 (2002). [DOI] [PubMed] [Google Scholar]
- 64.Buffie CG et al. Precision microbiome restoration of bile acid-mediated resistance to Clostridium difficile. Nature 517, 205–208 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Bajaj JS et al. Association Between Intestinal Microbiota Collected at Hospital Admission and Outcomes of Patients With Cirrhosis. Clin Gastroenterol H 17, 756–765.e3 (2019). [DOI] [PubMed] [Google Scholar]
- 66.Bajaj JS et al. Association of serum metabolites and gut microbiota at hospital admission with nosocomial infection development in patients with cirrhosis. Liver Transplant 28, 1831–1840 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Biggins SW et al. Diagnosis, Evaluation, and Management of Ascites, Spontaneous Bacterial Peritonitis and Hepatorenal Syndrome: 2021 Practice Guidance by the American Association for the Study of Liver Diseases. Hepatology 74, 1014–1048 (2021). [DOI] [PubMed] [Google Scholar]
- 68.Wiest R, Krag A & Gerbes A Spontaneous bacterial peritonitis: recent guidelines and beyond. Gut 61, 297 (2012). [DOI] [PubMed] [Google Scholar]
- 69.Shamsaddini A. et al. Impact of Antibiotic Resistance Genes in Gut Microbiome of Patients With Cirrhosis. Gastroenterology 161, 508–521.e7 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Piano S. et al. Epidemiology and Effects of Bacterial Infections in Patients With Cirrhosis Worldwide. Gastroenterology 156, 1368–1380.e10 (2019). [DOI] [PubMed] [Google Scholar]
- 71.Fernández J, Bert F & Nicolas-Chanoine M-H The challenges of multi-drug-resistance in hepatology. J Hepatol 65, 1043–1054 (2016). [DOI] [PubMed] [Google Scholar]
- 72.Wong F. et al. Clinical features and evolution of bacterial infection-related acute-on-chronic liver failure ☆. J Hepatol 74, 330–339 (2021). [DOI] [PubMed] [Google Scholar]
- 73.Dhiman RK et al. Probiotic VSL#3 Reduces Liver Disease Severity and Hospitalization in Patients With Cirrhosis: A Randomized, Controlled Trial. Gastroenterology 147, 1327–1337.e3 (2014). [DOI] [PubMed] [Google Scholar]
- 74.Holte K, Krag A & Gluud LL Systematic review and meta-analysis of randomized trials on probiotics for hepatic encephalopathy. Hepatol Res 42, 1008–1015 (2012). [DOI] [PubMed] [Google Scholar]
- 75.Bajaj JS et al. Long-term Outcomes of Fecal Microbiota Transplantation in Patients With Cirrhosis. Gastroenterology 156, 1921–1923.e3 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Bajaj JS et al. Microbial Functional Change is Linked with Clinical Outcomes after Capsular Fecal Transplant in Cirrhosis. Jci Insight 4, (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Bajaj JS et al. Fecal microbiota transplant from a rational stool donor improves hepatic encephalopathy: A randomized clinical trial. Hepatology 66, 1727–1738 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.DeFilipp Z. et al. Drug-Resistant E. coli Bacteremia Transmitted by Fecal Microbiota Transplant. New Engl J Med 381, 2043–2050 (2019). [DOI] [PubMed] [Google Scholar]
- 79.Zellmer C. et al. Shiga Toxin–Producing Escherichia coli Transmission via Fecal Microbiota Transplant. Clin Infect Dis 72, e876–e880 (2020). [DOI] [PubMed] [Google Scholar]
- 80.Bloom P, Tapper EB, Young VB & Lok AS Microbiome Therapeutics for Hepatic Encephalopathy. J Hepatol 75, 1452–1464 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Liu Q. et al. Synbiotic modulation of gut flora: Effect on minimal hepatic encephalopathy in patients with cirrhosis. Hepatology 39, 1441–1449 (2004). [DOI] [PubMed] [Google Scholar]
- 82.Malaguarnera M. et al. Bifidobacterium longum with Fructo-Oligosaccharide (FOS) Treatment in Minimal Hepatic Encephalopathy: A Randomized, Double-Blind, Placebo-Controlled Study. Digest Dis Sci 52, 3259 (2007). [DOI] [PubMed] [Google Scholar]
- 83.Button JE et al. Dosing a synbiotic of human milk oligosaccharides and B. infantis leads to reversible engraftment in healthy adult microbiomes without antibiotics. Cell Host Microbe 30, 712–725.e7 (2022). [DOI] [PubMed] [Google Scholar]
- 84.Barratt MJ et al. Bifidobacterium infantis treatment promotes weight gain in Bangladeshi infants with severe acute malnutrition. Sci Transl Med 14, eabk1107 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Sinha SR et al. Dysbiosis-Induced Secondary Bile Acid Deficiency Promotes Intestinal Inflammation. Cell Host Microbe 27, 659–670.e5 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Martino C. et al. Acetate reprograms gut microbiota during alcohol consumption. Nat Commun 13, 4630 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Bolger AM, Lohse M & Usadel B Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Blanco-Miguez A. et al. Extending and improving metagenomic taxonomic profiling with uncharacterized species with MetaPhlAn 4. Biorxiv 2022.08.22.504593 (2022) doi: 10.1101/2022.08.22.504593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Li D, Liu C-M, Luo R, Sadakane K & Lam T-W MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015). [DOI] [PubMed] [Google Scholar]
- 90.Buchfink B, Reuter K & Drost H-G Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat Methods 18, 366–368 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Schluter J. et al. The TaxUMAP atlas: Efficient display of large clinical microbiome data reveals ecological competition in protection against bacteremia. Cell Host Microbe (2023) doi: 10.1016/j.chom.2023.05.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Haak BW et al. Impact of gut colonization with butyrate producing microbiota on respiratory viral infection following allo-HCT. Blood 131, blood-2018-01-828996 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Cheng AG et al. Design, construction, and in vivo augmentation of a complex gut microbiome. Cell 185, 3617–3636.e19 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
In keeping with the Health Insurance Portability and Accountability Act (HIPAA) and the University of Chicago IRB policy, clinical data that we obtained through the Center for Research Informatics is stored on secure, encrypted and password protected servers and is not all publicly available. Select, non-identifiable clinical data is available on the data repository: https://github.com/DFI-Bioinformatics/DFILiverDiseaseMicrobiome. Metagenomic information is publicly available on NCBI under BioProject ID PRJNA912122 (liver disease patient cohort and murine 16S data) and BioProject ID PRJNA838648 (healthy donor cohort). Quantitative fecal metabolomic information paired to fecal metagenomic information is publicly available on NCBI under BioProject ID PRJNA912122 (liver disease cohort) and BioProject ID PRJNA838648 (healthy donor cohort). Raw data files are publicly available on MassIVE repository under Dataset IDs MSV000092750 and MSV000092751(both liver disease and healthy donor cohorts). In addition to the repositories specified above, all raw data included in the manuscript is publicly available at: https://github.com/DFI-Bioinformatics/DFILiverDiseaseMicrobiome.
All code used for data analysis is publicly available at: https://github.com/DFI-Bioinformatics/DFILiverDiseaseMicrobiome.