Abstract
Seeds are unique time capsules that can switch between 2 complex and highly interlinked stages: seed dormancy and germination. Dormancy contributes to the survival of plants because it allows to delay germination to optimal conditions. The switch between dormancy and germination occurs in response to developmental and environmental cues. In this review we provide a comprehensive overview of studies that have helped to unravel the molecular mechanisms underlying dormancy and germination over the last decades. Genetic and physiological studies provided a strong foundation for this field of research and revealed the critical role of the plant hormones abscisic acid and gibberellins in the regulation of dormancy and germination, and later natural variation studies together with quantitative genetics identified previously unknown genetic components that control these processes. Omics technologies like transcriptome, proteome, and translatomics analysis allowed us to mechanistically dissect these processes and identify new components in the regulation of seed dormancy and germination.
In this review we provide a comprehensive overview of studies that have helped to unravel the molecular mechanisms underlying dormancy and germination over the last decades.
Introduction
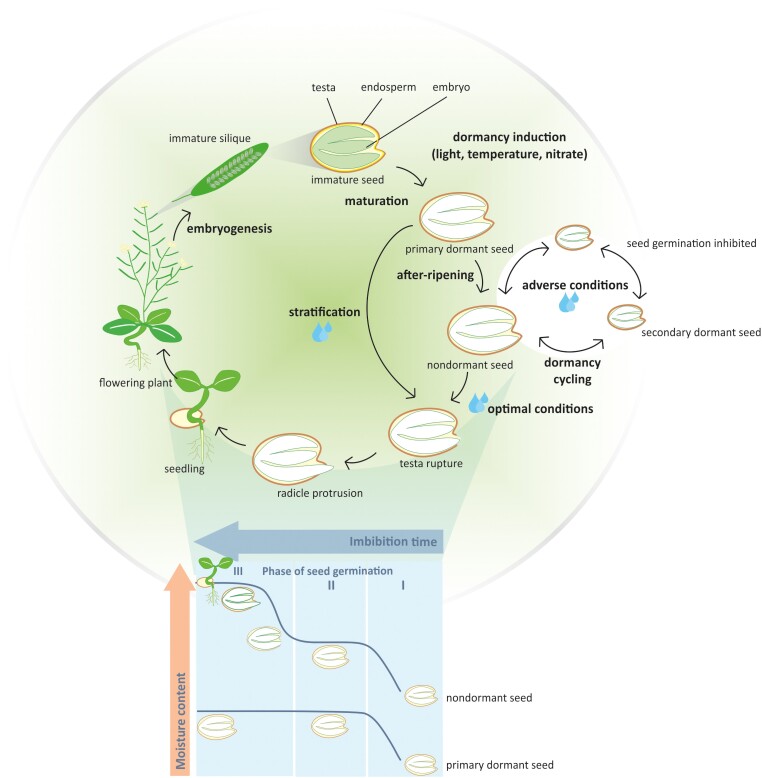
Seeds are complex entities that encompass all the genetic information needed to give rise to a new plant. Structurally, seeds are composed of 3 basic components: the embryo, which will form the future seedling; the energy-rich and nutritious tissue surrounding the embryo called the endosperm; and a protective layer on the outside called the testa or seed coat (Bewley et al. 2012) (Fig. 1). The seed coat also functions in producing protective metabolites, regulation of nutrient transport to the embryo, and defining the final seed size and shape (Francoz et al. 2018).
Figure 1.
The life cycle of a seed. The seed life cycle begins with the embryogenesis phase occurring in the ovary of the flower of the mother plant. The ovary develops into a silique. Here the immature seed consisting of the testa, the endosperm and the embryo, which will form the future seedling, is developing. This stage is followed by seed maturation during which dormancy, desiccation tolerance, and seed longevity are induced. The induction of dormancy is dependent on genetic factors as well as environmental factors like light, temperature, and nitrate availability that the mother plant experiences. Seed maturation ends with a dry mature seed in which primary dormancy has been established. After-ripening or stratification can release dormancy and make the seeds nondormant. Germination of a nondormant seed can be inhibited, or the seed can reinter a dormant state if they are imbibed in adverse environmental conditions. If the nondormant seed is imbibed in optimal conditions, germination progresses through the different stages of germination, including TR and RP, which leads to the establishment of the seedling. Germination can be divided into 3 phases of water uptake. Stage 1 involves rapid uptake of water, which plateaus at stage II. Dormant seeds will remain in stage II and not progress to stage III of water uptake, whereas nondormant seeds will continue water uptake in stage III and complete germination. Imbibed stages indicated by the blue drops and the blue shades in the lower panel of the figure.
The process by which plants establish their next generation is called seed germination. This establishment is a primal goal for plants. Furthermore, seeds can travel for thousands of miles or wait for hundreds of years to germinate. This ability to remain in a quiescent state is called seed dormancy (Bewley 1997).
In the past decades numerous studies have been performed to unravel the mechanisms of seed dormancy and germination; however, we still know little about how emergence is inhibited in dormant seeds and how precisely the embryo emerges from the nondormant seed. Because reproduction is a crucial process in the plant life cycle, the genes regulating seed dormancy and germination are under enormous evolutionary selection pressure. It is challenging to measure dormancy and germination independently because this very much depends on the environmental context in which these measurements are made. An additional complication is that germination is often measured at a population level; when a batch of seeds germinates 100% in optimal conditions, the seeds are said to be nondormant.
In this review, we will attempt to untangle and shed light on the highly inter-linked seed dormancy and germination processes by reporting the most important scientific breakthroughs through time in this field. Here, we emphasize the use of genetic mutants, natural variation, and various omics studies that helped us understand the mechanisms and regulators of seed dormancy and germination. We will mostly focus on the model plant Arabidopsis thaliana because most comprehensive studies have been and are still performed with this species.
The seed life cycle
The seed life cycle (Fig. 1) begins with the embryogenesis phase, in which a double fertilization event results in a single zygote. This zygote expands and grows in cell size and number to fill the embryo sac with the mature embryo (Bewley et al. 2012). The embryo growth is then arrested and followed by the maturation phase, in which the main goal is to accumulate resources and properties needed for successful germination and seedling establishment (Nonogaki et al. 2010). These resources can include mRNAs, sugars, and storage proteins required to kickstart germination. The maturation phase is also the critical phase wherein important traits like desiccation tolerance are established by intense moisture loss and seed dormancy and seed longevity are acquired. At the end of this phase, the seed is in a quiescent dry stage (Gutierrez et al. 2007; Holdsworth et al. 2008). Environmental conditions like light and temperature during the maturation phase, also termed as maternal or parental environment, are critical in determining the dormancy levels and the timing of germination in nature (Biere 1991; Fernandez-Pascual et al. 2013).
Seed dormancy: a blessing in disguise
Seed dormancy is defined as the inability of seeds to germinate in favorable conditions. Dormancy enables the mother plants to avoid risking germination of their entire progeny in an unpredictable and dynamic environment. Nikolaeva (1969) was the first seed biologist to develop a comprehensive classification scheme for the various types of seed dormancy. Dormancy can be broadly classified into 5 types of dormancies: physical, morphological, morphophysiological, physiological, and combinational dormancy (Fig. 2; Baskin and Baskin 2004). The most commonly studied type of dormancy is physiological dormancy. This type of dormancy is prevalent in gymnosperms and major classes of angiosperms, including Arabidopsis. The level of dormancy in seeds is highest at the end of the maturation phase and is called primary dormancy. Primary dormancy in Arabidopsis seeds can be released or alleviated by factors like after-ripening (AR; seed storage in the dry state) or stratification (cold or warm treatment in the imbibed state). The dormancy status also changes in response to environmental signals such as temperature, nitrate, and light quality exposure during seed development. Moreover, the sensitivity of the mature dry seeds to these signals depends on the AR time (Finch-Savage and Leubner-Metzger 2006; Nee et al. 2017a). If adverse environmental conditions prevail for extended periods of time, then after-ripened (AR) seeds can re-enter dormancy. This cycling of dormancy is known as secondary dormancy and can only occur when the seed is imbibed (Fig. 1) (Donohue 2005; Buijs 2020).
Figure 2.
Milestones in the seed dormancy and germination research field through time.This figure graphically represents the biggest insights in the seed dormancy and germination field according to our interpretation. The first milestone was achieved in 1969, when Nikolaeva (1969) classified the 5 types of dormancies. This was followed by the breakthrough study of Koornneef and van der Veen (1980), where they concluded that the ABA concentration within the seeds determines the level of seed dormancy. The graph shows the germination percentage of GA-deficient mutants ga1-2 and the wild-type seeds in increasing concentrations of exogenous GA. In 2004 it was discovered that the translation, but not transcription, is absolutely required for germination to occur (Rajjou et al. 2004). The next milestone arrived when Bentsink et al. (2006) cloned DOG1, the major quantitative trait locus governing seed dormancy in Arabidopsis and fine mapped it to the gene DOG1 (At5G45830). The graph here shows the germination percentages of 3 DOG1 alleles Cvi, Ler, and the mutant (dog1-1) in Arabidopsis during 0 to 150 days of AR. The graph shows the contribution of DOG1 to seed dormancy, as the dog1-1 germinates 100% without a need for AR. In 2011, Bassel et al. generated a network model of global transcriptional interactions called SeedNet. This network represents 2 state-dependent sets of interactions that associate with dormancy and germination but also an intermediate transition region between these processes. The most recent milestone presented is from 2022, when Krzyszton et al. (2022) revealed transcriptional heterogeneity by performing RNA-seq on individual seeds during secondary dormancy induction. The figure shows that the heterogeneity is the lowest in very dormant seeds and increases when dormancy is broken.
Seed germination: the birth of a seedling
When nondormant seeds are imbibed in optimal conditions, seeds are “woken up,” so to say, and germination-promoting processes are initiated. By optimal conditions during imbibition of Arabidopsis seeds, we mean the presence of sufficient moisture, oxygen, optimal temperature, light quality, and absence of environmental stressors such as salinity, drought, cold, or heat stress. Regarding light quality, red light and its perception by photoreceptors called phytochromes is an essential factor for promoting seed germination (Longo et al. 2020). Germination itself is divided into 3 phases (Fig. 1). In Phase I there is tremendous uptake of water by the seed leading to swelling, followed by a phase wherein water uptake plateaus (Phase II). Dormant seeds will not surpass this stage, whereas nondormant seeds will move on to Phase III, in which water uptake is resumed and the seed coat or the testa will rupture (TR). This is a critical physiologically visible stage during seed germination. After TR, cell elongation and expansion take place, which will eventually lead to the embryonic root or radicle to protrude through the endosperm (radicle protrusion [RP]). Once RP has occurred, germination is said to be complete (germination “sensu stricto”). The phase between TR and RP is a very sensitive one because the commitment to germinate into a seedling cannot be reversed once RP has occurred. This could be because seeds drastically lose their desiccation tolerance ability upon RP (Maia et al. 2011). Hence, nondormant seeds can stop or slow germination before RP if environmental conditions do support successful seedling establishment. Therefore, the final commitment to germination is made by integrating signals from the environment, and this process is tightly regulated at the genetic level.
Hormonal control of seed germination and dormancy
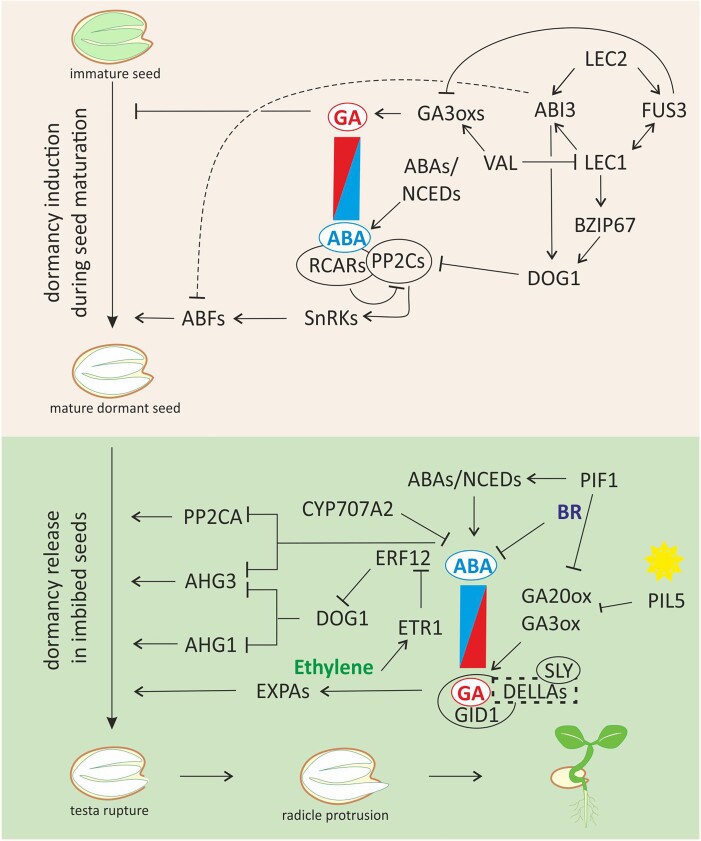
The absolute requirement of abscisic acid (ABA) and gibberellins (GAs) for the regulation of seed dormancy and germination was revealed in Arabidopsis and tomato mutants defective in the biosynthesis of these hormones in the early 1980s (Fig. 2) (Koornneef and van der Veen 1980; Groot and Karssen 1987). Most of these mutants were initially isolated on the basis of their altered seed germination phenotype. The requirement of GA for germination depends on the ABA concentration within seeds, as could be concluded from the observation that GA-deficient mutants can germinate in the absence of ABA. The latter led to the isolation of ABA-deficient mutants as revertants of GA-deficient mutants, which can germinate without GA or on high concentrations of the GA biosynthesis inhibitor paclobutrazol (Koornneef et al. 1982; Léon-Kloosterziel et al. 1996). The biosynthesis mutants (especially those in Arabidopsis, maize, and rice) were crucial in elucidating the biochemical steps and genes of the biosynthetic pathways for both plant hormones. For this the chemical analysis of the mutants was important. The cloning of the underlying genes was based on the mutants using map-based cloning and transposon tagging, also described as forward genetics. In the meantime, the signal transduction steps were also identified. Due to the fact that many of these steps are controlled by multiple genes, this required additional procedures such as chemical genetics and biochemical interaction studies [reviewed by Cutler et al. (2010)]. At this stage, increasing use was made of reverse genetics of candidate genes and the characterization of these mutants and double mutants with seed dormancy tests. Overall, the regulation of seed dormancy and germination and especially the role of ABA and GA in this has been extensively reviewed (Finch-Savage and Leubner-Metzger 2006; Finkelstein et al. 2008; Holdsworth et al. 2008; Bewley et al. 2012; Graeber et al. 2012; Shu et al. 2016). Below we aim at providing an overview of the essential roles of these 2 hormones. Gene interactions during the induction of seed dormancy in immature seeds and during dormancy release in the imbibed state are schematically presented in Fig. 3.
Figure 3.
The regulation of seed dormancy and germination during seed maturation and dormancy release in imbibed seeds. The figure represents a schematic overview of gene interactions during induction of seed dormancy in the immature seeds and during dormancy release in the imbibed state. Key for both stages is the balance between concentrations of GA and ABA, indicated by the red/blue rectangle. Both hormones are sensed during seeds maturation and dormancy release; however, in the scheme, factors involved in the perception of ABA (RCARs and PP2Cs) are indicated in more detail the upperpart of the figure, which represents factors involved in dormancy induction during seed maturation. Those involved in GA perception (GID1, DELLAs, and SLY) are depicted in the lower panel, which shows the interactions during the dormancy release in the imbibed seeds. Arrows indicate a promotive effect, bars a repressive effect. The line between ABI3 and ABFs is dashed because the role of ABI3 in ABA perception has remained unclear so far. The indicated interactions are also described in the main text.
ABA, a repressor of germination and inducer of seed dormancy
In addition to not needing GAs for germination, ABA-deficient aba1 mutants showed an absence of AR requirement in the shallow dormant Arabidopsis accessions Landsberg erecta (Ler) and Columbia (Col; Koornneef et al. 1982). Severe ABA biosynthesis mutants (aba1, aba2, aba3, and the nced6/9 double mutant) show an absolute lack of seed dormancy (Lefebvre et al. 2006). The ABA concentration in seeds is, in addition to the control of ABA biosynthesis, also determined by conversion of ABA into phasic acid leading to inactivation of ABA. Responsible for this conversion are the ABA-8’ hydroxylase(s), which are 2 P450 cytochromes encoded by CYP707A1 and A2. These cyp707a mutants have an increased dormancy phenotype (Kushiro et al. 2004).
ABA levels in mature seeds are the result of synthesis during seed maturation. Depending on the species, the ABA levels peak at 2 moments (Kozaki and Aoyanagi 2022). Detailed analysis of ABA concentrations during imbibition showed that although ABA levels initially decrease, they increase at later imbibition stages in dormant seeds (Ali-Rachedi et al. 2004). Moreover, it is ABA produced and released by the endosperm that represses embryo germination, as was revealed by a “seed coat bedding” assay (Lee et al. 2010). The expression of the DELLA protein RGL2 is required for this response. The importance of ABA in maintaining the dormant state in imbibed seeds is also confirmed by the germination-promoting effect of the ABA biosynthesis inhibitor fluoridone when applied during the imbibition of dormant seeds. ABA in seeds is mainly synthesized in the endosperm and transported to the embryo by 4 ATP-binding cassette (ABC) transporter genes (Kang et al. 2015). AtABCG25 and AtABCG31 export ABA from the endosperm, whereas AtABCG30 and AtABCG40 import ABA into the embryo (Kang et al. 2015). The role of ABA transported from the mother plant during dormancy induction is minor, as was shown by ABA-deficient mother plants with ABA-producing embryos and endosperms obtained by reciprocal crossing of aba1 mutants with its wild type (Karssen et al. 1983).
Seed-specific ABA signaling
The complex regulation of the ABA signaling pathway in seeds, with many gene copies present for its major steps, has recently been described by Née and Krüger (2023). Mutant screens for seeds germinating in the presence of wild type inhibiting concentrations of ABA identified several genes, among which were ABA insensitive 3 (ABI3), ABI4, and ABI5. These genes are important seed-specific downstream ABA targets and are considered repressors of germination (Koornneef et al. 1984; Finkelstein 1994). Based on their specific expression during seed development and the early stages of germination, they are considered to have seed-specific roles.
Studies using strong ABI3 mutants revealed, in addition to a lack of dormancy, many important defects during seed development, including the absence of chlorophyll degradation and desiccation intolerance (Ooms et al. 1993; Nambara et al. 1994). The ABI3 gene together with the seed maturation genes LEAFY COTYLEDON 1 and 2 (LEC1/2) and FUSCA3 (FUS3) form the LAFL (LEC1, ABI3, FUS3, LEC2) regulatory network (Carbonero et al. 2017). These genes are essential for seed maturation, and single gene mutants show partial overlap in their phenotypes (Raz et al. 2001; Jia et al. 2013). All these genes (except LEC1) are members of the AFL (ABI3/FUS3/LEC2) subfamily of B3 transcription factors (TFs) and bind G-box like sequences in the promoters of seed maturation genes, which are bound by basic leucine zipper (bZIP) TFs (Boulard et al. 2017; Carbonero et al. 2017). ABI3 has many direct targets, including ABI5, DELLAs, and the bHLH type TF PIL5 (PHYTOCHROME INTERACTING FACTOR-LIKE 5) (Tian et al. 2020). ABI3, ABI5, and DELLAs together activate the expression of SOMNUS, which encodes a C3H type zinc finger protein that regulates the expression of genes involved in the synthesis of ABA and in the catabolism of GA in imbibed seeds, thus inhibiting seed germination (Park et al. 2011; Lim et al. 2013). The molecular interaction of ABI3 with the primary ABA signaling pathway remains unclear and is based only on the extreme ABA insensitivity of its mutant. The latter is not observed for the fus3 mutants despite that they displayed a similar absence of dormancy as the abi3-3 mutant (Keith et al. 1994).
ABI4 and ABI5 were also identified in a mutant screen for ABA-insensitive mutants and thereby seemed also to be candidates in the ABA signal transduction (Finkelstein 1994). Their mutant phenotypes were much less severe than that of abi3 mutants; however, they showed a strong reduction of ABA/ABI3-induced genes, such as the downregulation of the LATE EMBRYO ABUNDANT proteins AtEM1 and AtEM6 gene expression (Finkelstein 1994; Carles et al. 2002). Conversely, the abi4 and 5 mutants did not show clear seed dormancy phenotypes. The ABI4 gene, reviewed by Chandrasekaran et al. (2020), encodes an APETALA2/ethylene response factor type TF that affects many processes by regulating gene expression, including that of the ABA degradation genes ABA 8’ hydroxylase(s) (Cyp707A1 and Cyp707A2), as well as genes involved in sugar signaling. Additionally, it downregulates several GA biosynthesis genes. The abi4 mutant has a small effect on the rate of germination, and the mutant is not affected in its seed development phenotype (Shu et al. 2013).
ABI5, reviewed by Skubacz et al. (2016), encodes a bZIP TF, which can both bind to the ABA Responsive Element in promotors and interact with ABI3 via its B1 domain. The protein also interacts with AFPs (ABA 5 binding proteins), which allows interaction with other proteins (Lynch et al. 2017). Overexpression of AFP1 and AFP2 resulted in extreme ABA tolerance and desiccation-intolerant seeds resembling strong abi3 mutant alleles and suggests that these genes are major actors in seed-specific ABA signaling. Deng et al. (2023) demonstrated that AFP2 negatively regulates primary seed dormancy by suppressing the expression of the major regulator of seed dormancy, DELAY OF GERMINATION 1 (DOG1). The TF WRKY36 can interact with AFP2 to transcriptionally co-repress TOPLESS RELATED PROTEIN 2 and epigenetically silence DOG1 by reducing histone acetylation. The abi5 mutants themselves show tolerance to germination inhibition by NaCl and osmotic stress (Carles et al. 2002). ABI5 can be phosphorylated by SnRK2 proteins as part of the ABA signal transduction chain. The phosphorylation of ABI5 enables interactions with other proteins, including EM1 promotors, suggesting a role in the acquisition of desiccation tolerance in seeds. Furthermore, ABI5 can interact with other hormones, including GA, jasmonic acid, and auxin pathways (Skubacz et al. 2016). Overall, it seems that ABI5 is crucial for part of the ABA transduction chain in seeds, early germination, and its response to stresses.
GA promoters of seed germination
GAs are essential for germination because GA biosynthesis mutants such as ga1/ga2/ga3 require application of exogenous GAs for germination (Fig. 2, Koornneef and van der Veen 1980). The GA requirement for germination is further indicated by the observation that GA biosynthesis inhibitors such as paclobutrazol and unicazol inhibit germination. GA biosynthesis and GA signaling in seeds has been reviewed by Urbanova and Leubner-Metzger (2018). Active GA binds to the receptor GID1 (encoded by AtGID1a, AtGID1b, and AtGID1c), and together they repress the DELLA proteins. The latter are repressors of GA signaling, and especially RGA2 is the seed-specific representative of this group of 5 proteins (Lee et al. 2002).
The role of GA seems mainly to overcome restraints that are partially related to (ABA induced) dormancy but also to overcome the resistance of the embryo surrounding tissues, including that conferred by the integuments (Debeaujon and Koornneef 2000) and the endosperm (Groot et al. 1988). The latter aspect relates to the induction of genes encoding cell wall–degrading enzymes by GA [reviewed by Nonogaki (2019)]. Focusing on expansin mediate cell-wall loosening, Xu et al. (2020) elucidated the pathway by which DELLA proteins and nitric oxide (NO) control TFs (including TCP14/15, RAP2.2/2.3/2.12, and ZML1), which in turn control expansin gene expression.
Hormonal crosstalk during dormancy and germination
ABA and GA do not regulate seed dormancy and germination on their own. Where interactions with jasmonic acid and auxin were already mentioned, brassinolides (BRs) and ethylene are also known to affect these traits. BRs stimulate germination by promoting TR and endosperm rupture in Arabidopsis in a similar way as GAs do. BRs can rescue the germination phenotypes of the GA biosynthetic mutant ga1-3 and the GA-insensitive mutant sleepy1 (Steber and McCourt 2001). Germination does not entirely depend on GA response but rather on GA and BR working in parallel. Two key basic helix–loop–helix transcription factors in the BR signaling pathway, BRASSINOSTEROID ENHANCED EXPRESSION 2 (BEE2) and BEE 2 INTERACTING with INCREASED LEAF INCLINATION 1 BINDING bHLH1 (HBI1), are involved in the regulation of endosperm rupture. Expression of BEE2 and HBI1 was induced in response to BR and GA treatment (Zhong et al. 2021). Furthermore, BRI1-EMS-SUPPRESSOR 1 (BES1) affecting BR signaling was shown to physically interact with ABI5 to hinder its DNA binding capacity, attenuating the ABA-mediated suppression of seed germination by lowering the expression of ABI5 targets. BIN2, a repressor of BR signaling, promotes the ABA responses (Zhao et al. 2019). BES1 and DELLAs were reported to mediate the crosstalk between BRs and GAs during cell elongation in Arabidopsis [reviewed by Ross and Quittenden (2016)].
Ethylene is another promoter of germination. The loss-of-function mutant of one of the ethylene receptors ETHYLENE RESPONSE FACTOR 1 (ETR1) was identified by its reduced dormancy phenotype and therefore named (reduced dormancy 3; Li et al. 2019). It was shown that ETR1 represses the expression of ERF12, which recruits TOPLESS to form a repressor complex and binds to the DOG1 promoter, thereby inhibiting the expression of DOG1 and leading to reduced seed dormancy by ethylene.
Another mechanism controlled by EIN2 (ETHYLENE INSENSITIVE 2), a component of ethylene signal transduction, was recently described by Guo et al. (2023). The authors concluded that EIN2 can function independently from the canonical ethylene pathway acting via EIN3/EIL1, by inhibition of a histone H3 acetylase encoded by HLS1 (HOOKLESS1). When active, HLS1 acetylates H3 histones in the region of ABI5 and ABI3, which represses their transcription. In the absence of EIN2, the expression of these ABA signaling molecules is upregulated and germination is inhibited.
Auxin has been shown to function both positively and negatively in seed germination depending on its dose. It has also been reported that high doses of auxins enhance seed dormancy by activating ABI3 mediated by the AUXIN RESPONSE FACTOR 10 TF (Liu et al. 2013) and IAA8 (Hussain et al. 2020). This leads to enhanced expression of ABI3, although not by binding to the ABI3 promotor. IAA8 expression is upregulated by cold and by ROS signaling and was found to associate with the ABI3 promotor (Hussain et al. 2020). In both cases the auxin effects were relatively minor in affecting the rate of germination. Wang et al. (2016) emphasized the specific role of auxin in the RP and thereby germination rate. They noticed that this is, to a large extent, controlled by the auxin transporter AUX1. AUX1 expression is negatively controlled by the histone acetylation genes SNL1/SNL2, which decrease in expression upon after dormancy breakage.
The power of natural variation, among others, a DOG1 story
Seed germination is a quantitative trait for which ample genetic variation is present in nature. Especially seed dormancy, which together with the timing of flower initiation is a main factor determining where and when a seed should germinate, has shown to be a key trait defining local adaptation. In European accessions a North-South cline was identified, where (primary) dormancy is lower in the North compared with the South. A similar pattern was found for accessions collected in the Iberian Peninsula (Debieu et al. 2013; Vidigal et al. 2016). Studies on the Iberian population also show that dormancy is generally lower at higher altitudes. Kronholm et al. (2012) described genetic signatures of local adaptation as well as a negative correlation between dormancy and summer precipitation. These clines are likely explained by the shorter growing season (in the Northern regions and higher altitudes) and the riskier environment with a lower chance of survival in the Southern regions and lower altitudes. The importance of the right dormancy levels is illustrated in the work of Huang et al. (2010). They showed with a garden experiment that genetic variation for fitness and germination phenology are associated with the genetic regions that were earlier detected for primary seed dormancy by testing AR requirement in the laboratory.
The use of natural variation for the identification of seed dormancy loci has led to the identification of the key regulator of seed dormancy, DOG1. Natural variation for dormancy was first explored in the Ler Col recombinant population (RIL) (Van der Schaar et al. 1997). However, the use of both these accessions with low dormancy levels only revealed minor quantitative trait loci (QTL). A breakthrough was made when more dormant accessions were used for these genetic approaches. The Ler, Cape Verde Island (Cvi) RIL population yielded 7 QTL that together explained more than 60% of the total variance (Alonso-Blanco et al. 2003). The main QTL identified in this study was the aforementioned DOG1 QTL. Experiments with different RIL populations mainly resulted in the identification of the already known DOG loci (Meng et al. 2008; Bentsink et al. 2010; Amiguet-Vercher et al. 2015; Footitt et al. 2020). Moreover, genome-wide association studies using Arabidopsis accessions from all over the world also identified the already known QTL (He 2014; Martínez-Berdeja et al. 2020), sometimes with a few additional loci (Chen et al. 2023).
So far we have been discussing mostly natural variation for seed dormancy, measured as AR requirement. However, there is also genetic variation for germinability in cold and dark (Meng et al. 2008) and germination variability (Abley et al. 2021), which is referred to as the heterogeneity in germination that occurs in genetically identical plants that have been grown in the same condition. The level of germination variability differs between Arabidopsis accessions, and QTL analyses using MAGIC lines revealed QTL in the DOG1 and DOG6 regions (Abley et al. 2021). The major Cold-tolerant Dark Germination QTL occurred at positions for which no QTL have been cloned so far (Meng et al. 2008).
Identifying the genes underlying the DOGs: an ongoing search
The identified DOG QTL underly genes of a different nature; so far DOG1, DOG6, and DOG18 are cloned, and these encode, respectively, an unknown protein, a NAC transcription factor, and a pseudo phosphatase (RDO5) (Bentsink et al. 2006; Xiang et al. 2016; Song et al. 2021). Although we can only speculate about the nature of the remaining DOG loci, genetic analyses and transcriptional profiling have indicated that they control dormancy by different additive genetic and molecular pathways (Bentsink et al. 2010). Recently, several reviews have appeared that describe the regulation of dormancy via DOG1 (Nee et al. 2017a; Carrillo-Barral et al. 2020; Li et al. 2020; Soppe and Bentsink 2020; Iwasaki et al. 2022). DOG1 was identified using a combined genetic approach involving the fine-mapping of the identified QTL region and a mutagenesis analyses in the background on the near isogenic line (NIL) that contained the DOG1 Cvi allele (Bentsink et al. 2006). The nondormant mutant identified in the NIL background could not be genetically separated from the DOG1 dormant allele and turned out to be a mutation in the DOG1 gene. Knowing that knocking-out DOG1 would lead to loss of dormancy allowed for screening knockout lines in the fine-mapped region of the QTL. As such, At5G45830, a gene of unknown function, was identified as the gene encoding DOG1. The availability of 3 alleles (Cvi, Ler, and the mutant dog1-1), which showed very strong dormancy phenotypes, was of great importance for the later studies (Fig. 2; Bentsink et al. 2006). It is the amount of DOG1 protein that determines the level of seed dormancy, and loss of function of the protein that occurs during AR leads to a reduction of seed dormancy (Nakabayashi et al. 2012). DOG1 interacts with clade A PP2C phosphatases, the ABA-HYPERSENSITIVE GERMINATION 1 and 3 (AHG1 and 3). DOG1 likely suppresses the action of AHG1 and 3 and as such inhibits the release of seed dormancy (Nee et al. 2017b).
There is a clear link between ABA and DOG1, which involves the above described interaction with the AHG proteins but is also based on the fact that DOG1 cannot confer dormancy in a ABA-deficient/-insensitive background (Bentsink et al. 2006). The fact that DOG1 is an α-helical heme-binding protein is another link to ABA (Nishimura et al. 2018). Heme binding proteins, which are reported to function as sensors for oxygen and NO, are required to counteract ABA during dormancy release (Arc et al. 2013). DOG1 is post-translationally modified as the result of AR (Nakabayashi et al. 2012). It is hypothesized that DOG1-bound heme could cause non-enzymatic oxidative cellular damage by promoting ROS formation during seed dry storage; this would be more severe in the presence of larger amounts of DOG1 protein (Chiabrando et al. 2014).
DOG1 expression is induced during seed maturation (Bentsink et al. 2006). In-depth analyses revealed that during seed development, DOG1 genetically interacts with ABI3 and as such affects other seed maturation traits as well (Dekkers et al. 2016b). Seeds of the dog1 mutant have reduced expression of genes encoding LATE EMBRYOGENESIS ABUNDANT and HEAT SHOCK PROTEINs and contain reduced amounts of certain primary metabolites and oligosaccharides. Interestingly, the abi3-1 dog1-1 double mutant has green seeds, which phenocopies the more severe abi3 mutants. Based on sequence alignment, DOG1 belongs to a family of 6 members, the DOG1-Like (DOGL) genes. None of these proteins affect seed dormancy; however, DOGL4 strongly induced the expression of seed maturation–specific genes, including the major seed reserve proteins ALBUMIN, CRUCIFERIN, and OLEOSIN (Sall et al. 2019).
The regulation of DOG1 has intensively been studied. Lower temperatures during seed maturation led to increased levels of DOG1 transcript and protein (Nakabayashi et al. 2012). This effect is mediated by an increased expression of the bZIP transcription factor bZIP67, which transactivates DOG1 expression. Genetic variation in the bZIP67 binding sites of the DOG1 promoter provides a mechanism to explain natural genetic variation in DOG1 expression (Bryant et al. 2019). Moreover, low temperatures, when perceived during seed maturation, are known to increase DOG1 expression, as well as dormancy (Nakabayashi et al. 2012; He et al. 2014). Gene-swapping experiments between Arabidopsis and the nondormant Lepidium sativa showed that the DOG1 function is conserved among these Brassicaceae (Graeber et al. 2014). Whereas biomechanical analyses that allow to investigate the material properties of the endosperm so far have never left the pilot phase in Arabidopsis, these are common ground in Lepidium (Muller et al. 2006; Voegele et al. 2012). Such analyses reveal that DOG1 inhibits the expression of cell-wall remodeling genes via temperature-dependent adjustments of key enzymes of the GA biosynthetic pathway (Graeber et al. 2014). How a role for DOG1 in the endosperm aligns with DOG1 expression in the embryo remains unknown.
In addition to external factors regulating the expression of DOG1, several mechanisms that involve the DOG1 gene structure have been identified to regulate DOG1 expression. These include the following: 1) alternative splicing and selective polyadenylation resulting in short DOG1 and long DOG1 isoforms (Cyrek et al. 2016; Fedak et al. 2016); 2) long noncoding RNAs in the 5’ region of the gene, which are referred to as PUPPIES and promote DOG1 expression in high-salt conditions (Montez et al. 2023), and an antisense RNA located in the 3’ region of the gene referred to as 1GOD that negatively affects DOG1 expression (Cyrek et al. 2016; Kowalczyk et al. 2017); and 3) epigenetic regulation as summarized by Ding et al. (2022).
REDUCED DORMANCY 5 (RDO5), originally identified in the same genetic screen as the dog1-1 mutant and underlying the natural variation at the DOG18 locus (Xiang et al. 2016), has also been identified to bind to DOG1. RDO5 is annotated as a so-called pseudo phosphatase; although it belongs to the PP2C phosphatase family, it lacks phosphatase activity (Xiang et al. 2014). RDO5, similarly to DOG1, is localized to the nucleus. Three members of the PUM family of RNA binding proteins were upregulated in the rdo5 mutant, suggesting that these genes promote germination. The PUM proteins are, in contrast to DOG1 or RDO5, localized in the cytoplasm; therefore the regulation of PUMs by RDO5 is indirect. In agreement with a role for PUMs in the translational regulation of seed germination, these might be downstream factors that regulate the onset of germination and not dormancy specifically. Downregulation of PUM9 and 11 by RNAi in the rdo5 mutant background resulted in increased dormancy levels. PUMs are RNA binding proteins and are therefore discussed in more detail in the section “Mechanisms underlying translational control: The role of RNA binding proteins.”
DOG6 encodes the NAC transcription factor ANAC60 (Song et al. 2021, 2022). Different natural alleles of DOG6 determine the localization of the protein due to the presence or absence of a membrane binding domain. Absence of the membrane binding domain allows the protein to enter the nucleus and as such attenuate seed dormancy.
Exploring transcriptome profiles
Transcriptome analyses have been a preferred method to address seed dormancy– and seed germination–related questions. In the early days, transcript profiling was based on hybridizing mRNAs to cDNA libraries that were constructed from specific tissues. One of the first reports revealing transcriptional changes during seed germination was that of Harada et al. (1988), who made use of nitrocellulose arrays containing a seedling cDNA library. These authors reported on the temporal and spatial specificity of these transcripts in Brassica napus cotyledons and seedling axes based on hybridization analyses of the identified clones. The nitrocellulose carriers used for hybridization of mRNA to cDNAs were replaced for coated slides in 1995 and around 2015 for RNA sequencing (RNA-seq). Before the switch to microarrays was made, Bove et al. (2005) reported a cDNA-AFLP study during dormancy breaking in Nicotiana plumbaginifolia seeds. Of the 15,000 cDNA fragments identified, 1,020 responded differentially to dormancy breaking or maintenance. The first seed germination–related study involving gene expression analysis using microarrays also dates from 2005 (Nakabayashi et al. 2005). This work performed on Arabidopsis seeds led to the identification of 12,470 seed-stored mRNAs. More than 10,000 of these transcripts were also detected at the different imbibition time-points investigated (6, 12, and 24 hours after imbibition). Differential expressed genes include those related to metabolism, transcription, cell cycle, and DNA processing. Moreover, the analyses of seeds at different stages during imbibition revealed multiple regulatory mechanisms, including epigenetic chromatin structures, co-regulated gene clusters, and cis-acting elements. The first RNA-seq study on seeds revealed 50% more differentially expressed genes then initially reported by Nakabayashi et al. (2005) based on microarray analyses (Narsai et al. 2017). In this section we will discuss transcriptome analyses in dormant and germinating seeds and focus mostly on genome-wide analyses.
Differences between dormant and nondormant seed transcriptomes
Seeds are full of stored mRNAs for which translation is considered important for germination to occur. The importance of stored transcripts for seed germination raised questions about the changes in the transcriptome, particularly when seeds transition from the dormant to the nondormant state during seed AR (dry storage). Although it has been debated whether transcriptional changes, especially upregulation, can occur in dry seeds, studies supporting such findings have been reported for N. plumbaginifolia, barley, and Arabidopsis seeds (Bove et al. 2005; Finch-Savage et al. 2007; Leymarie et al. 2007). More specifically, it has been suggested that specific mRNAs are being oxidized during AR. In sunflower seeds 24 oxidized transcripts have been identified that belong to the gene ontology classes metabolic processes, response to stress, and transport. Specific genes identified include protein phosphatase 2C PPH1, mitogen-activated protein kinase phosphatase 1, and phenyl ammonia lyase 1 (Bazin et al. 2011).
More obvious dormancy-specific transcriptomes can be revealed when seeds are investigated in the imbibed state. The comparison of dormant and AR Cvi seeds revealed an upregulation of genes related to translation, including genes that encode ribosomal proteins, translation initiation, and elongation factors in AR seeds (Cadman et al. 2006). This suggests that translation is turned down in dormant seeds, which has been confirmed by the analyses of translation by means of polysome profiling. In imbibed dormant seeds, translation starts like it does in AR seeds; however, it does not reach the same level as it does in nondormant seeds (Bai et al. 2018). Another attempt to identify differences between dormant and AR seeds revealed a set of 30 genes with similar expression profiles in the different physiological stages investigated; among these genes was the key dormancy regulator DOG1 (Finch-Savage et al. 2007). Although all these 30 genes in theory have the potential to be regulators of seed dormancy or seed germination, they are not reported as such. Only ERF1, which was associated with temperature-sensitive germination in lettuce and improved germination in saline conditions in rice, was described to be germination related (Yoong et al. 2016).
Investigating the temporal changes in dormant and nondormant seeds revealed that transcriptional patterns already differ at 12 hours of imbibition, when nondormant seeds prepare for germination (Dekkers et al. 2016a). After the initial imbibition, which leads to large transcriptional changes (up and down) in both dormant and nondormant seeds (approximately 1,000 and 2,000, respectively), most changes occur in nondormant seeds. Up to 2,000 transcripts change when comparing each time point with the previous, whereas in the dormant seeds this is limited to less than 100. The transcriptional profiles that discriminate dormant and nondormant seeds are found both in the embryo and endosperm and show great overlap with transcriptional changes in seeds in the soil seed bank that cycle in and out of dormancy, suggesting that laboratory-based experiments are a good alternative to study natural processes (Buijs et al. 2020). Transcriptional differences between physiological stages can be used to reveal genes that regulate processes of interest. An example of this is a reverse genetics approach that was chosen to identify the role of genes that are differentially expressed in 24-hour imbibed dormant and AR seeds of 5 different genotypes (Ler and 4 DOG NILs) (Yazdanpanah et al. 2017). Analyses in imbibed dormant and AR seeds in the same 5 genotypes revealed that dormancy maintenance and germination are likely to be very conserved processes (Bentsink et al. 2010). Of the total 3,689 differentially expressed genes, 1,720 were dormancy and 1,969 germination upregulated; of these 45 and 25 were common between all genotypes. Among these genes were the earlier identified germination related GA3OX2, PIF6, and ASG5. Moreover, for 50% of the genes for which T-DNA mutants were available, a germination phenotype could be observed.
Insights from network analyses
The initial transcriptome analyses did not yield new key regulators of seed dormancy and germination, and this became a reason for Bassel et al. (2011) to perform a coexpression analysis on publicly available expression data from imbibed mature seeds. The condition-dependent network SeedNet (http://netvis.ico2s.org/dev/seednet/#/) consisted of 1,583 transcripts associated with germination and 1,844 transcripts associated with nongermination (Fig. 2). The germination and nongermination subdomains were connected by a domain referred to as the intermediate transition region, which was enriched for genes involved in cellular phase transitions, including SERRATE and EARLY FLOWERING SHORT DAYS. Knockouts of the latter show both a flowering time and germination phenotype (Soppe et al. 1999). Overall, this work shows that network studies not only have the potential to identify co-regulated genes but also possible phase transitions.
Transcriptional phase transitions during seed germination were also identified by (Dekkers et al. 2013), who investigated spatial and temporal transcriptome changes during seed germination. Arabidopsis seeds were dissected into 4 sections: cotyledons, radicle and hypocotyl, micropylar and chalazal endosperm, and peripheral endosperm. These tissues were investigated from the start of seed imbibition until endosperm rupture at 10 time-points, of which at TR (25 HAI) and endosperm rupture (38 HAI), the transcriptomes of ruptured and nonruptured seeds had been separated. Whereas at 25 hours after imbibition (HAI) the transcriptome of the nonruptured seeds do not differ from those at 20 HAI, there are a large number of differential expressed genes, mostly upregulated, in the TR seeds compared with the nonruptured in both the embryo and endosperm tissues. These transcriptional changes separate germination into 2 transcriptional phases: before and after TR. The transcriptional changes over time have been captured by a coexpression network (Dekkers et al. 2013). The latter showed that coexpression networks form highly confident templates for hypothesis generation, which as such can be further scrutinized to unravel molecular networks. An example of this is the interaction mapping study performed for the cell wall–loosening EXPANSIN (EXPA) gene family (Xu et al. 2020). The EXPA gene family is associated with cell wall modification, and the expression of several family members is induced during Arabidopsis seed germination. One of these expansins, EXPA9 is regulated by the transcription factors TCP14 and TCP15. Xu et al. (2020) show that the reduced germination phenotype of the tcp14 tcp15 double mutant can be partially overcome by EXPA9 ectopic expression.
New developments in transcriptome analyses
Recently, we have entered a new era of transcriptome analyses: single-cell sequencing. For seeds this is not yet straightforward; however, a first attempt that includes imbibed and germinated seeds has recently been presented (Lee et al. 2023). In single-cell sequence analyses, cell types are annotated based on a priori known cell type–specific markers. For seeds, such a bona fide cell type–specific data set is not yet available; therefore the researchers made use of spatial transcriptome technologies, Slide-seqV2. The Slide-seqV2 method is based on a combination of detecting RNAs at a spatial resolution of 10 μm (Slide-seq) with single-cell trajectory analysis tools (Rodriques et al. 2019; Stickels et al. 2021). Using this methodology, it was revealed that the identified single-cell clusters spatially mapped to the cotyledons, root tip region, epidermis, seed coat, and provasculature. As such this dataset has the potential to function as foundation for future spatial transcriptome analyses in seeds. Important to consider when performing detailed transcriptome analyses is the material used for these investigations. Seeds are highly dynamic in their response to the environment, the changes they show over time (e.g. the differential expression when comparing TR vs nonruptured seeds from the same imbibition time-point) and on top of that show a large germination variability. With this we refer for example to the variation that occurs in a population of seeds harvested from a single plant (Mitchell et al. 2017). This variability likely contributes to the seed's role as a survival structure for seed plants and is nicely demonstrated by single-seed RNA-seq analyses performed on seeds during secondary dormancy induction (Krzyszton et al. 2022). Transcriptional variability is the lowest in seeds that have been exposed to the secondary dormancy, inducing stress, and is the largest in seeds that have recovered from this stress and have started germination (Fig. 2). Single-seed RNA-seq in combination with single-cell analyses might help to reveal the molecular basis of variability in seed pools at a cellular resolution.
Epigenetic control of seed dormancy and germination
An important aspect of gene regulation is that controlled by epigenetic mechanisms. It has been reported that this mechanism, which includes DNA methylation of cytosines, histone, and in general chromatin modifications, controls important genes during seed development that affect dormancy and subsequent seed germination (Sato et al. 2021; Ding et al. 2022; Iwasaki et al. 2022; Sato and Köhler 2022). The function of many regulators of these processes had been described before using methods such as methylC-sequencing and ChIP-sequencing applied on mutants of these regulators. Often the respective mutants were used to test their effect on the above-mentioned seed development, seed dormancy, and germination processes leading to conclusions on the role of epigenetic mechanisms.
Epigenetic regulation of seed maturation
Epigenetic control in seeds starts with the differential gene expression in the 2 gametes that together form the zygote and quicky develops into an embryo. This differential parental silencing distinguishes between maternal expressed genes (MEGs) and paternally expressed genes (PEGs). Two histone demethylases, RELATIVE OF EARLY FLOWERING 6 (REF6) and EARLY FLOWERING 6 (ELF6), control endosperm dormancy by activating the MEGs (Chen et al. 2020; Sato and Köhler 2022). A recent study analyzing natural variation for seed dormancy in Arabidopsis identified that VERNALIZATION5/VIN3-LIKE 3 (VEL3) maintains seed dormancy by establishing an epigenetic state in the central cell that primes the depth of primary seed dormancy, which is established later during seed maturation (Chen et al. 2023). VEL3 is required for deacetylation and the tri-methylation of lysine 27 on histone H3 (H3K27me3) established in the central cell. The authors propose that VEL3 maintains a repressive epigenetic state at ORE1 (OBSOLETE1) and other genes with a role in suppressing programmed cell death and inducing seed dormancy (Chen et al. 2023).
The role of epigenetic control during seed maturation included that of the major regulators of seed maturation LEC1, LEC2, FUS3, and ABI3 (Ding et al. 2022). Genetic studies subsequently have uncovered several loss-of-function mutations of chromatin-related genes that lead to ectopic expression of embryo-associated genes. These include mutants of the HISTON 2B MONO-UBIQUINATION genes HUB1 and HUB2, resulting in reduced seed dormancy and, among others, downregulation of the dormancy gene DOG1 (Liu et al. 2007). The proteins encoded by HUB1 and HUB2 most likely bind the polymerase II-associated factor 1 complex (PAF1C). This complex modulates the local structure of chromatin during transcription elongation and affects methylation of histone H3 at lysine 4 and 36 (H3K4me and H3K36me), respectively, which are activating epigenetic marks for transcription. Interestingly, the isolation and genetic analysis of additional factors associated with PAF1C, like REDUCED DORMANCY 2 in Arabidopsis, confirmed the important role of transcription elongation factors in seed maturation (Liu et al. 2007). Mutations in the KRYPTONITE (KYP)⁄SU(VAR)3-9 HOMOLOG 4 (SUVH4) gene, encoding the histone methyltransferase for H3K9me2, cause increased seed dormancy. In accordance, KYP⁄SUVH4-overexpressing Arabidopsis plants show decreased dormancy (Zheng et al. 2012). This indicates that this repressive chromatin mark influences seed maturation. Potential direct targets of KYP⁄SUVH4 are the seed maturation genes ABI3 and DOG1, which are upregulated in the kyp-2 mutant.
DELAY OF GERMINATION-LIKE 4 (DOGL4) is a central factor mediating reserve accumulation in seeds without strongly affecting seed dormancy and other processes controlled by DOG1 (Sall et al. 2019). Zhu et al. (2018) described that the DNA demethylase ROS1 regulates the imprinting of DOGL4. DOGL4 is expressed from the MEG and displays preferential methylation and suppression of the paternal allele. ROS1 negatively regulates imprinting by demethylating the paternal allele, preventing its hypermethylation and complete silencing. The dormancy effect of the dogl4 mutant described by these authors is relatively weak compared with what is described for the dog1 mutants (Sall et al. 2019). Epigenetic control of the dormancy master regulator DOG1 was recently summarized by Ding et al. (2022).
The process of epigenetic silencing during germination involves the POLYCOMB REPRESSIVE COMPLEX 2 (PRC2), which plays a role in the repression of flowering in young seedlings by mediating H3K27me3. Mutants in FERTILIZATION INDEPENDENT ENDOSPERM, an essential component of the PRC2 complex, displayed genome-wide abolishment of H3K27me3 and exhibited increased seed dormancy and germination defects (Bouyer et al. 2011). Therefore, it was concluded that PRC2 is required for termination of the embryonic transcriptional program to promote the phase transition from embryo to autotrophic seedling. PRC2 sustains the balance between ABA and GA responsiveness via H3K27me3-mediated inhibition of positive ABA and negative GA regulators in maturing seeds. Moreover, the seed dormancy regulator DOG1 is repressed through PRC2-catalyzed H3K27- trimethylation at its locus (Bouyer et al. 2011).
Histone acetylation is another modification that plays a role in silencing of embryonic traits and activating genes associated with seedling establishment and growth. In Arabidopsis, HDA6 and HDA19 act redundantly to repress embryonic properties after germination (Tanaka et al. 2008). The function of HDA9 is opposite to that of its homologous genes HDA6 and HDA19 (van Zanten et al. 2014). HDA9 negatively influences germination and is involved in the suppression of seedling traits in dry seeds. The HDA9 transcript is abundant in dry seeds and becomes reduced during imbibition in wild-type seeds.
Epigenetic effects during seed germination
Dry seeds contain nuclei with reduced size and highly compacted chromatin; this phenomenon occurs during maturation and is therefore related to the acquisition of desiccation tolerance and seems independent from the dormancy state (van Zanten et al. 2012). Transcriptional reprogramming during germination requires repression of embryonic properties and activation of genes involved in the progression into photoautotrophic growth. Independent papers demonstrated roles for chromatin-remodeling factors and epigenetic signaling mechanisms in these processes. Seedlings of mutants lacking the CHD3 class SWI/SNF chromatin-remodeling factor PICKLE (PKL) display embryonic properties, that is, “pickle” roots, with a swollen and greenish distal root tip and accumulation of neutral lipid bodies (Ogas et al. 1999). PKL represses the expression of ABI3 and ABI5 in an ABA-dependent manner, as pkl mutants showed high ABI3 and ABI5 transcript levels after ABA treatment and hypersensitivity to ABA-mediated repression of germination (Perruc et al. 2007). The abundance of these epigenetic silencing marks was reduced in pkl mutants compared with the corresponding wild types, which was further enhanced by the application of ABA (Perruc et al. 2007). A direct role for PKL in determining levels of H3K27me3 at these repressed loci during germination was demonstrated (Zhang et al. 2012). Taken together, PKL controls ABA-mediated repression of germination by affecting the abundance of silencing epigenetic histone marks associated with the ABI3 and ABI5 loci. A detailed time course of germination using RNA seq showed that changes in gene expression and large-scale demethylation was observed toward the end of germination during the transitions from an embryo-like to a vegetative seedling state (Narsai et al. 2017).
Translational regulation in seeds
As mentioned earlier, during seed maturation there is selective accumulation of mRNA also known as “stored mRNAs” or “long-lived mRNAs” (Ajtkhozhin et al. 1976; Ishibashi et al. 1990; Kuligowski et al. 1991). These stored mRNAs serve as a reservoir for rapid translation into proteins during germination (Ishibashi et al. 1990; Kuligowski et al. 1991; Rajjou et al. 2004; Sano et al. 2012). When nondormant Arabidopsis seeds were imbibed in a solution containing the transcriptional inhibitor α-amanitin, germination could still take place (Fig. 2). This indicated that the long-lived or stored mRNAs within the dry nondormant seed are sufficient to produce proteins required for successful germination. However, when Arabidopsis and rice seeds were imbibed in a solution containing the translational inhibitor cycloheximide, germination was blocked (Fig. 2; Rajjou et al. 2004; Sano et al. 2012, 2015). Therefore, the process of translation is crucial for germination to occur.
With the advent of technologies like ribosome profiling, studying the translational status of all mRNAs in seeds became possible. This technique employs sucrose-based fractions to separate mRNAs based on their association to no ribosomes (free mRNA), 1 ribosome (monosome), or multiple ribosomes (polysomes). mRNAs present in the polysome fraction most likely indicate active translation of those mRNAs into proteins. Using this technique, it was observed that dormant and nondormant seeds show differential association of mRNAs to polysomes (Layat et al. 2014). In the dormant embryos, negative regulators of germination seemed to be associated with polysomes, for example, transcription factors promoting ABA responses like ABI2 and LEA proteins, which should disappear during germination. Moreover, Bai et al. (2018) showed that translation was more abundant in AR Arabidopsis seeds than in dormant seeds at 24HAI. The addition of the transcriptional inhibitor cordycepin resulted in similar levels of translation in dormant and AR seeds, suggesting that transcriptional changes are required for the increased translational activity, which is seen in the nondormant seeds in control conditions. Based on this it was concluded that dormancy maintenance or alleviation is regulated at the transcriptional level (Bai et al. 2018).
Translational regulation, the control of the protein levels produced from its mRNA, is a highly efficient and reversible way of fine-tuning gene expression in response to developmental and environmental (Sablok et al. 2017; Chantarachot and Bailey-Serres 2018). To identify the translational efficiency of transcript one can determine the ratio of the mRNA abundance in the polysome fraction to the total RNA abundance of a given transcript between 2 timepoints or treatments (King and Gerber 2016). Using this approach, selective translation during seed maturation was reported (Bai et al. 2023). Among the translationally regulated genes are ABI2 and ABI5 (Finkelstein 1994; Finkelstein and Lynch 2000). It has also been reported that within the mature dry seed, most stored mRNAs are associated to monosomes and many are translationally upregulated upon imbibition. This indicates that these mRNAs are important for germination. mRNAs associated with polysomes in the dry seed are degraded during germination, suggesting that these are remnants from the maturation program and thus no longer needed for germination (Bai et al. 2020).
The first major study that implicated a role for translational regulation during germination came from Galland et al. (2014). In their study, they observed that differential accumulation of proteins occurred during early germination. The authors compared the transcript abundance, the rate of protein synthesis, and the protein abundance of 158 newly synthesized radiolabeled proteins during Arabidopsis seed germination. The protein abundance of these proteins, which included proteins involved in mRNA translation (10% of the identified proteins), Late Embryogenesis Abundant, and Heat Shock Proteins, correlated very well with the rate of protein synthesis and poorly with the amount of its transcript, thereby indicating that translational regulation occurs during germination for newly synthesized proteins (Galland et al. 2014). Another study performed in rice seeds showed that from the 20 proteins that were synthesized upon from the stored/long-lived mRNAs present dry seeds, only 12 of these were upregulated 2 days after imbibition, and the rest were upregulated 4 days after imbibition. Similarly, in Arabidopsis seeds, using ribosome profiling, Bai et al. (2017) showed that there is extensive sequence-specific translational regulation occurring during 2 shifts of germination, namely the hydration translational shift (HTS) and the germination translational shift (GTS). The HTS shift spans the transition from a dry seed to an imbibed seed, whereas the GTS marks the phase between 2 physiologically visible stages of germination, namely, TR and RP. The pools of mRNA that were translationally regulated were different in these 2 shifts, indicating independent mechanisms (Bai et al. 2017). The purpose of this regulation only in these 2 shifts of germination has not yet been established. However, during the HTS, the seed goes from a completely dry to a fully hydrated imbibed seed. Several metabolic processes must be kickstarted, and translation is initiated selectively from stored mRNA. This could be the reason why translational regulation occurs during the HTS, whereas during the GTS, the seed must make the all or nothing commitment to germinate and establish a seedling. This is a complex process based on several factors including a dynamically changing environment, which is why extensive translational regulation takes place during the GTS. Several mRNA features like upstream ORFs, transcript length, GC content, and secondary structures were correlated with an upregulation of translation.
Mechanisms underlying translational control: the role of RNA binding proteins
The precise mechanism involved in the differential translation of mRNAs during germination is not known; however, RNA binding proteins (RBPs) are emerging as key players in the translational control of gene expression in plants (Hentze et al. 2018; Cho et al. 2019; Lou et al. 2020; Sajeev et al. 2022). RBPs are proteins that can bind target mRNAs through their RNA binding domain and regulate the fate and functions of their target mRNA (Lorković 2009; Li et al. 2014; Köster et al. 2017; Lou et al. 2020). Examples of these domains include RNA recognition motif, Pumilio/FBF domain, DEAD box domain, the K homology domain, and the Pentatricopeptide repeat motif.
RBPs have been implicated in seed development and performance (extensively reviewed by Lou et al. 2020). Some examples include RBPs like GLYCINE RICH PROTEIN 2 that can promote seed germination in stress conditions like low temperature via an ABA-independent pathway, whereas the overexpression of RNA DEAD box (RHs) helicases like RH9 and RH29 inhibit germination in salt conditions (Kim et al. 2005, 2007). APUM9, a conventional RBP from the PUMILIO family, plays a role in promoting germination in optimal conditions possibly by targeting ABA signaling genes for degradation (Xiang et al. 2014; Nyikó et al. 2019; Huang et al. 2021), and APUM5 plays a role in inhibiting seed germination in salt and osmotic stress (Huh and Paek 2014). The RBPs VARICOSE (VCS) and EXORIBONUCLEASE4 also play a role in seed dormancy and germination. The mutant seeds of VCS were less dormant, and those of xrn-4 were more dormant (Basbouss-Serhal et al. 2017). Transcriptomic analyses of the vcs mutant seeds showed higher abundance of transcripts promoting germination like WRKY60 and WRKY63, which are TFs that negatively regulate ABA signaling, while xrn-4 seeds displayed higher transcript levels of genes involved in dormancy maintenance like ABA response elements ABF3 and ABP4 (Basbouss-Serhal et al. 2017). Interestingly, VCS and XRN are also components of higher-order RBP-mRNA complexes called processing bodies (P-bodies). These are membrane-less cytoplasmic granules that can transiently store or decay translationally repressed mRNAs (Maldonado-Bonilla 2014; Jang et al. 2020). Jang et al. (2019) reported that P-bodies control translation of certain mRNAs and are required for optimal seed to seedling transition.
Recently, the first repertoire of RBPs in germinating seeds that could provide potential candidates for future research was reported. This study employed an mRNA interactome capture approach that led to the identification of more than 200 high confidence RBPs present during the GTS (the phase between TR and RP) in seeds (GTS-RBPs) and 30 GTS-RBPs to be present specifically in seeds (Sajeev et al. 2022). Some of the identified RBPs were known to have a seed germination and dormancy phenotype. For instance, TUDOR SN1/2 has been shown to promote seed germination under salt stress by regulating the mRNA levels of a key GA biosynthesis enzyme, GA20ox3 (Maldonado-Bonilla 2014), and EIN2 has been reported to play a strong role in alleviating dormancy (Beaudoin et al. 2000; Ghassemian et al. 2000). This study also identified an RBP from the hyaluronan/mRNA, namely, AT5G47210, and showed that it plays a role in inhibiting germination under optimal conditions via an unknown mechanism (Sajeev et al. 2022).
Concluding remarks and perspectives
In this review we provided an overview of the research that led to the current knowledge on the regulation of seed dormancy and germination in the past century. Still major questions remain to be solved. At the level of inducing and overcoming seed dormancy, ABA and DOG1 are major components. Although a lot is known about the ABA perception, the role of ABI3 in this has remained unclear. The same holds for the mechanism of the balance between ABA and GA. On the whole, genetic and natural variation studies in Arabidopsis have contributed significantly to our current knowledge on the regulation of seed dormancy and germination. Moreover, although not yet fully proven, there are strong indications that oxidative processes during AR change the dormancy state, which upon seed imbibition allow the seeds to germinate. The onset of germination itself seems more complicated. Nondormant seeds do not germinate if the environment is not optimal; however, the on/off switch for the decision to germinate has not yet been identified. Factors that could play a role in this are downstream genes of GA signaling, like TCPs and EXPAs; however, seeds contain many copies of these genes, and knocking out a single component generally only results in a small change in germination rate (Xu et al. 2020). Is this redundancy required to allow spreading the risk associated with germination in time and space, even in a single population, and does that mean that such an on/off switch does not exist? Or do we so far not have the insights or tools to identify this component? The use of single-cell analyses and mathematical modeling at the (sub)cellular scale might allow us to address this question. The component(s) that we are looking for may very well lie at the level of translation, as can be concluded from the strong translational control of seed germination at 2 critical physiological phases during seed germination (Bai et al. 2017). Employing techniques like interactome capture and individual-nucleotide resolution UV crosslinking and immunoprecipitation in seeds will advance our understanding and contribute to revealing the role of translational regulation in the control of seed germination. So far, technical difficulties have hampered advances in applying these high-tech protocols in seeds.
Acknowledgments
N.S. and L.B. were supported the project Seeds4Ever: Protection of stored mRNAs to ensure seed survival (with project number 17047) of the research programme Vici which is financed by the Dutch Research Council (NWO).
Contributor Information
Nikita Sajeev, Wageningen Seed Science Centre, Laboratory of Plant Physiology, Wageningen University, 6708PB Wageningen, the Netherlands.
Maarten Koornneef, Laboratory of Genetics, Wageningen University, 6708PB Wageningen, the Netherlands; Max Planck Institute for Plant Breeding Research, Former Department of Plant Breeding and Genetics, Koeln 50829, Germany.
Leónie Bentsink, Wageningen Seed Science Centre, Laboratory of Plant Physiology, Wageningen University, 6708PB Wageningen, the Netherlands.
Author contributions
N.S., M.K., and L.B. conceptualized the review outline, and wrote the original draft, L.B. prepared the figures, M.K. and L.B. reviewed and edited the manuscript.
Dive Curated Terms
The following phenotypic, genotypic, and functional terms are of significance to the work described in this paper:
References
- Abley K, Formosa-Jordan P, Tavares H, Chan EYT, Afsharinafar M, Leyser O, Locke JCW. An ABA-GA bistable switch can account for natural variation in the variability of Arabidopsis seed germination time. eLife. 2021:10:e59485. 10.7554/eLife.59485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ajtkhozhin MA, Doschanov KJ, Akhanov AU. Informosomes as a stored form of mRNA in wheat embryos. FEBS Lett. 1976:66(1):124–126. 10.1016/0014-5793(76)80600-X [DOI] [PubMed] [Google Scholar]
- Ali-Rachedi S, Bouinot D, Wagner MH, Bonnet M, Sotta B, Grappin P, Jullien M. Changes in endogenous abscisic acid levels during dormancy release and maintenance of mature seeds: studies with the Cape Verde Islands ecotype, the dormant model of Arabidopsis thaliana. Planta. 2004:219(3):479–488. 10.1007/s00425-004-1251-4 [DOI] [PubMed] [Google Scholar]
- Alonso-Blanco C, Bentsink L, Hanhart CJ, Vries HBE, Koornneef M. Analysis of natural allelic variation at seed dormancy loci of Arabidopsis thaliana. Genetics. 2003:164(2):711–729. 10.1093/genetics/164.2.711 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amiguet-Vercher A, Santuari L, Gonzalez-Guzman M, Depuydt S, Rodriguez PL, Hardtke CS. The IBO germination quantitative trait locus encodes a phosphatase 2C-related variant with a nonsynonymous amino acid change that interferes with abscisic acid signaling. New Phytol. 2015:205(3):1076–1082. 10.1111/nph.13225 [DOI] [PubMed] [Google Scholar]
- Arc E, Galland M, Godin B, Cueff G, Rajjou L. Nitric oxide implication in the control of seed dormancy and germination. Front Plant Sci. 2013:4:346. 10.3389/fpls.2013.00346 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bai B, Novak O, Ljung K, Hanson J, Bentsink L. Combined transcriptome and translatome analyses reveal a role for tryptophan-dependent auxin biosynthesis in the control of DOG1-dependent seed dormancy. New Phytol. 2018:217(3):1077–1085. 10.1111/nph.14885 [DOI] [PubMed] [Google Scholar]
- Bai B, Peviani A, van der Horst S, Gamm M, Snel B, Bentsink L, Hanson J. Extensive translational regulation during seed germination revealed by polysomal profiling. New Phytol. 2017:214(1):233–244. 10.1111/nph.14355 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bai B, Schiffthaler B, van der Horst S, Willems L, Vergara A, Karlström J, Mähler N, Delhomme N, Bentsink L, Hanson J. SeedTransNet: a directional translational network revealing regulatory patterns during seed maturation and germination. J Exp Bot. 2023:74(7):2416–2432. 10.1093/jxb/erac394 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bai B, van der Horst S, Cordewener JHG, America T, Hanson J, Bentsink L. Seed-Stored mRNAs that are specifically associated to monosomes are translationally regulated during germination. Plant Physiol. 2020:182(1):378–392. 10.1104/pp.19.00644 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Basbouss-Serhal I, Pateyron S, Cochet F, Leymarie J, Bailly C. 5’ to 3’ mRNA decay contributes to the regulation of arabidopsis seed germination by dormancy. Plant Physiol. 2017:173(3):1709–1723. 10.1104/pp.16.01933 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baskin JM, Baskin CC. A classification system for seed dormancy. Seed Sci Res. 2004:14(1):1–16. 10.1079/SSR2003150 [DOI] [Google Scholar]
- Bassel GW, Lan H, Glaab E, Gibbs DJ, Gerjets T, Krasnogor N, Bonner AJ, Holdsworth MJ, Provart NJ. Genome-wide network model capturing seed germination reveals coordinated regulation of plant cellular phase transitions. Proc Natl Acad Sci U S A. 2011:108(23):9709–9714. 10.1073/pnas.1100958108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bazin J, Langlade N, Vincourt P, Arribat S, Balzergue S, El-Maarouf-Bouteau H, Bailly C. Targeted mRNA oxidation regulates sunflower seed dormancy alleviation during dry after-ripening. Plant Cell. 2011:23(6):2196–2208. 10.1105/tpc.111.086694 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beaudoin N, Serizet C, Gosti F, Giraudat J. Interactions between abscisic acid and ethylene signaling cascades. Plant Cell. 2000:12(7):1103–1115. 10.1105/tpc.12.7.1103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bentsink L, Hanson J, Hanhart CJ, Blankestijn-de Vries H, Coltrane C, Keizer P, El-Lithy M, Alonso-Blanco C, Teresa de Andres M, Reymond M, et al. Natural variation for seed dormancy in Arabidopsis is regulated by additive genetic and molecular pathways. Proc Natl Acad Sci U S A. 2010:107(9):4264–4269. 10.1073/pnas.1000410107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bentsink L, Jowett J, Hanhart CJ, Koornneef M. Cloning of DOG1, a quantitative trait locus controlling seed dormancy in Arabidopsis. Proc Natl Acad Sci U S A. 2006:103(45):17042–17047. 10.1073/pnas.0607877103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bewley JD. Seed germination and dormancy. Plant Cell. 1997:9:1055–1066. 10.1105/tpc.9.7.1055 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bewley JD, Bradford KJ, Hilhorst HW, Nonogaki H. Seeds. New York: Springer; 2012. [Google Scholar]
- Biere A. Parental effects in Lychnis flos-cuculi. I: Seed size, germination and seedling performance in a controlled environment. J Evol Biol. 1991:4(3):447–465. 10.1046/j.1420-9101.1991.4030447.x [DOI] [Google Scholar]
- Boulard C, Fatihi A, Lepiniec L, Dubreucq B. Regulation and evolution of the interaction of the seed B3 transcription factors with NF-Y subunits. Biochim Biophys Acta Gene Regul Mech. 2017:1860(10):1069–1078. 10.1016/j.bbagrm.2017.08.008 [DOI] [PubMed] [Google Scholar]
- Bouyer D, Roudier F, Heese M, Andersen ED, Gey D, Nowack MK, Goodrich J, Renou J-P, Grini PE, Colot V, et al. Polycomb Repressive Complex 2 controls the embryo-to-seedling phase transition. PLoS Genet. 2011:7(3):e1002014. 10.1371/journal.pgen.1002014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bove J, Lucas P, Godin B, Oge L, Jullien M, Grappin P. Gene expression analysis by cDNA-AFLP highlights a set of new signaling networks and translational control during seed dormancy breaking in Nicotiana plumbaginifolia. Plant Mol Biol. 2005:57(4):593–612. 10.1007/s11103-005-0953-8 [DOI] [PubMed] [Google Scholar]
- Bryant FM, Hughes D, Hassani-Pak K, Eastmond PJ. Basic LEUCINE ZIPPER TRANSCRIPTION FACTOR67 transactivates DELAY OF GERMINATION1 to establish primary seed dormancy in Arabidopsis. Plant Cell. 2019:31(6):1276–1288. 10.1105/tpc.18.00892 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buijs G. A perspective on secondary seed dormancy in Arabidopsis thaliana. Plants (Basel). 2020:9(6):749. 10.3390/plants9060749 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buijs G, Vogelzang A, Nijveen H, Bentsink L. Dormancy cycling: translation-related transcripts are the main difference between dormant and non-dormant seeds in the field. Plant J. 2020:102(2):327–339. 10.1111/tpj.14626 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cadman CS, Toorop PE, Hilhorst HW, Finch-Savage WE. Gene expression profiles of Arabidopsis Cvi seeds during dormancy cycling indicate a common underlying dormancy control mechanism. Plant J. 2006:46(5):805–822. 10.1111/j.1365-313X.2006.02738.x [DOI] [PubMed] [Google Scholar]
- Carbonero P, Iglesias-Fernández R, Vicente-Carbajosa J. The AFL subfamily of B3 transcription factors: evolution and function in angiosperm seeds. J Exp Bot. 2017:68(4):871–880. 10.1093/jxb/erw458 [DOI] [PubMed] [Google Scholar]
- Carles C, Bies-Etheve N, Aspart L, Léon-Kloosterziel KM, Koornneef M, Echeverria M, Delseny M. Regulation of Arabidopsis thaliana Em genes: role of ABI5. Plant J. 2002:30(3):373–383. 10.1046/j.1365-313X.2002.01295.x [DOI] [PubMed] [Google Scholar]
- Carrillo-Barral N, Rodríguez-Gacio MDC, Matilla AJ. Delay of Germination-1 (DOG1): a key to understanding seed dormancy. Plants (Basel). 2020:9(4):480. 10.3390/plants9040480 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chandrasekaran U, Luo X, Zhou W, Shu K. Multifaceted signaling networks mediated by Abscisic Acid Insensitive 4. Plant Commun. 2020:1(3):100040. 10.1016/j.xplc.2020.100040 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chantarachot T, Bailey-Serres J. Polysomes, Stress Granules, and Processing Bodies: a dynamic triumvirate controlling cytoplasmic mRNA fate and function. Plant Physiol. 2018:176(1):254–269. 10.1104/pp.17.01468 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen H, Tong J, Fu W, Liang Z, Ruan J, Yu Y, Song X, Yuan L, Xiao L, Liu J, et al. The H3K27me3 demethylase RELATIVE OF EARLY FLOWERING6 suppresses seed dormancy by inducing abscisic acid catabolism. Plant Physiol. 2020:184(4):1969–1978. 10.1104/pp.20.01255 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen X, MacGregor DR, Stefanato FL, Zhang N, Barros-Galvão T, Penfield S. A VEL3 histone deacetylase complex establishes a maternal epigenetic state controlling progeny seed dormancy. Nat Commun. 2023:14(1):2220. 10.1038/s41467-023-37805-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiabrando D, Vinchi F, Fiorito V, Mercurio S, Tolosano E. Heme in pathophysiology: a matter of scavenging, metabolism and trafficking across cell membranes. Front Pharmacol. 2014:5:61. 10.3389/fphar.2014.00061 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cho H, Cho HS, Hwang I. Emerging roles of RNA-binding proteins in plant development. Curr Plant Biol. 2019:51:51–57. 10.1016/j.pbi.2019.03.016 [DOI] [PubMed] [Google Scholar]
- Cutler SR, Rodriguez PL, Finkelstein RR, Abrams SR. Abscisic acid: emergence of a core signaling network. Annu Rev Plant Biol. 2010:61(1):651–679. 10.1146/annurev-arplant-042809-112122 [DOI] [PubMed] [Google Scholar]
- Cyrek M, Fedak H, Ciesielski A, Guo Y, Sliwa A, Brzezniak L, Krzyczmonik K, Pietras Z, Kaczanowski S, Liu F, et al. Seed dormancy in Arabidopsis is controlled by alternative polyadenylation of DOG1. Plant Physiol. 2016:170(2):947–955. 10.1104/pp.15.01483 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Debeaujon I, Koornneef M. Gibberelin requirement for Arabidopsis thaliana seed germination is determined both by testa characteristics and embryonic ABA. Plant Physiol. 2000:122(2):415–424. 10.1104/pp.122.2.415 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Debieu M, Tang C, Stich B, Sikosek T, Effgen S, Josephs E, Schmitt J, Nordborg M, Koornneef M, de Meaux J. Co-variation between seed dormancy, growth rate and flowering time changes with latitude in Arabidopsis thaliana. PLoS One. 2013:8(5):e61075. 10.1371/journal.pone.0061075 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dekkers BJ, He H, Hanson J, Willems LA, Jamar DC, Cueff G, Rajjou L, Hilhorst HW, Bentsink L. The Arabidopsis DELAY OF GERMINATION 1 gene affects ABSCISIC ACID INSENSITIVE 5 (ABI5) expression and genetically interacts with ABI3 during Arabidopsis seed development. Plant J. 2016b:85(4):451–465. 10.1111/tpj.13118 [DOI] [PubMed] [Google Scholar]
- Dekkers BJ, Pearce SP, van Bolderen-Veldkamp RP, Holdsworth MJ, Bentsink L. Dormant and after-ripened Arabidopsis thaliana seeds are distinguished by early transcriptional differences in the imbibed state. Front Plant Sci. 2016a:7:1323. 10.3389/fpls.2016.01323 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dekkers BJ, Pearce S, van Bolderen-Veldkamp RP, Marshall A, Widera P, Gilbert J, Drost HG, Bassel GW, Muller K, King JR, et al. Transcriptional dynamics of two seed compartments with opposing roles in Arabidopsis seed germination. Plant Physiol. 2013:163(1):205–215. 10.1104/pp.113.223511 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng G, Sun H, Hu Y, Yang Y, Li P, Chen Y, Zhu Y, Zhou Y, Huang J, Neill SJ, et al. A transcription factor WRKY36 interacts with AFP2 to break primary seed dormancy by progressively silencing DOG1 in Arabidopsis. New Phytol. 2023:238(2):688–704. 10.1111/nph.18750 [DOI] [PubMed] [Google Scholar]
- Ding X, Jia X, Xiang Y, Jiang W. Histone modification and chromatin remodeling during the seed life cycle. Front Plant Sci. 2022:13:865361. 10.3389/fpls.2022.865361 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donohue K. Seeds and seasons: interpreting germination timing in the field. Seed Sci Res. 2005:15(3):175–187. 10.1079/SSR2005208 [DOI] [Google Scholar]
- Fedak H, Palusinska M, Krzyczmonik K, Brzezniak L, Yatusevich R, Pietras Z, Kaczanowski S, Swiezewski S. Control of seed dormancy in Arabidopsis by a cis-acting noncoding antisense transcript. Proc Natl Acad Sci U S A. 2016:113(48):E7846–E7855. 10.1073/pnas.1608827113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fernandez-Pascual E, Jimenez-Alfaro B, Caujape-Castells J, Jaen-Molina R, Diaz TE. A local dormancy cline is related to the seed maturation environment, population genetic composition and climate. Ann Bot. 2013:112(5):937–945. 10.1093/aob/mct154 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finch-Savage WE, Cadman CS, Toorop PE, Lynn JR, Hilhorst HW. Seed dormancy release in Arabidopsis cvi by dry after-ripening, low temperature, nitrate and light shows common quantitative patterns of gene expression directed by environmentally specific sensing. Plant J. 2007:51(1):60–78. 10.1111/j.1365-313X.2007.03118.x [DOI] [PubMed] [Google Scholar]
- Finch-Savage WE, Leubner-Metzger G. Seed dormancy and the control of germination. New Phytol. 2006:171(3):501–523. 10.1111/j.1469-8137.2006.01787.x [DOI] [PubMed] [Google Scholar]
- Finkelstein RR. Mutations at two new Arabidopsis ABA response loci are similar to the abi3 mutations. Plant J. 1994:5(6):765–771. 10.1046/j.1365-313X.1994.5060765.x [DOI] [Google Scholar]
- Finkelstein RR, Lynch TJ. The Arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell. 2000:12(4):599–609. 10.1105/tpc.12.4.599 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finkelstein R, Reeves W, Ariizumi T, Steber C. Molecular aspects of seed dormancy. Annu Rev Plant Biol. 2008:59(1):387–415. 10.1146/annurev.arplant.59.032607.092740 [DOI] [PubMed] [Google Scholar]
- Footitt S, Walley PG, Lynn JR, Hambidge AJ, Penfield S, Finch-Savage WE. Trait analysis reveals DOG1 determines initial depth of seed dormancy, but not changes during dormancy cycling that result in seedling emergence timing. New Phytol. 2020:225(5):2035–2047. 10.1111/nph.16081 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Francoz E, Lepiniec L, North HM. Seed coats as an alternative molecular factory: thinking outside the box. Plant Reprod. 2018:31(3):327–342. 10.1007/s00497-018-0345-2 [DOI] [PubMed] [Google Scholar]
- Galland M, Huguet R, Arc E, Cueff G, Job D, Rajjou L. Dynamic proteomics emphasizes the importance of selective mRNA translation and protein turnover during Arabidopsis seed germination. Mol Cell Proteomics. 2014:13(1):252–268. 10.1074/mcp.M113.032227 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghassemian M, Nambara E, Cutler S, Kawaide H, Kamiya Y, McCourt P. Regulation of abscisic acid signaling by the ethylene response pathway in Arabidopsis. Plant Cell. 2000:12(7):1117–1126. 10.1105/tpc.12.7.1117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graeber K, Linkies A, Steinbrecher T, Mummenhoff K, Tarkowská D, Turečková V, Ignatz M, Sperber K, Voegele A, de Jong H, et al. DELAY OF GERMINATION 1 mediates a conserved coat-dormancy mechanism for the temperature- and gibberellin-dependent control of seed germination. Proc Natl Acad Sci U S A. 2014:111(34):E3571–E3580. 10.1073/pnas.1403851111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graeber K, Nakabayashi K, Miatton E, Leubner-Metzger G, Soppe WJ. Molecular mechanisms of seed dormancy. Plant Cell Environ. 2012:35(10):1769–1786. 10.1111/j.1365-3040.2012.02542.x [DOI] [PubMed] [Google Scholar]
- Groot SP, Karssen CM. Gibberellins regulate seed germination in tomato by endosperm weakening: a study with gibberellin-deficient mutants. Planta. 1987:171(4):525–531. 10.1007/BF00392302 [DOI] [PubMed] [Google Scholar]
- Groot SP, Kieliszewska-Rokicka B, Vermeer E, Karssen CM. Gibberellin-induced hydrolysis of endosperm cell walls in gibberellin-deficient tomato seeds prior to radicle protrusion. Planta. 1988:174(4):500–504. 10.1007/BF00634479 [DOI] [PubMed] [Google Scholar]
- Guo R, Wen X, Zhang W, Huang L, Peng Y, Jin L, Han H, Zhang L, Li W, Guo H. Arabidopsis EIN2 represses ABA responses during germination and early seedling growth by inactivating HLS1 protein independently of the canonical ethylene pathway. Plant J. 2023:115(6):1514–1527. 10.1111/tpj.16335 [DOI] [PubMed] [Google Scholar]
- Gutierrez L, Van Wuytswinkel O, Castelain M, Bellini C. Combined networks regulating seed maturation. Trends Plant Sci. 2007:12(7):294–300. 10.1016/j.tplants.2007.06.003 [DOI] [PubMed] [Google Scholar]
- Harada JJ, Baden CS, Comai L. Spatially regulated genes expressed during seed germination and postgerminative development are activated during embryogeny. Mol Gen Genet. 1988:212(3):466–473. 10.1007/BF00330851 [DOI] [Google Scholar]
- He H. Environmental regulation of seed performance. Wageningen, The Netherlands: Wageningen University; 2014. [Google Scholar]
- He H, de Souza Vidigal D, Snoek LB, Schnabel S, Nijveen H, Hilhorst H, Bentsink L. Interaction between parental environment and genotype affects plant and seed performance in Arabidopsis. J Exp Bot. 2014:65(22):6603–6615. 10.1093/jxb/eru378 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hentze MW, Castello A, Schwarzl T, Preiss T. A brave new world of RNA-binding proteins. Nat Rev Mol Cell Biol. 2018:19(5):327–341. 10.1038/nrm.2017.130 [DOI] [PubMed] [Google Scholar]
- Holdsworth MJ, Bentsink L, Soppe WJJ. Molecular networks regulating Arabidopsis seed maturation, after-ripening, dormancy and germination. New Phytol. 2008:179(1):33–54. 10.1111/j.1469-8137.2008.02437.x [DOI] [PubMed] [Google Scholar]
- Huang R, Liu M, Gong G, Wu P, Patra B, Yuan L, Qin H, Wang X, Wang G, Liao H, et al. The Pumilio RNA-binding protein APUM24 regulates seed maturation by fine-tuning the BPM-WRI1 module in Arabidopsis. J Integr Plant Biol. 2021:63(7):1240–1259. 10.1111/jipb.13092 [DOI] [PubMed] [Google Scholar]
- Huang X, Schmitt J, Dorn L, Griffith C, Effgen S, Takao S, Koornneef M, Donohue K. The earliest stages of adaptation in an experimental plant population: strong selection on QTLs for seed dormancy. Mol Ecol. 2010:19(7):1335–1351. 10.1111/j.1365-294X.2010.04557.x [DOI] [PubMed] [Google Scholar]
- Huh SU, Paek KH. APUM5, encoding a Pumilio RNA binding protein, negatively regulates abiotic stress responsive gene expression. BMC Plant Biol. 2014:14(1):75. 10.1186/1471-2229-14-75 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hussain S, Kim SH, Bahk S, Ali A, Nguyen XC, Yun D-J, Chung WS. The auxin signaling repressor IAA8 promotes seed germination through down-regulation of ABI3 transcription in Arabidopsis. Front Plant Sci. 2020:11:111. 10.3389/fpls.2020.00111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishibashi N, Yamauchi D, Minamikawa T. Stored mRNA in cotyledons of Vigna unguiculata seeds: nucleotide sequence of cloned cDNA for a stored mRNA and induction of its synthesis by precocious germination. Plant Mol Biol. 1990:15(1):59–64. 10.1007/BF00017724 [DOI] [PubMed] [Google Scholar]
- Iwasaki M, Penfield S, Lopez-Molina L. Parental and environmental control of seed dormancy in Arabidopsis thaliana. Annu Rev Plant Biol. 2022:73(1):355–378. 10.1146/annurev-arplant-102820-090750 [DOI] [PubMed] [Google Scholar]
- Jang GJ, Jang JC, Wu SH. Dynamics and functions of stress granules and processing bodies in plants. Plants (Basel). 2020:9(9):1122. 10.3390/plants9091122 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jang GJ, Yang JY, Hsieh HL, Wu SH. Processing bodies control the selective translation for optimal development of Arabidopsis young seedlings. Proc Natl Acad Sci U S A. 2019:116(13):6451–6456. 10.1073/pnas.1900084116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jia H, McCarty DR, Suzuki M. Distinct roles of LAFL network genes in promoting the embryonic seedling fate in the absence of VAL repression. Plant Physiol. 2013:163(3):1293–1305. 10.1104/pp.113.220988 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang J, Yim S, Choi H, Kim A, Lee KP, Lopez-Molina L, Martinoia E, Lee Y. Abscisic acid transporters cooperate to control seed germination. Nat Commun. 2015:6(1):8113. 10.1038/ncomms9113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karssen CM, Brinkhorst-van der Swan DLC, Breekland AE, Koornneef M. Induction of dormancy during seed development by endogenous abscisic acid: studies on abscisic acid deficient genotypes of Arabidopsis thaliana (L.) Heynh. Planta. 1983:157(2):158–165. 10.1007/BF00393650 [DOI] [PubMed] [Google Scholar]
- Keith K, Kraml M, Dengler NG, McCourt P. Fusca3: a heterochronic mutation affectting late embryo development in Arabidopsis. Plant Cell. 1994:6(5):589–600. 10.2307/3869865 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim Y-O, Kim JS, Kang H. Cold-inducible zinc finger-containing glycine-rich RNA-binding protein contributes to the enhancement of freezing tolerance in Arabidopsis thaliana. Plant J. 2005:42(6):890–900. 10.1111/j.1365-313X.2005.02420.x [DOI] [PubMed] [Google Scholar]
- Kim YO, Pan S, Jung CH, Kang H. A zinc finger-containing glycine-rich RNA-binding protein, atRZ-1a, has a negative impact on seed germination and seedling growth of Arabidopsis thaliana under salt or drought stress conditions. Plant Cell Physiol. 2007:48(8):1170–1181. 10.1093/pcp/pcm087 [DOI] [PubMed] [Google Scholar]
- King HA, Gerber AP. Translatome profiling: methods for genome-scale analysis of mRNA translation. Brief Funct Genomics. 2016:15(1):22–31. 10.1093/bfgp/elu045 [DOI] [PubMed] [Google Scholar]
- Koornneef M, Jorna ML, Brinkhorst-van der Swan DLC, Karssen CM. The isolation of abscisic acid (ABA) deficient mutants by selection of induced revertants in non-germinating gibberillin sensitive lines of Arabidopsis thaliana (L.) Heynh. Theor Appl Genet. 1982:61(4):385–393. 10.1007/BF00272861 [DOI] [PubMed] [Google Scholar]
- Koornneef M, Reuling G, Karssen CM. The isolation and characterization of abscisic acid-insensitive mutants of Arabidopsis thaliana. Physiol Plant. 1984:61(3):377–383. 10.1111/j.1399-3054.1984.tb06343.x [DOI] [Google Scholar]
- Koornneef M, van der Veen JH. Induction and analysis of gibberellin sensitive mutants in Arabidopsis thaliana (L.) heynh. Theor Appl Genet. 1980:58(6):257–263. 10.1007/BF00265176 [DOI] [PubMed] [Google Scholar]
- Köster T, Marondedze C, Meyer K, Staiger D. RNA-Binding proteins revisited—the emerging Arabidopsis mRNA interactome. Trends Plant Sci. 2017:22(6):512–526. 10.1016/j.tplants.2017.03.009 [DOI] [PubMed] [Google Scholar]
- Kowalczyk J, Palusinska M, Wroblewska-Swiniarska A, Pietras Z, Szewc L, Dolata J, Jarmolowski A, Swiezewski S. Alternative polyadenylation of the sense transcript controls antisense transcription of DELAY OF GERMINATION 1 in Arabidopsis. Mol Plant. 2017:10(10):1349–1352. 10.1016/j.molp.2017.07.011 [DOI] [PubMed] [Google Scholar]
- Kozaki A, Aoyanagi T. Molecular aspects of seed development controlled by gibberellins and abscisic acids. Int J Mol Sci. 2022:23(3):1876. 10.3390/ijms23031876 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kronholm I, Picó FX, Alonso-Blanco C, Goudet J, Meaux JD. Genetic basis of adaptation in Arabidopsis thaliana: local adaptation at the seed dormancy QTL DOG1. Evolution. 2012:66:2287–2302. [DOI] [PubMed] [Google Scholar]
- Krzyszton M, Yatusevich R, Wrona M, Sacharowski S, Adamska D, Swiezewski S. Single seeds exhibit transcriptional heterogeneity during secondary dormancy induction. Plant Physiol. 2022:190(1):211–225. 10.1093/plphys/kiac265 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuligowski J, Ferrand M, Chenou E. Stored mRNA in early embryos of a fern Marsilea vestita: a paternal and maternal origin. Mol Reprod Dev. 1991:30(1):27–33. 10.1002/mrd.1080300104 [DOI] [PubMed] [Google Scholar]
- Kushiro T, Okamoto M, Nakabayashi K, Yamagishi K, Kitamura S, Asami T, Hirai N, Koshiba T, Kamiya Y, Nambara E. The Arabidopsis cytochrome P450 CYP707A encodes ABA 8'-hydroxylases: key enzymes in ABA catabolism. EMBO J. 2004:23(7):1647–1656. 10.1038/sj.emboj.7600121 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Layat E, Leymarie J, El-Maarouf-Bouteau H, Caius J, Langlade N, Bailly C. Translatome profiling in dormant and nondormant sunflower (Helianthus annuus) seeds highlights post-transcriptional regulation of germination. New Phytol. 2014:204(4):864–872. 10.1111/nph.13002 [DOI] [PubMed] [Google Scholar]
- Lee KP, Piskurewicz U, Tureckova V, Strnad M, Lopez-Molina L. A seed coat bedding assay shows that RGL2-dependent release of abscisic acid by the endosperm controls embryo growth in Arabidopsis dormant seeds. Proc Natl Acad Sci U S A. 2010:107(44):19108–19113. 10.1073/pnas.1012896107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee S, Cheng H, King KE, Wang W, He Y, Hussain A, Lo J, Harberd NP, Peng J. Gibberellin regulates Arabidopsis seed germination via RGL2, a GAI/RGA-like gene whose expression is up-regulated following imbibition. Genes Dev. 2002:16(5):646–658. 10.1101/gad.969002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee TA, Nobori T, Illouz-Eliaz N, Xu J, Jow B, Nery JR, Ecker JR. A Single-nucleus atlas of seed-to-seed development in Arabidopsis. bioRxiv. 2023:2023.2003.2023.533992. 10.1101/2023.03.23.533992 [DOI] [Google Scholar]
- Lefebvre V, North H, Frey A, Sotta B, Seo M, Okamoto M, Nambara E, Marion-Poll A. Functional analysis of Arabidopsis NCED6 and NCED9 genes indicates that ABA synthesized in the endosperm is involved in the induction of seed dormancy. Plant J. 2006:45(3):309–319. 10.1111/j.1365-313X.2005.02622.x [DOI] [PubMed] [Google Scholar]
- Léon-Kloosterziel KM, Alvarez Gil M, Ruijs GJ, Jacobsen SE, Olszewski NE, Schwartz SH, Zeevaart JAD, Koornneef M. Isolation and characterization of abscisic acid-deficient Arabidopsis mutants at two new loci. Plant J. 1996:10(4):655–661. 10.1046/j.1365-313X.1996.10040655.x [DOI] [PubMed] [Google Scholar]
- Leymarie J, Bruneaux E, Gibot-Leclerc S, Corbineau F. Identification of transcripts potentially involved in barley seed germination and dormancy using cDNA-AFLP. J Exp Bot. 2007:58(3):425–437. 10.1093/jxb/erl211 [DOI] [PubMed] [Google Scholar]
- Li X, Chen T, Li Y, Wang Z, Cao H, Chen F, Li Y, Soppe WJJ, Li W, Liu Y. ETR1/RDO3 regulates seed dormancy by relieving the inhibitory effect of the ERF12-TPL complex on DELAY OF GERMINATION1 expression. Plant Cell. 2019:31(4):832–847. 10.1105/tpc.18.00449 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X, Kazan H, Lipshitz HD, Morris QD. Finding the target sites of RNA-binding proteins. WIREs RNA. 2014:5(1):111–130. 10.1002/wrna.1201 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li P, Ni H, Songbei Y, Wei J, Hu X. Teaching an old dog a new trick: multifaceted strategies to control primary seed germination by DELAY OF GERMINATION 1 (DOG1). Phyton (B Aires). 2020:89(1):1–12. 10.32604/phyton.2020.09817 [DOI] [Google Scholar]
- Lim S, Park J, Lee N, Jeong J, Toh S, Watanabe A, Kim J, Kang H, Kim DH, Kawakami N, et al. ABA-insensitive3, ABA-insensitive5, and DELLAs interact to activate the expression of SOMNUS and other high-temperature-inducible genes in imbibed seeds in Arabidopsis. Plant Cell. 2013:25(12):4863–4878. 10.1105/tpc.113.118604 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu X, Zhang H, Zhao Y, Feng Z, Li Q, Yang H-Q, Luan S, Li J, He Z-H. Auxin controls seed dormancy through stimulation of abscisic acid signaling by inducing ARF-mediated ABI3 activation in Arabidopsis. Proc Natl Acad Sci U S A. 2013:110(38):15485–15490. 10.1073/pnas.1304651110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Koornneef M, Soppe WJ. The absence of histone H2B monoubiquitination in the Arabidopsis hub1 (rdo4) mutant reveals a role for chromatin remodeling in seed dormancy. Plant Cell. 2007:19(2):433–444. 10.1105/tpc.106.049221 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Longo C, Holness S, De Angelis V, Lepri A, Occhigrossi S, Ruta V, Vittorioso P. From the outside to the inside: new insights on the main factors that guide seed dormancy and germination. Genes (Basel). 2020:12(1):52. 10.3390/genes12010052 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorković ZJ. Role of plant RNA-binding proteins in development, stress response and genome organization. Trends Plant Sci. 2009:14(4):229–236. 10.1016/j.tplants.2009.01.007 [DOI] [PubMed] [Google Scholar]
- Lou L, Ding L, Wang T, Xiang Y. Emerging roles of RNA-binding proteins in seed development and performance. Int J Mol Sci. 2020:21(18):6822. 10.3390/ijms21186822 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lynch TJ, Erickson BJ, Miller DR, Finkelstein RR. ABI5-binding proteins (AFPs) alter transcription of ABA-induced genes via a variety of interactions with chromatin modifiers. Plant Mol Biol. 2017:93(4–5):403–418. 10.1007/s11103-016-0569-1 [DOI] [PubMed] [Google Scholar]
- Maia J, Dekkers BJ, Provart NJ, Ligterink W, Hilhorst HW. The re-establishment of desiccation tolerance in germinated Arabidopsis thaliana seeds and its associated transcriptome. PLoS One. 2011:6(12):e29123. 10.1371/journal.pone.0029123 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maldonado-Bonilla LD. Composition and function of P bodies in Arabidopsis thaliana. Front Plant Sci. 2014:5:201. 10.3389/fpls.2014.00201 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martínez-Berdeja A, Stitzer MC, Taylor MA, Okada M, Ezcurra E, Runcie DE, Schmitt J. Functional variants of DOG1 control seed chilling responses and variation in seasonal life-history strategies in Arabidopsis thaliana. Proc Natl Acad Sci U S A. 2020:117(5):2526–2534. 10.1073/pnas.1912451117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meng PH, Macquet A, Loudet O, Marion-Poll A, North H. Analysis of natural allelic variation controlling Arabidopsis thaliana seed germinability in response to cold and dark: indentification of three major quantitative trait loci. Mol Plant. 2008:1(1):145–154. 10.1093/mp/ssm014 [DOI] [PubMed] [Google Scholar]
- Mitchell J, Johnston IG, Bassel GW. Variability in seeds: biological, ecological, and agricultural implications. J Exp Bot. 2017:68(4):809–817. 10.1093/jxb/erw397 [DOI] [PubMed] [Google Scholar]
- Montez M, Majchrowska M, Krzyszton M, Bokota G, Sacharowski S, Wrona M, Yatusevich R, Massana F, Plewczynski D, Swiezewski S. Promoter-pervasive transcription causes RNA polymerase II pausing to boost DOG1 expression in response to salt. EMBO J. 2023:42(5):e112443. 10.15252/embj.2022112443 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muller K, Tintelnot S, Leubner-Metzger G. Endosperm-limited Brassicaceae seed germination: abscisic acid inhibits embryo-induced endosperm weakening of Lepidium sativum (cress) and endosperm rupture of cress and Arabidopsis thaliana. Plant Cell Physiol. 2006:47(7):864–877. 10.1093/pcp/pcj059 [DOI] [PubMed] [Google Scholar]
- Nakabayashi K, Bartsch M, Xiang Y, Miatton E, Pellengahr S, Yano R, Seo M, Soppe WJJ. The time required for dormancy release in Arabidopsis is determined by DELAY OF GERMINATION1 protein levels in freshly harvested seeds. Plant Cell. 2012:24(7):2826–2838. 10.1105/tpc.112.100214 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakabayashi K, Okamoto M, Koshiba T, Kamiya Y, Nambara E. Genome-wide profiling of stored mRNA in Arabidopsis thaliana seed germination: epigenetic and genetic regulation of transcription in seed. Plant J. 2005:41(5):697–709. 10.1111/j.1365-313X.2005.02337.x [DOI] [PubMed] [Google Scholar]
- Nambara E, Keith K, McCourt P, Naito S. Isolation of an internal deletion mutant of the Arabidopsis thaliana ABI3 gene. Plant Cell Physiol. 1994:35:509–513. 10.1093/oxfordjournals.pcp.a078623 [DOI] [PubMed] [Google Scholar]
- Narsai R, Gouil Q, Secco D, Srivastava A, Karpievitch YV, Liew LC, Lister R, Lewsey MG, Whelan J. Extensive transcriptomic and epigenomic remodelling occurs during Arabidopsis thaliana germination. Genome Biol. 2017:18(1):172. 10.1186/s13059-017-1302-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nee G, Xiang Y, Soppe WJ. The release of dormancy, a wake-up call for seeds to germinate. Curr Opin Plant Biol. 2017a:35:8–14. 10.1016/j.pbi.2016.09.002 [DOI] [PubMed] [Google Scholar]
- Nee G, Kramer K, Nakabayashi K, Yuan B, Xiang Y, Miatton E, Finkemeier I, Soppe WJJ. DELAY OF GERMINATION1 requires PP2C phosphatases of the ABA signalling pathway to control seed dormancy. Nat Commun. 2017b:8(1):72. 10.1038/s41467-017-00113-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Née G, Krüger T. Dry side of the core: a meta-analysis addressing the original nature of the ABA signalosome at the onset of seed imbibition. Front Plant Sci. 2023:14:1192652. 10.3389/fpls.2023.1192652 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikolaeva MG. Physiology of deep dormancy in seeds. Leningrad, USSR: Nauka; 1969. [Google Scholar]
- Nishimura N, Tsuchiya W, Moresco JJ, Hayashi Y, Satoh K, Kaiwa N, Irisa T, Kinoshita T, Schroeder JI, Yates JR, 3rd et al. Control of seed dormancy and germination by DOG1-AHG1 PP2C phosphatase complex via binding to heme. Nat Commun. 2018:9(1):2132. 10.1038/s41467-018-04437-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nonogaki H. Seed germination and dormancy: the classic story, new puzzles, and evolution. J Integr Plant Biol. 2019:61(5):541–563. 10.1111/jipb.12762 [DOI] [PubMed] [Google Scholar]
- Nonogaki H, Bassel GW, Bewley JD. Germination-still a mystery. Plant Sci. 2010:179(6):574–581. 10.1016/j.plantsci.2010.02.010 [DOI] [Google Scholar]
- Nyikó T, Auber A, Bucher E. Functional and molecular characterization of the conserved Arabidopsis PUMILIO protein, APUM9. Plant Mol Biol. 2019:100(1–2):199–214. 10.1007/s11103-019-00853-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogas J, Kaufmann S, Henderson J, Somerville C. PICKLE is a CHD3 chromatin-remodeling factor that regulates the transition from embryonic to vegetative development in Arabidopsis. Proc Natl Acad Sci U S A. 1999:96(24):13839–13844. 10.1073/pnas.96.24.13839 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ooms JJJ, Léon-Kloosterziel KM, Bartels D, Koornneef M, Karssen CM. Acquisition of desiccation tolerance and longevity in seeds of Arabidopsis thaliana—a comparative study using abscisic acid- insensitive abi3 mutants. Plant Physiol. 1993:102(4):1185–1191. 10.1104/pp.102.4.1185 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park J, Lee N, Kim W, Lim S, Choi G. ABI3 and PIL5 collaboratively activate the expression of SOMNUS by directly binding to its promoter in imbibed Arabidopsis seeds. Plant Cell. 2011:23(4):1404–1415. 10.1105/tpc.110.080721 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perruc E, Kinoshita N, Lopez-Molina L. The role of chromatin-remodeling factor PKL in balancing osmotic stress responses during Arabidopsis seed germination. Plant J. 2007:52(5):927–936. 10.1111/j.1365-313X.2007.03288.x [DOI] [PubMed] [Google Scholar]
- Rajjou L, Gallardo K, Debeaujon I, Vandekerckhove J, Job C, Job D. The effect of alpha-amanitin on the Arabidopsis seed proteome highlights the distinct roles of stored and neosynthesized mRNAs during germination. Plant Physiol. 2004:134(4):1598–1613. 10.1104/pp.103.036293 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raz V, Bergervoet JH, Koornneef M. Sequential steps for developmental arrest in Arabidopsis seeds. Development. 2001:128(2):243–252. 10.1242/dev.128.2.243 [DOI] [PubMed] [Google Scholar]
- Rodriques SG, Stickels RR, Goeva A, Martin CA, Murray E, Vanderburg CR, Welch J, Chen LM, Chen F, Macosko EZ. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science. 2019:363(6434):1463–1467. 10.1126/science.aaw1219 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ross JJ, Quittenden LJ. Interactions between brassinosteroids and gibberellins: synthesis or signaling? Plant Cell. 2016:28(4):829–832. 10.1105/tpc.15.00917 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sablok G, Powell JJ, Kazan K. Emerging roles and landscape of translating mRNAs in plants. Front Plant Sci. 2017:8:1443. 10.3389/fpls.2017.01443 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sajeev N, Baral A, America AHP, Willems LAJ, Merret R, Bentsink L. The mRNA-binding proteome of a critical phase transition during Arabidopsis seed germination. New Phytol. 2022:233(1):251–264. 10.1111/nph.17800 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sall K, Dekkers BJW, Nonogaki M, Katsuragawa Y, Koyari R, Hendrix D, Willems LAJ, Bentsink L, Nonogaki H. DELAY OF GERMINATION1-LIKE4 acts as an inducer of seed reserve accumulation. Plant J. 2019:100(1):7–19. 10.1111/tpj.14485 [DOI] [PubMed] [Google Scholar]
- Sano N, Ono H, Murata K, Yamada T, Hirasawa T, Kanekatsu M. Accumulation of long-lived mRNAs associated with germination in embryos during seed development of rice. J Exp Bot. 2015:66(13):4035–4046. 10.1093/jxb/erv209 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sano N, Permana H, Kumada R, Shinozaki Y, Tanabata T, Yamada T, Hirasawa T, Kanekatsu M. Proteomic analysis of embryonic proteins synthesized from long-lived mRNAs during germination of rice seeds. Plant Cell Physiol. 2012:53(4):687–698. 10.1093/pcp/pcs024 [DOI] [PubMed] [Google Scholar]
- Sato H, Köhler C. Genomic imprinting regulates establishment and release of seed dormancy. Curr Opin Plant Biol. 2022:69:102264. 10.1016/j.pbi.2022.102264 [DOI] [PubMed] [Google Scholar]
- Sato H, Santos-González J, Köhler C. Combinations of maternal-specific repressive epigenetic marks in the endosperm control seed dormancy. eLife. 2021:10:e64593. 10.7554/eLife.64593 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shu K, Liu XD, Xie Q, He ZH. Two faces of one seed: hormonal regulation of dormancy and germination. Mol Plant. 2016:9(1):34–45. 10.1016/j.molp.2015.08.010 [DOI] [PubMed] [Google Scholar]
- Shu K, Zhang H, Wang S, Chen M, Wu Y, Tang S, Liu C, Feng Y, Cao X, Xie Q. ABI4 regulates primary seed dormancy by regulating the biogenesis of abscisic acid and gibberellins in arabidopsis. PLoS Genet. 2013:9(6):e1003577. 10.1371/journal.pgen.1003577 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skubacz A, Daszkowska-Golec A, Szarejko I. The role and regulation of ABI5 (ABA-insensitive 5) in plant development, abiotic stress responses and phytohormone crosstalk. Front Plant Sci. 2016:7:1884. 10.3389/fpls.2016.01884 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song S, He H, Gühl K, van Bolderen-Veldkamp M, Buijs G, Willems L, Bentsink L. DELAY OF GERMINATION 6 encoding the ANAC060 transcription factor, inhibits seed dormancy. bioRxiv. 2021:2021. 05.03.442418. 10.1101/2021.05.03.442418 [DOI]
- Song S, Willems LAJ, Jiao A, Zhao T, Eric Schranz M, Bentsink L. The membrane associated NAC transcription factors ANAC060 and ANAC040 are functionally redundant in the inhibition of seed dormancy in Arabidopsis thaliana. J Exp Bot. 2022:73(16):5514–5528. 10.1093/jxb/erac232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soppe W, Bentsink L, Koornneef M. The early-flowering mutant efs is involved in the autonomous promotion pathway of Arabidopsis thaliana. Development. 1999:126(21):4763–4770. 10.1242/dev.126.21.4763 [DOI] [PubMed] [Google Scholar]
- Soppe WJJ, Bentsink L. Seed dormancy back on track; its definition and regulation by DOG1. New Phytol. 2020:228(3):816–819. 10.1111/nph.16592 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steber CM, McCourt P. A role for brassinosteriods in germination in Arabidopsis. Plant Physiol. 2001:125(2):763–769. 10.1104/pp.125.2.763 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stickels RR, Murray E, Kumar P, Li J, Marshall JL, Di Bella DJ, Arlotta P, Macosko EZ, Chen F. Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nature Biotech. 2021:39(3):313–319. 10.1038/s41587-020-0739-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka M, Kikuchi A, Kamada H. The Arabidopsis histone deacetylases HDA6 and HDA19 contribute to the repression of embryonic properties after germination. Plant Physiol. 2008:146(1):149–161. 10.1104/pp.107.111674 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian R, Wang F, Zheng Q, Niza V, Downie AB, Perry SE. Direct and indirect targets of the Arabidopsis seed transcription factor ABSCISIC ACID INSENSITIVE3. Plant J. 2020:103(5):1679–1694. 10.1111/tpj.14854 [DOI] [PubMed] [Google Scholar]
- Urbanova T, Leubner-Metzger G. Gibberellins and seed germination. Annual plant reviews online; 2018. p. 253–284. https://onlinelibrary.wiley.com/doi/abs/10.1002/9781119312994.apr0538 [Google Scholar]
- Van der Schaar W, Alonso-Blanco C, Léon-Kloosterziel KM, Jansen RC, van Ooijen JW, Koornneef M. QTL analysis of seed dormancy in Arabidopsis using recombinant inbred lines and MQM mapping. Heredity (Edinburgh). 1997:79(2):190–200. 10.1038/hdy.1997.142 [DOI] [PubMed] [Google Scholar]
- van Zanten M, Carles A, Li Y, Soppe W. Control and consequences of chromatin compaction during seed maturation in Arabidopsis thaliana. Plant Signal Behav. 2012:7(3):338–341. 10.4161/psb.19281 [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Zanten M, Zoll C, Wang Z, Philipp C, Carles A, Li Y, Kornet NG, Liu Y, Soppe WJ. HISTONE DEACETYLASE 9 represses seedling traits in Arabidopsis thaliana dry seeds. Plant J. 2014:80(3):475–488. 10.1111/tpj.12646 [DOI] [PubMed] [Google Scholar]
- Vidigal DS, Marques AC, Willems LA, Buijs G, Mendez-Vigo B, Hilhorst HW, Bentsink L, Pico FX, Alonso-Blanco C. Altitudinal and climatic associations of seed dormancy and flowering traits evidence adaptation of annual life cycle timing in Arabidopsis thaliana. Plant Cell Environ. 2016:39(8):1737–1748. 10.1111/pce.12734 [DOI] [PubMed] [Google Scholar]
- Voegele A, Graeber K, Oracz K, Tarkowská D, Jacquemoud D, Turečková V, Urbanová T, Strnad M, Leubner-Metzger G. Embryo growth, testa permeability, and endosperm weakening are major targets for the environmentally regulated inhibition of Lepidium sativum seed germination by myrigalone A. J Exp Bot. 2012:63(14):5337–5350. 10.1093/jxb/ers197 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z, Chen F, Li X, Cao H, Ding M, Zhang C, Zuo J, Xu C, Xu J, Deng X, et al. Arabidopsis seed germination speed is controlled by SNL histone deacetylase-binding factor-mediated regulation of AUX1. Nat Commun. 2016:7(1):13412. 10.1038/ncomms13412 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiang Y, Nakabayashi K, Ding J, He F, Bentsink L, Soppe WJ. Reduced Dormancy5 encodes a protein phosphatase 2C that is required for seed dormancy in Arabidopsis. Plant Cell. 2014:26(11):4362–4375. 10.1105/tpc.114.132811 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiang Y, Song BX, Nee G, Kramer K, Finkemeier I, Soppe WJJ. Sequence polymorphisms at the REDUCED DORMANCY5 pseudophosphatase underlie natural variation in Arabidopsis dormancy. Plant Physiol. 2016:171(4):2659–2670. 10.1104/pp.16.00525 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu H, Lantzouni O, Bruggink T, Benjamins R, Lanfermeijer F, Denby K, Schwechheimer C, Bassel GW. A molecular signal integration network underpinning Arabidopsis seed germination. Curr Biol. 2020:30(19):3703–3712.e3704. 10.1016/j.cub.2020.07.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yazdanpanah F, Hanson J, Hilhorst HWM, Bentsink L. Differentially expressed genes during the imbibition of dormant and after-ripened seeds—a reverse genetics approach. BMC Plant Biol. 2017:17(1):151. 10.1186/s12870-017-1098-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoong FY, O'Brien LK, Truco MJ, Huo H, Sideman R, Hayes R, Michelmore RW, Bradford KJ. Genetic variation for thermotolerance in lettuce seed germination is associated with temperature-sensitive regulation of ETHYLENE RESPONSE FACTOR1 (ERF1). Plant Physiol. 2016:170(1):472–488. 10.1104/pp.15.01251 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang H, Bishop B, Ringenberg W, Muir WM, Ogas J. The CHD3 remodeler PICKLE associates with genes enriched for trimethylation of histone H3 lysine 27. Plant Physiol. 2012:159(1):418–432. 10.1104/pp.112.194878 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao X, Dou L, Gong Z, Wang X, Mao T. BES1 hinders ABSCISIC ACID INSENSITIVE5 and promotes seed germination in Arabidopsis. New Phytol. 2019:221(2):908–918. 10.1111/nph.15437 [DOI] [PubMed] [Google Scholar]
- Zheng J, Chen F, Wang Z, Cao H, Li X, Deng X, Soppe WJ, Li Y, Liu Y. A novel role for histone methyltransferase KYP/SUVH4 in the control of Arabidopsis primary seed dormancy. New Phytol. 2012:193(3):605–616. 10.1111/j.1469-8137.2011.03969.x [DOI] [PubMed] [Google Scholar]
- Zhong C, Patra B, Tang Y, Li X, Yuan L, Wang X. A transcriptional hub integrating gibberellin–brassinosteroid signals to promote seed germination in Arabidopsis. J Exp Bot. 2021:72(13):4708–4720. 10.1093/jxb/erab192 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu H, Xie W, Xu D, Miki D, Tang K, Huang CF, Zhu JK. DNA demethylase ROS1 negatively regulates the imprinting of DOGL4 and seed dormancy in Arabidopsis thaliana. Proc Natl Acad Sci U S A. 2018:115(42):E9962–E9970. 10.1073/pnas.1812847115 [DOI] [PMC free article] [PubMed] [Google Scholar]