Abstract
Background
Rising clarithromycin resistance undermines Helicobacter pylori (H. pylori) treatment efficacy. We aimed to determine clarithromycin's minimum inhibitory concentration (MIC) levels and identify specific mutation sites in the 23S ribosomal subunit (23S rRNA) that predict treatment outcomes in a 14-day regimen of clarithromycin bismuth quadruple therapy (amoxicillin 1g, clarithromycin 500 mg, rabeprazole 10 mg, and colloidal bismuth pectin 200 mg).
Materials and methods
We included adult H. pylori patients who hadn't previously undergone clarithromycin-based treatment, either as initial or rescue therapy. Exclusions were made for penicillin allergy, recent use of related medications, severe illnesses, or inability to cooperate. Patients underwent a 14-day clarithromycin bismuth quadruple therapy. Gastric mucosa specimens were obtained during endoscopy before eradication. MIC against amoxicillin and clarithromycin was determined using the E-test method. The receiver operating characteristic (ROC) curve helped to find the optimal clarithromycin resistance MIC breakpoint. Genetic sequences of H. pylori 23S rRNA were identified through Sanger Sequencing. (ChiCTR2200061476)
Results
Out of 196 patients recruited, 92 met the inclusion criteria for the per-protocol (PP) population. The overall intention-to-treat (ITT) eradication rate was 80.00 % (84/105), while the modified intention-to-treat (MITT) and PP eradication rates were 90.32 % (84/93) and 91.30 % (84/92) respectively. No amoxicillin resistance was observed, but clarithromycin resistance rates were 36.19 % (38/105), 35.48 % (33/93), and 34.78 % (33/92) in the ITT, MITT, and PP populations respectively. Compared with the traditional clarithromycin resistance breakpoint of 0.25 μg/mL, a MIC threshold of 12 μg/mL predicted better eradication. Among 173 mutations on 152 sites in the 23S rRNA gene, only the 2143A > G mutation could predict eradication outcomes (p < 0.000).
Conclusions
Interpretation of elevated MIC values is crucial in susceptibility testing, rather than a binary "susceptible" or "resistant" classification. The 2143A > G mutation has limited specificity in predicting eradication outcomes, necessitating further investigation into additional mutation sites associated with clarithromycin resistance.
Keywords: Helicobacter pylori, Clarithromycin, Antibiotic resistance, Minimum inhibitory concentration (MIC), Receiver operating characteristic (ROC), 23S rRNA mutation sites
1. Introduction
Helicobacter. pylori (H. pylori) is a microaerophilic gram-negative bacterium known to adversely impact both the structural integrity and physiological functions of the gastric mucosa [1]. Notably, in China, the prevalence of H. pylori infection reaches a strikingly high rate, estimated at 50–60 % [2].
Clarithromycin is considered to be the cornerstone of H. pylori eradication therapy [3]. In regions characterized by a high (>15 %) clarithromycin resistance prevalence or with unknown resistance rates, bismuth quadruple therapy is recommended as the first-line treatment option [4]. Antimicrobial resistance has been increasing recently, especially in clarithromycin [5], and studies have proved that clarithromycin resistance significantly impacts the success of eradication, even when employing bismuth quadruple therapy [6]. The overall clarithromycin resistance rate in China was 47.72 % [7], which was consistent with global data (43.1 %) [8]. Clinicians often consider giving up the use of clarithromycin once clarithromycin resistance occurs. We know that MIC>0.25 μg/mL has been defined as a clarithromycin resistance breakpoint according to the European Committee on Antimicrobial Susceptibility Testing (EUCAST). However, we found that few studies have investigated the MIC levels of clarithromycin in 14-day clarithromycin bismuth quadruple therapy. It has not been proven whether the traditional clarithromycin resistance MIC value is suitable for the 14-day clarithromycin bismuth quadruple therapy.
In 1996, Versalovic et al. found that resistance to clarithromycin is mediated by the point mutations in the peptidyl transferase region encoded in domain V of 23S rRNA [9], especially at the sites of 2142 (11.7 %) and 2143 [10]. Resistance to clarithromycin is all or none [11]. The detection of clarithromycin gene sites is widely used to evaluate whether clarithromycin is suitable for eradication therapy in the clinic. However, there are limited clinical studies about the effect of mutation sites on the eradication outcome of 14-day clarithromycin bismuth quadruple therapy.
We intended to investigate the clinical MIC breakpoint of clarithromycin in predicting the eradication outcome and discuss the effect of mutation sites on the eradication outcome in 14-day clarithromycin bismuth quadruple therapy.
2. Materials and methods
2.1. Patients recruitment
This study was a prospective, single-center trial conducted in the Gastroenterology Department of Beijing Hospital. It took place from June 2022 to November 2022. Patients infected with H. pylori who met the inclusion and exclusion criteria were consecutively enrolled.
The inclusion criteria included i) individuals ages 18–65 years old, ii) Current H. pylori infection was confirmed by 13C-urea Breath Test, stool H. pylori antigen test, rapid urease test, or histological staining before enrollment (including treatment-naive patients and those with a history of eradication). Additionally, the criteria required that treatment-rescue patients had not been previously exposed to a clarithromycin-containing regimen for H. pylori eradication, iii) Potential participants were willing to undergo endoscopy to facilitate H. pylori eradication and expressed their commitment by completing an informed consent form.
By contrast, patients with (a) a history of a penicillin allergy reaction,(b) a history of the therapy containing clarithromycin to eradicate H. pylori, (c) a history of alcohol abuse or drug addiction, (d) a history of upper gastrointestinal surgery, (e)pregnant or lactating women, (f) severe concurrent disease or malignancy, (g) inability to participate due to psychosis and severe neurocognitive impairment and (h) patients who were recent or ongoing PPI within 2 weeks, bismuth, hormones, or antibiotics within 4 weeks were excluded.
We used a structured questionnaire to collect information on the patient's demographic and clinical data, including age, sex, history of penicillin allergy, previous exposure to non-eradication related macrolides, previous eradication history and the specific eradication regiments as recorded by clinicians. Informed consent was obtained from all subjects at the time of enrollment. This study has been approved by the Ethic Committee of Beijing Hospital Medical Research Institutions (2021BJYYEC-274-01) and registered in the Chinese Clinical Trail Registry. Registration (ChiCTR2200061476).
2.2. The eradication regimen of H. pylori infection
A quadruple therapy regimen was prescribed for the patients, consisting of amoxicillin 1g, clarithromycin 500 mg, rabeprazole 10 mg, and colloidal bismuth pectin 200 mg. All medications were administered twice daily, beginning on the second day after the endoscopy. The duration of quadruple therapy was 14 days for all subjects. Simultaneously, patients were provided with detailed information on medication administration and potential adverse reactions before starting the treatment. 13C-urea Breath Test was performed 6 weeks after the completion of the prescribed treatment. Infections were considered eradicated when the 13C-urea Breath Test results were negative, as assessed by the investigators. Eradication rates, determined by the results of the 13C-urea Breath Test, were the primary outcomes to be analyzed.x3 Study compliance and adverse events were recorded as secondary endpoints.
2.3. Specimen collection and H. pylori culture
Two mucosal biopsy specimens were obtained from the gastric antrum and corpus using sterile biopsy forceps during endoscopy. The tissues were kept in sterile plastic tubes and transported to the laboratory immediately, frozen in a refrigerator at 4 °C for no more than 6 h with brain-heart infusion broth (CM1135, OXOID) containing 10 % glycerin as transport media. Endoscopic diagnosis and pathological results were recorded after endoscopy. The specimens were ground and directly streaked onto the surface of selective Columbia blood agar plates (CM0331B, OXOID), containing 15 % defibrinated sheep blood (FJ11933, FEIMOBIO) and H. pylori selective supplement (SR0147E, OXOID). The plates were incubated at 37 °C under microaerophilic conditions (5 % O2, 10 % CO2, 85 % N2, and 10%H2) in the incubator for no more than 10 days. The initial isolate colonies were checked every other day and identified as H. pylori based on colony morphology. The culture was considered positive if either the antrum or corpus grew bacteria. If no significant visible changes occurred after 10 days, the culture was considered H. pylori-negative and discarded. The bacteria were harvested when growth was vigorous for antimicrobial susceptibility tests or cryopreservation at −80 °C for long-term storage.
2.4. Amoxicillin and clarithromycin susceptibility test
An antimicrobial sensitivity test was performed on freshly prepared agar plates after subculture with vigorous bacterial growth. The phenotypic resistance of each H. pylori strain was determined by measuring the MIC against amoxicillin and clarithromycin using the E-test method. An E-test strip (Liofilchem, Italy) containing amoxicillin or clarithromycin with concentrations ranging from <0.016 to >256 μg/mL was applied to the agar surface after drying the smear plate. Following incubation, a symmetrical inhibition ellipse intersects the strip, and the MIC value is read directly from the scale in μg/mL at the point where the edge of the inhibition ellipse intersects the strip. The MIC value is read directly from the scale in terms of μg/mL at the point where the edge of the inhibition ellipse intersects the strip. A suspension of H. pylori in brain-heart infusion broth was adjusted to 3 McFarland. Each strain underwent independent repetitions of the antimicrobial sensitivity test until a consistent MIC value was obtained on more than two occasions. H. pylori 26695 (NR_076155.1) was used as the quality control strain once a month. According to EUCAST recommendations, H. pylori was considered resistant if the MIC values for amoxicillin and clarithromycin were above 0.125 μg/mL and 0.25 μg/mL, respectively.
2.5. Detection of mutations in the 23S rRNA genes
At least 200 μl of H. pylori genetic DNA samples were extracted from the cultured material for the full-length 23S rRNA gene sequencing assay using a DNA Extraction Kit (hooseen), following the manufacturer's instructions. For DNA amplification, the primer sequences were as follows: forward primer, 5′-GCGTAGATGTCCGAATGGGG-3’; reverse primer, 5′GCATTACTGCGCTCACACATC-3’. Following preliminary identification by electrophoresis, the Sanger technology platform was utilized for full-length 23S rRNA sequencing to detect SNP mutations. This technique was conducted by Newglife (Beijing, China), and we verified the full-length sequences using high-quality peak maps.
2.6. Statistical analysis
All subjects who met the inclusion and exclusion criteria and had positive culture results were considered as the intention-to-treat (ITT) population. At follow-up, patients who adhered to the prescribed medication regimen and had eradication outcome results were categorized as the modified intention-to-treat (MITT) population. Patients with poor medication compliance were excluded from the per-protocol (PP) population. Drug compliance was considered satisfactory if patients took more than 85 % of the prescribed medication.
All statistical analyses were carried out using SPSS Statistics 26.0 (IBM Corporation, Armonk, NY) and compared using the chi-square test or Fisher's exact test when appropriate. Statistical significance was defined as a two-sided P-value <0.05. We delineated the MIC distribution using Prism 9 (Version 9.4.1) Raw sequencing reads were mapped to the reference genetic sequence of H. pylori 26695. Peak maps were verified using SeqMan Version 11.1.0.59. All mutation sites and the ROC curve were shown using R version 4.3.0.
3. Results
3.1. Enrolled patients and baseline characteristics
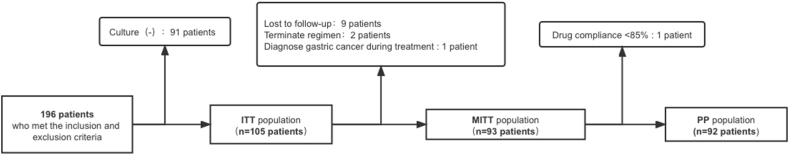
The recruitment outcomes are illustrated in Fig. 1. A total of 196 patients were enrolled initially. After the exclusion of 103 patients, both phenotype and genotype analysis were performed on the remaining eligible subjects as the MITT population (n = 93). One eradication failure patient with drug compliance of less than 85 % was excluded from the PP population. The remaining PP population including 92 patients were analyzed to explore the optimal MIC breakpoint and resistance mutations for predicting eradication outcome in 14-day clarithromycin bismuth quadruple therapy and the baseline characteristics of PP analysis patients were as shown in Table 1. The distribution of age, gender, previous exposure to non-eradication-related macrolides, endoscopic findings and previous eradication history among the two groups exhibited no statistically significant differences.
Fig. 1.
Flow chart of the patient recruitment process.
Table 1.
The baseline characteristics of enrolled patients.
| Characteristics | Total (n = 92) | Eradication Success (n = 84) | Eradication Failure (n = 8) | p-Value |
|---|---|---|---|---|
| Age±SDa(year) | 47.96 ± 12.183 | 48.00 ± 11.859 | 47.50 ± 16.186 | 0.912 |
| Sex(n) | 0.288 | |||
| Male | 40 | 35 | 5 | |
| Female | 52 | 49 | 3 | |
| previous non-eradication related macrolides exposure(n) | 0.187 | |||
| Yes | 39 | 33 | 6 | |
| No | 38 | 36 | 2 | |
| Unknownb | 15 | 15 | 0 | |
| Endoscopic findings(n) | 0.847 | |||
| Chronic non-atrophic gastritis | 67 | 60 | 7 | |
| Chronic atrophic gastritis | 13 | 12 | 1 | |
| Peptic ulcer | 11 | 11 | 0 | |
| Gastric cancer | 1 | 1 | 0 | |
| Integrity of eradication history(n) | 1.000 | |||
| Completec | 89 | 81 | 8 | |
| Incompleted | 3 | 3 | 0 | |
| Previous eradication history(n) | 1.000 | |||
| Yes | 6 | 6 | 0 | |
| No | 86 | 78 | 8 | |
| Eradication regimen(n)e | ||||
| Amoxicillin + Metronidazole | 2 | 2 | 0 | |
| Amoxicillin + Levofloxacin | 1 | 1 | 0 | |
| Unknown | 3 | 3 | 0 | |
Standard deviation.
Patients whose previous non-eradication related macrolides exposure was unknown.
All the previous eradication regimens were exactly provided in the eradication history (including naive patients with no eradication history).
The eradication history had missing information of antibiotics use or only the number of previous eradication therapies was provided.
The antibiotic combinations in previous eradication regimens.
There was a total of 86 treatment-naive patients and 6 treatment-rescue patients in PP population. In detail, 3 patients could only provide the number of previous eradication courses without specific information on antibiotic use in the process of questionnaire survey. 6 patients with eradication history have clarithromycin MIC values of 0.016 μg/mL, 0.047 μg/mL, 6 μg/mL, 8 μg/mL, 8 μg/mL, and 12 μg/mL, respectively. Among them, 3 patients with an unspecified eradication regimen have MIC values of 6 μg/mL, 8 μg/mL, and 12 μg/mL. The impact of different number of eradication courses on the eradication rate showed no statistical significance (Table 1).
3.2. The traditional resistance definition is incapable of predicting eradication outcomes
Each subgroup was evaluated for antibiotic resistance, 23S rRNA traditional mutation sites, and eradication rate as shown in Table 2. We analyzed the relationship between antibiotic resistance and eradication outcome by calculating the eradication rate in different subgroups. Resistance to amoxicillin (MIC>0.125 μg/mL) was not found in any of the groups. The incidence of severe adverse reactions necessitating premature cessation of medication in the ITT population was 1.90 % (2/105), with a regimen compliance rate of 87.62 % (92/105).
Table 2.
Antibiotic Resistance, 23S rRNA traditional mutation sites and H. pylori Eradication Rate in ITT, MITT, and PP population.
| Characteristics | ITTc (n = 105) | MITT (n = 93) | PPd (n = 92) |
|---|---|---|---|
| AMX-Ra | 0 | 0 | 0 |
| CLA-Sa | 63.81 % (67/105) | 64.52 % (60/93) | 65.22 % (60/92) |
| Eradication success | 89.55 % (60/67) | 100 % (60/60) | 100 % (60/60) |
| CLA-Ra | 36.19 % (38/105) | 35.48 % (33/93) | 34.78 % (32/92) |
| Eradication success | 63.16 % (24/38) | 72.73 % (24/33) | 75 % (24/32) |
| Eradication Rate | 80.00 % (84/105) | 90.32 % (84/93) | 91.30 % (84/92) |
| 23S rRNA traditional mutation sitesb | 35.24 % (37/105) | 35.48 % (33/93) | 34.78 % (32/92) |
| Eradication success | 78.38 %(29/37) | 75.76 %(25/33) | 75 %(24/32) |
AMX, amoxicillin; CLA, clarithromycin; R, resistant.
Based on the recommendations of the EUCAST, the resistance breakpoints of amoxicillin and clarithromycin were defined as >0.125 μg/mL and >0.25 μg/mL.
23S r RNA traditional mutation sites include A2143G, A2142G and A2142C.
Patients who were lost to follow-up and terminated regimen were counted as eradication failure.
Patients with eradication failure due to medication compliance of less than 85 % were counted as eradication failure.
Based on the conventional MIC threshold, patients exhibiting resistance to clarithromycin continue to demonstrate a high eradication rate (63.16 %, 72.73 %, and 75 % in ITT, MITT and PP population, respectively). The eradication rates remained high in the presence of traditional mutation sites among patients resistant to clarithromycin (78.37 %, 75.75 %, and 75 % in ITT, MITT and PP population, respectively). These findings prompt us to conduct further investigations into resistance MIC breakpoints and genetic mutation sites that can reliably predict the eradication outcomes in the context of a 14-day clarithromycin bismuth quadruple therapy.
3.3. The MIC distribution of amoxicillin and clarithromycin
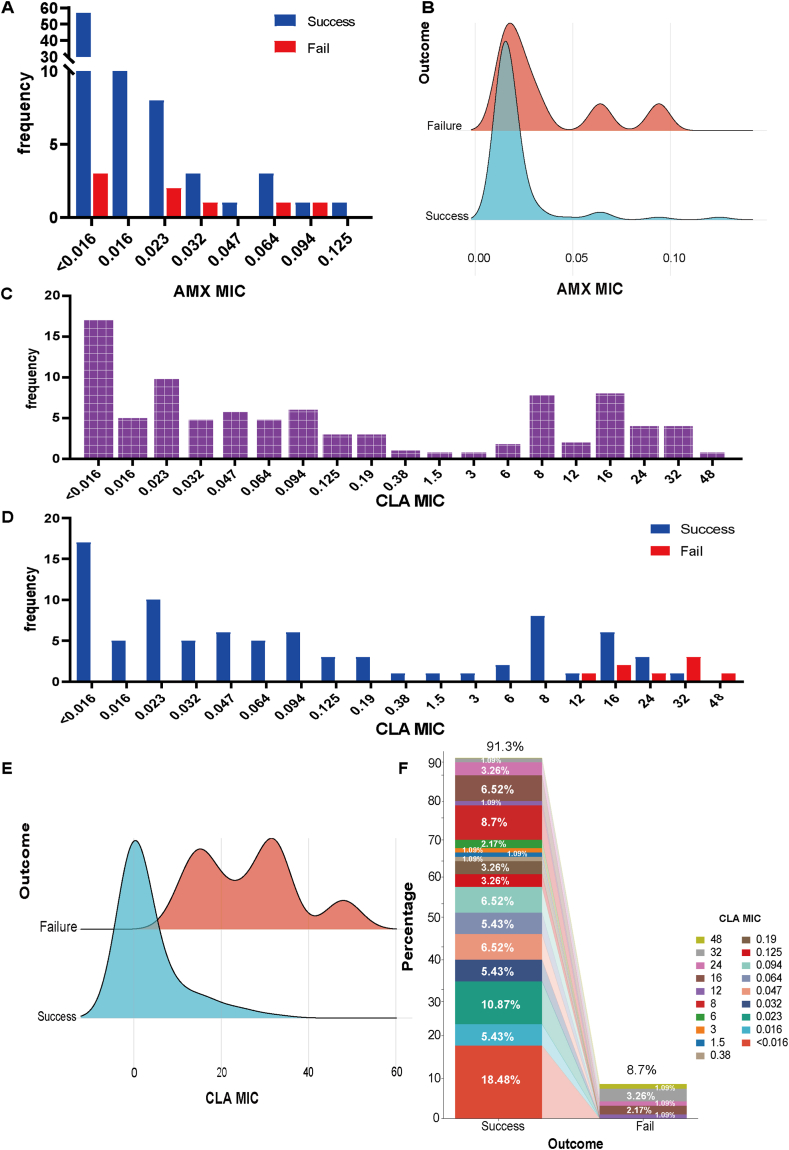
The MIC distribution of amoxicillin and clarithromycin was analyzed based on the PP population as shown in Fig. 2. As for amoxicillin, MIC values were positive skewness, and the main peaks were distributed in <0.016 μg/mL (Fig. 2A–B). Amoxicillin MIC ranged from <0.016 to 0.125 μg/mL and from <0.016 to 0.094 μg/mL in the eradication success and failure groups, respectively. The distribution of clarithromycin MIC exhibited a bimodal pattern (Fig. 2C–D). Fig. 2E provides a detailed depiction of the clarithromycin MIC distribution within the eradication success and failure groups. Among the 92 patients, 84 individuals (91.30 %) achieved successful eradication, with clarithromycin MIC distribution spanning from <0.016 to 32 μg/mL. Eight subjects failed eradication with MIC distribution ranging from 12 to 48 μg/mL. The MICs of patients with eradication failure were concentrated within the range of 12–48 μg/mL. The overall clarithromycin resistance rate was 34.78 % (32/92), yet eradication was still accomplished in 75.00 % (24/32) of these patients. The clarithromycin-resistant rate was 28.57 % (24/84) and 100.00 % (8/8) in eradication success and failure groups respectively (P < 00.000). Therefore, optimizing the clarithromycin resistance MIC breakpoint for predicting eradication outcomes in the context of 14-day clarithromycin bismuth quadruple therapy is essential.
Fig. 2.
The MIC distribution of amoxicillin and clarithromycin. (A) Frequency distribution of Amoxicillin MIC in different eradication outcome groups. (B) MIC distribution peaks of Amoxicillin in eradication success and failure groups. (C)Distribution of Clarithromycin MICs for 92 H. pylori strains. (D)Frequency distribution of Clarithromycin MIC in different eradication outcome groups. (E) MIC distribution peaks of Clarithromycin in eradication success and failure groups. (F) Correlation between the frequency proportion of different Clarithromycin MICs distribution in eradication success and failure groups.
3.4. Diagnostic test for predicting the optimal clarithromycin resistance MIC breakpoint in 14-day clarithromycin bismuth quadruple therapy
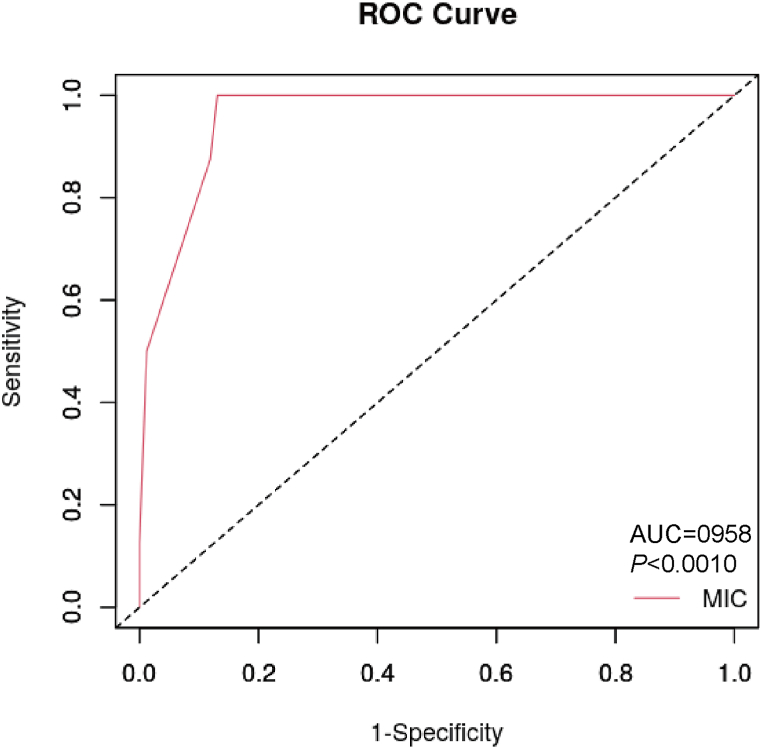
A substantial portion of patients, defined as resistant to clarithromycin based on the traditional MIC breakpoint, still achieved successful eradication. The actual eradication outcome of 14-day clarithromycin bismuth quadruple therapy was used as the gold standard to draw the ROC curve (Fig. 3) of the diagnostic ability of clarithromycin for each MIC level. Since MIC is determined through the method of multiple dilutions and lacks a continuous value, we established the optimal MIC breakpoint for precise prediction of the eradication outcome at 12 μg/mL. The area under the ROC curve (AUC) reached 0.958 (95%CI:0.914–1.000, P < 00.0001). The detailed data are described in Table 3.
Fig. 3.
The ROC curve of clarithromycin resistance MIC breakpoint in the PP population.
Table 3.
The detailed statistics of the ROC curve.
| Range of MIC values (μg/mL) | Sensitivity |
Specificity |
Youden index | LR+ | LR- |
|---|---|---|---|---|---|
| (95 % CI) | (95 % CI) | ||||
| >0.19 | 100.00(63.1–100.0) | 71.43(60.5–80.8) | 0.714 | 3.5 | 0 |
| >0.38 | 100.00(63.1–100.0) | 72.62(61.8–81.8) | 0.726 | 3.65 | 0 |
| >3 | 100.00(63.1–100.0) | 75.00(64.4–83.8) | 0.75 | 4 | 0 |
| >8 | 100.00(63.1–100.0) | 86.90(77.8–93.3) | 0.869 | 7.64 | 0 |
| >12 | 87.50(47.3–99.7) | 88.10(79.2–94.1) | 0.756 | 7.35 | 0.14 |
| >16 | 62.50(24.5–91.5) | 95.24(88.3–98.7) | 0.577 | 13.12 | 0.39 |
| >24 | 50.00(15.7–84.3) | 98.81(93.5–100.0) | 0.488 | 42 | 0.51 |
LR+, positive likelihood ratio; LR−, negative likelihood ratio.
3.5. 23S rRNA mutation sites and the outcome of eradication
The full-length 23S rRNA mutation in each clinical strain was assessed in reference to the standard strain 26695. Analyzing the connection between mutation sites and eradication outcomes posed a challenge when dealing with overlapping peaks. Among the data, 23 sites from eight patients showed overlapping peaks and they were all successfully eradicated. The overlapping peaks were regarded as mixed infections and they were all discarded from the data. The final peak maps were symmetrical, single, and complete in the remaining 91.30 % (84/92) patients.
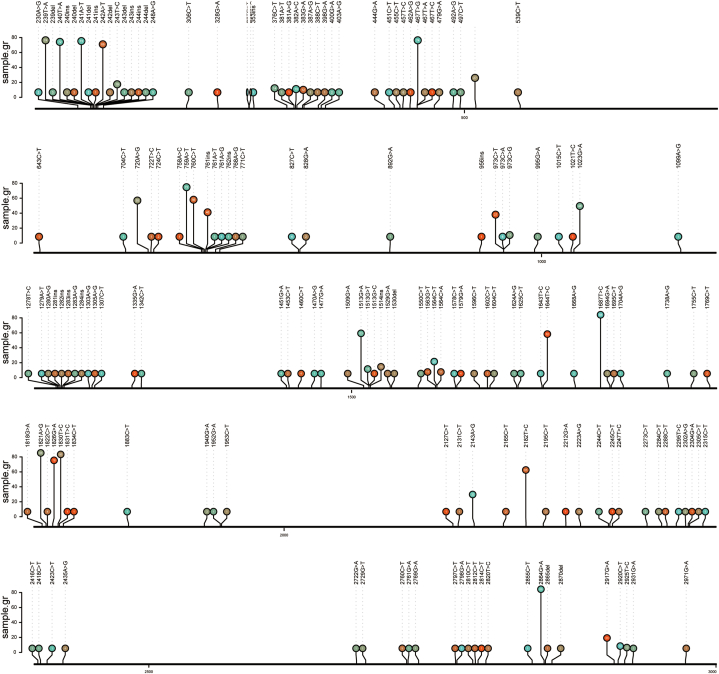
All 92 strains exhibited mutations in 23S rRNA gene sequences. Fig. 4 illustrates a total of 173 mutation sites identified within the 92 strains, including the traditional mutation 2143A > G but excluding 2142A > G or 2142A > C. We selected all the mutation sites that occurred in patients who experienced eradication failure and further discussed these sites in eradication success and failure groups separately.
Fig. 4.
The lollipop plot of 23S rRNA mutation sites in the PP population.
As shown in Table 4, all mutation sites in patients with eradication failure were monitored. Of these 50 sites as we have selected from the 173 mutation sites, only 2143A > G was statistically significantly different between successful and failed eradication groups (P < 0.000). The sensitivity and specificity of 2143A > G in predicting eradication outcome were 100 % (8/8) and 71.43 % (60/84) respectively.
Table 4.
The selected mutation sites in the eradication success and failure groups.
| Mutation | Frequency |
Eradication Success | Eradication Failure | p-Value |
|---|---|---|---|---|
| n(%) | ||||
| 239T > A | 74 (80.43 %) | 66 | 8 | 0.347 |
| 240T > A | 73 (79.35 %) | 65 | 8 | 0.198 |
| 241A > T | 72 (78.26 %) | 64 | 8 | 0.193 |
| 242A > T | 69 (75.00 %) | 62 | 7 | 0.674 |
| 243T > C | 21 (22.83 %) | 18 | 3 | 0.377 |
| 243ins | 1 (1.09 %) | 0 | 1 | 0.087 |
| 353ins | 1 (1.09 %) | 0 | 1 | 0.087 |
| 376C > T | 14 (15.22 %) | 12 | 2 | 0.351 |
| 381A > T | 1 (1.09 %) | 0 | 1 | 0.087 |
| 382A > C | 13 (14.13 %) | 12 | 1 | 1 |
| 383G > A | 12 (13.04 %) | 11 | 1 | 1 |
| 455C > T | 8 (8.70 %) | 7 | 1 | 0.532 |
| 457T > C | 6 (6.52 %) | 5 | 1 | 0.43 |
| 467T > G | 74 (80.43 %) | 67 | 7 | 1 |
| 492A > G | 7 (7.61 %) | 6 | 1 | 0.483 |
| 508C > T | 27 (29.35 %) | 25 | 2 | 1 |
| 720A > G | 55 (59.78 %) | 50 | 5 | 1 |
| 759A > T | 72 (78.26 %) | 65 | 7 | 0.681 |
| 760C > T | 56 (60.87 %) | 50 | 6 | 0.475 |
| 761ins | 40 (43.48 %) | 36 | 4 | 0.724 |
| 892G > A | 1 (1.09 %) | 0 | 1 | 0.087 |
| 973C > A | 6 (6.52 %) | 4 | 2 | 0.084 |
| 973C > T | 37 (40.22 %) | 34 | 3 | 1 |
| 1023G > A | 48 (52.17 %) | 42 | 6 | 0.271 |
| 1460C > T | 4 (4.35 %) | 3 | 1 | 0.309 |
| 1513G > A | 63 (68.48 %) | 57 | 6 | 1 |
| 1513G > T | 15 (16.30 %) | 13 | 2 | 0.612 |
| 1514ins | 18 (19.57 %) | 16 | 2 | 0.652 |
| 1563G > T | 11 (11.96 %) | 10 | 1 | 1 |
| 1564C > A | 11 (11.96 %) | 10 | 1 | 1 |
| 1564C > T | 25 (27.17 %) | 21 | 4 | 0.206 |
| 1604C > T | 1 (1.09 %) | 0 | 1 | 0.087 |
| 1644T > C | 62 (67.39 %) | 55 | 7 | 0.266 |
| 1687T > C | 88 (95.65 %) | 81 | 7 | 0.309 |
| 1704A > G | 1 (1.09 %) | 0 | 1 | 0.087 |
| 1755C > T | 2 (2.17 %) | 1 | 1 | 0.167 |
| 1821A > G | 88 (95.65 %) | 80 | 8 | 1 |
| 1826G > A | 78 (84.78 %) | 70 | 8 | 0.603 |
| 1830T > C | 86 (93.48 %) | 78 | 8 | 1 |
| 1883C > T | 1 (1.09 %) | 0 | 1 | 0.087 |
| 1953C > T | 1 (1.09 %) | 0 | 1 | 0.087 |
| 2143A > G | 32 (34.78 %) | 24 | 8 | 0 |
| 2182T > C | 65 (70.65 %) | 58 | 7 | 0.429 |
| 2245C > T | 1 (1.09 %) | 0 | 1 | 0.087 |
| 2435A > G | 4 (4.35 %) | 3 | 1 | 0.309 |
| 2760C > T | 9 (9.78 %) | 7 | 2 | 0.175 |
| 2797C > T | 1 (1.09 %) | 0 | 1 | 0.087 |
| 2864G > A | 89 (96.74 %) | 81 | 8 | 1 |
| 2917G > A | 23 (25.00 %) | 22 | 1 | 0.674 |
| 2925T > C | 10 (10.87 %) | 9 | 1 | 1 |
ins, insertion.
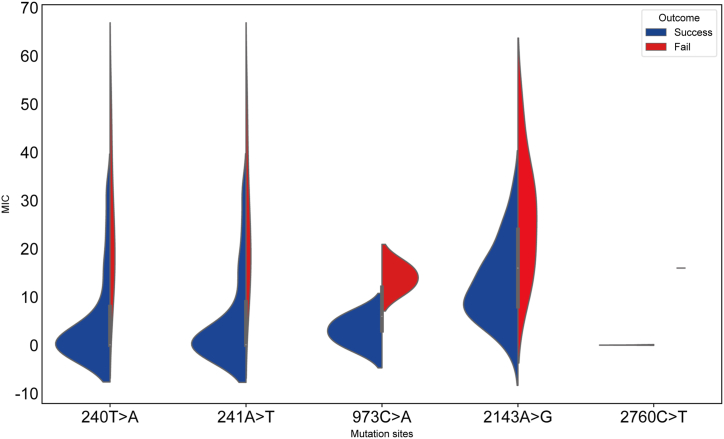
We identified a total of 16 sites with a p-value below 0.2, with 11 of them being unique to 1–2 patients who experienced eradication failure. Among the remaining 5 sites, which included 2143A > G, 973C > A, 2760C > T, 241A > T, and 240T > A, only 2143A > G (P < 00.000) exhibited an association with eradication outcomes (Table 4). Fig. 5 illustrates the corresponding frequency and MIC distribution of these five sites in both the eradication success and failure groups. The MIC values of strains with the 2143A > G mutation ranged from 0.016 to 48 μg/mL, with the MIC distribution peak centered at 8–32 μg/mL, indicating a relatively high MIC range.
Fig. 5.
Frequency and MIC distribution of selected five sites in eradication success and failure groups.
In summary, mutations of 243ins, 353ins, 381A > T, 892G > A, 1604C > T, 1704A > G, 1883C > T, 1953C > T, 2245C > T, and 2797C > T were observed in patients who experienced eradication failure, but the frequency of occurrence was infrequent, limited to only one patient. Other previously well-studied sites like 1821A > G and 2182T > C [[12], [13], [14]] were also found in our study, but their detection significance needs to be further explored. Apart from 2143A > G, no other site exhibited a significant association with eradication outcomes. However, it should be noted that the specificity of 2143A > G was not optimal to meet the clinical needs.
4. Discussion
Clarithromycin is routinely used to eradicate H. pylori. [3] Unfortunately, in many countries, H. pylori is increasingly developing resistance to clarithromycin, a major contributor to eradication failures [5,[15], [16], [17]]. A 15 % resistance rate serves as a threshold to differentiate regions with high and low clarithromycin resistance rates. As a result, empirical clarithromycin-based triple therapy is no longer recommended for use in high-prevalence areas [18]. Instead, bismuth-based quadruple therapy is now considered the first-line treatment option in regions with high clarithromycin resistance [4,19,]. Our study aimed to investigate precise methods for assessing clarithromycin resistance in 14-day clarithromycin bismuth quadruple therapy.
Our antibiotic susceptibility results reflected the epidemiological characteristics in Beijing, with a low rate (0 %) of resistance to amoxicillin and a relatively high rate (34.78 %) to clarithromycin. Our clarithromycin resistance rate (34.78 %) was lower than the global average of 50 % reported in a meta-analysis [20] and similar to rates observed in Nanjing (37.24 %) [21] and Japan (35.5 %) [22]. According to M. Miftahussurur et al.’ opinion [23], the agar dilution method (ADM) is the gold standard for the detection of H. pylori drug resistance, while E-test may overestimate the rates of resistance to clarithromycin. Thus, our results reliably revealed that the clarithromycin resistance rate in Beijing was lower than in other regions. Surprisingly, the eradication rate exceeded our expectations when we employed 14-day clarithromycin bismuth quadruple therapy empirically, reaching a remarkable rate of over 90 %. Additionally, severe adverse events were uncommon (1.90 %) and mostly mild (one had abdominal pain and one had nausea), and patient compliance was as high as 87.62 %.
The distribution of clarithromycin MIC showed a bimodal pattern as previously reported [24,25], which may be related to previous non-eradication-related macrolides. We assumed that excessive drug intake including self-medication and overmedication may further increase the MIC value, which in turn could indirectly affect the eradication rate. Hence, it is crucial to raise awareness about the importance of reducing drug misuse, particularly in developing countries [26]. Physicians should also consider inquiring about patients' previous medication history before prescribing antibiotics and staying informed about the local patterns of antibiotic resistance. These measures can significantly aid physicians in optimizing treatment strategies.
Among our enrolled patients, the highest observed MIC for clarithromycin was 48 μg/mL, and no strains exhibited an MIC of 256 μg/mL. This finding may be attributed to the fact that most of the enrolled patients were undergoing initial treatment. Notably, among the patients who experienced eradication failure, MIC values were less scattered below 12 μg/mL but notably concentrated at 32 μg/mL. This observation leads us to speculate that achieving higher MICs may involve more complex resistance mechanisms. To substantiate these hypotheses, further studies are warranted.
The traditional clarithromycin MIC breakpoint and traditional mutation sites are widely used to guide this regimen in clinical practice. Clarithromycin had all-or-none characteristics, such that if clarithromycin resistance is present, clarithromycin will not have any role in eradication [11,27]. We analyzed the efficacy of 14-day clarithromycin bismuth quadruple therapy to eradicate H. pylori at different clarithromycin MIC levels. The results of the ROC test showed that the optimal clarithromycin resistance MIC breakpoint was set at 12 μg/mL. It is worth noting that the MIC breakpoint for clarithromycin in EUCAST was established based on triple therapy. Given that the addition of bismuth can improve the eradication rate [11], current guidelines in China recommend quadruple therapy including bismuth as the first-line treatment for eradicating H. pylori, instead of triple therapy. This result indicated that the traditional clarithromycin MIC breakpoint provided by EUCAST was not suitable for 14-day clarithromycin bismuth quadruple therapy. Our study substantiates that, even in areas characterized by high clarithromycin resistance rates, the prescribed regimen can achieve commendably high eradication rates, thereby reinforcing clinicians' trust in this treatment approach. Consequently, it becomes imperative to use the MIC value of clarithromycin measured during quadruple therapy as a reference when quadruple therapy is used as the empirical treatment or the results of antibiotic susceptibility testing are used to guide treatment for H. pylori. It should be noted that there are two necessary conditions for the use of the clarithromycin MIC breakpoint in this study: (1) It is only suitable for 14-day clarithromycin bismuth quadruple therapy with a standard dose of clarithromycin (500mg/day). (2) The MIC breakpoint for clarithromycin resistance proposed in this study is only used for reference for results with the E-test method. Although 0.25 μg/mL can maximize the identification of patients who may fail to eradicate H. pylori, almost 75.00 % of patients who could have been successfully eradicated would lose the opportunity to use this regimen. Under the condition of ensuring patient compliance (87.62 %), MIC≥12 μg/mL had the highest sensitivity (100 %) and specificity (86.9 %) in predicting eradication outcomes in our study.
However, eradication was still successful in some patients with MIC ranging from 16 to 32 μg/mL, indicating that susceptibility testing in vitro could not fully predict susceptibility results in vivo. As many experts recommend, antibiotic susceptibility testing should be performed before treatment [13]. We believe that rather than using susceptibility results alone to guide medication regimens, confirming the MIC results of clarithromycin before treatment helps more to improve the eradication rate [28]. However, susceptibility testing relies on culture-based methods, which is difficult, time-consuming, and easily reduces the patient's medication compliance. We are supposed to use bismuth quadruple therapy in areas of high (>15 %) or unknown clarithromycin resistance when individual susceptibility testing is not available according to the Maastricht VI/Florence consensus report [4].
Furthermore, as pointed out by Enzo Ierardi et al. [29], geographical differences have an impact on antibiotic resistance, that is, contradictory resistance results may be obtained in different geographical areas or even in the same area. Therefore, it is necessary to take into account the resistance affected by geographical factors to further adjust the use of eradication therapies. Amoxicillin, for instance, has shown relatively low resistance levels in global studies. Nevertheless, recent findings indicate an upward trend in amoxicillin resistance, exemplified by rates of 8.42 % in South Korea [30] and 8.9 % in Southeastern Europe [31]. At present, there is a lack of updated regional studies on amoxicillin resistance in China. The situation is slightly more complicated for clarithromycin. A multi-center, cross-sectional study in China revealed a resistance rate of 50.83 % to clarithromycin among H. pylori infected patients [32]. The northeastern region exhibited the highest clarithromycin resistance at 75.28 %, significantly surpassing rates in North China, Northwest China, and East China (all exceeding 50 %). A prospective Spanish study showed 12.1 % [33] and an Australian study showed an almost 3.7 % annual increase in clarithromycin over the past 20 years, with an average resistance of over 20 % [34].
This study revealed that despite a clarithromycin resistance rate of 36.19 % based on traditional breakpoints, an impressive eradication rate of 91.3 % was achieved. Consequently, in regions with high clarithromycin resistance levels, such as Spain and Australia, successful eradication appears achievable with this therapeutic approach. These findings underscore the necessity for clinicians to understand the impact of geographical variations on resistance outcomes, influencing local eradication strategy selection. Moreover, it is crucial to monitor the MIC closely to accurately reassess antibiotic resistance and tailor the antibiotic regimen according to specific resistance patterns observed.
Now that we had found the new MIC breakpoint, we sought to further explore the resistance mutation sites that corresponded to the eradication outcome. C.-H. Lok al [35] believed that drug-resistant mutation sites could have accurate and stable detection results between different laboratories. Thus, finding statistically significant clarithromycin-resistant mutation sites is crucial to guide the eradication outcome. 2143A > G, 2142A > G, and 2142A > C mutation sites are the traditional resistance mutation sites, which are recognized as related to eradication failure in clinical [12,36,37]. However, we found that the specificity of predicting eradication only based on traditional mutation sites was poor and could not meet the clinical needs. Therefore, full-length 23S rRNA sequencing was used in our study. Our study performed a comprehensive screening and calculating of mutation sites for association with eradication outcomes by sequencing.
In our study, only 2143A > G was detected, with a rate of 34.78 %. We did not detect 2142A > C or 2142A > G in any of our cases in line with the results of many other studies owing to the small sample size [13,38,39]. Notably, all patients with eradication failure had 2143A > G. Though patients with 2143A > G did not necessarily have eradication failure, 2143A > G increased the likelihood of the chances of eradication failure. We found that 2143A > G was highly correlated with high MIC values, which was the same as in other studies [33,37,[40], [41], [42]].
50 mutation sites present in patients with eradication failure were analyzed in PP populations in our study. It was found that 10 mutation sites only occurred in eradication-failed strains, but these sites were sporadic and could not independently predict the eradication outcome. Above all, 49 sites were found to be mutated but not associated with outcome, including 1821A > G and 2182T > C [13,38], which has been studied many times before. Therefore, this was the first study put forward that monitoring clarithromycin-resistant mutation sites could not guide the eradication outcome of 14-day clarithromycin bismuth quadruple therapy accurately. Considering the diversity of point mutations in the region, the significance of these site mutations needs to be further studied [13,40].
Our study is the first report that measured MIC levels and the full-length 23S rRNA with high quality and accurately discussed their relationship with eradication outcomes in 14-day clarithromycin bismuth quadruple therapy. Few studies estimate phenotype and genotype simultaneously, let alone prospective analysis. Resistance to amoxicillin was not found in any of the groups. This allowed us to rule out the possibility of eradication failure due to amoxicillin resistance. Meanwhile, the high compliance and the low loss rate of follow-up guaranteed the quality of our study.
The most salient finding of the present study was to evaluate the clarithromycin resistance MIC breakpoint of 14-day clarithromycin bismuth quadruple therapy by analyzing MIC levels with eradication outcomes and to give specific values through ROC diagnostic test. Moreover, we detected all the corresponding genetic mutation sites and analyzed their relationship with eradication outcomes. Our study represents a pioneering endeavor in the exploration of the clarithromycin resistance MIC breakpoint tailored specifically for the 14-day clarithromycin bismuth quadruple therapy regimen.
Our study has several limitations. Firstly, the sample size is relatively small. Secondly, we did not consider influential factors such as smoking, gastric acid levels, and CYP2C19 status, which can impact eradication outcomes [3,43]. Thirdly, the clinical relevance of the clarithromycin resistance breakpoint we calculated in this study may be influenced by interactions with other drugs. Despite resistance in certain strains to clarithromycin, the efficacy of the quadruple therapy may still be promising due to the effectiveness of the two remaining active anti-infectives, amoxicillin and bismuth. Therefore, future research should conduct more comprehensive investigations.
5. Conclusion
The high eradication rate achieved with this quadruple regimen has made it a viable empirical treatment option for Chinese physicians. Nevertheless, conducting antimicrobial susceptibility tests remains imperative in guiding treatment decisions for patients with refractory H. pylori infections. It is important to emphasize the precise utilization of Minimum Inhibitory Concentration (MIC) values rather than solely relying on the binary classification of 'susceptible' or 'resistant.' Traditional MIC breakpoints and mutation sites have proven inadequate in accurately predicting H. pylori eradication outcomes, despite the notable association of the 2143A > G mutation with a heightened risk of treatment failure. Consequently, further research into MIC breakpoints and candidate genes is warranted in the context of 14-day clarithromycin bismuth quadruple therapy for H. pylori eradication.
Ethical approval
This study has been approved by the Ethic Committee of Beijing Hospital Medical Research Institutions (2021BJYYEC-274-01) and registered in the Chinese Clinical Trail Registry. Registration (ChiCTR2200061476).
Funding
This project was funded by the Chinese Academy of Medical Sciences (2021-I2M-C&T-B-091), the National Key R&D Program of China (2022YFC3602101), and the Chinese Academy of Medical Sciences (2021-I2M-1–050).
Data statement
The authors confirm that the data supporting the findings of this study are available within the article and publicly available repository. The data of this study are stored at National Center for Biotechnology Information (NCBI). GenBank submission: SUB13467252.
CRediT authorship contribution statement
Ying Chen: Writing – review & editing, Writing – original draft, Visualization, Software, Methodology, Investigation, Formal analysis, Data curation. Siyu Li: Writing – review & editing, Validation, Supervision, Software, Resources, Methodology, Investigation, Conceptualization. Wenbin Li: Validation, Supervision, Resources, Methodology, Investigation, Data curation, Conceptualization. Yiran Wang: Writing – review & editing, Visualization, Software, Methodology, Formal analysis, Conceptualization. Jihua Shi: Validation, Supervision, Methodology, Investigation, Conceptualization. Xue Xu: Validation, Methodology, Investigation. Pan Zhang: Validation, Methodology, Investigation. Dan Chen: Validation, Methodology. Rui Gang: Validation, Methodology. Qingfeng Luo: Writing – review & editing, Validation, Supervision, Resources, Project administration, Methodology, Investigation, Funding acquisition, Conceptualization.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- 1.Malfertheiner P., Camargo M.C., El-Omar E., et al. Helicobacter pylori infection. Nat. Rev. Dis. Prim. 2023;9(1):19. doi: 10.1038/s41572-023-00431-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ren S., Cai P., Liu Y., et al. Prevalence of Helicobacter pylori infection in China: a systematic review and meta-analysis. J gastroen hepatol. 2022;37(3):464–470. doi: 10.1111/jgh.15751. [DOI] [PubMed] [Google Scholar]
- 3.Pichon M., Pichard B., Barrioz T., et al. Diagnostic accuracy of a noninvasive test for detection of Helicobacter pylori and resistance to clarithromycin in stool by the amplidiag H. pylori+ClariR real-time PCR assay. J. Clin. Microbiol. 2020;58(4) doi: 10.1128/JCM.01787-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Malfertheiner P., Megraud F., Rokkas T., et al. Management of Helicobacter pylori infection: the Maastricht VI/Florence consensus report. Gut. 2022 doi: 10.1136/gutjnl-2012-302084. gutjnl-2022-327745. [DOI] [PubMed] [Google Scholar]
- 5.Savoldi A., Carrara E., Graham D.Y., et al. Prevalence of antibiotic resistance in Helicobacter pylori: a systematic review and meta-analysis in world health organization regions. Gastroenterology. 2018;155(5) doi: 10.1053/j.gastro.2018.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhang W., Chen Q., Liang X., et al. Bismuth, lansoprazole, amoxicillin and metronidazole or clarithromycin as first-line Helicobacter pylori therapy. Gut. 2015;64(11):1715–1720. doi: 10.1136/gutjnl-2015-309900. [DOI] [PubMed] [Google Scholar]
- 7.Song Z., Chen Y., Lu H., et al. Diagnosis and treatment of Helicobacter pylori infection by physicians in China: a nationwide cross-sectional study. Helicobacter. 2022;27(3) doi: 10.1111/hel.12889. [DOI] [PubMed] [Google Scholar]
- 8.Li Y., Choi H., Leung K., et al. Global prevalence of Helicobacter pylori infection between 1980 and 2022: a systematic review and meta-analysis. Lancet Gastroenterol Hepatol. 2023;8(6):553–564. doi: 10.1016/S2468-1253(23)00070-5. [DOI] [PubMed] [Google Scholar]
- 9.Versalovic J., Shortridge D., Kibler K., et al. Mutations in 23S rRNA are associated with clarithromycin resistance in Helicobacter pylori. Antimicrob agents ch. 1996;40(2):477–480. doi: 10.1128/aac.40.2.477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nezami B.G., Jani M., Alouani D., et al. Helicobacter pylori mutations detected by next-generation sequencing in formalin-fixed, paraffin-embedded gastric biopsy specimens are associated with treatment failure. J. Clin. Microbiol. 2019;57(7) doi: 10.1128/JCM.01834-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Dore M.P., Lu H., Graham D.Y. Role of bismuth in improving Helicobacter pylori eradication with triple therapy. Gut. 2016;65(5):870–878. doi: 10.1136/gutjnl-2015-311019. [DOI] [PubMed] [Google Scholar]
- 12.Marques A.T., Vítor J.M.B., Santos A., et al. Trends in Helicobacter pylori resistance to clarithromycin: from phenotypic to genomic approaches. Microb. Genom. 2020;6(3) doi: 10.1099/mgen.0.000344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Anis S., Farooqi S.R., Niaz S.K. Characterization of domain V mutations in clinical isolates of Helicobacter pylori in Pakistan and their effect on clarithromycin MIC. Infect. Drug Resist. 2021;14:3393–3403. doi: 10.2147/IDR.S306878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Tran V.H., Ha T.M.T., Le P.T.Q., et al. Characterisation of point mutations in domain V of the 23S rRNA gene of clinical Helicobacter pylori strains and clarithromycin-resistant phenotype in central Vietnam. Journal of Global Antimicrobial Resistance. 2019;16(null):87–91. doi: 10.1016/j.jgar.2018.09.012. [DOI] [PubMed] [Google Scholar]
- 15.Hong T.C., El-Omar E.M., Kuo Y.T., et al. Primary antibiotic resistance of Helicobacter pylori in the Asia-Pacific region between 1990 and 2022: an updated systematic review and meta-analysis. Lancet Gastroenterol Hepatol. 2024;9(1):56–67. doi: 10.1016/S2468-1253(23)00281-9. [DOI] [PubMed] [Google Scholar]
- 16.Megraud F., Bruyndonckx R., Coenen S., et al. Helicobacter pylori resistance to antibiotics in Europe in 2018 and its relationship to antibiotic consumption in the community. Gut. 2021;70(10):1815–1822. doi: 10.1136/gutjnl-2021-324032. [DOI] [PubMed] [Google Scholar]
- 17.Xu W., Yang B., Lin L., et al. Antibiotic resistance of Helicobacter pylori in Chinese children: a multicenter study from 2016 to 2023. Helicobacter. 2024;29(1) doi: 10.1111/hel.13038. [DOI] [PubMed] [Google Scholar]
- 18.Malfertheiner P., Megraud F., O'Morain C.A., et al. Management of Helicobacter pylori infection-the Maastricht V/florence consensus report. Gut. 2017;66(1) doi: 10.1136/gutjnl-2016-312288. [DOI] [PubMed] [Google Scholar]
- 19.Kim S.G., Jung H.-K., Lee H.L., et al. Guidelines for the diagnosis and treatment of Helicobacter pylori infection in Korea, 2013 revised edition. J. Gastroenterol. Hepatol. 2014;29(7):1371–1386. doi: 10.1111/jgh.12607. [DOI] [PubMed] [Google Scholar]
- 20.Hooi J.K.Y., Lai W.Y., Ng W.K., et al. Global prevalence of Helicobacter pylori infection: systematic review and meta-analysis. Gastroenterology. 2017;153(2):420–429. doi: 10.1053/j.gastro.2017.04.022. [DOI] [PubMed] [Google Scholar]
- 21.Wei W., Wang Z., Li C., et al. Antibiotic resistance of Helicobacter pylori in Nanjing, China: a cross-section study from 2018 to 2023. Front. Cell. Infect. Microbiol. 2023;13(null) doi: 10.3389/fcimb.2023.1294379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Okimoto T., Ando T., Sasaki M., et al. Antimicrobial-resistant Helicobacter pylori in Japan: report of nationwide surveillance for 2018-2020. Helicobacter. 2024;29(1) doi: 10.1111/hel.13028. [DOI] [PubMed] [Google Scholar]
- 23.Miftahussurur M., Fauzia K.A., Nusi I.A., et al. E-test versus agar dilution for antibiotic susceptibility testing of Helicobacter pylori: a comparison study. BMC Res. Notes. 2020;13(1):22. doi: 10.1186/s13104-019-4877-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kato S., Fujimura S., Udagawa H., et al. Antibiotic resistance of Helicobacter pylori strains in Japanese children. J. Clin. Microbiol. 2002;40(2):649–653. doi: 10.1128/JCM.40.2.649-653.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Inagaki T., Yagi T., Ichikawa K., et al. Evaluation of a rapid detection method of clarithromycin resistance genes in Mycobacterium avium complex isolates. J. Antimicrob. Chemother. 2011;66(4):722–729. doi: 10.1093/jac/dkq536. [DOI] [PubMed] [Google Scholar]
- 26.Chen Q., Long X., Ji Y., et al. Randomised controlled trial: susceptibility-guided therapy versus empiric bismuth quadruple therapy for first-line Helicobacter pylori treatment. Aliment. Pharmacol. Ther. 2019;49(11):1385–1394. doi: 10.1111/apt.15273. [DOI] [PubMed] [Google Scholar]
- 27.Graham D.Y. Helicobacter pylori update: gastric cancer, reliable therapy, and possible benefits. Gastroenterology. 2015;148(4) doi: 10.1053/j.gastro.2015.01.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.De Francesco V., Zullo A., Fiorini G., et al. Role of MIC levels of resistance to clarithromycin and metronidazole in Helicobacter pylori eradication. J. Antimicrob. Chemother. 2019;74(3):772–774. doi: 10.1093/jac/dky469. [DOI] [PubMed] [Google Scholar]
- 29.Ierardi E., Giorgio F., Losurdo G., et al. How antibiotic resistances could change Helicobacter pylori treatment: a matter of geography? World J. Gastroenterol. 2013;19(45):8168–8180. doi: 10.3748/wjg.v19.i45.8168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Park S.Y., Lee E.H., Kim D., et al. Novel mutations conferring amoxicillin resistance in Helicobacter pylori in South Korea. Antibiotics (Basel, Switzerland) 2023;12(4) doi: 10.3390/antibiotics12040748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Boyanova L., Kandilarov N., Hadzhiyski P., et al. Increase in amoxicillin resistance in Helicobacter pylori from Bulgarian patients over 15 years. Diagn. Microbiol. Infect. Dis. 2022;104(1) doi: 10.1016/j.diagmicrobio.2022.115746. [DOI] [PubMed] [Google Scholar]
- 32.Wang L., Li Z., Tay C.Y., et al. Multicentre, cross-sectional surveillance of Helicobacter pylori prevalence and antibiotic resistance to clarithromycin and levofloxacin in urban China using the string test coupled with quantitative PCR. Lancet Microbe. 2024;S2666–5247(24) doi: 10.1016/S2666-5247(24)00027-2. 00027-2. [DOI] [PubMed] [Google Scholar]
- 33.Mormeneo Bayo S., Belles Belles A., Vazquez Gomez D., et al. Antibiotic susceptibility and clarithromycin resistance determinants in Helicobacter pylori in the northeast of Spain: a one-year prospective study. Antibiotics (Basel, Switzerland) 2023;12(2):356. doi: 10.3390/antibiotics12020356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Schubert J.P., Warner M.S., Rayner C.K., et al. Increasing Helicobacter pylori clarithromycin resistance in Australia over 20 years. Intern. Med. J. 2022;52(9):1554–1560. doi: 10.1111/imj.15640. [DOI] [PubMed] [Google Scholar]
- 35.Lok C.-H., Zhu D., Wang J., et al. Phenotype and molecular detection of clarithromycin and levofloxacin resistance in Helicobacter pylori clinical isolates in beijing. Infect. Drug Resist. 2020;13:2145–2153. doi: 10.2147/IDR.S249370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Monno R., Giorgio F., Carmine P., et al. Helicobacter pylori clarithromycin resistance detected by Etest and TaqMan real-time polymerase chain reaction: a comparative study. APMIS. 2012;120(9):712–717. doi: 10.1111/j.1600-0463.2012.02896.x. [DOI] [PubMed] [Google Scholar]
- 37.Mégraud F. H pylori antibiotic resistance: prevalence, importance, and advances in testing. Gut. 2004;53(9):1374–1384. doi: 10.1136/gut.2003.022111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Gong E.J., Ahn J.Y., Kim J.M., et al. Genotypic and phenotypic resistance to clarithromycin in Helicobacter pylori strains. J. Clin. Med. 2020;9(6) doi: 10.3390/jcm9061930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kocazeybek B., Sakli M.K., Yuksel P., et al. Comparison of new and classical point mutations associated with clarithromycin resistance in Helicobacter pylori strains isolated from dyspeptic patients and their effects on phenotypic clarithromycin resistance. J. Med. Microbiol. 2019;68(4):566–573. doi: 10.1099/jmm.0.000944. [DOI] [PubMed] [Google Scholar]
- 40.Hussein R.A., Al-Ouqaili M.T.S., Majeed Y.H. Detection of clarithromycin resistance and 23SrRNA point mutations in clinical isolates of Helicobacter pylori isolates: phenotypic and molecular methods. Saudi J. Biol. Sci. 2022;29(1):513–520. doi: 10.1016/j.sjbs.2021.09.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.De Francesco V., Zullo A., Ierardi E., et al. Minimal inhibitory concentration (MIC) values and different point mutations in the 23S rRNA gene for clarithromycin resistance in Helicobacter pylori. Dig. Liver Dis. 2009;41(8):610–611. doi: 10.1016/j.dld.2009.01.001. [DOI] [PubMed] [Google Scholar]
- 42.Kageyama C., Sato M., Sakae H., et al. Increase in antibiotic resistant Helicobacter pylori in a university hospital in Japan. Infect. Drug Resist. 2019;12(null):597–602. doi: 10.2147/IDR.S196452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Thung I., Aramin H., Vavinskaya V., et al. Review article: the global emergence of Helicobacter pylori antibiotic resistance. Aliment. Pharmacol. Ther. 2016;43(4):514–533. doi: 10.1111/apt.13497. [DOI] [PMC free article] [PubMed] [Google Scholar]