ABSTRACT.
Delays in malaria diagnosis increase treatment failures and deaths. In endemic regions, standard diagnostic methods are microscopy and malaria rapid diagnostic tests (mRDTs) detecting Plasmodium falciparum histidine-rich protein 2/3 (PFHRP2/PFHRP3), but gene deletions can allow certain parasites to remain undetected. We enlisted a cohort comprising 207 symptomatic individuals, encompassing both children and adults, at a hospital in Nnewi, Nigeria. The prevalence of parasites was determined using a highly sensitive, species-specific quantitative polymerase chain reaction (SS-qPCR). Within a subset of 132 participants, we assessed the sensitivity and specificity of microscopy and HRP2-mRDTs in comparison to SS-qPCR for the detection of P. falciparum. We also investigated the prevalence of pfhrp2/pfhrp3 gene deletions. Greater sensitivity was achieved with mRDTs (95%) compared with microscopy (77%). Also, mRDTs exhibited greater specificity (68%) than microscopy (44%). The positive predictive value of mRDTs (89%) surpassed that of microscopy (80%), suggesting a greater probability of accurately indicating the presence of infection. The negative predictive value of mRDTs (82%) was far greater than microscopy (39%). Of the 165 P. falciparum–positive samples screened for pfhrp2/pfhrp3 gene deletions, one gene deletion was detected in one sample. Regarding infection prevalence, 84% were positive for Plasmodium spp. (by reverse transcription [RT]-qPCR), with P. falciparum responsible for the majority (97%) of positive cases. Thus, exclusive reliance on microscopy in endemic areas may impede control efforts resulting from false negatives, underscoring the necessity for enhanced training and advocating for high-throughput molecular testing such as RT-qPCR or qPCR at referral centers to address limitations.
INTRODUCTION
Among the six Plasmodium species that cause malaria in humans, Plasmodium falciparum is the most prevalent in sub-Saharan Africa, and is responsible for most deaths from malaria.1–3 According to the latest report by the WHO, there were an estimated 247 million malaria cases and 619,000 malaria deaths worldwide in 2021.1 Nigeria’s increased efforts and strategies to control malaria have not resulted in any reduction in disease incidence or mortality for several years.2,4,5 Thus, Nigeria accounted for most of the global mortality from malaria in 2021 (31%) compared with 2019 (24%).1,6,7 In addition, in 2021, 38% of global malaria deaths occurred in Nigerian children younger than 5 years old.1 Furthermore, alongside P. falciparum, other malaria-causing species (e.g., Plasmodium vivax, Plasmodium ovale wallikeri, Plasmodium ovale curtisi, and Plasmodium malariae) are also present.8–10
Fatal malaria, particularly in children, can be caused by a delay in malaria treatment that results from various factors, including misdiagnosis, severe coinfection, and self-medication. Malaria is conventionally diagnosed by microscopic examination of thick blood smears (TBSs). Because of the labor intensiveness and training needed to carry out this technique, and the issue of subjectivity,11 malaria rapid diagnostic tests (mRDTs) are now often used in the field.12 There are numerous mRDTs on the market, and almost all detect P. falciparum based on its exclusively expressed P. falciparum histidine-rich protein 2 (PFHRP2).13,14
Malaria rapid diagnostic tests have been shown to provide rapid and efficient diagnosis of malaria in field settings, making them a viable alternative to microscopy and helping to reduce the indiscriminate use of antimalarial drugs in endemic areas.15–17 However, recent reports indicate that, in some regions, P. falciparum escapes detection by mRDTs by pfhrp2/pfhrp3 deletion. This is widespread in the Amazon region,18 Ethiopia,19 and, more recently, in Equatorial Guinea.20 This could jeopardize the continued usefulness of HRP2-based mRDTs for malaria diagnosis. To overcome this challenge, nucleic acid amplification techniques (NAATs) could be used for the detection of Plasmodium infections.21 However, this is not a logistically suitable alternative, and unless new rapid diagnostic tests are developed, the routine use of TBSs will remain standard practice.
Most NAAT protocols detect Plasmodium spp. by using primers specific to the 18S RNA gene.22 Molecular methods can also be used to decipher the infecting Plasmodium species, but the cost of setting up and maintaining these techniques in hospitals in resource-limited settings is prohibitive. Extensive training is required before personnel can use this technique reliably to diagnose malaria, and the supply of required temperature-sensitive reagents is an additional barrier. Therefore, polymerase chain reaction (PCR)-based tests remain reference methods useful for validating other diagnostic tests and for aiding research in limited-resource environments. They can also be used in close-to-elimination settings to diagnose asymptomatic carriers.
In southeastern Nigeria, malaria diagnosis mainly relies on mRDTs and microscopy techniques.23,24 This study aims to assess the performance and validity of the commonly used diagnostic techniques (TBSs and mRDTs) in the Nnewi District, southeastern Nigeria, using highly sensitive species-specific quantitative PCR (SS-qPCR) as a reference technique. Poor performance of mRDTs could have many reasons; here, we evaluate the presence of pfhrp2/3 gene deletions that might impact the sensitivity of mRDTs. In this study, we also assess the presence and species composition of human Plasmodium infections in our cohorts.
MATERIALS AND METHODS
Patient population and sample collection.
Patients presenting with at least one of the symptoms of fever (>37.5°C), headache, vomiting, or malaise between February 2019 and January 2020 at Nnamdi Azikiwe University Teaching Hospital in Nnewi, Anambra State, southeastern Nigeria, were invited to be included in the study (Figure 1). In total, 207 patients were included (120 children of preschool and adolescent age and 87 adults).
Figure 1.
Geographic overview of the study sites in Nigeria. The shading represents the sampling site. This map was created using Mapchart, v. 4.2 (https://www.mapchart.net/africa.html).
Parasite detection by microscopy and mRDT.
From each participant, 2 mL of whole blood was collected into ethylenediaminetetraacetic acid containers. From all samples, one mRDT (SD BIOLINE, catalog no. 05FK90; Standard Diagnostics Inc., Seoul, Korea), which detects only HRP2, and one Giemsa-stained TBS was prepared. The remaining blood was frozen and stored until further processing. Thick blood smears were examined via microscopy by two independent readers blinded to each other, according to the WHO method, using standard procedures.25 Parasitemia was calculated as Parasite density = [(No. of parasites counted)/(White blood cells counted)] × 8,000 white blood cells/µL. Samples were designated negative if no parasites were detected after examining 200 fields through a 100× oil immersion objective on a thick blood film. The mRDT was performed according to the manufacturer’s instructions. For mRDTs, negative and positive results were reported as the appearance of a single or double band on the test strip, respectively.
Nucleic acid preparation.
According to the manufacturer’s protocol, total nucleic acids (DNA and RNA) were extracted from 200 µL thawed blood samples using the QIAamp Blood Mini Kit (Qiagen, Hilden, Germany). If the starting material was less than 200 µL, phosphate-buffered saline was added, and the elution volume was adjusted accordingly. The DNA yield was quantified using a NanoDrop 1000 (Thermo Fisher Scientific) and then stored at –20°C.
Pan-Plasmodium detection by reverse transcription qPCR.
To detect parasites, primers for all human Plasmodium species (conserved regions of 18S ribosomal DNA) were used in reverse transcription (RT) qPCR (pan-RT-qPCR) as described previously26 using TaqMan chemistry and hydrolysis probes. The reaction involved 2.5 µL of sample DNA (20–60 ng/µL), a 400-nM probe, 400 nM of each primer,26 5 µL TaqMan RT-PCR mix (Applied Biosystems), 0.25 µL TaqMan RT Enzyme mix (Applied Biosystems), and nuclease-free water in a total reaction volume of 10 µL. All primers were manufactured and purified by Eurofins Germany. The primers and TaqMan probe sequences used are listed in Supplemental Table 1. Reactions were performed using the LightCycler 480 Instrument II (Roche Diagnostics, Basel, Switzerland), with the following cycling conditions: RT at 48°C for 20 minutes, polymerase activation at 95°C for 10 minutes, 45 cycles of denaturation at 95°C for 15 seconds, annealing at 61°C for 1 minute, followed by a final cooling step at 40°C for 30 seconds. All tests were performed in a 384-well plate and technical duplicate. The reaction master mix and sample templates were distributed by an automated liquid handling system (QIAgility; Qiagen, Hilden, Germany). All assays included a nontemplate control and a positive control (P. falciparum strain 3D7 from cell culture) in duplicate. Positivity was considered after visual assessment of the amplification curves for variability between each sample replicate (deviation ≤1 cycle) and a cycle quantification (Cq) threshold value of ≤37.
Standard curve for quantification of parasite density by pan-RT-qPCR.
For pan-RT-qPCR, parasitemia (ranging from 4 parasites/mL to 27 × 106 parasites/mL) was estimated by correlating it to the nucleic acid extract of ring stages of synchronized P. falciparum 3D7 parasites. Parasitemia was quantified by extrapolating Cq values from the standard curve based on linear regression analysis. The standard curve was constructed with a dilution factor of 10. This quantification method may not be entirely comparable to microscopic readings of clinical infections, because discrepancies can arise as a result of the presence of gametocytes in clinical samples.
Identification of Plasmodium species.
Identification of Plasmodium species was done for all pan-RT-qPCR Plasmodium-positive samples. Plasmodium SS-qPCR identifying P. falciparum, P. malariae, P. vivax, P. ovale curtisi and P. ovale wallikeri targeted the 18S ribosomal DNA and involved a preamplification step to increase the concentration of the target genes in the sample, followed by SS-qPCR. For preamplification, a singleplex PCR reaction was performed using the Mastercycler nexus gradient (Eppendorf, Hamberg, Germany). The reaction involved 5 µL of sample DNA (20–60 ng/µL), 0.25 mM deoxynucleotide triphosphate, 2 µL magnesium chloride, 300 nM of each primer, 5 µL PCR Buffer (Qiagen, Hilden, Germany), 0.2 µL Taq DNA polymerase (Qiagen), and nuclease-free water in a total reaction volume of 50 µL. The cycling conditions were initial denaturation at 95°C for 5 minutes, then 20 or 30 cycles of amplification (denaturation at 95°C for 30 seconds, annealing at 58°C for 30 seconds, and elongation at 72°C for 90 seconds) followed by a final elongation at 72°C for 1 minute. The amplification was done at 20 or 30 cycles, and the criteria for cycling definition was based on pan-RT-qPCR: samples with cycle threshold (Ct) values were <30 or >30, respectively. Amplified samples were then stored at 4°C until their use in species identification qPCR. Primers and probe are listed in Supplemental Table 2.
The amplified samples were used as a template for a nested singleplex SS-qPCR in a reaction containing 2.5 µL of sample DNA (20–60 ng/µL), 400 nM of each speciation primer, 5 µL SensiFAST Probe No-ROX Mix (Bioline, Hamberg, Germany), and a 0.15-µL probe. The total reaction volume was 10 µL. The cycling program included initial denaturation at 95°C for 3 minutes, then 45 cycles of denaturation at 95°C for 10 seconds and annealing at 60°C for 30 seconds, followed by a final cooling at 40°C for 10 seconds. The primers and probes are as published27 and are shown in Supplemental Table 3. Their specificity was validated, and minimal formation of primers and dimers was confirmed. All primers were manufactured and high-purity, salt-free (HPSF)–purified by Eurofins. The reaction master mix and sample templates were distributed by an automated liquid handling system (QIAgility; Qiagen). Reactions were performed in technical duplicates. No-template, positive, and negative controls were included in this assessment. All the assays were tested and validated in accordance with the minimum information for publication of quantitative real-time PCR experiments guidelines28 for their specificity and showed no cross-species amplification.
Detection of pfhrp2 and pfhrp3 deletion.
Deletions of the pfhrp2/3 genes were investigated in 165 samples that tested positive for P. falciparum by SS-qPCR. This was done using a previously published fourplex qPCR protocol that detects simultaneously in one sample pfhrp2 and pfhrp3 genes as well as P. falciparum–specific cytochrome b (cytb) and P. falciparum β-tubulin (btub) as internal controls.29 The single-copy gene btub controls for sufficient template DNA for single-copy gene amplification such as pfhrp2 and pfhrpr3,29 ensuring accurate and reliable detection in the fourplex assay. The reaction included the following components: 1× TaqMan Multiplex Master Mix (Thermo Fisher Scientific, Darmstadt, Germany); 400 nM of pfhrp2, pfhrp3, and btub primers each; along with 100 nM of cytb primers each. In addition, there were 100 nM of btub probe, and 50 nM each of pfhrp2, pfhrp3, and cytb probes. The reaction mixture had 3 µL of DNA template, and nuclease-free water was added to achieve a final volume of 10 µL. The primers and probes were as published,29 and were manufactured and HPSF-purified by Eurofins (Supplemental Table 4). Primer specificity was validated. Reactions were performed using a LightCycler 480 Instrument II (Roche Diagnostics, Basel, Switzerland) with the following cycling conditions: initial activation step at 95°C for 20 seconds, followed by 45 cycles at 95°C for 3 seconds and at 60°C for 150 seconds. All tests were performed in a 384-well plate and technical triplicate. The reaction master mix and sample templates were distributed by an automated liquid handling system (QIAgility; Qiagen, Hilden, Germany). Appropriate no-template controls were included. The DNA from positive controls were included in the assay: P. falciparum laboratory strains 3D7 (pfhrp2 positive/pfhrp3 positive), Dd2 (pfhrp2 negative/pfhrp3 positive), and HB3 (pfhrp2 positive/pfhrp3 negative). The threshold for positivity for the respective target gene was Cq ≤35.
STATISTICAL ANALYSES
All data were entered and reviewed using Excel (Microsoft Corp., Redmond, WA). Further analysis was done with GraphPad Prism v. 6 (Boston, MA) for descriptive statistics. Diagnostic performance for microscopy, mRDTs, and RT-qPCR was analyzed using MedCalc v. 16.8 (Ostend, Belgium). Descriptive statistics were represented with frequencies and means. The chi-squared test was used to compare the diagnostic performance of microscopy and mRDTs using RT-qPCR as a standard. R v. 4.1.0 (R Foundation for Statistical Computing, Vienna, Austria) was used to perform correlation analysis of parasitemia with age.
RESULTS
Characteristics of the study population.
This was a hospital-based study, and 207 febrile patients were included, 120 (58%) of whom were children of preschool, school, and adolescent age; 87 (42%) were adults. The mean ages of children and adults were 7 years and 37.5 years, respectively. The age range for children spanned from 9 months to 18 years, whereas for adults, it extended from 19 to 93 years. Gender distribution was 47% female children and 53% male children, and 62% female adults and 38% male adults. Detailed demographic data are presented in Figure 2. Parasitemia and age relationship are in Supplemental Figure 1.
Figure 2.
Demographic characteristics of the study population.
Plasmodium spp. infection rate.
Plasmodium spp. infection prevalence evaluated by pan-RT-qPCR.
Initially, we conducted pan-RT-qPCR to detect Plasmodium spp. in all 207 samples. Of the febrile patients, 174 of 207 (84%) were infected with malaria parasites. Among them, children and adults had similar rates of Plasmodium spp. infection, with both groups showing an 84% infection rate.
Plasmodium species identification by SS-qPCR.
Plasmodium falciparum was the most common infecting species. Overall, P. falciparum was detected in 165 (80%) of the 207 febrile patients screened. There was a low frequency (6 of 207, 3%) of infection with non-falciparum Plasmodium species (three P. malariae, two P. ovale curtisi, and one P. ovale wallikeri), of which three were coinfected with P. falciparum (Figure 3).
Figure 3.
Distribution of Plasmodium species composition in the study population. Each circle represents 1% of a total 100%.
Performance evaluation of mRDTs, microscopy, and SS-qPCR.
For performance evaluation of the different malaria diagnostic methods, we included 132 individuals for which results of all three techniques (mRDTs, TBS microscopy, and SS-qPCR as a reference) were available. Of these 132 samples, 66 were from children and 66 were from adults. By SS-qPCR, 98 of 132 samples were positive for P. falciparum infection (Figure 4).
Figure 4.
Venn diagram for diagnostic tests. The included data comprise only those samples for which all three diagnostic tests were conducted. The number of positive (A) and negative (B) diagnostic tests results for microscopy, malaria rapid diagnostic tests (mRDTs), and species-specific quantitative polymerase chain reaction (SS-qPCR) are given. Overlapping sections represent the agreement of the results among the different tests. TBS = thick blood smear.
The diagnostic performance of the TBSs and the mRDTs versus SS-qPCR was compared (Table 1). Here, mRDTs demonstrated greater sensitivity (95%) in comparison with microscopy (77%), indicating a greater ability to identify P. falciparum–infected cases correctly. Similarly, mRDTs exhibited greater specificity (68%) than microscopy (44%), indicating a stronger accuracy in identifying negative cases correctly. The positive predictive value of mRDTs (89%) also surpassed that of microscopy (80%), suggesting a greater probability of indicating accurately the presence of the infection. Also, the negative predictive value of mRDTs (82%) was far greater than that of microscopy (39%).
Table 1.
Comparative performance of malaria rapid diagnostic tests and thick blood smear/microscopy vs. species-specific quantitative polymerase chain reaction
| SS-qPCR (ref.) | mRDT | 95% CI | P-Value | Microscopy | 95% Cl | P-Value |
|---|---|---|---|---|---|---|
| Sensitivity, % | 94.9 | 88.6–97.8 | – | 76.5 | 67.2–83.8 | – |
| Specificity, % | 67.6 | 50.8–80.8 | – | 44.1 | 28.8–60.5 | – |
| PPV, % | 89.4 | 82.1–93.9 | <0.0001* | 79.7 | 70.5–86.6 | 0.0284* |
| NPV, % | 82.1 | 64.4–92.1 | – | 39.4 | 25.6–55.2 | – |
| Likelihood ratio | 2.9 | – | – | 1.3 | – | – |
mRDT = malaria rapid diagnostic test; NPV = negative predictive value; PPV = positive predictive value; SS-qPCR = species-specific quantitative polymerase chain reaction.
Statistically significant at P <0.05.
Microscopy and SS-qPCR had concordant results for 75 positive samples and 15 negative samples, whereas mRDTs and SS-qPCR were concordant for 93 positive samples and 23 negative samples (Figure 4).
A positivity plot shows the performance of TBS and mRDT in relation to the Cq value of the Plasmodium spp. SS-qPCR (Figure 5). Most of the negative samples obtained through TBSs (TBS negative) and mRDTs (mRDT negative) consistently displayed Cq values predominantly >37 (0 parasites/mL). However, we observed some exceptions: two samples displaying Cq values within the range of 34 (33 parasites/mL) to 36 (10 parasites/mL) and two other samples with Cq values <30 (374 parasites/mL). Samples categorized as TBS negative/mRDT positive consistently exhibited Cq values <37, indicating their positive status and a failure in microscopy detection. Notably, there were six exceptions with Cq values >37, which may indicate false-positive results by mRDTs. For TBS positive/mRDT negative samples, all demonstrated Cq values >37. However, we observed two samples with Cq values <30 and one sample with a Cq value of 35.8 (8 parasites/mL). These samples could be a non-falciparum infection, a pfhrp2/pfhrp3 gene deletion, or a false diagnostic test result. Therefore, we tested all samples for pfhrp2 as well as pfhrp3 deletions, as described in the next section. Regarding the TBS-positive/mRDT-positive samples, most of them exhibited Cq values <37. Although many of the samples had Cq values <30, there were five exceptions within the range of 31 (219 parasites/mL) to 36 (10 parasites/mL) (Figure 5). Also, there were six samples greater than the threshold of 37.
Figure 5.
Qualitative agreement of thick blood smears (TBSs) and malaria rapid diagnostic tests (mRDTs) with species-specific quantitative polymerase chain reaction (qPCR) and the cycle quantification (Cq) value. The dotted line represents the threshold of positivity (Cq = 37). Samples with a greater Cq value are defined as being negative (–) for a Plasmodium spp. infection. + = positive infection.
Deletion of pfhrp2/pfhrp3 genes.
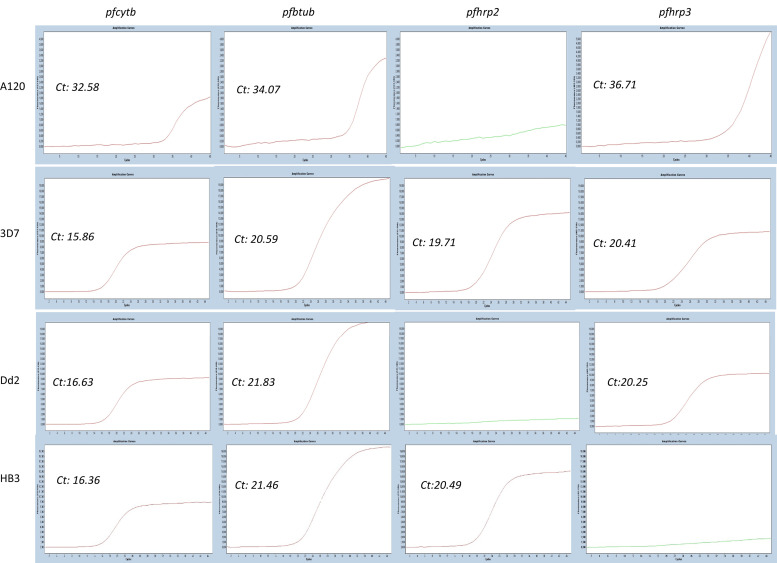
Of the 132 individuals with results available for all the three diagnostic tests (TBS, mRDT, and SS-qPCR), six were SS-qPCR positive but mRDT negative. Thus, these samples were potentially suspicious for pfhrp2/pfhrp3 deletion. Of these six samples, all were also confirmed for the presence of P. falciparum by the fourplex qPCR built-in pfcytb detection. All six samples passed the btub Cq quality control for DNA quantity for single-copy gene amplification. Of the six mRDT-negative samples, none had a deletion for pfhrp2 and/or pfhrp3 (mean Cq = 30.24, range = 22.00–36.08). However, of the 165 P. falciparum SS-qPCR–positive samples that we screened overall using fourplex qPCR, 158 were qualified by btub; of these, we found one sample with a pfhrp2 gene deletion (Figure 6). The sample came from a 4-year-old child (ID A120). The deletion assay was controlled with 3D7 (pfhrp2 positive/pfhrp3 positive), Dd2 (pfhrp2 negative/pfhrp3 positive), and HB3 (pfhrp2 positive/pfhrp3 negative) samples. The sample was positive by mRDT.
Figure 6.
Fourplex quantitative polymerase chain reaction Plasmodium falciparum histidine-rich 2 gene assay results of P. falciparum–positive samples. Ct = cycle threshold.
DISCUSSION
This study assessed the performance of the HRP2-based mRDT and TBS/microscopy for diagnosing malaria using P. falciparum–specific qPCR as reference diagnostics in samples from febrile patients in a hospital in southeastern Nigeria. In addition, Plasmodium species were identified, and the prevalence of pfhrp2/pfhrp3 gene deletions in P. falciparum–positive qPCR samples were evaluated.
Our results show that the diagnostic performance of mRDTs was better than microscopy in diagnosing P. falciparum malaria in this setting. The mRDTs demonstrated greater sensitivity compared with microscopy, indicating a greater ability to identify positive cases correctly. Similarly, mRDTs exhibited greater specificity than microscopy, indicating a stronger accuracy in identifying negative cases correctly. With SS-qPCR as the reference, microscopy recorded a false-negative rate greater than mRDTs. Also, microscopy had greater percentages of false-positive rates. In patient management, false-negative results are a significant problem because these individuals will return home without antimalarial treatment, potentially leading to fatal disease and continued parasite transmission. On the other hand, the occurrence of false positives may lead inadvertently to the unwarranted administration of antimalarial drugs. This is particularly concerning because the use of these drugs already faces heightened pressure resulting from the ongoing development of parasite resistance. Accurate diagnosis is a prerequisite for adequate disease control and prevention of the spread of resistance to antimalarial drugs. The observed differences in malaria diagnostic outcomes in the study region may be explained by low parasite densities, unexperienced laboratory staff, and inadequately stored and/or improperly conducted mRDTs.14,21,30 Also, variation in the amino acid sequence of the epitope targeted by the anti-HRP2 monoclonal antibody used in the rapid test31 could be a reason for our six mRDT false-negative samples.32
Our study shows that mRDTs perform better than microscopy. Thus, we recommend that, in addition to a TBS, an mRDT always be done. This is supported by other studies33–35 in this area that showed that mRDTs are adequate where well-trained microscopy personnel are not available. Enhanced diagnostics are particularly important to safeguard children from a potentially life-threatening P. falciparum infection if untreated. Furthermore, untreated P. falciparum infections can contribute to transmission in the region.
As expected, P. falciparum was the most prevalent species in our study population, with levels similar to what has been published for the WHO African region.6,36,37 Other species (P. malariae, P. ovale curtisi, and P. ovale wallikeri) were detected rarely, and P. vivax was absent, expectedly, as it is scarcely prevalent in the WHO African region.6 Parasitemia was, on average, slightly greater in children compared with adult and adolescent samples.
Our study provides valuable insights into the frequency of pfhrp2/pfhrp3 deletions in P. falciparum and its potential impact on mRDT diagnosis in southeastern Nigeria. A similar study38 in Central and West Africa (Nigeria inclusive) revealed a low prevalence of single pfhrp2/pfhrp3 deletions across multiple sites; however, no mRDTs were done in that study. In contrast, in our study, a large proportion of samples were assessed concomitantly by three diagnostic techniques. Reassuringly, the false-negative mRDT results of six samples were not a result of pfhrp2/3 deletions, but instead were most likely caused by too little parasitemia or operator issues. We identified one sample exhibiting a pfhrp2 deletion through the robustly established fourplex qPCR method, even though it tested positive with the mRDT. Mostly likely, the mRDT built-in anti-HRP2 monoclonal antibodies cross-reacted with the HRP3 protein, leading to a correct diagnostic result, or detected highly stable HRP2 that resulted from a prior infection.39,40 Eventually, the low parasitemia was at the threshold for detection, as can be seen by the Ct values, despite applying well-established and strict performance procedures of fourplex qPCR. Last, we cannot fully rule out a simple operator issue such as incorrect documentation of the mRDT result. Overall, the prevalence of pfhrp2-deleted P. falciparum strains circulating in the study area is very low.7 Deletions of pfhrp2/3 have been found predominantly in the Brazilian Amazon region,41 but reports of pfhrp2 or pfhrp3 deletions in African countries such as Mali,41 Kenya,42 the Democratic Republic of the Congo,43 Gabon,29 Ethiopia,3 and Eritrea44 have surfaced. More recently, there have also been reports of pfhrp2 deletions in symptomatic patients who traveled from Nigeria to Australia.45,46 Overall, pfhrp2 deletions currently do not have a considerable impact on the reliability of HRP2-based mRDTs. However, continuous monitoring of pfhrp2/pfhrp3 deletions as recommended by the WHO in endemic settings is warranted.
CONCLUSION
Our study demonstrated superior performance of HRP2-based mRDTs compared with microscopy for diagnosing malaria within our cohort in Nigeria. Nevertheless, in settings where well-trained personnel for microscopy are not readily available, we recommend the concurrent use of both methods. In addition, we emphasize the importance of regular training for health-care staff as a pivotal strategy to enhance diagnostic quality. Continuous training ensures that personnel are equipped with the latest knowledge and skills, ultimately contributing to more accurate and reliable malaria diagnoses. Differences in the diagnostic performance of microscopy, mRDT, and NAAT exist, and many factors may be implicated. One factor could be the deletion of P. falciparum pfhrp2/pfhrp3 genes, but the prevalence in our hospital cohort was extremely low and does not pose the HRP2-based mRDT as an immediate risk of failure.
Although PCR methodologies hold the greatest sensitivity levels, they are not feasible in daily clinical routine. However, in malaria research studies undertaken in Africa, they are of the utmost value. To improve diagnostic performance for malaria, investments in infrastructure and training are critical to combating malaria in a region known to have one of the greatest numbers of cases of malaria.
Supplemental Materials
ACKNOWLEDGMENTS
We thank the staff of the medical laboratory unit of Nnamdi Azikiwe University Teaching Hospital Nnewi, Anambra State, Nigeria. We also thank the Molecular Research Foundation for Students and Scientists (MRFSS) team for its invaluable contributions to sample collection and preliminary processing and analysis.
Note: Supplemental material appears at www.ajtmh.org.
Data Availability
The data sets used in this study are available at the repository of the Institute for Tropical Medicine, Germany.
REFERENCES
- 1. World Health Organization , 2022. World Malaria Report 2022. Geneva, Switzerland: WHO. [Google Scholar]
- 2. Rosenthal PJ, 2022. Malaria in 2022: Challenges and progress. Am J Trop Med Hyg 106: 1565–1567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Feleke SM. et al. , 2021. Plasmodium falciparum is evolving to escape malaria rapid diagnostic tests in Ethiopia. Nat Microbiol 6: 1289–1299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Gao L, Shi Q, Liu Z, Li Z, Dong X, 2023. Impact of the COVID-19 pandemic on malaria control in Africa: A preliminary analysis. Trop Med Infect Dis 8: 67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Jagannathan P, Kakuru A, 2022. Malaria in 2022: Increasing challenges, cautious optimism. Nat Commun 13: 2678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. World Health Organization , 2020. World Malaria Report 2020: 20 Years of Global Progress and Challenges. Geneva, Switzerland: WHO. [Google Scholar]
- 7. World Health Organization , 2019. World Malaria Report 2019. Geneva, Switzerland: WHO. [Google Scholar]
- 8. Aniweh Y. et al. , 2023. Comparative susceptibility of Plasmodium ovale and Plasmodium falciparum field isolates to reference and lead candidate antimalarial drugs in Ghana. Microbiol Spectr 11: e0491622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Herman C. et al. , 2023. Non-falciparum malaria infection and IgG seroprevalence among children under 15 years in Nigeria, 2018. Nat Commun 14: 1360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Sendor R. et al. , 2023. Similar prevalence of Plasmodium falciparum and non-P. falciparum malaria infections among schoolchildren, Tanzania. Emerg Infect Dis 29: 1143–1153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Kassile T, Lokina R, Mujinja P, Mmbando BP, 2014. Determinants of delay in care seeking among children under five with fever in Dodoma region, central Tanzania: A cross-sectional study. Malar J 13: 348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. White NJ, Watson JA, Uyongo S, Williams TN, Maitland KM, 2022. Substantial misdiagnosis of severe malaria in African children. Lancet 400: 807. [DOI] [PubMed] [Google Scholar]
- 13. Tangpukdee N, Duangdee C, Wilairatana P, Krudsood S, 2009. Malaria diagnosis: A brief review. Korean J Parasitol 47: 93–102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Wilson ML, 2012. Malaria rapid diagnostic tests. Clin Infect Dis 54: 1637–1641. [DOI] [PubMed] [Google Scholar]
- 15. Houzé S, 2017. Rapid diagnostic test for malaria. Bull Soc Pathol Exot 110: 49–54. [DOI] [PubMed] [Google Scholar]
- 16. Wanja EW, Kuya N, Moranga C, Hickman M, Johnson JD, Moseti C, Anova L, Ogutu B, Ohrt C, 2016. Field evaluation of diagnostic performance of malaria rapid diagnostic tests in western Kenya. Malar J 15: 456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Wongsrichanalai C, Barcus MJ, Muth S, Sutamihardja A, Wernsdorfer WH, 2007. A review of malaria diagnostic tools: Microscopy and rapid diagnostic test (RDT). Am J Trop Med Hyg 77 ( Suppl ): 119–127. [PubMed] [Google Scholar]
- 18. Góes L, Chamma-Siqueira N, Peres JM, Nascimento JM, Valle S, Arcanjo AR, Lacerda M, Blume L, Póvoa M, Viana G, 2020. Evaluation of histidine-rich proteins 2 and 3 gene deletions in Plasmodium falciparum in endemic areas of the Brazilian Amazon. Int J Environ Res Public Health 18: 123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Golassa L, Messele A, Amambua-Ngwa A, Swedberg G, 2020. High prevalence and extended deletions in Plasmodium falciparum hrp2/3 genomic loci in Ethiopia. PLoS One 15: e0241807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Berzosa P, González V, Taravillo L, Mayor A, Romay-Barja M, García L, Ncogo P, Riloha M, Benito A, 2020. First evidence of the deletion in the pfhrp2 and pfhrp3 genes in Plasmodium falciparum from Equatorial Guinea. Malar J 19: 99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Berzosa P. et al. , 2018. Comparison of three diagnostic methods (microscopy, RDT, and PCR) for the detection of malaria parasites in representative samples from Equatorial Guinea. Malar J 17: 333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Mangold KA, Manson RU, Koay ESC, Stephens L, Regner M, Thomson RB, Jr, Peterson LR, Kaul KL, 2005. Real-time PCR for detection and identification of Plasmodium spp. J Clin Microbiol 43: 2435–2440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Dozie U, Chukwuocha UM, 2016. Comparative evaluation of malaria rapid diagnostic test kits commercially available in parts of south eastern Nigeria. J Trop Dis 4: 201. [Google Scholar]
- 24. Okoro C, Chukwuocha UM, Nwakwuo GC, Ukaga CN, 2015. Presumptive diagnosis and treatment of malaria in febrile children in parts of south eastern Nigeria. J Infect Dis Ther 3: 2332-0766.100240. [Google Scholar]
- 25. Cheesbrough M, 2005. District Laboratory Practice in Tropical Countries, 2nd edition. Cambridge, United Kingdom: Cambridge University Press. [Google Scholar]
- 26. Groger M. et al. , 2018. Prospective clinical trial assessing species-specific efficacy of artemether-lumefantrine for the treatment of Plasmodium malariae, Plasmodium ovale, and mixed Plasmodium malaria in Gabon. Antimicrob Agents Chemother 62: e01758-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Woldearegai TG, Lalremruata A, Nguyen TT, Gmeiner M, Veletzky L, Tazemda-Kuitsouc GB, Matsiegui PB, Mordmüller B, Held J, 2019. Characterization of Plasmodium infections among inhabitants of rural areas in Gabon. Sci Rep 9: 9784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Bustin SA. et al. , 2009. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55: 611–622. [DOI] [PubMed] [Google Scholar]
- 29. Kreidenweiss A. et al. , 2019. Monitoring the threatened utility of malaria rapid diagnostic tests by novel high-throughput detection of Plasmodium falciparum hrp2 and hrp3 deletions: A cross-sectional, diagnostic accuracy study. EBioMedicine 50: 14–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. World Health Organization , 2017. False-Negative RDT Results and Implications of New Reports of P. falciparum Histidine-Rich Protein 2/3 Gene Deletions. Geneva, Switzerland: WHO. [Google Scholar]
- 31. Addai-Mensah O. et al. , 2020. Plasmodium falciparum histidine-rich protein 2 diversity in Ghana. Malar J 19: 256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Nolder D. et al. , 2021. Failure of rapid diagnostic tests in Plasmodium falciparum malaria cases among travelers to the UK and Ireland: Identification and characterisation of the parasites. Int J Infect Dis 108: 137–144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Ayogu EE, Ukwe CV, Nna EO, 2016. Assessing the reliability of microscopy and rapid diagnostic tests in malaria diagnosis in areas with varying parasite density among older children and adult patients in Nigeria. J Postgrad Med 62: 150–156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Azikiwe CC, Ifezulike CC, Siminialayi IM, Amazu LU, Enye JC, Nwakwunite OE, 2012. A comparative laboratory diagnosis of malaria: Microscopy versus rapid diagnostic test kits. Asian Pac J Trop Biomed 2: 307–310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Oboh MA, Oriero EC, Ndiaye T, Badiane AS, Ndiaye D, Amambua-Ngwa A, 2021. Comparative analysis of four malaria diagnostic tools and implications for malaria treatment in southwestern Nigeria. Int J Infect Dis 108: 377–381. [DOI] [PubMed] [Google Scholar]
- 36. Aribodor D, Njoku OO, Eneanya CI, Onyali IO, 2003. Studies on prevalence of malaria and management practices of the Azia community, Ihiala L.G.A., Anambra State, south-east Nigeria. Niger J Parasitol 24: 33–38. [Google Scholar]
- 37. Nwaorgu O, Orajaka B, 2011. Prevalence of malaria among children 1–10 years old in communities in Awka north local government area, Anambra State Southeast Nigeria. Afr Res Rev 5: 264–281. [Google Scholar]
- 38. Krueger T. et al. , 2023. Low prevalence of Plasmodium falciparum histidine-rich protein 2 and 3 gene deletions: A multiregional study in Central and West Africa. Pathogens 12: 455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Tian W, Yan P, Xu N, Chakravorty A, Liefke R, Xi Q, Wang Z, 2019. The HRP3 PWWP domain recognizes the minor groove of double-stranded DNA and recruits HRP3 to chromatin. Nucleic Acids Res 47: 5436–5448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Tran TM. et al. , 2013. An intensive longitudinal cohort study of Malian children and adults reveals no evidence of acquired immunity to Plasmodium falciparum infection. Clin Infect Dis 57: 40–47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Costa GL. et al. , 2021. A comprehensive analysis of the genetic diversity of Plasmodium falciparum histidine-rich protein 2 (PfHRP2) in the Brazilian Amazon. Front Cell Infect Microbiol 11: 742681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Koita OA. et al. , 2012. False-negative rapid diagnostic tests for malaria and deletion of the histidine-rich repeat region of the hrp2 gene. Am J Trop Med Hyg 86: 194–198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Beshir KB, Sepúlveda N, Bharmal J, Robinson A, Mwanguzi J, Busula AO, de Boer JG, Sutherland C, Cunningham J, Hopkins H, 2017. Plasmodium falciparum parasites with histidine-rich protein 2 (pfhrp2) and pfhrp3 gene deletions in two endemic regions of Kenya. Sci Rep 7: 14718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Parr JB. et al. , 2017. Pfhrp2-deleted Plasmodium falciparum parasites in the Democratic Republic of the Congo: A national cross-sectional survey. J Infect Dis 216: 36–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Berhane A, Russom M, Bahta I, Hagos F, Ghirmai M, Uqubay S, 2017. Rapid diagnostic tests failing to detect Plasmodium falciparum infections in Eritrea: An investigation of reported false negative RDT results. Malar J 16: 105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Prosser C, Gresty K, Ellis J, Meyer W, Anderson K, Lee R, Chng Q, 2021. Plasmodium falciparum histidine-rich protein 2 and 3 gene deletions in strains from Nigeria, Sudan, and South Sudan. Emerg Infect Dis 27: 471–479. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data sets used in this study are available at the repository of the Institute for Tropical Medicine, Germany.