Abstract
Lysosomes are single membrane-bound organelles containing acid hydrolases responsible for the degradation of cellular cargo and maintenance of cellular homeostasis. Lysosomes could originate from pre-existing endolysosomes or autolysosomes, acting as a critical juncture between autophagy and endocytosis. Stress that triggers lysosomal membrane permeabilization can be altered by ESCRT complexes; however, irreparable damage to the membrane results in the induction of a selective lysosomal degradation pathway, specifically lysophagy. Lysosomes play an indispensable role in different types of autophagy, including microautophagy, macroautophagy, and chaperone-mediated autophagy, and various cell death pathways such as lysosomal cell death, apoptotic cell death, and autophagic cell death. In this review, we discuss lysosomal reformation, maintenance, and degradation pathways following the involvement of the lysosome in autophagy and cell death, which are related to several pathophysiological conditions observed in humans.
Graphic abstract
Keywords: Lysosome, Autolysosome, Autophagic lysosome reformation, Autophagy, Autophagic cell death
Introduction
Lysosomes are single membrane-bound vesicular structures containing acid hydrolases and are known as the suicidal bag of cells. Lysosomes are characterized by a low internal pH in the approximate range of 4.5–5.5, which is maintained by v-ATPase [1–3]. This low pH facilitates the degradation of cargo present inside the cells, which may be internalized from the cell exterior via endocytosis or may be present inside the cells owing to the formation of autophagic vacuoles. Autophagy is an evolutionarily conserved lysosome-dependent catabolic process triggered by various forms of stress that contribute to the clearance of aggregated and damaged proteins and organelles to maintain cellular and metabolic homeostasis [4].
Lysosomes fuse with the outer membrane of mature autophagosomes and release acid hydrolases to degrade the autophagosomal content, thereby recycling cellular nutrients and metabolic intermediates. The final product of the autophagy network, namely, the autolysosome, contains cargo from autophagosomes and multivesicular bodies derived from the endosomal pathway [5, 6]. These are degraded by acid hydrolases, approximately 60 of which are present in lysosomes, and this fusion of autophagosomes and lysosomes occurs via a kiss-and-run mechanism [7, 8]. Conversely, lysosomal depletion by autophagy clearance adversely affects the lysosomal number inside the cell, which can be mitigated via alternative lysosomal biogenesis pathways such as reformation from endolysosomes and autolysosomes [9, 10]. In addition, intact lysosomal membrane permeability is critical in determining cellular survival or death, where lysosomal cathepsins play a significant role [11, 12].
This review discusses the role of lysosomes in autophagy and cell death, the various lysosomal biogenesis pathways, which contribute to the maintenance of lysosomal homeostasis, lysosome participation in autophagy, the involvement of lysosomes in mediating cell death, as well as the role of lysosomes in various human diseases, and as potential therapeutic targets for many autophagy and lysosomal storage disorders.
Lysosomal biogenesis
Lysosomes are involved in many cellular pathways, such as endocytosis and autophagy. Lysosomes can be formed directly as an end product of the endocytic pathway mediated via vesicular trafficking or indirectly by the lysosomal reformation pathway, which may originate from the endolysosomes or autolysosomes [13, 14]. The early endosomes receive cargo and mannose-6-phosphate receptors from the endocytic recycling endosomes or the trans-Golgi network [15, 16], while lysosomal proteins are produced in the endoplasmic reticulum. The lysosomes are transferred to the Golgi apparatus, from which they are directed towards the early endosomes or to the plasma membrane via a constitutive secretory pathway followed by endocytosis [17, 18]. The recycled cargo components are sorted into tubular sorting endosomes (TSE) and kept separate from the components marked for degradation. The early endosomes undergo maturation to form late endosomes or multivesicular bodies. The early endosomes and late endosomes are delineated by their specific marker proteins, such as Rab5 and EEA1 for early endosomes and Rab11 for late endosomes [16, 19, 21]. Early endosomes can undergo both fusion and fission events to form late endosomes. Moreover, the Rab cascade plays a critical role in endosomal biogenesis. The initial vesicles generated from the plasma membrane fuse with early endosomes with the help of Rab5 and are regulated by the guanine nucleotide exchange factor (GEF). Furthermore, these early endosomes can fuse with other early endosomes to form late endosomes with the activation of Rab7, which is mediated by the Rab5 effector, Mon1-Ccz1, and a mechanism involving the exchange between Rab5 and Rab7 via a positive feedback loop [20–22]. Subsequently, the lysosomes originate from matured late endosomes or from endolysosomes [21, 23]. These functionally active lysosomes are readily available for degradation processes involved in autophagy and endocytosis. Lysosomal biogenesis is further regulated by various transcription factors. The primary factors include the transcription factor EB (TFEB) and its regulatory molecules, such as mTORC1. TFEB is present in an inactive form in the cytoplasm as it is phosphorylated by mTORC1 at the Ser142 and Ser211 positions in the lysosomal membrane. Under stress conditions, mTORC1 is downregulated, resulting in the dephosphorylation and translocation of TFEB to the nucleus. Interestingly, TFEB enhances lysosomal biogenesis and transcription partners and subsequently regulates lysosomal biogenesis by binding to the promoter region of a 10-base E-box-like sequence called coordinated lysosomal expression and regulation (CLEAR). In addition to lysosomal biogenesis, TFEB also promotes autophagosome and autolysosome biogenesis by interaction with the CLEAR network. The function of TFEB is also regulated by other transcription factors such as PGC1α, FXR, CREB, and regulators such as MEK and cyclin-dependent kinases CDK4/6 [24–27].
Lysosomal reformation pathways
Lysosomes may also be formed from other subcellular structures, such as endolysosomes and autolysosomes (autophagic lysosome reformation). Fusion machinery plays a critical role in this regard and the relevant pathways are described below.
Reformation of lysosomes from endolysosomes
The formation of mature lysosome from the hybrid endolysosome reflects a unique conduit for lysosome reformation in the endolysosomal pathway. First, the late endosome or the multivesicular bodies fuse with the lysosome to form the endolysosome, a hybrid organelle similar to the autolysosome formed during autophagy [28]. Various proteins play an active role in these fusion events. The heterotypic fusion of endosomes and lysosomes is mediated by trans-SNARE, Syntaxin 7, Vti1b, Syntaxin 8, and VAMP7, whereas for homotypic fusion, VAMP7 is replaced by another R-SNARE protein, VAMP8 [29, 30]. Furthermore, N-ethylmaleimide sensitive factor (NSF) and soluble NSF attachment proteins (SNAPs) also play a vital role in these fusion events, along with the involvement of small Rab GTPase Rab7 [31, 32]. The actual mechanism involves SNARE insertion into the membrane, followed by tethering of the endosome and lysosomal lipid bilayer mediated by the HOPS complex. The endosome and lysosome then fuse to form endolysosomes, a process mediated at the interface of Rab7 with microtubule-dependent dynein via RILP and kinesins via FYCO1 [33]. v-ATPase and Ca2+ play essential roles in compartment acidification [34]. PIKfyve activity is crucial for the formation of lysosomes from endolysosomes. Furthermore, extensive membrane remodeling occurs after the synthesis of the lipid molecule PI(3,5)P2, and Ca2+ efflux is mediated by the effector TRPML1 [28]. Although the actual mechanism is unknown, the membrane is retrieved or buds off, followed by the condensation and recycling of SNAREs to form active functional lysosomes. This results in the longevity of lysosomal hydrolases and the recycling of lysosomes [34].
Reformation of lysosomes from autolysosomes
The process of regeneration of lysosomes from autolysosomes, termed autophagic lysosome reformation (ALR), is the termination of autophagy and an indirect method for lysosomal biogenesis [35]. The ALR process is inversely correlated with autophagy; that is, an increase in autophagy response reduces ALR and vice versa [36]. The overall ALR process involves the protrusion of tubules from autolysosomes, followed by the formation of protolysosomes and active lysosomes [10].
The first step in ALR is closely linked with autophagy and involves the reactivation of mTOR inside cells, which results in the upregulation of p70-S6 kinase 1 (p70S6K1) [36]. Inactivation of mTOR induces autophagy under stress conditions during the initial phases. However, prolonged stress causes reactivation of mTOR and initiation of the ALR response in the cells by decreasing autophagic flux and accumulation of LAMP1-LC3 dual positive vesicles called autolysosomes [36]. The activation of ATF4 (activating transcription factor 4), a transcription factor of the GAAC pathway or SLC7A5, a leucine transporter of the plasma membrane, facilitates mTOR reactivation [37]. Furthermore, a sugar transporter protein and putative lysosomal efflux permease, Spinster, regulate mTOR activation by affecting the nutrient status inside the cells [38]. mTOR activation is followed by tubulation of the autolysosome membrane. The endocytic protein, clathrin, and a member of the Wiskott-Aldrich syndrome protein (WASP) family, WHAMM, play a vital role in the tubulation process by forming a small bud from the membrane of the autolysosome [39, 40]. Subsequently, PtdIns(4)P-5-Kinase-1-Beta (PIP5K1B) facilitates the conversion of PI4P to PI(4,5)P2, which is the source of the budding vesicle. The clathrin on the surface recruits AP2 onto the PI(4,5)P2 to form the microdomain structure, and the motor protein KIF5B regulates the directive force along the microtubule [35]. Furthermore, dissociation of Rab7 and lipid molecules such as PI(4,5)P2, PI(4)P, and PI3P plays a prominent role in various stages of ALR progression [35, 41, 42]. In addition, PI3P, which is formed by VPS34 during the autophagy nucleation step, also has a potential role in tubulation events, primarily regulated by the VPS34-UVRAG complex [43]. The successful tubulation step is followed by a scission process aided by PIP5K1A. The large GTPase DNM2/dynamin 2 is also involved in the formation of protolysosomes that bind to PI(4,5)P2 [14, 35]. A protolysosome is formed after the scission of the budding tubule [44], which subsequently undergoes maturation (Fig. 1).
Fig. 1.
Lysosomal reformation pathways. Lysosomal proteins are synthesized in the ER and transferred to the Golgi apparatus. Subsequently, they are transferred to the early endosome directly or through a constitutive secretory pathway via the plasma membrane. Following this, Rab5 is replaced by Rab7, which results in early endosome maturation for the formation of the late endosome. Late endosomes undergo fusion events, either homotypic (VAMP8) or heterotypic (VAMP7) with HOPS and SNARE complex to form hybrid endolysosomes. Then PIKfyve converts PI(3)P to PI(3,5)P2 to form the budded tubule from the endolysosome, which subsequently breaks to form protolysosome. Autophagy nucleation occurs with the
source membrane derived from Golgi and ER. The phagophore is then elongated to form the autophagosome, which fuses with the lysosome to form the autolysosome. Subsequently, tubulation events occur with the help of the KIF5B and VPS34-UVRAG complex. PIP5K1B helps in converting PI(4)P to PI(4,5)P2. Then the budded tubule is fragmented with the help of DNM2 and PIP5K1A to form the protolysosome. The resultant protolysosomes undergo maturation to form the functional lysosomes, which are again available for the fusion events and degradation pathways
Lysosomal repair and lysophagy in lysosomal homeostasis
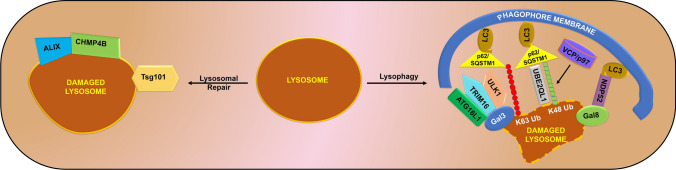
Specific stress signals damage lysosomes, resulting in the loss of lysosomal membrane potential. Several signaling cascades are activated and determine whether the lysosome should undergo a repair pathway to become functional again or be degraded via lysophagy [45]. The lysosomal membrane maintains its integrity with the assistance of Hsp70 by binding to bis(monoacylglycerol)phosphate, which leads to increased activity of acid sphingomyelinase and balances the lipid composition of the lysosomal membrane [46]. Furthermore, the endosomal sorting complexes required for transport (ESCRT) appear to play an active role in membrane repair. After permeabilization of the lysosomal membrane, Ca2+ ions released into the cytosol enhance the binding affinity of apoptosis-linked gene-2-interacting protein X and CHMP4B to the membrane for modulation of the repair process. In addition, Tsg101, another component of ESCRT, lacks Ca2+ binding affinity and helps recruit repair machinery to the lysosomal membrane [46, 47]. Small-scale damage can be repaired via the ESCRT pathway. Otherwise, lysosomes opt for self-degradation via lysophagy, a selective autophagy pathway [48, 49]. Lysophagy occurs only when the repair mechanism fails to be initiated due to extensive damage to the lysosomal membrane and the inability to recruit the ESCRT repair complexes. During lysophagy, galectins enter the lysosome, resulting in the exposure of lysosomal glycosylated proteins, and the ubiquitinated lysosomes are loaded onto expanded LC3 positive phagophores with the help of the adaptor protein p62/SQSTM1. The ubiquitinated lysosomes are subsequently engulfed into autophagosomes, which fuse with other intact lysosomes to degrade the inner cargo lysosomes [47, 50, 51]. Galectins (Gal) act as sensors for different types of lysosomal damage by binding to β-galactosides on the lumenal side of the lysosomal membrane. Galectin-1, -3, -8, and -9 are involved in lysosomal sensing. Gal3 recruits TRIM16, which helps in autophagic initiation, whereas Gal8 directly binds to NDP52, which recruits the LC3 positive phagophore. The TRIM16-Gal3 complex can sense the damaged state of lysosomes and recruit autophagic components to the damaged site on the lysosome via the K63-linked ubiquitination system of Ulk1 and ATG16L1. Furthermore, K63-linked chains originate from the lysosome within an hour of lysosomal damage and recruit p62/SQSTM1 for autophagic degradation. However, K48-linked chains emerge post 3–4 h after damage and are targeted by ubiquitin-directed AAA-ATPase VCP/p97 [49, 51–53]. In addition, the ubiquitin-conjugating enzyme UBE2QL1 helps maintain the integrity of lysosomes by translocating to the lysosomal damage site in the vicinity of galectins and ubiquitination targets. UBE2QL1 primarily regulates the K48-linked ubiquitination response in cells. Depletion of UBE2QL1 results in the abrogation of VCP/p97 recruitment and the accumulation of p62/SQSTM1 for inhibiting lysophagy [50]. This suggests that lysosomal damage can be compared to mitophagy, where only the damaged species is degraded. In the case of lysosomes, the autophagic components are recruited to the damaged site, and in the case of exhausted lysosomes, whole lysosomes are degraded by the process of lysophagy for damage clearance of the defective lysosomal pool (Fig. 2).
Fig. 2.
Lysosome repair or lysophagy. Stress response or excessive autophagy leads to the accumulation of exhausted lysosomes inside the cells. These lysosomal pools can be repaired via the recruitment of ALIX, CHMP4B, and ESCRT component Tsg101 in response to minor damage to the lysosomal membrane. In contrast, if the lysosomal membrane is extensively damaged, galactins sense this damage and are recruited onto the membrane-damaged site. Gal3 forms a complex with TRIM16 and recruits Ulk1 and ATG16L1 to the ubiquitination site, followed by the binding of p62/SQSTM1 to the LC3 positive phagophore membrane. Furthermore, K48 ubiquitination is regulated by the ubiquitin-directed AAA-ATPase VCP/p97 and the gene UBE2QL1. This complex binds to the autophagy adapter protein p62/SQSTM1 and subsequently binds to the LC3 positive phagophore membrane. Moreover, Gal8 can directly recruit NDP52 that can bind to LC3 positive phagophore to modulate lysophagy
Role of lysosome in the autophagic lysosomal pathway
Lysosomes play an active role in vesicular trafficking pathways, whereas autophagy acts as a dual regulator of mechanisms directing cellular fate. Hence, autophagy may serve as a signaling hub for cell survival or cell death in various scenarios [54]. At the basal level, the autophagy-initiation complex is generally inactive. However, under stress conditions, autophagy complexes are activated and modulate the autophagy-lysosomal axis to recycle nutrients by degrading macromolecules and cellular cargo [55]. In this regard, TFEB plays an essential role in the stage-specific regulation of autophagy. TFEB promotes the expression of the genes regulating autophagy initiation (BECN1, WIPI1, ATG9B, NRBF2), elongation (GABARAP, MAP1LC3B, ATG5), and fusion processes (UVRAG, RAB7) [56]. Furthermore, the TFEB activation and suppression machinery depend exclusively on the kinase activity of mTOR. TFEB remains a part of the mTORC1 complex, and mTORC1 helps in the phosphorylation of TFEB under amino acid-or nutrient-rich conditions. During nutrient depletion, mTOR is inactivated, and consequently, TFEB undergoes dephosphorylation and migrates towards the nucleus, where it activates the lysosomal biogenesis and autophagy regulatory machinery. TFEB activity is also regulated by signaling molecules such as ERK2, Akt, and PKCβ. ERK2 and mTORC1-mediated phosphorylation of TFEB prevent nuclear translocation, whereas Akt and PKCβ-mediated phosphorylation increases TFEB stability [56, 57].
Lysosomes play a critical role in controlling autophagic responses inside the cells. Autophagy can be divided into three types: microautophagy, macroautophagy, and chaperone-mediated autophagy (CMA) [58]. Microautophagy refers to the direct enwrapping of autophagic cargo into lysosomes. Here, the attachment of the cargo to the lysosomal membrane results in the invagination of the membrane, where the cargo is sequestered in the form of a bud. After internalization, cargo is degraded inside lysosomes, primarily by lysosomal hydrolases. Although the underlying molecular mechanism has not been extensively studied, the standard machinery involves ESCRTs and SNAREs in yeast and mammals [59–61]. This degradation may occur in three different patterns based on membrane dynamics: 1. Protrusion of the lysosomal membrane to engulf the cargo, 2. Invagination of the lysosomal membrane to entrap the cargo inside the lysosome, and 3. Invagination of the endosome to entrap the cargo inside endosomes [60]. Earlier, it was believed that the cargo undergoes macroautophagy or bulk autophagy for its degradation. However, recent advances have shown that prior to bulk degradation, selective protein degradation occurs via a process known as endosomal microautophagy, which is a prerequisite for the induction of bulk autophagy and the survival of the cells. This stage is mediated by degrading the selective proteins and autophagy receptors in mammals, plants, yeasts, and Drosophila [60, 62, 63].
Macroautophagy refers to the degradation of a wide range of subcellular materials inside cells and has been studied extensively. Macroautophagy is initiated by the formation of pre-autophagosomal structures called omegasomes in mammals. Phagophores can be formed from the membranes of various organelles, such as the ER, Golgi, mitochondria, and plasma membrane [64, 65]. Phagophore formation requires the activation of the initiation complex by unc-51 like autophagy activating kinase-1 (ULK1), which is autophosphorylated at Thr180 in response to mTORC1 and AMPK levels inside the cells. mTORC1 is localized to lysosomes under basal conditions and promotes growth by inhibiting the autophagic response and nuclear translocation of TFEB. During stress responses, mTORC1 dissociates from lysosomes and induces autophagy [66]. Following the activation, ULK1 phosphorylates its binding partner focal adhesion kinase (FAK)-family interacting protein of 200 kDa (FIP200) and ATG13. Downstream of this complex, ULK1 also phosphorylates Ser15 and other residues in Beclin-1 to activate the Beclin-1-PI3KC3-VPS15/p150 complex [4, 67]. Following these events, autophagy nucleation is completed by Beclin-1 activation. This is followed by membrane elongation and its closure by ESCRT components, particularly the ESCRT-III subunit [68, 69], to form a mature double membrane-bound autophagosome [70]. These mature autophagosomes fuse with lysosomes to form autolysosomes [71]. Autolysosomes are vesicular organelles that act as sites for cargo degradation, which takes place via lysosomal hydrolases, as mentioned previously [72]. SNARE insertion into the membrane recruits HOPS, a tethering complex that holds the autophagosome and lysosome together to facilitate the fusion process. Genes such as Rab7 and Syntaxin-17 (STX-17) also play an active role in the formation of fusion machinery. STX-17 interacts with SNAP29 and VAMP8 to form the STX-17-SNAP29-VAMP8 complex, which brings the vesicles close enough for fusion [73, 74]. Furthermore, TECPR1 interacts with PI3P and ATG12-ATG5 conjugates and facilitates the fusion of autophagosomes to lysosomes [75, 76]. After the completion of the fusion events, the autophagic flux process is triggered by lysosomal hydrolases, and the nutrients and energy are restored inside the cell [77, 78], thereby ensuring proper function (Fig. 3).
Fig. 3.
Role of lysosome in autophagy. a Microautophagy: Cargo gets sequestered onto lysosome with the help of ESCRT and SNARE assembly, which helps in the incorporation of the cargo inside the lysosome. After internalization of the cargo, it gets degraded with the help of lysosomal hydrolases. b Macroautophagy: Autophagy is initiated following the inhibition of mTOR by response initiated to stress, which leads to activation of the ULK1-FIP200-ATG13 complex. Post activation of the initiation complex, Beclin-1 gets activated, which remains in the complex form with VPS15 and PI3KC3. Subsequently, the phagophore is elongated, followed by the formation of a matured autophagosome. The lysosome fuses with the autophagosome to form autolysosome, where the sequestered cargo from the autophagosome gets degraded, and nutrients are recycled back into the cell. c Chaperone-mediated autophagy: Hsc70 remains in a complex form with other co-chaperones bag1, hsp40, hip, hop, and hsp90. This complex helps in the identification of cargo containing a KFERQ sequence. Then, it binds to the LAMP2A monomer on the lysosome. In this instance, LAMP2A forms a multimeric structure leading to the formation of a translocon. Translocon provides a passage for the cargo to move inside the lysosome with the help of Lys-Hsc70. Subsequently, the cargo undergoes degradation
In addition to macroautophagy, chaperone-mediated autophagy (CMA) also contributes to the degradation of protein molecules inside cells. This process is different from macroautophagy, as lysosomes are directly involved rather than depending on autophagosomes, as in the case of macroautophagy [79]. For degradation via the CMA pathway, a protein must have a KFERQ amino acid sequence in its structure [80]. The KFERQ sequence is present in 30% of cytosolic proteins. However, the incomplete sequence can behave like this motif after post-translational modification by mechanisms including acetylation, phosphorylation, and deamidation [81, 82]. The presence of the KFERQ sequence facilitates the binding of the proteins to the chaperone heat shock cognate 71 kDa protein (Hsc70), also known as HSPA8 in humans [79]. Hsc70 is stabilized by five co-chaperones: heat shock protein of 40 kDa (Hsp40), Hsc70 interacting protein (hip), hsc70‑hsp90 organizer protein (hop), heat shock protein 90 kDa (Hsp90), and Bcl2‑associated athanogene‑1 (bag‑1). Hsp40 aids the ATPase activity of Hsc70, which helps in substrate binding and release, whereas the hip protein helps to stabilize the binding of Hsc70, Hsp40, and target protein. Hop links Hsc70 and Hsp90. Hsp90 prevents the aggregation and/or refolding of unfolded proteins, and bag-1 acts as a regulator of Hsc70, which may be positive or negative [81]. After associating with the target protein, Hsc70 interacts with the cytosolic tail of LAMP2A in lysosomes. LAMP2A shares similar luminal regions with LAMP2B and LAMP2C but has a different transmembrane and cytosolic region. To date, only LAMP2A is believed to be involved in the CMA. LAMP2A has four positively charged residues in its C-terminal domain, which help in substrate binding [79, 82]. LAMP2A is a transmembrane protein existing in a monomeric form in the lysosomal membrane [83]. It forms a multimer after binding to the Hsc70-protein complex on the outer cytosolic side. This multimeric conversion requires the presence of a GXXG motif on the transmembrane sequence, and mutation of the sequence results in blockade to the multimerization process [82]. This multimeric LAMP2A forms a translocon that provides access to the attached protein to move through the membrane. The protein subsequently degrades with the help of lysosomal hydrolases [84]. Translocation of the protein across the membrane requires the presence of lys-hsc70 on the luminal side of the lysosomes, which may help in pulling the protein substrate inside the lysosome. Furthermore, during the conformational change of LAMP2A, it is stabilized by the lysosome-specific form of Hsp90 present on the luminal side. Once the substrate is degraded, LAMP2A reverts to its monomeric form [80, 85].
Lysosomes act as mediators in different modes of cell death
Although lysosomes are single membrane-bound organelles regarded as suicidal bags, they play a major role in cell death in that they determine cell fate and cellular survival. Hence, we may refer to lysosomes as a connecting link between cell survival and cell death. Starting from the initial organization of the living organism to maintain cellular homeostasis during stress and to check the proliferation of cancer cells, the various processes of regulated cell death play an essential role in determining cellular fates. Major forms of regulated cell death, such as apoptosis, pyroptosis, ferroptosis, and autophagy-dependent cell death, are cellular responses to uncontrolled stress [86]. However, in this review, we primarily discuss lysosomal cell death, apoptosis, and autophagy-dependent cell death.
Lysosomal cell death
Lysosomal cell death is caused by the permeabilization of the lysosomal membrane, which results in leakage of the internal milieus into the cytoplasm [87]. Lysosomal membrane integrity is compromised (lysosomal membrane permeabilization or LMP) upon degradation of proteins, such as HSP70, LAMP1, LAMP2, LIMP2, and CD63 [88]. Factors that induce lysosome-dependent cell death by triggering LMP, include the activity of caspases, calpains, and the destabilization of the lysosomal membrane due to the inhibition of v-ATPase [89]. ROS play a major role in lysosomal cell death, as elevated ROS levels result in lipid and protein peroxidation and subsequently lead to LMP. Hence, ROS-mediated LMP plays a significant role in cancer progression and metastasis [88]. Furthermore, all low extent of damage to the lysosomal membrane can be repaired by recruitment of ESCRT complexes onto the lysosomal membrane, which leads to the restoration of the acidic properties of lysosomes. When the lysosomal injury is significant and irreparable, lysosomes are degraded by lysophagy [45, 90], resulting in the activation of cell death pathways (Fig. 4).
Fig. 4.
Role of lysosome in cell death: Lysosome remains intact with the help of proteins such as LAMP1, LAMP2, CD63, HSP70, and LIMP2. Any damage to these molecular players results in lysosomal membrane permeabilization. It leads to the release of cathepsins to the cytosol and activates the cell death signaling pathways. Cathepsin B, H, K, L, and S inhibit Bcl-2, Bcl-xL, Bak, Mcl-1, and BimEL; however, they cannot inhibit Bax and Bid. Activation of Bax and Bid leads to cell death. Furthermore, DRAM1 is regulated by p53 and modulates cell death by triggering mitochondrial outer membrane permeabilization, lysosomal membrane permeabilization, and autophagy activation. EVA1A activates ATG16L1 to induce autophagic cell death. An association of Beclin-1 with Bcl-2 results in the inhibition of autophagy, which leads to cell death. However, when BNIP3 sequesters Bcl-2, Beclin-1 is free to induce autophagy, thereby resulting in autophagic cell death. Moreover, ROS generation inside the cells helps in the peroxidation of lipids and proteins, leading to lysosomal membrane permeabilization and, in turn, cell death. In contrast, the ESCRT complex can repair the damaged lysosomal membrane and prevents cell death
Apoptotic cell death and lysosomes
Apoptosis pathways can be divided into two major signaling cascades: extrinsic and intrinsic [91]. The extrinsic pathway is triggered by the binding of specific ligands to the transmembrane death receptors to activate cysteine-dependent aspartate-directed proteases (caspases), such as caspase-8 and caspase-10. This forms a death-inducing signaling complex (DISC), followed by the activation of executioner caspases, such as caspase-3, -6, and -7, to implement apoptotic cell death [91, 92]. In contrast, the intrinsic pathway of apoptosis, also called mitochondrial apoptosis, is initiated by a variety of cellular stresses such as oxidative or hypoxic stress, DNA damage, and deprivation of growth factors, which subsequently leads to the loss of mitochondrial membrane potential (MMP) and the release of cytochrome C into the cytosol [93, 94]. Cytochrome C with apoptotic protease-activating factor 1 (Apaf1) forms an apoptosome that activates caspase-9, which in turn activates caspase-3 to induce apoptosis. [91, 92]. Apoptotic cell death may be caspase-dependent or caspase-independent [12]. The process is mediated by releasing lysosomal proteases such as cysteine cathepsins and the aspartic protease cathepsin D into the cytosol. This process can be attributed to leakage in the lysosomal membrane or loss of lysosomal membrane integrity. Hence, lysosomal membrane permeabilization indirectly plays an active role in potentiating apoptosis [95]. LMP may be regulated by many factors, including the HSP70 chaperone protein family and various stimuli from ROS and mitochondria. Furthermore, different stimuli induce different types and/or patterns of LMP [96, 97]. The molecular targets for effective mediation of apoptosis with the help of cathepsins include molecules such as proapoptotic Bcl-2 protein Bax and Bid and the initiator caspase 8 in the case of neutrophil apoptotic cell death [12]. Furthermore, cysteine cathepsins B, L, S, K, and H cleave anti-apoptotic Bcl-2 family proteins such as Bcl-2, Bcl-xL, Mcl-1, Bak, and BimEL but cannot cleave the pro-apoptotic Bax. Hence, the release of these cathepsins results in the indirect activation of proapoptotic Bid and degradation of anti-apoptotic proteins to initiate a mitochondrial-dependent cell death pathway [98]. Various cathepsins such as aspartic cathepsin D and cysteine cathepsins B, C, F, H, K, L, O, S, V, W, and X are considered effector cathepsins that mediate apoptotic cell death [12]. Apoptotic caspases trigger cell death by activating secondary LMP. For instance, cleavage of caspases 2, 8, and 9 induces secondary LMP, leading to apoptosis. Furthermore, lysosomotropic detergents such as hydrophobic side-chains (e.g., imidazole and morpholine), ciprofloxacin o-methyl-serine dodecyl amine hydrochloride, sphingosine, and siramesine can damage the lysosomal membrane and induce LMP. This can be attributed to the weak nature of these agents, which can easily penetrate the lysosomes and become entrapped within to initiate the LMP response, leading to cell death. Moreover, sphingosine can activate lysosomal acid sphingomyelinase and acid ceramidase in TNF-treated rat hepatocytes and accumulate inside to trigger cell death by releasing cathepsin D [87].
Autophagic cell death and lysosomes
In addition to apoptosis, another form of cell death, autophagy-dependent cell death, is observed under specific circumstances. It is characterized by the excessive formation of autophagic vacuoles containing cytosolic components [99]. Although autophagy is mostly studied as a process promoting cell survival under stress conditions by regulating cellular homeostasis, it can favor cell death in several ways. In the case of autophagy-associated cell death, autophagy facilitates other forms of cell death, including apoptosis. Moreover, autophagy-dependent cell death is independent of different forms of cell death and is executed by the autophagy machinery [100]. In the absence of apoptosis, autophagy-dependent cell death plays a critical role in tissue remodeling during the developmental stages of mice [101, 102]. At the physiological level, autophagy-dependent cell death provides an alternative strategy for targeting cancer cell proliferation in apoptosis-resistant tumor cells [102]. For instance, DNA damage-regulated autophagy modulator 1 (DRAM1) is localized to the lysosome, activated by p53, and promotes p53-mediated apoptotic cell death by triggering LMP, cathepsin release, and subsequent mitochondrial outer membrane permeabilization (MOMP) [12]. Since DRAM1 regulates the autophagic response in cells, it can act as a master regulator of autophagic cell death, where autophagy induction leads to or supports the apoptotic pathways leading to cell death [103]. Furthermore, another gene, EVA1A/TMEM166, that interacts with ATG16L, is reported to enhance autophagosome biogenesis via the Beclin-1 independent pathway. EVA1A plays a dual role in cell death in that it can either trigger apoptosis via modulation of excessive autophagic flux with the help of autophagy inducers or assist or promote cell death due to the accumulation of autophagic vacuoles in the presence of autophagy fusion inhibitors such as CQ and Baf-A1 [104]. In addition, it has been reported that the compound Z18 can induce a non-canonical Beclin-1 independent autophagic death in HeLa cells. Moreover, it can interact with the BH3 binding groove of Bcl-XL/Bcl-2 and helps accumulate autophagic vacuoles, which subsequently leads to cell death in the case of HeLa cells [105]. In another study, hypoxia-triggered by cobalt chloride promotes Beclin-1 independent autophagy to induce cell death by activating ATG5/CHOP/Smac pathway through ER and mitochondrial stress in cancer cells [106].
The upregulation of HSP70 reduces the chances of LMP induction and, hence, abolishes caspase-independent and caspase-dependent cell death by interacting with and inhibiting Apaf1 and AIF [12]. Furthermore, the acetylation of HSP70 can induce an autophagic response and attenuate the likelihood of autophagic cell death by maintaining lysosomal integrity and modulating protective autophagy inside cells [107]. In addition, the major autophagosome marker protein, Beclin-1, was discovered as a Bcl2 interacting protein. When Beclin-1 dissociates from Bcl2, it is activated and induces autophagosome biogenesis under stress conditions. Hence, the sequestering agent of Bcl2, that is, BNIP3, can induce autophagy by inhibiting Beclin-1-Bcl2 complex formation and induce autophagic cell death. Moreover, death-associated protein kinase 1 (DAPK1) and protein phosphatase 2 (PP2A) induce autophagy via phosphorylation of Beclin-1, thereby increasing autophagic flux. This results in autophagic cell death, as demonstrated in the IM-9 cell line. Hence, the complex involving Beclin-1 and Bcl2 plays a pivotal role in autophagy and autophagy-dependent cell death [102, 108].
Role of lysosomes in human health and diseases
Cellular homeostasis plays a vital role in the viability and longevity of cellular systems. A dysfunction in any form of the homeostasis and degradative systems results in a deviation from the normal metabolic pathways and causes many pathophysiological disorders in humans. The lysosome is a crucial player in homeostasis maintenance and plays a significant role in various disease conditions. In addition to lysosomal action, the main pathways involved in the regulation and causative factors of these disorders include autophagy and apoptotic pathways.
Autophagy may be impaired due to treatment with exogenous compounds or genetic dysfunction. The autophagy-related cancer-suppressing peptide can induce autophagy via the AMPK/mTOR signaling pathway; however, it can also inhibit autophagic flux and alter lysosomal activity. Hence, the absence of fusion results in the accumulation of autophagic vacuoles, leading to the death of cervical cancer cells [109]. Fluvastatin nanoparticles induce autophagy by inhibiting mTORC1 signaling and autophagic flux by depleting lysosomes. Treatment with fluvastatin via a self-nano-emulsifying delivery system causes accumulation of autophagosomes, leading to the death of breast cancer cells [110]. Moreover, water-soluble yeast β-D-glucan (WSG) leads to an increase in lysosomal pH and a decrease in cathepsin activity in liver cancer. Owing to lysosomal dysfunction, autophagic flux remains impaired and causes activation of apoptotic signals, which results in the death of hepatocellular carcinoma (HCC) via ROS-dependent pathways [111]. Another synthetic compound, 163 N, inhibits the activity of ATG4B protein and promotes the alkalinization of lysosomes, resulting in the impairment of lysosomal protease activities. This novel synthetic lysosomal activity inhibitor alters autophagic flux and triggers anti-tumour activity in colorectal cancer [112]. Silver nanoparticles injure lysosomes in prostate cancer cells, as evidenced by the decreased intensity of LAMP1 staining in PC3 cells. In addition, these types of nanoparticles induce autophagy via the promotion of hypoxia but block autophagic flux, and hence, might act as novel anti-tumour agents against prostate cancer [113]. NDRG1 suppresses both basal and hypoxia-induced autophagy in pancreatic cancer cells, sensitizes the cells to the treatment of the Dp44mT-Cu complex and induces LMP. Furthermore, it synergistically enhances the effect of Dp44mT and gemcitabine in pancreatic cancer [114]. Prodigiosin inhibits autophagy in colorectal cancer cells by blocking autophagosome-lysosome fusion both in vitro and in vivo. It decreases the activity of cathepsins without altering the pH of lysosomes. Moreover, it enhances cell death in combination with 5-fluorouracil in colorectal cancer both in vitro and in vivo [115]. Co-treatment of simvastatin with temozolomide triggers apoptotic signaling in the case of glioblastoma cells, inhibits autophagic flux, and decreases the lysosomal biogenesis pathway, as indicated by LAMP-1 staining. Furthermore, the effect of these drugs cannot be prevented by ATG7 knockdown; hence, it might be an active inducer of autophagic cell death, which needs further investigation [116]. Furthermore, palmitic acid inhibits autophagic flux without altering lysosomal activity in hypothalamic neurons. It inhibits the fusion events of autophagy but triggers the enlargement of endolysosomal compartments by altering the activity of Rab7 and plays a prominent role in obesity-related cancers [117]. The amino acid transporter SLC38A9 has a critical role in maintaining essential amino acids in the lysosomes. Moreover, SLC38A9 also activates mTORC1 to promote Ras-driven pancreatic cancers, which depend on the macropinocytosis for nutrient balance inside the cells [118].
In addition to cancer, autophagy plays a vital role in multiple neurodegenerative diseases. Defective autophagy may lead to the accumulation of cargo, which triggers disease signals leading to the development of various neurodegenerative disorders, including Alzheimer’s disease (AD), Parkinson’s disease (PD), Huntington's disease, amyotrophic lateral sclerosis (ALS), and frontotemporal dementia [119]. AD patients demonstrate an accumulation of autophagosomes and a decrease in the number of lysosomes, as indicated by LC3 staining and cathepsin B and LAMP1 degradation, respectively. Moreover, AD cells attempt to degrade Aβ and tau proteins via the autophagic pathway. With age, autophagy becomes less pronounced in mammalian systems, leading to the accumulation of autophagosomes. Hence, the undegraded Aβ and tau proteins worsen AD symptoms in the elderly [120, 121]. Furthermore, apart from autophagic degradation, lysosomal and proteasomal degradation is essential in AD. Dysregulation of BACE2 in AD may be associated with the progression of AD, as AD leads to impairment of autophagic lysosomal degradation. Subsequently, impairment of lysosomal activity results in an increased half-life of BACE2. Hence, the cellular degradation system plays an essential role in the progression of AD [122]. Likewise, the accumulation of α-synuclein is a hallmark of PD. α-synuclein is degraded by the autophagic lysosomal pathway but not by proteasomal degradation. In addition to autophagic clearance, lysosomal cathepsin B, D, and L are essential for α-synuclein degradation. Moreover, a mutation in the GBA can result in Gaucher disease (GD), which increases the risk of PD in GD patients. As GD is a lysosomal storage disorder, it shows the importance of the autophagic lysosomal pathway in PD progression and therapeutics [123]. Deficiency of lysosomal acid alpha-glucosidase results in the accumulation of glycogen inside lysosomes, leading to Pompe disease in mammals [124]. Furthermore, the loss of lysosomal β-galactosidase results in the accumulation of GM1 ganglioside in the brain due to impaired autophagy in the case of GM1 gangliosidosis, indicating the critical role of autophagy in the case of lysosomal storage disorders [125]. Furthermore, mucopolysaccharidosis VI (MPS VI) or Maroteaux-Lamy syndrome is observed due to deficiency in N-acetylgalactosamine-4-sulfatase. Fibroblasts from the patients with this syndrome have shown impaired autophagic response due to the inhibition of cathepsin activity, which results in the accumulation of polyubiquitinated cargo and dysfunction of mitochondria. In contrast, autosomal recessive mucolipidosis type IV (MLIV) is caused by a mutation in the MCOLN1 gene (TRPML1). Accumulation of autophagosomes has been observed in fibroblast cells due to the inhibition of autolysosome formation, resulting in the accumulation of p62/SQSTM1 and dysfunctional mitochondria. Moreover, mucolipidosis type II (MLII) and mucolipidosis type III (MLIII) occur due to the absence of the enzyme N-acetylglucosamine 1-phosphotransferase (GlcNAc-phosphotransferase), which helps in the addition of mannose-6-phosphate to form lysosomal hydrolases. Mutations in this enzyme lead to missorting and loss of lysosomal enzymes, resulting in the accumulation of p62/SQSTM1 and dysfunctional mitochondria, as in MLIV [126]. In addition, Lafora disease is an autosomal recessive disorder in which mutations in EPM2A and EPM2B are observed, which encode laforin and malin, respectively. These mutations lead to an impaired autophagic response by activation of mTOR in fibroblast cells, as evidenced by the failure of autophagosome formation in EPM2A and EPM2B knockout mice. Furthermore, autophagy has been suppressed in Von Gierke’s disease, also known as glycogen storage disorder type Ia, due to activation of mTOR, the inability of TFEB translocation to the nucleus, and downregulation of SIRT1 expression [127]. As discussed, lysosomes play an essential role in the progressive modulation of different diseases, where lysosomal integrity and homeostasis are significantly affected. Hence, lysosomal stabilizing and autophagy modulating genes need to be targeted and sorted with natural and synthetic molecules to combat certain diseases, including autophagy and lysosome-related disorders, which provides the cue for the upcoming challenges in the future.
Conclusion and future perspectives
The lysosome, a key player in both autophagy and endosomal pathways, is a crucial platform for developing targeted therapeutic approaches for various diseases and basic research. In this review, we have discussed various pathways involved in the biogenesis of lysosomes and their role in autophagy and cell death, leading to the regulation of metabolic pathways. Several genes are involved in controlling lysosomal activity to modulate the fusion of autophagosomes and lysosomes. Internal acid hydrolases actively degrade inner cargo to facilitate nutrient recycling inside the cells. Moreover, the role of LMP has been extensively studied to establish the crosstalk between autophagy-dependent cell death and apoptosis modulated by lysosomes. This critical pathway provides a cue for determining a novel link between drug targets to control and manage the efficacy of various autophagy-related disorders, including neurological and lysosomal storage diseases. Furthermore, in this review, we demonstrate that the lysosome is a multipotent player in regulating the autophagy-lysosomal pathway, which can be further explored to find a solution for many autophagy and age-related disorders. These new insights can be integrated with endosomal trafficking pathways for the development of a new approach for cellular autophagic maintenance for stability, survival, and homeostasis.
Acknowledgements
KKM is highly acknowledged by the Government of India, Ministry of Science & Technology, Department of Science and Technology (No. DST/INSPIRE Fellowship/2017/IF170344). Research support was partly provided by the Science and Engineering Research Board (SERB) [EMR/2016/001246], Department of Science and Technology.
Abbreviations
- TFEB
Transcription factor EB
- mTORC1
Mechanistic target of rapamycin complex 1
- VAMP7
Vesicle associated membrane Protein 7
- VAMP8
Vesicle associated membrane Protein 8
- NSF
N-Ethylmaleimide sensitive factor
- SNAP
Soluble NSF attachment proteins
- RILP
Rab-interacting lysosomal protein
- p70S6K1
P70-S6 kinase 1
- LAMP1
Lysosomal-associated membrane protein 1
- ATF4
Activating transcription factor 4
- WASP
Wiskott-Aldrich syndrome protein
- PIP5K1B
PtdIns(4)P-5-kinase-1-baeta
- KIF5B
Kinesin family member 5B
- UVRAG
UV radiation resistance-associated gene
- DNM2
Dynamin2
- CHMP4B
Charged multivesicular body protein 4B
- TRIM16
Tripartite motif containing 16
- UBE2QL1
Ubiquitin conjugating enzyme E2 Q family like 1
- ULK1
Unc-51 like autophagy activating kinase-1
- FIP200
Focal adhesion kinase family interacting protein of 200 kDa
- BECN1
Beclin1
- WIPI1
WD repeat domain, phosphoinositide interacting 1
- NRBF2
Nuclear receptor-binding factor 2
- STX-17
Syntaxin-17
- TECPR1
Tectonin beta-propeller repeat containing 1
- Hsc70
Heat shock cognate 71 kDa protein
- Hsp40
Heat shock protein of 40 kDa
- hip
Hsc70 interacting protein
- hop
Hsc70-hsp90 organizer protein
- Hsp90
Heat shock protein of 90 kDa
- bag-1
Bcl2-associated athanogene-1
- caspase
Cysteine-dependent aspartate-directed proteases
- LIMP2
Lysosomal integral membrane protein type-2
- Apaf1
Apoptotic protease-activating factor 1
- DRAM1
DNA damage-regulated autophagy modulator
- DAPK1
Death-associated protein kinase 1
Declarations
Conflict of interest
The authors declare that they have no conflict of interest with this manuscript.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Koh J-Y, et al. Lysosomal dysfunction in proteinopathic neurodegenerative disorders: possible therapeutic roles of cAMP and zinc. Mol Brain. 2019;12(1):18–18. doi: 10.1186/s13041-019-0439-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Song Q, et al. The emerging roles of vacuolar-type ATPase-dependent Lysosomal acidification in neurodegenerative diseases. Transl Neurodegener. 2020;9(1):17–17. doi: 10.1186/s40035-020-00196-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Gyparaki M-T, Papavassiliou AG. Lysosome: the cell's ‘suicidal bag’ as a promising cancer target. Trends Mol Med. 2014;20(5):239–241. doi: 10.1016/j.molmed.2014.01.009. [DOI] [PubMed] [Google Scholar]
- 4.Mahapatra KK, et al. Molecular interplay of autophagy and endocytosis in human health and diseases. Biol Rev. 2019;94(4):1576–1590. doi: 10.1111/brv.12515. [DOI] [PubMed] [Google Scholar]
- 5.Schulze H, Kolter T, Sandhoff K. Principles of lysosomal membrane degradation: Cellular topology and biochemistry of lysosomal lipid degradation. Biochim Biophys Acta (BBA) 2009;1793(4):674–683. doi: 10.1016/j.bbamcr.2008.09.020. [DOI] [PubMed] [Google Scholar]
- 6.Lőrincz P, Juhász G. Autophagosome–lysosome fusion. J Mol Biol. 2020;432(8):2462–2482. doi: 10.1016/j.jmb.2019.10.028. [DOI] [PubMed] [Google Scholar]
- 7.Xu H, Ren D. Lysosomal physiology. Annu Rev Physiol. 2015;77:57–80. doi: 10.1146/annurev-physiol-021014-071649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jahreiss L, Menzies FM, Rubinsztein DC. The itinerary of autophagosomes: from peripheral formation to kiss-and-run fusion with lysosomes. Traffic (Copenhagen, Denmark) 2008;9(4):574–587. doi: 10.1111/j.1600-0854.2008.00701.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bright NA, Davis LJ, Luzio JP. Endolysosomes are the principal intracellular sites of acid hydrolase activity. Curr Biol. 2016;26(17):2233–2245. doi: 10.1016/j.cub.2016.06.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chen Y, Yu L. Autophagic lysosome reformation. Exp Cell Res. 2013;319(2):142–146. doi: 10.1016/j.yexcr.2012.09.004. [DOI] [PubMed] [Google Scholar]
- 11.Repnik U, Česen MH, Turk B. The endolysosomal system in cell death and survival. Cold Spring Harb Perspect Biol. 2013;5(1):a008755. doi: 10.1101/cshperspect.a008755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Mrschtik M, Ryan KM. Lysosomal proteins in cell death and autophagy. Febs j. 2015;282(10):1858–1870. doi: 10.1111/febs.13253. [DOI] [PubMed] [Google Scholar]
- 13.Luzio JP, et al. The biogenesis of lysosomes and lysosome-related organelles. Cold Spring Harb Perspect Biol. 2014;6(9):a016840–a016840. doi: 10.1101/cshperspect.a016840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chen Y, Yu L. Recent progress in autophagic lysosome reformation. Traffic. 2017;18(6):358–361. doi: 10.1111/tra.12484. [DOI] [PubMed] [Google Scholar]
- 15.Pfeffer SR. Multiple routes of protein transport from endosomes to the trans Golgi network. FEBS Lett. 2009;583(23):3811–3816. doi: 10.1016/j.febslet.2009.10.075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Saftig P, Klumperman J. Lysosome biogenesis and lysosomal membrane proteins: trafficking meets function. Nat Rev Mol Cell Biol. 2009;10(9):623–635. doi: 10.1038/nrm2745. [DOI] [PubMed] [Google Scholar]
- 17.Sun Z, Brodsky JL. Protein quality control in the secretory pathway. J Cell Biol. 2019;218(10):3171–3187. doi: 10.1083/jcb.201906047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zavodszky E, Hegde RS. Misfolded GPI-anchored proteins are escorted through the secretory pathway by ER-derived factors. eLife. 2019;8:e46740. doi: 10.7554/eLife.46740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hammerling BC, et al. A Rab5 endosomal pathway mediates Parkin-dependent mitochondrial clearance. Nat Commun. 2017;8(1):14050. doi: 10.1038/ncomms14050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Langemeyer L, et al. A conserved and regulated mechanism drives endosomal Rab transition. eLife. 2020;9:e56090. doi: 10.7554/eLife.56090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Poteryaev D, et al. Identification of the switch in early-to-late endosome transition. Cell. 2010;141(3):497–508. doi: 10.1016/j.cell.2010.03.011. [DOI] [PubMed] [Google Scholar]
- 22.Nagano M, et al. Rab5-mediated endosome formation is regulated at the trans-Golgi network. Commun Biol. 2019;2(1):419. doi: 10.1038/s42003-019-0670-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Hyttinen, J.M.T., et al., Maturation of autophagosomes and endosomes: A key role for Rab7. Biochimica et Biophysica Acta (BBA) - Molecular Cell Research, 2013. 1833(3): p. 503–510. [DOI] [PubMed]
- 24.Bala S, Szabo G. TFEB, a master regulator of lysosome biogenesis and autophagy, is a new player in alcoholic liver disease. Digest Med Res. 2018;1:16. doi: 10.21037/dmr.2018.09.03. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zhao B, et al. TFEB-mediated lysosomal biogenesis and lysosomal drug sequestration confer resistance to MEK inhibition in pancreatic cancer. Cell Death Discov. 2020;6(1):12. doi: 10.1038/s41420-020-0246-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Yin Q, et al. CDK4/6 regulate lysosome biogenesis through TFEB/TFE3. J Cell Biol. 2020;219(8):e201911036. doi: 10.1083/jcb.201911036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Palmieri M, et al. Characterization of the CLEAR network reveals an integrated control of cellular clearance pathways. Hum Mol Genet. 2011;20(19):3852–3866. doi: 10.1093/hmg/ddr306. [DOI] [PubMed] [Google Scholar]
- 28.Bissig C, et al. PIKfyve activity regulates reformation of terminal storage lysosomes from endolysosomes. Traffic. 2017;18(11):747–757. doi: 10.1111/tra.12525. [DOI] [PubMed] [Google Scholar]
- 29.Wang Y, et al. SNARE-mediated membrane fusion in autophagy. Semin Cell Dev Biol. 2016;60:97–104. doi: 10.1016/j.semcdb.2016.07.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Pryor PR, et al. Combinatorial SNARE complexes with VAMP7 or VAMP8 define different late endocytic fusion events. EMBO Rep. 2004;5(6):590–595. doi: 10.1038/sj.embor.7400150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Luzio JP, et al. Lysosome-endosome fusion and lysosome biogenesis. J Cell Sci. 2000;113(9):1515–1524. doi: 10.1242/jcs.113.9.1515. [DOI] [PubMed] [Google Scholar]
- 32.Luzio JP, Pryor PR, Bright NA. Lysosomes: fusion and function. Nat Rev Mol Cell Biol. 2007;8(8):622–632. doi: 10.1038/nrm2217. [DOI] [PubMed] [Google Scholar]
- 33.Inpanathan S, Botelho RJ. The lysosome signaling platform: adapting with the times. Front Cell Dev Biol. 2019;7:113. doi: 10.3389/fcell.2019.00113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Pryor PR, Luzio JP. Delivery of endocytosed membrane proteins to the lysosome. Biochim Biophys Acta (BBA) 2009;1793(4):615–624. doi: 10.1016/j.bbamcr.2008.12.022. [DOI] [PubMed] [Google Scholar]
- 35.Chen Y, Yu L. Development of research into autophagic lysosome reformation. Mol Cells. 2018;41(1):45–49. doi: 10.14348/molcells.2018.2265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yu L, et al. Termination of autophagy and reformation of lysosomes regulated by mTOR. Nature. 2010;465(7300):942–946. doi: 10.1038/nature09076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Chen R, et al. The general amino acid control pathway regulates mTOR and autophagy during serum/glutamine starvation. J Cell Biol. 2014;206(2):173–182. doi: 10.1083/jcb.201403009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Rong Y, et al. Spinster is required for autophagic lysosome reformation and mTOR reactivation following starvation. Proc Natl Acad Sci. 2011;108(19):7826–7831. doi: 10.1073/pnas.1013800108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Du W, et al. Kinesin 1 drives autolysosome tubulation. Dev Cell. 2016;37(4):326–336. doi: 10.1016/j.devcel.2016.04.014. [DOI] [PubMed] [Google Scholar]
- 40.Dai A, Yu L, Wang H-W. WHAMM initiates autolysosome tubulation by promoting actin polymerization on autolysosomes. Nat Commun. 2019;10(1):3699. doi: 10.1038/s41467-019-11694-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Magalhaes J, et al. Autophagic lysosome reformation dysfunction in glucocerebrosidase deficient cells: relevance to Parkinson disease. Hum Mol Genet. 2016;25(16):3432–3445. doi: 10.1093/hmg/ddw185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ktistakis NT, Tooze SA. PIPing on lysosome tubes. EMBO J. 2013;32(3):315–317. doi: 10.1038/emboj.2012.355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Munson MJ, et al. mTOR activates the VPS34-UVRAG complex to regulate autolysosomal tubulation and cell survival. EMBO J. 2015;34(17):2272–2290. doi: 10.15252/embj.201590992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Yu L, Chen Y, Tooze SA. Autophagy pathway: cellular and molecular mechanisms. Autophagy. 2018;14(2):207–215. doi: 10.1080/15548627.2017.1378838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Radulovic M, et al. ESCRT-mediated lysosome repair precedes lysophagy and promotes cell survival. EMBO J. 2018;37(21):e99753. doi: 10.15252/embj.201899753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Papadopoulos C, Kravic B, Meyer H. Repair or lysophagy: dealing with damaged lysosomes. J Mol Biol. 2020;432(1):231–239. doi: 10.1016/j.jmb.2019.08.010. [DOI] [PubMed] [Google Scholar]
- 47.Zhu S-Y, et al. Lysosomal quality control of cell fate: a novel therapeutic target for human diseases. Cell Death Dis. 2020;11(9):817. doi: 10.1038/s41419-020-03032-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hasegawa J, et al. Selective autophagy: lysophagy. Methods. 2015;75:128–132. doi: 10.1016/j.ymeth.2014.12.014. [DOI] [PubMed] [Google Scholar]
- 49.Papadopoulos C, Meyer H. Detection and clearance of damaged lysosomes by the endo-lysosomal damage response and lysophagy. Curr Biol. 2017;27(24):R1330–R1341. doi: 10.1016/j.cub.2017.11.012. [DOI] [PubMed] [Google Scholar]
- 50.Koerver L, et al. The ubiquitin-conjugating enzyme UBE2QL1 coordinates lysophagy in response to endolysosomal damage. EMBO Rep. 2019;20(10):e48014. doi: 10.15252/embr.201948014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Chauhan S, et al. TRIMs and galectins globally cooperate and TRIM16 and Galectin-3 co-direct autophagy in endomembrane damage homeostasis. Dev Cell. 2016;39(1):13–27. doi: 10.1016/j.devcel.2016.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Thurston TLM, et al. Galectin 8 targets damaged vesicles for autophagy to defend cells against bacterial invasion. Nature. 2012;482(7385):414–418. doi: 10.1038/nature10744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Fraiberg M, Elazar Z. A TRIM16-Galectin3 complex mediates autophagy of damaged endomembranes. Dev Cell. 2016;39(1):1–2. doi: 10.1016/j.devcel.2016.09.025. [DOI] [PubMed] [Google Scholar]
- 54.Nazio F, et al. Autophagy and cancer stem cells: molecular mechanisms and therapeutic applications. Cell Death Differ. 2019;26(4):690–702. doi: 10.1038/s41418-019-0292-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Kroemer G, Mariño G, Levine B. Autophagy and the integrated stress response. Mol Cell. 2010;40(2):280–293. doi: 10.1016/j.molcel.2010.09.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Di Malta C, Cinque L, Settembre C. Transcriptional regulation of autophagy: mechanisms and diseases. Front Cell Dev Biol. 2019;7:114. doi: 10.3389/fcell.2019.00114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Zhang W, et al. Regulation of TFEB activity and its potential as a therapeutic target against kidney diseases. Cell Death Discov. 2020;6(1):32. doi: 10.1038/s41420-020-0265-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Feng Y, et al. The machinery of macroautophagy. Cell Res. 2014;24(1):24–41. doi: 10.1038/cr.2013.168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Schuck S. Microautophagy—distinct molecular mechanisms handle cargoes of many sizes. J Cell Sci. 2020;133(17):246322. doi: 10.1242/jcs.246322. [DOI] [PubMed] [Google Scholar]
- 60.Oku M, Sakai Y. Three distinct types of microautophagy based on membrane dynamics and molecular machineries. BioEssays. 2018;40(6):1800008. doi: 10.1002/bies.201800008. [DOI] [PubMed] [Google Scholar]
- 61.Schäfer JA, et al. ESCRT machinery mediates selective microautophagy of endoplasmic reticulum in yeast. EMBO J. 2020;39(2):e102586. doi: 10.15252/embj.2019102586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Olsvik HL, et al. Endosomal microautophagy is an integrated part of the autophagic response to amino acid starvation. Autophagy. 2019;15(1):182–183. doi: 10.1080/15548627.2018.1532265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Mukherjee A, et al. Selective endosomal microautophagy is starvation-inducible in Drosophila. Autophagy. 2016;12(11):1984–1999. doi: 10.1080/15548627.2016.1208887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Abounit K, Scarabelli TM, McCauley RB. Autophagy in mammalian cells. World J Biol Chem. 2012;3(1):1–6. doi: 10.4331/wjbc.v3.i1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Rubinsztein DC, Shpilka T, Elazar Z. Mechanisms of autophagosome biogenesis. Curr Biol. 2012;22(1):29–34. doi: 10.1016/j.cub.2011.11.034. [DOI] [PubMed] [Google Scholar]
- 66.Yim WW-Y, Mizushima N. Lysosome biology in autophagy. Cell Discovery. 2020;6(1):6. doi: 10.1038/s41421-020-0141-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Hurley JH, Young LN. Mechanisms of autophagy initiation. Annu Rev Biochem. 2017;86:225–244. doi: 10.1146/annurev-biochem-061516-044820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Nakatogawa H. Mechanisms governing autophagosome biogenesis. Nat Rev Mol Cell Biol. 2020;21(8):439–458. doi: 10.1038/s41580-020-0241-0. [DOI] [PubMed] [Google Scholar]
- 69.Zhen Y, et al. ESCRT-mediated phagophore sealing during mitophagy. Autophagy. 2020;16(5):826–841. doi: 10.1080/15548627.2019.1639301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Hollenstein DM, Kraft C. Autophagosomes are formed at a distinct cellular structure. Curr Opin Cell Biol. 2020;65:50–57. doi: 10.1016/j.ceb.2020.02.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Hundeshagen P, et al. Concurrent detection of autolysosome formation and lysosomal degradation by flow cytometry in a high-content screen for inducers of autophagy. BMC Biol. 2011;9(1):38. doi: 10.1186/1741-7007-9-38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Hewitt G, Korolchuk VI. Repair, reuse, recycle: the expanding role of autophagy in genome maintenance. Trends Cell Biol. 2017;27(5):340–351. doi: 10.1016/j.tcb.2016.11.011. [DOI] [PubMed] [Google Scholar]
- 73.Liu R, Zhi X, Zhong Q. ATG14 controls SNARE-mediated autophagosome fusion with a lysosome. Autophagy. 2015;11(5):847–849. doi: 10.1080/15548627.2015.1037549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Yu S, Melia TJ. The coordination of membrane fission and fusion at the end of autophagosome maturation. Curr Opin Cell Biol. 2017;47:92–98. doi: 10.1016/j.ceb.2017.03.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Zheng P, et al. Recent advances in molecular mechanisms of autophagosome-lysosome fusion and impacts of exogenous factors. ES Food Agroforestry. 2020;1:53–76. [Google Scholar]
- 76.Chen D, et al. A mammalian autophagosome maturation mechanism mediated by TECPR1 and the Atg12-Atg5 conjugate. Mol Cell. 2012;45(5):629–641. doi: 10.1016/j.molcel.2011.12.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Pallet N, Livingston M, Dong Z. Emerging functions of autophagy in kidney transplantation. Am J Transplant. 2014;14(1):13–20. doi: 10.1111/ajt.12533. [DOI] [PubMed] [Google Scholar]
- 78.Tilija Pun N, Jang W-J, Jeong C-H. Role of autophagy in regulation of cancer cell death/apoptosis during anti-cancer therapy: focus on autophagy flux blockade. Arch Pharm Res. 2020;43(5):475–488. doi: 10.1007/s12272-020-01239-w. [DOI] [PubMed] [Google Scholar]
- 79.Alfaro IE, et al. Chaperone mediated autophagy in the crosstalk of neurodegenerative diseases and metabolic disorders. Front Endocrinol. 2019;9:778. doi: 10.3389/fendo.2018.00778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Cuervo AM, Wong E. Chaperone-mediated autophagy: roles in disease and aging. Cell Res. 2014;24(1):92–104. doi: 10.1038/cr.2013.153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Dice JF. Chaperone-mediated autophagy. Autophagy. 2007;3(4):295–299. doi: 10.4161/auto.4144. [DOI] [PubMed] [Google Scholar]
- 82.Kaushik S, et al. Chaperone-mediated autophagy at a glance. J Cell Sci. 2011;124(4):495–499. doi: 10.1242/jcs.073874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Rout AK, et al. Structure of transmembrane domain of lysosome-associated membrane protein type 2a (LAMP-2A) reveals key features for substrate specificity in chaperone-mediated autophagy. J Biol Chem. 2014;289(51):35111–35123. doi: 10.1074/jbc.M114.609446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Kaushik S, Cuervo AM. Chaperone-mediated autophagy: a unique way to enter the lysosome world. Trends Cell Biol. 2012;22(8):407–417. doi: 10.1016/j.tcb.2012.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Kaushik S, Cuervo AM. The coming of age of chaperone-mediated autophagy. Nat Rev Mol Cell Biol. 2018;19(6):365–381. doi: 10.1038/s41580-018-0001-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Tang D, et al. The molecular machinery of regulated cell death. Cell Res. 2019;29(5):347–364. doi: 10.1038/s41422-019-0164-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Aits S, Jäättelä M. Lysosomal cell death at a glance. J Cell Sci. 2013;126(9):1905–1912. doi: 10.1242/jcs.091181. [DOI] [PubMed] [Google Scholar]
- 88.Domagala A, et al. Typical and atypical inducers of lysosomal cell death: a promising anticancer strategy. Int J Mol Sci. 2018;19(8):2256. doi: 10.3390/ijms19082256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Johansson A-C, et al. Regulation of apoptosis-associated lysosomal membrane permeabilization. Apoptosis. 2010;15(5):527–540. doi: 10.1007/s10495-009-0452-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Wang F, Gómez-Sintes R, Boya P. Lysosomal membrane permeabilization and cell death. Traffic. 2018;19(12):918–931. doi: 10.1111/tra.12613. [DOI] [PubMed] [Google Scholar]
- 91.Mukhopadhyay S, et al. Autophagy and apoptosis: where do they meet? Apoptosis. 2014;19(4):555–566. doi: 10.1007/s10495-014-0967-2. [DOI] [PubMed] [Google Scholar]
- 92.Nirmala JG, Lopus M. Cell death mechanisms in eukaryotes. Cell Biol Toxicol. 2020;36(2):145–164. doi: 10.1007/s10565-019-09496-2. [DOI] [PubMed] [Google Scholar]
- 93.Galluzzi L, Kepp O, Kroemer G. Mitochondria: master regulators of danger signalling. Nat Rev Mol Cell Biol. 2012;13(12):780–788. doi: 10.1038/nrm3479. [DOI] [PubMed] [Google Scholar]
- 94.Westphal D, et al. Molecular biology of Bax and Bak activation and action. Biochim Biophys Acta (BBA) 2011;1813(4):521–531. doi: 10.1016/j.bbamcr.2010.12.019. [DOI] [PubMed] [Google Scholar]
- 95.Ivanova S, et al. Lysosomes in apoptosis. Methods Enzymol. 2008;442:183–199. doi: 10.1016/S0076-6879(08)01409-2. [DOI] [PubMed] [Google Scholar]
- 96.Boya P, Kroemer G. Lysosomal membrane permeabilization in cell death. Oncogene. 2008;27(50):6434–6451. doi: 10.1038/onc.2008.310. [DOI] [PubMed] [Google Scholar]
- 97.Fricker M, et al. Neuronal cell death. Physiol Rev. 2018;98(2):813–880. doi: 10.1152/physrev.00011.2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Droga-Mazovec G, et al. Cysteine cathepsins trigger caspase-dependent cell death through cleavage of bid and antiapoptotic Bcl-2 homologues. J Biol Chem. 2008;283(27):19140–19150. doi: 10.1074/jbc.M802513200. [DOI] [PubMed] [Google Scholar]
- 99.Galluzzi L, et al. Molecular mechanisms of cell death: recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 2018;25(3):486–541. doi: 10.1038/s41418-017-0012-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Denton D, Kumar S. Autophagy-dependent cell death. Cell Death Differ. 2019;26(4):605–616. doi: 10.1038/s41418-018-0252-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Arakawa S, et al. Role of Atg5-dependent cell death in the embryonic development of Bax/Bak double-knockout mice. Cell Death Differ. 2017;24(9):1598–1608. doi: 10.1038/cdd.2017.84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Bialik S, Dasari SK, Kimchi A. Autophagy-dependent cell death—where, how and why a cell eats itself to death. J Cell Sci. 2018;131(18):215152. doi: 10.1242/jcs.215152. [DOI] [PubMed] [Google Scholar]
- 103.Nagata M, et al. Dram1 regulates DNA damage-induced alternative autophagy. Cell Stress. 2018;2(3):55–65. doi: 10.15698/cst2018.03.127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Hu J, et al. TMEM166/EVA1A interacts with ATG16L1 and induces autophagosome formation and cell death. Cell Death Dis. 2016;7(8):e2323–e2323. doi: 10.1038/cddis.2016.230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Tian S, et al. Beclin 1-independent autophagy induced by a Bcl-XL/Bcl-2 targeting compound, Z18. Autophagy. 2010;6(8):1032–1041. doi: 10.4161/auto.6.8.13336. [DOI] [PubMed] [Google Scholar]
- 106.Sun L, et al. Beclin-1-independent autophagy mediates programmed cancer cell death through interplays with endoplasmic reticulum and/or mitochondria in colbat chloride-induced hypoxia. Am J Cancer Res. 2015;5(9):2626–2642. [PMC free article] [PubMed] [Google Scholar]
- 107.Park YH, et al. Hsp70 acetylation prevents caspase-dependent/independent apoptosis and autophagic cell death in cancer cells. Int J Oncol. 2017;51(2):573–578. doi: 10.3892/ijo.2017.4039. [DOI] [PubMed] [Google Scholar]
- 108.Decuypere J-P, Parys JB, Bultynck G. Regulation of the autophagic bcl-2/beclin 1 interaction. Cells. 2012;1(3):284–312. doi: 10.3390/cells1030284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Yang Y, et al. Lysosomal dysfunction and autophagy blockade contribute to autophagy-related cancer suppressing peptide-induced cytotoxic death of cervical cancer cells through the AMPK/mTOR pathway. J Exp Clin Cancer Res. 2020;39(1):197. doi: 10.1186/s13046-020-01701-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Elimam H, et al. Regulation of autophagy progress via lysosomal depletion by fluvastatin nanoparticle treatment in breast cancer cells. ACS Omega. 2020;5(25):15476–15486. doi: 10.1021/acsomega.0c01618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Wang N, et al. Yeast β-d-glucan exerts antitumour activity in liver cancer through impairing autophagy and lysosomal function, promoting reactive oxygen species production and apoptosis. Redox Biol. 2020;32:101495. doi: 10.1016/j.redox.2020.101495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Fu Y, et al. New anti-cancer strategy to suppress colorectal cancer growth through inhibition of ATG4B and lysosome function. Cancers. 2020;12(6):1523. doi: 10.3390/cancers12061523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Chen Y, et al. Silver nanoparticles regulate autophagy through lysosome injury and cell hypoxia in prostate cancer cells. J Biochem Mol Toxicol. 2020;34(5):e22474. doi: 10.1002/jbt.22474. [DOI] [PubMed] [Google Scholar]
- 114.Sahni S, et al. (2020) NDRG1 suppresses basal and hypoxia-induced autophagy at both the initiation and degradation stages and sensitizes pancreatic cancer cells to lysosomal membrane permeabilization. Biochim Biophys Acta (BBA) 1864;8:129625. doi: 10.1016/j.bbagen.2020.129625. [DOI] [PubMed] [Google Scholar]
- 115.Zhao C, et al. Prodigiosin impairs autophagosome-lysosome fusion that sensitizes colorectal cancer cells to 5-fluorouracil-induced cell death. Cancer Lett. 2020;481:15–23. doi: 10.1016/j.canlet.2020.03.010. [DOI] [PubMed] [Google Scholar]
- 116.Shojaei S, et al. Simvastatin increases temozolomide-induced cell death by targeting the fusion of autophagosomes and lysosomes. FEBS J. 2020;287(5):1005–1034. doi: 10.1111/febs.15069. [DOI] [PubMed] [Google Scholar]
- 117.Hernández-Cáceres MP, et al. Palmitic acid reduces the autophagic flux in hypothalamic neurons by impairing autophagosome-lysosome fusion and endolysosomal dynamics. Mol Cell Oncol. 2020;7(5):1789418. doi: 10.1080/23723556.2020.1789418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Wyant GA, et al. mTORC1 Activator SLC38A9 is required to efflux essential amino acids from lysosomes and use protein as a nutrient. Cell. 2017;171(3):642–654.e12. doi: 10.1016/j.cell.2017.09.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Finkbeiner S. The autophagy lysosomal pathway and neurodegeneration. Cold Spring Harb Perspect Biol. 2020;12(3):a033993. doi: 10.1101/cshperspect.a033993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Long Z, et al. Dynamic changes of autophagic flux induced by Abeta in the brain of postmortem Alzheimer's disease patients, animal models and cell models. Aging. 2020;12(11):10912–10930. doi: 10.18632/aging.103305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Funderburk SF, Marcellino BK, Yue Z. Cell "self-eating" (autophagy) mechanism in Alzheimer's disease. Mount Sinai J Med. 2010;77(1):59–68. doi: 10.1002/msj.20161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Qiu K, et al. BACE2 degradation is mediated by both the proteasome and lysosome pathways. BMC Mol Cell Biol. 2020;21(1):13. doi: 10.1186/s12860-020-00260-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Senkevich K, Gan-Or Z. Autophagy lysosomal pathway dysfunction in Parkinson's disease; evidence from human genetics. Parkinsonism Relat Disord. 2020;73:60–71. doi: 10.1016/j.parkreldis.2019.11.015. [DOI] [PubMed] [Google Scholar]
- 124.Meena NK, Raben N. Pompe disease: new developments in an old lysosomal storage disorder. Biomolecules. 2020;10(9):1339. doi: 10.3390/biom10091339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Kajihara R, et al. Novel drug candidates improve ganglioside accumulation and neural dysfunction in GM1 gangliosidosis models with autophagy activation. Stem Cell Rep. 2020;14(5):909–923. doi: 10.1016/j.stemcr.2020.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Lieberman AP, et al. Autophagy in lysosomal storage disorders. Autophagy. 2012;8(5):719–730. doi: 10.4161/auto.19469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Palhegyi AM, et al. Biomedical implications of autophagy in macromolecule storage disorders. Front Cell Dev Biol. 2019;7:179. doi: 10.3389/fcell.2019.00179. [DOI] [PMC free article] [PubMed] [Google Scholar]