Abstract
Objective:
To analyze the high expression of peroxisome membrane protein 4 (PXMP4) in hepatocellular carcinoma (HCC) and its clinical significance.
Methods:
The expression of PXMP4 mRNA in HCC tissues and corresponding adjacent tissues was detected by Q-PCR, and the expression of PXMP4 protein was detected by Western blot and immunohistochemistry. The correlation of PXMP4 protein expression with clinicopathological features and prognosis of HCC was analyzed.
Results:
The expression levels of PXMP4 mRNA and protein in HCC tissues were significantly higher than those in adjacent tissues (P < 0.05), and its high expression was significantly correlated with tumor differentiation, lymph node metastasis, depth of invasion and TNM stage (P < 0.05). Patients with high expression of PXMP4 had a poor prognosis (P < 0.05).
Conclusion:
The high expression of PXMP4 may promote the occurrence and development of HCC, and inhibition of PXMP4 may be one of the potential molecular targets for targeted therapy of HCC.
Key Words: Hepatocellular Carcinoma, PXMP4, prognosis
Introduction
The etiology of liver cancer is complex, and it is related to the long-term interaction of clinical genetics, viral hepatitis, heavy drinking and other factors [1]. Liver is an important organ of human metabolism, and its blood supply is extremely rich. Once the disease occurs, it will not only have a serious impact on the normal metabolism of the body, but also the tumor cells are prone to metastasis, resulting in organ dysfunction and reducing the overall survival rate [2]. China has a high prevalence of liver cancer [3]. At present, the radical treatment of HCC is still mainly based on surgery. Clinically, most of the patients with liver cancer are treated with comprehensive plans based on surgery, which can remove tumor cells in the body to the maximum extent and avoid postoperative recurrence and metastasis [4]. However, long-term follow-up showed that there were still some patients with poor prognosis. Most of the patients were diagnosed in the middle and late stages, and lost the chance of radical treatment. Other commonly used treatments, such as radiofrequency ablation, transarterial chemoembolization, and chemoradiotherapy, can benefit patients to a certain extent, but the long-term prognosis is still unsatisfactory [5, 3].
Peroxisomes are single-membrane-bound organelles involved in a variety of metabolic pathways [6]. PXMP4 is a peroxisome membrane protein that is widely expressed in cells and is one of the main components of peroxisome membrane with a molecular weight of 24kDa. PXMP4 expression is silenced in prostate cancer mainly due to intronic CpG dinucleotide mediated DNA methylation, while PXMP4 expression is inversely correlated with CpG island methylation value in non-small cell lung cancer (NSCLC). Moreover, PXMP4 is negatively correlated with proliferation marker Ki-67 protein in NSCLC [7] Our research group found that PXMP4 can promote the proliferation, invasion and migration of colorectal cancer cells [8]. It can be inferred that PXMP4 plays an important role in the occurrence and development of tumors, and also shows that PXMP4 has very important clinical significance in the diagnosis and prognosis of malignant tumors. However, the role of PXMP4 in hepatocellular carcinoma has not been reported [9].
The objective of this study was to analyze the expression of PXMP4 in HCC and its prognostic significance.
Materials and Methods
Materials
Case report
A total of 50 HCC and corresponding paracancerous tissues from patients who underwent surgical treatment in Henan Cancer Hospital from January 2022 to December 2022 were collected. Consent was obtained from the ethics committee of the hospital and patients were informed. There were 31 males and 19 females. 20 cases were less than 60 years old and 30 cases were more than 60 years old. Well differentiated, moderately differentiated and poorly differentiated cases were 13, 27 and 10, respectively. There were 13 cases with lymph node metastasis and 37 cases without lymph node metastasis. There were 19 cases with tumor diameter ≤5cm and 31 cases with tumor diameter >5cm. The surgically removed tissues were isolated and fixed in formalin or cryopreserved.
Reagent
PXMP4 antibody (Novus Biologicals, USA), PVDF membrane (Millipore, USA), pre-dye Maker (Thermo, USA), PXMP4 primer and β-Actin primer (Jierai Biological, Shanghai), Anti-gapdh rabbit monoclonal antibody, RIPA lysate, BCA protein quantification kit, ECL color development kit, Trizol, reverse transcription kit, UltraSYBR Mixture and rabbit/mouse Universal SP kit (DAB) (Kangyi).
Methods
The expression of PXMP4 mRNA was detected by quantitative real-time PCR
The samples of hepatocellular carcinoma and adjacent tissues frozen in liquid nitrogen were ground into powder and RNA was extracted by Trizol method [10]. Three μg of RNA was used for reverse transcription to synthesize cDNA. The sequence of human LSD1 specific primer was designed by using pubmed database. The sequence of human LSD1 specific primer was as follows: forward primer 5’-ATAACATCAACAGCCAGAT-3’, reverse primer R:5’-GGATGTAGCCCTTCTCTA-3’. The reference human β-actin-specific primer sequence was as follows: forward primer 5 ‘-TCGTCATACTCCTGCTTGCTG-3’ and reverse primer 5 ‘-CGGGACCTGACTGACTACCTCA-3’. 2 μL of cDNA synthesized by reverse transcription was used for Q-PCR, and the annealing temperature of both PXMP4 and β-Actin was 60℃. After 40 cycles of amplification, 2-ΔΔCt was calculated, and the value indicates the expression level of PXMP4 mRNA.
The expression of LSD1 protein was detected by Western blot
The above frozen tissues were ground into powder, and an appropriate amount of protein lysate was added to fully dissociate. After shaking and centrifugation, the supernatant was taken as the required protein extract, and BCA kit was used for protein quantification. Protein electrophoresis and membrane transfer were carried out on 12% SDS-polypropyl amide gel. The PVDF membrane was blocked with skim milk powder for 1h, and PXMP4 rabbit polyclonal antibody (1:1000) and the internal reference GAPDH anti-rabbit polyclonal antibody (1:2000) were incubated at 4℃ overnight, and the goat anti-rabbit secondary antibody (1:2000) was incubated at room temperature for 2h. The PVDF membrane was treated with ECL and the target protein bands were detected by gel imaging system, and the gray value of protein bands was detected by Image J software [11, 12]. The expression level of the target protein was expressed as the ratio of gray value of the target protein band to the gray value of the internal reference protein band.
The expression of PXMP4 protein was detected by immunohistochemistry
The fixed tissue was embedded in paraffin, sectioned, dissolved in wax at 60℃ for 1 h, deparaffinized with xylene, hydrated with gradient ethanol, and repaired with citric acid antigen. The LSD1 protein was detected by immunohistochemical SP high-sensitivity kit. The primary anti-PXMP4 rabbit polyclonal antibody (1:200) was incubated at room temperature. DAB was used for color development, followed by sulfoxide counterstained, dehydrated with gradient ethanol, transparent with xylene and sealed with neutral gum [13]. The PXMP4-positive cells showed brownish yellow granules in the nucleus. A total of 200 cells (400×) were randomly selected for observation under 4 high-power fields, and the staining results were judged by double-blind method. The immunohistochemical scores were analyzed by semi-quantitative method [9]. Immunohistochemical score = staining intensity score × positive cell count score. Staining intensity score: 0, no staining, negative; 1 point, light yellow, weakly positive; 2 points, yellow or dark yellow, moderately positive; 3 points, yellowish brown or dark brown, strongly positive; Positive cell count score: 0, no positive cells; 1 point, positive cell count ≤25%; 2 points, positive cell count > 25% and ≤50%; 3 points: positive cell count > 50%. The mean of four visual field immune scores was taken: 0-2 was considered negative; > 2 was defined as positive.
Statistical Analysis
SPSS21.0 was used to analyze the data. The difference between Q-PCR and Western blot quantitative results was analyzed by paired t test. Immunohistochemistry was used to detect the difference of PXMP4 protein expression in HCC and corresponding paracancerous tissues. Paired Chi-square test was used to analyze the difference of PXMP4 protein expression in HCC and corresponding paracancerous tissues. The correlation between PXMP4 protein expression and clinicopathological features of HCC was analyzed by chi-square test. Fifty patients with hepatocellular carcinoma were followed up for 30 months after surgery. Survival time was defined as the date of operation to the last follow-up date or death. Uncensored data were patients who died due to hepatocellular carcinoma, and censored data were patients who were still alive after 30 months of follow-up or patients died of other causes. The survival curve was drawn by Kaplan-Meier method and the difference of survival time was tested by Log-rank test. All measurement data were expressed as (x ®± s).
Results
Expression of PXMP4 in hepatocellular carcinoma and adjacent tissues
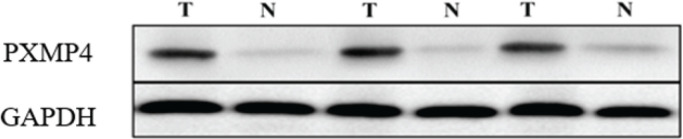
The quantitative detection of PXMP4 mRNA and protein expression levels by Q-PCR and Western blot (Figure 1 and Table 1) showed that the expression levels of PXMP4 mrna and protein in HCC tissues were significantly higher than those in the corresponding adjacent tissues (P <0.001).
Figure 1.
. Detect the PXMP4 Protein Expression in Hepatocellular Carcinoma and Its Adjacent Tissues by Western Blot
Table 1.
Detect the PXMP4 Expression in Hepatocellular Carcinoma and Its Adjacent
| Groups | PXMP4 mRNA |
PXMP4
protein |
|---|---|---|
| hepatocellular carcinoma | 3.25 ± 1.42 | 0.73 ± 0.26 |
| Adjacent cancer | 1.79 ± 1.07 | 0.30 ± 0.16 |
| t | 5.39 | 8.13 |
| P | <0.001 | <0.001 |
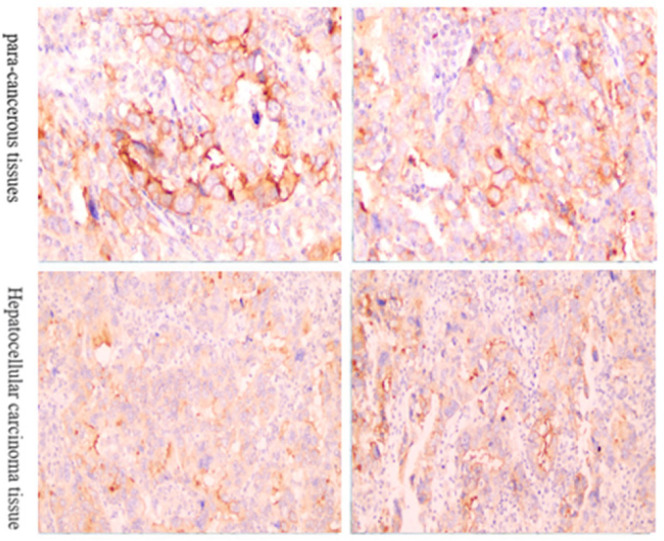
The immunohistochemical test results are shown in Figure 2 and Table 2. The expression of PXMP4 protein was mainly located in the nucleus of the tumor cells, which was brownish yellow granules, and some of the cytoplasm was also positive, which was light yellow. The expression was mainly negative in paracancerous tissues, while strongly positive in HCC tissues. The positive expression rate of PXMP4 in hepatocellular carcinoma tissues (64.00%) (32/50) was significantly higher than that in the corresponding adjacent tissues (18.00%) (9/50), and the difference was statistically significant (P <0.05).
Figure 2.
Detect the Expression of PXMP4 in Rectal Adenocarcinoma by Immunohistochemistry
Table 2.
Expression of PXMP4 Protein in Hepatocellular Carcinoma and Its Adjacent Tissues
| Adjacent cancer | hepatocellular carcinoma | |
|---|---|---|
| PXMP4 positive | PXMP4 negative | |
| PXMP4 positive | 20 | 0 |
| PXMP4 negative | 10 | 20 |
Correlation between PXMP4 protein expression level and clinicopathological features of hepatocellular carcinoma
Chi-square test was used to analyze the correlation between PXMP4 protein expression and clinicopathological features of HCC (Table 3), and it was found that PXMP4 protein expression was significantly correlated with gender, degree of tumor differentiation, and lymph node metastasis. The positive rate of PXMP4 protein expression in male HCC tissues (77.4%) was significantly higher than that in female HCC tissuest (42%, χ2=6.376, P=0.012). The positive rate of PXMP4 protein expression in poorly differentiated group (100%) was significantly higher than that in well differentiated group (53.8%) and moderately differentiated group (55.5%, χ2 =7.042, P=0.03). The positive rate of PXMP4 in the group with lymph node metastasis (92.3%) was significantly higher than that in the group without lymph node metastasis (54%, χ2=6.11, P=0.033). The expression of PXMP4 protein in HCC tissues was not significantly correlated with age (χ2=0.014, P=0.904) and tumor diameter (χ2=1.719, P=0.19).
Table 3.
Correlation between PXMP4 Protein Expression and Clinicopathological Features of Hepatocellular Carcinoma
| Pathological factors | n | PXMP4 positive | PXMP4 negative | P | |
|---|---|---|---|---|---|
| Hepatocellular carcinoma tissue | 50 | 32(64.0%) | 18(34.0%) | ||
| Adjacent to cancer tissue | 50 | 9(18.0%) | 41(82.0%) | 14.297 | 0 |
| Gender | |||||
| Male | 31 | 24(77.4%) | 7(22.6%) | ||
| Women | 19 | 8(42.0%) | 11(58.0%) | 6.376 | 0.012 |
| Age (years) | |||||
| ≥60 | 30 | 19(63.3%) | 11(36.7%) | ||
| <60 | 20 | 13(65.0%) | 7(35.0%) | 0.014 | 0.904 |
| Degree of differentiation | |||||
| High degree of differentiation | 13 | 7(53.8%) | 6(46.2%) | ||
| Intermediate differentiation | 27 | 15(55.5%) | 12(44.5%) | ||
| Low differentiation | 10 | 10(100.0%) | 0(0.0%) | 7.042 | 0.03 |
| Lymph node metastasis | |||||
| Positive | 13 | 12(92.3%) | 1(7.7%) | ||
| Negative | 37 | 20(54.0%) | 17(46.0%) | 6.11 | 0.033 |
| Diameter of tumor | |||||
| ≤5 | 19 | 10(52.6%) | 9(47.4%) | ||
| >5 | 31 | 22(71.0%) | 9(29.0%) | 1.719 | 0.19 |
Correlation between PXMP4 protein expression and prognosis of patients with hepatocellular carcinoma
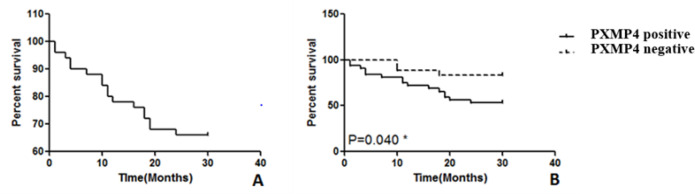
The overall survival rate of 50 patients with HCC at 30 months after surgery was 66.00%. The 30-month survival rate of the PXMP4 positive expression group (56.3%) was significantly lower than that of the negative group (83.3%, Log-rank test, χ2=4.202, P=0.040, Figure 3).
Figure 3.
Survival Curve of Kaplan-Meier in Patients with Hepatocellular Carcinoma
Discussion
As a kind of organelle, peroxisome exists in the nucleus, which affects the metabolic process of cells and then affects the biological function. However, peroxisome can affect different metabolic methods in cells or tissues due to different locations, and can be changed by changes in cellular regulation or physiological conditions [14, 15]. PXMP4 is a ubiquitously expressed peroxisomal membrane protein that is transcriptionally regulated by peroxisome proliferator-activated receptor α, although its function remains unclear [16]. The PXMP4 gene is located at 20q11.22 and is conserved in a variety of animals such as chimpanzees, dogs, cattle, and mice. It encodes an integral membrane protein associated with peroxisomes. Its only known binding partner is ex19, an intracellular chaperone and part of the peroxisome membrane insertion machinery. It is encoded by Nkt cell control gene Nkt1 [17-19]. Studies have shown that PXMP4 is closely related to the repair process of oocytes [20, 18]. Moreover, previous studies have found that the expression of PXMP4 has clinical significance in prostate cancer and non-small cell lung cancer [21], but the study of PXMP4 in hepatocellular carcinoma has not been reported.
The results of this study showed that the expression of PXMP4 mRNA and protein in HCC tissues was significantly higher than that in adjacent tissues. PXMP4 was mainly located in the nucleus of hepatocellular carcinoma cells. Its high expression was significantly correlated with the degree of differentiation, lymph node metastasis, depth of invasion and TNM stage of hepatocellular carcinoma, and was negatively correlated with the survival time of patients. The study found that the expression of PXMP4 protein and mRNA in colon cancer cells was higher than that in the control group, and the difference was statistically significant. PXMP4 was closely related to the malignant degree and metastatic potential of colon cancer [22], which was consistent with the results in this paper.
There are still some shortcomings in this study. For example, the study on the pathogenesis of colorectal cancer is limited to in vitro experiments on related genes, and the support of animal models is lacking. Since there are many aspects involved in the development of tumors, and in terms of genes, multiple genes contribute to the development of tumors, how do these genes interact with each other? Synergistic or antagonistic? And the related molecules or signaling pathways of genes in promoting tumor proliferation all point out the direction for future research.
In conclusion, the expression of PXMP4 is significantly increased in HCC, suggesting that PXMP4 may be closely related to the occurrence and development of HCC, and inhibition of PXMP4 may be one of the potential molecular targets for targeted therapy of HCC.
Author Contribution Statement
MingMing Zhao and Farra Aidah Binti Jumuddin designed and carried out the experiments. Zarina Awang and MingMing Zhao analyzed the data. MingMing Zhao wrote the manuscript. All authorscontributed to the article and approved the submitted version.
Acknowledgements
Ethical Clearance
This study was approved by the Ethics Committee of the Henan Cancer Hospital.
Data Availability statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author.
Conflict of Interest
The authors declare no conflict of interest.
References
- 1.Akinyemiju T, Abera S, Ahmed M, Alam N, Alemayohu MA, Allen C, et al. The burden of primary liver cancer and underlying etiologies from 1990 to 2015 at the global, regional, and national level: Results from the global burden of disease study 2015. JAMA Oncol. 2017;3(12):1683–91. doi: 10.1001/jamaoncol.2017.3055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin. 2018;68(6):394–424. doi: 10.3322/caac.21492. [DOI] [PubMed] [Google Scholar]
- 3.Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin. 2021;71(3):209–49. doi: 10.3322/caac.21660. [DOI] [PubMed] [Google Scholar]
- 4.Llovet JM, Montal R, Sia D, Finn RS. Molecular therapies and precision medicine for hepatocellular carcinoma. Nat Rev Clin Oncol. 2018;15(10):599–616. doi: 10.1038/s41571-018-0073-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wang D, Li C, Xu Y, Xing Y, Qu L, Guo Y, et al. Clinicopathological characteristics and prognosis of alpha-fetoprotein positive gastric cancer in chinese patients. Int J Clin Exp Pathol. 2015;8(6):6345–55. [PMC free article] [PubMed] [Google Scholar]
- 6.Youssef AA, Issa HA, Omar MZ, Behiry EG, Elfallah AA, Hasaneen A, et al. Serum human endothelial cell-specific molecule-1 (endocan) and vascular endothelial growth factor in cirrhotic hcv patients with hepatocellular carcinoma as predictors of mortality. Clin Exp Gastroenterol. 2018;11:431–8. doi: 10.2147/CEG.S171339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Park SJ, Jang JY, Jeong SW, Cho YK, Lee SH, Kim SG, et al. Usefulness of afp, afp-l3, and pivka-ii, and their combinations in diagnosing hepatocellular carcinoma. Medicine (Baltimore) 2017;96(11):e5811. doi: 10.1097/MD.0000000000005811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Wei T, Zhang W, Tan Q, Cui X, Dai Z. Electrochemical assay of the alpha fetoprotein-l3 isoform ratio to improve the diagnostic accuracy of hepatocellular carcinoma. Anal Chem. 2018;90(21):13051–8. doi: 10.1021/acs.analchem.8b04045. [DOI] [PubMed] [Google Scholar]
- 9.Shen YL, Li HZ, Hu YW, Zheng L, Wang Q. Loss of gins2 inhibits cell proliferation and tumorigenesis in human gliomas. CNS Neurosci Ther. 2019;25(2):273–87. doi: 10.1111/cns.13064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Rio DC, Ares M Jr, Hannon GJ, Nilsen TW. Purification of rna using trizol (tri reagent) Cold Spring Harb Protoc. 2010;2010(6):pdb.prot5439. doi: 10.1101/pdb.prot5439. [DOI] [PubMed] [Google Scholar]
- 11.Hirano S. Western blot analysis. Methods Mol Biol. 2012;926:87–97. doi: 10.1007/978-1-62703-002-1_6. [DOI] [PubMed] [Google Scholar]
- 12.Gallo-Oller G, Ordoñez R, Dotor J. A new background subtraction method for western blot densitometry band quantification through image analysis software. J Immunol Methods. 2018;457:1–5. doi: 10.1016/j.jim.2018.03.004. [DOI] [PubMed] [Google Scholar]
- 13.Ramos-Vara JA. Technical aspects of immunohistochemistry. Vet Pathol. 2005;42(4):405–26. doi: 10.1354/vp.42-4-405. [DOI] [PubMed] [Google Scholar]
- 14.Ye Y, Song YN, He SF, Zhuang JH, Wang GY, Xia W. Gins2 promotes cell proliferation and inhibits cell apoptosis in thyroid cancer by regulating cited2 and loxl2. Cancer Gene Ther. 2019;26(3-4):103–13. doi: 10.1038/s41417-018-0045-y. [DOI] [PubMed] [Google Scholar]
- 15.Yan T, Liang W, Jiang E, Ye A, Wu Q, Xi M. Gins2 regulates cell proliferation and apoptosis in human epithelial ovarian cancer. Oncol Lett. 2018;16(2):2591–8. doi: 10.3892/ol.2018.8944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zheng M, Zhou Y, Yang X, Tang J, Wei D, Zhang Y, et al. High gins2 transcript level predicts poor prognosis and correlates with high histological grade and endocrine therapy resistance through mammary cancer stem cells in breast cancer patients. Breast Cancer Res Treat. 2014;148(2):423–36. doi: 10.1007/s10549-014-3172-7. [DOI] [PubMed] [Google Scholar]
- 17.Peng L, Song Z, Chen D, Linghu R, Wang Y, Zhang X, et al. Gins2 regulates matrix metallopeptidase 9 expression and cancer stem cell property in human triple negative breast cancer. Biomed Pharmacother. 2016;84:1568–74. doi: 10.1016/j.biopha.2016.10.032. [DOI] [PubMed] [Google Scholar]
- 18.Waterham HR, Ferdinandusse S, Wanders RJ. Human disorders of peroxisome metabolism and biogenesis. Biochim Biophys Acta. 2016;1863(5):922–33. doi: 10.1016/j.bbamcr.2015.11.015. [DOI] [PubMed] [Google Scholar]
- 19.Blankestijn M, Bloks VW, Struik D, Huijkman N, Kloosterhuis N, Wolters JC, et al. Mice with a deficiency in peroxisomal membrane protein 4 (pxmp4) display mild changes in hepatic lipid metabolism. Sci Rep. 2022;12(1):2512 . doi: 10.1038/s41598-022-06479-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Farré JC, Mahalingam SS, Proietto M, Subramani S. Peroxisome biogenesis, membrane contact sites, and quality control. EMBO Rep. 2019;20(1) doi: 10.15252/embr.201846864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhang X, Yang H, Zhang J, Gao F, Dai L. Hsd17b4, acaa1, and pxmp4 in peroxisome pathway are down-regulated and have clinical significance in non-small cell lung cancer. Front Genet. 2020;11:273. doi: 10.3389/fgene.2020.00273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Fletcher JM, Jordan MA, Snelgrove SL, Slattery RM, Dufour FD, Kyparissoudis K, et al. Congenic analysis of the nkt cell control gene nkt2 implicates the peroxisomal protein pxmp4. J Immunol. 2008;181(5):3400–12. doi: 10.4049/jimmunol.181.5.3400. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author.