Abstract
Satellite RNA of bamboo mosaic potexvirus (satBaMV), a single-stranded positive-sense RNA encoding a nonstructural protein of 20 kDa (P20), depends on bamboo mosaic potexvirus (BaMV) for replication and encapsidation. A full-length cDNA clone of satBaMV was used to examine the sequences required for the synthesis of potexvirus subgenomic RNAs (sgRNAs). Subgenomic promoter-like sequences (SGPs), 107 nucleotides (nt) upstream from the capsid protein (CP) gene of BaMV-V, were inserted upstream of the start codon of the P20 gene of satBaMV. Insertion of SGPs gave rise to the synthesis of sgRNA of satBaMV in protoplasts of Nicotiana benthamiana and leaves of Chenopodium quinoa when coinoculated with BaMV-V genomic RNA. Moreover, both the satBaMV cassette and its sgRNA were encapsidated. From analysis of the SGPs by deletion mutation, we concluded that an SGP contains one core promoterlike sequence (nt −30 through +16), two upstream enhancers (nt −59 through −31 and −91 through −60), and one downstream enhancer (nt +17 through +52), when the transcription initiation site is taken as +1. Site-directed mutagenesis and compensatory mutation to disrupt and restore potential base pairing in the core promoter-like sequence suggest that the stem-loop structure is important for the function of SGP in vivo. Likewise, the insertion of a putative SGP of the BaMV open reading frame 2 gene or a heterologous SGP of potato virus X resulted in generation of an sgRNA. The satBaMV cassette should be a useful tool to gain insight into sequences required for the synthesis of potexvirus sgRNAs.
The production of subgenomic RNAs (sgRNAs) is one of the strategies by which internally located open reading frames (ORFs) of multicistronic RNA virus may be expressed and regulated during replication. The capsid proteins (CPs) of alpha-like superfamily viruses are expressed by sgRNA transcription (24). The proteins encoded by the triple gene block of potex-, carla-, furo-, and hordeiviruses and the movement proteins of tobamo- and tombusviruses are also expressed by the same strategy (24). Several mechanisms are proposed for the transcription of sgRNAs. The first is leader-primed transcription, which occurs in the production of coronavirus sgRNAs. This process involves fusion of noncontinuous sequences from the 5′ leader sequences to the sgRNA by a discontinuous process (14). Recently, a new model for coronavirus transcription was proposed. This model predicts that subgenome-length negative strands containing the common 5′ leader sequence were directly derived from genome RNA and served as a template for the production of sgRNA (27). The second mechanism proposed for transcription of sgRNAs is exemplified by transcription of RNA-mediated trans-activation. A 34-nucleotide (nt) trans-activator on red clover necrotic mosaic virus RNA2 was proposed to bind to the element upstream of the sgRNA initiation site on RNA1 and to prevent the synthesis of full-length minus-strand RNA1. This truncated minus-strand RNA1 may serve as a template for viral RNA-dependent RNA polymerase (RdRp) to produce the sgRNA (31). Another mechanism proposed is suitable for most plant viruses, as exemplified by brome mosaic virus (BMV). SgRNA transcription is directed by subgenomic promoter-like sequences (SGPs), located 100 to 200 nt upstream of the start codon of ORFs in the minus-strand template, and recognized by the viral RdRp (24). In some cases, additional long-distance RNA-RNA interactions between the SGP sequence and the 5′ terminus are required for the genomic and sgRNA accumulation (13, 41).
The SGPs of BMV (8, 25, 32) and alfalfa mosaic virus (AMV) (33, 34) have been extensively characterized in vivo and in vitro. All of these SGPs contain a “core promoter,” which is the minimum number of sequences required for the synthesis of the sgRNA, and several “enhancers,” which modulate the function of the core promoter. In addition, RdRp complexes of BMV (8) and AMV (34) have been shown to bind initially to two internal sites on the minus strand of RNA3 for sgRNA synthesis. Recently, sgRNA initiation has been studied on the small template of minus-strand BMV RNA3, using purified viral RdRp complexes, which suggests that recognition of the SGP by the BMV RdRp is sequence specific (1, 30). However, the secondary structure of the SGP may also facilitate efficient sgRNA transcription in vivo (1).
For the potexvirus group, two major sgRNAs of about 1.9 to 2.1 kb and 0.9 to 1.0 kb in size, which direct the expression of the ORF2 protein and the CP, respectively, were detected in the infected cells (7, 11, 15, 37). Another bicistronic sgRNA of 1.4 kb, which was responsible for the translation of ORF3 and ORF4 proteins, accumulated at a very low level in the infected cells (36). Alignment of the putative SGPs of ORF2 protein and CP among potexviruses revealed that the conserved octanucleotide motif (3′-CAAUUCAA-5′) was located upstream from the transcription initiation sites of the sgRNAs (12, 15, 37). However, the nucleotide sequences and the spacing between the transcription initiation site and the octanucleotide motif vary among potexviruses (12, 15). It has been shown that complementarity between the conserved octanucleotide sequence and the 3′ terminus of minus-strand potato virus X (PVX) RNA is important for the synthesis of sgRNA (13). Mutations that reduced the complementarity affected the viral RNA replication and accumulation of sgRNA (12, 13).
Bamboo mosaic potexvirus (BaMV) contains a single-stranded positive-sense RNA genome of 6.4 kb and belongs to the alphavirus-like superfamily (21, 40). The genome of BaMV contains five common ORFs (21, 40). ORF1 encodes a putative RdRp of 155 kDa (19, 21). ORF2 to ORF4, which code for triple gene block proteins of 28, 13, and 6 kDa (21, 40), respectively, are required for viral cell-to-cell movement (3, 39). The product of ORF5 is the CP of 25 kDa (15). A satellite RNA (satBaMV) was found to be associated with the BaMV-V isolate (22). The satBaMV is a linear molecule of 836 nt and encodes a 20-kDa protein (P20) (22). The ORF of P20 is not essential for satBaMV replication and can be replaced with a chloramphenicol acetyltransferase (CAT) reporter gene to express the CAT protein in coinoculated plants (23).
In this study, we used the satBaMV cassette to examine the sequences and structures required for the syntheses of potexvirus sgRNAs. The insertion of the putative SGP of the 1.0-kb sgRNA of BaMV into the satBaMV cassette resulted in a new abundant sgRNA of satBaMV. By analyzing the SGP activities of various mutants, the core promoter-like and enhancer-like sequences of the 1.0-kb sgRNA were mapped. The generation of an sgRNA of satBaMV was also confirmed by the insertion of the putative SGP of the 2.0-kb sgRNA or by a heterologous SGP of PVX by primer extension. The satBaMV cassette provides a useful tool for the analysis of sequences or structures required for the synthesis of potexvirus sgRNAs.
MATERIALS AND METHODS
Plasmid construction.
Plasmid pBSF4 is a full-length cDNA clone of satBaMV with a T7 RNA promoter at the 5′ end as previously described (23). pICF4 was constructed by insertion of 107 nt upstream of the start codon of the CP gene of BaMV-V into the start codon of the P20 gene of pBSF4 (Fig. 1A). The DNA fragments corresponding to the putative SGP of the CP gene were amplified from pBa19 (40) by PCR using primers B43 (5′-TATCCACGACGTTGGAAATAATAATAAAC) and B44 (5′-CGCCCATCGTCCTGGCTTTAGTGTTTAATT). After digestion with BstXI, the fragments were inserted into BstXI-cut pBSF4 to generate pICF4.
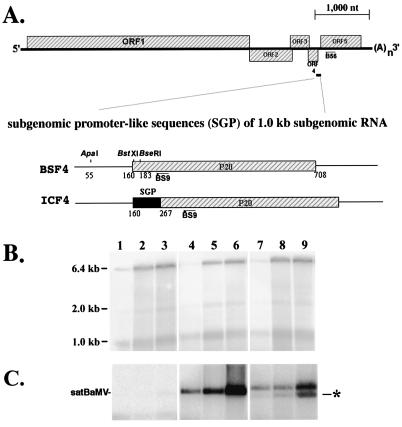
FIG. 1.
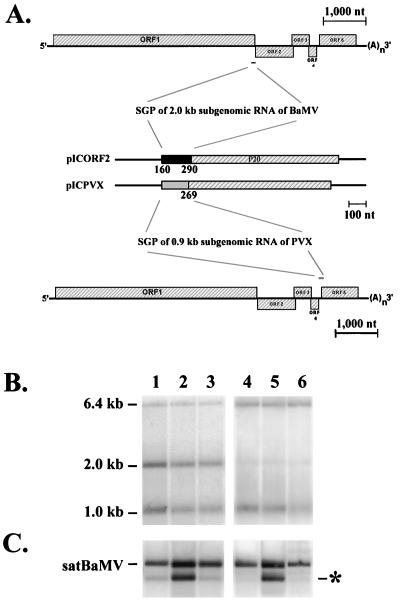
Genome organization of BaMV and satBaMV mutants (A) and the time course accumulation of BaMV-V genomic RNA (B) and satBaMV mutants (C) in inoculated protoplasts. (A) In these schematic diagrams of BaMV and satBaMV cassettes, the hatched regions represent the ORF regions of BaMV (designated ORFs 1 through 5) or satBaMV (designated P20); the black bar represents the 107 nt upstream of the CP gene of BaMV (designated SGP); restriction enzyme sites ApaI, BstXI, and BseRI in pBSF4 are also indicated as are the positions of primers B56 and BS9. (B and C) Protoplasts of barley were inoculated with BaMV-L RNA only (lanes 1 through 3) or coinoculated with pBSF4 (lanes 4 through 6) or pICF4 (lanes 7 through 9) transcripts. Total RNAs extracted from 5 × 104 protoplasts at 8 h.p.i. (lanes 1, 4, and 7), 16 h.p.i. (lanes 2, 5, and 8), and 24 h.p.i. (lanes 3, 6, and 9) were glyoxylated and fractionated in a 1% agarose gel, blotted to a nylon membrane, and hybridized to a BaMV-specific (20) (B) or satBaMV-specific (23) (C) riboprobe. The star indicates the position of the 0.7-kb sgRNA transcribed from pICF4.
Plasmids pICN9, pICN18, pICN36, and pICN54 are modified forms of pICF4 with the addition of the N-terminal 9, 18, 36, and 54 nt of the CP gene (40), respectively (Fig. 2A). They were constructed by site-directed mutagenesis using a double-PCR method (4). For the construction of pICN18, the first PCR fragment was synthesized from the pICF4 template by primers IC2 (5′-AATTAAACACTAAAGATGTCTGGAACTGGAACGATGGTTCGGAGGAGA) and BS26 (5′-CCTTCTCGAGTCAACTGGTTGGCGCACG) followed by the second PCR using pICF4 as a template with primer BS19 (5′-TGCCTGCAGTAATACGACTCACTATAGAAAACTCACCGCAACGA) and the first PCR fragments. The ApaI- and BseRI-digested secondary PCR fragments were ligated with the ApaI- and BseRI-cut pICF4 to generate pICN18. To generate pICN9, pICN36, and pICN54, a similar method was used except that the pICN18 was used as a template and the corresponding primers BSIC23 (5′-CCTCCGAACCATTCCAGACATCTTTAG), BSIC22 (5′-CCTCCGAACCATAGTCCCTCGCCCAGTTCCTGTTCCAGTTCCAG) and BSIC21 (5′-GACCTTGACCTTGTC CTTGACCCGCTCGACCTTGACCACGCCCTCCGTACCAAGCCTCC) were used instead, respectively.
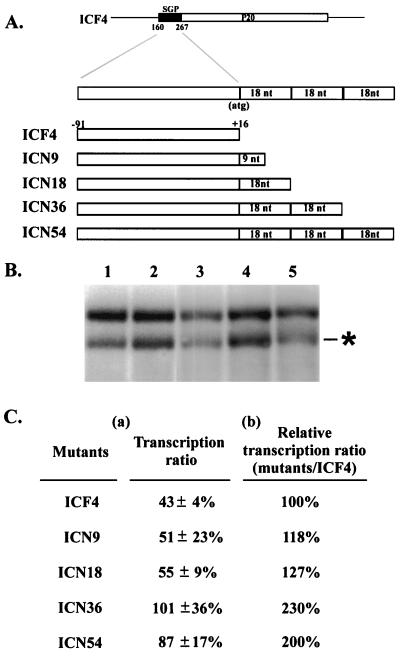
FIG. 2.
Schematic diagrams of pICF4 mutants (A) and their transcription activities (B and C). (A) pICF4 and its mutants carrying various lengths of SGP sequences extended downstream to the coding region of the CP gene. The black region represents the 107 nt upstream of the CP gene of BaMV (designated SGP). The start codon of the CP gene is indicated. (B) Northern blot analysis was performed on total RNAs extracted from 5 × 104 protoplasts of N. benthamiana coinoculated with BaMV-L RNA and pICF4 (lane 1), pICN9 (lane 2), pICN18 (lane 3), pICN36 (lane 4), and pICN54 (lane 5) transcripts at 24 h.p.i. with a satBaMV-specific riboprobe. The asterisk indicates the position of the 0.7-kb sgRNA transcribed from satBaMV cassette. (C) Quantitative analyses of the transcription ratios of pICF4 and its mutants in coinoculated protoplasts of N. benthamiana were performed. The transcription ratio was determined by quantifying the 0.7-kb sgRNA relative to that of satBaMV RNA (a), and the transcription ratio was the transcription ratio of ICF4 mutants over that of ICF4 at 24 h.p.i. (b). These data were collected in three independent experiments by PhosphorImager and using the ImageQuant version 3.3 program (Molecular Dynamics); standard deviations are indicated.
pIC3, pIC4, and pIC5 are the 5′ end deletion mutants of pICN18 deleted from nt −91 to −8, −91 to −31, and −91 to −60, respectively, with the transcription initiation site taken as +1 (Fig. 3B). pIC2 retained only the −30 to +3 sequences of the putative SGP of 1.0-kb sgRNA (Fig. 3B). pIC8-C/C, pIC9-G/G, pIC10-G/C, and pIC14-CCC are the stem-loop 2 (SL2) stem region mutants of pICN18 obtained by replacing the GGG (at positions −9 through 11)/CCC(−24 through −26) with GGC/CCC, GGG/GCC, GGC/GCC and CCC/CCC, respectively (Fig. 4A). pBSIC7 is the SL2 loop region mutant of pICN18 in which the conserved octanucleotide motif 3′-CAATTCAA-5′ is changed to 3′-CCTAAATA-5′ in the minus strand (Fig. 4A). The DNA fragments corresponding to the SGP mutants were amplified from pICN18 by double PCR as described for pICN18, except that the corresponding primers were as follows: BSIC24 (5′-GTGTTTAATTTACAAACACGTCGTGGTAAGCGTC) was used for pIC3; BSIC4 (5′-AAACTTAACAAACCCTAGCCGTCGTGGTAAGCGTC) was used for pIC4; BSIC5 (5′-GGTAGCAGTTGCCTCTCGTCGTGGTAAGCGTC) was used for pIC5; BSIC7 (5′-TAATTTACAAACAAGGGAAATAAATCCAAACCTAGCTGGAG) was used for pIC7; BSIC8 (5′-GTTTAATTTACAAACAAGGCAAACTTAACAAACCCTAG) was used for pIC8-C/C; BSIC9 (5′-CAAGGGAAACTTAACAAAGCCTAGCTGGAGGTAG) was used for pIC9-G/G; and BSIC14 (5′-GTGTTTAATTTACAAACAACCCAAACTTAACAAACCCTAG) was used for pIC14-CCC. To generate pIC2 and pIC10-G/C, the pIC4 and pIC9-G/G plasmids were used as templates, respectively. Their corresponding primers were BSIC18 (5′-CTCCTCCGAACCATGTTTACAAACAAGGGAAAC) for pIC2 and BSIC10 (5′-GTTTAATTTACAAACAAGGCAAACTTAACAAAGCCTAG) for pIC10-G/C.
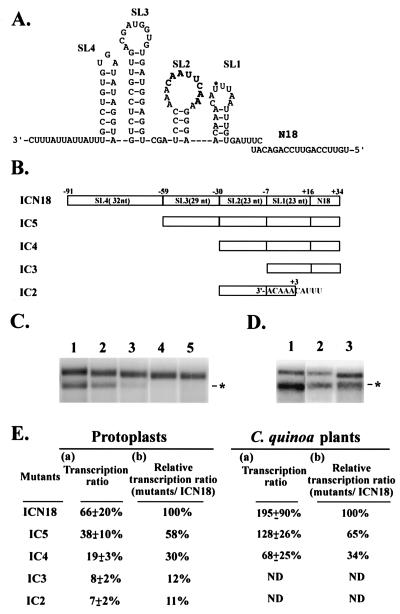
FIG. 3.
The secondary structure of the minus-strand SGP sequence of the 1.0-kb sgRNA of BaMV-V (A), schematic diagrams of satBaMV mutants (B), and their transcriptional-activity assays (C through E). (A) The secondary structure was predicted by the STAR computer program. The stem-loop structures, designated SL1 (ΔG = −1.3 kcal/mol), SL2 (ΔG = −2.5 kcal/mol), SL3 (ΔG = −8.3 kcal/mol), and SL4 (ΔG = −4.8 kcal/mol) and 18 nt of the N-terminal CP gene of BaMV (designated N18) are shown. The asterisk denotes the transcription initiation site, and the conserved octanucleotide motif is in bold. (B) For SatBaMV mutants, the numbers of nucleotides for each structure are given in parentheses. (C through E) For Northern (C and D) and quantitative (E) analyses of the sgRNAs transcribed by the mutants, total RNAs were extracted from 5 × 104 protoplasts of N. benthamiana at 24 h.p.i. (C) or 5-mg leaves of C. quinoa at 6 d.p.i. (D). These RNAs were coinoculated with BaMV-L RNA and the pICN18 (lane 1), pIC5 (lane 2), pIC4 (lane 3), pIC3 (lane 4), or pIC2 (lane 5) transcript and were subjected to Northern analyses with a satBaMV-specific riboprobe. The asterisk indicates the position of the 0.7-kb sgRNA transcribed from satBaMV cassettes. (E) For quantitative analyses of the transcription ratio of satBaMV mutants in protoplasts of N. benthamiana or in leaves of C. quinoa in three independent experiments, the transcription ratio (a) and the relative transcription ratio over that of pICN18 (b) were determined as described in the legend to Fig. 2.
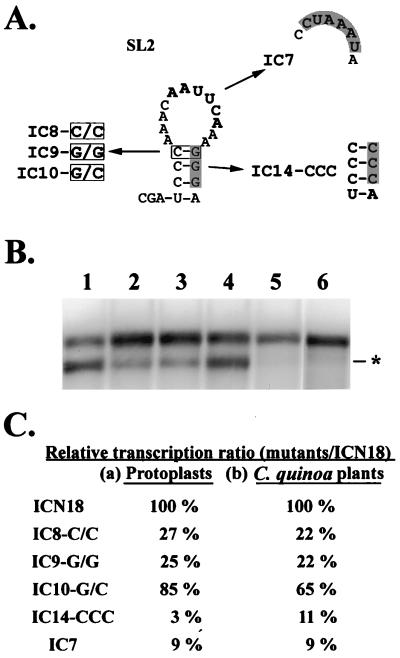
FIG. 4.
Predicted secondary structure of SL2 and the schematic diagrams of satBaMV mutants (A) and their transcription activities (B and C). (A) Shown are the predicted secondary structure of SL2 and the schematic diagrams of mutants introduced in SL2. The boxed and shaded letters represent the mutated nucleotides. The bold letters indicate the conserved nucleotide motif. (B) Northern blot analysis of total RNAs extracted from 5 × 104 protoplasts of N. benthamiana coinoculated with BaMV-L RNA and pICN18 (lane 1), pIC8-C/C (lane 2), pIC9-G/G (lane 3), pIC10-G/C (lane 4), pIC14-CCC (lane 5), and pIC7 (lane 6) transcripts at 24 h.p.i. with a satBaMV-specific riboprobe. The asterisk indicates the position of the 0.7-kb sgRNA transcribed from the satBaMV cassette. (C) The relative ratio was the transcription ratio of a mutant over that of pICN18 in protoplasts of N. benthamiana (a) or in leaves of C. quinoa (b). These data were collected three times from protoplasts and two times from plants.
pICORF2 and pICPVX were constructed by insertion of the putative SGPs of the ORF2 gene of the BaMV-V or of the CP gene of PVX just before the start codon of the P20 gene of pBSF4 (Fig. 5A). The DNA fragments corresponding to the putative SGPs of the ORF2 gene of BaMV-V were amplified from pBa19 (40) using primers BSICORF2-1 (5′-TATCCACGACGTTGGCTTTCCATTACCAAAC) and BSICORF2-2 (5′-CGCCCATCGTCCTGGTTAGTTATTAAACTAA) or for the CP gene of the PVX-Taiwan strain amplified from pPVX-T (Y. H. Hsu, unpublished data) using primers BSICPVX-1 (5′-TATCCACGACGTTGGAATCAATCACAGTG) and BSICPVX-2 (5′-CGCCCATCGTCCTGGCTTTCGAGTATCAATG) by PCR. After digestion with BstXI, the corresponding fragments were inserted into the BstXI site of pBSF4 to generate pICORF2 or pICPVX.
FIG. 5.
SgRNA transcription assays of pICORF2 and pICPVX. (A) Schematic diagrams of ICORF2 and ICPVX. Hatched regions represent the ORF regions of BaMV, PVX, and satBaMV. The black and dotted regions represent the SGP of the 2.0-kb sgRNA of BaMV and of the 0.9-kb sgRNA of PVX, respectively. (B and C) Northern analyses of total RNAs extracted from 5 × 104 protoplasts of N. benthamiana at 24 h.p.i. (lanes 1 through 3) or 5-mg leaves of C. quinoa at 6 d.p.i. (lanes 4 through 6) coinoculated with BaMV-L RNA and pICORF2 (lanes 1 and 4), pICF4 (lanes 2 and 5), and pICPVX (lanes 3 and 6) transcripts with a BaMV (B) or satBaMV (C) riboprobe. The asterisk indicates the position of the 0.7-kb sgRNA transcribed from satBaMV cassettes.
For all the constructs, the DNA sequences flanking the inserted or replaced fragment were confirmed by sequencing.
Virus purification and viral-RNA extraction.
BaMV-L is an isolate derived from BaMV-V and is free of satellite RNA (22, 23). Purification of BaMV virions from infected Chenopodium quinoa and extraction of viral RNA were previously described (17, 19). The purified RNAs were dissolved in sterile distilled water, quantitated by UV absorption, and stored at −150°C until use.
Protoplast and plant inoculation, RNA extraction, and Northern blot analyses.
Preparation and RNA inoculation of protoplasts from Nicotiana benthamiana or barley (Hordeum vulgare L. “Laker”) were as previously described (18). For each inoculation, protoplasts of 2 × 105 cells were inoculated with 1.0 μg of BaMV-L RNA only or coinoculated with 1.0 μg of satBaMV transcripts by the polyethylene glycol method (18). Coinoculation of C. quinoa plants with BaMV-L and satBaMV mutants was also as previously described (22, 23) except that each inoculum contained a mixture of 0.1 μg of BaMV-L RNA and 0.1 μg of satBaMV transcripts per leaf. Total RNA extraction, glyoxylation, and Northern blot analyses were carried out as previously described (23). Extraction of mRNA from total RNAs was according to the instructions of the Straight A's mRNA isolation system (Novagen). For Northern blot analyses, genomic and satBaMV RNAs were detected by 32P-labeled probes specific for the 3′ end of genomic RNA (20) and satBaMV RNA (23), respectively. The hybridized membranes were exposed by PhosphorImager (Molecular Dynamics) and quantified by ImageQuant, version 3.3 (Molecular Dynamics).
Primer extension and DNA sequencing.
Primer extensions were performed using total RNAs extracted from leaves of C. quinoa or using mRNA extracted from infected protoplasts of N. benthamiana as templates. Total RNA or mRNA was annealed with primer B56 (sequences complementary to nt 5631 through 5647, the CP region of the BaMV-V isolate) (40) or primer BS9 (sequences complementary to nt 338 through 352, the P20-coding region of satBaMV) (22). For all experiments, 500 pM primer was mixed with 5-μl RNA samples extracted from 5 mg of C. quinoa leaves or from 4 × 105 protoplasts. Primer extensions were carried out as previously described (15). Extension products were analyzed on 6% polyacrylamide–7 M urea gel containing 0.5× TBE (133 mM Tris-HCl [pH 8.3], 44 mM boric acid, and 2.5 mM EDTA) and exposed by PhosphorImager. The DNA sequences were determined by the chain termination procedure, using Sequenase, version 2.0 (Amersham).
RESULTS
Transcription of sgRNA via satBaMV cassette.
To explore the possibility of sgRNA being transcribed from the satBaMV cassette, a full-length satBaMV-derived cDNA clone, pICF4, was constructed, in which the putative SGP of the BaMV CP gene with 107 nt was inserted just before the start codon of the P20 gene of satBaMV (Fig. 1A). This putative SGP contains the whole intercistronic region of 41 nt between ORF4 and ORF5 of BaMV-V (40) and its upstream sequences extended downstream of the first AUG, thus avoiding the possible interference of ORFs from the expression of the P20 gene in pICF4 satBaMV. The 5′ end of the putative SGP lies within the C-terminal coding region of ORF4 and contains multiple stop codons. The activity of the SGP was analyzed by coinoculation of barley protoplasts with BaMV-V genomic RNA (designated BaMV-L RNA and free of satBaMV) and pICF4 transcripts. Protoplasts inoculated with BaMV-L RNA alone or coinoculated with wild-type pBSF4 transcripts were used as controls. Protoplast samples were harvested every 8 h postinoculation (h.p.i.), and total RNA was extracted. Northern blot analyses with the BaMV-specific probe revealed that all of the genomic RNAs accumulated to similar levels regardless of whether the inocula contained the satBaMV transcripts, indicating that the presence of the satBaMV transcripts did not significantly affect helper virus replication (Fig. 1B). Genomic RNAs were first detectable at 8 h.p.i., quickly increased from 8 to 16 h.p.i., and further continued to increase to 24 h.p.i. This pattern of increasing accumulation was also observed for pBSF4 and pICF4 by the satBaMV-specific probe (Fig. 1C). An additional RNA about 0.7 kb in length, presumably transcribed from the functional SGP, was detectable at 8 h.p.i. but increased greatly by 24 h.p.i. in pICF4 coinoculated protoplasts (Fig. 1C, lanes 7 through 9). No such RNA was detectable in the protoplasts coinoculated with pBSF4 transcripts (Fig. 1C, lanes 4 through 6). It should be noted that the level of satBaMV decreased when sgRNA was expressed (Fig. 1C, lanes 7 through 9). When the transcription ratio was determined by the quantity of 0.7-kb sgRNA transcribed over its satBaMV RNA, the transcription ratio of ICF4 was only 4% at 8 h.p.i., increased to 40% at 16 h.p.i., and reached a level of 70% at 24 h.p.i. (Fig. 1C).
When the same experiment was performed with C. quinoa plants and the leaves were harvested every 3 days postinoculation (d.p.i.), results similar to those with protoplasts were obtained (Fig. 6A and B). The 0.7-kb RNA detected in protoplasts was also detectable in C. quinoa leaves when coinoculated with pICF4 transcripts. However, the transcription ratios were higher than those in protoplasts, reaching levels of 90, 110, and 80% at 3, 6, and 9 d.p.i., respectively (Fig. 6B, lanes 3, 6, and 9). To determine whether the 0.7-kb RNA was encapsidated, viral RNAs were extracted from the purified virions of coinoculated leaves of C. quinoa. As previously shown (15), in addition to genomic RNA, BaMV-L contained the encapsidated sgRNAs which were detectable by ethidium bromide (EtBr) staining of the agarose gel (Fig. 6C, lane 1). pBSF4 and pICF4 were also visualized in virion RNA preparations stained with EtBr; however, the 0.7-kb sgRNA was only barely visible (Fig. 6C, lanes 2 and 3). The encapsidation of pICF4 and 0.7-kb sgRNA was further confirmed by Northern blot analysis with the satBaMV-specific probe (Fig. 6D). It is interesting to note that this sgRNA was rather poorly encapsidated compared to satBaMV.
FIG. 6.
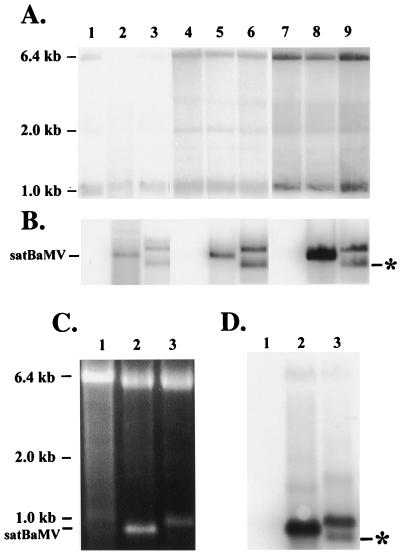
Time course accumulation of BaMV-V genomic RNA, satBaMV, and its derived sgRNA in infected leaves of C. quinoa. (A and B) Total RNAs extracted from 5-mg leaves inoculated with BaMV-L RNA only (lanes 1, 4, and 7) or together with pBSF4 (lanes 2, 5, and 8) or pICF4 (lanes 3, 6, and 9) transcripts at 3 d.p.i. (lanes 1 through 3), 6 d.p.i. (lanes 4 through 6), and 9 d.p.i. (lanes 7 through 9) were subjected to Northern blotting with a BaMV-specific (20) (A) or satBaMV-specific (23) (B) riboprobe. (C and D) For staining (C) and Northern blot analysis (D) of encapsidated satBaMV and its sgRNA, virion RNAs were purified from 0.2 g of C. quinoa leaves and inoculated with BaMV-L RNA only (lane 1) or with pBSF4 (lane 2), and pICF4 (lane 3) transcripts were separated by electrophoresis in a 1% agarose gel and stained with EtBr (C) or analyzed by Northern hybridization with the satBaMV probe (D). The asterisks indicate the position of the 0.7-kb sgRNA transcribed from pICF4.
Identification of the 5′ terminus of the 0.7-kb sgRNA.
Presumably, if the 0.7-kb RNA is synthesized via the SGP of 1.0-kb sgRNA in the satBaMV cassette, the 5′ terminus of 0.7-kb RNA should be identical to that of the 1.0-kb sgRNA, which is located 12 nt downstream from the octanucleotide motif (15; also Fig. 7A, lane 1). To map the 5′ terminus of the 0.7-kb RNA, primer extension was performed by using total RNAs extracted from C. quinoa leaves coinoculated with pICF4 transcripts as templates. The primer BS9 (complementary to the P20 gene of satBaMV [Fig. 1A]) extended a predominant and a minor product (Fig. 7B, lane 1). The former was at a position identical to that of the 1.0-kb sgRNA, indicating that the 0.7-kb RNA was indeed transcribed from the SGP in the satBaMV cassette. The minor product, with one nucleotide difference from the predominant one, may have arisen from insertion of a residue complementary to the cap structure as suggested by White and Mackie (38).
FIG. 7.
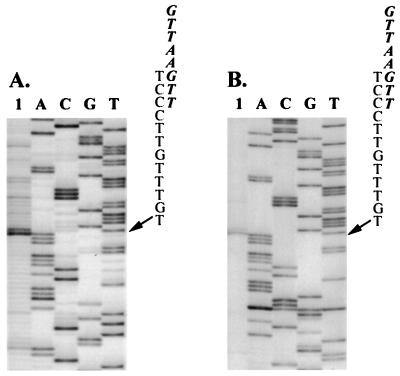
Identification of the 5′ termini of sgRNAs by primer extension. Total RNAs (lanes 1) extracted from C. quinoa leaves infected with BaMV-L RNA only (A) or together with pICF4 (B) were hybridized with primer B56 (complementary to the CP gene of BaMV) (A) or primer BS9 (complementary to the P20 gene of satBaMV) (B) for primer extension. The products were displayed on a 6% polyacrylamide–7 M urea sequencing gel. Lanes A, C, G, and T contained the products of a dideoxynucleotide-sequencing reaction performed on cDNA clone pBa19 of BaMV-V genomic RNA (40) with primer B56 (A) and on pICF4 with primer BS9 (B). Arrows indicate the position of the transcription initiation site. Letters represent the sequences between the octanucleotide motif (in italics) and the transcription initiation site.
Enhancement of sgRNA transcription by extension of the SGP to the CP coding region.
To test whether the extension of the SGP to the coding region of the CP gene can enhance sgRNA transcription, plasmids containing various extensions of SGP toward the CP-coding region were constructed. The first 9, 18, 36, and 54 nt of the CP gene were added downstream of the SGP sequences of pICF4 to make plasmids pICN9, pICN18, pICN36, and pICN54, respectively (Fig. 2A). The transcriptional activity of pICF4 and its variants was assayed in coinoculated protoplasts of N. benthamiana at 24 h.p.i (Fig. 2B). In three independent experiments, the transcription ratios of pICN9, pICN18, pICN36, and pICN54 were enhanced to levels that were 118, 127, 230, and 200%, respectively, over that of pICF4 (Fig. 2C). Similar results for pICN18 were also obtained with inoculated C. quinoa leaves (data not shown).
Deletion analyses of the SGP of 1.0-kb sgRNA.
As shown in Fig. 3A, the secondary structure of the minus-strand SGP of 1.0-kb sgRNA was predicted by the STAR computer program. This structure contained four stem-loops (nt −91 through −60, −59 through −31, −30 through −8, and −7 through +16, designated SL4, SL3, SL2, and SL1, respectively, when the transcription initiation site was taken as +1). SL1 contained the transcription initiation site, and SL2 contained the conserved octanucleotides among the potexviruses. To dissect the functional domain of SGP, four deletion mutants were generated in this region by site-directed mutagenesis (Fig. 3B). pIC5, pIC4, and pIC3 are mutants containing progressive SGP deletions from the 5′ end of pICN18, in which SL4 was deleted in pIC5; both SL4 and SL3 were deleted in pIC4, while SL4, SL3, and SL2 were deleted in pIC3. The pIC2 contained SL2 as well as the truncated SL1 (nt −8 through +3). The overall structures of these mutated SGP regions were predicted not to change, by the STAR program.
When assayed in protoplasts of N. benthamiana at 24 h.p.i., Northern blot analyses revealed that the transcription ratios of pICN18, pIC5, and pIC4, were 66, 38, and 19%, respectively, whereas the transcription of pIC3 and pIC2 had dropped to 8 and 7%, respectively (Fig. 3C and 3E). The relative transcription levels of pIC5 and pIC4 were 58 and 30% that of pICN18, respectively, while the transcription activities of pIC3 and pIC2 were significantly lower, with levels only 12 and 11% that of pICN18, respectively (Fig. 3E). Although similar relative transcription levels were also obtained in the coinoculated leaves of C. quinoa at 6 d.p.i., plasmids ICN18, IC5, and IC4 resulted in transcription ratios that were substantially higher in C. quinoa leaves than in N. benthamiana protoplasts in three independent experiments (Fig. 3D and E). For instance, the transcription ratios in C. quinoa leaves increased to 195% for pICN18, 128% for pIC5, and 68% for pIC4. In summary, the SL1 and SL2 of SGP maintained in pIC4 supported about one-third (30 to 34%) of the activity of pICN18 for sgRNA transcription, and one additional stem-loop in pIC5 supported about two-thirds (58 to 65%) of the activity in vivo. Mutants pIC3, with SL1 only, and pIC2, with SL2 and truncated SL1, both greatly reduced their transcription abilities. These data suggest that SL2 and SL1 are required for functional SGP and that SL3 and SL4 can enhance the sgRNA transcription level.
Mutational analyses of SL2.
From the results described above, the stem structure SL2 is essential for sgRNA transcription. Four mutants were constructed in the stem of SL2 (Fig. 4A). pIC8-C/C, pIC9-G/G, and pIC14-CCC disrupted the stem structure in the SL2 by replacing the GGG (nt −9 through −11)/CCC (nt −24 through −26) with GGC/CCC, GGG/GCC, and CCC/CCC, respectively. pIC10-G/C restored potential base pairing, which was disrupted in pIC8-C/C and pIC9-G/G by replacing the GGG/CCC with GGC/GCC. The predicted structures for the site-directed mutants pIC8, pIC9, and pIC14 revealed no existence of SL2 or small SL2 structures but no other stem-loop structures in SGP were altered. However, the SL2 structures were indeed restored in compensatory mutant pIC10.
The transcription ratios of these mutants were analyzed in inoculated protoplasts of N. benthamiana at 24 h.p.i. In three independent experiments, the relative transcription ratios of the base-pairing-disrupted mutants pIC8-C/C, pIC9-G/G, and pIC14-CCC dropped to 27, 25, and 3%, respectively, when compared to the transcription ratio of pICN18. However, the compensatory mutant pIC10-G/C increased to 85% (Fig. 4B and 4C). Similar relative transcription levels compared to pICN18 were also obtained in the coinoculated leaves of C. quinoa at 7 d.p.i. (Fig. 4C).
Since the loop structure of SL2 contains the conserved octanucleotides which are known to be important for potexviruses replication (14, 39), a plasmid, pIC7, derived from pICN18 was constructed in which the octanucleotide motif 3′-CAATTCAA-5′ in the minus strand was changed into 3′-CCTAAATA-5′ (Fig. 4A). A transcriptional-activity assay of pIC7 revealed that it exhibited only 9% activity compared with pICN18, both in coinfected protoplasts of N. benthamiana at 24 h.p.i. (Fig. 4B and 4C) and leaves of C. quinoa at 7 d.p.i. (Fig. 4C). In summary, the SL2 stem-disrupted mutants or octanucleotide mutant significantly decreased the SGP activity whereas the compensatory mutant restored the SGP activity. These data indicate that the conserved octanucleotides in the loop region and the stem structure of SL2 play an essential role in the functional SGP.
Transcription activity of SGP of 2.0-kb sgRNA.
In general, the two major 2.0- and 1.0-kb sgRNAs of BaMV were easily detected in the inoculated protoplasts, but the 2.0-kb RNA occurred in much lower levels in the infected plants (15) (see also Fig. 5B). To assay the transcription activity of the putative SGP of the BaMV 2.0-kb sgRNA, plasmid pICORF2 was constructed in which the satBaMV cassette contained the 130 nt upstream of the start codon of the ORF2 gene (Fig. 5A). Inoculation of pICORF2 also resulted in the generation of a 0.7-kb sgRNA, with a transcriptional level 20% that of pICF4, when protoplasts were coinoculated with BaMV-L RNA (Fig. 5C, lane 1). However, this 0.7-kb RNA was only barely detectable in coinoculated leaves of C. quinoa at 6 d.p.i. (Fig. 5C, lane 4) and also at 1, 2, or 3 d.p.i. (data not shown) by Northern blot analysis. This is consistent with our previous data that the 2.0-kb sgRNA accumulated substantially in protoplasts but much less in infected leaves (15) (Fig. 5B, lanes 1 and 4).
To analyze the transcription initiation site of the 0.7-kb RNA generated by pICORF2, primer extension was performed using total RNA extracted from infected leaves as a template. The primer BS9 extended a product terminating 12 nt downstream of the octanucleotide motif of SGP of the ORF2 gene (Fig. 8A, lane 2). The position of the product corresponded to the 5′ terminus of the BaMV 2.0-kb sgRNA as determined by Lee et al. (15), even though its quantity was only 5% of that of pICF4 (data not shown). This may explain the low accumulation of sgRNA transcribed by pICORF2 in leaves. Primer extension analysis of mRNA extracted from coinoculated N. benthamiana protoplasts showed similar results (Fig. 8A, lane 1). We concluded that the sgRNA was indeed transcribed by the SGP of the BaMV 2.0-kb sgRNA but was differentially expressed compared with the 1.0-kb sgRNA.
FIG. 8.
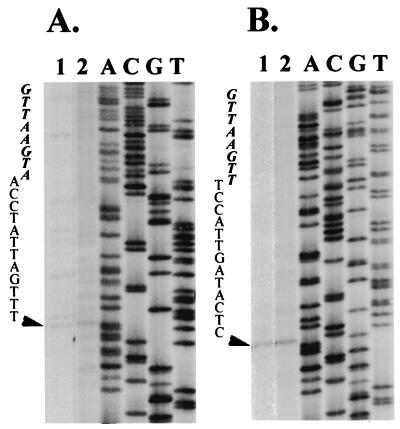
Primer extension analyses of the sgRNAs transcribed by pICORF2 (A) and pICPVX (B). Messenger RNAs extracted from 2 × 105 protoplasts of N. benthamiana at 24 h.p.i. (lane 1) or total RNAs extracted from 5-mg leaves of C. quinoa at 6 d.p.i. (lane 2) coinoculated with pICORF2 (A) or pICPVX (B) transcripts and BaMV-L RNA were hybridized with primer BS9 for primer extension. The products were displayed on a 6% polyacrylamide–7 M urea sequencing gel. Lanes A, C, G, and T contained the products of a dideoxynucleotide sequencing reaction performed on cDNA clone pICORF2 (A) or pICPVX (B) with primer BS9. Arrows indicate the positions of the transcription initiation sites, and letters represent the sequences between the octanucleotide motif (in italics) and the transcription initiation sites.
Transcription of sgRNA by a heterologous SGP in the satBaMV cassette.
The specificity and transcription efficiency of the sgRNA directed by a heterologous SGP were analyzed using the satBaMV cassette. Plasmid pICPVX was generated to carry the putative SGP of the 0.9-kb sgRNA of PVX. pICPVX gave rise to a 0.7-kb sgRNA at a transcription level of 20% of that in protoplasts coinoculated with pICF4 at 24 h.p.i. (Fig. 5C, lane 3). This 0.7-kb sgRNA was detected at a reduced level in leaves at 6 d.p.i. (Fig. 5C, lane 6). Analyses of primer extension indicated that the 5′ terminus of the 0.7-kb RNA mapped 13 nt downstream of the octanucleotide motif (Fig. 8B, lanes 1 and 2), a position which corresponds to that of the PVX 0.9-kb sgRNA as determined by Kim and Hemenway (12). Evidently, the heterologous SGP can be utilized in the satBaMV cassette with a reduced efficiency.
DISCUSSION
SatBaMV cassette as an assay system to characterize the SGP in vivo.
The mutation of viral RNA in the SGP region has added greatly to our understanding of the mechanism of sgRNA transcription. These mutants may affect the accumulation of gene products of sgRNA and thus interfere with viral replication. For instance, the SGP of AMV was characterized by the duplicated SGPs (34), and the position of duplicated SGPs may affect sgRNA transcription (5, 6). Moreover, the SGPs of coronavirus (35) and Sindbis virus (16) were characterized by insertion of SGP into the defective interfering (DI) RNA. The viability and stability of DI RNAs are dependent on the site of insertion or on the developmental stage of the DI RNAs (28). In this study, we used the satBaMV cassette to characterize the SGP in vivo. The advantages of using the satBaMV system are that (i) the satBaMV can replicate autonomously in the presence of a helper virus, (ii) the satBaMV does not interfere with the replication of the helper virus, and (iii) the satBaMV accumulates at a higher level than the helper virus in the infected cells (15). To the best of our knowledge, this is the first report that satellite RNA can provide an assay system to characterize SGPs in vivo.
Although sgRNA could be generated via the satBaMV cassette in the coinoculated protoplasts of barley or N. benthamiana and leaves of C. quinoa, the transcription ratio of the satBaMV cassette varied. The SGP of BaMV 1.0-kb sgRNA showed more activity in infected leaves than in infected protoplasts (Fig. 3D and E), whereas the SGP of BaMV 2.0-kb sgRNA had an adverse effect (Fig. 5C, lanes 1 and 4). Of special note in our study, SGP mutants exhibited similar relative-transcription levels to those of the wild-type pICN18 in coinoculated protoplasts and leaves (Fig. 3E). However, transcripts derived from infectious cDNA clones containing the modified SGP region upstream of the CP gene of PVX showed poor replication in infected plants, even though considerable accumulation of those mutants was observed in the infected protoplasts (12, 13). This may prove the additional advantage of using the satBaMV cassette to characterize the SGP.
Mapping the SGP of 1.0-kb sgRNA.
The secondary structure of the minus-strand SGP of 1.0-kb sgRNA was predicted by the computer STAR program to comprise four consecutive stem-loops (Fig. 3A). The existence of the predicted SL1, SL2, SL3, and SL4 was supported by enzymatic probing with RNase T1, RNase T2, RNase V1, and RNase A (N. S. Lin and C. P. Cheng, unpublished data). The results from satBaMV cassettes carrying the various SGP sequences showed that the sequences between nt −30 and +16 contained two stable RNA stem-loop structures, SL1 and SL2, required for basal SGP activity (Fig. 3A). The core promoter of BaMV 1.0-kb sgRNA contains 46 nt. Extension of the SGP to the 5′ border nt −59 and −91 enhanced the SGP activity two- and threefold, respectively. These results are consistent with those found in other alpha-like superfamily viruses which contain some modulated enhancers to stimulate and/or modulate gene expression in addition to the core promoter-like sequences (24). Similarly, extension of the SGP into the N-terminal 36 nt of the CP gene enhanced SGP activity twofold. Such a downstream enhancer extended into the ORF was known to sgRNAs coding for CP of a number of RNA viruses (2, 10, 29). The upstream and downstream enhancers of BaMV 1.0-kb sgRNA were located at the coding region of ORF4 and the CP gene, respectively. It is an intriguing question how the genetic information at these locations can be used as a coding region for protein and as a promoter region for sgRNA synthesis.
When the sequences of SL2 were further analyzed by site-directed mutagenesis, the results showed that the stem-region-disrupted mutants abolished the SGP activity, whereas the compensatory mutant restored the SGP activity, indicating that the stem structure was important for the SGP-directed sgRNA transcription in vivo. However, the sgRNA transcription of PVX required complementarity between the conserved octanucleotide motif and the 3′-terminal sequences of the minus-strand genomic RNA (13). This long-distance RNA-RNA interaction between the octanucleotide motif and the terminal sequences affected genomic RNA accumulation and the sgRNA transcription (12, 13). Growing evidence also shows that different functional long-distance interactions occur in a variety of plus-strand RNA viruses for the regulation of sgRNA synthesis (13, 41). In the pICF4 satBaMV, the octanucleotide motif of SGP may interact with the 3′ end of minus-strand BaMV genomic RNA in trans (complementarity of 8 nt in the first 11 nt) or with the 3′ end of minus-strand satBaMV RNA in cis (complementarity of 10 nt in the first 11 nt), which may facilitate the sgRNA synthesis via the RNA-RNA interaction. However, octanucleotide motif mutant pIC7, whose complementarity with the terminus of satBaMV was reduced to 5 bp, resulted in barely detectable sgRNA accumulation (Fig. 4).
The activity of SGP of 2.0-kb sgRNA.
Analyses of the SGP of 2.0-kb sgRNA with the satBaMV cassette showed that its activity is less efficient than that of the SGP of 1.0-kb sgRNA. The positional effect in the genomic RNA can be excluded since we inserted both SGPs into the same position. pICORF2 exhibited significant activity for sgRNA transcription in coinoculated protoplasts (Fig. 5C, lane 1) whereas much less activity was detected in leaves of C. quinoa (Fig. 5C, lane 4). A consistently detected level was found in 2.0-kb sgRNA in the BaMV-infected protoplasts and leaves of C. quinoa (15) (Fig. 5B, lanes 1 and 4). The mechanism for the down-regulation of the 2.0-kb sgRNA in infected leaves remains to be further investigated although the secondary structure of the SGP of the 2.0-kb sgRNA predicted by the STAR program revealed a similar structure to that of 1.0-kb SGP. The SL1 and SL2 structures displayed on the 1.0-kb SGP are present in 2.0-kb SGP. The conserved octanucleotide motif and transcription site of the 2.0-kb SGP are also located at the loop region of SL2 and SL1, respectively.
The activity of heterologous SGP in satBaMV cassette.
The pICPVX could direct sgRNA transcription by the SGP of PVX 0.9-kb sgRNA in this satBaMV cassette system. This transcription activity was low but significantly above the background (Fig. 5C). The SGPs of BaMV and PVX share 45% identity in sequence at the 3′ terminal 40 nt of SGPs. In pICPVX, there also exists a complementarity between the octanucleotide and the 3′ end of minus-strand satBaMV genomic RNA in cis (complementarity, 10 nt in the first 11 nt); however, the stable stem-loop structures were absent in the SGP of 0.9-kb sgRNA of PVX as predicted by the STAR program. In members of alphaviruses of plant and animal origin, the required nucleotides within the sgRNA promoter for sgRNA synthesis are highly conserved (24, 26, 30). The RdRp of alphaviruses, such as BMV (30) and Sindbis virus (9), can utilize the heterologous SGPs of other alphaviruses for sgRNA synthesis. Our results also favor the conserved mechanism of the recognition of RdRp for the synthesis of sgRNAs of alphaviruses.
ACKNOWLEDGMENTS
We thank Michael M. C. Lai for critical reading of the manuscript and Hsin-Chuan Chen for technical assistance.
This research was supported in part by National Science Council Project grants NSC 86-2311-B-001-034-B13 and NSC 87-2311-B-001-007-B13 and by the Academia Sinica, Taipei, Taiwan, Republic of China.
REFERENCES
- 1.Adkins S, Siegel R W, Sun J H, Kao C C. Minimal templates directing accurate initiation subgenomic RNA synthesis in vitro by the brome mosaic virus RNA-dependent RNA polymerase. RNA. 1997;3:634–647. [PMC free article] [PubMed] [Google Scholar]
- 2.Balmori E, Gilmer D, Richards K, Guilley H, Jonard G. Mapping the promoter for subgenomic RNA synthesis on beet necrotic yellow vein virus RNA 3. Biochimie. 1993;75:517–521. doi: 10.1016/0300-9084(93)90056-x. [DOI] [PubMed] [Google Scholar]
- 3.Beck D L, Guilford D M V, Andersen M T, Forster R L S. Triple gene block proteins of white clover mosaic potexvirus are required for transport. Virology. 1991;183:695–702. doi: 10.1016/0042-6822(91)90998-q. [DOI] [PubMed] [Google Scholar]
- 4.Chang R C, Chen J C, Shaw J F. Facile purification of highly active recombinant Staphylococcus hyicus lipase fragment and characterization of a putative Lid region. Biochem Biophy Res Commun. 1996;228:774–779. doi: 10.1006/bbrc.1996.1730. [DOI] [PubMed] [Google Scholar]
- 5.Chapman S, Kavanagh T A, Baulcombe D C. Potato virus X as a vector for gene expression in plants. Plant J. 1992;2:549–557. doi: 10.1046/j.1365-313x.1992.t01-24-00999.x. [DOI] [PubMed] [Google Scholar]
- 6.Culver J N, Lehto K, Close S M, Hilf M E, Dawson W O. Genomic position affects the expression of tobacco mosaic virus movement and coat protein genes. Proc Natl Acad Sci USA. 1993;90:2055–2059. doi: 10.1073/pnas.90.5.2055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Dolja V V, Grama D P, Morozov S Y, Atabekov J G. Potato virus X-related single- and double-stranded RNAs. FEBS Lett. 1987;214:308–312. [Google Scholar]
- 8.French R, Ahlquist P. Characterization and engineering of sequences controlling in vivo synthesis of brome mosaic virus subgenomic RNA. J Virol. 1988;62:2411–2420. doi: 10.1128/jvi.62.7.2411-2420.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hertz J M, Huang H V. Utilization of heterologous alphavirus junction sequences as promoters by Sindbis virus. J Virol. 1992;66:857–864. doi: 10.1128/jvi.66.2.857-864.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Johnston J C, Rochon D M. Deletion analysis of the promoter for the cucumber mosaic necrosis virus 0.9-kb subgenomic RNA. Virology. 1995;214:100–109. doi: 10.1006/viro.1995.9950. [DOI] [PubMed] [Google Scholar]
- 11.Kim K H, Hemenway C. The 5′ nontranslated region of potato virus X RNA affects both genomic and subgenomic RNA synthesis. J Virol. 1996;70:5533–5540. doi: 10.1128/jvi.70.8.5533-5540.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kim K H, Hemenway C. Mutations that alter a conserved element upstream of the potato virus X triple block and coat protein genes affect subgenomic RNA accumulation. Virology. 1997;232:187–197. doi: 10.1006/viro.1997.8565. [DOI] [PubMed] [Google Scholar]
- 13.Kim K H, Hemenway C L. Long-distance RNA-RNA interactions and conserved sequence elements affect potato virus X plus-strand RNA accumulation. RNA. 1999;5:636–645. doi: 10.1017/s1355838299982006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lai M M, Cavanagh D. The molecular biology of coronaviruses. Adv Virus Res. 1997;38:1–100. doi: 10.1016/S0065-3527(08)60286-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lee Y S, Lin B Y, Hsu Y H, Chang B Y, Lin N S. Subgenomic RNAs of bamboo mosaic potexvirus-V isolate are packaged into virions. J Gen Virol. 1998;79:1825–1832. doi: 10.1099/0022-1317-79-7-1825. [DOI] [PubMed] [Google Scholar]
- 16.Levis R, Schlesinger S, Huang H V. Promoter for Sindbis virus RNA-dependent subgenomic RNA transcription. J Virol. 1990;64:1726–1733. doi: 10.1128/jvi.64.4.1726-1733.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lin N S, Chen C C. Association of bamboo mosaic virus (BaMV) and BaMV-specific electron-dense crystalline bodies with chloroplasts. Phytopathology. 1991;81:1551–1555. [Google Scholar]
- 18.Lin N S, Huang T Y, Hsu Y H. Infection of barley protoplasts with bamboo mosaic virus RNA. Bot Bull Acad Sin. 1992;33:271–275. [Google Scholar]
- 19.Lin N S, Lin F Z, Huang T Y, Hsu Y H. Genome properties of bamboo mosaic virus. Phytopathology. 1992;82:731–734. [Google Scholar]
- 20.Lin N S, Chai Y J, Huang T Y, Chang T Y, Hsu Y H. Incidence of bamboo mosaic potexvirus in Taiwan. Plant Dis. 1993;77:448–450. [Google Scholar]
- 21.Lin N S, Lin B Y, Lo W W, Hu C C, Chow T Y, Hsu Y H. Nucleotide sequence of the genomic RNA of bamboo mosaic potexvirus. J Gen Virol. 1994;75:2513–2518. doi: 10.1099/0022-1317-75-9-2513. [DOI] [PubMed] [Google Scholar]
- 22.Lin N S, Hsu Y H. A satellite RNA associated with bamboo mosaic potexvirus. Virology. 1994;202:707–714. doi: 10.1006/viro.1994.1392. [DOI] [PubMed] [Google Scholar]
- 23.Lin N S, Lee Y S, Lin B Y, Lee C W, Hsu Y H. The open reading frame of bamboo mosaic potexvirus satellite RNA is not essential for its replication and can be replaced with a bacterial gene. Proc Natl Acad Sci USA. 1996;93:3138–3142. doi: 10.1073/pnas.93.7.3138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Maia I G, Seron K, Haenni A-L, Bernardi F. Gene expression from viral RNA genomes. Plant Mol Biol. 1996;32:367–391. doi: 10.1007/BF00039391. [DOI] [PubMed] [Google Scholar]
- 25.Marsh L E, Dreher T W, Hall T C. Mutational analysis of the core and modulator sequences of the BMV RNA3 subgenomic promoter. Nucleic Acids Res. 1988;16:981–995. doi: 10.1093/nar/16.3.981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ou J H, Rice C M, Dalgarno L, Strauss E G, Strauss J H. Sequence studies of several alphavirus genomic RNAs in the region containing the start of the subgenomic RNA. Proc Natl Acad Sci USA. 1982;79:5253–5239. doi: 10.1073/pnas.79.17.5235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Sawicki S G, Sawicki D L. A new model for coronavirus transcription. Adv Exp Med Biol. 1998;440:215–219. doi: 10.1007/978-1-4615-5331-1_26. [DOI] [PubMed] [Google Scholar]
- 28.Scholthof H B, Scholthof K-B G. Plant virus gene vectors for transient expression of foreign proteins in plants. Annu Rev Phytopathol. 1996;34:299–323. doi: 10.1146/annurev.phyto.34.1.299. [DOI] [PubMed] [Google Scholar]
- 29.Shivprasad S, Pogue G P, Lewandowski D J, Hidalgo J, Donson J, Grill L K, Dawson W O. Heterologous sequences greatly affect foreign gene expression in tobacco mosaic virus-based vectors. Virology. 1999;255:312–323. doi: 10.1006/viro.1998.9579. [DOI] [PubMed] [Google Scholar]
- 30.Siegel R W, Adkins S, Kao C C. Sequence-specific recognition of a subgenomic RNA promoter by a viral RNA polymerase. Proc Natl Acad Sci USA. 1997;94:11238–11243. doi: 10.1073/pnas.94.21.11238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sit T L, Vaewhongs A A, Lommel S A. RNA-mediated trans-activation of transcription from a viral RNA. Science. 1998;281:829–832. doi: 10.1126/science.281.5378.829. [DOI] [PubMed] [Google Scholar]
- 32.Smirnyagina E, Hsu Y H, Chua N, Ahlquist P. Second-site mutations in the brome mosaic virus RNA3 intercistronic region partially suppress a defect in coat protein mRNA transcription. Virology. 1994;198:427–436. doi: 10.1006/viro.1994.1054. [DOI] [PubMed] [Google Scholar]
- 33.Van der Kuyl A C, Langereis K, Houwing C J, Jaspars E M J, Bol J F. cis-acting elements involved in replication of alfalfa virus RNA in vitro. Virology. 1990;176:346–354. doi: 10.1016/0042-6822(90)90004-b. [DOI] [PubMed] [Google Scholar]
- 34.van der Vossen E A G, Notenboom T, Bol J F. Characterization of sequences controlling the synthesis of alfalfa mosaic virus subgenomic RNA in vivo. Virology. 1995;212:663–672. doi: 10.1006/viro.1995.1524. [DOI] [PubMed] [Google Scholar]
- 35.van Marle G, Luytjes W, van der Most R G, van der Straaten T, Spaan W J. Regulation of coronavirus mRNA transcription. J Virol. 1995;69:7851–7856. doi: 10.1128/jvi.69.12.7851-7856.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Verchot J, Angell S M, Baulcombe D C. In vivo translation of the triple gene block of potato virus X requires two subgenomic mRNAs. J Virol. 1998;72:8316–8320. doi: 10.1128/jvi.72.10.8316-8320.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.White K A, Bancroft J B, Mackie G A. Mutagenesis of a hexanucleotide sequence conserved in potexvirus RNAs. Virology. 1992;189:817–820. doi: 10.1016/0042-6822(92)90614-u. [DOI] [PubMed] [Google Scholar]
- 38.White K A, Mackie G A. Control and expression of 3′ open reading frames in clover yellow mosaic virus. Virology. 1990;179:576–584. doi: 10.1016/0042-6822(90)90124-a. [DOI] [PubMed] [Google Scholar]
- 39.Wung C H, Hsu Y H, Liou D Y, Huang W C, Lin N S, Chang B Y. Identification of the RNA-binding sites of the triple gene block protein 1 of bamboo mosaic potexvirus. J Gen Virol. 1999;80:1119–1126. doi: 10.1099/0022-1317-80-5-1119. [DOI] [PubMed] [Google Scholar]
- 40.Yang C C, Liu J S, Lin C P, Lin N S. Nucleotide sequence and phylogenetic analysis of a bamboo mosaic potexvirus isolate from common bamboo (Bambusa vulgaris McClure) Bot Bull Acad Sin. 1997;38:77–84. [Google Scholar]
- 41.Zhang G, Slowinski V, White K A. Subgenomic mRNA regulation by a distal RNA element in a (+)-strand RNA virus. RNA. 1999;5:550–561. doi: 10.1017/s1355838299982080. [DOI] [PMC free article] [PubMed] [Google Scholar]