Abstract
Recombinant multiepitope proteins (RMPs) are a promising alternative for application in diagnostic tests and, given their wide application in the most diverse diseases, this review article aims to survey the use of these antigens for diagnosis, as well as discuss the main points surrounding these antigens. RMPs usually consisting of linear, immunodominant, and phylogenetically conserved epitopes, has been applied in the experimental diagnosis of various human and animal diseases, such as leishmaniasis, brucellosis, cysticercosis, Chagas disease, hepatitis, leptospirosis, leprosy, filariasis, schistosomiasis, dengue, and COVID-19. The synthetic genes for these epitopes are joined to code a single RMP, either with spacers or fused, with different biochemical properties. The epitopes’ high density within the RMPs contributes to a high degree of sensitivity and specificity. The RMPs can also sidestep the need for multiple peptide synthesis or multiple recombinant proteins, reducing costs and enhancing the standardization conditions for immunoassays. Methods such as bioinformatics and circular dichroism have been widely applied in the development of new RMPs, helping to guide their construction and better understand their structure. Several RMPs have been expressed, mainly using the Escherichia coli expression system, highlighting the importance of these cells in the biotechnological field. In fact, technological advances in this area, offering a wide range of different strains to be used, make these cells the most widely used expression platform. RMPs have been experimentally used to diagnose a broad range of illnesses in the laboratory, suggesting they could also be useful for accurate diagnoses commercially. On this point, the RMP method offers a tempting substitute for the production of promising antigens used to assemble commercial diagnostic kits.
Keywords: Recombinant multiepitope proteins, Escherichia coli, Bioinformatics, Biophysical analysis, Immunodiagnosis
Introduction
Recombinant multiepitope proteins (RMP) are the result of epitopes joining to form a single molecule that does not exist in nature [1, 2]. A study by Dipti et al. (2006) is thought to be the first one to define an RMP as a molecule that contains linear, immunodominant, and conserved epitopes that are connected through linkers [1]. Since then, the commercial use of these molecules has gained market space with many applications related to human and animal health, such as the development of vaccines and diagnostic devices [3–7]. In fact, the global market of recombinant proteins is expected to grow by 12% from 2022 to 2030, with 2030 revenue estimated at USD 5.09 billion [8].
To form a new RMP, the first step involves selecting the epitopes to be used. This can be performed in several ways, such as choosing epitopes that have already been characterized as immunodominant in the literature [9–11], and through bioinformatics analyses [12–14]. Bioinformatics analysis identifies a pathogen’s antigens using computational analyses of its genome, without the need to manipulate the microorganism [15]. This implies a reduction in costs and research time, and minimizes the use of animals, proving to be an effective method for better targeting in in vitro and in vivo experiments [14, 16–19]. As much as it is an already consolidated area, and its importance demonstrated by several studies, the importance of bioinformatics became evident during the COVID-19 pandemic, during which it was necessary to select epitopes as quickly and inexpensively as possible to develop vaccines and diagnostic tests, increasing the number of studies using these analyses [20–24].
Designing the new RMP includes many steps, such as: (i) selection of how many epitopes will be used; (ii) selection of spacing linkers between each epitope; (iii) tag selection to allow for better heterologous expression and purification; and (iv), evaluation of the physicochemical RMPs parameters [2] (Fig. 1). The next step is to choose the most appropriate host organism for RMP expression. The recombinant protein technology has become available worldwide, and several expression platforms are now available [25]. Among them, Escherichia coli became the most popular expression platform due to its relative simplicity, quick and inexpensive cultivation, and the availability of various compatible biotechnological tools [26]. However, this expression system comes with several drawbacks, such as the production of the protein in inclusion bodies and the absence of post-translational modifications [26–28]. To overcome these problems, expression platforms, such as yeast, insect, and mammalian cells, have been developed, and the choice of the best platform depends on the characteristics of each RMP and its applicability [25–27, 29]. Nevertheless, the recombinant protein technology is an efficient method for obtaining antigens at a relatively low cost that favors a more cost-effective production.
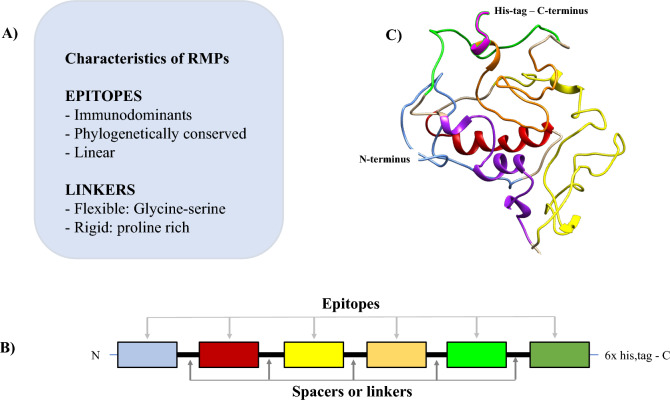
Fig. 1.
Design of a putative recombinant multiepitope protein. A General characteristics of majority RMPs available on literature. B According to the information presented in (A), a linear structure can be rationally drawn by the researchers. C The RMP 3D structure can be visualized from amino acid’s sequence by programs
Among their diverse applications, RMPs have been widely used, experimentally and commercially, to diagnose a wide range of human and animal diseases through tests, such as enzyme-linked immunosorbent assay (ELISA), immunofluorescence assays, and lateral flow tests. For example, RMPs has been used in Chagas IgG-ELISA® (NovaTec Immunodiagnostica GmbH; Dietzenbach, Germany) and Chagas Detect™ Plus (CDP) Rapid Test (InBios; Seattle, Washington, USA) commercial kits for Chagas disease detection. In this sense, these molecules offer advantages in the field of immunological diagnosis, such as increased sensitivity and specificity, which can improve diagnostic accuracy [1, 13]. Additionally, RMPs can be mass-produced, which facilitates diagnostic device standardization [13, 30–32]. The aim of this review was to select only studies using the nomenclature “recombinant multiepitope protein” and summarize all the research that used RMPs in diagnosing human and animal diseases, as well as explore significant issues surrounding this technology.
RMPs applied in disease diagnosis
RMPs applied in the diagnosis of diseases caused by bacteria
Currently, more than one thousand species of bacteria have been described as being pathogenic to vertebrate hosts [33, 34]. The most recent estimates have shown that more than seven million deaths were caused by bacterial infections in 2019, representing the second leading cause of global deaths [35]. Additionally, antibiotic resistance currently represents one of the greatest threats to global health [36, 37]. In this sense, the ability to provide a rapid and accurate diagnosis is essential for correct clinical conduct, improving the effectiveness of treatments and helping in antibiotic misuse [38]. RMPs has been applied in the experimental diagnoses of infections caused by bacteria and Table 1 summarizes the main points of these studies.
Table 1.
RMPs applied in the diagnosis of diseases caused by bacteria
| Disease/causative agent | RMP name | Host for protein expression | Serological assay | Samples (n) | Results | Reference/country |
|---|---|---|---|---|---|---|
| Human tuberculosis/Mycobacterium tuberculosis | TbF6 | E. coli cells | ELISA | 272 tuberculosis-positive serum samples/339 serum samples from healthy individuals |
Sensitivity: 71.2–86% Specificity: 94.7–98.3% |
[39]/USA |
| Human leptospirosis/Leptospira interrogans | r-LMP | E. coli) BL21(DE3) pLysS cells | ELISA | 156 leptospirosis-positive serum samples/10 serum samples from healthy individuals/24 serum samples from non-leptospirosis diseases | r-LMP was recognized by all positive samples, without cross-reactions | [40]/China |
| Human leprosy/Mycobacterium leprae | PADL | E. coli HMS-174 cells | ELISA | 54 leprosy- positive serum samples/18 serum samples from healthy individuals | PADL was recognized by positive samples | [41]/USA |
| Human tuberculosis/Mycobacterium tuberculosis | Fusion protein antigen | E. coli BL21(DE3) cells | ELISA | 171 tuberculosis-positive serum samples/48 serum samples from healthy individuals/38 serum samples from non-tuberculosis diseases |
Sensitivity: 42.1% Specificity: 89.5% |
[42]/China |
| Human tuberculosis/Mycobacterium tuberculosis | PstS1-LEP | E. coli BL21(DE3) cells | Indirect ELISA | 442 tuberculosis-positive serum samples/102 serum samples from healthy individuals/75 serum samples from non-tuberculosis diseases |
Sensitivity: 42.1–82% Specificity: 72.2–95.2% |
[43]/China |
| Human brucellosis/Brucella spp. | Recombinant protein | E. coli BL21(DE3) cells | Indirect ELISA | 146 brucellosis-positive serum samples/20 serum samples from healthy individuals/82 serum samples from non-brucellosis diseases |
Sensitivity: 88.89% Specificity: 85.54% |
[44]/China |
| Human Lyme disease/Borrelia burgdorferi | A/C-2, A/C-4, and A/C-7.1 | E. coli (XL1-Blue) cells | ELISA | 169 Lyme-positive serum samples/5 serum samples from healthy individuals |
A/C-2 Sensitivity: 80.17% Specificity: 52.83% A/C-4 results not shown A/C-7.1 Sensitivity: 91.37% Specificity: 73.58% |
[45]/Slovakia |
| Human and goat brucellosis/Brucella spp. | rOmp | E. coli BL21(DE3) cells | Indirect ELISA | 40 human and 31 goat brucellosis-positive serum samples/20 healthy human and 20 healthy goat serum samples | rOmp was recognized by human and goat positive samples | [46]/China |
| Goat brucellosis/Brucella spp. | rMEP | – | Indirect ELISA | 57 brucellosis-positive serum samples/36 healthy goat serum samples |
Sensitivity: 96.49% Specificity: 94.44% |
[47]/China |
| Bovine and goat brucellosis/Brucella spp. | Multi-epitope fusion protein | E. coli BL21 cells | Indirect ELISA | 116 and 140 goat brucellosis-positive serum samples/75 healthy bovine and 54 healthy goat serum samples |
Bovine Sensitivity: 97.85% Specificity: 96.61% Goat Sensitivity: 98.85% Specificity: 98.51% |
[48]/China |
| Human brucellosis/Brucella spp. | Fusion protein | E. coli BL21(DE3) cells | nano-p-ELISA | 121 brucellosis-positive serum samples/50 serum samples from healthy individuals/40 serum samples from non-brucellosis diseases |
Sensitivity: 92.38% Specificity: 98.35% |
[49]/China |
| Bovine tuberculosis/Mycobacterium bovis | BID109, TB1f, and TB2f | – | MAPIA and Dual Path Platform | 125 tuberculosis-positive serum samples/57 healthy bovine serum samples |
BD109 Sensitivity: 61.6% Specificity: 94.7% TB1f Sensitivity: 60.8% Specificity: 98.2% TB2f Sensitivity: 77.6% Specificity: 96.5% |
[50]/USA |
| Canine brucellosis/Brucella spp. | Multiepitope-based fusion protein | E. coli BL21(DE3) cells | Indirect ELISA | 34 brucellosis-positive serum samples/62 healthy dog serum samples |
Sensitivity: 97.06% Specificity: 100% |
[51]/China |
“–”: information not provided by the study
Houghton et al. (2002) [39] published the first study with RMPs applied to the diagnosis of bacterial disease. The authors designed TbF6 to diagnose tuberculosis, caused by Mycobacterium tuberculosis. To form TbF6, antigens were selected based on the published literature, and, sequentially, E. coli cells were used for heterologous protein expression. When conducting ELISA assays, TbF6 was combined with a a proline-rich antigen, with sensitivity values ranging from 71.2% to 86% and specificity values from 94.7% to 98.3%. Lin et al. (2008) [40] conducted a study to design a new RMP for the diagnosis of leptospirosis, caused by Leptospira interrogans. After applying bioinformatics analyses to the epitope selection, a new RMP, called r-LMP, was obtained using E. coli BL21(DE3) pLysS cells. Although the sensitivity and specificity values were not provided, results showed that all positive serum samples recognized r-LMP in an ELISA assay. Subsequently, Duthie et al. (2010) [41] developed a new RMP for diagnosing leprosy, caused by Mycobacterium leprae. The new RMP, called PADL, was formed by using peptides that reacted with positive serum samples. E. coli HMS-174 cells were used for PADL expression, and an ELISA assay was performed to verify PADL reactivity with positive human serum samples. Results showed that PADL was recognized by positive samples. However, sensitivity and specificity values were not shown.
Cheng et al. (2011) [42] developed a new RMP that could be used to diagnose human tuberculosis. Epitopes were selected based on published studies to form the fusion protein antigen, which was expressed in E. coli BL21(DE3) cells. An ELISA assay was performed to assess antigen reactivity with positive human serum samples, with sensitivity and specificity values of 42.1% and 89.5%, respectively. Later, Li et al. (2015) [43] designed a new RMP, named PstS1-LEP, to diagnose human tuberculosis. After selecting epitopes based on the literature, E. coli BL21(DE3) cells were used for PstS1-LEP production. After performing an indirect ELISA assay, results showed that sensitivity values ranged from 42.1% to 82%, depending on the antibody subclass and clinical form of the disease. Specificity values ranged from 72.2% to 95.2%. Yin et al. (2016) [44] conducted a study to construct a new RMP to diagnose human brucellosis, caused by Brucella spp. Bioinformatics analyses were performed to select epitopes and E. coli BL21(DE3) cells were used to express the new RMP. An indirect ELISA assay was performed, with sensitivity and specificity values of 88.89% and 85.54%, respectively.
Schreterova et al. (2017) [45] conducted a study aimed at developing new RMPs for diagnosing Lyme disease, caused by Borrelia burgdorferi. After using phage display and multiple alignments for epitope selection, the new RMPs, named A/C-2, A/C-,4, and A/C-7.1, were expressed in E. coli (XL1-Blue) cells. Among the tested RMPs, A/C-2 and A/C-7.1 showed the best results in the ELISA assay, with 80.17% and 91.37% sensitivity values, respectively. Furthermore, A/C-2 and A/C-7.1 had specificity values of 52.83% and 73.58%, respectively. Next, Yin and colleagues (2020) [46] developed a new RMP, called rOmp, for diagnosing human and goat brucellosis. Epitopes were selected through bioinformatics analyses and E. coli BL21(DE3) cells were used for heterologous protein expression. An indirect ELISA was performed to assess rOmp reactivity with positive human and goat serum samples. Results showed that rOmp was able to be recognized by both human and goat positive sera, although sensitivity and specificity values were not determined. Continuing the study, Yin et al. (2021) [47] tested rOmp for goat brucellosis diagnosis. After performing an indirect ELISA, 96.49% sensitivity and 94.44% specificity were obtained.
Yin et al. (2021) [48] developed a new RMP for diagnosing bovine and goat brucellosis. Epitopes were selected after bioinformatics analyses, and E. coli BL21 cells were used for heterologous protein expression. The multi-epitope fusion protein reactivity was verified through indirect ELISA. Sensitivity and specificity values for bovine brucellosis were determined as 97.85% and 96.61%, respectively, while 98.85% sensitivity and 98.51% specificity were observed using goat serum samples. In a similar approach, Yin et al. (2021) [49] selected epitopes through bioinformatics analyses to form a fusion protein for diagnosing human brucellosis. After performing a nano-p-ELISA assay, sensitivity and specificity values of 92.38% and 98.35, respectively, were determined.
Next, Lyashchenko et al. (2021) [50] developed three new RMPs for diagnosing bovine tuberculosis, caused by Mycobacterium bovis. To form these RMPs, named BID109, TB1f, and TB2f, antigens were selected based on the literature. Next, multiantigen print immunoassay and Dual Path Platform (DPP) assay were performed to assess the RMP’s reactivity with positive serum samples. A strong immunoreactivity was observed in the multiantigen print immunoassays for all tested RMPs. As for the DPP results, TB2f showed the best performance, with 77.6% sensitivity and 96.5% specificity. Lastly, Yao et al. (2022) [51] tested the multi-epitope fusion protein’s ability to diagnose canine brucellosis, which was previously designed by Yin et al. (2021) [49]. An indirect ELISA assay was performed, resulting in 97.06% sensitivity and 100% specificity.
RMPs applied in the diagnosis of diseases caused by fungus
The fungal kingdom has approximately six million species [52, 53], among which 200 have already been described as members of the human microbiome or as human pathogens [54], with 19 of them on the WHO’s fungal priority pathogens list [55]. Moreover, several species are also known to cause animal infections [56]. It is estimated that more than 150 million severe cases and 1.7 million human deaths occur worldwide annually [57]. Despite their importance, fungal diseases have largely been neglected over the years, and reliable diagnoses are only available for a small number of species [58]. Moreover, the diagnostic tests that do exist are not widely available [59, 60]. RMPs has been applied in the experimental diagnoses of infections caused by fungus and Table 2 summarizes the main points of these studies.
Table 2.
RMPs applied in the diagnosis of diseases caused by fungus
| Disease/causative agent | RMP name | Host for protein expression | Serological assay | Samples | Results | Reference/country |
|---|---|---|---|---|---|---|
| Human pneumonia/Pneumocystis jirovecii | RSA | E. coli BL21 Star(DE3) cells | Indirect ELISA | 88 pneumonia-positive serum samples/17 serum samples from healthy individuals |
Associated with clinical diagnosis-Sensitivity: 100% Specificity: 80.8% Without an associated clinical diagnosis-Sensitivity: 68% Specificity: 61.8% |
[61]/Portugal |
| Human cryptococcosis/Cryptococcus spp | Recombinant multiepitope proteins A, B, C, and D | E. coli BL21(DE3) cells | ELISA | 70 cryptococcosis-positive serum samples/10 serum samples from healthy individuals/68 serum samples from non-cryptococcosis diseases |
Protein A Sensitivity: 57.14% Specificity: 90% Protein B Sensitivity: 80% Specificity: 90% Protein C Sensitivity: 88.57% Specificity: 90% Protein D Sensitivity: 88.57% Specificity: 100% |
[62]/Brazil |
| Human pneumonia/Pneumocystis jirovecii | Kex1 RSA | E. coli XJb(DE3) cells | ELISA | 48 pneumonia-positive serum samples/104 serum samples from non-P. jirovecii infection |
Sensitivity: 70.8% Specificity: 75.0% |
[63]/Portugal |
Keeping in mind the need to develop new diagnostic tests, few studies in the literature have used RMPs in the diagnosis of fungus infections. A study published by Tomás et al. (2016) [61] was the first to use RMPs for this purpose. The authors developed a new RMP, called RSA, to diagnose human pneumonia, caused by Pneumocystis jirovecii. The epitopes comprising the new RMP were selected by studying the immunogenicity of a major surface glycoprotein, and RSA was obtained through heterologous expression in E. coli BL21 Star(DE3) cells. After performing an in-house ELISA, the diagnosis using RSA presented 100% sensitivity and 80.8% specificity when associated with a clinical diagnosis. When analyzed without associating it with a clinical diagnosis, the sensitivity and specificity values dropped to 68% and 61.8%, respectively.
Brandão et al. (2018) [62] conducted a study to develop a new RMP for the diagnosis of human cryptococcosis, caused by Cryptococcus spp. Authors selected epitopes through bioinformatics analyses to form four new RMPs, called recombinant multiepitope proteins A, B, C, and D. All genes were expressed in E. coli BL21(DE3) cells, and an in-house ELISA was used to evaluate their performance. Proteins C and D showed the best results, in which both demonstrated 88.57% sensitivity, and specificity of 90% and 100%, respectively. Lastly, Tomás et al. (2020) [63] developed and tested a new RMP for the diagnosis of human pneumonia, caused by P. jirovecii. The epitopes were selected by studying the immunogenicity of P. jirovecii’s kexin-like serine protease, with the new RMP being designated Kex1 RSA. After obtaining Kex1 RSA using E. coli XJb(DE3) cells, a sensitivity and specificity of 70.8% and 75.0%, respectively, was confirmed following an indirect ELISA.
RMPs applied in the diagnosis of diseases caused by protozoa
Protozoa species are found in all possible habitats [64], and although only a small percentage of species are known to be human and animal pathogens [65], they pose an important public health threat with a profound economic impact, being responsible for millions of deaths and significant morbidity worldwide [66]. Millions of human cases are reported annually related to protozoan diseases, such as malaria, leishmaniasis, and Chagas disease. In 2021, an estimated 247 million malaria cases were reported [67]. Moreover, leishmaniasis is responsible for causing nearly one million cases every year [68], and it is currently estimated that six to seven million people worldwide are afflicted with Chagas disease [69]. In view of such a profound economic impact, a rapid, effective, and accessible diagnosis is important for a better prognosis [70], and, in this regard, several studies have focused efforts on the development of new diagnostic tests. RMPs has been applied in the experimental diagnoses of infections caused by protoza and Table 3 summarizes the main points of these studies.
Table 3.
RMPs applied in the diagnosis of diseases caused by protozoa
| Disease/causative agent | RMP name | Host for protein expression | Serological assay | Samples | Results | Reference/country |
|---|---|---|---|---|---|---|
| Human chagas disease/Trypanosoma cruzi | ITC 8.2 | E. coli Rosetta2(DE3) pLysS cells | Dipstick assay | 118 Chagas-positive serum samples/106 serum samples from non-Chagas diseases |
Sensitivity: 99.2% Specificity: 99.1% |
[10]/USA |
| Human chagas disease/Trypanosoma cruzi | CP1 and CP2 | E. coli BL21(DE3) cells | ELISA | 141 Chagas-positive serum samples/164 Chagas-negative serum samples/15 cutaneous leishmaniasis-positive serum samples |
CP1 was recognized by positive serum samples (sensitivity and specificity values were not shown) CP2 Sensitivity: 98.6% Specificity: 99.4% |
[71]/Argentina |
| Human toxoplasmosis/Toxoplasma gondii | rMEP | E. coli BL21(DE3) cells | ELISA | 108 toxoplasmosis-positive serum samples/42 serum samples from healthy individuals |
Sensitivity: 94.4–96.9% Specificity: 96.9% |
[72]/China |
| Human toxoplasmosis/Toxoplasma gondii | rMEP | E. coli BL21(λDE3) cells | ELISA | 123 toxoplasmosis-positive serum samples/35 serum samples from healthy individuals |
Acute infection Sensitivity: 96.6% Specificity: 100% Past infection Sensitivity: 96.4% Specificity: 98.7% |
[30]/China |
| Human Chagas disease/Trypanosoma cruzi | CP2 | – | Immunoagglutination assay | 16 Chagas-positive serum samples/16 serum samples from healthy individuals |
Sensitivity: 92% Specificity: 84% |
[73]/Argentina |
| Canine visceral leishmaniasis/Leishmania infantum | PQ10 and PQ20 | E. coli cells | ELISA | 231 L. infantum-positive serum samples/131 healthy dog serum samples |
PQ10 Sensitivity: 88.8% Specificity: 80% PQ20 Sensitivity: 84.9% Specificity: 65% |
[74]/Brazil |
| Human toxoplasmosis/Toxoplasma gondii | USM.TOXO1 | E. coli BL21(DE3) pLysS cells | ELISA | 40 toxoplamosis-positive serum samples/40 serum samples from healthy individuals |
Sensitivity: 100% Specificity: 100% |
[31]/Malaysia |
| Human Chagas disease/Trypanosoma cruzi | TcF43 and TcF26 | E. coli cells | ELISA | 162 Chagas-positive serum samples/36 serum samples from healthy individuals | TcF43 and TcF26 were recognized by positive serum samples | [75]/USA |
| Canine visceral leishmaniasis/Leishmania infantum | PQ10 and PQ20 | E. coli cells | ELISA | 1450 L. infantum-positive serum samples/42 healthy dogs serum samples | PQ10 and PQ20 were recognized by positive serum samples at earlier time point | [76]/Brazil |
| Human toxoplasmosis/Toxoplasma gondii | USM.TOXO1 | E. coli cells | Indirect ELISA | 157 toxoplasmosis-positive serum samples/96 serum samples from healthy individuals/17 serum samples from non-toxoplasmosis diseases |
Sensitivity: 85.43% Specificity: 81.25% |
[77]/Malasya |
| Human Chagas disease/Trypanosoma cruzi | CP3 | E. coli BL21(DE3) cells | ELISA | 67 Chagas-positive serum samples/67 serum samples from healthy individuals |
Sensitivity: 100% Specificity: 90.2% |
[78]/Argentina |
| Canine visceral leishmaniasis/Leishmania infantum | PQ10 and PQ20 | E. coli cells | Chemiluminescent ELISA | 100 L. infantum-positive serum samples/30 healthy dog serum samples/32 serum samples from non-L. infantum diseases |
PQ10 Sensitivity: 93.1% Specificity: 80% PQ20 Sensitivity: 93.1% Specificity: 96.6% |
[79]/Brazil |
| Canine visceral leishmaniasis/Leishmania infantum | PQ10 | E. coli BL21(DE3) cells | ELISA/ Direct agglutination | 50 L. infantum-positive serum samples/50 healthy dog serum samples/30 serum samples from non-L. infantum diseases |
ELISA Sensitivity: 94% Specificity: 86% Direct agglutination PQ10 was recognized by 92% of asymptomatic and 96% of symptomatic infected dogs |
[80]/Iran |
| Human toxoplasmosis/Toxoplasma gondii | pQE30 | E. coli BL21(DE3) cells | Indirect ELISA | 95 toxoplasmosis-positive serum samples/25 serum samples from healthy individuals/6 serum samples from non-toxoplasmosis diseases |
Sensitivity: 72.6% Specificity: 90.3% |
[81]/Iran |
| Pig toxoplasmosis/Toxoplasma gondii | MAG | E. coli BL21(DE3) cells | ELISA | 82 toxoplasmosis-positive serum samples/127 healthy pig serum samples |
Sensitivity: 79.1% Specificity: 88.6% |
[82]/China |
| Canine visceral leishmaniasis/Leishmania infantum | P1P2P3 | E. coli BL21(DE3) cells | ELISA | 50 L. infantum-positive serum samples/50 healthy dog serum samples/14 serum samples from non-L. infantum diseases |
Sensitivity: 98% Specificity: 95.31% |
[83]/Iran |
| Human visceral leishmaniasis/Leishmania infantum | GRP-UBI-HSP | E. coli BL21 cells | ELISA | 30 L. infantum-positive serum samples/15 serum samples from healthy individuals |
Sensitivity: 70.6% Specificity: 84.1% |
[84]/Iran |
| Human visceral leishmaniasis/Leishmania infantum | PQ10 | E. coli BL21(DE3) cells | ELISA | 50 L. infantum-positive serum samples/50 serum samples from healthy individuals/20 serum samples from non-L. infantum diseases |
Sensitivity: 84% Specificity: 82% |
[85]/Iran |
| Human visceral leishmaniasis/Leishmania infantum | MRP | E. coli BL21(DE3) cells | ELISA | 35 L. infantum-positive serum samples/20 serum samples from healthy individuals/10 serum samples from non-L. infantum diseases |
Sensitivity: 93.1% Specificity: 77.4% |
[86]/Iran |
| Human and canine visceral leishmaniasis/Leishmania infantum | rMELEISH | E. coli BL21(DE3) pLysS cells | ELISA | 35 human and 45 dog L. infantum-positive serum samples/30 healthy human and 50 healthy dog serum samples/80 human and 45 dog serum samples from non-L. infantum diseases |
Human and canine diagnosis Sensitivity: 100% Specificity: 100% |
[4]/Brazil |
| Human Chagas disease/Trypanosoma cruzi | rTC | E. coli BL21(λDE3) pLysS cells | ELISA | 58 Chagas-positive serum samples/30 serum samples from healthy individuals/60 serum samples from non-Chagas diseases |
Sensitivity: 98.28% Specificity: 96.67% |
[5]/Brazil |
Houghton et al. (2009) [10] published the first study using RMP for the diagnosis of diseases caused by protozoa. The authors developed a new RMP, ITC 8.2, for human Chagas disease diagnosis, caused by Trypanosoma cruzi. ITC 8.2 was designed from the combination of another RMP, TcF, with immunodominant peptides. After obtaining ITC 8.2 through expression in E. coli Rosetta2(DE3) pLysS cells, reactivity with positive human sera sample was analyzed using a dipstick assay. Results showed sensitivity and specificity values of 99.2% and 99.1%, respectively. Also working with Chagas disease diagnosis, Camussone et al. (2009) [71] developed CP1 and CP2 antigens after performing an epitope junction that had shown promising results in an ELISA assay. These RMPs were obtained through heterologous expression using E. coli BL21(DE3) cells, and antigenicity was assessed during an ELISA assay. Results showed that CP1 and CP2 presented a greater antigenicity as compared to the mix of peptides that comprised each one. Moreover, CP2 showed better performance, with 98.6% sensitivity and 99.4% specificity.
Later, Dai et al. (2012) [72] worked with a new RMP for the diagnosis of human toxoplasmosis, caused by Toxoplasma gondii. Immunodominant epitopes were screened after bioinformatics analyses and selected to form rMEP, which was obtained through expression in E. coli BL21(DE3) cells. An ELISA assay was performed to access rMEP reactivity with human positive sera samples. Sensitivity values ranged from 94.4% to 96.9%, depending on the immunoglobulin class, with a specificity value of 100%. In addition, rMEP performance was superior to that of its constituent epitopes when analyzed separately. Continuing the work of the aforementioned study, Dai et al. (2013) [30] evaluated the rMEP’s capacity to differentiate recent from past toxoplasmosis infections. In their study, rMEP was also obtained through expression in E. coli BL21(DE3) cells and an in-house ELISA was performed. Results showed that rMEP could be used to differentiate acute from past infection, with sensitivity and specificity values ranging from 96.4% to 96.6% and 98.7% to 100%, respectively.
Garcia et al. (2013) [73] worked with an RMP, named CP2, aimed at diagnosing Chagas disease. This protein had been previously tested, and, in their study, the authors produced a latex-protein complex to be tested in an immunoagglutination assay. Sensitivity and specificity values were determined as 92% and 84%, respectively. In addition, CP2 was more efficient in discriminating between positive and negative serum samples as compared to single recombinant proteins. Next, Faria et al. (2015) [74] developed two new RMPs for the diagnosis of canine visceral leishmaniasis, caused by Leishmania infantum. Epitopes were selected based on the literature to form PQ10 and PQ20, and these antigens were obtained after heterologous expression in E. coli cells. An ELISA assay was performed to assess RMP reactivity with canine sera samples, where sensitivity values were determined as 88.8% and 84.9% for PQ10 and PQ20, respectively. These values were higher as compared to sensitivity values from DPP (Bio-Manguinhos/Fiocruz; Rio de Janeiro, Brazil) and EIE-LVC kit (Bio-Manguinhos/Fiocruz; Rio de Janeiro, Brazil) commercial tests, since sensitivity values for these commercial kits were calculated as 72.9% and 64.5%, respectively. However, PQ10 and PQ20 specificity values were lower compared to commercial tests, showing an 80% and 65% value, respectively, while DPP and EIE-LVC kit showed 90% and 100% specificity, respectively. Next, Hajissa et al. (2015) [31] produced a new RMP for diagnosing human toxoplasmosis infection. The authors selected epitopes based on bioinformatics analyses, and the new RMP antigen, USM.TOXO1, was obtained by heterologous expression in E. coli BL21(DE3) pLysS cells. Initially, a Western blotting was performed to verify USM.TOXO1 reactivity with positive serum samples, resulting in recognition by positive human serum samples. The reactivity was further confirmed by ELISA assay, with 100% sensitivity and specificity values.
Duthie et al. (2016) [75] developed two new RMPs, TcF43 and TcF26, for the purpose of diagnosing human Chagas disease. These proteins were expressed in E. coli cells and evaluated through an ELISA assay. Results showed that TcF43 and TcF26 proteins increased serum recognition as compared to TcF, an antigen used in commercial kits. However, sensitivity and specificity values were not provided. Faria et al. (2017) [76] continued the studies with the RMPs PQ10 and PQ20, previously cited, aimed at diagnosing canine visceral leishmaniasis. After heterologous expression in E. coli cells, an ELISA assay was performed to assess the antigens’ capacity to detect the disease at early stages. When compared to ELISA based on crude antigens, PQ10, and, especially, PQ20 were able to detect the infection at earlier time points. In addition, these recombinant antigens demonstrated high result concordances in relation to real-time PCR. Hajissa et al. (2017) [77] continued the work developed by Hajissa et al. (2015) [31], employing USM.TOXO1 for diagnosing human toxoplasmosis infection. USM.TOXO1 was obtained using E. coli cells, and, after an ELISA assay with human sera samples, sensitivity and specificity values were determined as 85.43% and 81.25%, respectively. With the objective of developing a new RMP for diagnosing human Chagas disease, Peverengo et al. (2018) [78] produced CP3 after selecting epitopes based on the published literature. CP3 was obtained through heterologous expression in E. coli BL21(DE3) cells, after which an ELISA assay was performed to assess protein antigenicity. Results showed 100% sensitivity and 90.2% specificity values.
In 2019, another study was performed using PQ10 and PQ20 to diagnose canine visceral leishmaniasis. After obtaining the proteins through expression in E. coli cells, Fonseca et al. (2019) [79] performed a chemiluminescent ELISA to evaluate the antigens’ reactivity with canine serum samples. PQ10 sensitivity and specificity values were determined as 93.1% and 80.0% respectively, while PQ20 showed 93.1% sensitivity and 96.6% specificity. Both PQ10 and PQ20 demonstrated better diagnostic performance as compared to crude antigen diagnostics results. PQ10 was also tested in a study conducted by Jameie et al. (2020) [80]. Following expression in E. coli BL21(DE3) cells, an ELISA assay was performed to evaluate protein reactivity with canine visceral leishmaniasis serum samples. Results showed sensitivity and specificity values of 94% and 86%, respectively. Moreover, a direct agglutination test assay was performed, in which PQ10 was able to detect 92% of asymptomatic and 96% of symptomatic infected dogs.
Alibakhshi et al. (2020) [81] developed a new RMP, pQE30, for diagnosing human toxoplasmosis. To construct pQE30, epitopes were selected using bioinformatics analyses and E. coli BL21(DE3) cells were used to obtain the recombinant antigen. An ELISA assay was then performed, with sensitivity and specificity values of 72.6% and 90.3%, respectively. While conducting a study aimed at diagnosing animal toxoplasmosis, caused by T. gondii, Song et al. (2021) [82] selected epitopes after performing bioinformatics analyses to form a new RMP, called MAG, which was expressed in E. coli BL21(DE3) cells. A Western blotting assay was then performed, with positive pig sera samples recognizing MAG. Reactivity was further confirmed by an ELISA assay, showing 79.1% sensitivity and 88.6% specificity.
Yaghoubi et al.. (2021) [83] conducted a study to construct a new RMP for canine visceral leishmaniasis diagnosis. Bioinformatics analyses were used to select epitopes for the new RMP development, named P1P2P3. This antigen was obtained after expression in E. coli BL21(DE3) cells, and an ELISA assay was performed to access antigen reactivity. Results showed a 98% sensitivity and 95.31% specificity, demonstrating agreement with the direct agglutination test, the gold standard test used in the study. Working with human visceral leishmaniasis, caused by L. infantum, Heidari et al. (2021) [84] developed a new RMP, called GRP-UBI-HSP, to be tested in ELISA assay. In their study, epitopes were selected from immunoreactive proteins through bioinformatic analyses, and GRP-UBI-HSP was obtained after expression in E. coli BL21 cells. The results of an ELISA assay showed 70.6% sensitivity and 84.1% specificity. A subsequent study, performed by Jameie et al. (2021) [85], aimed to verify PQ10’s diagnostic capacity for human visceral leishmaniasis, caused by L. infantum. PQ10 had already been tested in several studies for the diagnosis of canine visceral leishmaniasis, with promising results. In their study, the antigen was obtained through heterologous expression in E. coli BL21(DE3) cells, and an ELISA assay was performed to evaluate PQ10 antigenicity. Results showed a diagnostic performance of 84% sensitivity and 82% specificity.
Subsequently, Taherzadeh et al. (2021) [86] developed a new RMP for human visceral leishmaniasis diagnosis caused by L. infantum. Epitopes were selected based on bioinformatics analyses, and, after designing the recombinant antigen named MRP, the heterologous antigen was expressed using E. coli BL21(DE3) cells. Initially, MRP recognition by human positive serum samples was confirmed through Western blotting analyses. Results derived from an ELISA assay demonstrated 93.1% sensitivity and 77.4% specificity. Next, Dias et al. (2023) [4] conducted a study to evaluate the diagnostic efficiency of a new RMP, rMELEISH, for both human and canine visceral leishmaniasis, caused by L. infantum. After selecting epitopes based on the literature, rMELEISH was obtained through expression in E. coli BL21(DE3) pLysS cells and ELISA assays showed 100% sensitivity and specificity values. The same results were observed for canine sera sample reactivity. Moreover, rMELEISH demonstrated a better diagnostic performance as compared to results using soluble Leishmania antigen extract. Lastly, a novel RMP for diagnosing Chagas disease was developed by Machado et al. (2023) [5]. To compose rTC antigen, the epitopes were selected based on the literature. E. coli BL21(DE3) pLysS cells were used to obtain rTC, and an ELISA assay was performed. Results showed sensitivity and specificity values of 98.28% and 96.67%, respectively.
RMPs applied in the diagnosis of viral diseases
Among the species, 219 viral species are recognized as human pathogens [87, 88], and several species also affect animals [89, 90]. Viral infections represent a major public health problem and humanity has faced several lethal viral pandemics, such as Spanish flu and COVID-19, causing the death of millions of people worldwide [91]. Moreover, viral infections also cause thousands of deaths annually without being related to endemics or pandemics [92, 93]. Given its importance, much effort has been applied to the development of viral disease diagnoses, which are currently performed through several methods. Among them, the serological method is widely applied due to its high sensitivity and specificity, low cost, and rapid diagnosis [94, 95]. RMPs has been applied in the experimental diagnoses of infections caused by virus and Table 4 summarizes the main points of these studies.
Table 4.
RMPs applied in the diagnosis of viral diseases
| Disease/causative agent | RMP name | Host for protein expression | Serological assay | Samples | Results | Reference/country |
|---|---|---|---|---|---|---|
| Human dengue fever/dengue virus | r-DME-G | E. coli strain SG13009 cells | ELISA | 10 dengue-positive serum samples/10 serum samples from healthy individuals | Sensitivity: 100% Specificity: not provided | [9]/India |
| Human dengue fever/dengue virus | r-DME-M | E. coli cells | ELISA | 22 dengue-positive serum samples/150 dengue-negative serum samples | r-DME-M was recognized by all positive human samples | [96]/India |
| Human hepatitis C/hepatitis C virus | r-HCV-F-MEP | E. coli BL21(DE3) cells | ELISA/Lateral flow assay | > 200 hepatitis-positive serum samples/100 serum samples from healthy individuals |
ELISA Sensitivity: 99.8% Specificity: 100% Lateral flow 100% compatible with ELISA results |
[1]/India |
| Human dengue fever/dengue virus | rDME-M | E. coli cells | Dipstick ELISA | 81 dengue-positive serum samples/39 serum samples from healthy individuals/30 serum samples from non-dengue diseases | Sensitivity: 100% Specificity: 85% and 93% | [97]/India |
| Human dengue fever/dengue virus | rDME-G | E. coli cells | Dipstick ELISA | 50 dengue-positive serum samples/10 serum samples from healthy individuals |
Sensitivity: 100% and 95% Specificity: 100% |
[98]/India |
| Human HIV/human immunodeficincy virus | HIV-MEP | E. coli BL21(DE3) cells | Indirect ELISA | 57 HIV-positive serum samples/50 HIV-negative serum samples | High sensitivity and specificity values were observed | [99]/India |
| Human hepatitis C/hepatitis C virus | HCV | E. coli cells | Double-antigen sandwich ELISA | 259 hepatitis-positive serum samples/440 serum samples from healthy individuals/190 serum samples from non-hepatits C diseases | Sensitivity: 98.7% Specificity: 100% | [100]/China |
| Human hepatitis C/hepatitis C virus | rHCV-MEP | E. coli BL21(DE3) cells | ELISA | 56 hepatitis-positive serum samples/50 hepatitis C-negative serum samples | High sensitivity and specificity values were observed | [101]/India |
| Human hepatitis B/hepatitis B virus | rMEHB | E. coli BL21(DE3) pLysS cells | ELISA | – | rMEHB was recognized by human positive samples | [19]/Brazil |
| EBV-associated tumors/epstein-barr virus | EBV-LMP2m | E. coli BL21(DE3) cells | ELISA | 238 Epstein-Barr-positive serum samples/112 serum samples from healthy individuals | Sensitivity: 52.84% Specificity: 95.40% | [102]/China |
| Human hepatitis A/hepatitis A virus | H1 | E. coli BL21(DE3) cells | Double-antigen sandwich ELISA | 48 hepatitis-positive serum samples/96 serum samples from vaccinated individuals | Sensitivity: 93.75% Specificity: 93.75% | [103]/China |
| Human hepatitis C/hepatitis C virus | rMEHCV | E. coli BL21(λDE3) pLysS cells | ELISA | 17 hepatitis-positive serum samples/10 serum samples from healthy individuals | High sensitivity and specificity values were observed | [2]/Brazil |
| Human hepatitis C/hepatitis C virus | r-HCV-MEPs | E. coli BL21(DE3) cells | Secondary antibody/ Double-antigen immunoassays | 113 hepatitis-positive serum samples/58 hepatitis-negative serum samples |
Secondary antibody Sensitivity: 95.6% Specificity: 100% Double-antigen assay Sensitivity: 91.4% Specificity: 100% |
[104]/Finland |
| Swine foot-and-mouth disease/foot-and-mouth disease virus | B4 | E. coli cells | Indirect ELISA | 147 foot-and-mouth-positive serum samples/610 healthy swine serum samples | Sensitivity: 95.9% Specificity: 96.7% | [105]/China |
| Human hepatitis C/hepatitis C virus | Recombinant multiepitope HCV antigen | E. coli cells | ELISA | 88 hepatitis-positive serum samples/376 hepatitis C-negative serum samples/18 serum samples from non-hepatitis C diseases | Sensitivity: 100% Specificity: 99.73% | [106]/Brazil |
| Human cytomegalovirus/cytomegalovirus | rMEHCMV | E. coli BL21(DE3) cells | ELISA | 12 cytomegalovirus-positive serum samples/1 serum sample from healthy individual | rMEHCMV was recognized by human positive samples | [107]/Brazil |
| Canine coronavirus/SARS-CoV-2 | rSP | E. coli BL21(DE3) cells | Indirect ELISA | 64 coronavirus-positive serum samples/10 healthy dog serum samples | 82.81% positive rate | [108]/China |
| Human coronavirus/SARS-CoV-2 | Dx-SARS2-RBD and Dx-SARS2-noRBD | E. coli BL21(DE3) cells | ELISA | 185 coronavirus-positive serum samples/256 serum samples from healthy individuals/94 serum samples from non-coronovirus diseases |
Dx-SARS2-RBD Sensitivity: 100% Specificity: 99.51-100% Dx-SARS2-noRBD Sensitivity: 100% Specificity: 99.21-100% |
[109]/Brazil |
| African swine fever/African swine fever virus | reMeP72 | E. coli BL21(DE3) cells | Colloidal gold-based immunochromatographic assay | 139 swine serum samples | Sensitivity: 85.7% Specificity: 97.6% | [110]/China |
| African swine fever/African swine fever virus | m35 | E. coli BL21 cells | Indirect ELISA | 78 positive serum samples/215 negative serum samples | Sensitivity: 98.72% Specificity: 98.14% | [111]/China |
| Human mayaro fever/mayaro virus | Dx-MAYV-M | E. coli BL21(DE3) cells | ELISA | 18 mayaro-positive serum samples/40 serum samples from healthy individuals | Sensitivity: 99.6% Specificity: 100% | [6]/Brazil |
| Swine foot-and-mouth/foot-and-mouth disease virus | ME protein | E. coli BL21(DE3) cells | Indirect chemiluminescence immunoassay | 118 foot-and-mouth-positive serum samples/307 healthy swine serum samples | Sensitivity: 100% Specificity: 99.35% | [112]/China |
| Human hepatitis C/hepatitis C virus | MBP-rHCV | E. coli BL21(DE3) cells | Indirect ELISA | 142 hepatitis-positive serum samples/172 serum samples from healthy individuals | Sensitivity: 95% Specificity: 92% | [113]/Canada |
| Human rubella/rubella virus | rMERUB | E. coli BL21(λDE3) cells | ELISA | 22 rubella-positive serum samples/11 rubella-negative serum samples | Sensitivity: 100% Specificity: 90.91% | [114]/Brazil |
| Human HTLV-1 and HTLV-2/human T-lymphotropic viruses 1 and 2 | HTLV-1/HTLV-2 | E. coli Rosetta-gami 2(DE3) cells | ELISA | 162 HTLV-positive serum samples/297 serum samples from healthy individuals/92 serum samples from non-HTLV disease |
HTLV-1 and HTLV-2 samples Sensitivity: 82.41 to 92.36% Specificity: 90.09% to 95.19% HTLV-1 samples Sensitivity: 99.19% Specificity: 92.55% HTLV-2 samples Sensitivity: 57.14% Specificity: 94.61% |
[115]/Brazil |
| Human chikungunya/chikungunya virus | MULTREC | Binary system insect cell/baculovirus | ELISA | 161 chikungunya-positive serum samples/22 serum samples from healthy individuals/312 serum sample from non-chikungunya diseases | Sensitivity: 86.36% Specificity: 100% | [116]/Brazil |
“–”: information not provided by the study
Ananda Rao et al. (2005) [9] published the first study describing the use of an RMP for the diagnosis of a human viral infection. In their study, the epitopes were selected through phage display, pepscan, and computer analysis of three selected proteins of the dengue virus, responsible for causing dengue fever. The epitopes were combined to form a new RMP, identified as r-DME-G, which was obtained in E. coli strain SG13009 cells. An in-house ELISA was performed, resulting in 100% sensitivity. Specificity data were not provided. Next, Ananda Rao et al. (2006) [96] designed a new RMP, identified as r-DME-M, which was also applied to human dengue infection diagnosis. The epitopes were selected using the same approach as described in the 2005 study, and r-DME-M was obtained using E. coli cells. After performing an in-house ELISA, it was observed that r-DME-M was capable of detecting all seropositive human samples, demonstrating better performance than the commercial test PanBio (Pty Ltd.; Windsor, Australia), used for comparison. However, sensitivity and specificity values were not available.
Subsequently, Dipti et al. (2006) [1] developed a new RMP for human hepatitis C diagnosis. The epitopes were selected based on the literature, forming a new RMP, r-HCV-F-MEP, which was expressed in E. coli BL21(DE3) cells. The in-house ELISA showed a 99.8% sensitivity and 100% specificity. Moreover, a lateral flow assay was performed, and the results were fully compatible with those obtained in the in-house ELISA. Next, Tripathi et al. (2007) [97] developed a new RMP for human dengue diagnosis, identified as rDME-M, obtained through expression in E. coli cells. Initially, rDME-M reactivity was verified by Western blotting, in which a seropositive human sample recognized rDME-M. An in-house dipstick ELISA was then performed, with 93% and 85% specificity as compared to reference available ELISA and rapid immunochromatography tests, respectively. Moreover, the sensitivity of the in-house dipstick ELISA was calculated as 100% as compared to both reference tests. Tripathi et al. (2007) [98] also worked with a new RMP for diagnosing human dengue infection. In this study, the epitopes were selected based on phage display and computer predictions to form a new RMP, identified as rDME-G. After analyzing the reactivity of rDME-G with human sera in an in-house dipstick ELISA, the sensitivity was 100% and 95% as compared to reference available ELISA and rapid immunochromatography tests, respectively. Moreover, the specificity value was 100% as compared to both reference tests.
Talha et al. (2010) [99] developed HIV-MEP for the diagnosis of HIV infection. In their study, the authors selected the epitopes based on the literature, and a new RMP formed was obtained after expression in E. coli strain BL21 (DE3) cells. To assess HIV-MEP reactivity, an in-house indirect immunoassay was performed, and the results were similar to those obtained using Abbott HIV-1, Abbott HIV-1/2, Genetic Systems HIV-1, Genetic Systems HIV-1/2, and Organon Teknika HIV-1 commercial tests. Despite having high sensitivity and specificity, their values were not provided. Next, He et al. (2011) [100] constructed a new RMP for diagnosing human hepatitis C. After obtaining the new recombinant multiepitope HCV antigen through expression in E. coli cells, a double-antigen sandwich ELISA was performed to verify antigen reactivity. The results were similar to those obtained with Ortho ELISA 3.0 (GWK; Beijing, China) commercial test, where the sensitivity and specificity of both developed tests and commercial kit were 98.8% and 100%, respectively. Also working with the diagnosis of hepatitis C, Gurramkonda et al. (2012) [101] developed a new RMP, designated as rHCV-MEP. To compose this new RMP, epitopes were selected based on the literature, and rHCV-MEP was obtained using E. coli BL21(DE3) cells. An in-house ELISA assay was applied to evaluate performance, and a high sensitivity and specificity was observed, with results compatible with those obtained using Abbott HCV 2.0, Abbott HCV 3.0, Ortho HCV 2.0, and Ortho HCV 3.0 commercial tests. However, sensitivity and specificity values were not provided.
Subsequently, de Souza et al. (2013) [19] worked with a new RMP, identified as rMEHB, for the diagnosis of human hepatitis B. Conserved epitopes were selected and E. coli BL21(DE3) pLysS cells were used to obtain it. In their study, the TIEBK PLUS No 140—Diasorin commercial kit was used, with the commercial antigen being replaced by rMEHB. The results showed that rMEHB was recognized by antibodies in human positive samples and its performance was similar to that of the commercial test. Next, Lin et al. (2016) [102] developed a new RMP for the diagnosis of human Epstein-Barr virus-associated tumors. For epitope selection, bioinformatics analyses were conducted, and the new antigen was named EBV-LMP2m. Western blotting was used to verify EBV-LMP2m reactivity and confirm recognition by positive human serum. Moreover, the EBV-LMP2m performance was evaluated in an in-house ELISA, resulting in 52.84% sensitivity and 95.40% specificity. Su et al. (2016) [103] designed a new RMP aimed at diagnosing human hepatitis A. To form this new RMP, identified as H1, the authors selected immune-dominant epitopes based on previous studies. After obtaining H1 through expression in E. coli BL21(DE3), a double-antigen sandwich ELISA was performed to assess serum-RMP reactivity, with sensitivity and specificity values of 93.75%.
Galdino et al. (2016) [2] developed a new RMP for the diagnosis of human hepatitis C. In that study, the epitopes were selected based on the literature and the new RMP, rMEHCV, was obtained through expression in E. coli BL21(DE3) pLysS cells. Results of an in-house ELISA showed 100% agreement with those of the Hepanóstika HCV Ultra® (Beijing, China) commercial test, with high sensitivity and specificity values. However, these values were not provided. Also working with hepatitis C diagnosis, Salminen et al. (2016) [104] developed the antigen named r-HCV-MEPs, after epitope selection based on the literature. E. coli BL21(DE3) cells were used for heterologous protein expression. To assess the r-HCV-MEPs reactivity with positive serum samples, a secondary antibody and double-antigen immunoassays were used, resulting in sensitivity values of 95.6% and 91.4%, respectively. Moreover, specificity values were 100% in both immunoassays. Cao et al. (2018) [105] worked with a RMP, designated as B4, for the diagnosis of animal foot-and-mouth disease. B4 had already been developed in a previous study. After expression in E. coli cells, an indirect ELISA was performed using swine serum samples, and the results showed 95.9% sensitivity and 96.7% specificity values.
Subsequently, Thomasini et al. (2018) [106] developed a new RMP for diagnosing human hepatitis C. The epitopes were selected based on previously published studies and the new RMP was obtained after expression in E. coli cells. Initially, an immunoblot assay was performed to assess RMP reactivity, where a strong reaction with positive human samples was observed. After performing an ELISA assay, sensitivity and specificity values were defined as 100% and 99.73%, respectively. Later, Ribeiro et al. (2019) [107] worked with a new RMP, rMEHCMV, for the diagnosis of human cytomegalovirus infection. The authors selected conserved epitopes to form rMEHCMV, and E. coli BL21(DE3) cells were chosen for the heterologous expression. An in-house ELISA assay was performed, and results showed that rMEHCMV was recognized by human-infected samples, demonstrating a stronger reactivity as compared to those of the ETI-CYTOK-G PLUS (DiaSorin; Saluggia, Italy) commercial kit. However, sensitivity and specificity values were not provided. Hao et al. (2021) [108] developed a new RMP, rSP, aimed at diagnosing canine coronavirus diagnosis, caused by SARS-CoV-2. Epitopes were selected through bioinformatics analyses, and, after designing the new RMP, E. coli BL21(DE3) cells were used for heterologous protein expression. An indirect ELISA assay was performed, where the authors observed good sensitivity and specificity results. However, those values were not provided.
Also working with SARS-CoV-2 diagnosis, Gomes et al. (2021) [109] developed two new RMPs for the diagnosis of human coronavirus. Epitope selection was made through direct microsynthesis of phosphopeptides on membranes synthesis to form Dx-SARS2-RBD and Dx-SARS2-noRBD and both antigens were expressed using E. coli BL21(DE3) cells. An in-house ELISA assay was performed to assess reactivity and the results showed 100% sensitivity for both RMPs. Specificity values ranged from 99.51–100% for Dx-SARS2-RBD and 99.21–100% for Dx-SARS2-noRBD. Zhang et al. (2021) [110] worked with a new RMP for the diagnosis of African swine fever. The new antigen, designated reMeP72, was constructed based on epitopes selected by bioinformatics analyses. After obtaining the new antigen using E. coli BL21(DE3) cells, a colloidal gold-based immunochromatographic assay was performed. Results indicated 85.7% sensitivity and 97.6% specificity, showing an agreement rate of 96.4% with the ASFV indirect ELISA kit (INGENASA; Madrid, Spain) commercial kit. Also working with the diagnosis of African swine fever in animals, Gao et al. (2021) [111] developed a new RMP, m35, after selecting epitopes through bioinformatics analyses. The m35 protein was obtained after expression in E. coli BL21 cells, and an indirect ELISA assay was performed to verify reactivity with swine serum samples. Sensitivity and specificity values were defined as 98.72% and 98.14%, respectively.
In the same year, Napoleão‑Pêgo et al. (2021) [6] developed Dx-MAYV-M, a new RMP to be tested in human Mayaro fever diagnosis. After epitope mapping, selected epitopes were joined together and the recently formed Dx-MAYV-M was obtained using E. coli BL21(DE3) cells. Initially, Western blotting was performed to verify reactivity, in which antigen recognition by positive human sera was observed. An in-house ELISA assay was performed, resulting in estimated sensitivity and specificity values of 99.6% and 100%, respectively. Next, a new RMP for the diagnosis of animal foot-and-mouth disease was developed by Liu et al. (2021) [112]. In their work, the authors expressed this new RMP, identified as ME protein, using E. coli BL21(DE3) cells, and performed an indirect chemiluminescence immunoassay to evaluate ME reactivity with swine serum samples. Results showed 100% sensitivity and 99.35% specificity. Later, Pedersen et al. (2022) [113] designed a new RMP for hepatitis C diagnosis, labeled MBP-rHCV. To construct this protein, epitopes were selected based on published studies and E. coli BL21(DE3) cells were used for heterologous expression. An indirect ELISA assay was performed to determine MBP-rHCV reactivity with human sera samples, resulting in 95% sensitivity and 92% specificity.
Souza et al. (2022) [114] created a new RMP, named rMERUB, for the diagnosis of human rubella. Conserved epitopes were selected to construct rMERUB, which was obtained through expression in E. coli BL21(DE3) cells. After performing an in-house ELISA assay, sensitivity and specificity values were defined as 100% and 90.91%, respectively. Franco et al. (2022) [115] worked with a new RMP for human HTLV-1 and HTLV-2 infections diagnosis, caused by human T-lymphotropic viruses 1 and 2. The HTLV-1/HTLV-2 multiepitope protein was constructed based on previous studies and obtained through expression in E. coli Rosetta-gami 2 (DE3) cells. A Western blot was then performed, confirming multiepitope protein recognition by positive HTV-1 and HTV-2 serum samples. After an in-house ELISA assay, sensitivity and specificity values ranged from 82.41 to 92.36% and 90.09 to 95.19%, respectively, when considering positive samples for both HTLV1- and HTLV-2. When considering only HTLV-1 samples, sensitivity and specificity values were calculated as 99.19% and 92.55%, respectively. Considering the values when evaluating only HTLV-2 samples, sensitivity and specificity were determined as 57.14% and 94.61%, respectively. Lastly, da Silva et al. (2022) [116] developed a new RMP for diagnosing human chikungunya, caused by the chikungunya virus. This new RMP, MULTREC, was obtained through a binary system insect cell/baculovirus, after which an ELISA assay was performed to assess protein reactivity, with 86.36% sensitivity and 100% specificity.
RMPs applied in the diagnosis of diseases caused by worms
Infections caused by worms, also known as helminths, are one the most common diseases in the world, with estimates of approximately 1.5 billion people infected worldwide [117]. This group of diseases mainly affects people living in the world’s poorest countries and is associated with severe morbidity [118]. However, despite of the risk that these diseases pose to human and animal lives, they remain poorly studied compared to other disease groups [119]. If this scenario is to change, more efforts must be applied in scientific research, including the development of new diagnostic kits for helminths [120]. RMPs has been applied in the experimental diagnoses of infections caused by worms and Table 5 summarizes the main points of these studies.
Table 5.
RMPs applied in the diagnosis of diseases caused by worms
| Disease/causative agent | RMP name | Host for protein expression | Serological assay | Samples | Results | Reference/country |
|---|---|---|---|---|---|---|
| Goat schistosomiasis/Schistosoma japonicum | rBSjPGM-BSjRAD23-1-BSj23, rBSjRAD23-2-BSjPGM-BSj23, rBSjPGM-BSj23, and rBSjPGM-BSjRAD23-1 | E. coli cells | ELISA | 91 schistosomiasis-positive serum samples/44 healthy goat serum samples/49 serum samples from non-schistosomiasis diseases |
rBSjPGM-BSjRAD23-1-BSj23 Sensitivity: 97.8% Specificity: 100% rBSjRAD23-2-BSjPGM-BSj23 Sensitivity: 89.01% Specificity: 100% rBSjPGM-BSj23 Sensitivity: 93.41% Specificity: 100% rBSjPGM-BSjRAD23-1 Sensitivity: 59.34% Specificity: 97.73% |
[32]/China |
| Buffalo schistosomiasis/Schistosoma japonicum | rBSjPGM-BSjRAD23-1-BSj23 and rBSjRAD23-2-BSjPGM-BSj23 | E. coli cells | ELISA | 114 schistosomiasis-positive serum samples/92 healthy buffalo serum samples/14 serum samples from non-schistosomiasis diseases |
rBSjPGM-BSjRAD23-1-BSj23 Sensitivity: 95.61% Specificity: 97.83% rBSjRAD23-2-BSjPGM-BSj23 Sensitivity: 67.54% Specificity: 100% |
[121]/China |
| Bovine cysticercosis/Taenia saginata | rqTSA-25 | E. coli BL21-Codon-Plus(DE3)-RIL cells | ELISA/immunoblot assay | 60 cysticercosis-positive serum samples/30 healthy bovine serum samples/15 serum samples from non-cysticercosis diseases |
ELISA Sensitivity: 93.3% Specificity: 95.3% Immunoblot no false positive or false negative was observed |
[122]/Brazil |
| Sheep cystic echinococcosis/Echinococcus granulosus | reEg mefAg-1 | E. coli BL21(DE3) cells | Indirect ELISA | 86 echinococcosis-positive serum samples/30 echinococcosis-negative serum samples/21 serum samples from non- Echinococcus granulosus diseases | Sensitivity: 93.41% Specificity: 99.31% | [123]/China |
| Human onchocerciasis/Onchocerca volvulus | OvNMP-48 | – | ELISA | 101 onchocerciasis-positive serum samples/58 serum samples from healthy individuals/54 serum samples from non-onchocerciasis diseases | Sensitivity: 76.0% Specificity: 97.4% | [124]/Belgium |
| Human fascioliasis/Fasciola hepatica | rMEP | E. coli BL21 cells | Western blot | – | rMEP was recognized by human positive samples | [125]/Iran |
| Human lymphatic filariasis/Wuchereria bancrofti | Multiepitope antigen | E. coli Rosetta cells | Indirect ELISA | 70 lymphatic filariasis-positive serum samples/176 serum samples from healthy individuals/18 serum samples from non-lymphatic filariasis diseases | Sensitivity: 100% Specificity: 98.1% and 99.52% | [126]/India |
| Human cystic echinococcosis/Echinococcus granulosus | rMEP | E. coli BL21(DE3) cells | ELISA | 43 echinococcosis-positive serum samples/120 serum samples from healthy individuals/23 serum samples from non-echinococcosis diseases | Sensitivity: 95.3% Specificity: 95.0% | [127]/Iran |
| Human cystic echinococcosis/Echinococcus granulosus | DIPOL | E. coli cells | ELISA | 149 echinococcosis-positive serum samples/21 serum samples from healthy individuals/49 serum samples from non-echinococcosis diseases | Sensitivity: 75.4% for active and transitional cysts and 95.6% for inactive cysts Specificity: 97.71% | [128]/Turkey |
| Human onchocerciasis/Onchocerca volvulus | OvMCBL02 | – | Indirect ELISA |
63 onchocerciasis-positive serum samples/55 serum sample from healthy individuals/21 serum samples from non-onchocerciasis diseases / 54 onchocerciasis-negative serum samples from ivermectin treated individuals |
Sensitivity: 98.4% Specificity: 100% | [129]/Belgium |
“–”: information not provided by the study
Despite its importance, RMP application in the diagnosis of diseases caused by worms is recent. Lv et al. (2016) [32] evaluated the performance of four RMP molecules for diagnosing goat schistosomiasis, caused by Schistosoma japonicum. These new RMPs were designed using epitopes selected through bioinformatics analyses, and E. coli cells were used for heterologous expression. RMPs showed greater sensitivity as compared to single molecular recombinant antigens in ELISA assays, with emphasis on the rBSjPGM-BSjRAD23-1-BSj23 antigen, which showed 97.8% sensitivity and 100% specificity, as compared to soluble egg antigen. Continuing the studies with the RMPs cited above, Lv et al. (2018) [121] evaluated the diagnostic capacity of two RMPs for buffalo schistosomiasis caused by S. japonicum. Similar to the findings in the previous study, the rBSjPGM-BSjRAD23-1-BSj23 antigen showed the best performance in an ELISA assay, with sensitivity and specificity rates of 95.61% and 97.83%, respectively, again as compared to the soluble egg antigen results. Next, Guimarães-Peixoto et al. (2018) [122] conducted a study to develop a new RMP for diagnosing bovine cysticercosis, caused by Taenia saginata. For this purpose, bioinformatics analyses were used to select epitopes, and the new RMP, identified as rqTSA-25, was produced in E. coli BL21-Codon-Plus(DE3)-RIL cells. After an ELISA assay, rqTSA-25 showed 93.3% sensitivity and 95.3% specificity values. Furthermore, no false positive or false negative reaction was observed in the samples analyzed by the immunoblot test.
Tianli et al. (2019) [123] developed a new RMP for diagnosing sheep cystic echinococcosis, caused by Echinococcus granulosus. After conducting bioinformatics analyses for selecting epitopes, a new RMP was designed, named reEg mefAg-1. E. coli BL21(DE3) cells were used for antigen production, and an indirect ELISA assay was performed to assess reEg mefAg-1’s reactivity. Results showed 93.41% sensitivity and 99.31% specificity. Moreover, reEg mefAg-1-based ELISA results were similar to those found in the IgG ELISA Kit (ab108733, Abcam; Cambridge, Massachusetts, USA) commercial kit. In that same year, Lagatie et al. (2019) [124] conducted a study aimed at diagnosing human onchocerciasis, caused by Onchocerca volvulus. For this purpose, a new RMP, OvNMP-48, was constructed based on epitopes selected through proteome-wide screen as performed in previous studies. After an ELISA assay, sensitivity and specificity values were determined as 76.0% and 97.4%, respectively. Moreover, OvNMP-48-ELISA showed greater sensitivity values as compared to epitope-based ELISA. However, an OvNMP-48-based ELISA assay also demonstrated an increase in cross-reactions. Subsequently, Aghamolaei et al. (2020) [125] developed a new RMP for diagnosing human fascioliasis, caused by Fasciola hepatica. In their study, bioinformatics analyses were applied to select epitopes and a new RMP, designated as rMEP, was obtained after heterologous expression in E. coli BL21 cells. A Western blot was performed to assess rMEP’s reactivity with positive human serum samples. A strong band was observed using human positive serum samples, but no band was observed in poled serum with other helminths and healthy individuals. Despite the promising results, no further serological tests were performed.
Yasin et al. (2020) [126] conducted a study to develop a new RMP for diagnosing human lymphatic filariasis, caused by Wuchereria bancrofti. To form the new RMP, identified as an multiepitope antigen, epitopes were selected based on previous bioinformatics studies. The multiepitope antigen was obtained using E. coli Rosetta cells, and an ELISA assay was performed to assess the antigen’s reactivity with human-positive serum samples. Results showed 100% sensitivity, and a specificity range from 98.1 to 99.52%, depending on the antibody subclass detected. Mirzapour et al. (2020) [127] developed a new RMP aimed at diagnosing human cystic echinococcosis, caused by Echinococcus granulosus. Epitopes were selected through bioinformatics analyses, and, after designing the new RMP, named rMEP, E. coli BL21(DE3) cells were used for heterologous protein expression. ELISA assay results showed high sensitivity and specificity values, determined as 95.3% and 95.0%, respectively. However, the Euroimmun commercial kit showed better performance, with 100% for both sensitivity and specificity. More recently, Ozturk et al. (2022) [128], also working on diagnosing human cystic echinococcosis, tested an RMP, named as DIPOL. In their study, the authors obtained DIPOL through expression in E. coli cells and performed an ELISA assay to verify antigen reactivity. The DIPOL-based ELISA test showed sensitivity values of 75.4% for active and transitional cysts and 95.6% for inactive cysts. Moreover, specificity values were determined as 97.71%. Lastly, Yengo et al. (2022) [129] developed a new RMP for diagnosing human onchocerciasis. Epitopes were selected through bioinformatics analyses to form the new RMP, OvMCBL02. After performing an indirect ELISA assay, sensitivity and specificity values were determined as 98.4% and 100%, respectively.
Escherichia coli: the platform of choice for RMP production
Although various host cells described in the literature can serve as expression systems for recombinant protein production, almost all diagnostic RMP studies to date have used E. coli as the expression system of choice (Tables 1–5). What makes this host so well-suited for this purpose? This expression system boasts a long history and offers several well-established advantages, including ease of manipulation, low-cost culture, and rapid growth kinetics. The doubling time of 20 min facilitates the achievement of high cell density cultures, with a theoretical concentration limit of ~ 1 × 1013 viable bacteria/mL [26, 130]. Furthermore, E. coli stands out as the most cost-effective host, allowing for the attainment of high cellular densities with inexpensive culture media. Additionally, well-developed tools for molecular manipulations, coupled with in-depth knowledge of its biology [131], contribute to the versatility of bacterium as a protein expression host.
The E. coli cultivation process involves growing the bacteria in a culture medium, with selection antibiotics, until reaching an optical density600 (OD)600 between 0.6 and 0.8, a mid-log phase indicative. For that, the most widely used media are Luria–Bertani, Terrific Broth, and Super Broth, based on mixtures of tryptone/peptone and yeast extract in a saline solution, which can either be sodium chloride or sodium phosphate. To optimize growth, the bacterial culture must be kept at a temperature and rotation of 37 °C and 150–200 revolutions per minute (rpm). Upon reaching the desired absorbance, the inducer molecule must be added to the culture medium, typically the allolactose analog isopropyl β-D-1-thiogalactopyranose (IPTG), to start recombinant gene transcription. More details of this approach are well provided by Sambrook et al. (1989) [132].
The first E. coli isolate was deposited in the National Collection of Type Cultures (NCTC, UK) in 1920. Later on, Cohen et al. (1973) made a groundbreaking discovery of recombinant DNA technology, which marked a pivotal moment in the biotechnology field five decades ago [133]. Successful productions of human somatostatin [134] and insulin [135] were quickly achieved in those cells. From then on, its ease of genetic manipulation has allowed for the insertion and knockout of genes, resulting in several strains better suited for each recombinant protein profile. Although there is a clear preference for E. coli B derivative strains in RMP surveys (BL21 and BL21(DE3)), there are other useful lineages as well. For example, there are strains more effective in preventing the formation of inclusion bodies, a challenge frequently discussed in the literature concerning E. coli in protein production. The ideal strain to use will depend on the specific requirements of the protein being expressed.
The E. coli BL21 was developed in the work of Studier and Moffatt (1986) after various modifications of the parental B cell line [136]. Today, it is the most widely used strain for recombinant expression. Along the way to BL21 development, several mutations were introduced. Some of them were beneficial for recombinant protein production, such as a mutation in the hsdS gene that prevents plasmid loss from transformed bacteria (introduced in the parental B834 strain; [137]). However, a few mutations may not have a direct correlation with recombinant production or may even hamper cultivation in autoinduction media, an alternative to IPTG induction. This is the case of the inactivation of galK, galT and galE genes, which encode important enzymes for galactose use and the Leloir pathway (introduced in the parental strain B707; [138] likewise, the absence of flagellar biosynthesis genes fli, which renders the non-motility in B lineage. However, in this instance, it also has the benefit of saving energy that might otherwise be spent on recombinant yield [131]. In addition, a major advantage of the B strain comes from the higher expression of genes related to amino acid synthesis and decreased expression of those for degradation, indicating their suitability for efficient protein production [139].
The BL21 cells carry knockouts in the Lon and OmpT genes, which encode cytoplasmic and outer membrane proteases, respectively. Those components consistently hinder recombinant manufacturing by hydrolyzing proteins throughout the downstream process. The BL21(DE3) is a derivative strain version that contains a λ prophage that encodes the T7 RNA polymerase, which recognizes the widely-used T7 promoter and is five to eight times faster compared to native E. coli polymerases [140, 141]. Consequently, BL21 is used solely for protein expression by E. coli native RNA polymerase promoters, e.g. lac, tac, trc, ParaBAD, PrhaBAD, and T5, upstream of the gene.
Several strains have been derived from the BL21 focusing on overcoming common challenges encountered in laboratory routine. Two key issues to bear in mind when creating RMPs, as discussed in the third section, are the codon usage ratio and protein folding errors. In addition to the bioinformatics tools mentioned below, there are specific E. coli strains developed to mitigate those potential issues. In this regard, an E. coli Rosetta-derived lineage was used to express RMPs designed for T. cruzi, W. bancrofti, and human HTLV detection [10, 115, 126]. This lineage harbors extra copies of genes encoding rare tRNAs, including for AUA, AGG, AGA, CUA, CCC, and GGA codons [142]. The strain BL21-CodonPlus(DE3)-RIL, which contains similar modifications, was also used to express the RMP rqTSA-25 for the diagnosis of bovine tapeworm [122]. These examples demonstrate the ability to address codon usage issues by selecting suitable lineages without entirely replacing the gene codons planning.
A problem with folding errors that is routinely described in the literature is the inclusion bodies occurrence due to misfolded recombinant proteins. Inclusion bodies are aggregates of biomolecules, mostly proteins, to which the bacteria become more susceptible during the recombinant expression [143]. The physicochemical properties of amino acids, particularly the hydrophobic interactions, are key factors that govern the formation of inclusion bodies [144]. As the RMPs have not undergone natural selection, they might exhibit instability issues, being more prone to form inclusion bodies. Moreover, protein expression at high rates also triggers inclusion body formation, which is a common feature of BL21 derivative strains. For instance, T7 promoters are able to raise recombinant proteins to constitute 50% of total cell proteins within a few hours [145]. The inclusion bodies form more readily under metabolic stress as the production of recombinant and host proteins compete for resources. This contest arises from the overload on DNA replication, the rivalry for transcription and translation elements, and the supplementary energy [146].
Three leading ways to avoid inclusion body formation without involving the RMP redesigning are (‘) reduce recombinant protein production, (2) use stress-adapted strains, or (3) insert certain adjustments into the RMP’s plasmids. The first approach mainly consists of culturing E. coli at lower temperatures (30° to 25 °C), slowing down expression, and enhancing protein stability [147, 148]. Furthermore, strains containing plasmids, such as pLysS or pLysE, significantly benefit RMP production by using T7 promoters, preventing inclusion body formation [149]. These plasmids co-express T7 lysozyme, which suppresses transcriptional leak of recombinant genes, an approach adopted for expression of the following RMPs: r-LMP, ITC 8.2, USM.TOXO1, rMEHB, and rMEHCV [2, 10, 19, 31, 40].
Another example that is in line with better-controlled transcription of recombinant genes is the adoption of E. coli Tuner(DE3) strain. This BL21-derivative carries a Lac permease enzyme mutation, ensuring uniform IPTG uptake across all cells and leading to concentration-dependent induction levels [150, 151]. Also, the Evo21(DE3) strain, a recently developed cell line adapted to recombinant expression burden, is a promising candidate for future RMP production. It expresses 3.6-fold higher levels than BL21(DE3) eight hours post-induction, also dealing better with inclusion body formation [152].
A key factor in the widespread use of E. coli is its remarkable ability to readily incorporate foreign DNA, especially in a plasmid format [153, 154]. Thus, several tags and vector arrangements have been developed to strategically improve recombinant yield, including through the mitigation of inclusion bodies. Regarding vectors, the pET series of expression plasmids is by far the most commonly used for recombinant research (> 220,000 published research studies cited its use; [155, 156]). Its first generation was developed using a pBR322 backbone. Over 100 derivatives have since been developed, with pET28a and pET15b being the most commonly used. These vectors enable fusion with the histidine-tag (His-tag), which is highly effective for detection in immunochemical assays, e.g., ELISA and Western blot, and for purification via immobilized metal-affinity chromatography [157]. The His-tag effectiveness is not distinct in either the N- or C-terminal junction. However, depending on the protein's folding, the tag may enter a cryptic pocket and lose its utility. Additionally, the C-terminal location can be useful when verifying protein integrity in electrophoretic assays, such as the Western blot.
However, the His-tag is unlikely to interfere with solubility, especially for large proteins and, therefore, does not prevent inclusion body formation. To promote protein solubility, researchers have used a range of fusion tags, including thioredoxin (Trx), glutathione S-transferase (GST), small ubiquitin-related modifier (SUMO), and maltose binding protein (MBP). Several vectors are available that carry these fusion tags [158]. The SUMO tag, which is added to the end of proteins that have their genes cloned in the pSUMO or pET SUMO vectors, is particularly effective in this regard. It can act as a chaperone and facilitate folding, as well as increasing solubility [159]. Moreover, there are vectors, such as pET43 and pET44/pET32, which carry N-utilization substance A (NusA) and Trx, respectively, as well as more specialized ones. such as pGEX and pMAL, which bring the tags GST and MBP, respectively.
The K-lineage E. coli strains are an alternative to the aforementioned B-lineage. Their isolation occurred in 1922 from the stool of a diphtheria patient in Palo Alto (CA, USA; [138, 160]), and since then, many strains adapted to recombinant expression have been developed. For the production of PALD, a RMP for human leprosy, the authors used the HMS-174 strain [41], and E. coli SG13009 was used for r-DME-G expression, a RMP for dengue detection [9]. Notably, these K-lineage bacteria outperform BL21 strains in lactose induction scenarios (autoinduction component). The HMS-174(DE3) strain exhibits a nearly threefold higher lactose uptake rate compared to BL21(DE3), and it accumulates less galactose, minimizing osmotic stress. As a result, the specific product titer was twice as high in the HMS-174 (DE3) strain, as compared to BL21(DE3) strain [161].
Considering all the advantages of using E. coli in the production of recombinant proteins, it is not surprising that the vast majority of studies with RMPs mentioned above used this host for the expression of their target antigens, highlighting the great biotechnological contribution of this microorganism to scientific research.
Importance of bioinformatics for RMP analysis
Bioinformatics plays a crucial role in addressing challenges related to RMP production, particularly in cases of low heterologous overexpression and poor solubility [162, 163]. By employing bioinformatics tools, researchers can design optimized gene sequences for the target protein, including codon optimization to match the host organism's codon preferences [164, 165]. Codons are triplets of nucleotides in deoxyribonucleic acid (DNA) that code for specific amino acids in a protein [166]. Different organisms have variations in their codon usage preferences, with some being more frequently used than others [167]. When a gene from one organism is introduced into another, the differences in codon usage can lead to inefficient translation and lower protein expression levels [168, 169].
Bioinformatics tools analyze the codon usage patterns of both the source organism and the host organism [167, 170]. Researchers can then modify the gene sequence to replace rarely used codons with preferred codons of the host organism, a process known as codon optimization [171–173]. This adjustment improves the efficiency of translation and enhances protein expression levels in the host organism [171]. Besides, bioinformatics algorithms are used to identify the optimal codon replacements to maximize protein expression [174]. These algorithms consider factors such as codon frequency, transfer ribonucleic acid (tRNA) availability in the host organism, and messenger ribonucleic acid (mRNA) secondary structure [175]. By analyzing these factors, the bioinformatics tools generate a modified gene sequence that is more likely to be efficiently translated in the host [176]. Available bioinformatics tools for codon optimization and their features are shown in Table 6. Moreover, bioinformatics tools available from commercial entities can also provide gene synthesis services, being an additional option for researchers.
Table 6.
Bioinformatics tools for codon optimization
| Bioinformatics tool | Description |
|---|---|
| Gene designer | User-friendly software tool that allows researchers to design and optimize DNA sequences, including codon optimization. It provides a graphic interface for visualizing and editing gene sequences, and it can suggest codon substitutions based on codon usage preferences in the target organism [176] |
| JCat | Java Codon Adaptation Tool is an online tool that performs codon optimization for a target gene sequence. It takes into account the codon usage tables of both the source and target organisms and suggests codon replacements to enhance protein expression in the host organism [177] |
| Codon optimizer | Online tool that enables codon optimization for a wide range of organisms. Users can input their gene sequence and select the target organism, and the tool will provide a codon-optimized version of the gene [178] |
| DNA works | Software package that includes codon optimization functionality. It allows users to optimize gene sequences based on various criteria, such as codon frequency, secondary structure, and more [179] |
| COOL | Codon Optimization OnLine is an online tool that performs codon optimization for a variety of organisms. It provides options for specifying codon usage tables and other parameters to customize the optimization process [180] |
| OPTIMIZER | Program that optimizes codon usage based on a user-defined set of preferred codons. Researchers can input their codon usage table and specify codon preferences for optimization |
| ICOR | Codon optimization tool that uses recurrent neural networks to improve heterologous expression of synthetic genes [181] |
| EuGene | Comprehensive tool for analyzing and processing genetic information, ranging from codon usage to gene identification and structural analysis [182] |
| COStar | Algorithm that uses D-star Lite to address codon optimization by creating a specialized graph structure [183] |
| D-Tailor | DNA-Tailor is a versatile tool for designing DNA sequences with specific characteristics, using a customizable Monte Carlo approach [184] |
| CODA | Computationally Optimized DNA Assembly is an algorithm that leverages the redundancy in the genetic code to generate overlapping oligonucleotides with specific thermodynamic properties [185] |
| ATGme | Open-source web-based application that simplifies optimization through its user-friendly interface, offering three strategies: one-click, bulk based on codon types, and individualized custom optimization [185] |
| Codon wizard | Versatile tool for analyzing and optimizing codon usage in various types of input sequences. It allows users to freely combine algorithms, including both established and novel ones, to achieve desired results [186] |
Additionally, bioinformatics can predict and identify regions of the protein prone to misfolding or aggregation, enabling the design of modifications or fusion tags that enhance protein solubility and stability [187, 188]. Bioinformatics tools and algorithms analyze the amino acid sequence of a protein to predict regions that are susceptible to misfolding or aggregation [189]. These regions are often characterized by sequences that have a high propensity to form beta-sheet structures or expose hydrophobic residues [190]. Bioinformatics methods use algorithms and databases that incorporate data on protein structures, folding kinetics, and known aggregation-prone motifs [191, 192]. Once problematic regions are identified, bioinformatics can aid in the design of modifications to mitigate misfolding and aggregation issues [193]. One common approach is to introduce point mutations that disrupt or stabilize specific interactions within the protein structure [193, 194]. For example, mutations can be introduced to reduce the exposure of hydrophobic residues or promote more favorable intramolecular interactions [195].
Bioinformatics tools can also predict the impact of these mutations on the protein's stability and solubility. They guide the selection of fusion tags or chaperone proteins that enhance protein solubility and stability [188, 196]. Fusion tags are peptide sequences added to the target protein that improve its expression and solubility [197]. Bioinformatics helps choose appropriate fusion tags by considering such factors as size, charge, and affinity for purification [198]. Conversely, chaperone proteins can assist in the correct folding of the target protein [199]. Computational tools can suggest chaperone proteins known to interact with proteins of interest, facilitating proper folding during expression [200, 201]. Advanced bioinformatics tools can also perform in silico structural modeling to simulate how modifications or fusion tags will affect the protein's three-dimensional structure [202]. Available bioinformatic tools to predict misfolding or aggregation and their features are shown in Table 7.
Table 7.
Bioinformatics tools to predict misfolding or aggregation and design modifications or fusion
| Bioinformatics tool | Description |
|---|---|
| TANGO | Bioinformatics tool that predicts amyloidogenic regions in protein sequences. It calculates the aggregation propensity of different segments of a protein and helps identify regions that are likely to form aggregates. Researchers can use this information to design modifications to mitigate aggregation risks [203] |
| AGGRESCAN | Bioinformatics tool that predicts aggregation-prone regions in protein sequences based on the physicochemical properties of amino acids. It calculates aggregation propensity scores and identifies potential hotspots for aggregation [204] |
| FoldX | While primarily a tool for protein stability prediction, FoldX can be used to assess the impact of mutations on protein folding and aggregation propensity. Researchers can use it to evaluate the effects of mutations designed to enhance solubility and stability [205] |
| Rosetta FlexPepDock | Versatile suite of software tools for protein structure prediction and design. It can be used to model protein structures and assess the impact of mutations or modifications on protein stability and solubility [206] |
| I-TASSER | Protein structure and function prediction tool that can identify potential aggregation-prone regions based on structural modeling. It provides insights into the three-dimensional structure of a protein and can help design modifications to improve stability [207] |
| TIsigner | Computational tool in bioinformatics that is used to design and optimize translation start sites (TIS) in gene or mRNA sequences, allowing control and adjustment of gene expression [208] |
| SoDoPE | Valuable tool in protein engineering and recombinant protein production. It allows users to analyze a protein sequence and its domains to predict and enhance solubility [208] |
| ESPRESSO | System or tool designed to estimate the expression and solubility of proteins in different expression systems. Uses data analysis and bioinformatics approaches to predict the ability of a protein to be expressed and maintained in a soluble form in expression systems such as bacteria, yeast, or mammalian cells [209] |
| Aggrescan3D | Web server used to predict the aggregation properties of protein structures. It achieves this by incorporating 3D structural information and assessing the significance of solvent-exposed aggregation-prone regions [210] |
Moreover, bioinformatics analyses aid in selecting the most suitable host organism and expression system based on the protein's characteristics, ultimately streamlining the production process [155, 211]. Bioinformatics helps researchers choose the optimal host organism by considering factors such as the protein's size, complexity, post-translational modifications, the organism's genetic tools and available resources, and the desired protein yield [176, 212, 213]. For example, if a protein requires complex glycosylation, bioinformatics analysis may suggest using a eukaryotic expression system such as yeast or mammalian cells [214]. Conversely, if a simpler prokaryotic system, like E. coli, is sufficient, bioinformatics could confirm this choice based on the protein's features [162, 215].
Bioinformatics can assist in selecting the most suitable promoter for gene expression in the chosen host organism. By analyzing promoter databases and regulatory elements, researchers can identify promoters that match the protein's expression requirements, including inducible or constitutive expression, high-level production, or tissue-specific expression [216, 217]. In the case of secreted proteins, computational tools can predict signal peptides that target the protein for secretion in host organisms, such as yeast or bacteria. This ensures efficient secretion into the extracellular space [218, 219]. Bioinformatics tools can aid in designing plasmids for cloning and expression. This includes selecting appropriate vectors, resistance markers, and other genetic elements necessary for gene expression. Optimized plasmid design can enhance protein production efficiency [156, 220]. In addition, possible post-translational modifications, such as glycosylation or phosphorylation sites, can be predicted, allowing researchers to plan appropriate quality control and subsequent processing steps [221–223]. Available bioinformatic tools to select the most suitable host organism for RMPs expression and their features are shown in Table 8.
Table 8.
Bioinformatic tools to select the most suitable host organism and expression system
| Bioinformatics tool | Description |
|---|---|
| Host designer | Web-based tool that assists in choosing the most appropriate host organism for recombinant protein expression based on user-defined criteria and protein characteristics [224] |
| Vector NTI | This software offers features for sequence analysis and vector design, aiding in the selection of host organisms and plasmids for protein expression [225] |
| Geno CAD | Web-based platform that assists in designing genetic constructs, including promoter selection, for recombinant protein expression in various hosts [226] |
| Regulon DB | Database containing information on transcriptional regulation in Escherichia coli, aiding in promoter selection for bacterial expression [227] |
| Signal P | Tool for predicting signal peptides in protein sequences, which is essential for secreted protein expression [228] |
| Deep Sig | Tool for predicting signal peptides and their cleavage sites in proteins [229] |
| ApE | Versatile multi-platform application used to design plasmids and other constructs by simulating cloning methods such as PCR, Gibson assembly, restriction-ligation assembly, and Golden Gate assembly in silico [230] |
| Plas mapper | Web server that enables users to create, modify, add annotations, and interactively display high-quality plasmid maps [231] |
| Opt flux | Computational tool for metabolic engineering and optimization that can be used to predict optimal conditions for recombinant protein production [232] |
| Phospho site | Web-based bioinformatics resource specifically dedicated to cataloging the sites where proteins undergo phosphorylation in humans and mice [233] |
In summary, by leveraging the power of computational techniques and data-driven insights, the field of bioinformatics plays a crucial role in the synthesis of RMPs. It acts as a vital link between genetic engineering and protein manufacturing, providing detailed answers to some of the most difficult problems involved in this procedure. Bioinformatics enables researchers to make informed choices about host organisms, expression systems, and modifications, ensuring that proteins are effectively synthesized, correctly folded, and easily soluble. Codon optimization, structural analysis, and predictive modeling provide these choices by speeding up the production process and raising the likelihood of successfully producing functional recombinant proteins overall. The biotechnology and pharmaceutical sectors continue to rely on the adaptability of recombinant proteins for therapeutic and commercial applications, and bioinformatics continues to play a crucial role in advancing innovation and the ability to fully realize the potential of these proteins for the advancement of science and society.
Importance of biophysical analysis of the RMP´s structure for diagnosis
Biophysical analysis, in general, is applied to characterize biological systems, providing important information on the conformational stability of the molecules. These techniques have several advantages, such as obtaining structural information about the protein of interest in the most diverse experimental conditions. Normally, these techniques require a small amount of sample and provide rapid data acquisition [234, 235]. Among these techniques, circular dichroism (CD) can be applied for the characterization of chiral systems, including peptides, proteins, carbohydrates, and nucleic acids, based on the optical phenomenon of absorption of circularly polarized light. The study of proteins using this technique allows the identification and estimation of secondary structures and provides information about the tertiary structure from the Far-UltraViolet (Far-UV) and near-Ultra-Violet (near-UV) CD spectra, respectively. Furthermore, conformational changes can be analyzed under different experimental conditions, such as pH, temperature, and salt concentration in the solvent, as well as interactions between protein-protein, protein-membrane, or several other molecules [235–239].
The CD technique has been applied in several studies to estimate the secondary structure profile of the RMPs in solution for application in diagnostic tests [2, 4, 19, 107, 114]. Typically, the RMPs are rationally designed containing solvent-exposed bound epitopes. Therefore, to determine the secondary structure content of the alpha helix, beta-sheet, beta-turn, and random coil, the Far-UV spectra of the RMPs are recorded in solution, and the ellipticities are converted to molar ellipticity [θ] (deg.cm2/dmol) [240, 241]. These results can be used to confirm whether the RMPs recombinant preserved the predicted secondary structure or defined the stability under different environments.
Several RMPs developed for disease diagnosis, such as rMEHB for hepatitis B [19], rMEHCMV for human cytomegalovirus [107], rMERUB for rubella [114], and rMELEISH for visceral canine and human leishmaniasis [4], showed a Far-UV CD spectra with a pronounced negative dichroic band at 200 nm, compatible with an unordered structure (≥ 40%), followed by β-sheets content (≥ 35%) and a lower α-helix percentage (≤ 15%). In contrast, rMEHCV developed for hepatitis C diagnosis showed a negative dichroic band at 208 nm, indicating a higher content of β-sheets (≥ 56%) [2]. In these studies, CD results showed that proteins presented structural stability at 25 °C under neutral and basic pH. In addition, changes in the CD signal as a function of temperature ranging from 25 to 95 °C indicated that the structural stability of proteins decreases at temperature above 40 °C [2, 19, 107]. Taken together, the results from the biophysical analysis were fundamental to establishing the ideal structural conditions of the RMP, in which their biological functions would be preserved.
It is known that greater exposure of epitopes would be the ideal condition for better recognition of the molecule by antibodies present in serum [1]. Therefore, the structural study of RMPs in solution is one of the most important steps for evaluating their diagnostic capacity, being a useful tool aimed at increasing the efficiency of diagnostic tests, since the CD technique is an effective strategy for determining the secondary structure and folding properties of proteins (Greenfield, 2009).
Conclusion
In the present review, since only studies using the nomenclature “recombinant multiepitope protein” were selected, despite of the large number of studies using RMPs for immunodiagnostics, this number may be underestimated, since some authors use the nomenclature “chimera” instead of RMP. Ever since the first description of the development and use of RMPs for immunodiagnostics, these molecules have been widely used in the diagnosis of multiple animal and human diseases. It is evident that the use of RMPs for immunological diagnosis has increased significantly over the past ten years, most likely as a result of the benefits they have over other diagnostic kits that have been covered here. In reality, they are molecules of considerable scientific interest due to the high sensitivity and specificity values they offer in immunological diagnoses. In this regard, a significant amount of the next generation of diagnostic technologies will likely include these compounds.
Acknowledgements
The authors would like to thank to the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior-CAPES (Finance Code 001), Conselho Nacional de Desenvolvimento Científico e Tecnológico-CNPq, and Fundação de Amparo à Pesquisa do Estado de Minas Gerais- FAPEMIG (APQ-02704-23, BPD-00647-22, RED-00067-23, RED-00193-23), UFSJ, UFMG and UCSM. EAFC, RCG, SMF, and ASG would like to thank CNPq for their PQ/DT fellowship. The authors are grateful to Randall Johnson for revising the English of our manuscript.
Author contributions
All author’s contributed in writing, design and figures of this review. All authors read and approved the final manuscript.
Funding
None.
Availability of data and materials
Not applicable.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Dipti CA, Jain SK, Navin K. A novel recombinant multiepitope protein as a hepatitis C diagnostic intermediate of high sensitivity and specificity. Prot Exp Purific. 2006;47:319–328. doi: 10.1016/j.pep.2005.12.012. [DOI] [PubMed] [Google Scholar]
- 2.Galdino AS, Santos JC, Souza MQ, Nóbrega YKM, Xavier MAE, Felipe MSS, et al. A novel structurally stable multiepitope protein for detection of HCV. Hepat Res Treat. 2016 doi: 10.1155/2016/6592143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Agallou M, Margaroni M, Kotsakis SD, Karagouni E. A canine-directed chimeric multi-epitope vaccine induced protective immune responses in BALB/c mice infected with Leishmania infantum. Vaccines. 2020 doi: 10.3390/vaccines8030350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Dias DS, Machado JM, Ribeiro PAF, Machado AS, Ramos FF, Nogueira LM, et al. rMELEISH: a novel recombinant multiepitope-based protein applied to the serodiagnosis of both canine and human visceral leishmaniasis. Pathogens. 2023;12:302. doi: 10.3390/pathogens12020302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Machado JM, Pereira IAG, Maia ACG, Francisco MFC, Nogueira LM, Gandra IB, et al. Proof of concept of a novel multiepitope recombinant protein for the serodiagnosis of patients with chagas disease. Pathogens. 2023;12:312. doi: 10.3390/pathogens12020312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Napoleão-Pêgo P, Carneiro FRG, Durans AM, Gomes LR, Morel CM, Provance DW, Jr, et al. Performance assessment of a multi-epitope chimeric antigen for the serological diagnosis of acute mayaro fever. Sci Rep. 2021;11:15374. doi: 10.1038/s41598-021-94817-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Naz A, Shahid F, Butt TT, Awan FM, Ali A, Malik A. Designing multi-epitope vaccines to combat emerging coronavirus disease 2019 (COVID-19) by employing immuno-informatics approach. Front Immunol. 2020;11:1663. doi: 10.3389/fimmu.2020.01663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gran View Research. Recombinant proteins market size, share & trends analysis report by host cell (insect cells, mammalian), by application (research, therapeutics), by product & services, by end-user, by region, and segment forecasts, 2022–2030. 2022. https://www.grandviewresearch.com/industry-analysis/recombinant-proteins-market-report. 15 May 2023.
- 9.AnandaRao R, Swaminathan S, Fernando S, Jana AM, Khanna N. A custom-designed recombinant multiepitope protein as a dengue diagnostic reagent. Protein Expr Purif. 2005;41:136–147. doi: 10.1016/j.pep.2005.01.009. [DOI] [PubMed] [Google Scholar]
- 10.Houghton RL, Stevens YY, Hjerrild K, Guderian J, Okamoto M, Kabir M, et al. Lateral flow immunoassay for diagnosis of Trypanosoma cruzi infection with high correlation to the radioimmunoprecipitation assay. Clin Vaccine Immunol. 2009;16:515–520. doi: 10.1128/CVI.00383-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Palatnik-de-Sousa I, Wallace ZS, Cavalcante SC, Ribeiro MPF, Silva JABM, Cavalcante RC, et al. A novel vaccine based on SARS-CoV-2 CD4+ and CD8+ T cell conserved epitopes from variants alpha to omicron. Sci Rep. 2022;12:16731. doi: 10.1038/s41598-022-21207-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chávez-Fumagalli MA, Martins VT, Testasicca MC, Lage DP, Costa LR, Lage PS, et al. Sensitive and specific serodiagnosis of Leishmania infantum infection in dogs by using peptides selected from hypothetical proteins identified by an immunoproteomic approach. Clin Vaccine Immunol. 2013;20:835–841. doi: 10.1128/CVI.00023-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ebrahimi M, Seyyedtabaei SJ, Ranjbar MM, Tahvildar-biderouni F, Mamaghani AJ. Designing and modeling of multi-epitope proteins for diagnosis of Toxocara canis infection. Int J Pept Res Ther. 2020;26:1371–1380. doi: 10.1007/s10989-019-09940-1. [DOI] [Google Scholar]
- 14.Lemes MR, Rodrigues TCV, Jaiswal AK, Tiwari S, Sales-Campos H, Andrade-Silva LE, et al. In silico designing of a recombinant multi-epitope antigen for leprosy diagnosis. J Genet Eng Biotechnol. 2022;20:128. doi: 10.1186/s43141-022-00411-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Flower DR, Davies MN, Doytchinova IA. Identification of candidate vaccine antigens in silico. Immunomic Discov Adjuv Candidate Subunit Vaccines. 2012;28(5):39–71. [Google Scholar]
- 16.Acevedo GR, Juiz NA, Ziblat A, Perri LP, Girard MC, Ossowski MS, et al. In Silico guided discovery of novel class I and II Trypanosoma cruzi epitopes recognized by T cells from chagas’ disease patients. J Immunol. 2020;204:1571–1581. doi: 10.4049/jimmunol.1900873. [DOI] [PubMed] [Google Scholar]
- 17.Carvalho GBF, Resende DM, Siqueira LMV, Lopes MD, Lopes DO, Coelho PMZ, et al. Selecting targets for the diagnosis of Schistosoma mansoni infection: an integrative approach using multi-omic and immunoinformatics data. PLoS ONE. 2017;12:e0182299. doi: 10.1371/journal.pone.0182299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Davies MN, Flower DR. Harnessing bioinformatics to discover new vaccines. Drug Discov Today. 2007;12:389–395. doi: 10.1016/j.drudis.2007.03.010. [DOI] [PubMed] [Google Scholar]
- 19.de Souza MQ, Galdino AS, dos Santos JC, Soares MV, de Nóbrega YC, Alvares AC, et al. A recombinant multiepitope protein for hepatitis B diagnosis. Biomed Res Int. 2013;2013:148317. doi: 10.1155/2013/148317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Adam KM. Immunoinformatics approach for multi-epitope vaccine design against structural proteins and ORF1a polyprotein of severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) Trop Dis Travel Med Vaccines. 2021;7:22. doi: 10.1186/s40794-021-00147-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Baloch Z, Ikram A, Shamim S, Obaid A, Awan FM, Naz A, et al. Human coronavirus spike protein based multi-epitope vaccine against covid-19 and potential future zoonotic coronaviruses by using immunoinformatic approaches. Vaccines. 2022;10:1150. doi: 10.3390/vaccines10071150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Dar HA, Waheed Y, Najmi MH, Ismail S, Hetta HF, Ali A, et al. Multiepitope subunit vaccine design against covid-19 based on the spike protein of sars-cov-2: an in silico analysis. J Immunol Res. 2020;2020:8893483. doi: 10.1155/2020/8893483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Obaidullah AJ, Alanazi MM, Alsaif NA, Albassam H, Almehizia AA, Alqahtani AM, et al. Immunoinformatics-guided design of a multi-epitope vaccine based on the structural proteins of severe acute respiratory syndrome coronavirus 2. Trop Dis Travel Med Vaccines. 2021;7:22. doi: 10.1186/s40794-021-00147-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Singh H, Jakhar R, Sehrawat N. Designing spike protein (S-protein) based multi-epitope peptide vaccine against SARS COVID-19 by immunoinformatics. Heliyon. 2020;6:e05528. doi: 10.1016/j.heliyon.2020.e05528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Pollet J, Chen WH, Strych U. Recombinant protein vaccines, a proven approach against coronavirus pandemics. Adv Drug Deliv Rev. 2021;170:71–82. doi: 10.1016/j.addr.2021.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rosano GL, Ceccarelli EA. Recombinant protein expression in Escherichia coli: advances and challenges. Front Microbiol. 2014;5:172. doi: 10.3389/fmicb.2014.00172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Cabal ABS, Wu T-Y. Recombinant protein technology in the challenging era of coronaviruses. Processes. 2022;10:946. doi: 10.3390/pr10050946. [DOI] [Google Scholar]
- 28.Bhatwa A, Wang W, Hassan YI, Abraham N, Li XZ, Zhou T. Challenges associated with the formation of recombinant protein inclusion bodies in Escherichia coli and strategies to address them for industrial applications. Front Bioeng Biotechnol. 2021;9:630551. doi: 10.3389/fbioe.2021.630551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Rabert C, Weinacker D, Pessoa A, Jr, Farías JG. Recombinants proteins for industrial uses utilization of Pichia pastoris expression system. Braz J Microbiol. 2013;44:351–356. doi: 10.1590/S1517-83822013005000041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Dai JF, Jiang M, Qu LL, Sun L, Wang YY, Gong LL, et al. Toxoplasma gondii: enzyme-linked immunosorbent assay based on a recombinant multi-epitope peptide for distinguishing recent from past infection in human sera. Exp Parasitol. 2013;133:95–100. doi: 10.1016/j.exppara.2012.10.016. [DOI] [PubMed] [Google Scholar]
- 31.Hajissa K, Zakaria R, Suppian R, Mohamed Z. Design and evaluation of a recombinant multi-epitope antigen for serodiagnosis of Toxoplasma gondii infection in humans. Parasit Vectors. 2015;8:315. doi: 10.1186/s13071-015-0932-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lv C, Hong Y, Fu Z, Lu K, Cao X, Wang T, et al. Evaluation of recombinant multi-epitope proteins for diagnosis of goat schistosomiasis by enzyme-linked immunosorbent assay. Parasit Vectors. 2016;9:135. doi: 10.1186/s13071-016-1418-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Bartlett A, Padfield D, Lear L, Bendall R, Vos M. A comprehensive list of bacterial pathogens infecting humans. Microbiology. 2022;168:12. doi: 10.1099/mic.0.001269. [DOI] [PubMed] [Google Scholar]
- 34.Shaw LP, Wang AD, Dylus D, Meier M, Pogacnik G, Dessimoz C, et al. The phylogenetic range of bacterial and viral pathogens of vertebrates. Mol Ecol. 2020;29:3361–3379. doi: 10.1111/mec.15463. [DOI] [PubMed] [Google Scholar]
- 35.GBD 2019 Antimicrobial Resistance Collaborators Global mortality associated with 33 bacterial pathogens in 2019: a systematic analysis for the Global Burden of Disease Study 2019. Lancet. 2022;400:2221–2248. doi: 10.1016/S0140-6736(22)02185-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Dadgostar P. Antimicrobial resistance: implications and costs. Infect Drug Resist. 2019;12:3903–3910. doi: 10.2147/IDR.S234610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Larsson DGJ, Flach CF. Antibiotic resistance in the environment. Nat Rev Microbiol. 2022;20:257–269. doi: 10.1038/s41579-021-00649-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Peri AM, Stewart A, Hume A, Irwin A, Harris PNA. New microbiological techniques for the diagnosis of bacterial infections and sepsis in ICU including point of care. Curr Infect Dis Rep. 2021;23:12. doi: 10.1007/s11908-021-00755-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Houghton RL, Lodes MJ, Dillon DC, Reynolds LD, Day CH, McNeill PD, et al. Use of multiepitope polyproteins in serodiagnosis of active tuberculosis. Clin Diagn Lab Immunol. 2002;9:883–891. doi: 10.1128/CDLI.9.4.883-891.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Lin X, Chen Y, Yan J. Recombinant multiepitope protein for diagnosis of leptospirosis. Clin Vaccine Immunol. 2008;15:1711–1714. doi: 10.1128/CVI.00189-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Duthie MS, Hay MN, Morales CZ, Carter L, Mohamath R, Ito L, et al. Rational design and evaluation of a multiepitope chimeric fusion protein with the potential for leprosy diagnosis. Clin Vaccine Immunol. 2010;17:298–303. doi: 10.1128/CVI.00400-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Cheng Z, Zhao JW, Sun ZQ, Song YZ, Sun QW, Zhang XY, et al. Evaluation of a novel fusion protein antigen for rapid serodiagnosis of tuberculosis. J Clin Lab Anal. 2011;25:344–349. doi: 10.1002/jcla.20483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Li JL, Huang XY, Chen HB, Wang XJ, Zhu CZ, Zhao M, et al. Simultaneous detection of IgG and IgM antibodies against a recombinant polyprotein PstS1-LEP for tuberculosis diagnosis. BMC Infect Dis. 2015;47:643–649. doi: 10.3109/23744235.2015.1043941. [DOI] [PubMed] [Google Scholar]
- 44.Yin D, Li L, Song X, Li H, Wang J, Ju W, et al. A novel multi-epitope recombined protein for diagnosis of human brucellosis. BMC Infect Dis. 2016;16:219. doi: 10.1186/s12879-016-1552-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Schreterova E, Bhide M, Potocnakova L, Borszekova PL. Design, construction and evaluation of multi-epitope antigens for diagnosis of Lyme disease. Ann Agric Environ Med. 2017;24:696–701. doi: 10.26444/aaem/80699. [DOI] [PubMed] [Google Scholar]
- 46.Yin D, Bai Q, Zhang J, Xu K, Li J. A novel recombinant multiepitope protein candidate for the diagnosis of brucellosis: a pilot study. J Microbiol Method. 2020;174:105964. doi: 10.1016/j.mimet.2020.105964. [DOI] [PubMed] [Google Scholar]
- 47.Yin D, Bai Q, Li L, Xu K, Zhang J. Study on immunogenicity and antigenicity of a novel brucella multiepitope recombined protein. Biochem Biophys Res Commun. 2021;540:37–41. doi: 10.1016/j.bbrc.2020.12.098. [DOI] [PubMed] [Google Scholar]
- 48.Yin D, Bai Q, Wu X, Li H, Shao J, Sun M, et al. A multi-epitope fusion protein-based p-elisa method for diagnosing bovine and goat brucellosis. Front Vet Sci. 2021;8:708008. doi: 10.3389/fvets.2021.708008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Yin D, Bai Q, Wu X, Li H, Shao J, Sun M, et al. Paper-based ELISA diagnosis technology for human brucellosis based on a multiepitope fusion protein. PLoS Negl Trop Dis. 2021;15:e0009695. doi: 10.1371/journal.pntd.0009695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lyashchenko KP, Sikar-Gang A, Sridhara AA, Johnathan-Lee A, Elahi R, Lambotte P, et al. Novel polyprotein antigens designed for improved serodiagnosis of bovine tuberculosis. Vet Immunol Immunopathol. 2021;240:110320. doi: 10.1016/j.vetimm.2021.110320. [DOI] [PubMed] [Google Scholar]
- 51.Yao M, Liu M, Chen X, Li J, Li Y, Wei YR, et al. Comparison of BP26, Omp25 and Omp31 and a multiepitope-based fusion protein in the serological detection of canine brucellosis. Infect Drug Resist. 2022;15:5301–5308. doi: 10.2147/IDR.S374432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Firacative C. Invasive fungal disease in humans: are we aware of the real impact? Mem Inst Oswaldo Cruz. 2020;115:e200430. doi: 10.1590/0074-02760200430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Taylor DL, Hollingsworth TN, McFarland JW, Lennon NJ, Nusbaum C, Ruess RW. A first comprehensive census of fungi in soil reveals both hyperdiversity and fine-scale niche partitioning. Ecol Monogr. 2014;84:3–20. doi: 10.1890/12-1693.1. [DOI] [Google Scholar]
- 54.Fisher MC, Gurr SJ, Cuomo CA, Blehert DS, Jin H, Stukenbrock EH, et al. Threats posed by the fungal kingdom to humans, wildlife, and agriculture. MBio. 2020 doi: 10.1128/mBio.00449-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Who. Report. WHO fungal priority pathogens list to guide research, development and public health action. 2022. https://www.who.int/publications/i/item/9789240060241. 20 May 2023.
- 56.Seyedmousavi S, Bosco SMG, Hoog S, Ebel F, Elad D, Gomes RR, et al. Fungal infections in animals: a patchwork of different situations. Med Mycol. 2018;56:165–S187. doi: 10.1093/mmy/myx104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Kainz K, Bauer MA, Madeo F, Carmona-Gutierrez D. Fungal infections in humans: the silent crisis. Microb Cell. 2020;7:143–145. doi: 10.15698/mic2020.06.718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Rodrigues ML, Albuquerque PC. Searching for a change: The need for increased support for public health and research on fungal diseases. PLoS Negl Trop Dis. 2018;12:e0006479. doi: 10.1371/journal.pntd.0006479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Fisher MC, Izquierdo AA, Berman J, Bicanic T, Bignell EM, Bowyer P, et al. Tackling the emerging threat of antifungal resistance to human health. Nat Rev Microbiol. 2022;20:557–571. doi: 10.1038/s41579-022-00720-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Parums DV. Editorial: the World Health Organization (WHO) fungal priority pathogens list in response to emerging fungal pathogens during the COVID-19 pandemic. Med Sci Monit. 2022;28:e939088. doi: 10.12659/MSM.939088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Tomás AL, Cardoso F, Esteves F, Matos O. Serological diagnosis of pneumocystosis: production of a synthetic recombinant antigen for immunodetection of Pneumocystis jirovecii. Sci Rep. 2016;6:36287. doi: 10.1038/srep36287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Brandão RMSS, Faria AR, de Andrade HM, Soares Martins LM, da Silva AS, do Monte SJH. Novel recombinant multiepitope proteins for the detection of anti-Cryptococcus antibodies. Futur Microbiol. 2018;13:429–436. doi: 10.2217/fmb-2017-0184. [DOI] [PubMed] [Google Scholar]
- 63.Tomás AL, Cardoso F, de Sousa B, Matos O. Detection of anti-Pneumocystis jirovecii antibodies in human serum using a recombinant synthetic multi-epitope kexin-based antigen. Eur J Clin Microbiol Infect Dis. 2020;39:2205–2209. doi: 10.1007/s10096-020-03936-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Yaeger RG. Protozoa structure, classification, growth, and development. In: Baron S, editor. Medical microbiology. 4. Texas: Galveston; 1996. [PubMed] [Google Scholar]
- 65.Aronson NE, Magill AJ. General principles. In: Ryan ET, Hill DR, Solomon T, Aronson NE, Endy TP, editors. Hunter's tropical medicine and emerging infectious diseases. 10. Amsterdam: Elsevier; 2020. pp. 696–698. [Google Scholar]
- 66.Andrews KT, Fisher G, Skinner-Adams TS. Drug repurposing and human parasitic protozoan diseases. Int J Parasitol Drugs Drug Resist. 2014;4:95–111. doi: 10.1016/j.ijpddr.2014.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Who. Malaria. 2023. https://www.who.int/news-room/fact-sheets/detail/malaria. 31 May 2023.
- 68.Who. Leishmaniasis. 2023. https://www.who.int/news-room/fact-sheets/detail/leishmaniasis. 31 May 2023.
- 69.Who. Chagas disease (American trypanosomiasis). 2023. https://www.who.int/health-topics/chagas-disease#tab=tab_1. 31 May 2023.
- 70.Gazel D, Ekşi F. Novel methods for diagnosis of blood-borne protozoa. Eur J of Therap. 2020;26:141–149. doi: 10.5152/EurJTher.2019.18085. [DOI] [Google Scholar]
- 71.Camussone C, Gonzalez V, Belluzo MS, Pujato N, Ribone ME, Lagier CM, et al. Comparison of recombinant Trypanosoma cruzi peptide mixtures versus multiepitope chimeric proteins as sensitizing antigens for immunodiagnosis. Clin Vaccine Immunol. 2009;16:899–905. doi: 10.1128/CVI.00005-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Dai J, Jiang M, Wang Y, Qu L, Gong R, Si J. Evaluation of a recombinant multiepitope peptide for serodiagnosis of Toxoplasma gondii infection. Clin Vaccine Immunol. 2012;19:338–342. doi: 10.1128/CVI.05553-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Garcia VS, Gonzalez VD, Caudana PC, Vega JR, Marcipar IS, Gugliotta LM. Synthesis of latex-antigen complexes from single and multiepitope recombinant proteins. application in immunoagglutination assays for the diagnosis of Trypanosoma cruzi infection. Coll Surf B Biointerfac. 2013;101:384–391. doi: 10.1016/j.colsurfb.2012.07.018. [DOI] [PubMed] [Google Scholar]
- 74.Faria AR, Veloso LC, Coura-Vital W, Reis AB, Damasceno LM, Gazzinelli RT, et al. Novel recombinant multiepitope proteins for the diagnosis of asymptomatic Leishmania infantum-infected dogs. PLoS Negl Trop Dis. 2015;9:e3429. doi: 10.1371/journal.pntd.0003429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Duthie MS, Guderian JA, Vallur AC, Misquith A, Liang H, Mohamath R, et al. Multi-epitope proteins for improved serological detection of Trypanosoma cruzi infection and chagas disease. Diagn Microbiol Infect Dis. 2016;84:191–196. doi: 10.1016/j.diagmicrobio.2015.11.006. [DOI] [PubMed] [Google Scholar]
- 76.Faria AR, Pires SDF, Reis AB, Coura-Vital W, Silveira JAGD, Sousa GM, et al. Canine visceral leishmaniasis follow-up: a new anti-IgG serological test more sensitive than ITS-1 conventional PCR. Vet Parasitol. 2017;248:62–67. doi: 10.1016/j.vetpar.2017.10.020. [DOI] [PubMed] [Google Scholar]
- 77.Hajissa K, Zakaria R, Suppian R, Mohamed Z. An evaluation of a recombinant multiepitope based antigen for detection of Toxoplasma gondii specific antibodies. BMC Infect Dis. 2017;17:807. doi: 10.1186/s12879-017-2920-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Peverengo LM, Garcia V, Rodeles LM, Mendicino D, Vicco M, Lagier C, et al. Development and assessment of an improved recombinant multiepitope antigen-based immunoassay to diagnose chronic chagas disease. Parasitology. 2018;145:1594–1599. doi: 10.1017/S0031182018000458. [DOI] [PubMed] [Google Scholar]
- 79.Fonseca THS, Faria AR, Leite HM, da Silveira JAG, Carneiro CM, Andrade HM. Chemiluminescent ELISA with multi-epitope proteins to improve the diagnosis of canine visceral leishmaniasis. Vet J. 2019;253:105387. doi: 10.1016/j.tvjl.2019.105387. [DOI] [PubMed] [Google Scholar]
- 80.Jameie F, Dalimi A, Pirestani M, Mohebali M. Detection of Leishmania infantum infection in reservoir dogs using a multiepitope recombinant protein (PQ10) Arch Razi Inst. 2020;75:327–338. doi: 10.22092/ARI.2019.126524.1346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Alibakhshi A, Bandehpour M, Sharifnia Z, Kazemi B. The development and evaluation of a multi-epitope antigen as a serodiagnostic marker of Toxoplasma gondii infection. Adv Clin Exp Med. 2020;29:669–675. doi: 10.17219/acem/104554. [DOI] [PubMed] [Google Scholar]
- 82.Song Y, Zhao Y, Pan K, Shen B, Fang R, Hu M, et al. Characterization and evaluation of a recombinant multiepitope peptide antigen MAG in the serological diagnosis of Toxoplasma gondii infection in pigs. Parasit Vectors. 2021;14:408. doi: 10.1186/s13071-021-04917-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Yaghoubi P, Bandehpour M, Mohebali M, Akhoundi B, Kazemi B. Designing and evaluation of a recombinant multiepitope protein by using ELISA for diagnosis of Leishmania infantum infected in dogs. Iran J Parasitol. 2021;16:377–385. doi: 10.18502/ijpa.v16i3.7090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Heidari S, Hajjaran H, Kazemi B, Gharechahi J, Mohebali M, Ranjbar MM, et al. Identification of immunodominant proteins of Leishmania infantum by immunoproteomics to evaluate a recombinant multi-epitope designed antigen for serodiagnosis of human visceral leishmaniasis. Exp Parasitol. 2021;222:108065. doi: 10.1016/j.exppara.2021.108065. [DOI] [PubMed] [Google Scholar]
- 85.Jameie F, Dalimi A, Pirestani M, Mohebali M. Development of a multi-epitope recombinant protein for the diagnosis of human visceral leishmaniasis. Iran J Parasitol. 2021;16:1–10. doi: 10.18502/ijpa.v16i1.5506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Taherzadeh M, Fouladvand M, Kazemi B. Evaluation of a new multi-epitope sequence of eight known Leishmania infantum antigens for HVL diagnosis by ELISA and western blot. J Vector Borne Dis. 2021;58:289–296. doi: 10.4103/0972-9062.318310. [DOI] [PubMed] [Google Scholar]
- 87.Mirza AZ, Shamshad H, Osra FA, Habeebullah TM, Morad M. An overview of viruses discovered over the last decades and drug development for the current pandemic. Eur J Pharmacol. 2021;890:173746. doi: 10.1016/j.ejphar.2020.173746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Woolhouse M, Scott F, Hudson Z, Howey R, Chase-Topping M. Human viruses: discovery and emergence. Philos Trans R Soc Lond B Biol Sci. 2012;367:2864–2871. doi: 10.1098/rstb.2011.0354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Murcia P, Donachie W, Palmarini M. Viral pathogens of domestic animals and their impact on biology, medicine and agriculture. Encycl Microbiol. 2009 doi: 10.1016/B978-012373944-5.00368-0. [DOI] [Google Scholar]
- 90.Okeleji OL, Ajayi LO, Odeyemi AN, Amos V, Ajayi HO, Akinyemi AO, et al. Viral zoonotic diseases of public health importance and their effect on male reproduction. Zoonotic Dis. 2022;2:291–300. doi: 10.3390/zoonoticdis2040023. [DOI] [Google Scholar]
- 91.Piret J, Boivin G. Pandemics Throughout History. Front Microb. 2020;11:631736. doi: 10.3389/fmicb.2020.631736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Cdc. Disease burden of flu. 2022. https://www.cdc.gov/flu/about/burden/index.html. 19 May 2023.
- 93.Who. HIV, Number of people dying from HIV-related causes. 2023. https://www.who.int/data/gho/data/indicators/indicator-details/GHO/number-of-deaths-due-to-hiv-aids. 19 May 2023.
- 94.Cassedy A, Parle-McDermott A, O'Kennedy R. Virus detection: a review of the current and emerging molecular and immunological methods. Front Mol Biosci. 2021;8:637559. doi: 10.3389/fmolb.2021.637559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Dronina J, Samukaite-Bubniene U, Ramanavicius A. Advances and insights in the diagnosis of viral infections. J Nanobiotechnol. 2021 doi: 10.1186/s12951-021-01081-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Anandarao R, Swaminathan S, Fernando S, Jana AM, Khanna N. Recombinant multiepitope protein for early detection of dengue infections. Clin Vaccine Immunol. 2006;13:59–67. doi: 10.1128/CVI.13.1.59-67.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Tripathi NK, Shrivastva A, Pattnaik P, Parida M, Dash PK, Gupta N, et al. Production of IgM specific recombinant dengue multiepitope protein for early diagnosis of dengue infection. Biotechnol Prog. 2007;23:488–493. doi: 10.1021/bp0602698. [DOI] [PubMed] [Google Scholar]
- 98.Tripathi NK, Shrivastva A, Pattnaik P, Parida M, Dash PK, Jana AM, et al. Production, purification and characterization of recombinant dengue multiepitope protein. Biotechnol Appl Biochem. 2007;46:105–113. doi: 10.1042/BA20060090. [DOI] [PubMed] [Google Scholar]
- 99.Talha SM, Salminen T, Chugh DA, Swaminathan S, Soukka T, Pettersson K, et al. Inexpensive designer antigen for anti-HIV antibody detection with high sensitivity and specificity. Clin Vaccine Immunol. 2010;17:335–341. doi: 10.1128/CVI.00283-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.He J, Xiu B, Wang G, Chen K, Feng X, Song X, et al. Double-antigen sandwich ELISA for the detection of anti-hepatitis C virus antibodies. J Virol Method. 2011;171:163–168. doi: 10.1016/j.jviromet.2010.10.019. [DOI] [PubMed] [Google Scholar]
- 101.Gurramkonda C, Talha SM, Gudi SK, Gogineni VR, Rao KRSS. Fed-batch cultivation of Escherichia coli expressed designer hepatitis C virus diagnostic intermediate and its evaluation. Biotechnol Appl Biochem. 2012;59:437–444. doi: 10.1002/bab.1044. [DOI] [PubMed] [Google Scholar]
- 102.Lin X, Chen S, Xue X, Lu L, Zhu S, Li W, et al. Chimerically fused antigen rich of overlapped epitopes from latent membrane protein 2 (LMP2) of epstein-barr virus as a potential vaccine and diagnostic agent. Cell Mol Immunol. 2016;13:492–501. doi: 10.1038/cmi.2015.29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Su Q, Guo M, Jia Z, Qiu F, Lu X, Gao Y, et al. Epitope-based recombinant diagnostic antigen to distinguish natural infection from vaccination with hepatitis A virus vaccines. J Virol Method. 2016;233:41–45. doi: 10.1016/j.jviromet.2016.02.014. [DOI] [PubMed] [Google Scholar]
- 104.Salminen T, Juntunen E, Khanna N, Pettersson K, Talha SM. Anti-HCV immunoassays based on a multiepitope antigen and fluorescent lanthanide chelate reporters. J Virol Method. 2016;228:67–73. doi: 10.1016/j.jviromet.2015.11.015. [DOI] [PubMed] [Google Scholar]
- 105.Cao Y, Zhou W, Xing X, Zhang J, Fu Y, Li K, et al. Indirect ELISA using a multi-epitope recombinant protein to detect antibodies against foot-and-mouth disease virus serotype O in pigs. J Virol Method. 2018;262:26–31. doi: 10.1016/j.jviromet.2018.09.008. [DOI] [PubMed] [Google Scholar]
- 106.Thomasini RL, Souza HGA, Bruna-Romero O, Totola AH, Gonçales NSL, Lima CX, et al. Evaluation of a recombinant multiepitope antigen for diagnosis of hepatitis C virus: a lower cost alternative for antigen production. J Clin Lab Anal. 2018;32:e22410. doi: 10.1002/jcla.22410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Ribeiro PAF, Souza MQ, Dias DS, Álvares ACM, Nogueira LM, Machado JM, et al. A custom-designed recombinant multiepitope protein for human cytomegalovirus diagnosis. Recent Pat Biotechnol. 2019;13:316–328. doi: 10.2174/1872208313666190716093911. [DOI] [PubMed] [Google Scholar]
- 108.Hao YF, Li SH, Zhang GZ, Xu Y, Long GZ, Lu XX, et al. Establishment of an indirect ELISA-based method involving the use of a multiepitope recombinant S protein to detect antibodies against canine coronavirus. Arch Virol. 2021;166:1877–1883. doi: 10.1007/s00705-021-05072-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Gomes LR, Durans AM, Napoleão-Pêgo P, Waterman JA, Freitas MS, De Sá NBR, et al. Multiepitope proteins for the differential detection of IgG antibodies against RBD of the spike protein and Non-RBD regions of SARS-CoV-2. Vaccines. 2021;9:986. doi: 10.3390/vaccines9090986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Zhang X, Guo J, Wang L, Li Z, Liu Y, Tian L, et al. Development and evaluation of multi-epitope protein p72 (MeP72) for the serodiagnosis of African swine fever. Acta Virol. 2021;65:273–278. doi: 10.4149/av_2021_304. [DOI] [PubMed] [Google Scholar]
- 111.Gao Z, Shao JJ, Zhang GL, Ge SD, Chang YY, Xiao L, et al. Development of an indirect ELISA to specifically detect antibodies against African swine fever virus: bioinformatics approaches. Virol J. 2021;18:97. doi: 10.1186/s12985-021-01568-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Liu W, Shao J, Zhang G, Chang Y, Ge S, Sun Y, et al. Development of an indirect chemiluminescence immunoassay using a multiepitope recombinant protein to specifically detect antibodies against foot-and-mouth disease virus serotype O in swine. J Clin Microbiol. 2021;59:e02464–e2520. doi: 10.1128/JCM.02464-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Pedersen J, Moukandja IP, Ndidi S, Sørensen AL, Koumakpayi IH, Lekana-Douki JB, et al. An adaptable platform for in-house hepatitis C serology. J Virol Method. 2022;308:114586. doi: 10.1016/j.jviromet.2022.114586. [DOI] [PubMed] [Google Scholar]
- 114.Souza M, Machado J, da Silva J, Ramos L, Nogueira L, Ribeiro P, et al. Rational design and evaluation of the recombinant multiepitope protein for serodiagnosis of rubella. Curr Pharm Biotechnol. 2022;23:1094–1100. doi: 10.2174/1389201022666210907170921. [DOI] [PubMed] [Google Scholar]
- 115.Franco GM, da Rocha AS, Cox LJ, Daian E Silva DSO, da Silveira E, Santos DM, Martins ML, et al. Multi-epitope protein as a tool of serological diagnostic development for HTLV-1 and HTLV-2 infections. Front Publ Health. 2022;10:884701. doi: 10.3389/fpubh.2022.884701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.da Silva LAD, Lima MDRQ, de Camargo BR, Guimarães DKDSC, Barbastefano AAL, Lima RC, et al. A chikungunya virus multiepitope recombinant protein expressed from the binary system insect cell/recombinant baculovirus is useful for laboratorial diagnosis of chikungunya. Microorganisms. 2022;10:1451. doi: 10.3390/microorganisms10071451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Who. Soil-transmitted helminth infections. 2023. https://www.who.int/news-room/fact-sheets/detail/soil-transmitted-helminth-infections. 6 June 2023.
- 118.Wright JE, Werkman M, Dunn JC, Anderson RM. Current epidemiological evidence for predisposition to high or low intensity human helminth infection: a systematic review. Parasit Vectors. 2018;11:65. doi: 10.1186/s13071-018-2656-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Majewska AA, Huang T, Han B, Drake JM. Predictors of zoonotic potential in helminths. Philos Trans R Soc Lond B Biol Sci. 2021;376:20200356. doi: 10.1098/rstb.2020.0356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Ngwese MM, Manouana GP, Moure PAN, Ramharter M, Esen M, Adégnika AA. Diagnostic Techniques of soil-transmitted helminths: impact on control measures. Trop Med Infect Dis. 2020;5:93. doi: 10.3390/tropicalmed5020093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Lv C, Zhiqiang F, Ke L, Ruili Y, Tao W, Xiaodan C, et al. A perspective for improving the sensitivity of detection: the application of multi-epitope recombinant antigen in serological analysis of buffalo schistosomiasis. Acta Trop. 2018;183:14–18. doi: 10.1016/j.actatropica.2018.03.025. [DOI] [PubMed] [Google Scholar]
- 122.Guimarães-Peixoto RPM, Pinto PSA, Santos MR, Zilch TJ, Apolinário PF, Silva-Júnior A. Development of the multi-epitope chimeric antigen rqTSA-25 from Taenia saginata for serological diagnosis of bovine cysticercosis. PLoS Negl Trop Dis. 2018;12:e0006371. doi: 10.1371/journal.pntd.0006371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Tianli L, Xifeng W, Zhenzhong T, Lixia W, Xingxing Z, Jun Q, et al. Multi-epitope fusion protein Eg mefag-1 as a serodiagnostic candidate for cystic echinococcosis in sheep. Korean J Parasitol. 2019;57:61–67. doi: 10.3347/kjp.2019.57.1.61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Lagatie O, Verheyen A, Nijs E, Batsa Debrah L, Debrah YA, Stuyver LJ. Performance evaluation of 3 serodiagnostic peptide epitopes and the derived multi-epitope peptide OvNMP-48 for detection of Onchocerca volvulus infection. Parasitol Res. 2019;18:2263–2270. doi: 10.1007/s00436-019-06345-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Aghamolaei S, Kazemi B, Bandehpour M, Ranjbar MM, Rouhani S, Javadi Mamaghani A, et al. Design and expression of polytopic construct of cathepsin-L1, SAP-2 and FhTP16.5 proteins of Fasciola hepatica. J Helminthol. 2020 doi: 10.1017/S0022149X20000140. [DOI] [PubMed] [Google Scholar]
- 126.Yasin N, Laxmanappa HS, Muddapur UM, Cheruvathur J, Prakash SMU, Thulasiram HV. Design, expression, and evaluation of novel multiepitope chimeric antigen of Wuchereria bancrofti for the diagnosis of lymphatic filariasis-a structure-based strategy. Int Immunopharmacol. 2020;83:106431. doi: 10.1016/j.intimp.2020.106431. [DOI] [PubMed] [Google Scholar]
- 127.Mirzapour A, Tabaei SJS, Bandehpour M, Haghighi A, Kazemi B. Designing a recombinant multi-epitope antigen of echinococcus granulosus to diagnose human cystic echinococcosis. Iran J Parasitol. 2020;15:1–10. [PMC free article] [PubMed] [Google Scholar]
- 128.Ozturk EA, Manzano-Román R, Sánchez-Ovejero C, Caner A, Angın M, Gunduz C, et al. Comparison of the multi-epitope recombinant antigen DIPOL and hydatid fluid for the diagnosis of patients with cystic echinococcosis. Acta Trop. 2022;225:106208. doi: 10.1016/j.actatropica.2021.106208. [DOI] [PubMed] [Google Scholar]
- 129.Yengo BN, Shintouo CM, Hotterbeekx A, Yaah NE, Shey RA, Quanico J, et al. Immunoinformatics design and assessment of a multiepitope antigen (OvMCBL02) for onchocerciasis diagnosis and monitoring. Diagnostics. 2022;12:1440. doi: 10.3390/diagnostics12061440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Clark DJ, Maalře O. DNA replication and the division cycle in Escherichia coli. J Mol Biol. 1967;23:99–112. doi: 10.1016/S0022-2836(67)80070-6. [DOI] [Google Scholar]
- 131.Jeong H, Barbe V, Lee CH, Vallenet D, Yu DS, Choi SH, et al. Genome sequences of Escherichia coli B strains REL606 and BL21 (DE3) J Mol Biol. 2009;394:644–652. doi: 10.1016/j.jmb.2009.09.052. [DOI] [PubMed] [Google Scholar]
- 132.Sambrook J, Fritsch ER, Maniatis T. Molecular cloning: a laboratory manual. 2. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- 133.Cohen SN, Chang AC, Boyer HW, Helling RB. Construction of biologically functional bacterial plasmids in vitro. Proc Natl Acad Sci U S A. 1973;11:3240–3244. doi: 10.1073/pnas.70.11.3240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Itakura K, Tadaaki H, Crea R, Riggs AD, Heyneker HL, Bolivar F, et al. Expression in Escherichia coli of a chemically synthesized gene for the hormone somatostatin. Biotechnology. 1977;24:84–91. [PubMed] [Google Scholar]
- 135.Goeddel DV, Kleid DG, Bolivar F, Heyneker HL, Yansura DG, Crea R, et al. Expression in Escherichia coli of chemically synthesized genes for human insulin. Proc Natl Acad Sci U S A. 1979;76:106–110. doi: 10.1073/pnas.76.1.106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Studier FW, Moffatt BA. Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol. 1986;189:113–130. doi: 10.1016/0022-2836(86)90385-2. [DOI] [PubMed] [Google Scholar]
- 137.Wood WB. Host specificity of DNA produced by Escherichia coli: bacterial mutations affecting the restriction and modification of DNA. J Mol Biol. 1966;16:118–133. doi: 10.1016/S0022-2836(66)80267-X. [DOI] [PubMed] [Google Scholar]
- 138.Daegelen P, Studier FW, Lenski RE, Cure S, Kim JF. Tracing ancestors and relatives of Escherichia coli B, and the derivation of B strains REL606 and BL21 (DE3) J Mol Biol. 2009;394:634–643. doi: 10.1016/j.jmb.2009.09.022. [DOI] [PubMed] [Google Scholar]
- 139.Yoon SH, Han MJ, Jeong H, Lee CH, Xia XX, Lee DH, et al. Comparative multi-omics systems analysis of Escherichia coli strains B and K-12. Genome Biol. 2012;13:R37. doi: 10.1186/gb-2012-13-5-r37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Iost I, Guillerez J, Dreyfus M. Bacteriophage T7 RNA polymerase travels far ahead of ribosomes in vivo. J Bacteriol. 1992;174:619–622. doi: 10.1128/jb.174.2.619-622.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Lewicki BT, Margus T, Remme J, Nierhaus KH. Coupling of rRNA transcription and ribosomal assembly in vivo. formation of active ribosomal subunits in Escherichia coli requires transcription of rRNA genes by host RNA polymerase which cannot be replaced by bacteriophage T7 RNA polymerase. J Mol Biol. 1993;231:581–593. doi: 10.1006/jmbi.1993.1311. [DOI] [PubMed] [Google Scholar]
- 142.Tegel H, Tourle S, Ottosson J, Persson A. Increased levels of recombinant human proteins with the Escherichia coli strain rosetta (DE3) Protein Expr Purif. 2010;69:159–167. doi: 10.1016/j.pep.2009.08.017. [DOI] [PubMed] [Google Scholar]
- 143.Tsumoto K, Ejima D, Kumagai I, Arakawa T. Practical considerations in refolding proteins from inclusion bodies. Protein Expr Purif. 2003;28:1–8. doi: 10.1016/S1046-5928(02)00641-1. [DOI] [PubMed] [Google Scholar]
- 144.Winkler J, Seybert A, König L, Pruggnaller S, Haselmann U, Sourjik V, et al. Quantitative and spatio-temporal features of protein aggregation in Escherichia coli and consequences on protein quality control and cellular ageing. EMBO J. 2010;29:910–923. doi: 10.1038/emboj.2009.412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Mierendorf RC, Morris BB, Hammer B, Novy RE. Expression and purification of recombinant proteins using the pET system. Methods Mol Med. 1998;13:257–292. doi: 10.1385/0-89603-485-2:257. [DOI] [PubMed] [Google Scholar]
- 146.Zeng H, Yang A. Quantification of proteomic and metabolic burdens predicts growth retardation and overflow metabolism in recombinant Escherichia coli. Biotechnol Bioeng. 2019;116:1484–1495. doi: 10.1002/bit.26943. [DOI] [PubMed] [Google Scholar]
- 147.Hunke S, Betton JM. Temperature effect on inclusion body formation and stress response in the periplasm of Escherichia coli. Mol Microbiol. 2003;50:1579–1589. doi: 10.1046/j.1365-2958.2003.03785.x. [DOI] [PubMed] [Google Scholar]
- 148.Strandberg L, Enfors SO. Factors influencing inclusion body formation in the production of a fused protein in Escherichia coli. Appl Environ Microbiol. 1991;57:1669–1674. doi: 10.1128/aem.57.6.1669-1674.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Studier FW. Use of bacteriophage T7 lysozyme to improve an inducible T7 expression system. J Mol Biol. 1991;219:37–44. doi: 10.1016/0022-2836(91)90855-Z. [DOI] [PubMed] [Google Scholar]
- 150.Marbach A, Bettenbrock K. lac operon induction in Escherichia coli: systematic comparison of IPTG and TMG induction and influence of the transacetylase LacA. J Biotechnol. 2012;157:82–88. doi: 10.1016/j.jbiotec.2011.10.009. [DOI] [PubMed] [Google Scholar]
- 151.Mühlmann MJ, Forsten E, Noack S, Büchs J. Prediction of recombinant protein production by Escherichia coli derived online from indicators of metabolic burden. Biotechnol Prog. 2018;34:1543–1552. doi: 10.1002/btpr.2704. [DOI] [PubMed] [Google Scholar]
- 152.Heyde SAH, Nørholm MHH. Tailoring the evolution of BL21 (DE3) uncovers a key role for RNA stability in gene expression toxicity. Commun Biol. 2021 doi: 10.1038/s42003-021-02493-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983;166:557–580. doi: 10.1016/S0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]
- 154.Warren RL, Freeman JD, Levesque RC, Smailus DE, Flibotte S, Holt RA. Transcription of foreign DNA in Escherichia coli. Genome Res. 2008;18:1798–1805. doi: 10.1101/gr.080358.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Mital S, Christie G, Dikicioglu D. Recombinant expression of insoluble enzymes in Escherichia coli: a systematic review of experimental design and its manufacturing implications. Microb Cell Fact. 2021;20:208. doi: 10.1186/s12934-021-01698-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Shilling PJ, Mirzadeh K, Cumming AJ, Widesheim M, Köck Z, Daley DO. Improved designs for pET expression plasmids increase protein production yield in Escherichia coli. Commun Biol. 2020;3:214. doi: 10.1038/s42003-020-0939-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Porath J, Carlsson J, Olsson I, Belfrage G. Metal chelate affinity chromatography, a new approach to protein fractionation. Nature. 1975;258:598–599. doi: 10.1038/258598a0. [DOI] [PubMed] [Google Scholar]
- 158.Terpe K. Overview of tag protein fusions: from molecular and biochemical fundamentals to commercial systems. Appl Microbiol Biotechnol. 2003;60:523–533. doi: 10.1007/s00253-002-1158-6. [DOI] [PubMed] [Google Scholar]
- 159.Kuo D, Nie M, Courey AJ. SUMO as a solubility tag and in vivo cleavage of SUMO fusion proteins with Ulp1. Method Mol Biol. 2014;1177:71–80. doi: 10.1007/978-1-4939-1034-2_6. [DOI] [PubMed] [Google Scholar]
- 160.Mairhofer J, Krempl PM, Thallinger GG, Striedner G. Finished genome sequence of Escherichia coli K-12 Strain HMS174 (ATCC 47011) Genome Announc. 2014;2:e00975–e1014. doi: 10.1128/genomeA.00975-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Hausjell J, Weissensteiner J, Molitor C, Halbwirth H, Spadiut OE. coli HMS174 (DE3) is a sustainable alternative to BL21 (DE3) Microb Cell Fact. 2018;17:169. doi: 10.1186/s12934-018-1016-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Chang CCH, Song J, Tey BT, Ramanan RN. Bioinformatics approaches for improved recombinant protein production in Escherichia coli: protein solubility prediction. Brief Bioinform. 2014;1:953–962. doi: 10.1093/bib/bbt057. [DOI] [PubMed] [Google Scholar]
- 163.Ouellet S, Ferguson L, Lau AZ, Lim TKY. CysPresso: a classification model utilizing deep learning protein representations to predict recombinant expression of cysteine-dense peptides. BMC Bioinform. 2023;24:200. doi: 10.1186/s12859-023-05327-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Şen A, Kargar K, Akgün E, Pınar MÇ. Codon optimization: a mathematical programing approach. Bioinformatics. 2020;36:4012–4020. doi: 10.1093/bioinformatics/btaa248. [DOI] [PubMed] [Google Scholar]
- 165.Karaşan O, Şen A, Tiryaki B, Cicek AE. A unifying network modeling approach for codon optimization. Bioinformatics. 2022;38:3935–3941. doi: 10.1093/bioinformatics/btac428. [DOI] [PubMed] [Google Scholar]
- 166.Ahmad M, Jung LT, Bhuiyan A-A. From DNA to protein: Why genetic code context of nucleotides for DNA signal processing? A review Biomed Signal Process Control. 2017;34:44–63. doi: 10.1016/j.bspc.2017.01.004. [DOI] [Google Scholar]
- 167.Athey J, Alexaki A, Osipova E, Rostovtsev A, Santana-Quintero LV, Katneni U, et al. A new and updated resource for codon usage tables. BMC Bioinform. 2017;18:391. doi: 10.1186/s12859-017-1793-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Lipinszki Z, Vernyik V, Farago N, Sari T, Puskas LG, Blattner FR, et al. Enhancing the translational capacity of E. coli by resolving the codon bias. ACS Synth Biol. 2018;7:2656–2664. doi: 10.1021/acssynbio.8b00332. [DOI] [PubMed] [Google Scholar]
- 169.Liu Y. A code within the genetic code: codon usage regulates co-translational protein folding. Cell Commun Signal. 2020;18:145. doi: 10.1186/s12964-020-00642-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Panda A, Tuller T. Determinants of associations between codon and amino acid usage patterns of microbial communities and the environment inferred based on a cross-biome metagenomic analysis. npj Biofilm Microbiome. 2023;9:5. doi: 10.1038/s41522-023-00372-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Fu H, Liang Y, Zhong X, Pan Z, Huang L, Zhang H, et al. Codon optimization with deep learning to enhance protein expression. Sci Rep. 2020;10:17617. doi: 10.1038/s41598-020-74091-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Al-Hawash AB, Zhang X, Ma F. Strategies of codon optimization for high-level heterologous protein expression in microbial expression systems. Gene Rep. 2017;9:46–53. doi: 10.1016/j.genrep.2017.08.006. [DOI] [Google Scholar]
- 173.Fox DM, Branson KM, Walker RC. mRNA codon optimization with quantum computers. PLoS ONE. 2021;16:e0259101. doi: 10.1371/journal.pone.0259101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Trösemeier J-H, Rudorf S, Loessner H, Hofner B, Reuter A, Schulenborg T, et al. Optimizing the dynamics of protein expression. Sci Rep. 2019;9:7511. doi: 10.1038/s41598-019-43857-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Parvathy ST, Udayasuriyan V, Bhadana V. Codon usage bias. Mol Biol Rep. 2022;49:539–565. doi: 10.1007/s11033-021-06749-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Watts A, Sankaranarayanan S, Watts A, Raipuria RK. Optimizing protein expression in heterologous system: strategies and tools. Meta Gene. 2021;29:100899. doi: 10.1016/j.mgene.2021.100899. [DOI] [Google Scholar]
- 177.Grote A, Hiller K, Scheer M, Munch R, Nortemann B, Hempel DC, et al. JCat: a novel tool to adapt codon usage of a target gene to its potential expression host. Nucleic Acids Res. 2005;33:W526–W531. doi: 10.1093/nar/gki376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 178.Fuglsang A. Codon optimizer: a freeware tool for codon optimization. Protein Expr Purif. 2003;31:247–249. doi: 10.1016/S1046-5928(03)00213-4. [DOI] [PubMed] [Google Scholar]
- 179.Hoover DM. DNAWorks: an automated method for designing oligonucleotides for PCR-based gene synthesis. Nucl Acid Res. 2002;30:43e–43. doi: 10.1093/nar/30.10.e43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Chin JX, Chung BKS, Lee DY. Codon optimization online (COOL): a web-based multi-objective optimization platform for synthetic gene design. Bioinformatics. 2014;30:2210–2212. doi: 10.1093/bioinformatics/btu192. [DOI] [PubMed] [Google Scholar]
- 181.Kunjapur AM, Pfingstag P, Thompson NC. Gene synthesis allows biologists to source genes from farther away in the tree of life. Nat Commun. 2018;9:4425. doi: 10.1038/s41467-018-06798-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182.Gaspar P, Oliveira JL, Frommlet J, Santos MAS, Moura G. EuGene: maximizing synthetic gene design for heterologous expression. Bioinformatics. 2012;28:2683–2684. doi: 10.1093/bioinformatics/bts465. [DOI] [PubMed] [Google Scholar]
- 183.Liu X, Deng R, Wang J, Wang X. COStar: a d-star lite-based dynamic search algorithm for codon optimization. J Theor Biol. 2014;344:19–30. doi: 10.1016/j.jtbi.2013.11.022. [DOI] [PubMed] [Google Scholar]
- 184.Guimaraes JC, Rocha M, Arkin AP, Cambray G. D-tailor: automated analysis and design of DNA sequences. Bioinformatics. 2014;30:1087–1094. doi: 10.1093/bioinformatics/btt742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Larsen LSZ, Wassman CD, Hatfield GW, Lathrop RH. Computationally optimised DNA assembly of synthetic genes. Int J Bioinform Res Appl. 2008;4:324. doi: 10.1504/IJBRA.2008.019578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186.Rehbein P, Berz J, Kreisel P, Schwalbe H. “CodonWizard”–an intuitive software tool with graphical user interface for customizable codon optimization in protein expression efforts. Protein Expr Purif. 2019;160:84–93. doi: 10.1016/j.pep.2019.03.018. [DOI] [PubMed] [Google Scholar]
- 187.Jo BH. An intrinsically disordered peptide tag that confers an unusual solubility to aggregation-prone proteins. Appl Environ Microbiol. 2022;88:e0009722. doi: 10.1128/aem.00097-22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 188.Qing R, Hao S, Smorodina E, Jin D, Zalevsky A, Zhang S. Protein design: from the aspect of water solubility and stability. Chem Rev. 2022;122:14085–14179. doi: 10.1021/acs.chemrev.1c00757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Prabakaran R, Rawat P, Kumar S, Gromiha MM. Evaluation of in silico tools for the prediction of protein and peptide aggregation on diverse datasets. Brief Bioinform. 2021 doi: 10.1093/bib/bbab240. [DOI] [PubMed] [Google Scholar]
- 190.Louros N, Orlando G, De Vleeschouwer M, Rousseau F, Schymkowitz J. Structure-based machine-guided mapping of amyloid sequence space reveals uncharted sequence clusters with higher solubilities. Nat Commun. 2020;11:3314. doi: 10.1038/s41467-020-17207-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Navarro S, Ventura S. Computational methods to predict protein aggregation. Curr Opin Struct Biol. 2022;73:102343. doi: 10.1016/j.sbi.2022.102343. [DOI] [PubMed] [Google Scholar]
- 192.Pallarés I, Ventura S. Advances in the prediction of protein aggregation propensity. Curr Med Chem. 2019;26:3911–3920. doi: 10.2174/0929867324666170705121754. [DOI] [PubMed] [Google Scholar]
- 193.Kundu D, Prerna K, Chaurasia R, Bharty MK, Dubey VK. Advances in protein misfolding, amyloidosis and its correlation with human diseases. Biotech. 2020;10:193. doi: 10.1007/s13205-020-2166-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Musil M, Konegger H, Hon J, Bednar D, Damborsky J. Computational design of stable and soluble biocatalysts. ACS Catal. 2019;9:1033–1054. doi: 10.1021/acscatal.8b03613. [DOI] [Google Scholar]
- 195.Staller MV, Ramirez E, Kotha SR, Holehouse AS, Pappu RV, Cohen BA. Directed mutational scanning reveals a balance between acidic and hydrophobic residues in strong human activation domains. Cell Syst. 2022;13:334–345.e5. doi: 10.1016/j.cels.2022.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Köppl C, Lingg N, Fischer A, Kröß C, Loibl J, Buchinger W, et al. Fusion tag design influences soluble recombinant protein production in Escherichia coli. Int J Mol Sci. 2022;23:7678. doi: 10.3390/ijms23147678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197.Esposito D, Chatterjee DK. Enhancement of soluble protein expression through the use of fusion tags. Curr Opin Biotechnol. 2006;17:353–358. doi: 10.1016/j.copbio.2006.06.003. [DOI] [PubMed] [Google Scholar]
- 198.Remans K, Lebendiker M, Abreu C, Maffei M, Sellathurai S, May MM, et al. Protein purification strategies must consider downstream applications and individual biological characteristics. Microb Cell Fact. 2022;21:52. doi: 10.1186/s12934-022-01778-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 199.Saibil H. Chaperone machines for protein folding, unfolding and disaggregation. Nat Rev Mol Cell Biol. 2013;14:630–642. doi: 10.1038/nrm3658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Bui LM, Geraldi A, Nguyen TT, Lee JH, Lee JY, Cho B-K, et al. mRNA engineering for the efficient chaperone-mediated co-translational folding of recombinant proteins in Escherichia coli. Int J Mol Sci. 2019;20:3163. doi: 10.3390/ijms20133163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201.Serapian SA, Triveri A, Marchetti F, Castelli M, Colombo G. Exploiting folding and degradation machineries to target undruggable proteins: what can a computational approach tell us? ChemMedChem. 2021;16:1593–1599. doi: 10.1002/cmdc.202000960. [DOI] [PubMed] [Google Scholar]
- 202.Chakravarty N, Priyanka SJ, Singh RP. A potential type-II L-asparaginase from marine isolate bacillus australimaris NJB19: statistical optimization, in silico analysis and structural modeling. Int J Biol Macromol. 2021;174:527–539. doi: 10.1016/j.ijbiomac.2021.01.130. [DOI] [PubMed] [Google Scholar]
- 203.Lu X, Brickson CR, Murphy RM. TANGO-inspired design of anti-amyloid cyclic peptides. ACS Chem Neurosci. 2016;7:1264–1274. doi: 10.1021/acschemneuro.6b00150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Conchillo-Solé O, de Groot NS, Avilés FX, Vendrell J, Daura X, Ventura S. AGGRESCAN: a server for the prediction and evaluation of “hot spots” of aggregation in polypeptides. BMC Bioinform. 2007;8:65. doi: 10.1186/1471-2105-8-65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 205.Delgado J, Radusky LG, Cianferoni D, Serrano L. FoldX 5.0: working with RNA, small molecules and a new graphical interface. Bioinformatics. 2019;35:4168–4169. doi: 10.1093/bioinformatics/btz184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.London N, Raveh B, Cohen E, Fathi G, Schueler-Furman O. Rosetta FlexPepDock web server—high resolution modeling of peptide–protein interactions. Nucleic Acid Res. 2011;39:W249–W253. doi: 10.1093/nar/gkr431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207.Zhang Y. I-TASSER server for protein 3D structure prediction. BMC Bioinform. 2008;9:40. doi: 10.1186/1471-2105-9-40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 208.Bhandari BK, Lim CS, Gardner PP. TISIGNER com web services for improving recombinant protein production. Nucleic Acid Res. 2021 doi: 10.1093/nar/gkab175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 209.Hirose S, Noguchi T. ESPRESSO: a system for estimating protein expression and solubility in protein expression systems. Proteomics. 2013;13:1444–1456. doi: 10.1002/pmic.201200175. [DOI] [PubMed] [Google Scholar]
- 210.Kuriata A, Iglesias V, Pujols J, Kurcinski M, Kmiecik S, Ventura S. Aggrescan3D (A3D) 2.0: prediction and engineering of protein solubility. Nucleic Acid Res. 2019;47:W300–W307. doi: 10.1093/nar/gkz321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Lozano Terol G, Gallego-Jara J, Sola Martínez RA, Martínez Vivancos A, Cánovas Díaz M, de Diego PT. Impact of the expression system on recombinant protein production in Escherichia coli BL21. Front Microbiol. 2021;12:682001. doi: 10.3389/fmicb.2021.682001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 212.Mahmoudi E, Kiltschewskij D, Fitzsimmons C, Cairns MJ. Depolarization-associated CircRNA regulate neural gene expression and in some cases may function as templates for translation. Cells. 2019;9:25. doi: 10.3390/cells9010025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.Graw S, Chappell K, Washam CL, Gies A, Bird J, Robeson MS, et al. Multi-omics data integration considerations and study design for biological systems and disease. Mol Omics. 2021;17:170–185. doi: 10.1039/D0MO00041H. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 214.Dell A, Galadari A, Sastre F, Hitchen P. Similarities and differences in the glycosylation mechanisms in prokaryotes and eukaryotes. Int J Microbiol. 2010;2010:1–14. doi: 10.1155/2010/148178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215.Peleg Y, Unger T. Resolving bottlenecks for recombinant protein expression in E. coli. Method Mol Biol. 2012;800:173–186. doi: 10.1007/978-1-61779-349-3_12. [DOI] [PubMed] [Google Scholar]
- 216.Chitayat Levi L, Rippin I, Ben Tulila M, Galron R, Tuller T. Modulating gene expression within a microbiome based on computational models. Biology. 2022;1:1301. doi: 10.3390/biology11091301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217.Timerbaev V, Dolgov S. Functional characterization of a strong promoter of the early light-inducible protein gene from tomato. Planta. 2019;250:1307–1323. doi: 10.1007/s00425-019-03227-x. [DOI] [PubMed] [Google Scholar]
- 218.Maffei B, Francetic O, Subtil A. Tracking proteins secreted by bacteria: what’s in the toolbox? Front Cell Infect Microbiol. 2017;7:221. doi: 10.3389/fcimb.2017.00221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 219.Juibari AD, Ramezani S, Rezadoust MH. Bioinformatics analysis of various signal peptides for periplasmic expression of parathyroid hormone in E.coli. J Med Life. 2019;12:184–191. doi: 10.25122/jml-2018-0049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 220.Yano H, Shintani M, Tomita M, Suzuki H, Oshima T. Reconsidering plasmid maintenance factors for computational plasmid design. Comput Struct Biotechnol J. 2019;17:70–81. doi: 10.1016/j.csbj.2018.12.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 221.Beygmoradi A, Homaei A, Hemmati R, Fernandes P. Recombinant protein expression: challenges in production and folding related matters. Int J Biol Macromol. 2023;233:123407. doi: 10.1016/j.ijbiomac.2023.123407. [DOI] [PubMed] [Google Scholar]
- 222.Mary B, Maurya S, Arumugam S, Kumar V, Jayandharan GR. Post-translational modifications in capsid proteins of recombinant adeno-associated virus (AAV) 1-rh10 serotypes. FEBS J. 2019;286:4964–4981. doi: 10.1111/febs.15013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 223.Leutert M, Entwisle SW, Villén J. Decoding post-translational modification crosstalk with proteomics. Mol Cell Proteomics. 2021;20:100129. doi: 10.1016/j.mcpro.2021.100129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 224.Hay BP, Firman TK. Host designer: a program for the de novo structure-based design of molecular receptors with binding sites that complement metal ion guests. Inorg Chem. 2002;41:5502–5512. doi: 10.1021/ic0202920. [DOI] [PubMed] [Google Scholar]
- 225.Lu G. Vector NTI, a balanced all-in-one sequence analysis suite. Brief Bioinform. 2004;5:378–388. doi: 10.1093/bib/5.4.378. [DOI] [PubMed] [Google Scholar]
- 226.Czar MJ, Cai Y, Peccoud J. Writing DNA with GenoCADTM. Nucl Acid Res. 2009;37:W40–W47. doi: 10.1093/nar/gkp361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 227.Santos-Zavaleta A, Salgado H, Gama-Castro S, Sánchez-Pérez M, Gómez-Romero L, Ledezma-Tejeida D, et al. RegulonDB v 10.5: tackling challenges to unify classic and high throughput knowledge of gene regulation in E. coli K-12. Nucl Acid Res. 2019;47:D212–D220. doi: 10.1093/nar/gky1077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 228.Teufel F, Armenteros JJA, Johansen AR, Gíslason MH, Pihl SI, Tsirigos KD, et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat Biotechnol. 2022;40:1023–1025. doi: 10.1038/s41587-021-01156-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 229.Savojardo C, Martelli PL, Fariselli P, Casadio R. DeepSig: deep learning improves signal peptide detection in proteins. Bioinformatics. 2018;34:1690–1696. doi: 10.1093/bioinformatics/btx818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 230.Davis MW, Jorgensen EM. ApE, a plasmid editor: a freely available DNA manipulation and visualization program. Front Bioinform. 2022;2:818619. doi: 10.3389/fbinf.2022.818619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 231.Wishart DS, Ren L, Leong-Sit J, Saha S, Grant JR, Stothard P, et al. PlasMapper 30—a web server for generating, editing, annotating and visualizing publication quality plasmid maps. Nucleic Acids Res. 2023;51:W459–W467. doi: 10.1093/nar/gkad276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 232.Rocha I, Maia P, Evangelista P, Vilaça P, Soares S, Pinto JP, et al. OptFlux: an open-source software platform for in silico metabolic engineering. BMC Syst Biol. 2010;4:45. doi: 10.1186/1752-0509-4-45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 233.Hornbeck PV, Chabra I, Kornhauser JM, Skrzypek E, Zhang B. PhosphoSite: a bioinformatics resource dedicated to physiological protein phosphorylation. Proteomics. 2004;4:1551–1561. doi: 10.1002/pmic.200300772. [DOI] [PubMed] [Google Scholar]
- 234.Lakowicz JR. Quenching of fluorescence. In: Lakowicz JR, editor. Principles of fluorescence spectroscopy. Boston: Springer; 1983. pp. 257–301. [Google Scholar]
- 235.Wallace BA, Janes RW. Modern techniques for circular dichroism and synchrotron radiation circular dichroism spectroscopy. Amsterdam: Ios Press Bv; 2009. [Google Scholar]
- 236.Corrêa DHA, Ramos CHI. The use of circular dichroism spectroscopy to study protein folding, form and function. Afr J of Biochem Res. 2009;3:164–173. [Google Scholar]
- 237.Berova N, Polavarapu PL, Nakanish K, Woody RW. Comprehensive Chiroptical Spectroscopy: applications in stereochemical analysis of synthetic compounds, natural products, and biomolecules. 1ed. New Jersey: John Wiley & Sons, Inc.; 2012.
- 238.Souza AA, Leitão VO, Ramada MHS, Mehdad A, Georg RC, Ulhôa CJ, Freitas SM. Trichoderma harzianum produces a new thermally stable acid phosphatase, with potential for biotechnological application. PLoS ONE. 2016;11:1–18. doi: 10.1371/journal.pone.0150455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 239.Oliveira ICM, Garay AV, Souza AA, Valadares NF, Barbosa JARG, Faria FP, Freitas SM. Structural and biochemical analysis reveals how ferulic acid improves catalytic efficiency of Humicola grisea xylanase. Sci Rep. 2022;12:11409. doi: 10.1038/s41598-022-15175-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 240.Greenfield NJ. Using circular dichroism spectra to estimate protein secondary structure. Nat Protoc. 2006;1:2876–2890. doi: 10.1038/nprot.2006.202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241.Miles AJ, Janes RW, Wallace BA. Tools and methods for circular dichroism spectroscopy of proteins: a tutorial review. Chem Soc Rev. 2021;50:8400–8413. doi: 10.1039/D0CS00558D. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.