Abstract
MicroRNAs (miRNAs) are natural, single-stranded, small RNA molecules which subtly control gene expression. Several studies indicate that specific miRNAs can regulate heart function both in development and disease. Despite prevention programs and new therapeutic agents, cardiovascular disease remains the main cause of death in developed countries. The elevated number of heart failure episodes is mostly due to myocardial infarction (MI). An increasing number of studies have been carried out reporting changes in miRNAs gene expression and exploring their role in MI and heart failure. In this review, we furnish a critical analysis of where the frontier of knowledge has arrived in the fields of basic and translational research on miRNAs in cardiac ischemia. We first summarize the basal information on miRNA biology and regulation, especially concentrating on the feedback loops which control cardiac-enriched miRNAs. A focus on the role of miRNAs in the pathogenesis of myocardial ischemia and in the attenuation of injury is presented. Particular attention is given to cardiomyocyte death (apoptosis and necrosis), fibrosis, neovascularization, and heart failure. Then, we address the potential of miR-diagnosis (miRNAs as disease biomarkers) and miR-drugs (miRNAs as therapeutic targets) for cardiac ischemia and heart failure. Finally, we evaluate the use of miRNAs in the emerging field of regenerative medicine.
Keywords: miRNAs, myomiRs, Myocardial ischemia, Heart failure
Introduction
The impact of ischemic heart disease
Human heart disease remains the main cause of death and disability in children and adults in the developed countries. According to the American Heart Association [1], in every year since 1900 (except 1918) cardiovascular disease (CVD) accounted for more deaths than any other single cause in the United States, including cancer, chronic lower respiratory diseases, accidents, and diabetes mellitus. An expected 80,000,000 American adults (1 in 3) have one or more types of CVD. Among these, 7,900,000 are estimated to have an acute myocardial infarction (MI) [1]. This number keeps increasing despite the emphasis placed on prevention and new therapeutic agents, rendering heart disease the health epidemic of the twenty-first century.
Ischemic heart disease is the most common underlying cause of heart failure. Myocardial ischemia may arise from several aetiologies, the most prevalent being coronary artery disease (CAD). Coronary artery disease causes severe impairment of the coronary blood supply and may lead to MI and heart failure. Resumption of blood flow in ischemic tissues, by coronary angioplasty or thrombolytic treatments, is currently recommended for the immediate early approach to acute MI. Reperfusion can clearly limit the extent of cardiomyocyte loss. However, the resupply of blood may give rise to mitochondrial reactive oxygen species (ROS) and mitochondrial Ca2+ accumulation and finally elicit apoptotic pathways [2–4].
The complexity of microRNAs (miRNAs) biology
The pathogenesis of heart disease is often associated with the altered expression of pivotal specific genes [5]. A number of transcriptome studies have been applied to CVD, with the aim of improving diagnosis, prognosis, and therapeutic assessment [5–7]. Myocardial biopsies of patients suffering from acute/chronic myocardial ischemia, as well as animal models of ischemia and reperfusion (I/R), have revealed changes in gene expression which seem to be specific for the cardiac ischemic injury [8–10]. MicroRNAs (miRNAs) are important regulators of gene expression, constituting a major area of investigation in cardiovascular research [11]. MicroRNAs are a class of endogenous, non-coding RNA molecules of approximately 21 nucleotides, which result from sequential processing of primary transcripts mediated by the RNase III enzymes, Drosha and Dicer. They post-transcriptionally regulate gene expression by means of an imperfect complementarity to the 3’ untranslated region of target mRNAs. The most stringent requirement for specifically targeting mRNAs is a contiguous and perfect base-pairing of the miRNA’s nucleotides 2–8 (seed sequence), which nucleates the interaction with the target messenger. MicroRNAs can regulate mRNA translation at the initiation, elongation, and termination process, or can affect mRNA stability. In eukaryotes, mRNA enzymatic degradation can follow two pathways, each of which is initiated by shortening of the mRNA poly-A tail [12]. Proteolytic cleavage of nascent polypeptides has also been described [13]. MicroRNAs may also act through mechanisms which go beyond complementarity. Indeed, they can regulate pre-mRNA processing in the nucleus, act as chaperones to modify structure, or modulate mRNA-protein interactions [13]. While miRNAs generally function in the cytoplasm, they can be imported in the nucleus [14], in the mitochondrium [15], and even be secreted from the cell [16] or transferred from one cell to another [17, 18]. Nevertheless, the relevance of subcellular localization for miRNA regulation is still a matter of study [19].
Currently, the estimated number of miRNA genes is as high as ~1,500 in the human genome (1–3 % of known genes are represented by miRNAs) and about 750 in the mouse [20], and they are supposed to regulate about 10,000 target genes (excluding alternative splicing variants), which is 30 % of the coding genes [21]. Indeed, each miRNA is assumed to regulate tens to thousands of targets. Moreover, they most likely act as an ensemble (often synergistically). Indeed, “miRNAs working together” interact at the level of single targets, set of target genes, and complex systems [22]. Critically, miRNAs often attenuate the expression of their targets only moderately [23]. The network which derives is quite complex and many pieces in unravelling miRNAs biology are missing. Therefore, it is not surprising that the picture of miRNA regulation is still highly fragmented and sometimes incoherent, due to gaps in untangling the coordinated action of such a number of molecules [13].
MicroRNAs in cardiovascular disease
Since miRNAs have proved to be key regulators of cardiovascular development [24], misregulation of miRNA function is likely to be involved in the onset of many human diseases, including CVD [25–28]. Since the initial implication of miRNAs in controlling heart development [29] and the identification of the pathological role of miRNAs during cardiac hypertrophy and failure [30–32], more than 800 articles have been published examining miRNAs in the heart (more than 400 in 2012 alone). Recent studies have compared mRNA and miRNA expression profiles in the same myocardial samples obtained from patients suffering from heart failure, with or without mechanical left ventricular assist device (LVAD) support. In contrast to mRNA, which varied moderately between the two groups, a specific set of miRNAs was significantly upregulated in heart failure and returned to normal in the LVAD recovery group [33, 34]. These findings suggest that miRNAs expression reflects the functional status of the heart more accurately than mRNAs.
In this review, we will briefly introduce muscle-specific miRNAs and describe the interplay occurring between miRNAs and their targets. Then, we will focus on miRNA regulation in cardiac ischemia. We will illustrate the role played by specific miRNAs in the different pathological aspects of myocardial ischemia: cardiomyocyte cell death, fibrosis, neovascularization, and heart failure. We will focus on the possible use of specific miRNAs as diagnostic and prognostic biomarkers and as new potential targets/tools for the treatment of heart disease. Finally, we will discuss the potential of miRNAs in the key promising field of regenerative medicine.
Feedback loops in the regulation of myomiRs
The myomiRs are a subset of miRNAs highly enriched in cardiac and/or skeletal muscle [35]. They include miR-1, miR-133, miR-206 (present only in skeletal muscle), miR-208, miR-486, and miR-499 [36–38]. Several of these miRNAs are organized in bicistronic clusters on the same chromosome (i.e. miR-1 and miR-133) and are transcribed together [35]. Myogenic regulatory factors bind to upstream sequences and promote the transcription of these myomiRs in skeletal (MyoD, Myogenin) and cardiac (Mef2, Srf) tissues. The myomiRs, in turn, affect multiple developmental and functional aspects in both muscle and heart through the post-transcriptional regulation of genes controlling myogenesis [29, 39].
MyomiRs and myogenic factors are linked by reciprocal interactions, often based on the onset of feedback circuits. For example, a feedforward loop is established in skeletal muscle cells, where miR-1 derepresses the activity of Mef2 through suppression of Hdac4. Activated Mef2, in turn, induces the expression of miR-1 [39]. Mef2 is also regulated by miR-494. In myoblasts, miR-494 directly regulates mtTfa and Foxj3, which are upstream to Mef2. Mef2 promotes the expression of mitochondrial proteins that, in turn, induce mtTfa [40]. Intriguingly, the presence of several myomiRs translocated from the cytosol to mitochondria has been demonstrated in human skeletal muscle cells [15]. Among these, miR-133 was predicted to target the mitochondrial NADH dehydrogenase subunit 1 and was implicated in mitochondrial mRNA silencing [15]. The key role of miR-133 is further supported by the switch-off mechanisms regulating its expression. In cardiac muscle cells, Srf regulates the expression of many miRNAs, including miR-133 [29], which in turn represses expression of Srf [35, 41]. The tight modulation of forces driven by miR-1 and miR-133 finally contributes to the balance between proliferation and differentiation in muscles [39].
Feedback regulatory loops also exist between myomiRs and intracellular signaling pathways. MiR-1 and miR-378 are two representative examples. In fact, both the Met receptor and its ligand Hgf were identified as candidate targets of miR-1 [42]. Notably, Met is also downregulated by miR15b [43]. A reciprocal interplay between miR-1 and Met has been shown (in cancer), since miR-1 downregulation results in Met overexpression and, in turn, Met negatively controls miR-1 expression [44]. We have recently demonstrated the involvement of Met signaling in cardiac hypertrophy [45], and the importance of Hgf/Met axis in the response to MI is well documented [46]. It is therefore likely that a negative feedback loop is also established between miR-1 and Met in myocardial disease, constituting a circuit of potential therapeutic interest. Moreover, miR-1 targets Igf1 and Igf1 receptor (Igf1R). In turn, the Igf1 pathway downregulates miR-1 expression through the inhibitory phosphorylation, mediated by Akt, of Foxo3a, which directly acts on miR-1 promoter [47]. Through an analogous reciprocal control, miR-378 expression is inhibited by Igf1 in cardiomyocytes [48].
The comprehension of the role and regulation of miRNAs during cardiac development, as well as in pathological conditions in the adult, constitutes the basis for new targeted therapies for cardiac disease.
MicroRNAs in the attenuation of myocardial injury
The oxygen deprivation and ROS production which follow cardiac ischemia and I/R cause cardiomyocyte cell death via both apoptosis and necrosis. Subsequently, cardiac fibroblasts infiltrate the damaged tissue to increase the myocardial tensile strength by deposition of extracellular matrix. Finally, new blood vessels are formed in the border zone, resulting in partial restoration of blood supply. If the mechanisms aimed at attenuating injury are not sufficient, ultimately heart failure supervenes. In the following paragraphs, the most interesting results on the role/regulation of miRNAs in these processes are reported and discussed in the view of possible therapeutic exploitations.
Cardiomyocyte cell death: apoptosis and necrosis
Ischemia and reperfusion are powerful inducers of cell death programs [49, 50]. The critical role of cardiac myocytes death in the progression towards cardiac dilation and heart failure indicates that improvement of myocyte survival could represent an important therapeutic strategy. Very few studies involve miRNAs in the autophagic death, in particular during I/R [51]. On the contrary, a large body of work suggests a role for miRNAs in the regulation of apoptosis (Fig. 1a). Intriguingly, miRNAs also regulate necrotic cell death.
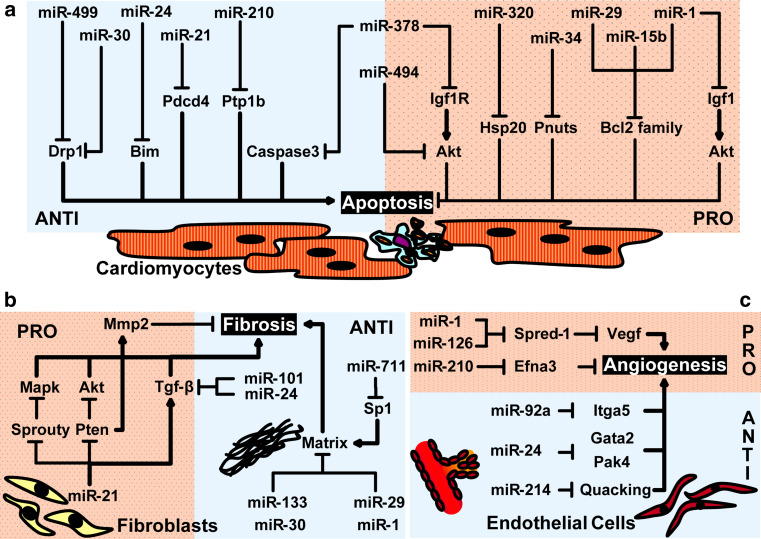
Fig. 1.
Target mRNAs and functional role of miRNAs related to myocardial ischemia in cardiomyocytes (a), fibroblasts (b), and endothelial cells (c). Arrows and bars at the end of the lines indicate whether the target is activated (arrow) or inhibited (bar). Dotted pink- and light blue-shaded regions indicate, respectively, stimulation or repression of a process (apoptosis, fibrosis, and angiogenesis) in myocardial ischemia
A number of miRNAs exert an antiapoptotic function by targeting important proapoptotic proteins (Fig. 1a). miR-499 and miR-30 family members diminish apoptosis in the injured heart by attenuating activation of dynamin-related protein-1 (Drp1) and thus inhibiting mitochondrial fission [52, 53]. Similarly, miR-24 and miR-21 inhibit cardiomyocyte apoptosis [54, 55], via the repression of the proapoptotic proteins Bim [54, 56] and programmed cell death 4 (Pdcd4) [57, 58]. Interestingly, ex vivo miR-24 enrichment, together with miR-21 and miR-221, improves the therapeutic potential of cardiacprogenitor cells upon transplantation in ischemic rodents [59]. Suppression of miR-24 increases cardiomyocytes apoptosis [60]. Nevertheless, miR-24 inhibition is beneficial for angiogenesis (see below), raising some concerns about its therapeutic use. Notably, miR-24 and miR-21 were shown to also reduce the rates of necrosis in ischemic cardiomyocyte cultures [56, 57]. Another important antiapoptotic miRNA is miR-210, which is considered the master hypoxamiR [61, 62]. In hypoxic cardiomyocytes, miR-210 is upregulated by p53 in a Hif-dependent, and by Akt in a Hif-independent way [63]. Increased survival of cardiomyocytes and of heart-engrafted bone marrow-derived stem cells in the ischemic myocardium is promoted by miR 210 by targeting, respectively, Ptp1b and caspase-8-associated protein 2 [64, 65]. Furthermore, miR-210 regulates mitochondrial metabolism by targeting crucial proteins of the electron transport chain, at least in cancer cells [66–68]. miR-210 may also have a previously unrecognized role in the metabolic adaptation to hypoxic conditions in the infarcted myocardium.
On the other hand, a number of miRNAs have been shown to exert proapoptotic effects by targeting key cardioprotective proteins (Fig. 1a), like, for example, heat-shock protein 20 (Hsp20), which is the target of miR-320 [69]. Among others, miR-34 family members promote growth arrest and apoptosis [70]. Indeed, therapeutic inhibition of miR-34 attenuates ischemia-induced remodeling and improves cardiac recovery [71]. A protein phosphatase 1 nuclear targeting subunit (Pnuts) was identified as a novel miR-34 target [72].
Bcl2 family is a major hub on which multiple signals converge. Indeed, the proapoptotic miR-29 targets Mcl-1, a Bcl2 family-member [73]. In addition, targeting miR-15 family members renders cultured cardiomyocytes resistant to hypoxic death and ameliorates cardiac response to myocardial ischemia by inhibiting repression of Bcl2 [74]. Finally, Bcl2, together with Hsp60 and Hsp70, is also the target of miR-1, which is upregulated in response to ischemic injury in vivo [75–77]. Akt signaling emerges as a further central node for improving cardioprotection against apoptosis and necrosis in myocardial ischemia and I/R. miR-1 as well as miR-378 both target the antiapoptotic IGF1 pathway [47, 48, 75] and are therefore involved in the apoptotic cell death induced by cardiac ischemia. However, the overexpression of miR-378 attenuates both apoptosis and necrosis in hypoxic cardiomyocytes by inhibiting caspase 3 [78]. Like miR-378, miR-494 lies at the edge between pro- and anti-apoptotic miRNAs, ultimately tilting the balance in favor of the activation of Akt [79].
Fibroblast proliferation and activation: fibrosis
Interstitial fibrosis is a major aspect of myocardial remodeling following MI, which contributes to loss of contractility and function. The extracellular deposition of collagen by fibroblasts contributes to this adverse remodeling after MI. Therefore, inhibiting fibroblasts proliferation, differentiation, and secretion of matrix proteins may be clinically relevant for the prevention and treatment of heart failure after MI. Specific antifibrotic drugs are not currently available. Therefore, efforts should be focused on exploring new possible therapeutic targets.
Interestingly, an anti-miR-21 was shown to prevent myocardial fibrosis in a model of pressure overload [80]. Indeed, miR-21 represses the expression of Sprouty, a negative regulator of Mapk, thus enhancing proliferation of fibroblasts [80, 81] (Fig. 1b). Moreover, miR-21-dependent targeting of sprouty homolog 1 (Spry1) and Pdcd4 was shown to promote the fibroblastoid phenotype in epicardial mesothelial cells undergoing epithelial-to-mesenchymal transition [82]. Moreover, phosphatase and tensin homologue (Pten) is a direct target of miR-21 [83] in cardiac fibroblasts. Pten upregulates the expression of matrix metalloproteinase 2 (Mmp2), promoting fibrosis in the infarcted heart. However, opposite results were obtained by Olson’s group [84], leaving still debated whether targeting miR-21 in the heart may be beneficial for the treatment of myocardial ischemia and infarction.
On the other hand, miR-133, miR-1, miR-30, and miR29 family members directly (or indirectly in the case of miR-711, which targets Sp1) downregulate key profibrotic proteins [85–88] (Fig. 1b). In particular, miR-29 family members are downregulated in the infarct area [86] and during collagen accumulation upon experimental fibrosis [89], while elevated miR-29 levels and repressed collagen synthesis follow stimulation with HGF [90], a powerful antifibrotic factor [46]. However, the proapoptotic effect exerted by miR-29 on cardiomyocytes generates an important criticism in the therapeutic use of miR-29 to reduce fibrosis in MI.
A major pathway implicated in cardiac fibrosis involves Tgf-β, which induces fibroblasts to synthesize and contract extracellular matrix. MicroRNAs targeting the Tgf-β pathway may offer a therapeutic tool to interfere with fibrosis following MI. In this respect, miR-21 has been proposed as a regulator of Tgf-β pathway. While inducing the proliferation of fibroblasts, miR-21 simultaneously increases matrix deposition. Notably, miR-21 itself is upregulated by Tgf-β [91]. Also, miR-24 and miR-101 have been suggested to influence Tgf-β pathway. Upregulation of miR-24, which is downregulated in MI heart, can reduce fibrosis by decreasing differentiation and migration of cardiac fibroblasts through the control of Tgf-β activation [92]. Overexpression of miR-101 suppresses both proliferation and collagen deposition in cardiac fibroblasts, resulting in inhibited post-infarct cardiac fibrosis and improved cardiac function in vivo, by targeting the Tgf-β pathway [93].
Endothelial cells proliferation and migration: neovascularization
Neoangiogenesis mainly occurs in the MI border zone to sustain cardiomyocytes survival and repair with oxygen and nutrients. A number of miRNAs are involved in vascular biology [94]. The most convincing evidence for the participation of miRNAs in the angiogenic response to myocardial ischemia concerns the proangiogenic miR-126, miR-1, and miR-210, and the antiangiogenic miR-92a, miR-24, and miR-214 (Fig. 1c).
MiR-126 is upregulated in the border zone of MI [95]. Consistently, neovascularization after MI is inhibited in miR-126 knockout mice [95]. The proangiogenic effect of miR-126 following myocardial infarction has been attributed to the repression of Spred-1 [95], an intracellular negative regulator of Mapks (Fig. 1c). Targeting of Spred1 leads to the increase in vascular endothelial growth factor (Vegf), favoring proliferation and migration of endothelial cells. Notably, miR-1 has also been recently shown to target Spred-1 [96], thus suggesting that the activation of Mapks in endothelial cells is a hub for the convergence of proangiogenic miRNA stimuli. Mesenchymal stem cells [97, 98] and endothelial progenitors [99] transduced with miR-126 have been proposed as a novel therapeutic approach for the improvement of post-ischemic angiogenesis. Finally, intramyocardial injection of a non-viral vector minicircle DNA carrying the miR-210 precursor reduces infarct size by enhancing angiogenesis [64], through the downregulation of ephrin-A3 (Efna3) [62]. Thus, delivery of proangiogenic miR-126 and miR-210 might be considered as a therapeutic approach in ischemic heart disease.
Conversely, miR-92a, miR-24 and miR-214 suppress neovascularization in myocardial infarction by inhibition of proangiogenic proteins, such as α5 integrin (Itga5) [100], Gata2 and Pak4 [101], and Quacking [102], respectively (Fig. 1c). Accordingly, blocking of miR-24 limits myocardial infarct size in mice [101] and increases angiogenesis and blood perfusion in the peri-infarct myocardium, by prompting endothelial cells survival, proliferation, and networking in capillary-like tubes [60].
Other miRNAs, such as miR-130, miR-320, miR-221/222, miR-296, and miR-378 have been shown to regulate physiological vascular function and tumor angiogenesis [36]. However, a function in the regulation of angiogenesis in cardiac ischemia has not yet been investigated.
MicroRNAs in heart failure
Ischemic cardiomyopathy is a common cause of congestive heart failure. In hypoxic conditions, the heart reactivates a fetal gene program, involving extensive remodeling, decreased aerobic metabolism, and increased cardiac efficiency. Among others, the fetal isoform of myosin heavy chain (β-MHC) and the natriuretic factor genes are invariably upregulated in the failing heart. However, this adaptive response is the first step of the subsequent heart failure. A subset of miRNAs which are up- and downregulated in experimental heart failure [31] are also modulated in failing human hearts [31, 103–106]. A number of miRNAs have been shown to be either upregulated (miR-21, miR-29b, miR-129, miR-210, miR-211, miR-212, miR-423, miR-199 family, miR-379, miR-503, and miR34b and c) or downregulated (miR-30, miR-182, and miR-526) [106, 107]. Consistent with the reexpression of the fetal gene program, a high degree of similarity was found between the miRNA expression patterns occurring in human failing hearts and those observed in human fetal hearts, thus indicating that a fetal miRNA program is also reactivated by cardiac stress [106].
Besides reactivation of fetal miRNAs and genes, impaired Ca2+ homeostasis is a hallmark of failing hearts. Myocardial sarcoplasmic reticulum Ca2+ ATPase 2a (SERCA2a) is downregulated in heart failure and SERCA2a gene therapy improves cardiac function in both animals and patients [108]. Intriguingly, miR-1 is downregulated in heart failure and SERCA2a gene therapy is able to restore miR-1 expression in heart failure via an Akt/Foxo3a-dependent pathway [109]. Notably, in the adult heart, both miR-1 and miR-133, which are in the same bicistronic unit, display an anti-hypertrophic activity [110, 111]. These data might encourage the use of miR-1 for treating myocardial ischemia; however, the negative effects exerted on cardiomyocytes survival might hamper any other beneficial effect.
The abnormal increase in intracellular Ca2+ observed during myocardial reperfusion is a major concern. In fact, it is thought to be the cause of cardiomyocyte death and consequent loss of cardiac function, leading to heart failure. Indeed, therapeutic miRNA-mediated modulation of Ca2+ handling may provide some cardioprotection against the paradoxical effects of reperfusion. In this respect, it was shown that miR-214 is upregulated during ischemic injury and heart failure [112], and that genetic deletion of miR-214 in mice worsens the heart response to I/R. The cardioprotective role of miR-214 during I/R injury is most likely due to the repression of sodium/calcium exchanger 1, a key regulator of Ca2+ influx, and to the repression of downstream apoptotic and necrotic effectors of Ca2+ signaling [112]. Among these, CaMKIIδ is a common target of both miR-214 and miR-145 [113]. MiR-145 concomitantly protects cardiomyocytes from ROS by targeting Bnip3 [114]. Boosting miR-214 and miR-145 levels to attenuate Ca2+ overload and cardiac cell death may provide a valuable therapeutic benefit for the treatment or prevention of heart failure after I/R injury. However, some caution must be adopted, due to the antiangiogenic effects of miR-214, which reduces sprouting of endothelial cells by targeting Quacking [115].
Future directions for therapy
miR-diagnosis and miR-drugs
The discovery of specific miRNAs as important regulators of the cardiac response to ischemia has opened new perspectives for clinical research. Recent results obtained in cancer suggest that the profiles of blood circulating miRNAs might mirror the changes observed in the cancerous tissue [116]. This concept has also proved valid in heart failure [117], and the significance of circulating miRNAs in comparison to conventional biomarkers has just begun to be investigated in the cardiovascular field [118, 119]. Circulating cardiac miRNAs are indeed released as a specific response to myocardial injury and can be used in the diagnosis of MI [120–124]. Recently, it was discovered that most of the extracellular miRNAs circulate in the blood of both healthy and diseased patients as secretory molecules, contained in apoptotic bodies, microvesicles, and exosomes or bound to RNA-binding proteins [16]. New challenges for miRNA biology include unraveling the secretory mechanism, as well as the significance and biological function of extracellular miRNAs. Moreover, large sample sizes are required to give an accurate estimate of the diagnostic and prognostic utility of miRNAs as CVD biomarkers. New frontiers are expected to be delineated for miR-diagnosis.
As important regulators of heart function, miRNAs represent attractive targets/tools for treating heart disease with miR drugs [125–131]. The momentum in miRNA field is hyped by the large investment announced by the Colorado-based Miragen, which will initially focus on identifying targets related to CVD, primarily heart failure. A number of in vivo pre-clinical studies have been conducted using anti-miRNAs strategies, such as antagomiRs, 2’O-methyl-modified cholesterol-conjugated single-strand RNA oligos with perfect complementarity to target miRNAs. Treatment with antagomiRs in mice has been used to prove the role of specific miRNAs in determined phenotypes [69, 111]. For example, a miR-21-directed antagomiR has been used to prevent myocardial fibrosis after pressure overload [80]. It must be noted that miR-21 is expressed in a broad array of tissues, including the vasculature, thus making the discrimination between primary and secondary cardiac effects induced by intravenous injection of the antagomiR difficult. The use of highly tissue-specific adeno-associated viral vectors should be taken into account in order to improve delivery and specificity. In this perspective, AAV9 might represent the best choice for cardiac interventions [132]. Mimics of angiomiRs (or antagomiRs of anti-angiomiRs) might also be used to improve neoangiogenesis in cardiac ischemia [64, 95].
Another intriguing therapeutic approach might be the use of miRNAs in preconditioning. Indeed, it is possible to stimulate the intrinsic resistance to hypoxia through repeated short episodes of ischemia. During normoxia, Hif1α is targeted by miR-199a, while, during hypoxia, miR-199a is downregulated (by a still undefined post-transcriptional mechanism), thus contributing to the expression of Hif1α [133]. The therapeutic knockdown of miR-199a during normoxia might therefore be used to mirror hypoxic preconditioning and protect cardiomyocytes against subsequent hypoxic damage, as suggested by recent results [134].
The drugs which are typically administered to treat heart failure mostly aim at reducing blood pressure and improve heart pumping. The rapid advancement in the identification of new miRNAs associated with heart failure will certainly increase our arsenal for the treatment of such a threatening disease.
MicroRNAs and regenerative applications
Regeneration of an injured heart can be carried out by new muscle cells coming from two sources: replication of preexisting cardiomyocytes and expansion, followed by proper differentiation of stem cells or precursors (resident or injected). The field of regenerative medicine is expanding. The exploitation of miRNAs to this purpose is a certain target for future research.
Reactivation of cardiomyocytes proliferation
In mammals, the transition from the proliferative to the hypertrophic phenotype is strictly regulated [135]. In the neonate, 3–15 % of the cardiomyocytes are still cycling [136–138]. However, cardiomyocyte proliferation declines after birth [139]. After cardiac injury, a low level of cardiomyocyte replication occurs, suggestive of a partial attempt of the myocardium to regenerate [140–143]. Nevertheless, the proliferative capacity of adult cardiomyocytes is limited and insufficient to replace the ≈1 billion cardiomyocytes typically lost in MI. Recent studies have provided evidence for supporting therapeutic reactivation of cardiomyocyte proliferation by means of miRNAs.
MiR-15 family has an important role in governing cardiomyocyte cell cycle withdrawal. During neonatal development, miR-15 family is upregulated in the heart, in concomitance with the irreversible exit of cardiomyocytes from the cell cycle [144]. Consistently, post-natal knockdown of miR-15 family members through in vivo delivery of anti-miR oligonucleotides has been associated with an increased number of mitotic cardiomyocytes [144]. Moreover, overexpression of miR-195 (member of miR-15 family) in the neonatal heart reduced the regenerative potential after myocardial ischemia [145].
Also, the depletion of miR-133 (which has an important role in the proliferation and differentiation of cardiac progenitors during cardiomyogenesis [29, 146]), enhances the cardiac regenerative responses in zebrafish cardiomyocytes after injury [147]. Indeed, cardiac regeneration in zebrafish, albeit unclear, appears to be at least partially based on cardiomyocyte proliferation [148–150]. In contrast, transgenic overexpression of miR-17-92 in cardiomyocytes is sufficient to induce cardiomyocyte proliferation in embryonic, post-natal, and adult hearts. Moreover, overexpression of miR-17-92 in adult cardiomyocytes protects the heart from myocardial infarction-induced injury [151].
The recent publication from Eulalio et al. [152] produced excitement and open questions [153, 154]. Giacca’s group has shown that hsa-miR-590 and hsa-miR-199a can trigger cell cycle reentry of adult cardiomyocytes ex vivo and promote cardiomyocyte proliferation in both neonatal and adult animals. When these miRNAs are administered to ischemic hearts, they induce the replication of cardiomyocytes and stimulate repair of damage through the formation of new heart cells [152].
These observations hint at the possibility of identifying and targeting the molecular regulators that maintain cardiomyocytes in the quiescent state after injury. However, care must be taken in forcing cell cycle progression in cardiomyocytes, since these may undergo hypertrophy or apoptosis, both detrimental to cardiac function [139, 155]. Thus, therapeutic strategies aimed at stimulating cardiomyocyte proliferation should require the concomitant expression of cell cycle regulators and cardioprotective molecules to ensure full activation of the cell cycle and protection from apoptosis. In this sense, the peculiar capacity of miRNAs to exert multiple coordinate effects represents a clear advantage in preventing catastrophic mitosis.
Stimulation of cardiac progenitors
The identification of cardiac cells with stem cell properties changed the paradigm of the heart as a post-mitotic organ [156–158]. These cells proliferate and differentiate into cardiomyocytes and endothelial and vascular smooth muscle cells, providing for cardiac homeostasis and regeneration. However, the regenerative potential of resident cardiac progenitors is limited. Some recent reviews extensively explored the possibility of manipulating this regenerative potential by the use of miRNAs [153, 154, 159, 160]. Indeed, miRNAs act as regulators of proliferation and differentiation of adult cardiac stem cells and progenitors [39, 161–164].
The use of exogenous stem cells for cellular therapy constitutes a major avenue of investigation [158]. The potential of miRNAs in stimulating expansion and differentiation of embryonic and induced pluripotent stem cells has been reviewed extensively elsewhere [153]. Survival and functional integration of transplanted cells as well as their proper differentiation are major concerns to improve cardiac function. Indeed, one of the major hurdles to achieve long-term improvement of heart function through stem cell therapies is the low survival rate of the injected stem cells in the hostile environment of the damaged myocardium. A miRNA cocktail has recently been shown to increase viability of transplanted cells in the infarcted heart by targeting Bim and, thus, induce better recovery [59].
An important regulator of cardiac progenitor cells (CPCs) differentiation is miR-499. Although barely detectable in undifferentiated precursors, it is strongly induced in post-mitotic cardiomyocytes. In human CPCs, miR-499 enhances cardiogenesis by repressing Sox6 and Rod1. Importantly, CPCs overexpressing miR-499 have increased potential to regenerate the damaged myocardium in an animal model of myocardial infarction [17].
Notably, miRNAs can act as paracrine messengers. For example, miR-499 has been shown to traverse gap junction channels and translocate to structurally coupled CPCs favoring their differentiation into functionally competent cells [17]. An inverse flux of miR-210 from transplanted stem cells of mesenchymal origin to host cardiomyocytes has been shown to functionally recover the ischemic heart, meanwhile promoting post-engraftment survival [18].
Conclusions and take-away message
Many miRNAs have been implicated in the control of cardiac apoptosis, fibrosis, and neovascularization following myocardial ischemia and I/R injury and in the adaptation of the heart to damage. Therefore, analyzing (miR-diagnosis) and manipulating (miR-drugs) miRNA biology represents an attractive, albeit ambitious, approach for diagnosis and therapeutics. Moreover, new evidence supports a role for miRNAs in the induction of proliferation of cardiomyocytes and improvement of survival, renewal, and differentiation of precursor and stem cells. One major advantage in using miRNAs is certainly the possibility to simultaneously target multiple proteins and even entire pathogenic pathways. The main problem is to globally obtain a coherent and beneficial effect. Indeed, in some cases, this huge potential might represent a double-edged sword. As emerges from Table 1, some miRNAs exerting a positive effect on one particular cell type might simultaneously have a deleterious influence on another cardiac cell population. The outbalancing force should therefore be identified with caution. Indeed, such a spectrum of actions implicates the need for a comprehensive knowledge of miRNAs networks and interplay in order to predict (and prevent) possible side effects.
Table 1.
miRNAs associated with cardiac ischemia which might be manipulated for attenuating several aspects of myocardial injury (apoptosis, necrosis, fibrosis, and neovascularization) or improving regenerative therapies
| miRNA | Attenuation of myocardial injury | Regenerative applications | |||
|---|---|---|---|---|---|
| Cardiomyocyte apoptosis | Necrosis | Fibrosis | Neovascularization | ||
| miR-1 | Repression of antiapoptotic Bcl2 [76], Igf1 [47, 75], Hsp60, Hsp70 [77] | Targeting of profibrotic Fibullin-2 [87] | Enhanced tubulogenesis by repressing Spred-1 [96] | ||
| miR-15 | Repression of antiapoptotic Bcl2 [74] | Inhibition of cardiomyocyte proliferation [144, 145] | |||
| miR-21 | Protection against apoptosis by targeting Pdcd4 [55, 57, 58] | Reduced necrosis in cultured cardiomyocytes [57] | Promotion of fibroblast proliferation and inhibition of metalloprotease [80, 81, 83], promotion of TGFβ [91] | Improved engraftment of transplanted cardiac progenitor cells [59] | |
| miR-24 | Suppression of apoptosis, by repression of Bim [54] | Reduced necrosis in cultured ischemic cardiomyocytes [56] | Decreased fibroblast differentiation and migration by targeting Tgfβ [92] | Inhibition of proangiogenic Gata2 and Pak4 [101] | Improved engraftment of transplanted cardiac progenitor cells [59] |
| miR-29 | Proapoptotic by targeting Mcl-1 [73] | Targeting of fibrotic proteins[86] | |||
| miR-30 | Inhibition of mitochondrial fission by suppressing Drp1 [53] | Decreased production of collagens[85] | |||
| miR-34 | Proapoptotic targeting of Pnuts [72] | ||||
| miR-92 | Inhibition of the proangiogenic Itga5 [100] | Induction of cardiomyocyte proliferation [151] | |||
| miR-101 | Decreased fibroblast proliferation and collagen deposition by targeting Tgf-β signaling [93] | ||||
| miR-126 | Increased proliferation and migration of endothelial cells by repressing Spred-1 [95] | Increased angiogenesis after mesenchymal stem cells [97, 98] and endothelial progenitors [99] trasplantation | |||
| miR-133 | Repressing proapoptotic Caspase 9 [77] | Decreased production of collagens [85] | Increased regeneration in zebrafish [147] | ||
| miR-145 | Attenuation of Calcium overload by repressing CaMKIIδ [113] and Bnip3 [114] | ||||
| miR-199 and 590 | Induction of cell cycle reentry [152] | ||||
| miR-210 | Inhibition of apoptosis by targeting Ptp1b [64] | Enhanced angiogenesis by repression of Efna3 [62] | Increased engraftment of bone marrow-derived stem cells [65] and increased survival of mesenchymal stem cells [18] | ||
| miR-214 | Attenuation of calcium overload by repressing Ncx1 and CaMKIIδ [112] | Attenuation of calcium overload [112] | Inhibition of endothelial cells sprouting [102] | ||
| miR-221 | Improved engraftment of transplanted cardiac progenitor cells [59] | ||||
| miR-320 | Proapoptotic by targeting Hsp20 [69] | ||||
| miR-378 | Targeting antiapoptotic Igf1R [48] but also apoptotic Caspase 3 [78] | Inhibition of necrosis in hypoxic cardiomyocytes [78] | |||
| miR-494 | Activation of Akt [79] | Reduced necrotic death after I/R [79] | |||
| miR-499 | Inhibition of mitochondrial fission [52] | Increased regenerative potential and differentiation of cardiac progenitors[17] | |||
| miR-711 | Targeting Sp1, reducing collagen production[88] | ||||
Detrimental effects are shown in italics
Acknowledgments
This work was supported by funds from the Association Francaise contre les Myopathies (AFM) to T.C. This work and a fellowship to V.S. were made possible by EU grant FP7-SST 265772 EM-Safety to A.P.
References
- 1.Roger VL, Go AS, Lloyd-Jones DM, Benjamin EJ, Berry JD, Borden WB, et al. Heart disease and stroke statistics-2012 update a report from the American Heart Association. Circulation. 2012;125:E2–E220. doi: 10.1161/CIR.0b013e31823ac046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Buja LM, Weerasinghe P. Unresolved issues in myocardial reperfusion injury. Cardiovasc Pathol. 2010;19:29–35. doi: 10.1016/j.carpath.2008.10.001. [DOI] [PubMed] [Google Scholar]
- 3.de Groot H, Rauen U. Ischemia-reperfusion injury: processes in pathogenetic networks: a review. Transpl Proc. 2007;39:481–484. doi: 10.1016/j.transproceed.2006.12.012. [DOI] [PubMed] [Google Scholar]
- 4.Frangogiannis NG. Regulation of the inflammatory response in cardiac repair. Circ Res. 2012;110:159–173. doi: 10.1161/CIRCRESAHA.111.243162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Nanni L, Romualdi C, Maseri A, Lanfranchi G. Differential gene expression profiling in genetic and multifactorial cardiovascular diseases. J Mol Cell Cardiol. 2006;41:934–948. doi: 10.1016/j.yjmcc.2006.08.009. [DOI] [PubMed] [Google Scholar]
- 6.Huang XH, Pan W, Grindle S, Han XQ, Chen YJ, Park SJ, et al. A comparative study of discriminating human heart failure etiology using gene expression profiles. BMC Bioinform. 2005;6:205–218. doi: 10.1186/1471-2105-6-205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Sanoudou D, Vafiadaki E, Arvanitis DA, Kranias E, Kontrogianni-Konstantopoulos A. Array lessons from the heart: focus on the genome and transcriptome of cardiomyopathies. Physiol Genomics. 2005;21:131–143. doi: 10.1152/physiolgenomics.00259.2004. [DOI] [PubMed] [Google Scholar]
- 8.Gabrielsen A, Lawler PR, Wang YZ, Steinbruchel D, Blagoja D, Paulsson-Berne G, et al. Gene expression signals involved in ischemic injury, extracellular matrix composition and fibrosis defined by global mRNA profiling of the human left ventricular myocardium. J Mol Cell Cardiol. 2007;42:870–883. doi: 10.1016/j.yjmcc.2006.12.016. [DOI] [PubMed] [Google Scholar]
- 9.Kittleson MM, Minhas KM, Irizarry RA, Ye SQ, Edness G, Breton E, et al. Gene expression analysis of ischemic and nonischemic cardiomyopathy: shared and distinct genes in the development of heart failure. Physiol Genomics. 2005;21:299–307. doi: 10.1152/physiolgenomics.00255.2004. [DOI] [PubMed] [Google Scholar]
- 10.Prat-Vidal C, Galvez-Monton C, Nonell L, Puigdecanet E, Astier L, Sole F, et al. Identification of temporal and region-specific myocardial gene expression patterns in response to infarction in swine. PLoS ONE. 2013;8:e54785. doi: 10.1371/journal.pone.0054785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Mendell JT, Olson EN. MicroRNAs in stress signaling and human disease. Cell. 2012;148:1172–1187. doi: 10.1016/j.cell.2012.02.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Parker R, Song HW. The enzymes and control of eukaryotic mRNA turnover. Nat Struct Mol Biol. 2004;11:121–127. doi: 10.1038/nsmb724. [DOI] [PubMed] [Google Scholar]
- 13.Filipowicz W, Bhattacharyya SN, Sonenberg N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet. 2008;9:102–114. doi: 10.1038/nrg2290. [DOI] [PubMed] [Google Scholar]
- 14.Hwang HW, Wentzel EA, Mendell JT. A hexanucleotide element directs microRNA nuclear import. Science. 2007;315:97–100. doi: 10.1126/science.1136235. [DOI] [PubMed] [Google Scholar]
- 15.Barrey E, Saint-Auret G, Bonnamy B, Damas D, Boyer O, Gidrol X. Pre-microRNA and mature microRNA in human mitochondria. PLoS ONE. 2011;6:e20220. doi: 10.1371/journal.pone.0020220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kosaka N, Ochiya T. Unraveling the mystery of cancer by secretory microRNA: horizontal microRNA transfer between living cells. Front Genet. 2011;2:97. doi: 10.3389/fgene.2011.00097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hosoda T, Zheng HQ, Cabral-da-Silva M, Sanada F, Ide-Iwata N, Ogorek B, et al. Human cardiac stem cell differentiation is regulated by a mircrine mechanism. Circulation. 2011;123(12):1287–1296. doi: 10.1161/CIRCULATIONAHA.110.982918. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 18.Kim HW, Jiang SJ, Ashraf M, Haider KH. Stem cell-based delivery of Hypoxamir-210 to the infarcted heart: implications on stem cell survival and preservation of infarcted heart function. J Mol Med (Berl) 2012;90:997–1010. doi: 10.1007/s00109-012-0920-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Liu JD, Valencia-Sanchez MA, Hannon GJ, Parker R. MicroRNA-dependent localization of targeted mRNAs to mammalian P-bodies. Nat Cell Biol. 2005;7(7):719–723. doi: 10.1038/ncb1274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 2006;34:D140–D144. doi: 10.1093/nar/gkj112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. doi: 10.1016/s0092-8674(04)00045-5. [DOI] [PubMed] [Google Scholar]
- 22.Friedman Y, Balaga O, Linial M. Working together: combinatorial regulation by microRNAs. Adv Exp Med Biol. 2013;774:317–337. doi: 10.1007/978-94-007-5590-1_16. [DOI] [PubMed] [Google Scholar]
- 23.Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature. 2005;433:769–773. doi: 10.1038/nature03315. [DOI] [PubMed] [Google Scholar]
- 24.Boettger T, Braun T. A new level of complexity: the role of microRNAs in cardiovascular development. Circ Res. 2012;110:1000–1013. doi: 10.1161/CIRCRESAHA.111.247742. [DOI] [PubMed] [Google Scholar]
- 25.Divakaran V, Mann DL. The emerging role of microRNAs in cardiac remodeling and heart failure. Circ Res. 2008;103(10):1072–1083. doi: 10.1161/CIRCRESAHA.108.183087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Han M, Toli J, Abdellatif M. MicroRNAs in the cardiovascular system. Curr Opin Cardiol. 2011;26:181–189. doi: 10.1097/HCO.0b013e328345983d. [DOI] [PubMed] [Google Scholar]
- 27.Kukreja RC, Yin C, Salloum FN. MicroRNAs: new players in cardiac injury and protection. Mol Pharmacol. 2011;80:558–564. doi: 10.1124/mol.111.073528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ono K, Kuwabara Y, Han JH. MicroRNAs and cardiovascular diseases. FEBS J. 2011;278:1619–1633. doi: 10.1111/j.1742-4658.2011.08090.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zhao Y, Samal E, Srivastava D. Serum response factor regulates a muscle-specific microRNA that targets Hand2 during cardiogenesis. Nature. 2005;436:214–220. doi: 10.1038/nature03817. [DOI] [PubMed] [Google Scholar]
- 30.Tatsuguchi M, Seok HY, Callis TE, Thomson JM, Chen JF, Newman M, et al. Expression of microRNAs is dynamically regulated during cardiomyocyte hypertrophy. J Mol Cell Cardiol. 2007;42:1137–1141. doi: 10.1016/j.yjmcc.2007.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.van Rooij E, Sutherland LB, Liu N, Williams AH, McAnally J, Gerard RD, et al. A signature pattern of stress-responsive microRNAs that can evoke cardiac hypertrophy and heart failure. Proc Natl Acad Sci USA. 2006;103:18255–18260. doi: 10.1073/pnas.0608791103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.van Rooij E, Sutherland LB, Qi XX, Richardson JA, Hill J, Olson EN. Control of stress-dependent cardiac growth and gene expression by a microRNA. Science. 2007;316:575–579. doi: 10.1126/science.1139089. [DOI] [PubMed] [Google Scholar]
- 33.Margulies KB, Matiwala S, Cornejo C, Olsen H, Craven WA, Bednarik D. Mixed messages: transcription patterns in failing and recovering human myocardium. Circ Res. 2005;96:592–599. doi: 10.1161/01.RES.0000159390.03503.c3. [DOI] [PubMed] [Google Scholar]
- 34.Matkovich SJ, Van Booven DJ, Youker KA, Torre-Amione G, Diwan A, Eschenbacher WH, et al. Reciprocal regulation of myocardial microRNAs and messenger RNA in human cardiomyopathy and reversal of the microRNA signature by biomechanical support. Circulation. 2009;119:1263–1271. doi: 10.1161/CIRCULATIONAHA.108.813576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Liu N, Olson EN. MicroRNA regulatory networks in cardiovascular development. Dev Cell. 2010;18:510–525. doi: 10.1016/j.devcel.2010.03.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Small EM, Frost RJA, Olson EN. MicroRNAs add a new dimension to cardiovascular disease. Circulation. 2010;121:1022–1032. doi: 10.1161/CIRCULATIONAHA.109.889048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.van Rooij E, Liu N, Olson EN. MicroRNAs flex their muscles. Trends Genet. 2008;24:159–166. doi: 10.1016/j.tig.2008.01.007. [DOI] [PubMed] [Google Scholar]
- 38.van Rooij E, Quiat D, Johnson BA, Sutherland LB, Qi XX, Richardson JA, et al. A family of microRNAs encoded by myosin genes governs myosin expression and muscle performance. Dev Cell. 2009;17:662–673. doi: 10.1016/j.devcel.2009.10.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Chen JF, Mandel EM, Thomson JM, Wu QL, Callis TE, Hammond SM, et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat Genet. 2006;38:228–233. doi: 10.1038/ng1725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Czubryt MP, McAnally J, Fishman GI, Olson EN. Regulation of peroxisome proliferator-activated receptor gamma coactivator 1 alpha (PGC-1 alpha) and mitochondrial function by MEF2 and HDAC5. Proc Natl Acad Sci USA. 2003;100:1711–1716. doi: 10.1073/pnas.0337639100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Liu N, Bezprozvannaya S, Williams AH, Qi XX, Richardson JA, Bassel-Duby R, et al. microRNA-133a regulates cardiomyocyte proliferation and suppresses smooth muscle gene expression in the heart. Gene Dev. 2008;22:3242–3254. doi: 10.1101/gad.1738708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.McCarthy JJ, Esser KA. MicroRNA-1 and microRNA-133a expression are decreased during skeletal muscle hypertrophy. J Appl Phys. 2007;102:306–313. doi: 10.1152/japplphysiol.00932.2006. [DOI] [PubMed] [Google Scholar]
- 43.Hua Z, Lv Q, Ye WB, Wong CKA, Cai GP, Gu DY, et al. miRNA-directed regulation of VEGF and other angiogenic factors under hypoxia. PLoS ONE. 2006;1:e116. doi: 10.1371/journal.pone.0000116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Migliore C, Martin V, Leoni VP, Restivo A, Atzori L, Petrelli A, et al. MiR-1 downregulation cooperates with MACC1 in promoting MET overexpression in human colon cancer. Clin Cancer Res. 2012;18:737–747. doi: 10.1158/1078-0432.CCR-11-1699. [DOI] [PubMed] [Google Scholar]
- 45.Leo C, Sala V, Morello M, Chiribiri A, Riess I, Mancardi D, et al. Activated Met signalling in the developing mouse heart leads to cardiac disease. PLoS ONE. 2011;6:e14675. doi: 10.1371/journal.pone.0014675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Sala V, Crepaldi T. Novel therapy for myocardial infarction: can HGF/Met be beneficial? Cell Mol Life Sci. 2011;68:1703–1717. doi: 10.1007/s00018-011-0633-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Elia L, Contu R, Quintavalle M, Varrone F, Chimenti C, Russo MA, et al. Reciprocal regulation of microRNA-1 and insulin-like growth factor-1 signal transduction cascade in cardiac and skeletal muscle in physiological and pathological conditions. Circulation. 2009;120:2377–2385. doi: 10.1161/CIRCULATIONAHA.109.879429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Knezevic I, Patel A, Sundaresan NR, Gupta MP, Solaro RJ, Nagalingam RS, et al. A novel cardiomyocyte-enriched microRNA, miR-378, targets insulin-like growth factor 1 receptor Implications in postnatal cardiac remodeling and cell survival. J Biol Chem. 2012;287:12913–12926. doi: 10.1074/jbc.M111.331751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Eltzschig HK, Eckle T. Ischemia and reperfusion-from mechanism to translation. Nat Med. 2011;17:1391–1401. doi: 10.1038/nm.2507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Oerlemans MIFJ, Koudstaal S, Chamuleau SA, de Kleijn DP, Doevendans PA, Sluijter JPG. Targeting cell death in the reperfused heart: pharmacological approaches for cardioprotection. Int J Cardiol. 2013;165:410–422. doi: 10.1016/j.ijcard.2012.03.055. [DOI] [PubMed] [Google Scholar]
- 51.Xiao J, Zhu XY, He B, Zhang YF, Kang B, Wang ZN, et al. miR-204 regulates cardiomyocyte autophagy induced by ischemia-reperfusion through LC3-II. J Biomed Sci. 2011;18:35. doi: 10.1186/1423-0127-18-35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Wang JX, Jiao JQ, Li QA, Long B, Wang K, Liu JP, et al. miR-499 regulates mitochondrial dynamics by targeting calcineurin and dynamin-related protein-1. Nat Med. 2011;17:71–78. doi: 10.1038/nm.2282. [DOI] [PubMed] [Google Scholar]
- 53.Li JC, Donath S, Li YR, Qin D, Prabhakar BS, Li PF. miR-30 regulates mitochondrial fission through targeting p53 and the dynamin-related protein-1 pathway. PLoS Genet. 2010;6:e1000795. doi: 10.1371/journal.pgen.1000795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Qian L, van Laake LW, Huang Y, Liu SY, Wendland MF, Srivastava D. miR-24 inhibits apoptosis and represses Bim in mouse cardiomyocytes. J Exp Med. 2011;208:549–560. doi: 10.1084/jem.20101547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Dong SM, Cheng YH, Yang J, Li JY, Liu XJ, Wang XB, et al. MicroRNA expression signature and the role of microRNA-21 in the early phase of acute myocardial infarction. J Biol Chem. 2009;284:29514–29525. doi: 10.1074/jbc.M109.027896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Li DF, Tian J, Guo X, Huang LM, Xu Y, Wang CC, et al. Induction of microRNA-24 by HIF-1 protects against ischemic injury in rat cardiomyocytes. Physiol Res. 2012;61:555–565. doi: 10.33549/physiolres.932270. [DOI] [PubMed] [Google Scholar]
- 57.Cheng YH, Liu XJ, Zhang SO, Lin Y, Yang J, Zhang CX. MicroRNA-21 protects against the H2O2-induced injury on cardiac myocytes via its target gene PDCD4. J Mol Cell Cardiol. 2009;47:5–14. doi: 10.1016/j.yjmcc.2009.01.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Cheng YH, Zhu P, Yang JA, Liu XJ, Dong SM, Wang XB, et al. Ischaemic preconditioning-regulated miR-21 protects heart against ischaemia/reperfusion injury via anti-apoptosis through its target PDCD4. Cardiovasc Res. 2010;87:431–439. doi: 10.1093/cvr/cvq082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Hu SJ, Huang M, Nguyen PK, Gong YQ, Li ZJ, Jia FJ, et al. Novel microRNA prosurvival cocktail for improving engraftment and function of cardiac progenitor cell transplantation. Circulation. 2011;124:S27–S34. doi: 10.1161/CIRCULATIONAHA.111.017954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Meloni M, Marchetti M, Garner K, Littlejohns B, Sala-Newby G, Xenophontos N, et al. Local inhibition of microRNA-24 improves reparative angiogenesis and left ventricle remodeling and function in mice with myocardial infarction. Mol Ther. 2013 doi: 10.1038/mt.2013.89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Chan YC, Banerjee J, Choi SY, Sen CK. miR-210: the master hypoxamir. Microcirculation. 2012;19:215–223. doi: 10.1111/j.1549-8719.2011.00154.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Fasanaro P, D’Alessandra Y, Di Stefano V, Melchionna R, Romani S, Pompilio G, et al. MicroRNA-210 modulates endothelial cell response to hypoxia and inhibits the receptor tyrosine kinase ligand Ephrin-A3. J Biol Chem. 2008;283:15878–15883. doi: 10.1074/jbc.M800731200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Mutharasan RK, Nagpal V, Ichikawa Y, Ardehali H. microRNA-210 is upregulated in hypoxic cardiomyocytes through Akt- and p53-dependent pathways and exerts cytoprotective effects. Am J Physiol Heart Circ Physiol. 2011;301:H1519–H1530. doi: 10.1152/ajpheart.01080.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hu SJ, Huang M, Li ZJ, Jia FJ, Ghosh ZM, Lijkwan MA, et al. MicroRNA-210 as a novel therapy for treatment of ischemic heart disease. Circulation. 2010;122:S124–S131. doi: 10.1161/CIRCULATIONAHA.109.928424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Kim HW, Haider HK, Jiang SJ, Ashraf M. Ischemic preconditioning augments survival of stem cells via miR-210 Expression by targeting caspase-8-associated protein 2. J Biol Chem. 2009;284:33161–33168. doi: 10.1074/jbc.M109.020925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Chan SY, Zhang YY, Hemann C, Mahoney CE, Zweier JL, Loscalzo J. MicroRNA-210 controls mitochondrial metabolism during hypoxia by repressing the iron-sulfur cluster assembly proteins ISCU1/2. Cell Metab. 2009;10:273–284. doi: 10.1016/j.cmet.2009.08.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Chen Z, Li Y, Zhang H, Huang P, Luthra R. Hypoxia-regulated microRNA-210 modulates mitochondrial function and decreases ISCU and COX10 expression. Oncogene. 2010;29:4362–4368. doi: 10.1038/onc.2010.193. [DOI] [PubMed] [Google Scholar]
- 68.Favaro E, Ramachandran A, McCormick R, Gee H, Blancher C, Crosby M, et al. MicroRNA-210 regulates mitochondrial free radical response to hypoxia and krebs cycle in cancer cells by targeting iron sulfur cluster protein ISCU. PLoS ONE. 2010;5:e10345. doi: 10.1371/journal.pone.0010345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Ren XP, Wu JH, Wang XH, Sartor MA, Qian J, Jones K, et al. MicroRNA-320 is involved in the regulation of cardiac ischemia/reperfusion injury by targeting heat-shock protein 20. Circulation. 2009;119:2357–2366. doi: 10.1161/CIRCULATIONAHA.108.814145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Hermeking H. p53 enters the microRNA world. Cancer Cell. 2007;12:414–418. doi: 10.1016/j.ccr.2007.10.028. [DOI] [PubMed] [Google Scholar]
- 71.Bernardo BC, Gao XM, Winbanks CE, Boey EJH, Tham YK, Kiriazis H, et al. Therapeutic inhibition of the miR-34 family attenuates pathological cardiac remodeling and improves heart function. Proc Natl Acad Sci USA. 2012;109:17615–17620. doi: 10.1073/pnas.1206432109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Boon RA, Iekushi K, Lechner S, Seeger T, Fischer A, Heydt S, et al. MicroRNA-34a regulates cardiac ageing and function. Nature. 2013;495:107–110. doi: 10.1038/nature11919. [DOI] [PubMed] [Google Scholar]
- 73.Ye YM, Hu ZY, Lin Y, Zhang CF, Perez-Polo JR. Downregulation of microRNA-29 by antisense inhibitors and a PPAR-gamma agonist protects against myocardial ischaemia-reperfusion injury. Cardiovasc Res. 2010;87:535–544. doi: 10.1093/cvr/cvq053. [DOI] [PubMed] [Google Scholar]
- 74.Hullinger TG, Montgomery RL, Seto AG, Dickinson BA, Semus HM, Lynch JM, et al. Inhibition of miR-15 protects against cardiac ischemic injury. Circ Res. 2012;110:71–81. doi: 10.1161/CIRCRESAHA.111.244442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Shan ZX, Lin QX, Fu YH, Deng CY, Zhou ZL, Zhu JN, et al. Upregulated expression of miR-1/miR-206 in a rat model of myocardial infarction. Biochem Bioph Res Co. 2009;381:597–601. doi: 10.1016/j.bbrc.2009.02.097. [DOI] [PubMed] [Google Scholar]
- 76.Tang YH, Zheng JY, Sun Y, Wu ZG, Liu ZM, Huang GZ. MicroRNA-1 regulates cardiomyocyte apoptosis by targeting Bcl-2. Int Heart J. 2009;50:377–387. doi: 10.1536/ihj.50.377. [DOI] [PubMed] [Google Scholar]
- 77.Xu CQ, Lu YJ, Pan ZW, Chu WF, Luo XB, Lin HX, et al. The muscle-specific microRNAs miR-1 and miR-133 produce opposing effects on apoptosis by targeting HSP60, HSP70 and caspase-9 in cardiomyocytes. J Cell Sci. 2011;124:3187. doi: 10.1242/jcs.098830. [DOI] [PubMed] [Google Scholar]
- 78.Fang J, Song XW, Tian J, Chen HY, Li DF, Wang JF, et al. Overexpression of microRNA-378 attenuates ischemia-induced apoptosis by inhibiting caspase-3 expression in cardiac myocytes. Apoptosis. 2012;17:410–423. doi: 10.1007/s10495-011-0683-0. [DOI] [PubMed] [Google Scholar]
- 79.Wang XH, Zhang XW, Ren XP, Chen J, Liu HZ, Yang JQ, et al. MicroRNA-494 targeting both proapoptotic and antiapoptotic proteins protects against ischemia/reperfusion-induced cardiac injury. Circulation. 2010;122:1308–1318. doi: 10.1161/CIRCULATIONAHA.110.964684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Thum T, Gross C, Fiedler J, Fischer T, Kissler S, Bussen M, et al. MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signaling in fibroblasts. Nature. 2008;456:980–984. doi: 10.1038/nature07511. [DOI] [PubMed] [Google Scholar]
- 81.Sayed D, Rane S, Lypowy J, He MZ, Chen IY, Vashistha H, et al. MicroRNA-21 targets sprouty2 and promotes cellular outgrowths. Mol Biol Cell. 2008;19:3272–3282. doi: 10.1091/mbc.E08-02-0159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Bronnum H, Andersen DC, Schneider M, Sandberg MB, Eskildsen T, Nielsen SB, et al. miR-21 promotes fibrogenic epithelial-to-mesenchymal transition of epicardial mesothelial cells involving programmed cell death 4 and sprouty-1. PLoS ONE. 2013;8(2):e56280. doi: 10.1371/journal.pone.0056280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Roy S, Khanna S, Hussain SRA, Biswas S, Azad A, Rink C, et al. MicroRNA expression in response to murine myocardial infarction: miR-21 regulates fibroblast metalloprotease-2 via phosphatase and tensin homologue. Cardiovasc Res. 2009;82:21–29. doi: 10.1093/cvr/cvp015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Patrick DM, Montgomery RL, Qi XX, Obad S, Kauppinen S, Hill JA, et al. Stress-dependent cardiac remodeling occurs in the absence of microRNA-21 in mice. J Clin Invest. 2010;120:3912–3916. doi: 10.1172/JCI43604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Duisters RF, Tijsen AJ, Schroen B, Leenders JJ, Lentink V, van der Made I, et al. miR-133 and miR-30 regulate connective tissue growth factor Implications for a role of microRNAs in myocardial matrix remodeling. Circ Res. 2009;104:170–178. doi: 10.1161/CIRCRESAHA.108.182535. [DOI] [PubMed] [Google Scholar]
- 86.van Rooij E, Sutherland LB, Thatcher JE, Dimaio JM, Naseem RH, Marshall WS, et al. Dysregulation of microRNAs after myocardial infarction reveals a role of miR-29 in cardiac fibrosis. Proc Natl Acad Sci USA. 2008;105:13027–13032. doi: 10.1073/pnas.0805038105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Karakikes I. Therapeutic cardiac-targeted delivery of miR-1 reverses pressure overload-induced cardiac hypertrophy and attenuates pathological remodeling. J Am Heart Assoc. 2013;2(2):e000078. doi: 10.1161/JAHA.113.000078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Zhao N, Yu HY, Yu HT, Sun M, Zhang YY, Xu M, et al. miRNA-711-SP1-collagen-I pathway is involved in the anti-fibrotic effect of pioglitazone in myocardial infarction. Sci China-Ser C. 2013;56:431–439. doi: 10.1007/s11427-013-4477-1. [DOI] [PubMed] [Google Scholar]
- 89.Cushing L, Kuang PP, Qian J, Shao FZ, Wu JJ, Little F, et al. miR-29 is a major regulator of genes associated with pulmonary fibrosis. Am J Resp Cell Mol Biol. 2011;45:287–294. doi: 10.1165/rcmb.2010-0323OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Kwiecinski M, Noetel A, Elfimova N, Trebicka J, Schievenbusch S, Strack I, et al. Hepatocyte growth factor (HGF) inhibits collagen I and IV synthesis in hepatic stellate cells by miRNA-29 induction. PLoS ONE. 2011;6:e24568. doi: 10.1371/journal.pone.0024568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Liang HH, Zhang C, Ban T, Liu Y, Mei L, Piao XM, et al. A novel reciprocal loop between microRNA-21 and TGF beta RIII is involved in cardiac fibrosis. Int J Biochem Cell B. 2012;44:2152–2160. doi: 10.1016/j.biocel.2012.08.019. [DOI] [PubMed] [Google Scholar]
- 92.Wang J, Huang WC, Xu RX, Nie Y, Cao XQ, Meng J, et al. MicroRNA-24 regulates cardiac fibrosis after myocardial infarction. J Cell Mol Med. 2012;16:2150–2160. doi: 10.1111/j.1582-4934.2012.01523.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Pan ZW, Sun XL, Shan HL, Wang N, Wang JH, Ren JS, et al. MicroRNA-101 inhibited postinfarct cardiac fibrosis and improved left ventricular compliance via the FBJ osteosarcoma oncogene/transforming growth factor-beta 1 pathway. Circulation. 2012;126:840–850. doi: 10.1161/CIRCULATIONAHA.112.094524. [DOI] [PubMed] [Google Scholar]
- 94.Yamakuchi M. MicroRNAs in vascular biology. Int J Vasc Med. 2013 doi: 10.1155/2012/794898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Wang SS, Aurora AB, Johnson BA, Qi XX, McAnally J, Hill JA, et al. The endothelial-specific microRNA miR-126 governs vascular integrity and angiogenesis. Dev Cell. 2008;15:261–271. doi: 10.1016/j.devcel.2008.07.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.van Mil A, Vrijsen KR, Goumans MJ, Metz CH, Doevendans PA, Sluijter JP. MicroRNA-1 enhances the angiogenic differentiation of human cardiomyocyte progenitor cells. J Mol Med. 2013;91(8):1001–1012. doi: 10.1007/s00109-013-1017-1. [DOI] [PubMed] [Google Scholar]
- 97.Chen JJ, Zhou SH. Mesenchymal stem cells overexpressing MiR-126 enhance ischemic angiogenesis via the AKT/ERK-related pathway. Cardiol J. 2011;18:675–681. doi: 10.5603/cj.2011.0032. [DOI] [PubMed] [Google Scholar]
- 98.Huang F, Zhu X, Hu XQ, Fang ZF, Tang L, Lu XL, et al. Mesenchymal stem cells modified with miR-126 release angiogenic factors and activate Notch ligand Delta-like-4, enhancing ischemic angiogenesis and cell survival. Int J Mol Med. 2013;31:484–492. doi: 10.3892/ijmm.2012.1200. [DOI] [PubMed] [Google Scholar]
- 99.Jakob P, Doerries C, Briand S, Mocharla P, Krankel N, Besler C, et al. Loss of AngiomiR-126 and 130a in angiogenic early outgrowth cells from patients with chronic heart failure Role for impaired in vivo neovascularization and cardiac repair capacity. Circulation. 2012;126:2962–2975. doi: 10.1161/CIRCULATIONAHA.112.093906. [DOI] [PubMed] [Google Scholar]
- 100.Bonauer A, Carmona G, Iwasaki M, Mione M, Koyanagi M, Fischer A, et al. MicroRNA-92a controls angiogenesis and functional recovery of ischemic tissues in mice. Science. 2009;324:1710–1713. doi: 10.1126/science.1174381. [DOI] [PubMed] [Google Scholar]
- 101.Fiedler J, Jazbutyte V, Kirchmaier BC, Gupta SK, Lorenzen J, Hartmann D, et al. MicroRNA-24 regulates vascularity after myocardial infarction. Circulation. 2011;124:720–730. doi: 10.1161/CIRCULATIONAHA.111.039008. [DOI] [PubMed] [Google Scholar]
- 102.van Mil A, Grundmann S, Goumans MJ, Lei ZY, Oerlemans MI, Jaksani S, et al. MicroRNA-214 inhibits angiogenesis by targeting Quaking and reducing angiogenic growth factor release. Cardiovasc Res. 2012;93:655–665. doi: 10.1093/cvr/cvs003. [DOI] [PubMed] [Google Scholar]
- 103.Ikeda S, Pu WT. Expression and function of microRNAs in heart disease. Curr Drug Targets. 2010;11:913–925. doi: 10.2174/138945010791591304. [DOI] [PubMed] [Google Scholar]
- 104.Sayed D, Hong C, Chen IY, Lypowy J, Abdellatif M. MicroRNAs play an essential role in the development of cardiac hypertrophy. Circ Res. 2007;100:416–424. doi: 10.1161/01.RES.0000257913.42552.23. [DOI] [PubMed] [Google Scholar]
- 105.Sucharov C, Bristow MR, Port JD. miRNA expression in the failing human heart: functional correlates. J Mol Cell Cardiol. 2008;45:185–192. doi: 10.1016/j.yjmcc.2008.04.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Thum T, Galuppo P, Wolf C, Fiedler J, Kneitz S, van Laake LW, et al. MicroRNAs in the human heart: a clue to fetal gene reprogramming in heart failure. Circulation. 2007;116:258–267. doi: 10.1161/CIRCULATIONAHA.107.687947. [DOI] [PubMed] [Google Scholar]
- 107.Reddy S, Zhao MM, Hu DQ, Fajardo G, Hu SJ, Ghosh Z, et al. Dynamic microRNA expression during the transition from right ventricular hypertrophy to failure. Physiol Genomics. 2012;44:562–575. doi: 10.1152/physiolgenomics.00163.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Jessup M, Greenberg B, Mancini D, Cappola T, Pauly DF, Jaski B, et al. Calcium upregulation by percutaneous administration of gene therapy in cardiac disease (CUPID) a phase 2 trial of intracoronary gene therapy of sarcoplasmic reticulum Ca2+-ATPase in patients with advanced heart failure. Circulation. 2011;124:304–313. doi: 10.1161/CIRCULATIONAHA.111.022889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Kumarswamy R, Lyon AR, Volkmann I, Mills AM, Bretthauer J, Pahuja A, et al. SERCA2a gene therapy restores microRNA-1 expression in heart failure via an Akt/FoxO3A-dependent pathway. Eur Heart J. 2012;33:1067–1075. doi: 10.1093/eurheartj/ehs043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Ikeda S, He AB, Kong SW, Lu J, Bejar R, Bodyak N, et al. MicroRNA-1 negatively regulates expression of the hypertrophy-associated calmodulin and Mef2a genes. Mol Cell Biol. 2009;29:2193–2204. doi: 10.1128/MCB.01222-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Care A, Catalucci D, Felicetti F, Bonci D, Addario A, Gallo P, et al. MicroRNA-133 controls cardiac hypertrophy. Nat Med. 2007;13:613–618. doi: 10.1038/nm1582. [DOI] [PubMed] [Google Scholar]
- 112.Aurora AB, Mahmoud AI, Luo X, Johnson BA, van Rooij E, Matsuzaki S, et al. MicroRNA-214 protects the mouse heart from ischemic injury by controlling Ca2+ overload and cell death. J Clin Invest. 2012;122:1222–1232. doi: 10.1172/JCI59327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Cha MJ. MicroRNA-145 suppresses ROS-induced Ca(2+) overload of cardiomyocytes by targeting CaMKII? Biochem Biophys Res Commun. 2013;435:720–726. doi: 10.1016/j.bbrc.2013.05.050. [DOI] [PubMed] [Google Scholar]
- 114.Li RT, Yan GJ, Li QL, Sun HX, Hu YL, Sun JX, et al. MicroRNA-145 protects cardiomyocytes against hydrogen peroxide (H2O2)-induced apoptosis through targeting the mitochondria apoptotic pathway. PLoS ONE. 2012;7:e44907. doi: 10.1371/journal.pone.0044907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.van Mil A, Grundmann S, Goumans MJ, Lei ZY, Oerlemans MI, Jaksani S, et al. MicroRNA-214 inhibits angiogenesis by targeting Quaking and reducing angiogenic growth factor release. Cardiovasc Res. 2012;93:655–665. doi: 10.1093/cvr/cvs003. [DOI] [PubMed] [Google Scholar]
- 116.Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA. 2008;105:10513–10518. doi: 10.1073/pnas.0804549105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Tijsen AJ, Creemers EE, Moerland PD, De Windt LJ, van der Wal AC, Kok WE, et al. MiR423-5p as a circulating biomarker for heart failure. Circ Res. 2010;106:1035–1039. doi: 10.1161/CIRCRESAHA.110.218297. [DOI] [PubMed] [Google Scholar]
- 118.Kumarswamy R, Thum T, Anker SD. MicroRNAs as circulating biomarkers for heart failure: questions about MiR-423-5p. Circ Res. 2010;106:E8. doi: 10.1161/CIRCRESAHA.110.220616. [DOI] [PubMed] [Google Scholar]
- 119.Wang GK, Zhu JQ, Zhang JT, Li Q, Li Y, He J, et al. Circulating microRNA: a novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur Heart J. 2010;31:659–666. doi: 10.1093/eurheartj/ehq013. [DOI] [PubMed] [Google Scholar]
- 120.Ai J, Zhang R, Li Y, Pu JL, Lu YJ, Jiao JD, et al. Circulating microRNA-1 as a potential novel biomarker for acute myocardial infarction. Biochem Biophys Res Commun. 2010;391:73–77. doi: 10.1016/j.bbrc.2009.11.005. [DOI] [PubMed] [Google Scholar]
- 121.Corsten MF, Dennert R, Jochems S, Kuznetsova T, Devaux Y, Hofstra L, et al. Circulating microRNA-208b and microRNA-499 reflect myocardial damage in cardiovascular disease. Circ Cardiovasc Gene. 2010;3:499–506. doi: 10.1161/CIRCGENETICS.110.957415. [DOI] [PubMed] [Google Scholar]
- 122.Kuwabara Y, Ono K, Horie T, Nishi H, Nagao K, Kinoshita M, et al. Increased MicroRNA-1 and MicroRNA-133a levels in serum of patients with cardiovascular disease indicate myocardial damage. Circ Cardiovasc Gene. 2011;4:446–454. doi: 10.1161/CIRCGENETICS.110.958975. [DOI] [PubMed] [Google Scholar]
- 123.Long GW, Wang F, Duan QL, Yang SL, Chen FQ, Gong W, et al. Circulating miR-30a, miR-195 and let-7b associated with acute myocardial infarction. PLoS ONE. 2012;7:e24568. doi: 10.1371/journal.pone.0050926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Oerlemans MIFJ, Mosterd A, Dekker MS, de Vrey EA, van Mil A, Pasterkamp G, et al. Early assessment of acute coronary syndromes in the emergency department: the potential diagnostic value of circulating microRNAs. EMBO Mol Med. 2012;4:1176–1185. doi: 10.1002/emmm.201201749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Abdellatif M. Differential expression of microRNAs in different disease states. Circ Res. 2012;110:638–650. doi: 10.1161/CIRCRESAHA.111.247437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Cai BZ, Pan ZW, Lu YJ. The roles of microRNAs in heart diseases: a novel important regulator. Curr Med Chem. 2010;17:407–411. doi: 10.2174/092986710790226129. [DOI] [PubMed] [Google Scholar]
- 127.Fasanaro P, Greco S, Ivan M, Capogrossi MC, Martelli F. MicroRNA: emerging therapeutic targets in acute ischemic diseases. Pharmacol Therap. 2010;125:92–104. doi: 10.1016/j.pharmthera.2009.10.003. [DOI] [PubMed] [Google Scholar]
- 128.Frost RJA, van Rooij E. miRNAs as therapeutic targets in ischemic heart disease. J Cardiovasc Transl Res. 2010;3:280–289. doi: 10.1007/s12265-010-9173-y. [DOI] [PubMed] [Google Scholar]
- 129.Heyn J, Hinske C, Mohnle P, Luchting B, Beiras-Fernandez A, Kreth S. MicroRNAs as potential therapeutic agents in the treatment of myocardial infarction. Curr Vasc Pharmacol. 2011;9:733–740. doi: 10.2174/157016111797484143. [DOI] [PubMed] [Google Scholar]
- 130.Montgomery RL, van Rooij E. MicroRNA regulation as a therapeutic strategy for cardiovascular disease. Curr Drug Targets. 2010;11:936–942. doi: 10.2174/138945010791591368. [DOI] [PubMed] [Google Scholar]
- 131.van Rooij E, Purcell AL, Levin AA. Developing microRNA therapeutics. Circ Res. 2012;110:496–507. doi: 10.1161/CIRCRESAHA.111.247916. [DOI] [PubMed] [Google Scholar]
- 132.Bish LT, Morine K, Sleeper MM, Sanmiguel J, Wu D, Gao GP, et al. Adeno-associated virus (aav) serotype 9 provides global cardiac gene transfer superior to AAV1, AAV6, AAV7, and AAV8 in the mouse and rat. Hum Gene Ther. 2008;19:1359–1368. doi: 10.1089/hum.2008.123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Rane S, He MZ, Sayed D, Vashistha H, Malhotra A, Sadoshima J, et al. Downregulation of MiR-199a derepresses hypoxia-inducible factor-1 alpha and sirtuin 1 and recapitulates hypoxia preconditioning in cardiac myocytes. Circ Res. 2009;104:879–886. doi: 10.1161/CIRCRESAHA.108.193102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Chan YC, Roy S, Huang Y, Khanna S, Sen CK. The microRNA miR-199a-5p down-regulation switches on wound angiogenesis by derepressing the v-ets erythroblastosis virus E26 oncogene homolog 1-matrix metalloproteinase-1 pathway. J Biol Chem. 2012;287:41032–41043. doi: 10.1074/jbc.M112.413294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Ahuja P, Sdek P, MaClellan WR. Cardiac myocyte cell cycle control in development, disease, and regeneration. Physiol Rev. 2007;87:521–544. doi: 10.1152/physrev.00032.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Collesi C, Zentilin L, Sinagra G, Giacca M. Notch1 signaling stimulates proliferation of immature cardiomyocytes. J Cell Biol. 2008;183:117–128. doi: 10.1083/jcb.200806091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Li FQ, Wang XJ, Capasso JM, Gerdes AM. Rapid transition of cardiac myocytes from hyperplasia to hypertrophy during postnatal development. J Mol Cell Cardiol. 1996;28:1737–1746. doi: 10.1006/jmcc.1996.0163. [DOI] [PubMed] [Google Scholar]
- 138.Soonpaa MH, Kim KK, Pajak L, Franklin M, Field LJ. Cardiomyocyte DNA synthesis and binucleation during murine development. Am J Physiol-Heart C. 1996;271:H2183–H2189. doi: 10.1152/ajpheart.1996.271.5.H2183. [DOI] [PubMed] [Google Scholar]
- 139.Bicknell KA, Coxon CNH, Brooks G. Can the cardiomyocyte cell cycle be reprogrammed? J Mol Cell Cardiol. 2007;42:706–721. doi: 10.1016/j.yjmcc.2007.01.006. [DOI] [PubMed] [Google Scholar]
- 140.Senyo SE, Steinhauser ML, Pizzimenti CL, Yang VK, Cai L, Wang M, et al. Mammalian heart renewal by pre-existing cardiomyocytes. Nature. 2013;493(7432):433–436. doi: 10.1038/nature11682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Beltrami AP, Urbanek K, Kajstura J, Yan SM, Finato N, Bussani R, et al. Evidence that human cardiac myocytes divide after myocardial infarction. New Engl J Med. 2001;344:1750–1757. doi: 10.1056/NEJM200106073442303. [DOI] [PubMed] [Google Scholar]
- 142.Kajstura J, Zhang X, Reiss K, Szoke E, Li P, Lagrasta C, et al. Myocyte cellular hyperplasia and myocyte cellular hypertrophy contribute to chronic ventricular remodeling in coronary-artery narrowing-induced cardiomyopathy in rats. Circ Res. 1994;74:383–400. doi: 10.1161/01.res.74.3.383. [DOI] [PubMed] [Google Scholar]
- 143.Reiss K, Kajstura J, Capasso JM, Marino TA, Anversa P. Impairment of myocyte contractility following coronary-artery narrowing is associated with activation of the myocyte Igf(1) autocrine system, enhanced expression of late growth-related genes, dna-synthesis, and myocyte nuclear mitotic division in rats. Exp Cell Res. 1993;207:348–360. doi: 10.1006/excr.1993.1202. [DOI] [PubMed] [Google Scholar]
- 144.Porrello ER, Johnson BA, Aurora AB, Simpson E, Nam YJ, Matkovich SJ, et al. miR-15 family regulates postnatal mitotic arrest of cardiomyocytes. Circ Res. 2011;109:670–679. doi: 10.1161/CIRCRESAHA.111.248880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Porrello ER, Mahmoud AI, Simpson E, Johnson BA, Grinsfelder D, Canseco D, et al. Regulation of neonatal and adult mammalian heart regeneration by the miR-15 family. Proc Natl Acad Sci USA. 2013;110:187–192. doi: 10.1073/pnas.1208863110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Zhao Y, Ransom JF, Li A, Vedantham V, von Drehle M, Muth AN, et al. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell. 2007;129:303–317. doi: 10.1016/j.cell.2007.03.030. [DOI] [PubMed] [Google Scholar]
- 147.Yin VP, Lepilina A, Smith A, Poss KD. Regulation of zebrafish heart regeneration by miR-133. Dev Biol. 2012;365:319–327. doi: 10.1016/j.ydbio.2012.02.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Jopling C, Sleep E, Raya M, Marti M, Raya A, Belmonte JCI. Zebrafish heart regeneration occurs by cardiomyocyte dedifferentiation and proliferation. Nature. 2010;464(7288):606–609. doi: 10.1038/nature08899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Kikuchi K, Holdway JE, Werdich AA, Anderson RM, Fang Y, Egnaczyk GF, et al. Primary contribution to zebrafish heart regeneration by gata4(+) cardiomyocytes. Nature. 2010;464:601–605. doi: 10.1038/nature08804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Poss KD, Nechiporuk A, Hillam AM, Johnson SL, Keating MT. Mps1 defines a proximal blastemal proliferative compartment essential for zebrafish fin regeneration. Development. 2002;129:5141–5149. doi: 10.1242/dev.129.22.5141. [DOI] [PubMed] [Google Scholar]
- 151.Chen J, Huang ZP, Seok HY, Ding J, Kataoka M, Zhang Z, et al. Mir-17-92 cluster is required for and sufficient to induce cardiomyocyte proliferation in postnatal and adult hearts. Circ Res. 2013;112:1557–1566. doi: 10.1161/CIRCRESAHA.112.300658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Eulalio A, Mano M, Dal Ferro M, Zentilin L, Sinagra G, Zacchigna S, et al. Functional screening identifies miRNAs inducing cardiac regeneration. Nature. 2012;492:376–381. doi: 10.1038/nature11739. [DOI] [PubMed] [Google Scholar]
- 153.Porrello ER. microRNAs in cardiac development and regeneration. Clin Sci (Lond) 2013;125:151–166. doi: 10.1042/CS20130011. [DOI] [PubMed] [Google Scholar]
- 154.Srivastava D. Small solutions to big problems: micrornas for cardiac regeneration. Circ Res. 2013;112:1412–1414. doi: 10.1161/CIRCRESAHA.113.301409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.van Amerongen MJ, Engel FB. Features of cardiomyocyte proliferation and its potential for cardiac regeneration. J Cell Mol Med. 2008;12:2233–2244. doi: 10.1111/j.1582-4934.2008.00439.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Anversa P, Nadal-Ginard B. Myocyte renewal and ventricular remodelling. Nature. 2002;415:240–243. doi: 10.1038/415240a. [DOI] [PubMed] [Google Scholar]
- 157.van Vliet P, Roccio M, Smits AM, van Oorschot AAM, Metz CHG, van Veen TAB, et al. Progenitor cells isolated from the human heart: a potential cell source for regenerative therapy. Neth Heart J. 2008;16:163–169. doi: 10.1007/BF03086138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Segers VFM, Lee RT. Stem-cell therapy for cardiac disease. Nature. 2008;451:937–942. doi: 10.1038/nature06800. [DOI] [PubMed] [Google Scholar]
- 159.Kuppusamy KT. MicroRNA regulation and role in stem cell maintenance, cardiac differentiation and hypertrophy. Curr Mol Med. 2013;13:757–764. doi: 10.2174/1566524011313050007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Scott E (2013) miRNA regulation of endothelial homeostasis and commitment: implications for vascular regeneration strategies using stem cell therapies. Free Radic Biol Med (in press) [DOI] [PubMed]
- 161.Brás-Rosário L, Matsuda A, Pinheiro AI, Gardner R, Lopes T, Amaral A, Gama-Carvalho M. Expression profile of microRNAs regulating proliferation and differentiation in mouse adult cardiac stem cells. PLoS ONE. 2013;8(5):e63041. doi: 10.1371/journal.pone.0063041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Sirish P, Lopez JE, Li N, Wong A, Timofeyev V, Young JN, et al. MicroRNA profiling predicts a variance in the proliferative potential of cardiac progenitor cells derived from neonatal and adult murine hearts. J Mol Cell Cardiol. 2012;52:264–272. doi: 10.1016/j.yjmcc.2011.10.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Xiao JJ, Liang DD, Zhang H, Liu Y, Zhang DS, Liu Y, et al. MicroRNA-204 is required for differentiation of human-derived cardiomyocyte progenitor cells. J Mol Cell Cardiol. 2012;53:751–759. doi: 10.1016/j.yjmcc.2012.08.024. [DOI] [PubMed] [Google Scholar]
- 164.Heinrich EM, Dimmeler S. MicroRNAs and stem cells control of pluripotency, reprogramming, and lineage commitment. Circ Res. 2012;110:1014–1022. doi: 10.1161/CIRCRESAHA.111.243394. [DOI] [PubMed] [Google Scholar]