Abstract
Transglutaminase 2 (TG2) is a ubiquitously expressed member of an enzyme family catalyzing Ca2+-dependent transamidation of proteins. It is a multifunctional protein having several well-defined enzymatic (GTP binding and hydrolysis, protein disulfide isomerase, and protein kinase activities) and non-enzymatic (multiple interactions in protein scaffolds) functions. Unlike its enzymatic interactions, the significance of TG2’s non-enzymatic regulation of its activities has recently gained importance. In this review, we summarize all the partners that directly interact with TG2 in a non-enzymatic manner and analyze how these interactions could modulate the crosslinking activity and cellular functions of TG2 in different cell compartments. We have found that TG2 mostly acts as a scaffold to bridge various proteins, leading to different functional outcomes. We have also studied how specific structural features, such as intrinsically disordered regions and embedded short linear motifs contribute to multifunctionality of TG2. Conformational diversity of intrinsically disordered regions enables them to interact with multiple partners, which can result in different biological outcomes. Indeed, ID regions in TG2 were identified in functionally relevant locations, indicating that they could facilitate conformational transitions towards the catalytically competent form. We reason that these structural features contribute to modulating the physiological and pathological functions of TG2 and could provide a new direction for detecting unique regulatory partners. Additionally, we have assembled all known anti-TG2 antibodies and have discussed their significance as a toolbox for identifying and confirming novel TG2 regulatory functions.
Keywords: Transglutaminase 2, Non-enzymatic interactions, Intrinsically disordered regions, TRANSDAB database, Scaffolding, Anti-TG2 antibodies
Introduction
Transglutaminases (TGs, EC 2.3.2.13) are a family of nine enzymes (TG1–7, F13, and Band 4.3) that covalently crosslink a wide array of substrate proteins with available glutamine and lysine residues via a transamidation reaction except Band 4.3, which is inactive. This crosslinking reaction leads to the formation of protease-resistant isopeptide bonds between the substrate proteins [1–3]. Crosslinking of proteins helps in a multitude of functions such as extracellular matrix (ECM) stabilization, cornified cell envelope formation during keratinocyte differentiation, blood clotting, wound healing, bone growth, scaffolding, apoptosis, cell adhesion and cell-survival signaling functions [1]. Besides crosslinking, TGs mediate incorporation of primary amines into proteins, deamidation of glutamine residues in the absence of substrates, and isopeptidase activities. TG2 is the most ubiquitous TG family member expressed in almost all cell compartments such as the cytoplasm, mitochondria, recycling endosomes, and nucleus. It is also present on the cell surface and gets secreted to the ECM via non-classical mechanisms [4].
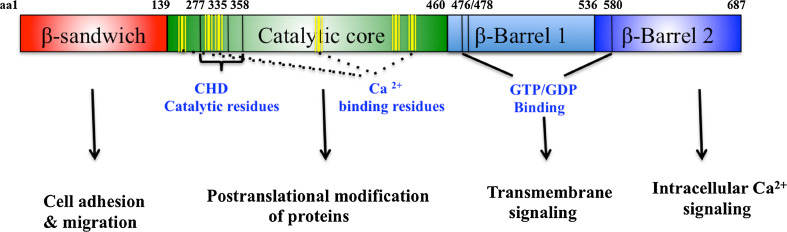
TG2 has a conserved 3D structure and catalytic triad shared by other family members but its multifunctionality is unique. The crystal structure of TG2 reveals four important domains: (a) N-terminal β sandwich domain, (b) catalytic core domain, and (c) two C-terminal β1 and β2 barrel domains (Fig. 1). Each of these domains has several essential functional characteristics, which are unique to TG2, as illustrated in Fig. 1. The N-terminal domain is involved in cell adhesion and migration function. The core domain has six calcium binding sites, which regulates the transamidation activity of TG2. The β1 barrel domain of TG2 binds and hydrolyzes GTP/ATP [5–7]. The β1, β2, and core domain together regulate the signal transducing function acting as a G-protein. These domain-specific functions are mediated by binding to different types of interacting proteins. TG2 also has protein disulfide isomerase [8, 9] and protein kinase [10, 11] activities besides its transamidation activities. In addition to these biochemical functions, TG2 has been implicated in several physiological and pathological processes as will be discussed herein. We hypothesize that non-enzymatic protein–protein interactions contribute to the unprecedented multifunctionality of TG2.
Fig. 1.
Scheme of TG2 with its domains along with its proposed functions. TG2 domains are color coded. Domain boundaries have been indicated by amino acid numbers. Calcium binding residues, catalytic residues, and GTP/GDP binding residues have also been indicated. Amino acid numbers have been provided for the catalytic residues and GTP/GDP binding residues. This figure is adapted from Mehta et al. [181]
TG2 participates in both enzymatic and non-enzymatic interactions. Enzymatic interactions are formed between TG2 and its substrate proteins containing the glutamine donor and lysine donor groups in the presence of calcium. Substrates of TG2 are known to affect TG2 activity, which enables it to subsequently execute diverse biological functions in the cell. Most of the substrates of TG2, as well as those of the other transglutaminase family members, can be found in the TRANSDAB database [12]. However, the importance of non-enzymatic interactions in regulating TG2 activities is yet to be revealed. Non-enzymatic interactions of TG2 do not involve a transamidation reaction and most probably occur at sites other than its catalytic site.
Recent studies indicate that non-enzymatic interactions play physiological roles and enable diverse TG2 functions in a context-specific manner [13, 14]. For example, interaction of TG2 with retinoblastoma (Rb) protein is pro-apoptotic when the TG2/Rb complex is present in the cytoplasm, but it is anti-apoptotic when the complex is present in the nucleus [15]. The TRANSDAB database also includes some non-enzymatic interaction partners but it is not an exhaustive list. It appears that protein–protein interaction databases, such as STRING or the human protein interaction database also lack most of the non-enzymatic partners of TG2. In some cases, the proteins that are bound non-covalently also serve as enzymatic substrates for transamidation/crosslinking (for example, fibronectin, Bcr, Rac1, and angiocidin). Thus, in addition to its enzymatic functions, its wide variety of non-covalent interactions implicates TG2 in a plethora of adapter and signaling functions both inside and outside cells, enabling it to impinge on a number of signaling pathways [4].
In this review, we have aimed to identify further interaction partners (not substrates) of TG2 from database analysis and a review of the literature. We collected almost double the number of interacting partners by manual data collection through a literature search compared with the partners of TG2 identified by online database analysis. This review focuses on the outcomes from the literature survey on how non-enzymatic interactions modulate and expand the functionality of TG2 in different cell compartments and how they can potentially interfere with the crosslinking activity of TG2.
Non-enzymatic interactions of TG2 and their physiological and pathological outcomes
In this section, we have grouped together all the biological functions in which TG2 is reported to be involved. Additionally, how non-covalent interactions regulate TG2-mediated cellular and extracellular functions are discussed.
Cell adhesion, migration, and stabilization
Secretion and oxidation of TG2
Although TG2 is predominantly present in the cytosol, it is secreted outside cells where it is involved in various extracellular activities such as cell attachment, motility, stabilization, and survival [16, 4]. However, it is not fully understood how TG2 is secreted because of the absence of any secretory leader sequence, hydrophobic, or transmembrane domains. Several non-conventional mechanisms of TG2 secretion have been reported [17–19]. A recent review describes plausible scenarios of TG2 externalization [4]. Amongst these mechanisms, TG2 was proposed to use the long loop of the endosomal recycling pathway for its secretion. This process most likely occurs in two steps whereby TG2 initially tethers to the endosomal phosphoinositides and then subsequently binds tightly to the endosomal membrane through unidentified membrane proteins. Future identification of these unknown surface proteins should help to further elucidate the TG2 secretion mechanism [20].
TG2 promotes cell adhesion, migration, and stabilization of the ECM by crosslinking several ECM proteins as well as interacting with them in a calcium-independent manner [21–23]. Though numerous substrates and interacting proteins have been identified in the ECM, surprisingly, extracellular TG2 was found to be inactive in vivo, but could be activated via mechanical injury or inflammatory stimuli.
The activity of TG2 in an extracellular environment also depends on its oxidation state. Reversible oxidation of the key cysteine triad (Cys230, Cys370, and Cys371) results in the formation of disulfide bonds between Cys230 and Cys370 and Cys370 and Cys371, which inactivates TG2. Oxidized TG2 adopts an open conformation [24]. Interestingly, the active form of TG2 also adopts an open conformation, but the structural differences between these two forms are not yet known. Calcium and substrate could regulate the oxidation state of TG2, suggesting that the local environment can modulate and fine-tune oxidative inactivation of TG2 in the extracellular space [24].
TG2 and fibronectin interaction
Fibronectin (FN) is one of the best-characterized ECM proteins. It forms a complex with TG2 [25] and is a substrate for TG2 activity. However, TG2-mediated formation of FN matrix fibrils does not require crosslinking activity. Instead, it depends on a direct non-covalent interaction with fibronectin [25, 26]. TG2 binds to the 42-kD gelatin binding domain of FN [21, 27], whereas the specific recognition sequence for FN binding is located within the N-terminal beta sandwich domain of TG2 (Figs. 1, 2; Table 1). Aspartate at position 94 and 97 within the TG2 recognition sequence, 88WTATDDVQQDCTLSLQLTT106, was found to be critical for the FN-TG2 interaction [28]. Also, TG2 was found to bind FN with very high affinity (Kd ~8–10 nM) and in a stoichiometric ratio of 2:1 [25, 26].
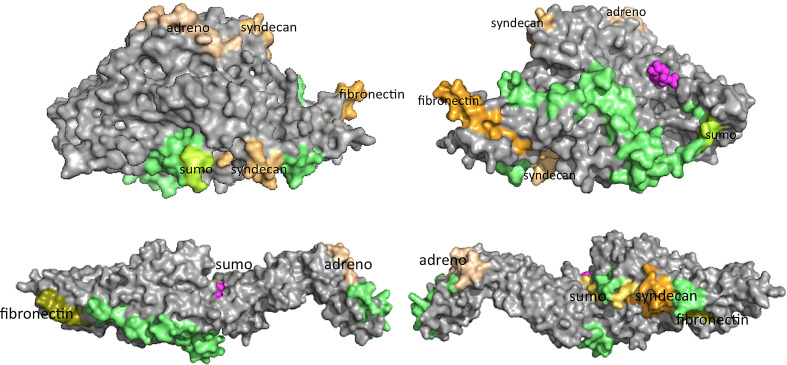
Fig. 2.
Interaction sites of TG2 binding proteins and ID regions in the TG2 crystal structure. The left panel and right panel show the front and back views of the TG2 crystal structure in the closed (upper half, 1kv3) and open form (lower half, 2q3z). Intrinsically disordered regions are shown in lime green. Interacting sites overlapping with disordered protein regions with linear interaction motifs are displayed: fibronectin (olive-green), syndecan (orange), SUMO 1 (dark yellow, light green), α1-adrenoceptor (beige)
Table 1.
Detailed list of TG2-interacting partners along with their proposed functions
| List of TG2 interacting proteins | |||||||
|---|---|---|---|---|---|---|---|
| Name of the protein | Site of interaction | Type of cell line | Technique used | Interacting domains and sequences | Proposed functions | STRING database score | References |
| Fibronectin P02751* | ECM, plasma | NIH3T3, WI38 fibroblast, HEL, REF52 | Direct cell adhesion assay, GST pull-down, SPR | N-terminal domain of TG2 (88WTATVVDQQD CTLSLQLTT106), critical residue in bold and 42-kDa gelatin binding domain of fibronectin | Cell adhesion and migration | 0.99 | Akimov et al. [23], Hang et al. [28] |
| Integrins β1 and β3 P05556/P05106* | Membrane | Swiss 3T3, WI38 fibroblast, REF52, HEL | Co-IP, IF | First and fourth domains of TG2 | Cell adhesion and migration | 0.96 | Akimov et al. [23] |
| Syndecan-4 AAH30805 | Membrane, ECM | HOB, human fibroblast cells, HEK | Co-IP | 202KFLKNAGRDCSRRS SPVYVGR222 on TG2 | Cell adhesion and migration | 0.833 | Wang et al. [43, 169] |
| PDGFR AAA60049 | Membrane | NHDFs, NIH3T3 | Co-IP, IF, in vitro binding assays | Not known | Cell adhesion and migration | Not present | Zemskov et al. [50] |
| LRP 5 NP002326, LRP 6 AAI43726 | Transmembrane (smooth muscle cells) | Vascular smooth muscle cells (VSMC) | GST pull-down, Co-IP, yeast two-hybrid screening (Y2H) | Not known | Nuclear translocation of β catenin, which leads to activation of Wnt/β catenin signaling pathways | Not present | Faverman et al. [51] |
| LRP1 Q07954 | Membrane | Fibroblast cells | Co-IP | Catalytic domain of TG2 | Cell adhesion, migration, receptor-mediated signaling | Not present | Zemskov et al. [14] |
| ACAP 13 Q12802 | Perinuclear-cytoplasmic areas | Prostate cancer cells | Co-IP, IF, Y2H | Catalytic domain of TG2 | Involved in Rho-mediated change in cell polarity, potentiate interaction between PKA and TG2 | 0.621 | Lewis et al. [61] |
| PTEN (tumor suppressor phosphatase) AAD13528 | Cytoplasm | PDAC, Pan28 cells | Co-IP, IF | Not known | Regulation of cell-survival signaling | Not present | Verma et al. [80] |
| HSP 70 P08107 | Membrane | HELA, MDAMB231 | Co-IP | Not known | Cell migration by targeting TG2 to cell surface | 0.207 | Boroughs et al. [81] |
| Endostatin NP085059 | ECM | HEK293-EBNA cells | SPR binding assays, double immunostaining | GTP binding site of TG2 and R 27 and/or R139 of endostatin | Regulation of angiogenesis and tumor growth | Not present | Faye et al. [86] |
| BCR CAA26441 and ABR Q12979 | Cytoplasm | COS-1, Swiss 3T3 | Co-IP, IF, GST pull-down | Core domain of Tg2 and GAP domain of BCR and ABR | Cytoskeleton rearrangement through Rac activation | Not present | Yi et al. [82] |
| GPR56 AAQ88766 | ECM | MC1, 293T | Affinity chromatography, far Western | C terminal β barrel of TG2 and N terminal domain of GPR56 | Tumor progression | 0.326 | Xu et al. [85] |
| MMP2 P08253 | ECM | MCF7, fibrosarcoma HT 1080, glioma U-MT cells | Co-IP, SPR, ELISA | Core domain of TG2 and catalytic domain of MMP2 | Regulation of matrix composition and migration/invasion of malignant cells | 0.24 | Belkin et al. [59] |
| Angiocidin NP002801 | Cytoplasm | HUVE cells, MD-MBA-231 | Co-IP, IF, ELISA, IHC | Not known | Regulates cell migration and adhesion by inhibiting FN incorporation to ECM | Not present | L’Heureux et al. [88] |
| α1 adrenoceptor P25100 | Membrane | COS-1 | Co-IP | Multiple sites, L547-I561, R564-D581, Q633-E646 on TG2 and third TM loop of α1 adrenoceptor | Regulates intracellular Ca2+ signaling | 0.297 | Feng et al. [89], Chen et al. [90] |
| PLC δ1 P51178* | Cytoplasm | COS-1, DDT1-MF2 | Co-IP, synthetic peptide approach | V665-K672 onTG2 and C2 domain 721TIPWNSLKQGYRH VHLL736 of PLCδ1 | Regulates intracellular Ca2+ signaling | 0.88 | Hwang et al. [94], Kang et al. [95] |
| Oxytocin receptor NP000907 | Membrane | Human myometrium | Co-IP | Not known | Intracellular calcium mobilization and muscle contraction | Not present | Baek et al. [93] |
| Thromboxane receptor BAA07274 | Membrane | COS-7 | Co-IP | Not known | Transmembrane signaling | 0.872 | Vezza et al. [6] |
| Calreticulin AAB51176 | Cytoplasm | Rat liver membrane | Co-IP, protein sequencing | Not known | Down-regulate both TGase and GTPase activities of TG2 | 0.235 | Hwang et al. [94] |
| Importin α3 AAC25605* | Cytoplasm, nucleus | NCL-H596 | Co-IP, IHC, Y2H | Not known | Active transport of TG2 into the nucleus | 0.625 | Peng et al. [109] |
| Eukaryotic initiation factor (eIF5A) AAH80196* | Nucleus, cytoplasm | HELA | Co-IP | Not known | TGases influence the cellular localization of eIF-5A | 0.623 | Singh et al. [110] |
| c-Jun P05412 | Nucleus | H9c2 | Co-IP | Not known | ECM regulation and turnover | Not present | Ahn et al. [112] |
| cSrc AAC50104 | Membrane | SKOV3, IGROV1 | Co-IP, IF | Not known | Cell–matrix interactions and Wnt signaling | Not present | Condello et al. [113] |
| Hypoxia-inducible factor (HIF1β) P27540 | Cytoplasm, nucleus | CHO, SH-SY5Y | GST pull-down, Co-IP, Y2H | Not known | Protective role in the brain during its response to ischemia and stroke | Not present | Filiano et al. [106] |
| Reitnoblastoma P06400* | Nucleus, cytoplasm | HEK | Co-IP | Not known | Transcriptional regulation | 0.66 | Milakovic et al. [15] |
| SUMO 1 (small ubiquitin-like modifier) NP001005781 | Nasal epithelium | IB3-1, 16HBE | Co-IP, IFe, FRET | 3 SUMO motifs detected on TG2 sequence (327–329, 364–366, 468–470) | Regulating oxidative stress and inflammation | 0.619 | Luciani et al. [108] |
| PIASy (SUMO 1 ligase) Q8N2W9 | Nasal epithelium | IB3-1, 16HBE | Co-IP, IF | Not known | Regulating oxidative stress and inflammation | Not present | Luciani et al. [108] |
| RAC1 CAB53579 | Cytoplasm, membrane | RBL2H3 | Co-IP | Not known | Regulates allergic inflammation | Not present | Kim et al. [182] |
| Tubulin β1 NP110400* | Nucleus | SK-N-BE, TGA | Co-IP | Not known | Regulates intracellular Ca2+ signaling | 0.619 | Piredda et al. [129] |
| Calmodulin AAD45181 | Nucleus | HD brain tissue | IF, Co-IP | Not known | Regulation of TG2-mediated crosslinking of huntingtin and formation of stable aggregates in HD brain | Not present | Zainelli et al. [133] |
| Histone H3, CAA58540 H2B CAB02542* | Nucleus | SK-N-BE | Co-IP, protein sequencing | Not known | Chromatin remodeling | 0.928 | McConoughey et al. [127], Piredda et al. [129] |
| BAX and BAK Q07812/AAA74466 | Cytoplasm | SK-N-BE(2), TGA | IF, Co-IP | 204–212 of TG2 | Stabilize efficient regulation of apoptosis | 0.226 | Rodolfo et al. [141] |
| 14-3-3 binding protein NP006817 | Cytoplasm | MEF | Pull-down assays | 209–223 of TG2 | Regulation of apoptosis | Not present | Mishra et al. [144] |
| MFG E8 P21956 | Cell surface | COS-1, MF2 | SPR | Not known | Phagocytosis of apoptotic cells | 0.85 | Toth et al. [38] |
| P62/SQSTM 1 AAA59990 | Cytoplasm | 2fTGH, HEK293 | Co-IP, IF | Not known | Recognition and recruitment of ubiquitinated proteins and organelles to pre-autophagic vesicles just before its degradation by autophagolysosomes | Not present | D’Eletto et al. [150] |
| NBR1 AAH09808 | Cytoplasm | 2fTGH, HEK293 | Co-IP, immuno-fluorescence | Not known | Recognition and recruitment of ubiquitinated proteins and organelles to pre-autophagic vesicle just before its degradation by autophagolysosomes | Not present | D’Eletto et al. [150] |
| HSP20 O14558 | Whole cell lysate | SH-SY5Y | Co-IP | Not known | Regulation of apoptosis by interacting with Hsp20/Hsp27 complex | Not present | Caccamo et al. [145] |
| Paxillin AAC50104 | Whole cell lysate | Corneal epithelial cells | Co-IP | Not known | Regulation of cell adhesion and migration by paxillin incorporation in focal adhesion complexes | Not present | Png and Tong (unpublished, GORDON research conference abstract) |
Interacting partners of TG2 collected from a literature search have been assembled in this table. The type of cell line and cellular locations, sequence of interaction, methodology used, and their proposed functions have been provided for each interacting partner. The proteins marked with an asterisk were also present in the integrated human protein interaction database [151]. The protein accession number is provided below the protein’s name
The TG2 and FN interaction is suggested to stabilize the ECM by enhancing FN matrix formation and crosslinking of other extracellular matrix proteins [16, 29–33]. Binding of cell-surface TG2 to the 42-kD gelatin binding domain of FN has been shown to promote stable cell adhesion, migration, and formation of specialized adhesive structures (focal adhesions) at the cell substratum interface in different cell types (e.g., human and rat fibroblast cells, human erythroleukemia, and endothelial cells). An affinity purified rabbit polyclonal antibody against the NH2-fragment of TG2 [23] as well as the monoclonal 4G3 antibody against the TG2 fibronectin binding domain were shown to interfere with the TG2 and FN interaction. These observations suggest that cell-surface TG2 serves as a principal adhesion receptor for this portion of the FN molecule [23, 34].
Association of TG2 with integrin
Integrins are a superfamily of cell-adhesion receptors that bind to both ECM and cell-surface ligands and connect them to the cytoskeleton. The sequence arginine–glycine–aspartic (RGD) is a general integrin binding motif that is present on several cell-adhesion proteins including FN [35]. Integrins recognize this RGD motif within FN and regulate cell adhesion and migration by facilitating cell–matrix interactions [36]. Because TG2 also shares similar functions [37], its probable mechanisms were explored. TG2 expressed on the surface of human erythroleukemia cells as well as rat fibroblast cells was reported to interact with the β1, β3, and β5 subunits of integrin through direct non-covalent interactions, forming stable ternary complexes with integrin and FN, thus facilitating enhanced cell adhesion and migration [23, 38].
The exact mapping of the integrin binding site on TG2 has not yet been reported but it has been proposed that it involves both the first and fourth domains of TG2 [4] (Table 1). When TG2 was simultaneously bound to integrin, it could also interact with the 42-kD gelatin binding domain [39] of FN. The gelatin binding domain of FN lacks integrin binding sites and thus TG2 serves as a bridge between integrin and FN, providing additional binding sites for the FN–integrin interaction [40]. The adhesive functions of TG2 strictly depend upon its association with integrin. Integrin is a low-affinity receptor for ECM proteins including FN [41], whereas TG2 is a high affinity binding partner of FN [40]. Therefore, formation of the TG2-FN-integrin ternary complex has a significant impact on cell-adhesion and migration signaling events. For example, an increase in the tyrosine phosphorylation of FAK (focal adhesion kinase) was observed in TG2-overexpressing cells [23].
Interaction of TG2 with syndecan-4
Another important binding partner for TG2 is the heparin sulfate proteoglycan (HSPG), syndecan-4. TG2 has very high affinity for a heparin sulfate (HS) analogue; heparin as TG2 from erythrocytes could be purified on heparin Sepharose columns [42]. The biological relevance of the heparin-TG2 interaction was studied by co-immunoprecipitation analysis using human osteoblast cells and Swiss 3T3 fibroblast cells with tetracycline-regulated inducible expression of TG2, wherein TG2 was shown to interact directly with syndecan-4. Later, two independent groups identified two sites for HS interaction on TG2 [22, 43] (Table 1, Fig. 2).
HSPGs bind extracellular ligands through HS and influence their biological activity by affecting their protein stability, activity, and conformation. The best example is the activation of the serine protease inhibitor antithrombin. The negatively charged HS chains of heparin interact with the positively charged amino acid residues within antithrombin leading to a conformational change accompanied by an increase in the rate of inactivation of the serine proteases (e.g., Factor IIa and Xa) involved in coagulation [44, 45]. HSPGs regulate other functions such as cell-surface localization, membrane secretion/internalization, and protein interactions [46, 47]. Indeed, TG2 also interacted with the HS chain of syndecan-4, which influenced trafficking of TG2 to the cell surface, subsequently affecting its cell-surface crosslinking activity. Mouse dermal fibroblast cells with targeted deletions of syndecan-4 were used to explore the role of syndecan-4 in regulating TG2 activity and trafficking [17].
Syndecans also act as co-receptors for both ECM components and soluble ligands. In the absence of FN, TG2 can promote cell adhesion by simultaneously associating with integrin and syndecan [48].
Growth factor receptors and low-density lipoprotein receptors (LDL) as TG2-interacting partners
Extracellular TG2 was shown to interact with two growth factor receptors, platelet-derived growth factor receptor (PDGFR) and vascular endothelial growth factor receptor (VEGFR). TG2 crosslinks VEGFR but it is not yet known if it crosslinks with PDGFR [49]. PDGFR interacts with integrin-associated TG2 on the surface of human fibroblast cells, thus TG2 serves as a scaffolding protein bringing two cell-surface receptors together. TG-mediated scaffolding of PDGFR and integrin contribute to amplifying joint PDGFR/integrin signaling from the membrane to the cell interior and stimulating cell migration and adhesion [50].
In addition to binding to integrin and growth factor receptors, extracellular TG2 is also capable of signaling through low-density lipoprotein receptor related proteins 5 and 6 (LRP 5/6). TG2 was shown to interact directly with LRP5/6 on smooth muscle cells, which in turn activate the β-catenin pathway and regulate calcification and osteoblastic transformations in vascular smooth muscle cells [51].
LRP1, a member of the LDL receptor superfamily, was also recently shown to interact directly with TG2 expressed on the surface of the fibroblast cells and with extracellular TG2 [14]. LRP1 is the major endocytic receptor that regulates internalization and intracellular trafficking of numerous ligands [52], including those of TG2-interacting partners, β1 and β3 integrin, and fibronectin [53, 54]. LRP1 regulates cell adhesion and migration and participates in outside-in signaling by internalizing and degrading TG2. In the absence of LRP1, TG2 gets upregulated on the cell surface leading to increased adhesion to the ECM [14]. LRP1 can weakly bind β1 integrin, however, with TG2 overexpression, this interaction became more prominent, indicating that TG2 associated with LRP1 and promoted the formation of ternary complexes with LRP1 and integrin [14]. These observations highlight the ability of TG2 to act as a scaffold to modulate the functions of proteins via their complexes, ultimately affecting cell–matrix adhesion, migration, receptor-mediated signaling, and endocytosis [14].
Role of ECM proteolysis in regulating TG2-mediated cell–matrix interactions
ECM and cell–matrix interactions are regulated by crosslinking of ECM proteins and also by proteolysis of these proteins. Matrix metalloproteinases (MMPs) are proteinases that participate in ECM degradation, thus serving as important molecular players in regulating the ECM macromolecules and their interactions with the cell surface [55]. Recent findings show that in addition to matrix breakdown, some MMPs are also involved in degradation of the TG2 present on tumor cell surfaces. MT1-MMP, a membrane-bound type 1 metalloprotease, was shown to proteolyze TG2 in vitro and in cultured fibrosarcoma cells [56]. MT1-MMP is also an activator of MMP2, a secreted and soluble metalloprotease [57, 58]. Activated MMP-2 was reported to interact with the core domain of TG2 and direct its cleavage. This accelerates TG2 degradation because of the synergistic/cooperative effect of both soluble and membrane-bound MMPs [59]. MMP2-mediated cleavage leads to elimination of the catalytic and adhesion activity of TG2. Interestingly, cells overexpressing TG2 showed reduced levels of active MMP2 when compared to cells expressing anti-sense TG2 or vector control. It seems that during overexpression, TG2 interacts with a MMP2 activation intermediate rather than the inactive zymogen or fully active MMP2, thus protecting itself from proteolytic cleavage by the fully mature MMP2 enzyme [59]. Because TG2 regulates cell–matrix interactions and migration/invasion of malignant and host cells, degradation of this multifunctional protein by MMP2 or its protection via inhibition of MMP2 activity likely plays a significant role in adhesion/migration-related physiopathological conditions.
Cancer cell adhesion, migration, and proliferation
Quantitative downregulation of TG2 in prostate cancer tissue sections is considered to be a strong biomarker for prostate cancer [60, 61]. To understand the potential functions of TG2 in prostate cancer, its interacting partners were explored using the PC3 prostate cancer cell line. TG2 was reported to interact with protein kinase A (PKA) anchor protein 13 (AKAP13) [61]. AKAPs, with more than 50 members known to date, are structurally diverse but functionally similar proteins defined on the basis of their ability to bind to PKA [62, 63]. They are capable of anchoring PKA at specific cellular locations, where they are responsible for phosphorylating local substrates in response to cAMP activation, enabling organelle-specific PKA function [63]. Analysis of the TG2 amino acid sequence revealed the presence of a site (rRRrS) that has a very high specificity for PKA-mediated phosphorylation [64–66].
It was recently reported that TG2 is phosphorylated by PKA at serine 216, which regulates TG2 function and mediates protein–protein interactions [67]. Because AKAP13 acts as a scaffolding protein anchoring PKA, it is potentially involved in the interaction between PKA and TG2, which in turn leads to substrate phosphorylation and could contribute to regulating prostate cancer. AKAP can also bind other signaling proteins besides PKA [68]. For example, AKAP binds to RhoA protein, which is overexpressed in the highly invasive PC3 prostate cancer cell line and plays a significant role in cancer progression [69]. RhoA protein is also a substrate of TG2, which gets cross-linked through TG2 activation, leading to stress fiber formation [70, 71]. Because stress fiber formation leads to cellular polarity and migration, which is altered during cancer progression [72, 73], assembly of RhoA, TG2, PKA, and AKAP13 possibly could contribute to cancer cell polarity and migration [61].
TG2 is highly expressed in drug-resistant and metastatic cancer cells [74–76]. Increased expression of TG2 in breast and pancreatic cancer cells leads to constitutive activation of nuclear factor-κB, making the cells drug resistant [77]. TG2 was found to interact with PTEN (a phosphatase and tensin homologue deleted on chromosome 10, also called MMAC1 or TEP1), a tumor suppressor protein in pancreatic ductal adenocarcinoma cell lines (PDAC). Expression of TG2 in PDAC cells leads to constitutive activation of focal adhesion kinase and its downstream PI3K/AKT signaling pathway [77]. PTEN regulates cell growth, invasion, migration, and focal adhesion functions by negatively regulating the PI3K/AKT pathway [78, 79]. Therefore, it was proposed that TG2 could affect FAK/PI3K/AKT pathways by regulating PTEN expression and function. In accordance with this idea, aberrant expression of TG2 inversely regulates PTEN expression by inhibiting PTEN phosphorylation, leading to the subsequent ubiquitination and degradation of PTEN by the proteasomal pathway [80]. Down-regulation of PTEN expression in the presence of TG2 leads to increased tumor growth in vivo in nude mice and to the poor survival of PDAC patients [80].
Recently, heat shock protein 70 (Hsp70) was reported as a novel interacting partner of TG2. Hsp70 co-localized with plasma membrane-associated TG2 to the leading edges of EGF-stimulated HeLa cells, as well as to the leading edges of constitutively migrating MDAMB231 breast cancer cells [81]. A TG2 active site mutant (Cys277Ser) was also capable of immunoprecipitating Hsp70, indicating that crosslinking activity was not needed for this interaction. HSP70 inhibitors, which block the ATP hydrolytic activity of Hsp70, inhibited the migration of TG2 to the leading edges of EGF-induced HeLa cells and MDAMB231 breast cancer cells. This confirms that the chaperoning activity of heat shock protein can regulate cell migration by targeting TG2 to leading edges of migrating cells. This also demonstrates that proper localization of TG2 to cellular leading edges is essential for its ability to promote cancer cell migration [81].
The Rho family of small GTPases, including Rho, Rac, and Cdc42, are critical regulators of a variety of cellular functions including cytoskeleton rearrangement and membrane trafficking, which are essential during cell adhesion and migration processes [82]. GTPase-activating proteins (GAPs), such as Bcr and Abr, regulate the GTPase activity of these proteins. TG2 was found to interact directly with the C-terminal GAP domain of Bcr and Abr using COS-1 and Swiss 3T3 cells. The interaction was more prominent when TG2 was in an open confirmation because the addition of calcium or GTP binding mutant forms of TG2 both increased the level of the TG2–Bcr/Abr interaction [82]. Bcr was later reported to serve as a substrate of TG2 in human pulmonary artery endothelial cell lines (HPAECs) but crosslinking was only observed under specific and extreme stress conditions. The crosslinking function required the N-terminal oligomerization domain of Bcr while the GAP domain was important for direct interaction with TG2. This suggests a dual function of Bcr both as a substrate and as a non-covalent interacting partner. Abr was not crosslinked with TG2 because of the absence of this oligomerization domain [83]. Because Bcr binds to its substrate Rac through its GAP domain, TG2 was found to efficiently block Rac binding due to the presence of overlapping binding sites and inhibited Bcr GAP activity towards Rac. These studies show that TG2 regulates Rac activity by modulating Bcr activation, which subsequently could affect Rac-mediated cellular processes [82, 83]. Because the interaction between Bcr and TG2 occurs under physiological conditions, whereas crosslinking takes place under extreme stress conditions, Bcr is likely regulated by TG2 via different mechanisms depending upon the physiological state of the cell.
Endostatin, which is a C-terminal fragment of the α1 chain of collagen XVIII, inhibits angiogenesis and tumor growth [84], and GPR56, which is a newly described family of G protein-coupled receptors, also suppresses melanoma metastasis and tumor growth [85]. Endostatin and GPR56 proteins were shown to interact with extracellular TG2 in endothelial cells and in the lung tissue, respectively. Endostatin interacts with TG2 in a calcium-dependent manner and most probably binds at the GTP binding site (Fig. 2) [86]. At the endothelial cell surface, endostatin inhibits both the extracellular activation of proMMP-2 by inhibition of MT1-MMP as well as the catalytic activity of MMP-2, thus blocking the invasiveness of tumor cells. As we discussed above, TG2 interacts with the MMP2 activation intermediate thereby inhibiting MMP2 activity in a manner similar to that of endostatin. Therefore, it is speculated that TG2 and endostatin together assist in ECM remodeling. The biological relevance of the GPR56 and TG2 interaction was recently explored wherein GPR56 was reported to internalize and degrade TG2 subsequently affecting ECM protein deposition and cell-matrix adhesion functions [87]. TG2 and its crosslinking activity promote melanoma growth but GPR56 antagonizes this affect by degrading TG2 [87]. GPR56 binds to the C-terminal domain of TG2 [85].
Angiocidin is another protein, which like fibronectin and Bcr, serves as a non-covalent binding partner and transamidating substrate of TG2. Angiocidin is an ECM protein expressed by endothelial and tumor cells. Recently, angiocidin was shown to colocalize with TG2 in the ECM of endothelial cells and interact non-covalently with TG2 via its C-terminal integrin and collagen-binding domain [88]. Interaction of angiocidin and TG2 led to inhibition of the TG2-fibronectin interaction as well as inhibition of fibronectin deposition in the ECM of endothelial and tumor cells, which ultimately contributed to modulation of adhesion and migration activities of TG2 [88].
Trans-membrane signaling
Transglutaminase 2 is also referred to as Ghα because of its GTPase activity combined with its ability to interact and signal through receptors. Ghα was shown to interact with several receptors, thus facilitating hormone-receptor mediated transmembrane signaling pathways like those involving the classical G-protein-coupled receptors. Involvement of TG2 in hormone receptor-mediated signaling was first demonstrated by isolation and characterization of an α1-agonist α1B-adrenoceptor (AR)-TG2 complex from rat liver membranes [5]. De novo purification of TG2/Ghα revealed that it is associated with Ghβ and α1-AR, which modulates the GTP binding and GTPase function of Ghα, facilitating receptor-mediated signaling [5]. There are three alpha-1 adrenergic receptor subtypes: α1A-AR, α1B-AR, and α1D-AR. Interaction between α1-AR and TG2 was subtype-specific, as α1B-AR [89, 90] and α1D-AR [90] were shown to interact with TG2/Ghα while α1A-AR did not interact [90]. This interaction was independent of its transamidation activity because the active site mutant (Cys277Ser) could not interact with α1B and α1D adrenoceptor [90]. Peptide screening and site-directed mutagenesis identified multiple binding sites for α1B-AR including L547–I561, R564–D581, and Q633–E646 on TG2 [89] (Table 1, Fig. 2). Because the regions L547–I561 and R564–D581 are close to the GTP binding motif of TG2, it clearly indicates that α1-AR interacts with GTP-TG2 to facilitate combined signaling events.
TG2 and α1-AR-mediated signaling leads to activation of phospholipase C δ1 (PLC δ1), which in turn stimulates the release of two secondary messengers, inositol-1,4,5-triphosphate (IP3) and diacylglycerol (DAG), by hydrolyzing phosphatidylinositol-4-biphosphate (PIP2) [91] [92, 93]. PLC δ1 is the best-known effector molecule in TG2-mediated signaling to date. There are three subtypes of phospholipase C: PLC β, γ, and δ, of which PLC δ1 has been shown to directly interact with TG2 [92]. A region of 12 amino acids between Leu661 and Lys672 in TG2 was identified as the PLC interaction site via a series of co-immunoprecipitation studies, while the TG2 interaction site in PLC δ1 was shown to be located within the C2 domain of the enzyme (Fig. 2) [94, 95]. PLC δ1 not only serves as a PIP2-hydrolyzing enzyme but also acts as a guanine nucleotide exchange factor (GEF) and GTP hydrolysis-inhibiting factor (GHIF) for TG2. These functions of PLC δ1 for TG2 facilitate TG2-mediated signaling [95].
Other TG2 coupled receptors, which interact with TG2 and mediate transmembrane signaling, are the thromboxane receptor [6] and the oxytocin receptor [7]. The thromboxane receptor-TG2 interaction was identified in cardiovascular cells [6] while the oxytocin receptor-TG2 interaction was identified in human myometrium [96]. Both of the interactions are regulated by activation of PLC and release of IP3.
Calreticulin (CRT) is a multifunctional chaperone protein involved in protein folding, maturation, and trafficking. It was found to interact with the GDP bound form of TG2, downregulating both its GTPase and TGase activities [97, 98]. The identity of calreticulin as a TG2 binding partner was recognized when Gβh, which consistently co-purified with Ghα/TG2, was sequenced. Amino acid sequencing and immunological studies strongly suggested that Gβh was nothing more than the CRT protein [98]. When the regulatory role of CRT in the GTPase activity of TG2 was examined, CRT inhibited GTP-binding to TG2 and TG2-mediated GTP hydrolysis in a concentration-dependent manner [97]. Because CRT inhibits GTP binding to TG2, these results also indicate that CRT induces TG2 to rearrange into an inactive conformation. CRT modulated the TGase activity of TG2 by shifting the calcium requirement to higher concentrations and also by inducing the inactive conformation of TG2. Another mechanism by which CRT influenced TGase activity was related to its allosteric hindrance of CRT. It was clearly shown that upon treating GDP-bound TG2 with increasing CRT concentrations, there was a decrease in the Vmax of the TG2 enzymatic activity without an increase in the substrate or the calcium requirement [97]. Though calreticulin is a calcium binding protein, its interaction with TG2 occurred in the absence of calcium, and was not disrupted upon increasing the calcium concentration [97]. Like TG2, calreticulin is also involved in cell growth, differentiation, and adhesion. Calreticulin regulates cell adhesion by interacting with integrin and participating in integrin-mediated signaling [99–101]. It could be possible that both proteins act in concert to regulate these functions.
Nuclear translocation and gene expression regulation
Accumulating evidence shows that cytoplasmic TG2 under specific physiological conditions translocates to the nucleus where it either crosslinks nuclear proteins or interacts with them non-covalently [13, 77, 102–104]. A recent review from Kuo and Kojima provides new insights into the molecular mechanisms of TG2 localization and its function in the nucleus [105]. Nuclear TG2 regulates gene expression by interacting with transcriptional factors and related proteins, including hypoxia inducible factor (HIF) 1, Sp1, and histones. These interactions control cell growth, survival, differentiation, apoptosis, and signaling events in several pathological disorders [15, 106–108].
Nuclear translocation
The transamidation activity of nuclear TG2 increases with a rise in the calcium ion concentration. The presence of maitotoxin or calcium ionophores increases the intracellular calcium concentration and results in the translocation of TG2 to the nucleus and a subsequent increase in transamidation activity [13, 77]. Importin α3, a nuclear transport protein, was identified as one of the binding partners of TG2 in a non-small cell lung cancer (NSCLC) cell line. Co-localization of TG2 with importin α3 was initially detected in the cytosol but treating the NSCLC cells with retinoic acid led to an increase in the amount of TG2/importin α3 complex in the nucleus, which indicates that importin α3 can mediate active transport of TG2 into the nucleus [109]. Primary sequence analysis of TG2 showed the presence of a putative bipartite nuclear localization sequence (NLS). This is in line with the active transport of TG2 into the nucleus [109] by proteins, which interact with the NLS motifs. Nuclear TG2 has been suggested to be involved in apoptosis and cell-cycle progression because of its interaction with nuclear histones and Rb. Thus, it is possible that importin or other mechanisms mediating active transport of TG2 are executed only when a cell needs nuclear TG2 during specific physiological conditions (for example during apoptosis).
The GDP bound form of TG2 interacts with eukaryotic initiation factor 5A (eIF5A) and regulates its intracellular localization [110]. Upon GDP/GTP exchange, eIF5A is released into the cytoplasm, perhaps to bind other cytosolic target proteins involved in protein synthesis or to return to the nucleus to participate in RNA trafficking. Thus, it seems that the GTP binding/GTPase cycle of TG2 serves as a timing device to regulate the changes in the cellular localization of eIF5A, thereby facilitating eIF5A to participate in protein synthesis or RNA trafficking between the cytosol and nucleus [110].
Transcriptional regulation of genes involved in tumor metastasis, hypoxia, and inflammation
As mentioned previously, members of the MMP family have been implicated in the metastasis of tumor cells [111]. TG2 was shown to down-regulate MMP-9 transcriptional activity by blocking the AP1 binding site present on the MMP-9 gene promoter. MMP9 gene transcription is initiated by binding of the c-Fos and c-Jun dimer to the AP1 binding site. TG2 was shown to interact non-covalently with c-Jun in the nucleus of cardiomyoblasts, which prevented c-Fos and c-Jun dimerization and subsequent downregulation of MMP9 gene transcription [112]. This transcriptional regulation of the MMP-9 gene supposedly regulates ECM remodeling.
TG2 was shown to regulate β-catenin signaling by associating with c-Src. β-catenin is a transcriptional regulator with oncogenic activity, which is associated with increased Wnt signaling. In normal cells, β-catenin is found to be transcriptionally inactive because it is often associated with E-cadherins. In the tumor cells where TG2 expression is often observed to be very high, it is reported that increased TG2 activity leads to enhanced TG2-FN-β-integrin complexation and subsequent recruitment and activation of c-Src. Activation of c-Src then leads to phosphorylation of β-catenin, allowing it to detach from the cytoplasmic domain of E-cadherin and rendering it transcriptionally active. Disruption of the β-catenin-E cadherin interaction, which was responsible for stabilizing the cell–cell interaction, now allows cells to become motile and invasive. At the same time, β-catenin becomes available in the cytosol, translocates to the nucleus, and activates the Wnt transcriptional machinery [113].
TG2 was shown to be up-regulated and neuroprotective in response to oxygen and glucose deprivation (OGD) in both primary rat cortical neurons and in a human neuroblastoma cell line leading to delayed neuronal cell death [106]. Under these conditions, TG2 was reported to bind non-covalently to hypoxia-inducible factor 1β (HIF1β), which prevented HIF1β and HIF1α heterodimerization to generate functional HIF1 transcription factor. Functional HIF1 transcription factor regulates the expression of the Bnip3 gene under hypoxic conditions by binding to the hypoxic response elements (HRE) located in the promoter region of the Bnip3 gene. Bnip3 (BCL/adenovirus E1B 19-kDa protein-interacting protein) is a proapoptotic gene whose transcriptional activity ultimately is reduced in neuronal cells in hypoxic conditions because of TG2 upregulation and subsequent unavailability of functional HIF1 transcription factor. Therefore, TG2 indirectly plays a protective role in the brain in response to OGD and hypoxia [114, 115].
Retinoblastoma (Rb), an anti-apoptotic protein, could also attenuate the up-regulation of the pro-apoptotic, HIF-dependent Bnip3, as well as bind HIF1α and enhance the HIF response [116]. Rb was identified as a potential substrate of TG2 [107], however it was also shown to interact directly with a transamidation inactive mutant of TG2 in the nuclear fraction of HEK293 cells transiently transfected with TG2 wild-type and mutant constructs [15]. Because TG2 serves as a scaffold for both Rb and HIF1β, it could contribute to cellular protection at the transcriptional level [105].
TG2 is also proposed to be involved in wound healing and inflammation [117–119]. Major cellular components that trigger inflammation are histamine, prostaglandins, and several inflammatory cytokines (TNF-α, IL-1, IL-8, IFNγ). However, peroxisome proliferator-activated receptor (PPAR)γ, which belongs to the family of nuclear receptors, decreases inflammation. Cystic fibrosis (CF) is characterized by chronic airway inflammation and is caused by mutations in the transmembrane conductance regulator (CFTR) gene, which encodes for cAMP-regulated chloride channels [108, 120, 121]. TG2 was reported to be highly up-regulated in nasal polyp mucosa from CF patients and CFTR-defective cell lines [108] and accelerated inflammation most likely because of crosslinking of PPARγ and subsequent ubiquitination and degradation by the proteasome [121]. CF airways released high levels of reactive oxygen species (ROS), which was the main cause of increased TG2 activity [121]. ROS not only increased TG2 activity but also mediated TG2 SUMOylation by facilitating interaction of TG2 with Small ubiquitin-like modifiers (SUMO1) and PIASy, a SUMO E3 ligase in a human CF bronchial epithelial cell line (IB3-1) [108]. SUMOylation ultimately led to increased TG2 levels in the cell by inhibiting ubiquitination and degradation of TG2 by the proteasome [121]. This indicates that there is reduced inflammation when TG2 undergoes ubiquitination and degradation by the proteasome but TG2 accelerates inflammation when it is SUMOylated. This leads to increased TG2 levels in the cell, causing crosslinking and degradation of the anti-inflammatory molecule PPARγ [108].
Another example of TG2-regulated allergic inflammation was observed in antigen-stimulated basophilic leukemia cells (RBL2H3). Antigen stimulation of RBL2H3 cells increased TG2 expression and concomitantly increased TG2 activity. TG2 expression could be down-regulated upon treating the cells with the anti-allergen, hyaluronic acid [122]. Antigen stimulation of the cells also led to an increase in EGFR, NF-κB, and ROS levels, which are important mediators of allergic inflammation. Treating the cells with inhibitors of EGFR and NF-κB led to a decrease in TG2 expression, which suggested that TG2 contributes to allergic inflammation via a combination of EGFR and NF-κB signaling pathways [122, 123]. Moreover, TG2 was also shown to interact with rac1 in the antigen-stimulated cells. Because rac1 has been implicated in generating ROS in several cell types by activating NADPH-oxidase, it is possible that TG2 regulates allergic inflammation by interacting with rac1 and regulating its activity. This speculation was supported by observation of an inhibition of rac1 activity coupled with inhibition of inflammation after treating RBL2H3 cells with a TG2 inhibitor [124].
Role of nuclear TG2 in neurological pathologies
TG2 has been reported to be involved in several neurological diseases such as Alzheimer’s [125, 126] and Huntington’s disease (HD) [127, 128]. To understand the role of TG2 in neurological disorders, neuroblastoma cell lines (SK-N-BE) were used to identify cellular proteins, which might interact with TG2 and contribute to pathogenesis. Glutathione S-transferase (GSTP1) and β-tubulin were identified as interacting partners of TG2 in these neural cells. GSTP1 acted as a substrate because it could form high molecular weight complexes in the presence of TG2 in vitro while β-tubulin was found to interact in a transamidation-independent manner [129]. Aggregates of huntingtin, a cytosolic protein, are a neuropathological hallmark of HD. Huntingtin is found to be primarily associated with microtubules as well as other huntingtin-associated proteins [130, 131]. Because TG2 co-localizes with huntingtin and β-tubulin, it was proposed that microtubules could serve as a binding site, bringing together TG2, huntingtin, and other TG2 substrates for TG2-dependent crosslinking [129]. Another protein reported to interact with TG2 and contribute to HD pathology was a calcium binding protein, calmodulin. Calmodulin was found to co-localize with TG2 and mutant huntingtin in the intranuclear inclusions in the HD brain. TG2 and mutant huntingtin could also be immunoprecipitated using anti-calmodulin Sepharose columns. Previously, it had been shown that calmodulin probably increased transglutaminase activity by regulating the calcium concentration [132]. Because mutant huntingtin interacts with both TG2 and calmodulin in HD brain, it was suggested that huntingtin might be involved in increasing TG2 activity and subsequently its own crosslinking by bringing calmodulin and TG2 in close proximity. Thus, calmodulin indirectly regulates TG2-mediated crosslinking of huntingtin and formation of stable aggregates and inclusions in HD brain [133].
Interaction of TG2 with histones
Histone H3 and H2B were shown to interact directly with TG2. Whether this interaction is a classical substrate–enzyme interaction or they are true interacting partners of TG2 is not known [127, 129]. Although in vitro crosslinking assays confirmed all core histones to be excellent glutaminyl substrates of TG2, it is not yet confirmed whether this crosslinking is observed in vivo as well [134, 135]. Temperature greatly affects the crosslinking pattern of TG2, and this crosslinking pattern is known to be different for intact nucleosomes when compared with free histones [134], therefore it is important to explore whether all the four core histones behave strictly as substrates or if they also interact with TG2 in a calcium-independent manner. In vivo crosslinking has been shown only between H2B and H4 in the sperm of the starfish Asterina pectinifera [136]. The different crosslinking patterns observed for free histones and intact nucleosomes indicate that TG2 may be involved in modulating the conformation of nucleosomes. Not much information is available on the physiological relevance of the TG2–histone interaction but it is proposed to be involved in the condensation of chromatin in apoptotic nuclei [134]. Apoptotic bodies show chromatin condensation in their perinuclear space and because we know that TG2 regulates apoptosis by crosslinking proteins involved in apoptosis, it has been speculated that the TG2–histone interaction may contribute to this condensation reaction. TG2 was also reported to catalyze crosslinking of polyamine binding proteins [137, 138] and polyamines are involved in modulation of chromatin structure and function [139]. Because putrescine, a polyamine, is readily incorporated by histones via a TGase reaction, TG2 could contribute to polyamine-mediated chromatin condensation by direct interaction and covalent modification of histones.
Apoptosis and autophagy
TG2 was shown to regulate apoptosis and it has been proposed that the type of cell, the kind of stressor, intracellular localization, and transamidation activity of TG2 determine whether it promotes pro-apoptotic or anti-apoptotic responses [140]. Transient transfection of wild-type TG2 targeted to different intracellular compartments confirmed the pro-apoptotic nature of cytosolic TG2. On the contrary, a TG2 active site mutant localized to the nuclear compartment reduced apoptosis [15]. Membrane-targeted TG2 had neither a pro-apoptotic effect nor anti-apoptotic effect, thus confirming that cellular localization of TG2 was essential for determining its influence on cell death and survival.
TG2 was shown to promote cell death by interacting with the pro-apoptotic Bcl2 family member Bax. It was proposed that TG2 possesses a functional BH3 motif in the catalytic core domain of TG2 [141]. Expression of mutated TG2 where the BH3 motif was either entirely deleted or mutated resulted in cells that were insensitive to cell death compared to cells expressing wild-type TG2. Co-IP experiments confirmed that TG2 itself interacts with the pro-apoptotic Bcl2 family members Bax and Bak. However, the level of interaction increased in the presence of apoptotic stimuli only in the case of the TG2–Bax interaction. Although the TG2–Bax interaction takes place in the cytosol, it is speculated that upon interaction, Bax undergoes conformational change and aggregation, mediating translocation of the entire complex to the mitochondria.
14-3-3, a serine/threonine binding protein, is involved in the regulation of apoptosis through multiple interactions with proteins of the core mitochondrial machinery, pro-apoptotic transcription factors, and their upstream signaling pathways [142]. 14-3-3 impacts cellular processes by binding to the specific phosphorylated sites on the target protein, thereby driving conformational changes or directing interactions between its target proteins and other molecules [143]. This protein was found to interact with TG2 in vitro, as well as in TG2-expressing cells. The interaction was more prominent when TG2 undergoes PKA-mediated phosphorylation compared with when it was not phosphorylated [144]. Two 14-3-3 binding motifs were identified in the TG2 protein sequence, and although not completely identical to the consensus sequences identified previously, the TG2 peptide fragments (TG2 209–223) containing these two binding motifs could pull down 14-3-3 protein from cell lysate. The interaction occurred only when the peptide fragment was phosphorylated. Phosphorylation at serine 215 facilitated 14-3-3 binding while phosphorylation at serine 216 was absolutely essential for the interaction. Thus, PKA-mediated phosphorylation creates a 14-3-3 protein binding motif on TG2, which is speculated to affect various activities and functions of TG2 including regulation of apoptosis [144]. Because the BH3 motif on TG2 overlaps with the 14-3-3 binding motif and the 14-3-3 binding protein was also reported to target the BH3 only proteins BAD and BIM, it is possible that 14-3-3 regulates apoptosis by interacting with TG2 and BH3 only protein family members.
TG2 was recently reported to protect retinoic acid (RA)-treated neuroblastoma SH-SY5Y cells from apoptosis induced by NMDA-evoked excitotoxic stress [145]. TG2 was found to interact with small heat shock protein Hsp20 in the RA-treated SH-SY5Y cells but the interaction was strongly reduced upon treating the cells with NMDA. This was likely because of degradation of Hsp20. NMDA-treated cells also demonstrated reduced levels of the pro-apoptotic enzyme caspase 3. However, specific inhibition of NMDA-induced TG2 activation by R283 (an active site TG2 inhibitor) reversed these effects. In the same cell model, Hsp20 was found to interact with Hsp27. We know from previous studies that Hsp27 acts as a “scaffolding” protein, leading to the activation of pro-survival molecules (e.g., MAPK-activated protein kinase-2 and serine/threonine kinase Akt) and inhibition of pro-apoptotic molecules (e.g., caspase 3). Hsp27-mediated activation/inhibition of these molecules results in cytoprotection in different models of neuronal injury [146]. These results indicated that the overexpression of TG2 because of NMDA treatment led to modulation of the anti-apoptotic functions of the Hsp20/Hsp27 complex and reduction in the activity of caspase 3, ultimately protecting the cells from apoptotic damage [145]. In a previous study by the same authors, TG2 was shown to promote apoptosis when the cells were treated with a low dose (100 µM) of NMDA [147, 148], however, in the present study, TG2 inhibited apoptosis when the same cells were treated with a high dose (1 mM) of NMDA. The authors hypothesized that NMDA increases the intracellular calcium level, which may contribute to activation of other TG2 functions different from crosslinking, playing a protective role against excitotoxicity [145].
TG2 was also shown to regulate cell death by initiating the phagocytosis of apoptotic cells [38]. For the efficient engulfment of apoptotic cells, TG2 is required to interact with integrin β3 and milk fat globule EGF factor 8 (MFG-E8). MFG-E8 is released by the activated macrophages, and in turn binds to the phosphatidylserine of apoptotic cells, acting as a bridge between apoptotic cells and the phagocyte receptors, integrin β3 and TG2 [38, 149]. TG2 cross-linking activity is not required for this process [23, 38].
Recently, a role of TG2 in autophagosome maturation, formation, and clearance of aggresomes by autophagy was suggested [150]. Autophagy is a cellular process that is responsible for removal of protein aggregates that cannot be processed by proteasomes. TG2 was found to interact with p62 and NBR1 in 2fTGH and HEK293 cells overexpressing TG2 [150]. Both of these interacting partners are autophagy cargo proteins, which bind to ubiquitinated proteins and mediate their delivery into autophagosomes. Whether TG2 activity is essential for its interaction with p62 and NBR1 has not yet been explored [150].
General comments
We assembled 39 interacting partners, which impact cellular functions of TG2 using information from a literature search and database (STRING [151] and Integrated human protein interaction network [152]) analysis. Most partners were identified from the literature search and via text mining in STRING. The latter have relatively low scores, indicating experimental difficulties and uncertainty in detecting such PPIs. Although the present list (Table 1) is far from complete, we analyzed those structural features which could contribute to further interactions and thereby the multi-functionality of TG2.
The listed interaction partners could bind to either an open (e.g., Bcr and Abr) or closed (e.g., calreticulin) form of TG2. The 14-3-3 protein binding motif was exposed only when TG2 was phosphorylated, thus phosphorylation of TG2 creates a novel protein binding site, which could further mediate protein–protein interactions and TG2 regulation. This scenario clearly illustrates the significance of unique structural features which influence protein–protein interactions and hence, TG2 functions. Structural characterization of TG2 would undoubtedly provide new insights from which to explore interactions of TG2 with various proteins.
Significance of short linear motifs and intrinsically disordered regions in TG2 for non-enzymatic TG2–protein interactions
Sequences of all TGs are highly conserved, which implies structural similarities among TG family members. The known TG structures for fXIIIA [153], sea bream TG [154], human TG2 [155, 156], and human TG3 are composed of four domains, as detailed previously [157]. TG2 differs from other transglutaminases in its number of calcium binding pockets and GTP/GDP binding regions. Crystal structures of Factor XIIIA [153, 158] and TG3 [157] reveal one and three calcium binding pockets, respectively, and no GTP binding regions. The crystal structure of TG2 with bound calcium is not available yet, but biochemical experiments revealed six calcium ion binding sites, which were confirmed by site-directed mutagenesis [159, 160]. Calcium and GTP differentially regulate TG2 activities. Calcium is reported to activate the protein by keeping TG2 in an open conformation, while GTP inactivates TG2 by keeping it in a closed conformation, making the catalytic region inaccessible for substrates and subsequent catalysis [155]. It is possible that binding of six calcium ions and GTP fine-tune TG2 activities in different cellular compartments in vivo. Because TG2 adopts different conformations in the cell, transitions between these structural states could be influenced by several transient interacting partners.
We assembled a list of 39 proteins that are not substrates of TG2 but, via non-enzymatic interactions, interfere with its activities in different cell compartments as reviewed above. The binding regions of non-enzymatic TG2 partners are only defined for eight proteins, fibronectin, syndecan, endostatin, SUMO 1, PLC-δ1, α1-adrenoceptor, Bax/Bak, and the 14-3-3 protein. Most of these proteins contact with short (3–20 AA) peptide regions of low amino acid complexity, often referred to as short linear motifs (SLiMs). In general, SLiMs mediate highly specific protein interactions with moderate affinities [161], which are often exploited in signaling pathways. The majority of SLiMs are embedded in protein segments, which lack a well-defined three-dimensional structure [162]. These protein regions are intrinsically disordered (ID) and are present in ~40 % of the human proteome [163]. We hypothesized that non-enzymatic interactions of TG2 are also located in ID regions.
To probe this tenet, we determined ID regions in TG2 using two bioinformatics predictors: PONDR VSL1 [164] and IUPred [165, 166] with two window sizes. A total of 13 ID regions containing more than five amino acids in TG2 were identified (listed in Table 2). In both the open (PDB: 1kv3) and closed form (PDB 2q3z), these are located along a spiral surrounding the GDP binding site (Fig. 2). SLiMs were predicted using the server of the eukaryotic linear motif (ELM) database [167] and those motifs which overlap with the ID regions are presented in Table 2. As we investigated potential binding motifs for non-enzymatic partners, we excluded consensus protease cleavage sites.
Table 2.
ID regions embedding ELMs in TG2
| ID region | PONDR | IUPred short (>0.5) | IUpred long (−0.44) | ELM | Explanation | Interacting partner |
|---|---|---|---|---|---|---|
| 1–8 | + | + [1–6] | − |
LIG_BIR_II_1 [1–5] LIG_SUMO_SBM_1 [5–8] LIG_SUMO_SBM_2 [3–7] |
Inhibitor of apoptosis binding motif; SUMO binding site | |
| 14–21 | + | − | + [12–19] | LIG_integrin_isoDGR_1 [17–19] | Integrin binding site | Integrin β1, β3 |
| 65–74 | + | + [72–78] | + [61–78] |
LIG_TRAF2_1 [67–70] LIG_SH3_3 [59–65] |
TRAF2 binding site, SH3 binding domain | |
| 122–125 | + | − | − |
MOD_GSK3_1 [115–122] MOD_NEK2_1 [119–124] |
NEK kinase and GSK3 phosphorylation motif | |
| 151–158 | + | − | − |
LIG_TRAF2_1 [152–155] LIG_SH2_STAT5 [149–152] LIG_SH2_SRC [149–152] |
TRAF2 binding site; SH2 domain binding motif; Src SH2 domain binding motif | |
| 208–217 | + | − | − |
DOC_MAPK_1 [213–220] DOC_WW_Pin1_4 [213–218] MOD_GSK3_1 [209–216] MOD_CK1_1 [212–218] MOD_PKA_2 [212–218] MOD_ProDKin_1 [213–219] |
Docking interaction in MAP kinase cascade (exemplified cJun); binds WW domains, involved in proline directed phosphorylation signaling pathways; phosphorylation motifs (CK1, GSK3,PKA, proline-directed kinase) | Syndecan-4 14-3-3 BAX, BAK |
| 358–367 | + | + [353–368] | + [351–368] |
DOC_WW_Pin1_4 [357–362] LIG_FHA_2 [358–364] LIG_TRAF2_1 [360–363] LIG_TRAF6 [359–367] MOD_CK2_1 [357–363] MOD_ProDKin_1 [357–363] |
Binds WW domains, involved in proline-directed phosphorylation signaling pathways; TRAF2 and TRAF6 binding motif; CK2 and proline-directed kinase phosphorylation motif | SUMO1 |
| 411–414 | + | − | − |
LIG_ACTIN_WH2_2 [409–427] MOD_GSK3_1 [408–415] MOD_PLK LIG_SUMO_SBM_2 [419–422] |
Actin binding motif (WH2 domains); GSK3 and PLK phosphorylation site; SUMO binding motif |
|
| 428–473 | + | + [437–467] | + [433–463] |
DOC_USP7_1 [446–450] LIG_TRAF2_1 [449–452] MOD_CK1_1 [427–433] MOD_CK2_1 [446–452] |
USP7 binding motif; TRAF2 binding motif; CK1 and CK2 phosphorylation motif | |
| 597–602 | + | − | − |
TRG_LysEnd_APsAcLL_1 [599–604] TRG_NLS_MonoExtN_4 [597–604] DOC_CYCLIN_1 [601–604] DOC_MAPK_1 [601–609] |
Sorting and directing signal to lysosomal endosomal compartment; NLS; cyclin recognition signal; MAPK docking motif | |
| 626–647 | + | + [632–644] | + [633–642] |
DEG_APCC_DBOX_1 [650–658] DOC_MAPK_1 [649–657] LIG_FHA_1 [633–639] |
APCC binding destruction signal; MAPK docking motif; FHA binding motif | α-1 adrenoceptor |
| 685–687 | + | + [685–687] | + [685–687] | DOC_MAPK_1 [674–684] | MAPK docking motif |
The italic regions are the ID regions where some known interacting partners bind to as indicated in the table. There are several ID regions (non-italic regions), which reveal short linear motifs with no interacting partners yet assigned to them. The first column shows the start and end residues of the ID region. Columns 2–4 are the results of different disorder prediction programs, column 2 is from Pondr VSL1, column 3 is from the IUPred long version, and column 4 is from the IUPred short version. The presence of an ID region in the given predictor is indicated by a + sign, − designates their absence according to the given predictor. ELM is the eukaryotic linear motif, first the names are displayed and in square brackets are the coordinates. Column 5 explains the function of the given ELM, and the last column indicates the overlap with the experimentally known binding partner
Six out of the eight interaction partners with known sites contact TG2 in an ID region of embedded linear motifs. Integrin b1 and b3 contact the N-terminal domain of TG2, which contains a DGR motif. An aspartate residue in the DGR motif has been reported to undergo posttranslational modification to form an iso-DGR motif that mimics a well-known integrin binding RGD motif [168]. Syndecan-4 targets the 208–217 AA ID region, which contains several phosphorylation sites. PKCα serves as a crucial link between TG2-mediated syndecan-4 and β1 integrin co-signaling [48]. PKCα interacts with integrin and facilitates cell adhesion and migration of cells seeded on FN-TG2 in the presence of RGD peptides [169]. TG2 is phosphorylated at AA 216 by PKA, which creates a 14-3-3 protein binding motif [144]. Because the interaction of TG2 and 14-3-3 protein was hypothesized to regulate apoptosis and syndecan-4 was also recently shown to inhibit apoptosis [170], it is possible that PKA-mediated phosphorylation of TG2 contributes to both syndecan and 14-3-3-mediated regulation of apoptosis. The α1-adrenoceptor interacts with the 626–647 AA ID region, which contains a destruction signal and a MAPK docking motif. G-protein-coupled receptors can signal independently of G-proteins and can activate MAP kinases through a family of adaptor proteins called arrestins [171]. MAPK activation by G-proteins can occur through phospholipase C-β, which in turn activates Ras and/or protein kinase C. Because we know that TG2 is involved in Ras and/or protein kinase C signaling, it is possible that TG2 interacts with AR and MAPK and modulates these signaling functions [172].
The N-terminal region of TG2 contains several SUMO binding motifs. SUMO1 was observed to bind to the 358–367 AA region, which also contains several putative phosphorylation sites. All these motifs are displayed in Fig. 2. Other known interaction sites are located in the proximity of the ID regions. For example, fibronectin contacts the 88–106 AA TG2 segment, which is spatially close to the 14–21 AA integrin binding site.
We also aimed to provide a mechanistic interpretation for how short motif-mediated interactions could interfere with TG2 activities. TG2 exists in two conformational states: a GDP-bound closed form and an open form, which is proposed to be the active state. Flexible, dynamic regions can serve as hinges to facilitate the transition to the open form. The conversion between these two conformations exposes a huge surface area (almost 3000 Å2) [173], which facilitates TG2 interacting partners to bind to short linear motifs embedded in these areas. Binding of TG2-interacting partners with SLiMs could further modulate the conformational equilibrium and facilitate formation of the active form of TG2.
Owing to its very large surface area (27,328.8 Å2), TG2 serves as a bridge and scaffold for its non-enzymatic partners. For example, interaction of TG2 with syndecan, fibronectin and integrins, with calmodulin and huntingtin, with SUMO-1 and PIASy, with calreticulin and integrin, with integrin and MFG-E8 and last with RhoA, PKA, AKAP13 and TG2, all require the bridging and scaffolding function of TG2 (Fig. 3). The affinity and lifetime of these contacts could be affected by the neighboring ID regions, which may regulate and govern the surface area exposure. For example, TG2 regulates receptor-mediated signaling by coordinating the α1-adrenoceptor and the PLC δ1 binding partner, whose binding sites are spatially close to each other.
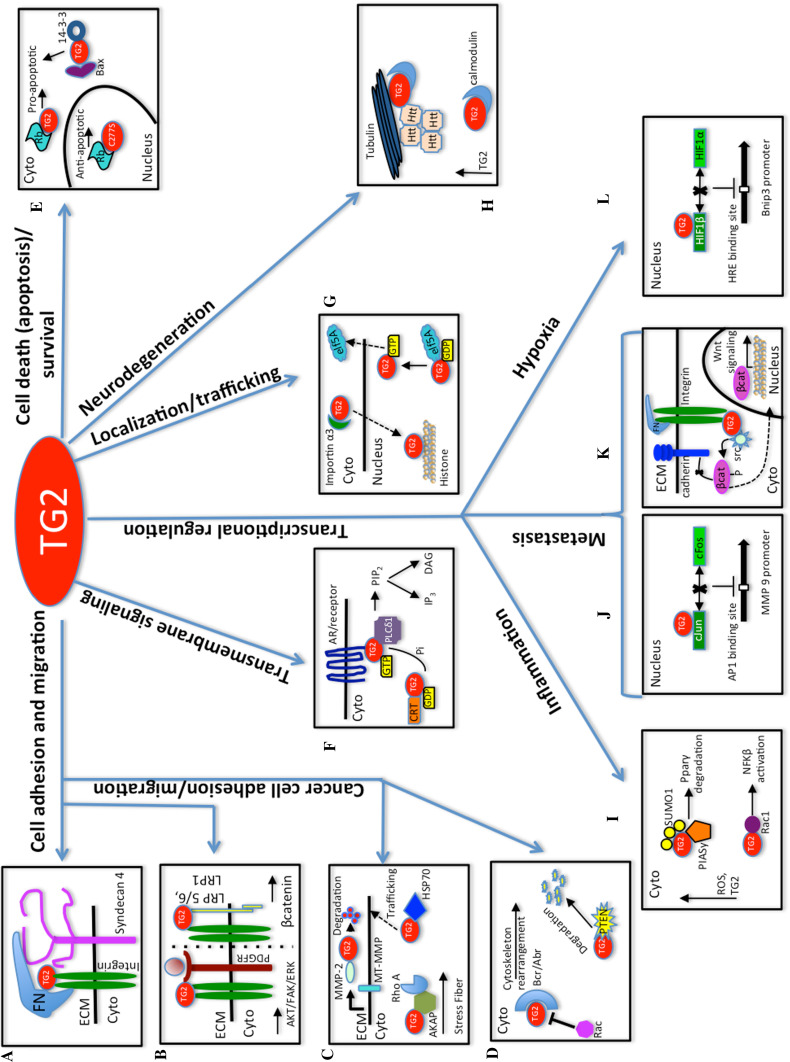
Fig. 3.
Scheme of TG2 and its non-enzymatic partners in regulating TG2 functions. a TG2 facilitates cell adhesion and migration by binding to cell-surface receptors, integrin, and syndecan-4 along with cell matrix protein fibronectin (FN). The interaction takes place in the extracellular matrix (ECM); b TG2 serves as a scaffolding protein by bridging integrin and platelet-derived growth factor receptor (PDGFR), thus participating in amplifying joint PDGFR/integrin signaling via activation of downstream signaling cascades (protein kinase B also called Akt, focal adhesion kinase (FAK), and extracellular signal regulated kinase (ERK); TG2 was shown to form a complex with integrin and low-density lipoprotein receptor related protein (LRP5/6), which in turn activates the β catenin pathway. TG2 was also shown to interact with LRP1, thus participating in LRP1-mediated cell adhesion and migration functions; c membrane metalloprotease (MT MMP) activates soluble metalloprotease (MMP2), which in turn binds to TG2 and mediates its degradation (solid black arrow represents activation), heat shock protein (HSP70) interacts with cytoplasmic (Cyto) TG2 and facilitates its migration to the cell periphery (dotted arrow represents migration of TG2 to the cell periphery), TG2 interacts with protein kinase A anchor protein 13 in the cytoplasm, which in turn interacts with RhoA, thereby potentially facilitating stress fiber formation, cancer cell polarity, and migration; d TG2 interacts with a tumor suppressor protein PTEN (phosphatase and tensin homologue deleted on chromosome 10) and facilitates its ubiquitination and simultaneous degradation by the proteasomal pathway, TG2 modulates Rac (Rho family of small GTPases) activity and its function by blocking the Rac binding site of GTPase-activating proteins such as Bcr/Abr; e TG2 interacts with retinoblastoma protein (Rb) and mediates the pro-apoptotic response when the interaction takes place in the cytoplasm but mediates the anti-apoptotic response when the interaction takes place in the nucleus. Similarly, TG2 mediates the pro-apoptotic response upon interacting with the Bcl2 family member Bax and 14-3-3, a serine/threonine binding protein; f calreticulin modulates TG2 activity by interacting with TG2 and downregulating its TGase and GTPase activities, TG2 facilitates transmembrane signaling by interacting with several transmembrane receptors like adrenoceptor (AR) and along with phospholipase C δ1 (PLCδ1), mediates the release of two second messengers, inositol 4,5 triphosphate (IP3) and diacylglycerol (DAG); g importin α3 interacts with TG2 and mediates its active transport into the nucleus where it can interact with histones, GDP and GTP-bound TG2 regulate the changes in the cellular localization of eukaryotic elongation factor elf5A; h calmodulin interacts with TG2 and increases its TGase activity. Activated TG2 simultaneously binds to tubulin and crosslinks huntingtin (Htt), thus contributing to Huntington’s disease pathology; i intracellular reactive oxygen species (ROS) mediates TG2 SUMOylation by facilitating interaction of TG2 with small ubiquitin-like modifiers 1 (SUMO1) (Fig. 2) and PIASy (a SUMO E3 ligase). ROS-mediated TG2 SUMOylation leads to an increase in TG2 activity, which subsequently affects inflammation by crosslinking and degrading PPARγ (peroxisome proliferator-activator receptor, a negative regulator of inflammation). TG2 was also shown to interact with Rac 1 and regulate inflammation by activating the NF-kβ pathway; j nuclear TG2 was shown to regulate ECM remodeling and metastasis by modulating MMP9 gene transcription. It does so by interacting with cJun, thereby inhibiting dimerization of cJun and cFos and subsequently blocking the AP1 transcription factor binding site on the MMP9 gene; k TG2 regulates metastasis by mediating phosphorylation of β catenin via the bound src. Phosphorylated β catenin dissociates from cadherin and translocates to the nucleus, where it activates Wnt signaling machinery; l TG2 regulates hypoxia by preventing the dimerization of hypoxia-inducible factor 1β and 1α (HIF1β, HIF1α), subsequently inhibiting functional HIF transcription factor binding to hypoxia response elements (HRE) present in the promoter of the Bnip3 gene
Anti-TG2 antibodies, a tool for identifying TG2 functions and their significance in TG2-mediated biological processes
As complexes of TG2 have not been structurally characterized, antibodies and their specific binding location to the TG2 molecule could provide important information regarding the conformational states of TG2 and its interacting proteins. Most techniques employed for detecting protein–protein interactions make use of antibodies and hence specific antibodies are essential for revealing interactions in cells. Because TG2 is present in different conformational states depending upon the physiology of the cellular compartments, it is possible that antibody-binding epitopes are masked, and hence we need to test different antibodies to confirm the experimental data. In this review, we provide an overview of all known TG2-specific antibodies along with their proposed binding sites and their significance (Table 3).
Table 3.
List of anti-TG2 antibodies with their epitopes
| Antibody | Epitope on TG2 | Sources |
|---|---|---|
| 4G3 hybridoma cells | Fibronectin binding site | Santa Cruz Biotechnology |
| G92 | Fibronectin binding site | Trejo Skalli et al. [183] |
| H23 | Ca2+ binding site (433–438) | Korponay-Szabó et al. [184] |
| TG 100 | Core domain (447–538) | Life Technologies |
| CUB7402 | Core domain (447–478) | Abcam |
| A033 | Catalytic core domain | Zedira |
| A034 | Beta sheet domain | Zedira |
| A036 | Beta barrel 1 domain | Zedira |
| A037 | Beta barrel 2 domain | Zedira |
| PAB0062-P | Not known | Covalab |
| PAB0063 | Not known | Covalab |
| MAB0024-P | Not known | Covalab |
| PA5-23219 | 623–645 AA | Thermo Scientific/Pierce |
| EP2957 | Not known | Abcam |
| Ab421 | Against full-length protein | Abcam |
| OAAF01946 | N terminal domain (1–50 AA) | Aviva Systems Biology |
| Celiac anti-TG2 autoantibodies | Glu 153, Glu 154, Arg 19, and Met 659, epitopes are conformational | Simon-Vecsei et al. [159] |
| Celiac anti-TG2 monoclonal autoantibodies | N-terminal domain (overlapping fibronectin binding site), two of the epitopes were conformational | Iversen et al. [185] |
Antibodies generated against TG2 available through commercial sources or in-house antibodies are listed. Their epitopes on TG2 have also been provided. References citing the antibody or suppliers information are also indicated
Antibodies targeting the linear epitope on TG2
Several different monoclonal and polyclonal antibodies against TG2 have been synthesized, targeting different functional domains of TG2. They have been extensively used to improve our understanding of TG2-mediated biological processes. Table 3 summarizes all the antibodies available and their proposed binding sites on TG2. These antibodies target linear epitopes, thus their binding is not influenced by TG2-conformational changes. CUB7402 and TG100 are the most frequently used TG2 antibodies, both targeting the core domain of TG2. 4G3 and G92 bind to the fibronectin binding region of TG2 and hence can contribute to understanding the fibronectin binding property as well as the associated functional aspects of the enzyme (Table 3).
For many years, none of the commercial TG2-specific antibodies or anti-TG2 autoantibodies was able to recognize cell-surface TG2. One of the monoclonal TG2 antibodies, 6B9, was claimed to recognize cell-surface TG2, but later it was determined that it actually recognized CD44 on the cell surface and not TG2 [174]. Recent work from our group showed that in contrast to other commercially available TG2 antibodies, TG100 was the only antibody that recognized cell-surface TG2 on monocyte-derived dendritic cells and macrophages [175], emphasizing the need for testing several antibodies to have a complete overview of the biochemical properties of the enzyme.
Celiac antibodies targeting conformational epitopes on TG2
Autoantibodies generated against TG2, found specifically in celiac disease patients, are reported to mediate celiac pathogenesis and affect several other biological processes. Some of the biological effects include (a) inhibition of cell adhesion and migration, most probably by modulating the heparin sulfate proteoglycan binding site [176] and by enhancing the expression of matrix metallo-proteases, (b) inhibition of epithelial cell differentiation by interfering with TG2 mediated TGF-β1 activation [177], (c) inhibition of several steps in angiogenesis [178], and (d) increasing the permeability of blood–brain barrier.
Serious efforts have recently been made to identify these autoantibodies and map their epitopes on TG2 [179, 180]. One such study from our group revealed that the epitopes of these TG2 antibodies were conformational. They included glutamine 153 and 154 in the first alpha helix of the core domain, and arginine 19 in the first alpha helix of the N-terminal domain [179]. In another study of this kind, TG2-autoantibodies were generated from single plasma cells of celiac disease lesions. Their epitope mapping also revealed that they recognize a few conformational epitopes, which are localized in the N-terminal region (Table 3). Because these antibodies inhibited fibronectin binding, and also the epitopes were inaccessible when TG2 was in the cell surface-bound form, it confirmed that they overlap with the fibronectin binding region of TG2 [180]. Monoclonal mouse antibodies, which overlapped with celiac antibody epitopes, resulted in the release of celiac antibodies from tissues, antagonizing their harmful effects in cell culture experiments. This observation further emphasizes the need for such antibodies to perform functional studies and to explore their therapeutic benefits.
Conclusions and perspectives
Although TG2 was discovered decades ago, this enigmatic protein still intrigues researchers. It is remarkable how a single protein could catalyze such a vast array of biochemical reactions and regulate several physiological and pathological processes. We speculate that the compartment-dependent regulation of the transamidation activity and non-enzymatic interactions unique to TG2 profoundly impact its multiple cell functions. Therefore, they are likely to contribute to a number of pathological states. In this review, we have focused on non-covalent interaction-mediated regulatory functions of TG2. Although we have tried to list only those proteins that interact via non-enzymatic interactions, in most cases sufficient evidence is not available to confirm whether they are true interacting partners or if they could also serve as a substrate of TG2. Much work needs to be done to define the interacting regions and sequences on both TG2 and its respective interacting partner. It also has yet to be revealed whether the interacting proteins bind to the open or closed form of TG2. Answers to these unresolved questions should help us in the future to evaluate the relevance of such interactions on TG2 function in more detail. We have used a rather novel approach to analyze unique structural features, such as differences between the 3D structure of TG2 and the rest of its family members, the presence of IDRs, SLiMs, and the surface area of the closed and open forms of the protein, to determine how these features could regulate TG2 function. Bioinformatic analysis revealed several IDRs along with SLiMs in the TG2 sequence. Some of the interacting protein binding sites overlapped these ID regions. Owing to the importance of these regions in mediating additional protein–protein interaction and its role in regulating diverse protein functions, this review provides a new direction for future TG2 research.
Acknowledgments
This work was supported by the Hungarian Scientific Research Fund (OTKA NK 105046], the New Hungary Development Plan via the TAMOP-4.2.2.A-11/1/KONV-2012-0023 “V´ED ELEM” project, European Union Framework Programme 7 TRANSCOM-IAPP 251506 and TRANSPATH ITN 289964. The support of the Momentum program (LP2012-41) of the Hungarian Academy of Sciences is gratefully acknowledged (M. F.). We thank Dr. Máte Demény for critical reading of this manuscript.
Conflict of interest
The authors declare that they have no conflict of interest.
References
- 1.Lorand L, Graham RM. Transglutaminases: crosslinking enzymes with pleiotropic functions. Nat Rev Mol Cell Biol. 2003;4(2):140–156. doi: 10.1038/nrm1014. [DOI] [PubMed] [Google Scholar]
- 2.Kiraly R, Demeny M, Fesus L. Protein transamidation by transglutaminase 2 in cells: a disputed Ca2+-dependent action of a multifunctional protein. FEBS J. 2011;278(24):4717–4739. doi: 10.1111/j.1742-4658.2011.08345.x. [DOI] [PubMed] [Google Scholar]
- 3.Griffin M, Casadio R, Bergamini CM. Transglutaminases: nature’s biological glues. Biochem J. 2002;368(Pt 2):377–396. doi: 10.1042/BJ20021234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Belkin AM. Extracellular TG2: emerging functions and regulation. FEBS J. 2011;278(24):4704–4716. doi: 10.1111/j.1742-4658.2011.08346.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Nakaoka H, Perez DM, Baek KJ, Das T, Husain A, Misono K, Im MJ, Graham RM. Gh: a GTP-binding protein with transglutaminase activity and receptor signaling function. Science. 1994;264(5165):1593–1596. doi: 10.1126/science.7911253. [DOI] [PubMed] [Google Scholar]
- 6.Vezza R, Habib A, FitzGerald GA. Differential signaling by the thromboxane receptor isoforms via the novel GTP-binding protein, Gh. J Biol Chem. 1999;274(18):12774–12779. doi: 10.1074/jbc.274.18.12774. [DOI] [PubMed] [Google Scholar]
- 7.Baek KJ, Kwon NS, Lee HS, Kim MS, Muralidhar P, Im MJ. Oxytocin receptor couples to the 80-kDa Gh alpha family protein in human myometrium. Biochem J. 1996;315(Pt 3):739–744. [PMC free article] [PubMed] [Google Scholar]
- 8.Hasegawa G, Suwa M, Ichikawa Y, Ohtsuka T, Kumagai S, Kikuchi M, Sato Y, Saito Y. A novel function of tissue-type transglutaminase: protein disulphide isomerase. Biochem J. 2003;373(Pt 3):793–803. doi: 10.1042/BJ20021084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mastroberardino PG, Farrace MG, Viti I, Pavone F, Fimia GM, Melino G, Rodolfo C, Piacentini M. “Tissue” transglutaminase contributes to the formation of disulphide bridges in proteins of mitochondrial respiratory complexes. Biochim Biophys Acta. 2006;1757(9–10):1357–1365. doi: 10.1016/j.bbabio.2006.07.007. [DOI] [PubMed] [Google Scholar]
- 10.Mishra S, Murphy LJ. Tissue transglutaminase has intrinsic kinase activity: identification of transglutaminase 2 as an insulin-like growth factor-binding protein-3 kinase. J Biol Chem. 2004;279(23):23863–23868. doi: 10.1074/jbc.M311919200. [DOI] [PubMed] [Google Scholar]
- 11.Mishra S, Saleh A, Espino PS, Davie JR, Murphy LJ. Phosphorylation of histones by tissue transglutaminase. J Biol Chem. 2006;281(9):5532–5538. doi: 10.1074/jbc.M506864200. [DOI] [PubMed] [Google Scholar]
- 12.Csosz E, Mesko B, Fesus L. Transdab wiki: the interactive transglutaminase substrate database on web 2.0 surface. Amino Acids. 2009;36(4):615–617. doi: 10.1007/s00726-008-0121-y. [DOI] [PubMed] [Google Scholar]
- 13.Lesort M, Attanavanich K, Zhang J, Johnson GV. Distinct nuclear localization and activity of tissue transglutaminase. J Biol Chem. 1998;273(20):11991–11994. doi: 10.1074/jbc.273.20.11991. [DOI] [PubMed] [Google Scholar]
- 14.Zemskov EA, Mikhailenko I, Strickland DK, Belkin AM. Cell-surface transglutaminase undergoes internalization and lysosomal degradation: an essential role for LRP1. J Cell Sci. 2007;120(Pt 18):3188–3199. doi: 10.1242/jcs.010397. [DOI] [PubMed] [Google Scholar]
- 15.Milakovic T, Tucholski J, McCoy E, Johnson GV. Intracellular localization and activity state of tissue transglutaminase differentially impacts cell death. J Biol Chem. 2004;279(10):8715–8722. doi: 10.1074/jbc.M308479200. [DOI] [PubMed] [Google Scholar]
- 16.Aeschlimann D, Thomazy V. Protein crosslinking in assembly and remodelling of extracellular matrices: the role of transglutaminases. Connect Tissue Res. 2000;41(1):1–27. doi: 10.3109/03008200009005638. [DOI] [PubMed] [Google Scholar]
- 17.Scarpellini A, Germack R, Lortat-Jacob H, Muramatsu T, Billett E, Johnson T, Verderio EA. Heparan sulfate proteoglycans are receptors for the cell-surface trafficking and biological activity of transglutaminase-2. J Biol Chem. 2009;284(27):18411–18423. doi: 10.1074/jbc.M109.012948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Gaudry CA, Verderio E, Aeschlimann D, Cox A, Smith C, Griffin M. Cell surface localization of tissue transglutaminase is dependent on a fibronectin-binding site in its N-terminal beta-sandwich domain. J Biol Chem. 1999;274(43):30707–30714. doi: 10.1074/jbc.274.43.30707. [DOI] [PubMed] [Google Scholar]
- 19.Balklava Z, Verderio E, Collighan R, Gross S, Adams J, Griffin M. Analysis of tissue transglutaminase function in the migration of Swiss 3T3 fibroblasts: the active-state conformation of the enzyme does not affect cell motility but is important for its secretion. J Biol Chem. 2002;277(19):16567–16575. doi: 10.1074/jbc.M109836200. [DOI] [PubMed] [Google Scholar]
- 20.Nurminskaya MV, Belkin AM. Cellular functions of tissue transglutaminase. International review of cell and molecular biology. 2012;294:1–97. doi: 10.1016/B978-0-12-394305-7.00001-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Akimov SS, Belkin AM. Cell-surface transglutaminase promotes fibronectin assembly via interaction with the gelatin-binding domain of fibronectin: a role in TGFbeta-dependent matrix deposition. J Cell Sci. 2001;114(Pt 16):2989–3000. doi: 10.1242/jcs.114.16.2989. [DOI] [PubMed] [Google Scholar]
- 22.Lortat-Jacob H, Burhan I, Scarpellini A, Thomas A, Imberty A, Vives RR, Johnson T, Gutierrez A, Verderio EA. Transglutaminase-2 interaction with heparin: identification of a heparin binding site that regulates cell adhesion to fibronectin-transglutaminase-2 matrix. J Biol Chem. 2012;287(22):18005–18017. doi: 10.1074/jbc.M111.337089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Akimov SS, Krylov D, Fleischman LF, Belkin AM. Tissue transglutaminase is an integrin-binding adhesion coreceptor for fibronectin. J Cell Biol. 2000;148(4):825–838. doi: 10.1083/jcb.148.4.825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Stamnaes J, Pinkas DM, Fleckenstein B, Khosla C, Sollid LM. Redox regulation of transglutaminase 2 activity. J Biol Chem. 2010;285(33):25402–25409. doi: 10.1074/jbc.M109.097162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lorand L, Dailey JE, Turner PM. Fibronectin as a carrier for the transglutaminase from human erythrocytes. Proc Natl Acad Sci USA. 1988;85(4):1057–1059. doi: 10.1073/pnas.85.4.1057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Turner PM, Lorand L. Complexation of fibronectin with tissue transglutaminase. Biochemistry. 1989;28(2):628–635. doi: 10.1021/bi00428a032. [DOI] [PubMed] [Google Scholar]
- 27.Radek JT, Jeong JM, Murthy SN, Ingham KC, Lorand L. Affinity of human erythrocyte transglutaminase for a 42-kDa gelatin-binding fragment of human plasma fibronectin. Proc Natl Acad Sci USA. 1993;90(8):3152–3156. doi: 10.1073/pnas.90.8.3152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hang J, Zemskov EA, Lorand L, Belkin AM. Identification of a novel recognition sequence for fibronectin within the NH2-terminal beta-sandwich domain of tissue transglutaminase. J Biol Chem. 2005;280(25):23675–23683. doi: 10.1074/jbc.M503323200. [DOI] [PubMed] [Google Scholar]
- 29.Forsprecher J, Wang Z, Nelea V, Kaartinen MT. Enhanced osteoblast adhesion on transglutaminase 2-crosslinked fibronectin. Amino Acids. 2009;36(4):747–753. doi: 10.1007/s00726-008-0125-7. [DOI] [PubMed] [Google Scholar]
- 30.Haroon ZA, Hettasch JM, Lai TS, Dewhirst MW, Greenberg CS. Tissue transglutaminase is expressed, active, and directly involved in rat dermal wound healing and angiogenesis. FASEB J Off Publ Fed Am Soc Exp Biol. 1999;13(13):1787–1795. doi: 10.1096/fasebj.13.13.1787. [DOI] [PubMed] [Google Scholar]
- 31.Martinez J, Chalupowicz DG, Roush RK, Sheth A, Barsigian C. Transglutaminase-mediated processing of fibronectin by endothelial cell monolayers. Biochemistry. 1994;33(9):2538–2545. doi: 10.1021/bi00175a024. [DOI] [PubMed] [Google Scholar]
- 32.Kleman JP, Aeschlimann D, Paulsson M, van der Rest M. Transglutaminase-catalyzed cross-linking of fibrils of collagen V/XI in A204 rhabdomyosarcoma cells. Biochemistry. 1995;34(42):13768–13775. doi: 10.1021/bi00042a007. [DOI] [PubMed] [Google Scholar]
- 33.Kaartinen MT, Pirhonen A, Linnala-Kankkunen A, Maenpaa PH. Transglutaminase-catalyzed cross-linking of osteopontin is inhibited by osteocalcin. J Biol Chem. 1997;272(36):22736–22741. doi: 10.1074/jbc.272.36.22736. [DOI] [PubMed] [Google Scholar]
- 34.Akimov SS, Belkin AM. Cell surface tissue transglutaminase is involved in adhesion and migration of monocytic cells on fibronectin. Blood. 2001;98(5):1567–1576. doi: 10.1182/blood.v98.5.1567. [DOI] [PubMed] [Google Scholar]
- 35.Takada Y, Ye X, Simon S. The integrins. Genome Biol. 2007;8(5):215. doi: 10.1186/gb-2007-8-5-215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.D’Souza SE, Ginsberg MH, Plow EF. Arginine–glycine–aspartic acid (RGD): a cell adhesion motif. Trends Biochem Sci. 1991;16(7):246–250. doi: 10.1016/0968-0004(91)90096-e. [DOI] [PubMed] [Google Scholar]
- 37.Verderio E, Nicholas B, Gross S, Griffin M. Regulated expression of tissue transglutaminase in Swiss 3T3 fibroblasts: effects on the processing of fibronectin, cell attachment, and cell death. Exp Cell Res. 1998;239(1):119–138. doi: 10.1006/excr.1997.3874. [DOI] [PubMed] [Google Scholar]
- 38.Toth B, Garabuczi E, Sarang Z, Vereb G, Vamosi G, Aeschlimann D, Blasko B, Becsi B, Erdodi F, Lacy-Hulbert A, Zhang A, Falasca L, Birge RB, Balajthy Z, Melino G, Fesus L, Szondy Z. Transglutaminase 2 is needed for the formation of an efficient phagocyte portal in macrophages engulfing apoptotic cells. J Immunol. 2009;182(4):2084–2092. doi: 10.4049/jimmunol.0803444. [DOI] [PubMed] [Google Scholar]
- 39.Jones RA, Nicholas B, Mian S, Davies PJ, Griffin M. Reduced expression of tissue transglutaminase in a human endothelial cell line leads to changes in cell spreading, cell adhesion and reduced polymerisation of fibronectin. J Cell Sci. 1997;110(Pt 19):2461–2472. doi: 10.1242/jcs.110.19.2461. [DOI] [PubMed] [Google Scholar]
- 40.Zemskov EA, Janiak A, Hang J, Waghray A, Belkin AM. The role of tissue transglutaminase in cell–matrix interactions. Front Biosci J Virtual Libr. 2006;11:1057–1076. doi: 10.2741/1863. [DOI] [PubMed] [Google Scholar]
- 41.Leiss M, Beckmann K, Giros A, Costell M, Fassler R. The role of integrin binding sites in fibronectin matrix assembly in vivo. Curr Opin Cell Biol. 2008;20(5):502–507. doi: 10.1016/j.ceb.2008.06.001. [DOI] [PubMed] [Google Scholar]
- 42.Signorini M, Bortolotti F, Poltronieri L, Bergamini CM. Human erythrocyte transglutaminase: purification and preliminary characterisation. Biol Chem Hoppe Seyler. 1988;369(4):275–281. doi: 10.1515/bchm3.1988.369.1.275. [DOI] [PubMed] [Google Scholar]
- 43.Wang Z, Collighan RJ, Pytel K, Rathbone DL, Li X, Griffin M. Characterization of heparin-binding site of tissue transglutaminase: its importance in cell surface targeting, matrix deposition, and cell signaling. J Biol Chem. 2012;287(16):13063–13083. doi: 10.1074/jbc.M111.294819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Huntington JA. Mechanisms of glycosaminoglycan activation of the serpins in hemostasis. J Thromb Haemost JTH. 2003;1(7):1535–1549. doi: 10.1046/j.1538-7836.2003.00305.x. [DOI] [PubMed] [Google Scholar]
- 45.Bishop JR, Schuksz M, Esko JD. Heparan sulphate proteoglycans fine-tune mammalian physiology. Nature. 2007;446(7139):1030–1037. doi: 10.1038/nature05817. [DOI] [PubMed] [Google Scholar]
- 46.Verderio E, Scarpellini A. Significance of the syndecan-4-transglutaminase-2 interaction. Sci World J. 2010;10:1073–1077. doi: 10.1100/tsw.2010.102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Bishop JR, Schuksz M, Esko JD. Heparan sulphate proteoglycans fine-tune mammalian physiology. Nature. 2007;446:1030–1037. doi: 10.1038/nature05817. [DOI] [PubMed] [Google Scholar]
- 48.Telci D, Wang Z, Li X, Verderio EA, Humphries MJ, Baccarini M, Basaga H, Griffin M. Fibronectin-tissue transglutaminase matrix rescues RGD-impaired cell adhesion through syndecan-4 and beta1 integrin co-signaling. J Biol Chem. 2008;283(30):20937–20947. doi: 10.1074/jbc.M801763200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Dardik R, Inbal A. Complex formation between tissue transglutaminase II (tTG) and vascular endothelial growth factor receptor 2 (VEGFR-2): proposed mechanism for modulation of endothelial cell response to VEGF. Exp Cell Res. 2006;312(16):2973–2982. doi: 10.1016/j.yexcr.2006.05.019. [DOI] [PubMed] [Google Scholar]
- 50.Zemskov EA, Loukinova E, Mikhailenko I, Coleman RA, Strickland DK, Belkin AM. Regulation of platelet-derived growth factor receptor function by integrin-associated cell surface transglutaminase. J Biol Chem. 2009;284(24):16693–16703. doi: 10.1074/jbc.M109.010769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Faverman L, Mikhaylova L, Malmquist J, Nurminskaya M. Extracellular transglutaminase 2 activates beta-catenin signaling in calcifying vascular smooth muscle cells. FEBS Lett. 2008;582(10):1552–1557. doi: 10.1016/j.febslet.2008.03.053. [DOI] [PubMed] [Google Scholar]
- 52.Herz J, Strickland DK. LRP: a multifunctional scavenger and signaling receptor. J Clin Investig. 2001;108(6):779–784. doi: 10.1172/JCI13992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Salicioni AM, Gaultier A, Brownlee C, Cheezum MK, Gonias SL. Low-density lipoprotein receptor-related protein-1 promotes beta1 integrin maturation and transport to the cell surface. J Biol Chem. 2004;279(11):10005–10012. doi: 10.1074/jbc.M306625200. [DOI] [PubMed] [Google Scholar]
- 54.Salicioni AM, Mizelle KS, Loukinova E, Mikhailenko I, Strickland DK, Gonias SL. The low-density lipoprotein receptor-related protein mediates fibronectin catabolism and inhibits fibronectin accumulation on cell surfaces. J Biol Chem. 2002;277(18):16160–16166. doi: 10.1074/jbc.M201401200. [DOI] [PubMed] [Google Scholar]
- 55.Nagase H, Woessner JF., Jr Matrix metalloproteinases. J Biol Chem. 1999;274(31):21491–21494. doi: 10.1074/jbc.274.31.21491. [DOI] [PubMed] [Google Scholar]
- 56.Belkin AM, Akimov SS, Zaritskaya LS, Ratnikov BI, Deryugina EI, Strongin AY. Matrix-dependent proteolysis of surface transglutaminase by membrane-type metalloproteinase regulates cancer cell adhesion and locomotion. J Biol Chem. 2001;276(21):18415–18422. doi: 10.1074/jbc.M010135200. [DOI] [PubMed] [Google Scholar]
- 57.Strongin AY, Collier I, Bannikov G, Marmer BL, Grant GA, Goldberg GI. Mechanism of cell surface activation of 72-kDa type IV collagenase. Isolation of the activated form of the membrane metalloprotease. J Biol Chem. 1995;270(10):5331–5338. doi: 10.1074/jbc.270.10.5331. [DOI] [PubMed] [Google Scholar]
- 58.Murphy G, Stanton H, Cowell S, Butler G, Knauper V, Atkinson S, Gavrilovic J. Mechanisms for pro matrix metalloproteinase activation. APMIS Acta Pathol Microbiol Immunol Scand. 1999;107(1):38–44. doi: 10.1111/j.1699-0463.1999.tb01524.x. [DOI] [PubMed] [Google Scholar]
- 59.Belkin AM, Zemskov EA, Hang J, Akimov SS, Sikora S, Strongin AY. Cell-surface-associated tissue transglutaminase is a target of MMP-2 proteolysis. Biochemistry. 2004;43(37):11760–11769. doi: 10.1021/bi049266z. [DOI] [PubMed] [Google Scholar]
- 60.Birckbichler PJ, Bonner RB, Hurst RE, Bane BL, Pitha JV, Hemstreet GP., 3rd Loss of tissue transglutaminase as a biomarker for prostate adenocarcinoma. Cancer. 2000;89(2):412–423. doi: 10.1002/1097-0142(20000715)89:2<412::aid-cncr29>3.0.co;2-o. [DOI] [PubMed] [Google Scholar]
- 61.Lewis TE, Milam TD, Klingler DW, Rao PS, Jaggi M, Smith DJ, Hemstreet GP, Balaji KC. Tissue transglutaminase interacts with protein kinase A anchor protein 13 in prostate cancer. Urol Oncol. 2005;23(6):407–412. doi: 10.1016/j.urolonc.2005.04.002. [DOI] [PubMed] [Google Scholar]
- 62.Edwards AS, Scott JD. A-kinase anchoring proteins: protein kinase A and beyond. Curr Opin Cell Biol. 2000;12(2):217–221. doi: 10.1016/s0955-0674(99)00085-x. [DOI] [PubMed] [Google Scholar]
- 63.Michel JJ, Scott JD. AKAP-mediated signal transduction. Annu Rev Pharmacol Toxicol. 2002;42:235–257. doi: 10.1146/annurev.pharmtox.42.083101.135801. [DOI] [PubMed] [Google Scholar]
- 64.Nishikawa K, Toker A, Johannes FJ, Songyang Z, Cantley LC. Determination of the specific substrate sequence motifs of protein kinase C isozymes. J Biol Chem. 1997;272(2):952–960. doi: 10.1074/jbc.272.2.952. [DOI] [PubMed] [Google Scholar]
- 65.Songyang Z, Lu KP, Kwon YT, Tsai LH, Filhol O, Cochet C, Brickey DA, Soderling TR, Bartleson C, Graves DJ, DeMaggio AJ, Hoekstra MF, Blenis J, Hunter T, Cantley LC. A structural basis for substrate specificities of protein Ser/Thr kinases: primary sequence preference of casein kinases I and II, NIMA, phosphorylase kinase, calmodulin-dependent kinase II, CDK5, and Erk1. Mol Cell Biol. 1996;16(11):6486–6493. doi: 10.1128/mcb.16.11.6486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Blom N, Gammeltoft S, Brunak S. Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J Mol Biol. 1999;294(5):1351–1362. doi: 10.1006/jmbi.1999.3310. [DOI] [PubMed] [Google Scholar]
- 67.Wang Y, Ande SR, Mishra S. Phosphorylation of transglutaminase 2 (TG2) at serine-216 plays a role in TG2-mediated activation of nuclear factor-kappa B and in the downregulation of PTEN. BMC Cancer. 2012;12(1):277. doi: 10.1186/1471-2407-12-277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.McConnachie G, Langeberg LK, Scott JD. AKAP signaling complexes: getting to the heart of the matter. Trends Mol Med. 2006;12(7):317–323. doi: 10.1016/j.molmed.2006.05.008. [DOI] [PubMed] [Google Scholar]
- 69.Hodge JC, Bub J, Kaul S, Kajdacsy-Balla A, Lindholm PF. Requirement of RhoA activity for increased nuclear factor kappaB activity and PC-3 human prostate cancer cell invasion. Cancer Res. 2003;63(6):1359–1364. [PubMed] [Google Scholar]
- 70.Singh US, Pan J, Kao YL, Joshi S, Young KL, Baker KM. Tissue transglutaminase mediates activation of RhoA and MAP kinase pathways during retinoic acid-induced neuronal differentiation of SH-SY5Y cells. J Biol Chem. 2003;278(1):391–399. doi: 10.1074/jbc.M206361200. [DOI] [PubMed] [Google Scholar]
- 71.Diviani D, Soderling J, Scott JD. AKAP-Lbc anchors protein kinase A and nucleates Galpha 12-selective Rho-mediated stress fiber formation. J Biol Chem. 2001;276(47):44247–44257. doi: 10.1074/jbc.M106629200. [DOI] [PubMed] [Google Scholar]
- 72.Bakin AV, Safina A, Rinehart C, Daroqui C, Darbary H, Helfman DM. A critical role of tropomyosins in TGF-beta regulation of the actin cytoskeleton and cell motility in epithelial cells. Mol Biol Cell. 2004;15(10):4682–4694. doi: 10.1091/mbc.E04-04-0353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Birukova AA, Smurova K, Birukov KG, Usatyuk P, Liu F, Kaibuchi K, Ricks-Cord A, Natarajan V, Alieva I, Garcia JG, Verin AD. Microtubule disassembly induces cytoskeletal remodeling and lung vascular barrier dysfunction: role of Rho-dependent mechanisms. J Cell Physiol. 2004;201(1):55–70. doi: 10.1002/jcp.20055. [DOI] [PubMed] [Google Scholar]
- 74.Verma A, Wang H, Manavathi B, Fok JY, Mann AP, Kumar R, Mehta K. Increased expression of tissue transglutaminase in pancreatic ductal adenocarcinoma and its implications in drug resistance and metastasis. Cancer Res. 2006;66(21):10525–10533. doi: 10.1158/0008-5472.CAN-06-2387. [DOI] [PubMed] [Google Scholar]
- 75.Mangala LS, Fok JY, Zorrilla-Calancha IR, Verma A, Mehta K. Tissue transglutaminase expression promotes cell attachment, invasion and survival in breast cancer cells. Oncogene. 2007;26(17):2459–2470. doi: 10.1038/sj.onc.1210035. [DOI] [PubMed] [Google Scholar]
- 76.Mehta K, Fok JY, Mangala LS. Tissue transglutaminase: from biological glue to cell survival cues. Front Biosci J Virtual Libr. 2006;11:173–185. doi: 10.2741/1789. [DOI] [PubMed] [Google Scholar]
- 77.Mann AP, Verma A, Sethi G, Manavathi B, Wang H, Fok JY, Kunnumakkara AB, Kumar R, Aggarwal BB, Mehta K. Overexpression of tissue transglutaminase leads to constitutive activation of nuclear factor-kappaB in cancer cells: delineation of a novel pathway. Cancer Res. 2006;66(17):8788–8795. doi: 10.1158/0008-5472.CAN-06-1457. [DOI] [PubMed] [Google Scholar]
- 78.Tamura M, Gu J, Matsumoto K, Aota S, Parsons R, Yamada KM. Inhibition of cell migration, spreading, and focal adhesions by tumor suppressor PTEN. Science. 1998;280(5369):1614–1617. doi: 10.1126/science.280.5369.1614. [DOI] [PubMed] [Google Scholar]
- 79.Maehama T, Dixon JE. The tumor suppressor, PTEN/MMAC1, dephosphorylates the lipid second messenger, phosphatidylinositol 3,4,5-trisphosphate. J Biol Chem. 1998;273(22):13375–13378. doi: 10.1074/jbc.273.22.13375. [DOI] [PubMed] [Google Scholar]
- 80.Verma A, Guha S, Wang H, Fok JY, Koul D, Abbruzzese J, Mehta K. Tissue transglutaminase regulates focal adhesion kinase/AKT activation by modulating PTEN expression in pancreatic cancer cells. Clin Cancer Res. 2008;14(7):1997–2005. doi: 10.1158/1078-0432.CCR-07-1533. [DOI] [PubMed] [Google Scholar]
- 81.Boroughs LK, Antonyak MA, Johnson JL, Cerione RA. A unique role for heat shock protein 70 and its binding partner tissue transglutaminase in cancer cell migration. J Biol Chem. 2011;286(43):37094–37107. doi: 10.1074/jbc.M111.242438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Yi SJ, Groffen J, Heisterkamp N. Transglutaminase 2 regulates the GTPase-activating activity of Bcr. J Biol Chem. 2009;284(51):35645–35651. doi: 10.1074/jbc.M109.062240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Yi SJ, Groffen J, Heisterkamp N. Bcr is a substrate for transglutaminase 2 cross-linking activity. BMC Biochem. 2011;12:8. doi: 10.1186/1471-2091-12-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Folkman J. Antiangiogenesis in cancer therapy—endostatin and its mechanisms of action. Exp Cell Res. 2006;312(5):594–607. doi: 10.1016/j.yexcr.2005.11.015. [DOI] [PubMed] [Google Scholar]
- 85.Xu L, Begum S, Hearn JD, Hynes RO. GPR56, an atypical G protein-coupled receptor, binds tissue transglutaminase, TG2, and inhibits melanoma tumor growth and metastasis. Proc Natl Acad Sci USA. 2006;103(24):9023–9028. doi: 10.1073/pnas.0602681103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Faye C, Inforzato A, Bignon M, Hartmann DJ, Muller L, Ballut L, Olsen BR, Day AJ, Ricard-Blum S. Transglutaminase-2: a new endostatin partner in the extracellular matrix of endothelial cells. Biochem J. 2010;427(3):467–475. doi: 10.1042/BJ20091594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Yang L, Friedland S, Corson N, Xu L. GPR56 inhibits melanoma growth by internalizing and degrading its ligand TG2. Cancer Res. 2014;74(4):1022–1031. doi: 10.1158/0008-5472.CAN-13-1268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.L’Heureux DZ, Rothman VL, Tuszynski GP. The interaction of angiocidin with tissue transglutaminase. Exp Mol Pathol. 2010;88(1):15–25. doi: 10.1016/j.yexmp.2009.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Feng JF, Gray CD, Im MJ. Alpha 1B-adrenoceptor interacts with multiple sites of transglutaminase II: characteristics of the interaction in binding and activation. Biochemistry. 1999;38(7):2224–2232. doi: 10.1021/bi9823176. [DOI] [PubMed] [Google Scholar]
- 90.Chen S, Lin F, Iismaa S, Lee KN, Birckbichler PJ, Graham RM. Alpha1-adrenergic receptor signaling via Gh is subtype specific and independent of its transglutaminase activity. J Biol Chem. 1996;271(50):32385–32391. doi: 10.1074/jbc.271.50.32385. [DOI] [PubMed] [Google Scholar]
- 91.Im MJ, Russell MA, Feng JF. Transglutaminase II: a new class of GTP-binding protein with new biological functions. Cell Signal. 1997;9(7):477–482. doi: 10.1016/s0898-6568(97)00049-1. [DOI] [PubMed] [Google Scholar]
- 92.Feng JF, Rhee SG, Im MJ. Evidence that phospholipase delta1 is the effector in the Gh (transglutaminase II)-mediated signaling. J Biol Chem. 1996;271(28):16451–16454. doi: 10.1074/jbc.271.28.16451. [DOI] [PubMed] [Google Scholar]
- 93.Baek KJ, Das T, Gray C, Antar S, Murugesan G, Im MJ. Evidence that the Gh protein is a signal mediator from alpha 1-adrenoceptor to a phospholipase C. I. Identification of alpha 1-adrenoceptor-coupled Gh family and purification of Gh7 from bovine heart. J Biol Chem. 1993;268(36):27390–27397. [PubMed] [Google Scholar]
- 94.Hwang KC, Gray CD, Sivasubramanian N, Im MJ. Interaction site of GTP binding Gh (transglutaminase II) with phospholipase C. J Biol Chem. 1995;270(45):27058–27062. doi: 10.1074/jbc.270.45.27058. [DOI] [PubMed] [Google Scholar]
- 95.Kang SK, Kim DK, Damron DS, Baek KJ, Im MJ. Modulation of intracellular Ca(2+) via alpha(1B)-adrenoreceptor signaling molecules, G alpha(h) (transglutaminase II) and phospholipase C-delta 1. Biochem Biophys Res Commun. 2002;293(1):383–390. doi: 10.1016/S0006-291X(02)00197-3. [DOI] [PubMed] [Google Scholar]
- 96.Park ES, Won JH, Han KJ, Suh PG, Ryu SH, Lee HS, Yun HY, Kwon NS, Baek KJ. Phospholipase C-delta1 and oxytocin receptor signalling: evidence of its role as an effector. Biochem J. 1998;331(Pt 1):283–289. doi: 10.1042/bj3310283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Feng JF, Readon M, Yadav SP, Im MJ. Calreticulin down-regulates both GTP binding and transglutaminase activities of transglutaminase II. Biochemistry. 1999;38(33):10743–10749. doi: 10.1021/bi9905009. [DOI] [PubMed] [Google Scholar]
- 98.Baek KJ, Das T, Gray CD, Desai S, Hwang KC, Gacchui R, Ludwig M, Im MJ. A 50-kDa protein modulates guanine nucleotide binding of transglutaminase II. Biochemistry. 1996;35(8):2651–2657. doi: 10.1021/bi9522965. [DOI] [PubMed] [Google Scholar]
- 99.Coppolino MG, Woodside MJ, Demaurex N, Grinstein S, St-Arnaud R, Dedhar S. Calreticulin is essential for integrin-mediated calcium signalling and cell adhesion. Nature. 1997;386(6627):843–847. doi: 10.1038/386843a0. [DOI] [PubMed] [Google Scholar]
- 100.Dedhar S. Novel functions for calreticulin: interaction with integrins and modulation of gene expression? Trends Biochem Sci. 1994;19(7):269–271. doi: 10.1016/0968-0004(94)90001-9. [DOI] [PubMed] [Google Scholar]
- 101.Dedhar S, Rennie PS, Shago M, Hagesteijn CY, Yang H, Filmus J, Hawley RG, Bruchovsky N, Cheng H, Matusik RJ, et al. Inhibition of nuclear hormone receptor activity by calreticulin. Nature. 1994;367(6462):480–483. doi: 10.1038/367480a0. [DOI] [PubMed] [Google Scholar]
- 102.Takeuchi Y, Ohashi H, Birckbichler PJ, Ikejima T. Nuclear translocation of tissue type transglutaminase during sphingosine-induced cell death: a novel aspect of the enzyme with DNA hydrolytic activity. Z Naturforsch C. 1998;53(5–6):352–358. doi: 10.1515/znc-1998-5-609. [DOI] [PubMed] [Google Scholar]
- 103.Campisi A, Caccamo D, Raciti G, Cannavo G, Macaione V, Curro M, Macaione S, Vanella A, Ientile R. Glutamate-induced increases in transglutaminase activity in primary cultures of astroglial cells. Brain Res. 2003;978(1–2):24–30. doi: 10.1016/s0006-8993(03)02725-2. [DOI] [PubMed] [Google Scholar]
- 104.Balajthy Z, Csomos K, Vamosi G, Szanto A, Lanotte M, Fesus L. Tissue-transglutaminase contributes to neutrophil granulocyte differentiation and functions. Blood. 2006;108(6):2045–2054. doi: 10.1182/blood-2004-02-007948. [DOI] [PubMed] [Google Scholar]
- 105.Kuo TF, Tatsukawa H, Kojima S. New insights into the functions and localization of nuclear transglutaminase 2. FEBS J. 2011;278(24):4756–4767. doi: 10.1111/j.1742-4658.2011.08409.x. [DOI] [PubMed] [Google Scholar]
- 106.Filiano AJ, Bailey CD, Tucholski J, Gundemir S, Johnson GV. Transglutaminase 2 protects against ischemic insult, interacts with HIF1beta, and attenuates HIF1 signaling. FASEB J. 2008;22(8):2662–2675. doi: 10.1096/fj.07-097709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Oliverio S, Amendola A, Di Sano F, Farrace MG, Fesus L, Nemes Z, Piredda L, Spinedi A, Piacentini M. Tissue transglutaminase-dependent posttranslational modification of the retinoblastoma gene product in promonocytic cells undergoing apoptosis. Mol Cell Biol. 1997;17(10):6040–6048. doi: 10.1128/mcb.17.10.6040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Luciani A, Villella VR, Vasaturo A, Giardino I, Raia V, Pettoello-Mantovani M, D’Apolito M, Guido S, Leal T, Quaratino S, Maiuri L. SUMOylation of tissue transglutaminase as link between oxidative stress and inflammation. J Immunol. 2009;183(4):2775–2784. doi: 10.4049/jimmunol.0900993. [DOI] [PubMed] [Google Scholar]
- 109.Peng X, Zhang Y, Zhang H, Graner S, Williams JF, Levitt ML, Lokshin A. Interaction of tissue transglutaminase with nuclear transport protein importin-alpha3. FEBS Lett. 1999;446(1):35–39. doi: 10.1016/s0014-5793(99)00018-6. [DOI] [PubMed] [Google Scholar]
- 110.Singh US, Li Q, Cerione R. Identification of the eukaryotic initiation factor 5A as a retinoic acid-stimulated cellular binding partner for tissue transglutaminase II. J Biol Chem. 1998;273(4):1946–1950. doi: 10.1074/jbc.273.4.1946. [DOI] [PubMed] [Google Scholar]
- 111.Bernhard EJ, Gruber SB, Muschel RJ. Direct evidence linking expression of matrix metalloproteinase 9 (92-kDa gelatinase/collagenase) to the metastatic phenotype in transformed rat embryo cells. Proc Natl Acad Sci USA. 1994;91(10):4293–4297. doi: 10.1073/pnas.91.10.4293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Ahn JS, Kim MK, Hahn JH, Park JH, Park KH, Cho BR, Park SB, Kim DJ. Tissue transglutaminase-induced down-regulation of matrix metalloproteinase-9. Biochem Biophys Res Commun. 2008;376(4):743–747. doi: 10.1016/j.bbrc.2008.09.048. [DOI] [PubMed] [Google Scholar]
- 113.Condello S, Cao L, Matei D. Tissue transglutaminase regulates beta-catenin signaling through a c-Src-dependent mechanism. FASEB J Off Publ Fed Am Soc Exp Biol. 2013;27(8):3100–3112. doi: 10.1096/fj.12-222620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Tolentino PJ, Waghray A, Wang KK, Hayes RL. Increased expression of tissue-type transglutaminase following middle cerebral artery occlusion in rats. J Neurochem. 2004;89(5):1301–1307. doi: 10.1111/j.1471-4159.2004.02436.x. [DOI] [PubMed] [Google Scholar]
- 115.Ientile R, Caccamo D, Marciano MC, Curro M, Mannucci C, Campisi A, Calapai G. Transglutaminase activity and transglutaminase mRNA transcripts in gerbil brain ischemia. Neurosci Lett. 2004;363(2):173–177. doi: 10.1016/j.neulet.2004.04.003. [DOI] [PubMed] [Google Scholar]
- 116.Tracy K, Dibling BC, Spike BT, Knabb JR, Schumacker P, Macleod KF. BNIP3 is an RB/E2F target gene required for hypoxia-induced autophagy. Mol Cell Biol. 2007;27(17):6229–6242. doi: 10.1128/MCB.02246-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Verderio EA, Johnson TS, Griffin M. Transglutaminases in wound healing and inflammation. Prog Exp Tumor Res. 2005;38:89–114. doi: 10.1159/000084235. [DOI] [PubMed] [Google Scholar]
- 118.Verderio EA, Johnson T, Griffin M. Tissue transglutaminase in normal and abnormal wound healing: review article. Amino Acids. 2004;26(4):387–404. doi: 10.1007/s00726-004-0094-4. [DOI] [PubMed] [Google Scholar]
- 119.Kim SY. Transglutaminase 2 in inflammation. Front Biosci J Virtual Libr. 2006;11:3026–3035. doi: 10.2741/2030. [DOI] [PubMed] [Google Scholar]
- 120.Luciani A, Villella VR, Esposito S, Brunetti-Pierri N, Medina D, Settembre C, Gavina M, Pulze L, Giardino I, Pettoello-Mantovani M, D’Apolito M, Guido S, Masliah E, Spencer B, Quaratino S, Raia V, Ballabio A, Maiuri L. Defective CFTR induces aggresome formation and lung inflammation in cystic fibrosis through ROS-mediated autophagy inhibition. Nat Cell Biol. 2010;12(9):863–875. doi: 10.1038/ncb2090. [DOI] [PubMed] [Google Scholar]
- 121.Maiuri L, Luciani A, Giardino I, Raia V, Villella VR, D’Apolito M, Pettoello-Mantovani M, Guido S, Ciacci C, Cimmino M, Cexus ON, Londei M, Quaratino S. Tissue transglutaminase activation modulates inflammation in cystic fibrosis via PPARgamma down-regulation. J Immunol. 2008;180(11):7697–7705. doi: 10.4049/jimmunol.180.11.7697. [DOI] [PubMed] [Google Scholar]
- 122.Kim Y, Lee YS, Hahn JH, Choe J, Kwon HJ, Ro JY, Jeoung D. Hyaluronic acid targets CD44 and inhibits FcepsilonRI signaling involving PKCdelta, Rac1, ROS, and MAPK to exert anti-allergic effect. Mol Immunol. 2008;45(9):2537–2547. doi: 10.1016/j.molimm.2008.01.008. [DOI] [PubMed] [Google Scholar]
- 123.Sohn J, Kim TI, Yoon YH, Kim JY, Kim SY. Novel transglutaminase inhibitors reverse the inflammation of allergic conjunctivitis. J Clin Investig. 2003;111(1):121–128. doi: 10.1172/JCI15937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Cheng G, Diebold BA, Hughes Y, Lambeth JD. Nox1-dependent reactive oxygen generation is regulated by Rac1. J Biol Chem. 2006;281(26):17718–17726. doi: 10.1074/jbc.M512751200. [DOI] [PubMed] [Google Scholar]
- 125.Kim SY, Grant P, Lee JH, Pant HC, Steinert PM. Differential expression of multiple transglutaminases in human brain. Increased expression and cross-linking by transglutaminases 1 and 2 in Alzheimer’s disease. J Biol Chem. 1999;274(43):30715–30721. doi: 10.1074/jbc.274.43.30715. [DOI] [PubMed] [Google Scholar]
- 126.Wilhelmus MM, Grunberg SC, Bol JG, van Dam AM, Hoozemans JJ, Rozemuller AJ, Drukarch B. Transglutaminases and transglutaminase-catalyzed cross-links colocalize with the pathological lesions in Alzheimer’s disease brain. Brain Pathol. 2009;19(4):612–622. doi: 10.1111/j.1750-3639.2008.00197.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.McConoughey SJ, Basso M, Niatsetskaya ZV, Sleiman SF, Smirnova NA, Langley BC, Mahishi L, Cooper AJ, Antonyak MA, Cerione RA, Li B, Starkov A, Chaturvedi RK, Beal MF, Coppola G, Geschwind DH, Ryu H, Xia L, Iismaa SE, Pallos J, Pasternack R, Hils M, Fan J, Raymond LA, Marsh JL, Thompson LM, Ratan RR. Inhibition of transglutaminase 2 mitigates transcriptional dysregulation in models of Huntington disease. EMBO Mol Med. 2010;2(9):349–370. doi: 10.1002/emmm.201000084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Munsie L, Caron N, Atwal RS, Marsden I, Wild EJ, Bamburg JR, Tabrizi SJ, Truant R. Mutant huntingtin causes defective actin remodeling during stress: defining a new role for transglutaminase 2 in neurodegenerative disease. Hum Mol Genet. 2011;20(10):1937–1951. doi: 10.1093/hmg/ddr075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Piredda L, Farrace MG, Lo Bello M, Malorni W, Melino G, Petruzzelli R, Piacentini M. Identification of ‘tissue’ transglutaminase binding proteins in neural cells committed to apoptosis. FASEB J Off Publ Fed Am Soc Exp Biol. 1999;13(2):355–364. doi: 10.1096/fasebj.13.2.355. [DOI] [PubMed] [Google Scholar]
- 130.Gutekunst CA, Levey AI, Heilman CJ, Whaley WL, Yi H, Nash NR, Rees HD, Madden JJ, Hersch SM. Identification and localization of huntingtin in brain and human lymphoblastoid cell lines with anti-fusion protein antibodies. Proc Natl Acad Sci USA. 1995;92(19):8710–8714. doi: 10.1073/pnas.92.19.8710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Li SH, Gutekunst CA, Hersch SM, Li XJ. Interaction of huntingtin-associated protein with dynactin P150Glued. J Neurosci. 1998;18(4):1261–1269. doi: 10.1523/JNEUROSCI.18-04-01261.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Billett HH, Puszkin EG. The red cell membrane contains calmodulin-regulated crosslinking and proteolytic activity. Hematol Pathol. 1991;5(4):185–193. [PubMed] [Google Scholar]
- 133.Zainelli GM, Ross CA, Troncoso JC, Fitzgerald JK, Muma NA. Calmodulin regulates transglutaminase 2 cross-linking of huntingtin. J Neurosci. 2004;24(8):1954–1961. doi: 10.1523/JNEUROSCI.4424-03.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Ballestar E, Abad C, Franco L. Core histones are glutaminyl substrates for tissue transglutaminase. J Biol Chem. 1996;271(31):18817–18824. doi: 10.1074/jbc.271.31.18817. [DOI] [PubMed] [Google Scholar]
- 135.Kim JH, Nam KH, Kwon OS, Kim IG, Bustin M, Choy HE, Park SC. Histone cross-linking by transglutaminase. Biochem Biophys Res Commun. 2002;293(5):1453–1457. doi: 10.1016/S0006-291X(02)00393-5. [DOI] [PubMed] [Google Scholar]
- 136.Shimizu T, Hozumi K, Horiike S, Nunomura K, Ikegami S, Takao T, Shimonishi Y. A covalently crosslinked histone. Nature. 1996;380(6569):32. doi: 10.1038/380032a0. [DOI] [PubMed] [Google Scholar]
- 137.Hand D, Perry MJ, Haynes LW. Cellular transglutaminases in neural development. Int J Dev Neurosci Off J Int Soc Dev Neurosci. 1993;11(6):709–720. doi: 10.1016/0736-5748(93)90060-q. [DOI] [PubMed] [Google Scholar]
- 138.Del Duca S, Beninati S, Serafini-Fracassini D. Polyamines in chloroplasts: identification of their glutamyl and acetyl derivatives. Biochem J. 1995;305(Pt 1):233–237. doi: 10.1042/bj3050233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Matthews HR. Polyamines, chromatin structure and transcription. BioEssays News Rev Mol Cell Dev Biol. 1993;15(8):561–566. doi: 10.1002/bies.950150811. [DOI] [PubMed] [Google Scholar]
- 140.Fesus L, Szondy Z. Transglutaminase 2 in the balance of cell death and survival. FEBS Lett. 2005;579(15):3297–3302. doi: 10.1016/j.febslet.2005.03.063. [DOI] [PubMed] [Google Scholar]
- 141.Rodolfo C, Mormone E, Matarrese P, Ciccosanti F, Farrace MG, Garofano E, Piredda L, Fimia GM, Malorni W, Piacentini M. Tissue transglutaminase is a multifunctional BH3-only protein. J Biol Chem. 2004;279(52):54783–54792. doi: 10.1074/jbc.M410938200. [DOI] [PubMed] [Google Scholar]
- 142.Porter GW, Khuri FR, Fu H. Dynamic 14-3-3/client protein interactions integrate survival and apoptotic pathways. Semin Cancer Biol. 2006;16(3):193–202. doi: 10.1016/j.semcancer.2006.03.003. [DOI] [PubMed] [Google Scholar]
- 143.Mackintosh C. Dynamic interactions between 14-3-3 proteins and phosphoproteins regulate diverse cellular processes. Biochem J. 2004;381(Pt 2):329–342. doi: 10.1042/BJ20031332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Mishra S, Murphy LJ. Phosphorylation of transglutaminase 2 by PKA at Ser216 creates 14-3-3 binding sites. Biochem Biophys Res Commun. 2006;347(4):1166–1170. doi: 10.1016/j.bbrc.2006.07.041. [DOI] [PubMed] [Google Scholar]
- 145.Caccamo D, Condello S, Ferlazzo N, Curro M, Griffin M, Ientile R. Transglutaminase 2 interaction with small heat shock proteins mediate cell survival upon excitotoxic stress. Amino Acids. 2013;44(1):151–159. doi: 10.1007/s00726-011-1083-z. [DOI] [PubMed] [Google Scholar]
- 146.Stetler RA, Gao Y, Signore AP, Cao G, Chen J. HSP27: mechanisms of cellular protection against neuronal injury. Curr Mol Med. 2009;9(7):863–872. doi: 10.2174/156652409789105561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Caccamo D, Curro M, Cusumano G, Crisafulli G, Ientile R. Excitotoxin-induced changes in transglutaminase during differentiation of cerebellar granule cells. Amino Acids. 2004;26(2):197–201. doi: 10.1007/s00726-003-0007-y. [DOI] [PubMed] [Google Scholar]
- 148.Ientile R, Caccamo D, Macaione V, Torre V, Macaione S. NMDA-evoked excitotoxicity increases tissue transglutaminase in cerebellar granule cells. Neuroscience. 2002;115(3):723–729. doi: 10.1016/s0306-4522(02)00482-7. [DOI] [PubMed] [Google Scholar]
- 149.Hanayama R, Tanaka M, Miwa K, Shinohara A, Iwamatsu A, Nagata S. Identification of a factor that links apoptotic cells to phagocytes. Nature. 2002;417(6885):182–187. doi: 10.1038/417182a. [DOI] [PubMed] [Google Scholar]
- 150.D’Eletto M, Farrace MG, Rossin F, Strappazzon F, Giacomo GD, Cecconi F, Melino G, Sepe S, Moreno S, Fimia GM, Falasca L, Nardacci R, Piacentini M. Type 2 transglutaminase is involved in the autophagy-dependent clearance of ubiquitinated proteins. Cell Death Differ. 2012;19(7):1228–1238. doi: 10.1038/cdd.2012.2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Snel B, Lehmann G, Bork P, Huynen MA. STRING: a web-server to retrieve and display the repeatedly occurring neighbourhood of a gene. Nucleic Acids Res. 2000;28(18):3442–3444. doi: 10.1093/nar/28.18.3442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Bossi A, Lehner B. Tissue specificity and the human protein interaction network. Mol Syst Biol. 2009;5:260. doi: 10.1038/msb.2009.17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Yee VC, Pedersen LC, Le Trong I, Bishop PD, Stenkamp RE, Teller DC. Three-dimensional structure of a transglutaminase: human blood coagulation factor XIII. Proc Natl Acad Sci USA. 1994;91(15):7296–7300. doi: 10.1073/pnas.91.15.7296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Noguchi K, Ishikawa K, Yokoyama K, Ohtsuka T, Nio N, Suzuki E. Crystal structure of red sea bream transglutaminase. J Biol Chem. 2001;276(15):12055–12059. doi: 10.1074/jbc.M009862200. [DOI] [PubMed] [Google Scholar]
- 155.Liu S, Cerione RA, Clardy J. Structural basis for the guanine nucleotide-binding activity of tissue transglutaminase and its regulation of transamidation activity. Proc Natl Acad Sci USA. 2002;99(5):2743–2747. doi: 10.1073/pnas.042454899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Pinkas DM, Strop P, Brunger AT, Khosla C. Transglutaminase 2 undergoes a large conformational change upon activation. PLoS Biol. 2007;5(12):e327. doi: 10.1371/journal.pbio.0050327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Ahvazi B, Boeshans KM, Idler W, Baxa U, Steinert PM. Roles of calcium ions in the activation and activity of the transglutaminase 3 enzyme. J Biol Chem. 2003;278(26):23834–23841. doi: 10.1074/jbc.M301162200. [DOI] [PubMed] [Google Scholar]
- 158.Fox BA, Yee VC, Pedersen LC, Le Trong I, Bishop PD, Stenkamp RE, Teller DC. Identification of the calcium binding site and a novel ytterbium site in blood coagulation factor XIII by X-ray crystallography. J Biol Chem. 1999;274(8):4917–4923. doi: 10.1074/jbc.274.8.4917. [DOI] [PubMed] [Google Scholar]
- 159.Kiraly R, Csosz E, Kurtan T, Antus S, Szigeti K, Simon-Vecsei Z, Korponay-Szabo IR, Keresztessy Z, Fesus L. Functional significance of five noncanonical Ca2+-binding sites of human transglutaminase 2 characterized by site-directed mutagenesis. FEBS J. 2009;276(23):7083–7096. doi: 10.1111/j.1742-4658.2009.07420.x. [DOI] [PubMed] [Google Scholar]
- 160.Bergamini CM, Dondi A, Lanzara V, Squerzanti M, Cervellati C, Montin K, Mischiati C, Tasco G, Collighan R, Griffin M, Casadio R. Thermodynamics of binding of regulatory ligands to tissue transglutaminase. Amino Acids. 2010;39(1):297–304. doi: 10.1007/s00726-009-0442-5. [DOI] [PubMed] [Google Scholar]
- 161.Davey NE, Van Roey K, Weatheritt RJ, Toedt G, Uyar B, Altenberg B, Budd A, Diella F, Dinkel H, Gibson TJ. Attributes of short linear motifs. Mol Biosyst. 2012;8(1):268–281. doi: 10.1039/c1mb05231d. [DOI] [PubMed] [Google Scholar]
- 162.Fuxreiter M, Tompa P, Simon I. Local structural disorder imparts plasticity on linear motifs. Bioinformatics. 2007;23(8):950–956. doi: 10.1093/bioinformatics/btm035. [DOI] [PubMed] [Google Scholar]
- 163.Ward JJ, Sodhi JS, McGuffin LJ, Buxton BF, Jones DT. Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J Mol Biol. 2004;337(3):635–645. doi: 10.1016/j.jmb.2004.02.002. [DOI] [PubMed] [Google Scholar]
- 164.Vucetic S, Brown CJ, Dunker AK, Obradovic Z. Flavors of protein disorder. Proteins. 2003;52(4):573–584. doi: 10.1002/prot.10437. [DOI] [PubMed] [Google Scholar]
- 165.Dosztanyi Z, Csizmok V, Tompa P, Simon I. IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics. 2005;21(16):3433–3434. doi: 10.1093/bioinformatics/bti541. [DOI] [PubMed] [Google Scholar]
- 166.Dosztanyi Z, Csizmok V, Tompa P, Simon I. The pairwise energy content estimated from amino acid composition discriminates between folded and intrinsically unstructured proteins. J Mol Biol. 2005;347(4):827–839. doi: 10.1016/j.jmb.2005.01.071. [DOI] [PubMed] [Google Scholar]
- 167.Dinkel H, Van Roey K, Michael S, Davey NE, Weatheritt RJ, Born D, Speck T, Kruger D, Grebnev G, Kuban M, Strumillo M, Uyar B, Budd A, Altenberg B, Seiler M, Chemes LB, Glavina J, Sanchez IE, Diella F, Gibson TJ. The eukaryotic linear motif resource ELM: 10 years and counting. Nucleic Acids Res. 2014;42(Database issue):D259–D266. doi: 10.1093/nar/gkt1047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Corti A, Curnis F. Isoaspartate-dependent molecular switches for integrin-ligand recognition. J Cell Sci. 2011;124(Pt 4):515–522. doi: 10.1242/jcs.077172. [DOI] [PubMed] [Google Scholar]
- 169.Wang Z, Telci D, Griffin M. Importance of syndecan-4 and syndecan-2 in osteoblast cell adhesion and survival mediated by a tissue transglutaminase–fibronectin complex. Exp Cell Res. 2011;317(3):367–381. doi: 10.1016/j.yexcr.2010.10.015. [DOI] [PubMed] [Google Scholar]
- 170.Echtermeyer F, Harendza T, Hubrich S, Lorenz A, Herzog C, Mueller M, Schmitz M, Grund A, Larmann J, Stypmann J, Schieffer B, Lichtinghagen R, Hilfiker-Kleiner D, Wollert KC, Heineke J, Theilmeier G. Syndecan-4 signalling inhibits apoptosis and controls NFAT activity during myocardial damage and remodelling. Cardiovasc Res. 2011;92(1):123–131. doi: 10.1093/cvr/cvr149. [DOI] [PubMed] [Google Scholar]
- 171.Shenoy SK, Lefkowitz RJ (2005) Seven-transmembrane receptor signaling through beta-arrestin. Sci STKE Signal Transduct Knowl Environ 308:cm10. doi:10.1126/stke.2005/308/cm10 [DOI] [PubMed]
- 172.Hao N, Behar M, Elston TC, Dohlman HG. Systems biology analysis of G protein and MAP kinase signaling in yeast. Oncogene. 2007;26(22):3254–3266. doi: 10.1038/sj.onc.1210416. [DOI] [PubMed] [Google Scholar]
- 173.Braun RFaW. Exact and efficient analytical calculation of the accessible surface areas and their gradients for macromolecules. J Comput Chem. 1998;19(3):319–333. [Google Scholar]
- 174.Stamnaes J, Fleckenstein B, Lund-Johansen F, Sollid LM. The monoclonal antibody 6B9 recognizes CD44 and not cell surface transglutaminase 2. Scand J Immunol. 2008;68(5):534–542. doi: 10.1111/j.1365-3083.2008.02173.x. [DOI] [PubMed] [Google Scholar]
- 175.Hodrea J, Demeny MA, Majai G, Sarang Z, Korponay-Szabo IR, Fesus L. Transglutaminase 2 is expressed and active on the surface of human monocyte-derived dendritic cells and macrophages. Immunol Lett. 2010;130(1–2):74–81. doi: 10.1016/j.imlet.2009.12.010. [DOI] [PubMed] [Google Scholar]
- 176.Teesalu K, Panarina M, Uibo O, Uibo R, Utt M. Autoantibodies from patients with celiac disease inhibit transglutaminase 2 binding to heparin/heparan sulfate and interfere with intestinal epithelial cell adhesion. Amino Acids. 2012;42(2–3):1055–1064. doi: 10.1007/s00726-011-1020-1. [DOI] [PubMed] [Google Scholar]
- 177.Halttunen T, Maki M. Serum immunoglobulin A from patients with celiac disease inhibits human T84 intestinal crypt epithelial cell differentiation. Gastroenterology. 1999;116(3):566–572. doi: 10.1016/s0016-5085(99)70178-2. [DOI] [PubMed] [Google Scholar]
- 178.Myrsky E, Kaukinen K, Syrjanen M, Korponay-Szabo IR, Maki M, Lindfors K. Coeliac disease-specific autoantibodies targeted against transglutaminase 2 disturb angiogenesis. Clin Exp Immunol. 2008;152(1):111–119. doi: 10.1111/j.1365-2249.2008.03600.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Simon-Vecsei Z, Kiraly R, Bagossi P, Toth B, Dahlbom I, Caja S, Csosz E, Lindfors K, Sblattero D, Nemes E, Maki M, Fesus L, Korponay-Szabo IR. A single conformational transglutaminase 2 epitope contributed by three domains is critical for celiac antibody binding and effects. Proc Natl Acad Sci USA. 2012;109(2):431–436. doi: 10.1073/pnas.1107811108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Iversen R, Di Niro R, Stamnaes J, Lundin KE, Wilson PC, Sollid LM. Transglutaminase 2-specific autoantibodies in celiac disease target clustered, N-terminal epitopes not displayed on the surface of cells. J Immunol. 2013;190(12):5981–5991. doi: 10.4049/jimmunol.1300183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Mehta K, Kumar A, Kim HI. Transglutaminase 2: a multi-tasking protein in the complex circuitry of inflammation and cancer. Biochem Pharmacol. 2010;80(12):1921–1929. doi: 10.1016/j.bcp.2010.06.029. [DOI] [PubMed] [Google Scholar]
- 182.Kim Y, et al. Transglutaminase II interacts with rac1, regulates production of reactive oxygen species, expression of snail, secretion of Th2 cytokines and mediates in vitro and in vivo allergic inflammation. Mol Immunol. 2010;47:1010–1022. doi: 10.1016/j.molimm.2009.11.017. [DOI] [PubMed] [Google Scholar]
- 183.Trejo Skalli AV, et al. Association of a transglutaminase-related antigen with intermediate filaments. Proc Natl Acad Sci USA. 1995;92:48940–48944. doi: 10.1073/pnas.92.19.8940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 184.Korponay-Szabó IR, et al. J Pediatr Gastroenterol Nutr. 2008;46:253–261. doi: 10.1097/MPG.0b013e31815ee555. [DOI] [PubMed] [Google Scholar]
- 185.Iversen R, et al. J Immunol. 2013;190:5981–5991. doi: 10.4049/jimmunol.1300183. [DOI] [PMC free article] [PubMed] [Google Scholar]