Abstract
We recently identified a cDNA encoding a putative isovaleryl-coenzyme A (CoA) dehydrogenase in Arabidopsis (AtIVD). In animals, this homotetrameric enzyme is located in mitochondria and catalyzes the conversion of isovaleryl-CoA to 3-methylcrotonyl-CoA as an intermediate step in the leucine (Leu) catabolic pathway. Expression of AtIVD:smGFP4 fusion proteins in tobacco (Nicotiana tabacum) protoplasts and biochemical studies now demonstrate the in vivo import of the plant isovaleryl-CoA dehydrogenase (IVD) into mitochondria and the enzyme in the matrix of these organelles. Two-dimensional separation of mitochondrial proteins by blue native and SDS-PAGE and size determination of the native and overexpressed proteins suggest homodimers to be the dominant form of the plant IVD. Northern-blot hybridization and studies in transgenic Arabidopsis plants expressing Ativd promoter:gus constructs reveal strong expression of this gene in seedlings and young plants grown in the absence of sucrose, whereas promoter activity in almost all tissues is strongly inhibited by exogeneously added sucrose. Substrate specificity tests with AtIVD expressed in Escherichia coli indicate a strong preference toward isovaleryl-CoA but surprisingly also show considerable activity with isobutyryl-CoA. This strongly indicates a commitment of the enzyme in Leu catabolism, but the activity observed with isobutyryl-CoA also suggests a parallel involvement of the enzyme in the dehydrogenation of intermediates of the valine degradation pathway. Such a dual activity has not been observed with the animal IVD and may suggest a novel connection of the Leu and valine catabolism in plants.
The synthesis of branched-chain amino acids in higher plants has been intensively investigated and cDNAs and genes encoding most of the enzymes involved are well characterized (Singh and Shaner, 1995; Singh, 1999). In contrast, much less is known about the degradation of this group of amino acids in plants. Several reports, however, indicate a metabolization of at least Leu. Stewart and Beevers (1967) reported gluconeogenesis from several amino acids including Leu in the castor bean (Ricinus communis) endosperm. It was also found that Leu, which is in large excess contained in maize (Zea mays) storage proteins, is respired and converted to starch, sugars, and organic acids (Sodek and Wilson, 1973). These observations were substantiated by the observation that branched-chain 2-oxo acids, intermediates of branched-chain amino acid catabolism, are degraded in peroxisomes of mung bean (Vigna radiata). It was also speculated that additional extra-peroxisomal Val and Leu degradation pathways might exist (Gerbling and Gerhardt, 1988, 1989). An initial clue that such a pathway does exist in plant mitochondria was the identification of a 3-methylcrotonyl-coenzyme A (CoA) carboxylase (MCCase) in several plant species (Alban et al., 1993; Song et al., 1994; Wang et al., 1994; Weaver et al., 1995; Aubert et al., 1996). Biochemical studies of this enzyme in sycamore (Acer pseudoplatanus) cells and in soybean (Glycine max) showed that this mitochondrial enzyme is involved in Leu catabolism. Several other enzyme activities implicated in Leu degradation have been more recently identified in soybean and maize. This includes an isovaleryl-CoA dehydrogenase (IVD; EC 1.3.99.10) activity, which catalyzes the conversion of isovaleryl-CoA to 3-methylcrotonyl-CoA (Anderson et al., 1998; Bode et al., 1999). We recently described a cDNA encoding a protein with high similarity to the mammalian IVDs, thus belonging to the acyl-CoA dehydrogenase (ACDH) gene family. This putative IVD was shown to cofractionate with the mitochondrial protein porin, suggesting a mitochondrial localization of this protein (Däschner et al., 1999). To better understand the function and significance of this enzyme in plants, we investigated the exact localization of this enzyme, the conformation of its native form and its substrate specificity, as well as its expression in different tissues and developmental stages.
RESULTS
Transiently Expressed AtIVD:smGFP4 Fusion Protein Is Imported into Mitochondria of Tobacco (Nicotiana tabacum) Protoplasts
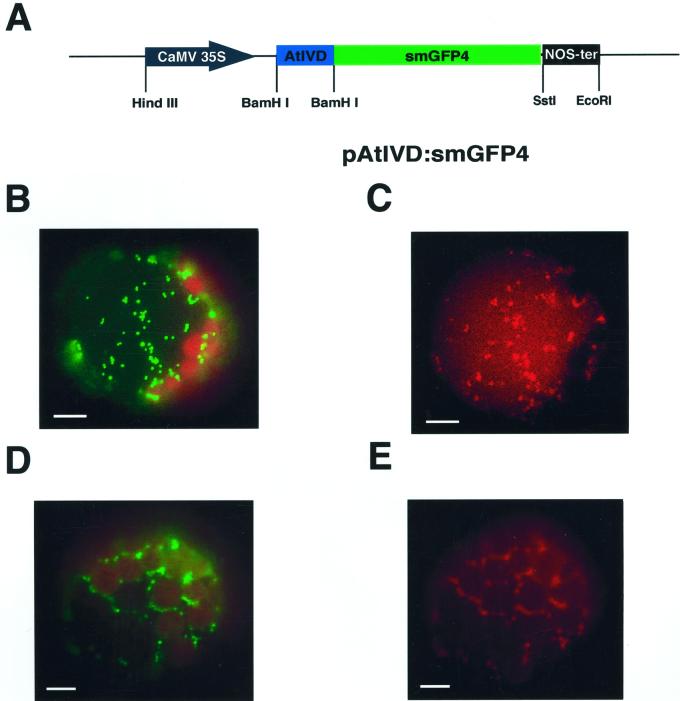
We recently found that a potential plant IVD cofractionates with the mitochondrial protein porin, suggesting a mitochondrial location (Däschner et al., 1999). To unambiguously determine the subcellular localization of the AtIVD in vivo and the functionality of the putative target sequence, a cDNA fragment encoding the 50-amino acid-long N-terminal part of the IVD was fused to the cDNA of smGFP4 (Fig. 1A). This ivd cDNA fragment comprises the N-terminal 25 amino acids most likely representing the mitochondrial targeting sequence as deduced from N-terminal protein sequencing data of the homologous protein in pea (Pisum sativum; Däschner et al., 1999). The cDNA fusion construct was cloned downstream of the cauliflower mosaic virus 35S promoter and transiently transformed into tobacco protoplasts. Transformed protoplasts were simultaneously incubated with MitoTracker Red for specific staining of mitochondria. Inspection of the protoplasts transformed with the AtIVD:smGFP4 construct by fluorescence microscopy revealed the GFP fluorescence to be present in particles with sizes of about 1 μm, which corresponds well with the size of mitochondria. The colocalization of the GFP fluorescence with the red fluorescence of the MitoTracker dye confirms the mitochondrial targeting of the respective GFP fusion (Fig. 1, B and C). A similar distribution of the green GFP and the red MitoTracker fluorescence was likewise observed with a GFP fusion protein containing the N-terminal part of the mitochondrial chaperonin CPN-60 protein, whose mitochondrial localization has been documented previously (Fig. 1, D and E; Logan and Leaver, 2000). These data clearly demonstrate the mitochondrial targeting of the AtIVD:smGFP4 protein in vivo and thus unambiguously establish the mitochondrial localization of the plant IVD.
Figure 1.
An AtIVD:smGFP4 fusion protein is imported into mitochondria of tobacco protoplasts. A, Construct carrying the gene encoding AtIVD:smGFP4 fusion protein. The N-terminal part of the Ativd cDNA encoding the first 50 amino acids (AtIVD, blue box) is cloned upstream of the smGFP4 reading frame (smGFP4, green box). Expression of the resulting fusion protein is controlled by the cauliflower mosaic virus 35S promoter (black arrow) and the NOS terminator (black box). Restriction sites are indicated for HindIII, BamHI, SstI, and EcoRI. B through E, Tobacco protoplasts expressing the AtIVD:smGFP4 fusion protein (B and C) and CPN-60:mGFP5 (D and E). Images B and D were taken with flourescein isothiocyanat (BP 450–490/LP515) and C and E were analyzed with MitoTracker (HQ545/30/HQ 610/75) filter sets, respectively. The green fluorescent protein (GFP) is observed in particles with sizes of about 1 μm. The colocalization of the GFP and MitoTracker Red fluorescence confirms the localization of the fusion proteins in mitochondria. The bars given in the individual frames correspond to 10 μm.
The Plant IVD Is Located in the Mitochondrial Matrix
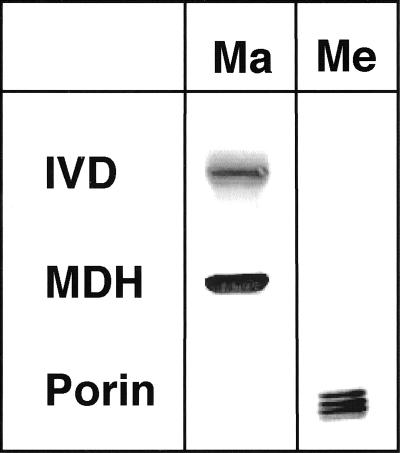
To further determine the localization of the plant IVD within these organelles, different pea mitochondrial subfractions were tested with antibodies against human mitochondrial IVD. Decoration of proteins with sizes of 34 and 30 kD with antibodies against the matrix located mitochondrial malate dehydrogenase (MDH) and the outer membrane polypeptide porin confirmed the clear separation of membrane and matrix fractions. The identification of a 43-kD protein in the matrix fraction with the IVD-specific antibody demonstrates the localization of the plant IVD in the mitochondrial matrix space (Fig. 2).
Figure 2.
The AtIVD is located in the mitochondrial matrix. Mitochondria isolated from etiolated pea seedlings were fractionated into membrane (Me) and matrix (Ma) fractions. The quality of the fractionation was confirmed by immunodetection with antibodies against the mitochondrial matrix protein malate dehydrogenase from watermelon (Citrullus lanatus; MDH) and the outer membrane protein porin of potato (Solanum tuberosum; Porin). Immunodetection using an antibody against the human mitochondrial IVD decorates a protein of 43 kD only in the matrix fraction confirming the presence of the IVD in the mitochondrial matrix space.
Isovaleryl-CoA and Isobutyryl-CoA Are Substrates of the AtIVD
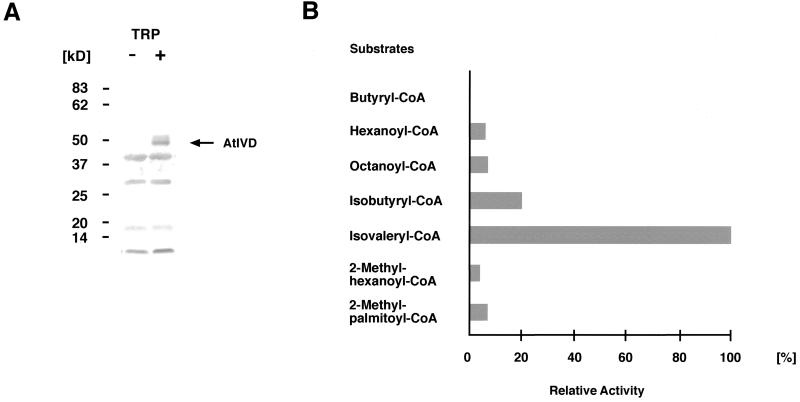
The IVD of Arabidopsis and other plant species share more than 60% identical amino acids with the homologous proteins from human and rat (Rattus norvegicus), but only about 30% or less with other ACDHs. Although these similarities suggest an assignment as IVD, it is nevertheless necessary to determine the exact substrate specificity of the plant enzymes. To this end the AtIVD was overexpressed in Escherichia coli and crude protein lysates were assayed for activity toward various acyl-CoA substrates (Fig. 3). In repeated tests with different overexpressing clones, highest activity is clearly found with isovaleryl-CoA, an intermediate of the Leu catabolism. Significant activity, however, is also found with isobutyryl-CoA, a breakdown product of Val. In enzyme activity tests with substrate concentrations varying between 9 and 70 μm, AtIVD showed typical Michaelis-Menten kinetics toward isobutyryl-CoA and isovaleryl-CoA. Apparent Km and Vmax values for these substrates were 50 (±30) μm and 4 (±1) nmol/mg/min for isovaleryl-CoA and 500 (±200) μm and 6 (±2) nmol mg−1 min−1 for isobutyryl-CoA. Although minor activities are also repeatedly observed with 2-methylhexanoyl-CoA and 2-methylpalmitoyl-CoA, differing activities are measured with the short straight-chain acyl-CoA substrates. Weak activities were seen with octanoyl- and hexanoyl-CoA, whereas butyryl-CoA was not metabolized at all (Table I, Fig. 3). No activity with any of these substrates was observed in control experiments with E. coli lysates from clones expressing the vector without Ativd insert, confirming the measured activities to originate from the expressed AtIVD.
Figure 3.
Substrate specificity of the overexpressed AtIVD. A, Western-blot analysis of the overexpressed AtIVD with an IVD-specific antibody against the homologous protein from human. A clear signal corresponding to the expected size of about 43 kD (highlighted by an arrow and designated AtIVD) is specifically detected only after induction (lane +) with Trp (TRP) but is not observed in noninduced E. coli cells (lane −). B, Relative activities (given in %) measured with different acyl-CoA substrates at 50-μm final concentration in a typical enzyme test. Highest activity as found with isovaleryl-CoA is arbitrarily set at 100%. Values correspond to those given in Table I, first column.
Table I.
Substrate specificity of the AtIVD
| Substrate | Relative Activity | Average Relative Activity |
|---|---|---|
| % | ||
| Isovaleryl-CoA | 100 | 100 |
| Isobutyryl-CoA | 20 | 22 ± 2 |
| 2-Methyl-palmitoyl-CoA | 7 | 6 ± 2 |
| 2-Methyl-hexanoyl-CoA | 4 | 5 ± 2 |
| Octanoyl-CoA | 7 | 7 ± 3 |
| Hexanoyl-CoA | 6 | 4 ± 2 |
| Butyryl-CoA | 0 | 0 |
These data strongly suggest that the previously described potential ivd reading frame from Arabidopsis encodes an IVD, which is most likely involved in the degradation of Leu in higher plants. The surprising oxidation of isobutyryl-CoA furthermore indicates a potential involvement of this enzyme in the breakdown of Val.
The Arabidopsis IVD Is a Homodimer
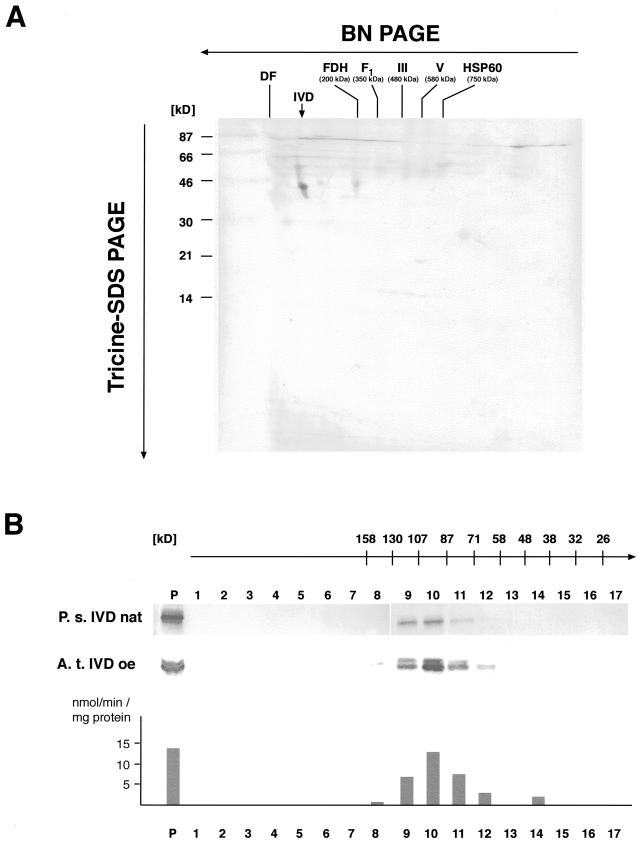
The IVD is a member of the ACDH protein family, which catalyzes the β-oxidative degradation of fatty acids and carbon moieties of branched-chain amino acids in animal mitochondria. In these organisms ACDHs usually form homotetramers with each subunit binding flarin adenine dinucleotide as a prosthetic group (Ikeda and Tanaka, 1983b). To gain more information about the native form of the IVD in plants, Arabidopsis mitochondrial proteins were separated by two-dimensional gel electrophoresis including a separation under native conditions by blue native (BN)-PAGE in the first dimension. Western-blot analysis using an antibody against the human IVD detects a single 43-kD protein corresponding to the mature mass of the monomer (Fig. 4A). The migration behavior of this protein under native conditions in the first dimension indicates that the IVD is either present as a monomer or dimer but reveals no trace of a tetramer.
Figure 4.
The plant IVD is a homodimer. A, Two-dimensional separation of Arabidopsis mitochondrial proteins. Mitochondrial proteins obtained from an Arabidopsis tissue culture were separated under native conditions in the presence of Coomassie Blue in the first dimension (BN-PAGE). A lane containing the size-separated protein complexes was cut out and transferred to Tricine-SDS-PAGE in the second dimension. Proteins were subsequently transferred to polyvinylidene difluoride membranes and investigated by immunostaining. A spot corresponding to the IVD polypeptide with 43 kD is detected with the IVD-specific antibody (indicated by a vertical arrow). The migration behavior of the protein in the first dimension indicates a native size of about 60 to 100 kD, suggesting a mono- or dimeric native structure of the protein. Masses of native respiratory chain complexes in the first dimension are given for fumarate dehydrogenase (FDH), the F1 part of complex V (F1), complexes III (III) and V (V), and the CPN-60 complex. The position of the dye front (DF) is also indicated. Molecular masses of marker proteins co-electrophoresed in the second dimension are given in kD on the left-hand side. B, Gel filtration chromatography of pea mitochondrial proteins and E. coli proteins containing overexpressed AtIVD. The apparent sizes (given in kD above the numbering of the fractions) expected in the individual fractions are deduced from the calibration of the column with marker proteins. Proteins of the individual fractions are investigated by western-blot/immunostaining analysis with an IVD-specific antibody. The native pea IVD (P.s. IVD nat, upper) as well as overexpressed AtIVD (A. t. IVD oe, lower) are detected in the protein fraction prior to separation (P) and in fractions corresponding to proteins with molecular masses between 130 and 71 kD with the majority of the protein eluting in fraction 10 corresponding to proteins between 107 and 87 kD. This is about twice the molecular mass of the plant IVD corroborating the dimeric form of the plant IVD. Enzyme activity (given in nmol min−1 mg−1 protein) of eluted fractions containing overexpressed AtIVD was measured with isovaleryl-CoA. Levels of activity correspond to the amounts of detected IVD protein indicating that the protein remained in its active state during the gel filtration analysis.
To evaluate this observation with a different unrelated technique, total pea mitochondrial proteins were size fractionated by gel-filtration chromatography. The IVD-specific antibody detects the respective protein in fractions corresponding to molecular sizes between 130 and 71 kD. The majority of the IVD elutes in a fraction corresponding to proteins with masses between 107 and 87 kD. This is about twice the molecular mass of the IVD, which substantiates the dimeric quaternary structure of this protein in plants (Fig. 4B).
An almost identical result was obtained in an analogous analysis of the AtIVD overexpressed in E. coli, which elutes in the same fractions as the pea IVD. Furthermore, enzyme activity tests of the eluted protein with isovaleryl-CoA showed a good correlation of the enzyme activity with the intensity of AtIVD revealed by immunodetection in the individual fractions. These results show that the proteins are retained in their active state during the gel filtration chromatography and thus confirm that the observed dimer is the active form of the IVD in plants (Fig. 4C).
Expression of the Arabidopsis IVD Is Repressed by Suc
To study transcription of the ivd gene in Arabidopsis, a northern-blot hybridization was performed as an initial experiment. Hybridization with a probe representing the complete Ativd cDNA to total RNA prepared from aboveground portions of about 2- to 3-week-old plants harvested in the early rosette stadium detects a signal corresponding to an mRNA of 1.4 to 1.5 kb. This covers well the 1,230-bp-large Ativd reading frame and indicates that the 5′- and 3′-untranslated parts together account for about 150 to 300 nucleotides of the mRNA (Fig. 5).
Figure 5.

Northern-blot analysis of total Arabidopsis RNAs from aboveground tissues. The hybridization with a probe corresponding to the complete Ativd cDNA detects a single RNA species of about 1.4 to 1.5 kb. Sizes of co-electrophoresed RNA marker molecules are given in kb.
To determine the expression of the ivd gene in more detail a potential promoter region covering 1.8 kb upstream of the translation start codon of the gene was fused to the gus reporter gene and transformed into Arabidopsis plants. Inspection of plants of different developmental stages obtained from the T2 generation grown on Murashige and Skoog (MS) medium with Suc by glucuronidase (GUS) staining of different lines revealed expression of the ivd gene almost exclusively in roots. GUS staining is visible in a zone immediately above the root tips and in the specialization zone. Weak, residual staining was occasionally observed in aboveground portions such as leaves or the apical meristem (Fig. 6, A–D).
Figure 6.
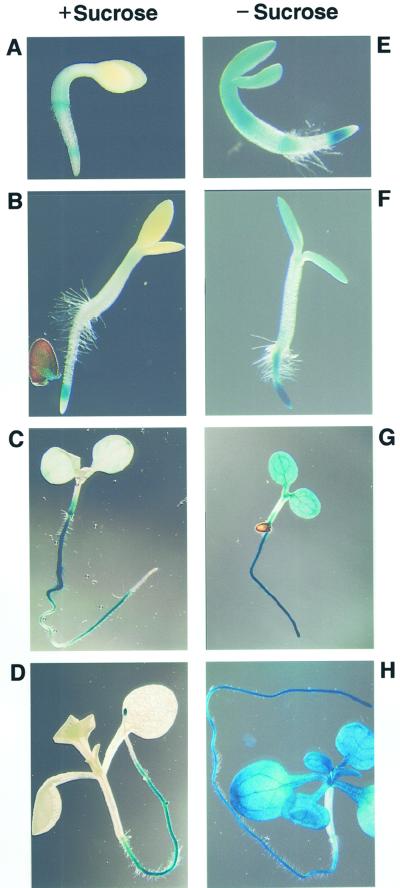
Ativd promoter activity is significantly inhibited by Suc. A through H, GUS staining of Arabidopsis seedlings grown in the presence (A–D) or absence (E–H) of 0.5% (w/v) Suc. In early developmental stages (A and B), expression observed in seedlings grown in the absence of Suc is rather low. In later stages, however, clear differences in Ativd promoter activity become apparent between seedlings grown in the absence (G and H) or presence (C and D) of Suc. No influence of Suc is observed in root tissues. Seedlings correspond to stages 2 (A and E), 3 (B and F), 7 (C and G), and 15 (D and H) d after sowing.
The general absence of the ivd promoter activity in aboveground tissues is in contrast to the result of the northern analysis, where a relatively strong signal indicates a high expression level in these parts of the plant. Because the plants used in the northern-blot analysis were grown on soil in the absence of Suc, we speculated whether exogeneously added Suc might inhibit promoter activity. Therefore, analogous GUS staining experiments were performed with plants grown on MS medium without Suc. After 2 to 3 d of germination, significant differences in promoter activity already became apparent in aboveground tissues of plants grown in the presence or absence of Suc (Fig. 6, A, B, F, and G). Suc repression of Ativd transcription is most obvious in about 2- to 3-week-old plants with an almost complete absence of promoter activity in aboveground parts of plants grown in the presence of Suc (Fig. 6, C and D), but a strong activity indicated by deep blue staining in seedlings grown without exogeneously added Suc (Fig. 6, G and H).
A northern-blot analysis of total RNA obtained from aboveground tissues of plants grown in the presence of Suc does not detect any ivd mRNA, confirming the results of the promoter gus analysis detailed above (data not shown).
DISCUSSION
Direct Evidence of Mitochondrial Localization of AtIVD, a Member of the ACDH Family in Higher Plants
Previous studies based on subcellular fractionation strongly indicated that some ACDH enzyme activities were associated with mitochondria in higher plants (Bode et al., 1999; Däschner et al., 1999). The data presented here now provide direct evidence that the AtIVD, a member of the ACDH enzyme family, is directed to the mitochondria. Moreover, its subcellular location in the mitochondrial matrix is clearly demonstrated, which is in accordance with what is known of the mammalian ACDH. This unambiguous localization of the AtIVD in mitochondria, together with the mitochondrial localization of the MCCase and the putative mitochondrial targeting signal of two subunits of the branched-chain ketoacid dehydrogenase, strongly support the importance of the plant mitochondrion for Leu metabolization (Fujiki et al., 2000). The AtIVD is so far the sole mitochondrial member of the plant ACDH family. Another homology-based putative Arabidopsis short-chain ACDH has been characterized as a peroxisomal acyl-CoA oxidase (Hayashi et al., 1999). However, other ACDH activities with varying substrate specificities have been described in different tissues of various plants, suggesting the presence of fatty acid β-oxidation pathways in plant mitochondria and thus the presence of additional members of the ACDH gene family in mitochondria of plants (Bode et al., 1999; Masterson and Wood, 2000). At present, a second, peroxisomal degradation pathway suggested by Gerbling and Gerhardt (1988, 1989) cannot be excluded; however, the respective genes from mung bean have not been isolated and the complete genomic sequence of Arabidopsis presents no clear candidate genes for such peroxisomal enzymes.
The Arabidopsis IVD Shows Considerable Activity toward Isobutyryl-CoA
The recently identified putative IVD from Arabidopsis shows more than 60% amino acid identity with the respective enzymes from human and rat, which suggested an analogous function of the plant enzyme (Däschner et al., 1999). Detailed substrate specificity analysis with the overexpressed AtIVD now confirms the strong activity of the plant enzyme toward isovaleryl-CoA. The apparent Km value of 50 μm for this substrate is similar to the Km value of 33 μm of the rat IVD measured under the same assay conditions (Ikeda and Tanaka, 1983a). These results strongly suggest a participation of the plant enzyme in Leu catabolism analogous to the function of this enzyme in animal mitochondria, which now justifies the plant enzyme to be termed IVD (Fig. 7).
Figure 7.
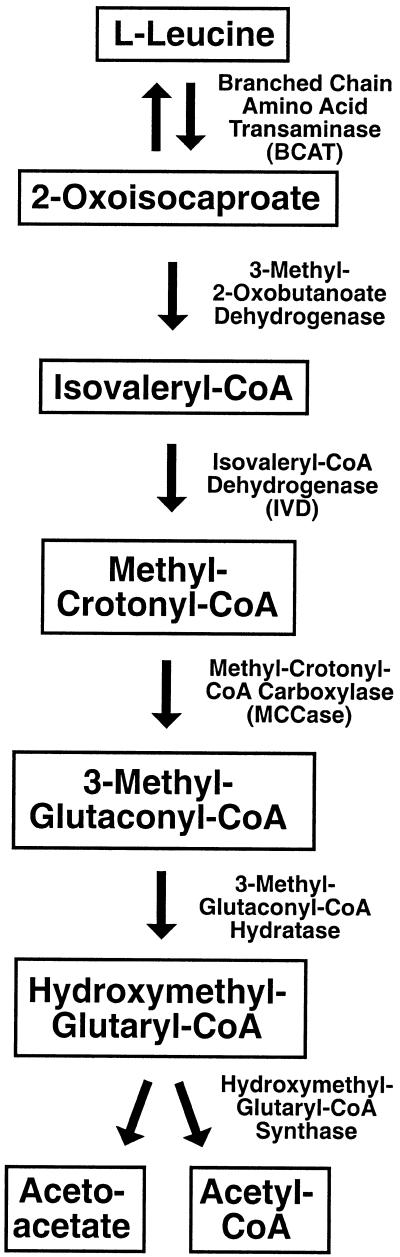
Scheme of the catabolic pathway of Leu. Intermediates are given in boxes. All enzymatic activities (given at the right-hand sides of the vertical arrows) have been crudely detected in soybean (Anderson et al., 1998) and are well characterized in animals.
Although with a much higher Km than that for isovaleryl-CoA, considerable activity is also observed with isobutyryl-CoA, a breakdown intermediate of Val. Thus, AtIVD also appears to convert 2-methyl-substituted substrates. This is in sharp contrast to the animal mitochondrial enzyme, which does not convert the latter substrate but instead shows significant activity with n-valeryl-CoA (relative activity 32%) and some background activity with n-butyryl-CoA (Ikeda and Tanaka, 1983a). All other substrates tested, mostly straight-chain acyl-CoA compounds, are not converted by the rat IVD. In animals, isobutyryl-CoA as well as 2-methyl-butyryl-CoA are oxidized by a separate enzyme, the 2-methyl-branched-chain ACDH, which can be clearly distinguished from other acyl-CoA enzymes by the differing substrate specificities (Ikeda and Tanaka, 1983b; Ikeda et al., 1983, Izai et al., 1992). This seems to be different in plants where a single enzyme, the IVD, can convert both isovaleryl-CoA and isobutyryl-CoA, which may indicate an involvement of this enzyme in the degradation of all branched-chain amino acids. Although enzyme activities with 2-methyl-butyryl-CoA, a breakdown product of iso-Leu, has not been detected so far, some activity on isobutyryl-CoA has been reported from a peroxisomal short-chain acyl-CoA oxidase (Hayashi at al., 1999).
Although the mobilization of Leu has been reported previously, very little is known about the degradation of iso-Leu and Val in plants. Indirect evidence for the degradation of Val is given by the investigation of carbohydrate-starved maize roots, where Val and Leu reach similar levels and show the same kinetics of accumulation and utilization (Brouquisse et al., 1992). Thus, it will be necessary to analyze and identify other enzyme activities involved in the breakdown of these branched-chain amino acids and to detect the respective degradation intermediates to elaborate the catabolic pathways for iso-Leu and Val in plants.
Besides the strong activities with isovaleryl-CoA and isobutyryl-CoA, weak activities are also observed with longer 2-methyl-acyl-CoA and some short straight-chain acyl-CoA substrates. These activities are very low and their significance is not yet clear. They may, however, be indicative for the relaxed and broader carbon chain specificity of the plant IVD. The substrate specificity tests of the plant as well as the animal enzymes were carried out under identical experimental conditions, suggesting the differing substrate specificities not to be the result of varying assay conditions (Ikeda and Tanaka, 1983a).
The Plant IVD Has a Dimeric Quaternary Structure
Beside the extended substrate specificity, the dominant aggregation of the plant IVD as a homodimer is another feature that significantly differentiates the plant ACDH from the homologous mammalian proteins. Although the latter frequently form homotetramers, this quaternary structure is not observed with the Arabidopsis and pea IVD. In contrast to all results presented above, the BN-PAGE and the gel filtration analyses indicate a homodimer to be the preferred native structure of the plant IVD. Stable dimers were also observed during the SDS PAGE analysis of the animal 2-methyl-branched-chain ACDH, which migrates as a dimer both in the presence and absence of 2-mercaptoethanol and without boiling prior to gel electrophoresis (Ikeda and Tanaka, 1983b). Although usually detected as a homotetramer, the dimeric form also has been observed recently in in vitro import assays with wild type and mutated forms of the human IVD. These most likely represent intermediates of the formation of the tetramer as a dimer of a dimer (Volchenboum and Vockley, 2000).
Dimers are also the native form of the very-long-chain ACDH from rat liver. But this enzyme, whose monomer has a molecular mass of 71 kD, also shows other substantial differences in comparison to the ACDHs with molecular masses around 43 kD (Izai et al., 1992).
What Is Responsible for the Altered Substrate Specificity and Quaternary Structure?
Although more than 60% of the amino acids are identical between the Arabidopsis and the human IVD, here we report significant differences in substrate specificity and quaternary structure between the two enzymes. The amino acids L95, A99, L103, T168, L258, L370, Y371, G374, and A375, recently identified to line the substrate-binding pocket in the human IVD, are all conserved in the Arabidopsis enzyme. This includes the G at position 374, which is responsible for the wider binding pocket when compared with the medium-chain ACDH (Tiffany et al., 1997). Thus, other features may be responsible for the altered substrate specificity of the Arabidopsis IVD.
Primary structural differences are seen, for example, between the C termini of the Arabidopsis and human IVD amino acid sequences. The crystal structure of the human IVD identified this part of the protein to extend into the neighboring dimer (Tiffany et al., 1997). Moreover, it was shown that even small deletions at the C terminus (four and more amino acids) dramatically reduce the stability of the protein and result in an almost complete loss of the activity. Replacement of the C-terminal nine amino acids of the human IVD by the respective sequence of medium-chain ACDH completely inactivates the enzyme, although this mutated protein was at least stable in vitro. These results clearly demonstrate the significance of the C terminus of the human IVD both for quaternary structure and enzyme activity (Volchenboum and Vockley, 2000). Because the last six amino acid residues differ significantly between the Arabidopsis IVD and both human and rat homologs, this part of the protein may be responsible for the altered quaternary structure and for the altered substrate specificity of the plant IVD.
Tissue-Dependent Suc Repression of Ativd Gene Expression in Seedlings
It has been reported previously that Leu is metabolized during germination of maize, soybean, and castor bean seeds. This amino acid is recruited from the breakdown of storage proteins and accumulates significantly during post-germination in cotyledons of soybean seedlings (Anderson et al., 1998). In castor bean, gluconeogenesis from Leu was observed in the endosperm of germinating seeds indicating the importance of Leu as a carbon source (Stewart and Beevers, 1967; Sodek and Wilson, 1973). Although we have not observed a strong transcription of the ivd gene in early developmental stages, our results of the transcription analysis of the ivd gene point in the same direction as these biochemical observations. Initially expressed at rather low levels, ivd mRNA accumulation increases with highest expression in 2- to 3-week-old seedlings (Fig. 6, G and H) and drops to a very reduced transcription level in adult plants (data not shown). This temporal transcription pattern resembles those of MCCase subunits A and B, whose mRNA steady-state levels increase until d 24 after planting and decrease significantly after d 27. Also, the spatial expression patterns are very similar between these MCCase genes and the ivd, which were both expressed in almost all cell types, at least in seedlings grown in the absence of Suc (McKean et al., 2000).
The availability of Suc, which serves as carbon source, strongly reduces the expression of the ivd gene, suggesting a potential major role of the Leu carbon skeleton as an alternative carbon source. Similar observations have also been made during the carbohydrate starvation of sycamore cells. In the absence of Suc, Leu accumulates after Asn to the second most abundant amino acid released from a massive breakdown of proteins and a significant increase in MCCase activity was observed (Génix et al., 1990; Aubert et al., 1996). The accumulation of transcripts of the E1β and E2 subunits of the branched-chain α-ketoacid dehydrogenase, an enzyme catalyzing the second step in Leu catabolism, is in Arabidopsis similarly inhibited by Suc (Fujiki et al., 2000a, 2000b). This is consistent with a general down-regulation of genes coding for enzymes that are part of the Leu catabolic pathway by Suc. In addition, our results show that Suc repression of ivd expression is tissue dependent because strong repression was observed in cotyledons and first true leaves, but not in root tissues. High expression of ivd in root tissues even in the presence of exogenous Suc may be related to the existence and biochemical requirement of proteolytic processes during normal root development.
MATERIALS AND METHODS
Protoplast Preparation and Transformation
Using primers IVDS3.H (5′-TAAGGATCCTATGCAGACGTTTTTCTC) and IVDGFP.R (5′-GAGGGATCCTATCTTGCGCAAAC), a 164-nucleotide-long Ativd cDNA fragment was amplified in a PCR on the complete Ativd cDNA as DNA template. The reaction was carried out with KlenTaq polymerase using a buffer supplied by the manufacturer (CLONTECH, Palo Alto, CA). Cycles of 1 min at 94°C, 1.5 min at 52°C, and 1 min at 68°C were repeated 30 times. The PCR fragment was digested with BamHI and cloned into the respective site of vector psmGFP4 containing the smGFP4 reading frame (Davis and Viestra, 1998). Correct assembly of the reading frame in the resulting plasmid pAtivd:smgfp4 was inspected by sequence analysis. The plasmid for expression of the CPN-60:mGFP5 fusion protein was kindly provided by David C. Logan (St Andrews; Logan and Leaver, 2000). About 500,000 protoplasts were prepared and transformed with 50 μg of DNA of clones pAtivd:smgfp4 and pcpn60:gfp in the presence of polyethylene glycol as described by Koop et al. (1996). MitoTracker staining was performed following the protocol given by the manufacturer (Molecular Probes, Eugene, OR).
Fluorescence Microscopy
GFP and MitoTracker Red fluorescence was visualized with a Carl Zeiss Axioplan I microscope equipped with flourescein isothiocyanat (HQ 470/40/HQ 500 LP) and MitoTracker (HQ545/30/HQ 610/75) filter sets obtained from AHF (Tübingen, Germany). Images were taken using the axiovision software (Carl Zeiss, Oberkochen, Germany).
Preparation and Sub-Fractionation of Mitochondria
Mitochondria from pea (Pisum sativum) seedlings and Arabidopsis suspension culture were isolated as described by Binder and Brennicke (1993) and Klein et al. (1998), respectively. Pea mitochondria were lysed and subfractionated into membrane and matrix fractions by sonification and ultracentrifugation following protocols given by Linke and Weiss (1986). Western blotting was performed as described elsewhere (Däschner et al., 1999). Selective protein detection with different antibodies was done with a Vectastain kit as recommended by the manufacturer (Vector Laboratories, Burlingame, CA).
BN-PAGE and Gel-Filtration Chromatography
Arabidopsis mitochondrial protein was two dimensionally separated by BN-PAGE and SDS-PAGE as described elsewhere (Jänsch et al., 1996). Proteins were blotted onto polyvinylidene difluoride membranes and the IVD was selectively detected as described above. A Superdex 75 column (Amersham Pharmacia Biotech, Freiburg, Germany) was used for size determination of native IVD. Pea mitochondria were resuspended in a buffer containing 200 mm Tris-HCl (pH 7.3), 25 mm KCl, 1 mm phenylmethylsulfonyl fluoride, and 1 mm EDTA and were sonicated for 3 × 10 s and 1 × 20 s. Non-solubilized debris was pelleted by a centrifugation step at 150,000g for 90 min at 4°C and the supernatant containing about 800 μg mitochondrial protein was loaded onto the column. Escherichia coli cells overexpressing AtIVD were lysed by sonification for 60 s interrupted by two short breaks. Cell debris was collected with a centrifugation at 20,000g for 20 min at 4°C. About 2.7 mg of total E. coli protein was loaded onto the column. Elution was carried out at a flow rate of 0.5 mL min−1 and fractions of 0.42 mL were collected. Eluted proteins were analyzed with a human IVD-specific antibody as described above. The Superdex 75, with a size exclusion of 100 kD, was calibrated with dextran blue, aldolase (158 kD), bovine serum albumine (65.2 kD), ovalbumine (43 kD), chymotrypsinogen A (25 kD), and ribonuclease A (13.7 kD).
Overexpression of AtIVD in E. coli and Substrate Specificity Tests
The Ativd cDNA representing the putative mature protein without the 25-amino-acid presequence, as deduced from N-terminal sequencing of the respective pea protein, was amplified by PCR (1 min at 94°C, 1 min at 54°C, and 1.5 min at 68°C) with primers IEX.H (5′-TCTCCCGGGTCTTCGTCTCTCC) and IQE.R (5′-AGAGTCGACTCTATTCTTCTTTGAAAAGCTC) using KlenTaq polymerase according to the supplier's instructions (CLONTECH). After digestion with SmaI and SalI, the DNA fragment was cloned into the respective sites of expression vector pAL-781 (Invitrogen, Groningen, The Netherlands). Overexpression was performed following the instructions given by the manufacturer. ACDH activities were assayed in terms of the reduction of 2,6-dichloroindophenol as electron acceptor and phenazine methosulfate as intermediate electron carrier (Izai et al., 1992). Reactions were carried out under the conditions described by Bode et al. (1999) in the presence of either 50 or 100 μm flavin adenine dinucleotide. The reaction was started by the addition of the acyl-CoA substrate. Reduction was followed spectrophotometrically by the decrease in absorption at 600 nm. Straight-chain, isobutyryl-CoA, and isovaleryl-CoA were purchased from Sigma (St. Louis). 2-Methyl-hexanoyl-CoA and 2-methyl-palmitoyl-CoA were synthesized as described by Vanhove et al. (1991). Activities were linear with respect to time for at least 4 min and were proportional to the amount of protein added to the assay. Blanks in the absence of protein showed no rate of reduction by contaminating coenzyme A-thioester. Data of enzyme activity versus substrate concentration were analyzed by linearization according to the Lineweaver-Burk method and subsequent linear regression.
Plant Transformation and GUS Staining
Primers IVDP2.H (5′-AAACCCGGGTGTGAATGAGG) and IVDP.R (5′-CTGCCCGGGCATATCTTCGTTATTAC) were used to amplify a 1.8-kb upstream region of the ivd gene from total Arabidopsis DNA. The amplification reaction was performed using KlenTaq polymerase (CLONTECH) in a buffer supplied by the manufacturer under the following conditions: five cycles of 1 min at 94°C, 1 min at 47°C, and 2.5 min at 68°C; and 30 cycles of 1 min at 94°C, 1 min at 45°C, and 2.5 min at 68°C. The PCR fragment was digested with SmaI and cloned into the respective site in vector pBecks19/101.3 generated by cloning the GUS expression cassette of pBI101.3 into the EcoRI site of pBecks19 (McCormac et al., 1997). After transformation into Agrobacterium tumefaciens strain GV2260 Arabidopsis plants were transformed by vacuum infiltration (Bechtold et al., 1993). GUS staining of different tissues of several individual transgenic Arabidopsis lines was performed as described elsewhere (Hull and Devic, 1995). Plant were grown on MS medium containing Gamborg vitamins with or without 0.5% (w/v) Suc with a 16-h photoperiod at 22°C or on soil in the greenhouse.
Miscellaneous Methods
All routine methods of molecular biology were performed following protocols given by Sambrook et al. (1989). Northern analysis was performed using the digoxigenin detection system according to protocols given by the supplier (Roche Diagnostics, Mannheim, Germany). DNA sequencing reactions were carried out with the Thermo Sequenase fluorescent labeling kit (Amersham Pharmacia Biotech) following the instructions given by the manufacturer.
ACKNOWLEDGMENTS
We are very grateful to Drs. David C. Logan (University of St. Andrews, UK) and Christopher J. Leaver (University of Oxford, UK) for the gift of a clone encoding the CPN-60:mGFP5 fusion protein (pBINmgfp5-cpn60). For the kind gift of antibodies we thank Drs. Christine Gietl (Technische Universität Munich, Munchen, Germany; anti-MDH), Hans-Peter Braun, Universität Hannover, Germany; anti-porin), and R. Jerry Vockley (Mayo Clinic and Mayo Foundation, Rochester, MN; anti-IVD). The generous donation of membrane carrying two dimensionally separated Arabidopsis proteins from the Braun lab is gratefully acknowledged. We also thank Holger Loose, Waltrand Kofer, and Ulrich Koop (Ludwig-Maximilians-Universität Munchen, Germany) for their help with protoplast preparation and transformation. 2-Methyl-hexanoyl-CoA and 2-methyl-palmitoyl-CoA were kind gifts from Prof. Guy P. Mannaerts and Prof. Paul P. Van Veldhoven (University of Leuven, Belgium). We also thank Claudia Gautsch (Universität Ulm, Germany) for excellent technical assistance.
Footnotes
This work was supported by the Fonds der Chemischen Industrie and by the Anfangsförderung der Universität Ulm.
LITERATURE CITED
- Alban C, Baldet P, Axiotis S, Douce R. Purification and characterization of 3-methylcrotonyl-coenzyme A carboxylase from higher plant mitochondria. Plant Physiol. 1993;102:957–965. doi: 10.1104/pp.102.3.957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anderson MD, Che P, Song J, Nikolau BJ, Wurtele ES. 3-Methylcrotonyl-coenzyme A carboxylase is a component of the mitochondrial leucine catabolic pathway in plants. Plant Physiol. 1998;118:1127–1138. doi: 10.1104/pp.118.4.1127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aubert S, Alban C, Bligny R, Douce R. Induction of β-methylcrotonyl-coenzyme A carboxylase in higher plant cells during carbohydrate starvation: evidence for a role of MCCase in leucine catabolism. FEBS Lett. 1996;383:175–180. doi: 10.1016/0014-5793(96)00244-x. [DOI] [PubMed] [Google Scholar]
- Bechtold N, Ellis J, Peletier G. In planta Agrobacterium mediated gene transfer by infiltration of adult Arabidopsis thaliana plants. C R Acad Sci. 1993;316:1194–1199. [Google Scholar]
- Binder S, Brennicke A. Transcription initiation sites in mitochondria of Oenothera berteriana. J Biol Chem. 1993;268:7849–7855. [PubMed] [Google Scholar]
- Bode K, Hooks MA, Couée I. Identification, separation and characterization of acyl-coenzyme A dehydrogenase involved in mitochondrial β-oxidation in higher plants. Plant Physiol. 1999;119:1305–1314. doi: 10.1104/pp.119.4.1305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brouquisse R, James F, Pradet A, Raymond P. Asparagine metabolism and nitrogen distribution during protein degradation in sugar-starved maize root tips. Planta. 1992;188:384–395. doi: 10.1007/BF00192806. [DOI] [PubMed] [Google Scholar]
- Däschner K, Thalheim C, Guha C, Brennicke A, Binder S. In plants a putative isovaleryl-CoA is located in mitochondria. Plant Mol Biol. 1999;39:1275–1282. doi: 10.1023/a:1006129220778. [DOI] [PubMed] [Google Scholar]
- Davis SJ, Viestra RD. Soluble, highly fluorescent variants of the green fluorescent protein (GFP) for use in higher plants. Plant Mol Biol. 1998;36:521–528. doi: 10.1023/a:1005991617182. [DOI] [PubMed] [Google Scholar]
- Fujiki Y, Ito M, Nishida I, Watanabe A. Multiple signaling pathways in gene expression during sugar starvation. Pharmacological analysis of din gene expression in suspension culture cells of Arabidopsis. Plant Physiol. 2000b;124:1139–1147. doi: 10.1104/pp.124.3.1139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiki Y, Sato T, Ito M, Watanabe A. Isolation and characterization of cDNA clones for the E1β and E2 subunits of the branched-chain α-ketoacid dehydrogenase complex in Arabidopsis. J Biol Chem. 2000a;275:6007–6013. doi: 10.1074/jbc.275.8.6007. [DOI] [PubMed] [Google Scholar]
- Génix P, Bligny R, Martin JB, Douce R. Transient accumulation of asparagine in sycamore cells after a long period of sucrose starvation. Plant Physiol. 1990;94:717–722. doi: 10.1104/pp.94.2.717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerbling H, Gerhardt B. Oxidative decarboxylation of branched-chain 2-oxo fatty acids by higher plant peroxisomes. Plant Physiol. 1988;88:13–15. doi: 10.1104/pp.88.1.13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerbling H, Gerhardt B. Peroxisomal degradation of branched-chain 2-oxo acids. Plant Physiol. 1989;91:1387–1392. doi: 10.1104/pp.91.4.1387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayashi H, De Bellis L, Ciurli A, Kondo M, Hayashi M, Nishimura M. A novel acyl-CoA oxidase that can oxidise short-chain acyl-CoA in plant peroxisomes. J Biol Chem. 1999;274:12715–12721. doi: 10.1074/jbc.274.18.12715. [DOI] [PubMed] [Google Scholar]
- Hull GA, Devic M. The β-glucuronidase (gus) reporter gene system. In: Jones J, editor. Plant Gene Transfer and Expression Protocols, Methods in Molecular Biology. Vol. 49. Totowa, NJ: Humana Press Inc; 1995. pp. 125–141. [DOI] [PubMed] [Google Scholar]
- Ikeda Y, Dabrowski C, Tanaka K. Separation and properties of five distinct acyl-CoA dehydrogenases from rat liver mitochondria. J Biol Chem. 1983;258:1066–1076. [PubMed] [Google Scholar]
- Ikeda Y, Tanaka K. Purification and characterization of isovaleryl-coenzyme A dehydrogenase from rat liver mitochondria. J Biol Chem. 1983a;258:1077–1085. [PubMed] [Google Scholar]
- Ikeda Y, Tanaka K. Purification and characterization of 2-methyl-branched-chain acyl-coenzyme A dehydrogenase, an enzyme involved in the isoleucine and valine metabolism, from rat liver mitochondria. J Biol Chem. 1983b;258:9477–9487. [PubMed] [Google Scholar]
- Izai K, Uchida Y, Orii T, Yamamoto S, Hashimoto T. Novel fatty acid β-oxidation enzymes in rat liver mitochondria. J Biol Chem. 1992;267:1027–1033. [PubMed] [Google Scholar]
- Jänsch L, Kruft V, Schmitz UK, Braun HP. New insights into the composition, molecular mass and stoichiometry of the protein complexes of plant mitochondria. Plant J. 1996;9:357–368. doi: 10.1046/j.1365-313x.1996.09030357.x. [DOI] [PubMed] [Google Scholar]
- Klein M, Binder S, Brennicke A. Purification of mitochondria from Arabidopsis. In: Martinez-Zapater JM, Salinas J, editors. Arabidopsis Protocols, Methods in Molecular Biology. Vol. 82. Totowa, NJ: Humana Press Inc.; 1998. pp. 49–53. [DOI] [PubMed] [Google Scholar]
- Koop HU, Steinmüller K, Wagner H, Rossler C, Eibl C, Sacher L. Integration of foreign sequences into the tobacco plastome via polyethylene glycol-mediated protoplast transformation. Planta. 1996;199:193–201. doi: 10.1007/BF00196559. [DOI] [PubMed] [Google Scholar]
- Linke P, Weiss H. Reconstitution of ubiquinol-cytochrome-c reductase from Neurospora mitochondria with regard to subunits I and III. Methods Enzymol. 1986;126:201–210. doi: 10.1016/s0076-6879(86)26022-x. [DOI] [PubMed] [Google Scholar]
- Logan DC, Leaver CJ. Mitochondria-targeted GFP highlights the heterogeneity of mitochondrial shape, size and movement within living plant cells. J Exp Bot. 2000;51:865–871. [PubMed] [Google Scholar]
- Masterson C, Wood C. Mitochondrial β-oxidation of fatty acids in higher plants. Physiol Plant. 2000;109:217–224. [Google Scholar]
- McCormac AC, Elliott MC, Chen DF. pBecks –A flexible series of binary vectors for Agrobacterium-mediated plant transformation. Mol Biotechnol. 1997;8:199–213. doi: 10.1007/BF02760773. [DOI] [PubMed] [Google Scholar]
- McKean AL, Ke J, Song J, Che P, Achenbach S, Nikolau BJ, Wurtele ES. Molecular characterization of the non-biotin-containing subunit of 3-methylcrotonyl-CoA carboxylase. J Biol Chem. 2000;275:5582–5590. doi: 10.1074/jbc.275.8.5582. [DOI] [PubMed] [Google Scholar]
- Sambrook J, Fritsch EF, Maniatis T. Molecular Cloning: A Laboratory Manual. Ed 2. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- Singh BK. Biosynthesis of valine, leucine and isoleucine. In: Singh BK, editor. Plant Amino Acids: Biochemistry and Biotechnology. New York: Marcel Dekker; 1999. pp. 227–247. [Google Scholar]
- Singh BK, Shaner DL. Biosynthesis of branched-chain amino acids: from test tube to field. Plant Cell. 1995;7:935–944. doi: 10.1105/tpc.7.7.935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sodek L, Wilson CM. Metabolism of lysine and leucine derived from storage protein during the germination of maize. Biochim Biophys Acta. 1973;304:353–362. doi: 10.1016/0304-4165(73)90253-5. [DOI] [PubMed] [Google Scholar]
- Song J, Wurtele ES, Nikolau BJ. Molecular cloning and characterization of the cDNA for the biotin-containing subunit of 3-methylcrotonyl-coenzyme A carboxylase: identification of the biotin carboxylase and biotin-carrier domains. Proc Natl Acad Sci USA. 1994;91:5779–5783. doi: 10.1073/pnas.91.13.5779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stewart CR, Beevers H. Gluconeogenesis from amino acids in germinating castor bean endosperm and its role in transport to the embryo. Plant Physiol. 1967;42:1587–1595. doi: 10.1104/pp.42.11.1587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tiffany KA, Roberts DL, Wang M, Paschke R, Mohlsen A-WA, Vockley J, Kim J-JK. Structure of human isovaleryl-CoA dehydrogenase at 2.6 Å resolution: structural basis for substrate specificity. Biochemistry. 1997;36:8455–8464. doi: 10.1021/bi970422u. [DOI] [PubMed] [Google Scholar]
- Vanhove G, Van Veldhoven PP, Vanhoutte F, Parmentier G, Eyssen HJ, Mannaerts GP. Mitochondrial and peroxisomal beta-oxidation of the branched chain fatty acid 2-methylpalmitate in rat liver. J Biol Chem. 1991;266:24670–24675. [PubMed] [Google Scholar]
- Volchenboum SL, Vockley J. Mitochondrial import and processing of wild type and type III mutant isovaleryl-CoA dehydrogenase. J Biol Chem. 2000;275:7958–7963. doi: 10.1074/jbc.275.11.7958. [DOI] [PubMed] [Google Scholar]
- Wang X, Wurtele ES, Keller G, McKean AL, Nikolau BJ. Molecular cloning of cDNAs and genes coding for β-methylcrotonyl-coenzyme A carboxylase of tomato. J Biol Chem. 1994;269:11760–11769. [PubMed] [Google Scholar]
- Weaver LM, Lebrun L, Franklin A, Huang L, Hoffman N, Wurtele ES, Nikolau BJ. Molecular cloning of the biotinylated subunit of 3-methylcrotonyl-coenzyme A carboxylase of Arabidopsis thaliana. Plant Physiol. 1995;107:1013–1014. doi: 10.1104/pp.107.3.1013. [DOI] [PMC free article] [PubMed] [Google Scholar]