Abstract
Firefly luciferase is a member of the acyl-adenylate/thioester-forming superfamily of enzymes and catalyzes the oxidation of firefly luciferin with molecular oxygen to emit light. Knowledge of the luminescence mechanism catalyzed by firefly luciferase has been gathered, leading to the discovery of a novel catalytic function of luciferase. Recently, we demonstrated that firefly luciferase has a catalytic function of fatty acyl-CoA synthesis from fatty acids in the presence of ATP, Mg2+ and coenzyme A. Based on identification of fatty acyl-CoA genes in firefly, Drosophila, and non-luminous click beetles, we then proposed that the evolutionary origin of firefly luciferase is a fatty acyl-CoA synthetase in insects. Further, we succeeded in converting the fatty acyl-CoA synthetase of non-luminous insects into functional luciferase showing luminescence activity by site-directed mutagenesis.
Keywords: Bioluminescence, Luciferin analogue, Inhibitors, Fatty acyl-CoA synthetase, Molecular evolution
Introduction
The phenomenon of bioluminescence has been observed in many different organisms including bacteria, fungi, algae, fish, squid, shrimp and insects [1–5]. Luminous organisms produce light by an enzymatic reaction of a luciferase (=enzyme) with a luciferin (=substrate). Light-emitting reactions are quite distinct among luminous organisms, but in each case the reaction is an oxidation process with molecular oxygen and is a conversion of chemical energy into light [6–9]. The various luciferase genes have been isolated and adapted as reporter genes [10, 11]. In insects, the luminous species are mainly found in three families: fireflies (Lampyridae), railroad worms (Phengodidae) and click beetles (Elateridae) [12–14]. The bioluminescence systems of these insects are essentially the same with an identical luciferin, ATP, and Mg2+, and with similar luciferases. Firefly luciferase has been studied for the last 50 years and proven to be a useful enzyme. It has been extensively used in molecular and cell biology, in particular for the quantification of ATP and as a reporter enzyme of gene expression [10, 11]. The purpose of this review is to reevaluate the basic information on firefly luciferase and luciferin including the luminescence reaction, and to reconsider the catalytic function of firefly luciferase and other unresolved problems, based on recent investigations. Further, the molecular evolution of firefly luciferase from a fatty acyl CoA synthetase in insects is described on the basis of our recent reports.
Chemical basis of firefly bioluminescence
Bioluminescence reaction of firefly luciferase
Firefly luciferase (EC. 1.13.12.7) is classified as a mono-oxygenase and catalyzes the emission of yellow-green light from the substrate d-luciferin, Mg2+-ATP, and oxygen, according to the following equations:
 |
1 |
 |
2 |
 |
3 |
The initial reaction catalyzed by firefly luciferase (Luc) is the formation of luciferase-bound luciferyl adenylate (Luc:LH2-AMP) in the presence of Mg2+ and ATP by the release of inorganic pyrophosphate (PPi) Eq. 1. The carboxyl group of d-luciferin (LH2) is adenylated. The second step involves the oxygenation of LH2-AMP with molecular oxygen (O2) to produce the excited state of oxyluciferin (Oxyluciferin*), adenosine monophosphate (AMP) and carbon dioxide (CO2) (Eq. 2). The emission of light is produced from the relaxation of excited state oxyluciferin to the corresponding ground state (Eq. 3). The luminescence reaction of firefly luciferin with other luciferases from luminous beetles and railroad worms is identical to that of firefly luciferase [13, 14].
On the other hand, firefly luciferase also catalyzes the formation of luciferyl-CoA (LH2-CoA) from LH2-AMP in the presence of coenzyme A (CoA) under anaerobic conditions (Eq. 4).
 |
4 |
The thiol-ester of CoA is formed at the carboxyl group of luciferin. l-Luciferyl-CoA is formed from l-luciferin in the same process as LH2-AMP formation [15].
Dehydroluciferyl-CoA (L-CoA) is obtained from dehydroluciferin (L) in the presence of ATP, Mg2+ and CoA (Eqs. 5 and 6).
 |
5 |
 |
6 |
In 1967, on the basis of above results, McElroy et al. proposed that firefly luciferase is one of the family of acyl-CoA synthetases, similar to a fatty acid acyl-CoA synthetase and an amino-acyl tRNA synthetase [16]. However, nobody knew whether firefly luciferase can synthesize the fatty acyl-CoA from free fatty acids until we demonstrated it in 2003 [17].
The catalytic reaction of acyl-CoA synthetase is as follows:
 |
7 |
 |
8 |
Thus, firefly luciferase is a member of the acyl-adenylate/thioester-forming superfamily of enzymes [16].
Chemical structures of related compounds involved in the firefly luciferin–luciferase reaction
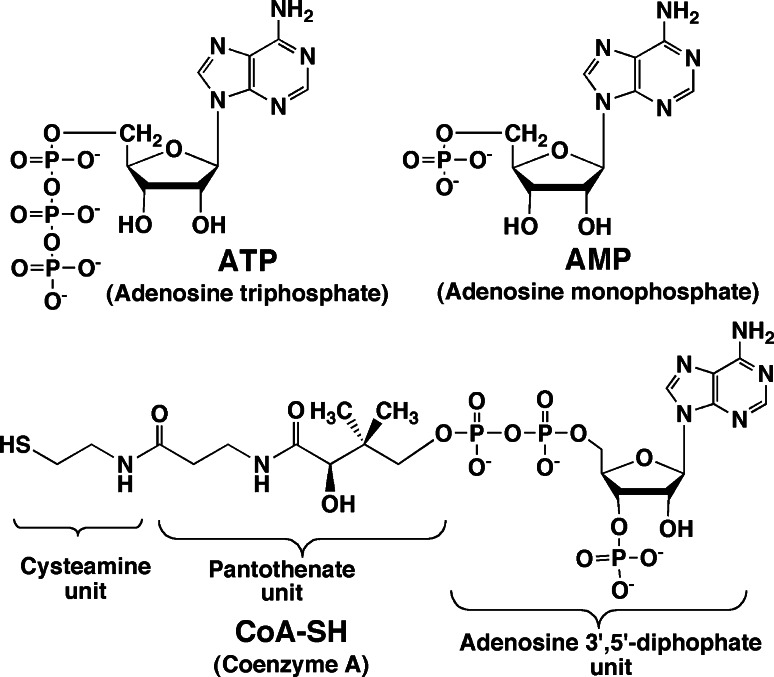
To understand the catalytic mechanism of firefly luciferase, it is necessary to know the chemical structures of ATP, AMP and coenzyme A (CoA) (Fig. 1). The chemical structures of firefly luciferin including its related compounds are shown in Fig. 2 and their properties are summarized as follows.
Fig. 1.
Chemical structures of ATP, AMP and coenzyme A (CoA)
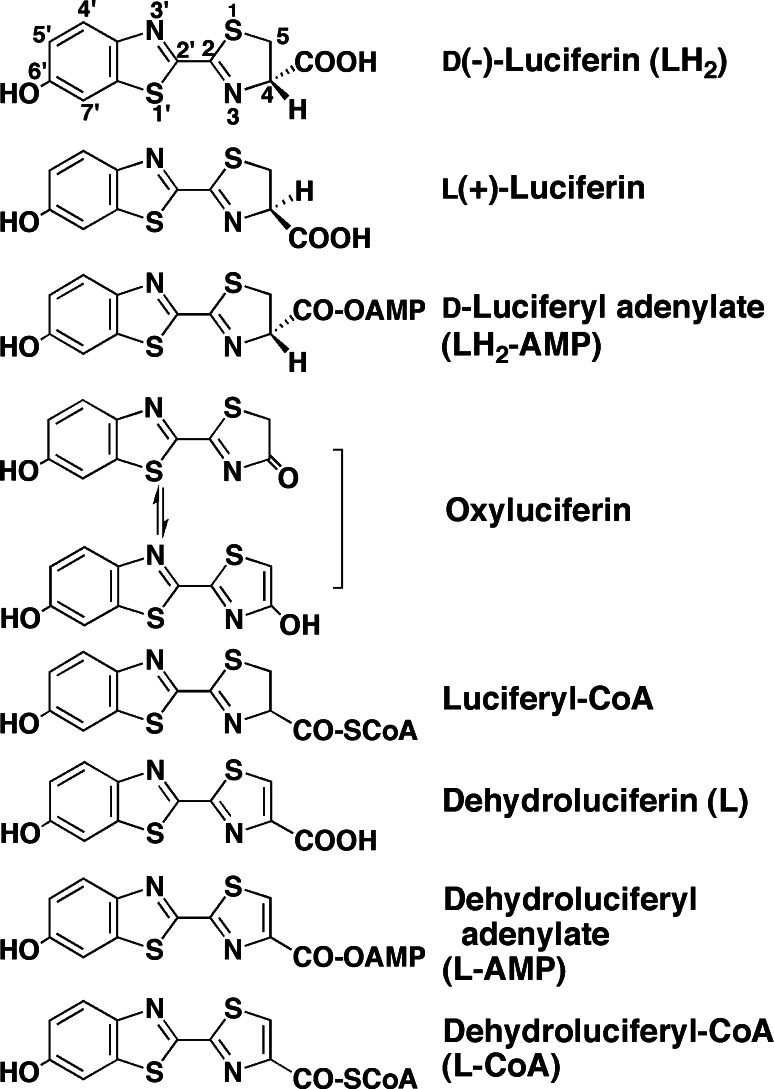
Fig. 2.
Chemical structures of firefly luciferin and its related compounds
Firefly luciferin = d(−)-luciferin (d-LH2)
In 1957, approximately 9 mg of crystalline firefly luciferin was isolated from 15,000 firefly lanterns (70 g of acetone powder) of the North American firefly Photinus pyralis [18]. From the Japanese firefly, Luciola cruciata, approximately 5.5 mg of luciferin was crystallized from 12,000 adult abdomens (233 g of acetone powder) [19]. The structure of firefly luciferin, D(−)-Luciferin, (S)-2-(6′-hydroxy-2′-benzothiazolyl)-2-thiazoline-4-carboxylic acid, was determined by chemical synthesis [20, 21]. The absolute configuration of the carboxyl group at the C4 position was also determined to be the S form. This chirality of luciferin is crucial to the luminescence reaction with luciferase. Thus, L(+)-luciferin is practically inactive for the luminescence reaction. The molecular configuration and confirmation of synthetic luciferin including intraatomic bond distances and angles were determined by X-ray structure analysis and were confirmed to be identical to the structure by chemical synthesis [22]. The K m value of recombinant P. pyralis luciferase for luciferin and ATP were 15 and 160 μM, respectively [23]. Recently, the contents of firefly luciferin in luminous and non-luminous beetles were re-examined and confirmed by the methods of HPLC with fluorescence detection and the luminescence reaction of luciferin–firefly luciferase (L–L reaction) [24]. In Japanese fireflies, the amount of luciferin is at the range of 0.5–3.5 nmol (140–980 ng) per specimen, and these values were similar to those previously reported [19]. On the other hand, no d-luciferin was detected in the non-luminous beetles, as far as was examined [24].
l(+)-Luciferin (l-LH2)
l-Luciferin is an enantiomer of d-luciferin and was synthesized in order to identify the chemical structure of natural d-luciferin [25]. In the standard reaction mixtures, l-luciferin is not used in the luminescence reaction [25]. l-Luciferin is a potent competitive inhibitor of luciferase and the Ki value is between 3 and 4 μM [26]. In 1996, weak light production from l-luciferin with luciferase was reported, and the mechanism of racemization from l-luciferin to d-luciferin through luciferyl adenylate formation was proposed [26]. In larvae of the Japanese firefly Luciola cruciata, approximately 1:1 mixtures of d- and l-luciferin were detected by an HPLC analysis using a chiral column [27]. Further, the extracts of light organs of L. cruciata showed a catalytic activity to convert l-luciferin to d-luciferin in the presence of ATP, Mg2+ and CoA [27]. However, these results did not explain the biosynthetic pathway of firefly luciferin and the synthesis process of luciferin was still unclear (see “Biosynthesis of firefly luciferin in vivo”).
Dehydroluciferin (L)
Dehydroluciferin was firstly identified as one of the products formed during the biological oxidation of luciferin. Dehydroluciferin is present in small quantities in the firefly and can be prepared from luciferin by chemical oxidation. l-Luciferin and d-luciferin can be converted to dehydroluciferin by oxidation with ferricyanide or heating in alkaline solution. Dehydroluciferin is known to be potent inhibitor of the luminescence reaction [18].
Oxyluciferin
In the early 1960s, dehydroluciferin referred to firefly oxyluciferin [25, 28, 29]. The chemical structure of oxyluciferin was proposed by the fluorescence spectral data of the synthetic compounds [30–33]. Oxyluciferin from the reaction mixture of the luciferin–luciferase reaction was only detected by HPLC analysis [34], because oxyluciferin is an unstable compound in solution. Oxyluciferin is a carbonyl compound with a hydrogen atom on its α carbon and shows a keto–enol tautomerism that is catalyzed by both acids and bases. The keto form and enol form of oxyluciferin in solution show red fluorescence and yellow-green fluorescence, respectively. The K i value of oxyluciferin for light production is 0.5–2 μM [35].
Luciferyl adenylate (LH2-AMP)
Luciferyl adenylates are formed from d-luciferin and l-luciferin by luciferase in the presence of ATP and Mg2+. The reversibility of this reaction has been demonstrated [28]. The K m values of native luciferase and recombinant luciferase for light production are 2.4 and 4.7 μM, respectively [36]. Luciferyl adenylate has been chemically synthesized and characterized. Luciferyl adenylate is extremely unstable in aqueous solution [25, 29, 37] and is hydrolyzed to luciferin and adenylic acid very rapidly at pH 8 [28, 37]. Luciferyl adenylate racemized easily in neutral to basic buffer [15, 37, 38]. On the other hand, d- and l-luciferyl adenylates emit a red chemiluminescence in an aprotic solvent upon addition of strong base [29]. d-Luciferyl adenylate is used for the luciferase reaction, but not l-luciferyl adenylate.
Dehydroluciferyl adenylate (L-AMP)
Dehydroluciferyl adenylate is an inhibitor of the luminescence reaction [28]. The K i value for light production is 0.0025–1.25 μM [35].
Mechanism of firefly luminescence reaction
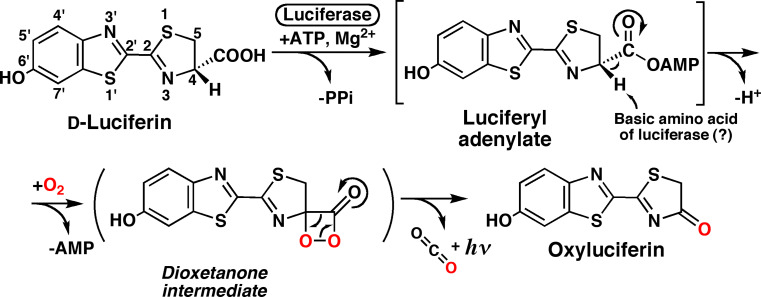
The mechanism of the bioluminescence reaction of firefly luciferin catalyzed by firefly luciferase is shown in Fig. 3. The luminescence reaction involves the formation of a dioxetanone intermediate [38]. The formation of dioxetanone was confirmed by 18O-labeling experiments [39]. Thus, one oxygen atom of molecular oxygen was incorporated into oxyluciferin and one into CO2. The decomposition of dioxetanone results in light emission from the excited state of oxyluciferin by the chemically initiated electron-exchange luminescence (CIEEL) mechanism [40, 41].
Fig. 3.
Mechanism of the bioluminescence reaction of firefly luciferin by firefly luciferase
Light emitter and color variation in firefly bioluminescence
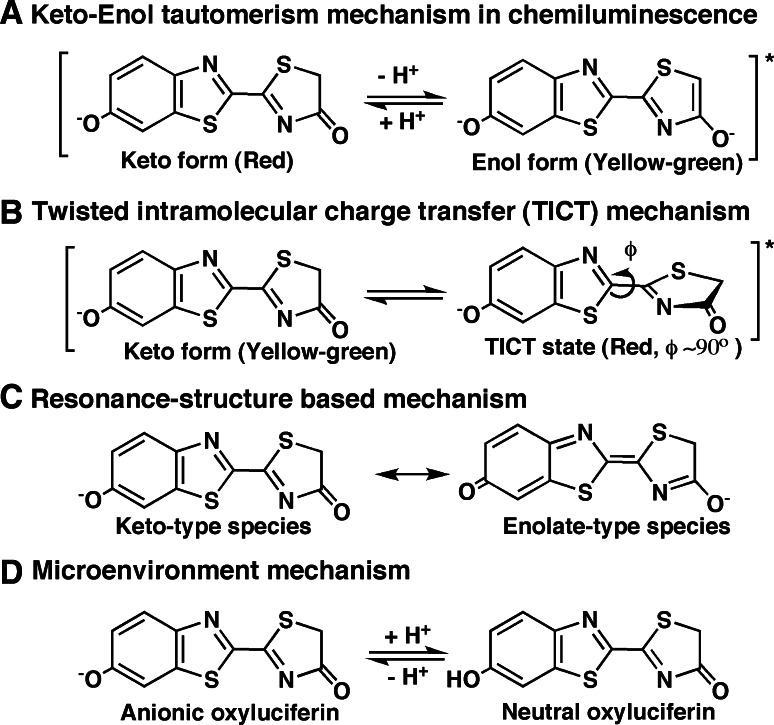
In P. pyralis luciferase in vitro, the luminescence color at pH 7.6 is yellow-green (λmax 565 nm), but a red luminescence (λmax 620 nm) is observed under acidic conditions at pH 5.4 [42]. To explain the multiple colors of firefly bioluminescence, the light emitter species of oxyluciferin have been proposed. Until early 1990, the relationship between the light emitter of oxyluciferin and the color of luminescence was explained by the keto–enol tautomerism (Fig. 4a). Under acidic and basic conditions, the fluorescence spectra showed red and yellow-green colors that corresponded to the keto form and the enol form of oxyluciferin, respectively [30–33]. To understand the species of light emitter, the fluorescence properties of the 5-methyl and O-methyl derivatives of oxyluciferin were investigated [43]. In 1991, a single amino acid mutant of L. cruciata luciferase, Ser286 to Asn286 (S286N), was prepared, which showed red bioluminescence (λmax 607 nm) with natural firefly luciferin. In addition, the different colors including red, orange-yellow and yellow-green were also produced by other luciferase mutants [44]. These results suggested that the light emitter could not be explained by the keto–enol tautomerism of oxyluciferin. Further, in 2002, d-5, 5-dimethylluciferin and its adenylate were chemically synthesized and examined as a substrate for the luciferase reaction. With the adenylate of d-5,5-dimethylluciferin as a substrate, the luciferases of the firefly (P. pyralis) and click beetle (Pyrophorus plagiophthalamus) gave yellow-green and red colors of luminescence, respectively [45]. This result suggested keto–enol tautomerism could not explain two colors of luminescence, since d-5,5-dimethyloxyluciferin has no hydrogen atom at the C5 position and cannot take the keto–enol tautomerism. Consequently, the twisted intramolecular charge transfer (TICT) mechanism was proposed [46, 47]. The benzothiazole and thiazolone rings in the oxyluciferin molecule are rotated around of the C2–C2′ bond (Fig. 4b), suggesting that different conformations of excited oxyluciferin give multicolor emission. Further, other mechanisms including the resonance-structure mechanism [48] (Fig. 4c) and microenvironment mechanism (Fig. 4d) have been proposed [48]. The discrepancy of emission color in the emitter structures of the keto form of oxyluciferin is preserved between Fig. 4a (red) and b (yellow-green). Several theoretical studies on the light emitter species of oxyluciferin combined with fluorescence spectral data of analogues have been reported [49–52]. However, the actual light emitter species in firefly bioluminescence is not yet conclusively known.
Fig. 4.
Proposed mechanisms for light emission in the firefly luciferase reaction
Luminescence pattern and emission spectrum of firefly luciferase
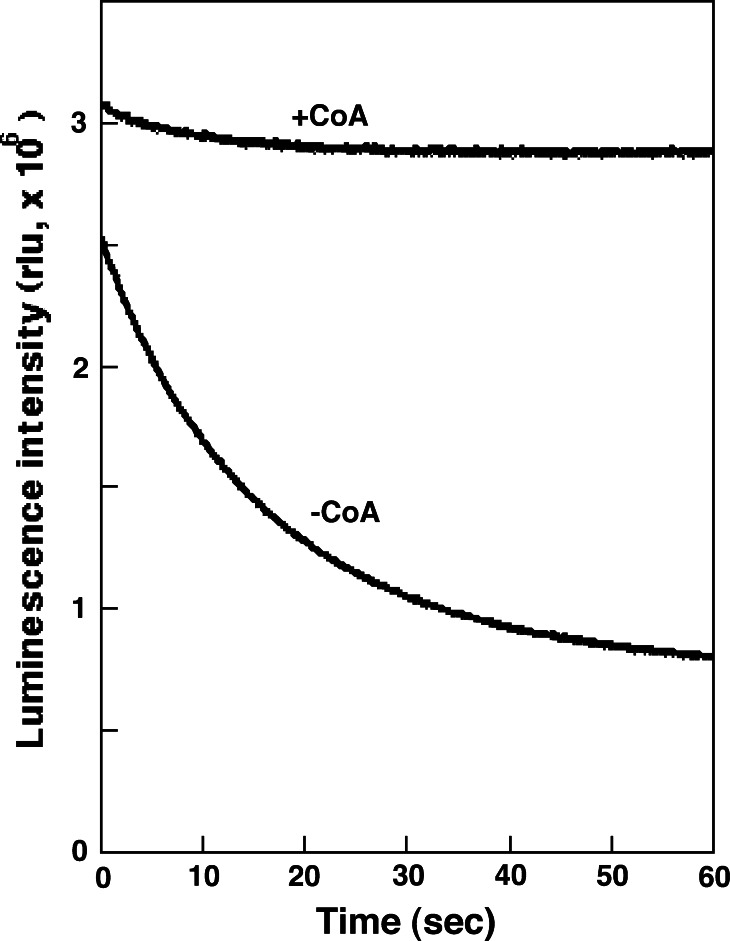
The kinetics of the firefly luciferase reaction in vitro are complicated, due to the fact that luciferin, ATP, Mg2+ and molecular oxygen are essential for the luminescence reaction [53]. Also, the luminescence pattern and the luminescence emission spectrum are affected by the reaction conditions such as pH, metal ion and temperature [54–57]. Further, the luminescent products including oxyluciferin and dehydroluciferin inhibit the luminescence reaction of firefly luciferase. As shown in Fig. 5, in the presence of saturating amounts of ATP, light emission by firefly luciferase rises quickly followed by a rapid decay, and then continuous weak-glow luminescence is observed. The rapid decay of light emission can be prevented by the addition of coenzyme A [58]. This luminescence enhancement may be explained by the formation of dehydroluciferyl-CoA, resulting in release of dehydroluciferin from the luciferase. This result suggests that firefly luciferase has CoA synthetic activity of dehydroluciferin through to dehydroluciferyl adenylate.
Fig. 5.
Luminescence pattern of firefly luciferase in the presence and absence of coenzyme A. The reaction mixture (100 μl) contained firefly luciferin (10 μM), ATP (250 μM) and MgCl2 (5 mM) in 100 mM Tris–HCl (pH 7.8). In the presence and absence of CoA (250 μM), the reaction was started by the addition of recombinant P. pyralis luciferase (1 pmol: Promega) and the light intensity of luminescence was measured at 22–23°C
Emission spectra with λmax between 546 and 618 nm were observed in various luciferases and their mutants [14, 59]. Some pH-sensitive and pH-insensitive luciferases have been reported, and several mutants of pH-sensitive luciferases were converted to pH-insensitive forms with a different color of light emission [14]. There are no clear explanations for these phenomena, because the emission spectrum of luciferase is dependent on the excitation species of oxyluciferin. Further, the quantum yield of the firefly luciferase reaction is dependent on the emission spectrum. In an early report, the quantum yield was estimated to be 0.88 ± 0.25 at pH 7.6 in Tris-buffer [42, 60]. Recently, the quantum yield was revised to 0.41 ± 0.074 at pH 8.5 in Tris-buffer [61]. To avoid similar historical confusion, it is necessary to confirm these values by other methods, and the quantum yields of other luciferases should be re-examined by the same procedures.
Firefly luciferase reaction with firefly luciferin
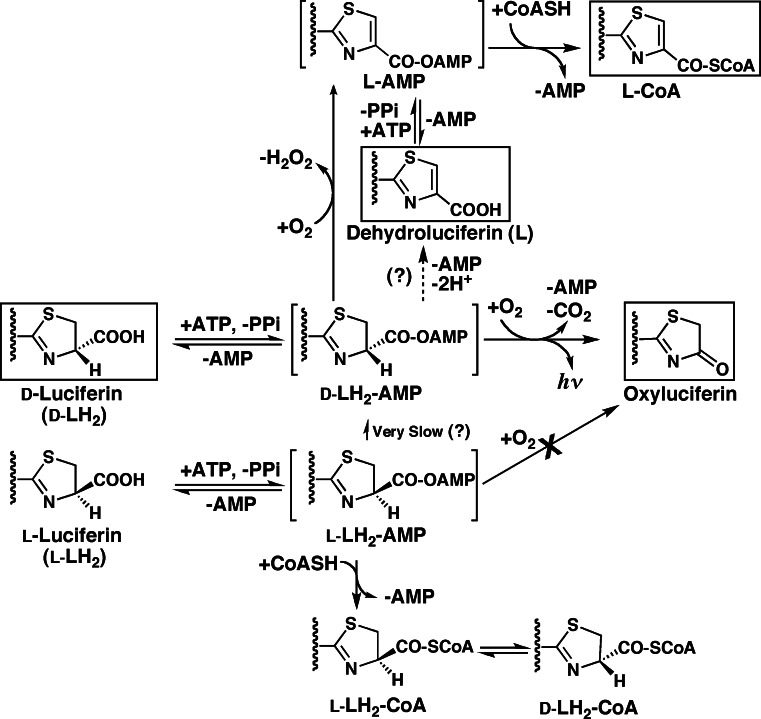
Under atmospheric conditions, the reaction scheme of firefly luciferin is summarized in Fig. 6, but the reaction catalyzed by firefly luciferase is still not completely described. In the firefly luciferase reaction (Fig. 3), the deprotonation and O2 addition at the C4 position of d-luciferyl adenylate is the limiting step to light emission. The racemization between d-LH2-AMP and l-LH2-AMP might not occur in the luciferase molecule, but in the solution non-enzymatically after releasing luciferyl adenylate from the luciferase molecule. d-Luciferyl adenylate bonded within luciferase reacts with O2 to emit light. Recently, the formation of dehydroluciferin from d-luciferin was proposed by reacting d-luciferyl adenylate with O2 to give H2O2 as a side reaction [62]. This reaction seems to be competitive with the luminescence reaction, and it is necessary to confirm the conversion from d-/l-luciferyl adenylate to dehydroluciferin in the presence and absence of firefly luciferase.
Fig. 6.
Formation of dehydroluciferin, dehydroluciferyl CoA and d-luciferyl CoA from d- and l-luciferin during the luminescence reaction by firefly luciferase
Biosynthesis of firefly luciferin in vivo
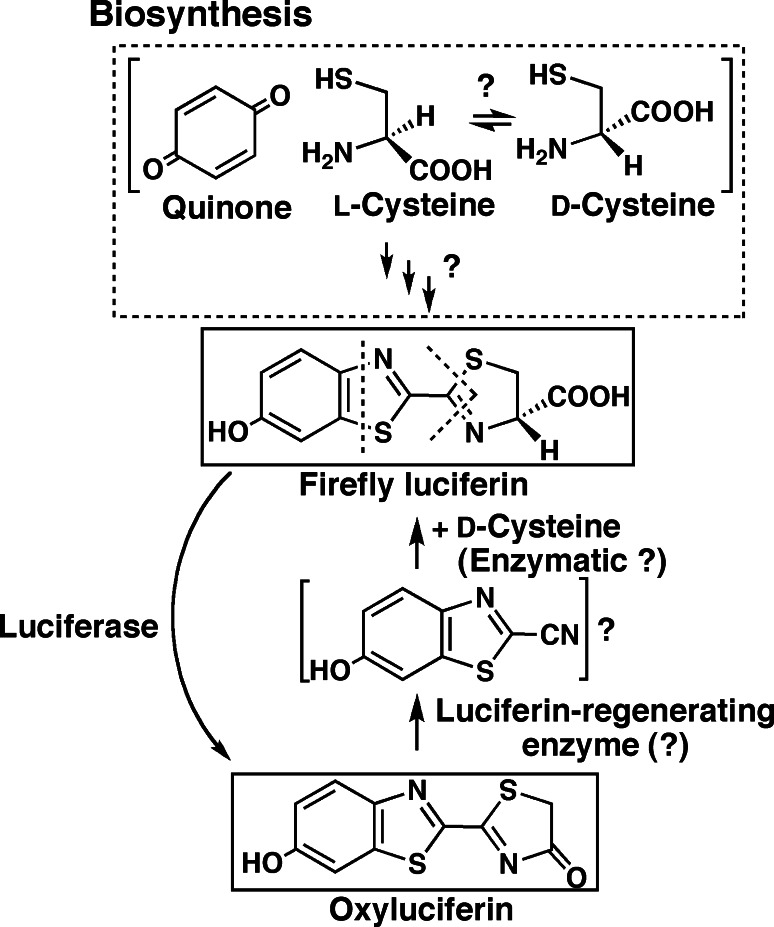
Regarding the biosynthesis of firefly luciferin in vivo, the synthetic pathway and precursor(s) are not solved. In the 1970s, studies on the biosynthesis of firefly luciferin were carried out by incorporation experiments using 14C-labeled compounds such as cysteine and quinone (Fig. 7). Biosynthetic pathways have been proposed, but based on insufficient information with inadequate experiments [63–66]. The only clear evidence is that cysteine is a synthetic unit of firefly luciferin [63, 66]. In 2001, a luciferin-regeneration enzyme of firefly luciferin from oxyluciferin was reported [67, 68]. Later, it was found that the protein has high amino acid sequence similarity to the senescence marker protein-30, that is a hydrolase, gluconolactonase (EC. 3.1.1.17). It is necessary to reconsider the previous results and re-examine these experiments [69, 70]. In some cases, mass spectral analysis using a stable isotope labeled compound could be useful, as reported in Cypridina luciferin biosynthesis [71, 72].
Fig. 7.
Biosynthesis of firefly luciferin and conversion to firefly luciferin form oxyluciferin
Luciferin analogues for luciferase substrate
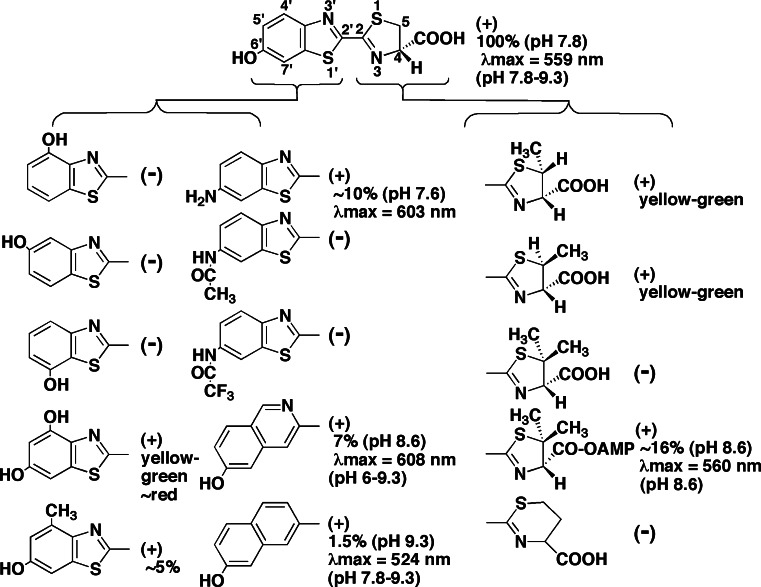
White et al. first synthesized the luciferin analogues in the 1960s [73–76] and showed narrow substrate specificity for luciferase. A convenient method of firefly luciferin synthesis was reported by Seto et al. [77]. The main structures of luciferin analogues and its luminescence activity are shown in Fig. 8. The luciferin esters of the hydroxyl group at the 6′ position of benzothiazole ring and the carboxyl group at the 4 position of thiazoline ring were not utilized for the luciferase reaction. Thus, the phenolic hydroxyl group of benzothiazole ring might interact with luciferase for the luminescence reaction. 6′-Amino-d-luciferin, a substituent of the hydroxyl group, showed lower luminescence activity (~10%) with red luminescence [74]. 4′-Methyl-d-luciferin showed ~5% activity [78]. N-Alkylated luciferins of 6′-aminoluciferin were synthesized and the luminescence activity was characterized [79]. d-Quinolylluciferin and d-naphthylluciferin, with substitutes of the benzothiazole ring with quinoly- and naphthyl-groups, showed 7 and 1.5% luminescence activities, respectively [80]. 5-Methylluciferin can be used for the luciferase reaction, but not 5,5-dimethylluciferin [76]. However, 5,5-dimethylluciferyl adenylate showed ~16% activity [45]. These findings by Branchini et al. gave insight into the light emitter of oxyluciferin (see "Light emitter and color variation in firefly bioluminescence", above).
Fig. 8.
Firefly luciferin analogues for the luciferase reaction and its luminescence activity. Plus and minus in parentheses show the positive and negative activities of luminescence as a substrate
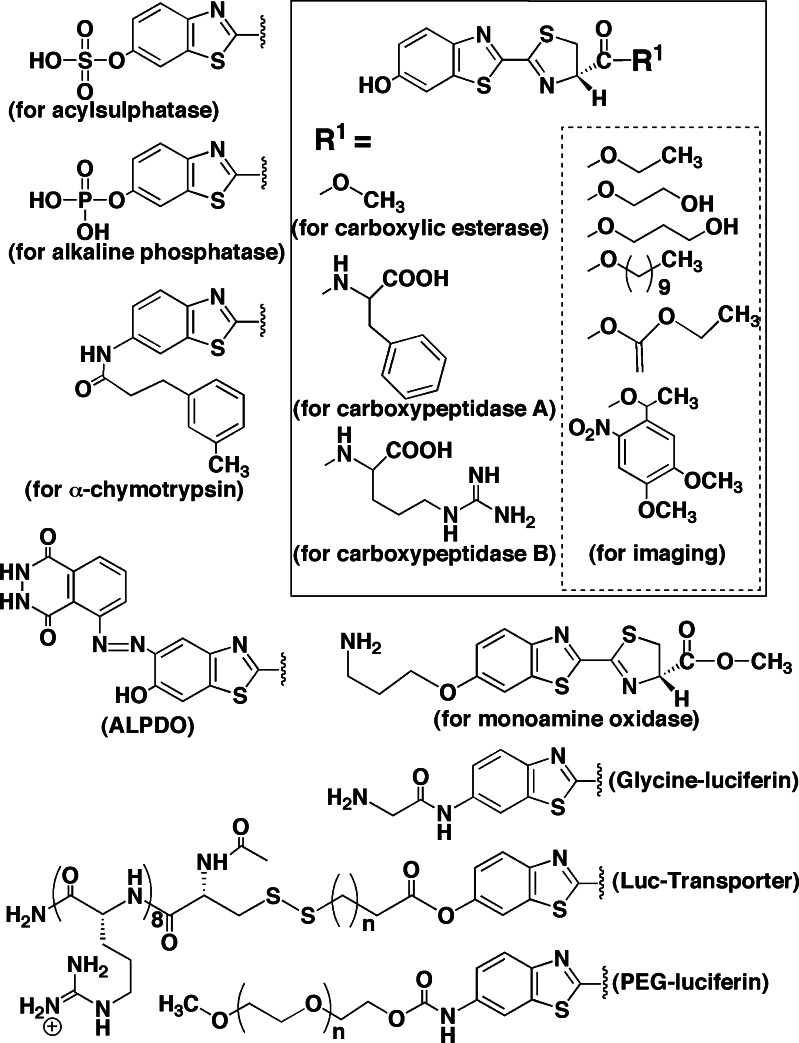
Luciferin derivatives for bioluminescent assays
To develop bioluminescence enhanced enzyme assays and immunoassays, derivatives of luciferin ester and amide have been synthesized. These enzymes hydrolyze the luciferin derivatives and the released d-luciferin is utilized for the firefly luciferase reaction. The derivatives are shown in Fig. 9. The sulfate and phosphate of d-luciferin were used as a substrate for acylsulphatase and alkaline phosphatase, respectively [81–85]. Also, d-luciferin glycosides were used to determine the glycohydrolase activity [86]. Further, the esters and amide at the carboxyl group of d-luciferin were used for carboxylic esterase and carboxypeptidase, respectively [81–83]. The amide of 6′-amino-d-luciferin was used for α-chymotrypsin assay [87]. A bifunctional luminogenic substrate (ALPDO) for both firefly luciferase and horseradish peroxidase was also reported [88]. To determine monoamine oxidase activity in a homogeneous assay system, acylic amine derivatives of luciferin were synthesized and applied [89, 90]. For bioluminescence imaging in mammalian cells using firefly luciferase as a reporter gene, membrane-permeable luciferin esters were reported [79, 91, 92], and 6′-amino-d-luciferin amides including glycine [93] and the transporter conjugate [94] were synthesized and applied in mice [95]. To improve the circulatory half-life time of luciferin for in vivo tumor imaging, 6′-amino-d-luciferin modified with polyethylene glycol (PEG-luciferin) was prepared [96].
Fig. 9.
Luciferin derivatives for bioluminescence assays using firefly luciferase
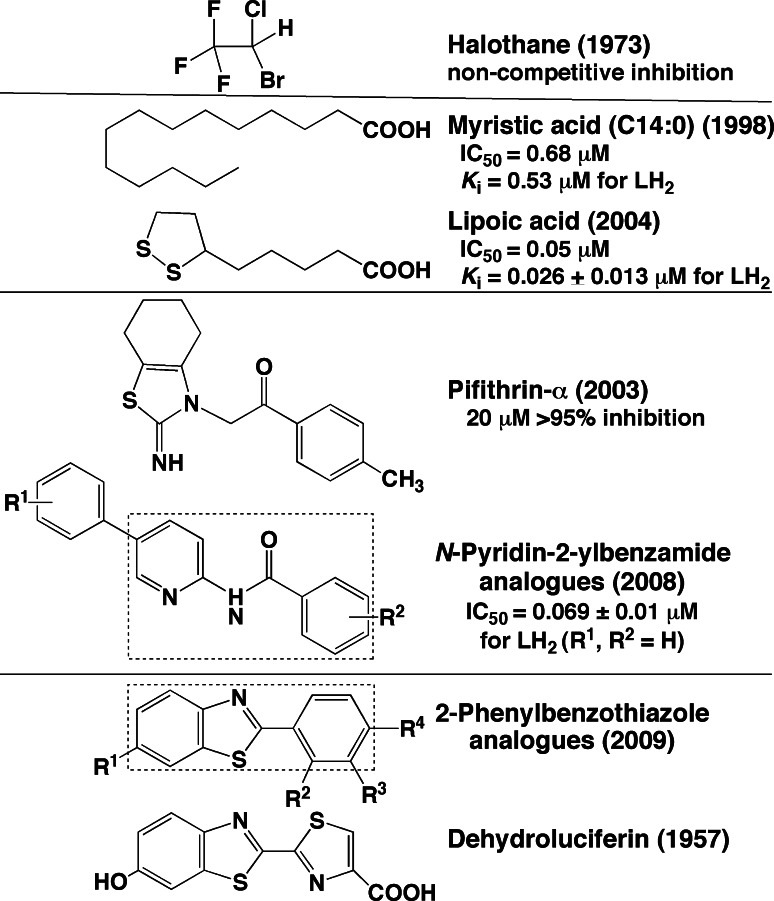
Inhibitors of firefly luciferase
Firefly luciferase has been used as a reporter protein in various assay systems including gene expression and was applied in the process of high-throughput screening for drug discovery. To avoid false positives in the chemical screening assays, it is valuable to know the inhibitors of the luciferin–luciferase reaction. Substrate-related compounds including dehydroluciferin [18], l-luciferin [26], oxyluciferin [97], pyrophosphate [98], AMP [99] and ATP analogues [100] were known as inhibitors. In contrast, after initiating the luminescence reaction with ATP, the addition of pyrophosphate and triphosphate stimulates light production under some conditions [57, 98]. As shown in Fig. 10, structurally unrelated compounds such as halothane [101], fatty acids [102, 103], pifithrin-α [104], lipoic acid [105], N-pyridin-2-ylbenzamide analogues [106], and 2-phenylbenzothiazole analogue [107] also inhibit luciferase activity. In cell-based luciferase reporter assays, the stabilization of luciferase by inhibitors results in the counterintuitive phenomenon of signal stimulation [108, 109]. Among competitive inhibitors, the luminescence inhibition of fatty acids reported by Ueda and Suzuki [102, 103] suggested to me that firefly luciferase has a catalytic function of fatty acyl-CoA synthesis.
Fig. 10.
Inhibitors of the firefly luciferase–luciferin reaction
Protein chemistry of firefly luciferase
Purification of firefly luciferase
In 1956, firefly luciferase was isolated from firefly lanterns of P. pyralis by gel filtration and was crystallized yielding 30 mg protein from ~6,000 specimens [53, 110]. Later, to purify native firefly luciferase more conveniently, several affinity chromatographies were performed using benzylamine [111] and Cibachoron Blue F3GA [112] as a ligand. The benzylamine group functions through hydrophobic interaction with luciferase, and 6.2 mg of luciferase was obtained from 2.2 g of acetone powder of lanterns. Cibachoron Blue F3GA (Blue Dextran, Blue Sepharose), which binds enzymes requiring adenylyl-type cofactors such as ATP, NADH and FAD, was applied for an affinity purification of luciferase. To remove adenylate kinases and prevent interference with ATP assays by firefly luciferase, Dyematrex Orange gel was used [113]. The Japanese firefly luciferases, L. cruciata and L. lateralis, were purified by ammonium sulfate precipitation, gel filtration, followed by hydroxyapatite HPLC [114] to afford ~0.2 mg protein from 1.5 g lanterns (wet weight). In a complementary approach, firefly luciferase gene was expressed in prokaryotic and eukaryotic cells. To express recombinant firefly luciferase in E. coli cells, the induction temperature is most effective when kept between 20 and 25°C to refold the active luciferase [115, 116]. Recently, firefly luciferase fused to IgG binding-domain of protein A was efficiently expressed in E. coli cells under the control of cold shock protein A promoter at 15°C [117].
Primary structure of firefly luciferase
The gene for P. pyralis luciferase was cloned [118] and the nucleotide sequences of cDNA and genomic DNA were determined in 1987 [119]. The primary structure of P. pyralis luciferase consists of 550 amino acid residues in a single polypeptide chain with a peroxisome targeting signal sequence of -Ser-Lys-Leu (-SKL) at C-terminus [120]. The calculated average molecular weight and theoretical pI were 60,745.17 and 6.42, respectively. Genomic sequence analysis of P. pyralis luciferase revealed that six short introns are present [119]. Interestingly, the identical positions of the introns were found in L. lateralis luciferase [121]. In 1989, the sequence similarity of firefly luciferase to plant 4-coumarate:CoA ligase (EC 6.2.1.12) was firstly reported [122] and the significant homology to rat long-chain acyl-CoA synthetase (EC 6.2.1.3) was also reported [123]. Further, the conserved motif sequences for the characteristic adenylate formation in the phosphate-binding sites of the phosphoproteins and the nucleotide-binding proteins were found by a homology search and it was proposed that acetyl-CoA synthetases were identified as members of firefly luciferase family [124]. The putative AMP-binding domain motif of [S/T/S]–[S/T/G]–G–[S/T]–[T/S/E]–[G/S]–x–[P/A/L/I/V/M]–K was found as the sequence of [198S–S–G–S–T–G–L–P–K206] [124]. After the cDNA clone of P. pyralis luciferase was isolated, many other firefly luciferase (beetle luciferase) genes including click beetle (Pyrophorus sp.) and railroad warm (Phrixothrix sp.) were readily cloned, sequenced, and characterized (Table 1). All luciferases are a single polypeptide chain and show high sequence identities among them with 48–99% identity [125, 126].
Table 1.
List of cDNA sequences of beetle luciferases from insects
| Insect species name | GenBank (Nucleotide) Accession no. | References |
|---|---|---|
| Photinus pyralis | M15077 | [119] |
| Luciola cruciata | M26194 | [139] |
| Luciola lateralis | X66919 | [140] |
| Luciola lateralis | Z49891 | [121] |
| Luciola mingrelica | S61961 | [115] |
| Photuris pennsylvanica | D25415 | [141] |
| Photuris pennsylvanica | U31240 | [125] |
| Pyrocoelia miyako | L39928 | [142] |
| Pyrocoelia rufa | AF328553 | [143] |
| Hotaria parvula | L39929 | [142] |
| Hotaria unmunsana | AF420006 | [144] |
| Lampyris noctiluca | X89479 | [145] |
| Lampyris turkestanicus | AY742225 | [146] |
| Cratomorphus distinctus | AY633557 | [147] |
| Lampyroidea maculata | DQ137139 | [148] |
| Pyrophorus plagiophthalamus | – | [149] |
| Pyrearinus termitilluminans | AF116843 | [150] |
| Pyrophorus plagiophthalamus | AF543368 | [151] |
| Pyrophorus mellifluus | AF545854 | [151] |
| Phrixothrix viviani | AF139644 | [152] |
| Phrixothrix hirtus | AF139645 | [152] |
Three-dimensional structure of firefly luciferase
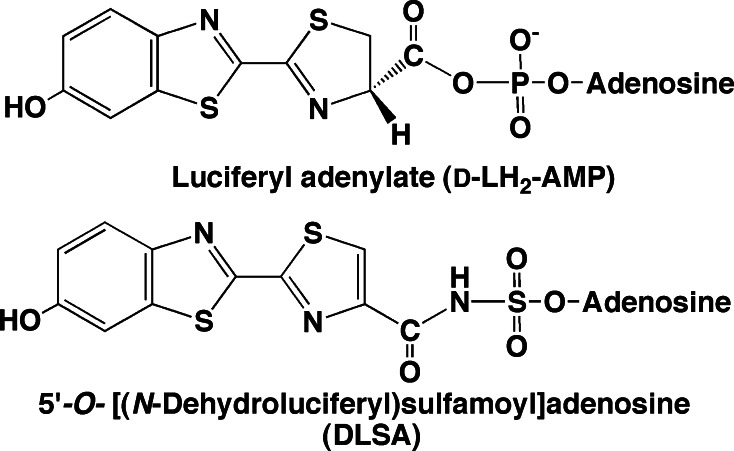
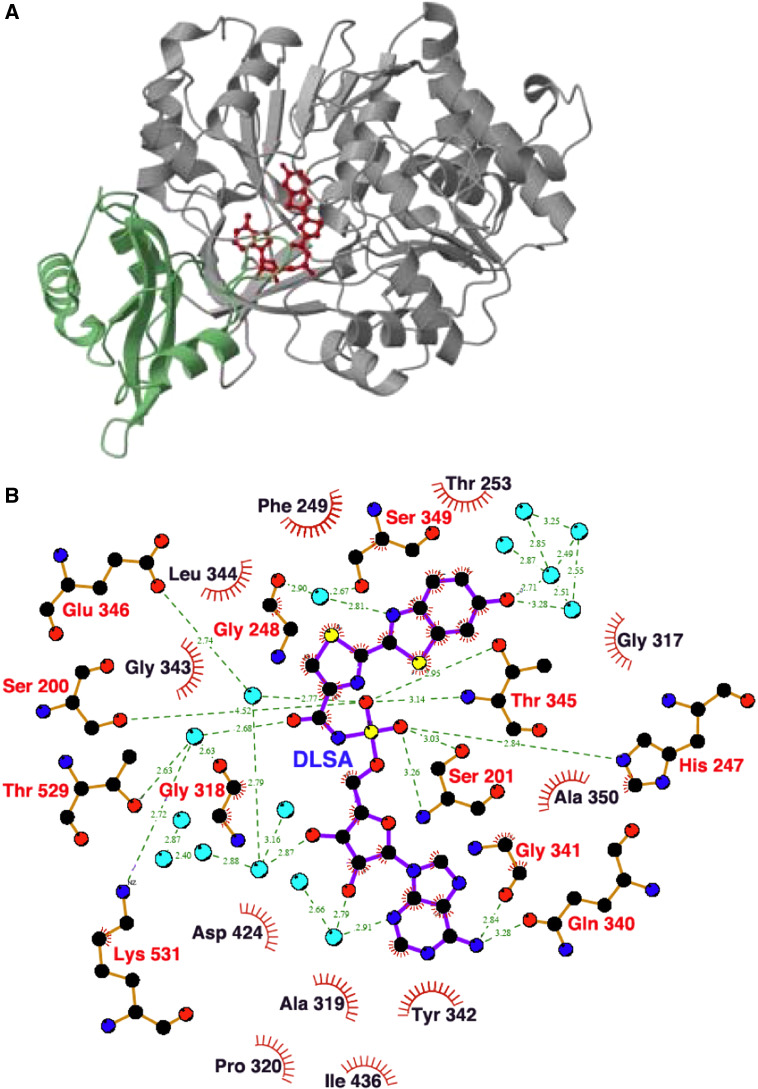
In 1996, the three-dimensional structure of P. pyralis luciferase was firstly determined as an unbounded form without any substrate or analogue [127]. The luciferase molecule folds two distinct domains, a large N-terminal domain (1–436 aa) and a small C-terminal domain (440–550 aa), joined by a flexible linker peptide (PDB ID: 1LC1). The N-terminal domain consists of an antiparallel β-barrel and two β-sheets. The two β-sheets are assembled to form a five-layered αβαβ tertiary structure. Bromoform (CHBr3) is known as an anesthetic compound that inhibits the luminescence reaction of firefly luciferase. The crystal structure of bromoform-bounded firefly luciferase was determined, suggesting that two bromoform molecules might bind in the possible substrate interacting area (PDB ID: 1BA3) [128]. Later, to elucidate the conformational changes of the enzyme–substrate complex in the firefly luciferase reaction, an intermediate analogue of the natural luciferyl adenylate, 5′-O-[N-(dehydroluciferyl)sulfamoyl]adenosine (DLSA), was synthesized (Fig. 11) [129]. The stable adenylate analogue DLSA is a potent competitive inhibitor for the luminescence reaction; K i = 34 ± 5 nM for LH2, K i = 41 ± 3 nM for Mg2+-ATP, and K i = 340 ± 50 nM for LH2-AMP. In 2006, DLSA was synthesized by another synthetic route and used for determining the crystal structure of L. cruciata luciferase in complex with DLSA (PDB ID: 2D1S) [130] (Fig. 12a). The results showed that two domain structures of L. cruciata luciferase are essentially the same as P. pyralis luciferase [127, 128], and a relationship between color variation and luciferase structure was proposed [130]. It is interesting that many water molecules could interact with the luciferyl adenylate analogue (DLSA) and oxyluciferin (PDB ID: 2D1Q) (Fig. 12b). However, the color variations were not explained by these structural analyses including the oxyluciferin–luciferase complex. The attack of molecular oxygen (O2) to d-luciferyl adenylate might be from a specific direction, because l-luciferyl adenylate could not react with O2 to produce light. An important point is how molecular oxygen accesses to the luciferyl adenylate to emit light from outside the luciferase molecule [Fig. 6]. An open space for O2 to reach luciferyl adenylate has been found in the structure of the luciferase–DLSA complex (A. Nakagawa, personal communication).
Fig. 11.
Comparison of chemical structures between luciferyl adenylate and its analogue DLSA
Fig. 12.
Structure of the firefly L. cruciata luciferase–DLSA complex. a Ribbon diagram of firefly luciferase in complex with DLSA (red). The N-terminal large and C-terminal small domains are drawn in gray and green, respectively, b A schematic representation of the DLSA binding area in firefly luciferase. The possible hydrogen bonds are shown by a green dashed line, and light blue circles show water molecules. The amino acid residues labeled with red and black show the hydrophobic contacts with DLSA
Multiple functions of luciferase as an adenylate-forming enzyme
Firefly luciferase is a fatty acyl-CoA synthetase
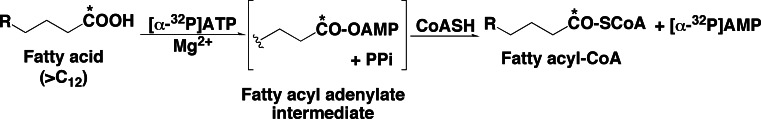
We predicted that firefly luciferase has the catalytic function of fatty acyl-CoA synthesis from free fatty acids, based on the following background information: (1) the luminescence reaction involves in an adenylation step [28]; (2) dehydroluciferin, a potent inhibitor for luciferase, might be released from luciferase molecule as dehydroluciferyl-CoA, suggesting that luciferase has the catalytic function of a acyl-CoA synthesis [58]; (3) the primary structure of firefly luciferase showed high sequence similarity to a long-chain acyl-CoA synthetase [123]; and (4) the long fatty acids (C12–C20) inhibited the luminescence reaction in competition with firefly luciferin [102, 103]. As shown in Fig. 13, the catalytic function of fatty acyl-CoA synthesis by firefly luciferase was determined [17]. The initial step of the acyl-adenylate formation was examined by monitoring the formation of [α-32P]AMP from [α-32P]ATP in the presence of Mg2+, CoA and various carboxylic acids; acetic acid (C2), propionic acid (C3), palmitic acid (C16:0), oleic acid (C18:1), linoleic acid (C18:2), linolenic acid (C18:3), arachidonic acid (C20:4), and p-coumaric acid [17]. Based on these results, long-chain unsaturated fatty acids (C16–C20) are suitable substrates for the adenylation reaction by P. pyralis luciferase, but other substrates of aromatic acids for plant p-coumarate CoA:ligase and short chain acids were not utilized efficiently by P. pyralis firefly luciferase. The identification of fatty acyl-CoA formation via fatty acyl-adenylate was performed using [1-14C]oleic acid as a substrate (Fig. 13). The formation of [1-14C]oleoyl-CoA was confirmed by TLC analysis, and the essential cofactors for the fatty acyl-CoA synthesis are ATP, Mg2+ and CoA. Other nucleotides including GTP, CTP, TTP, UTP, and ITP did not stimulate the formation of fatty acyl-CoA significantly. Further, fatty acyl-CoA synthesized from fatty acid (C18:1, C18:2, C18:3,C20:4) by P. pyralis and L. cruciata luciferase was isolated by reversed-phase HPLC and identified by MALDI-TOF–MS. Finally, we demonstrated that firefly luciferase is a bifunctional enzyme that catalyzes not only the luminescence reaction but also long-chain fatty acyl-CoA synthesis [17]. The substrate specificity of P. pyralis and L. cruciata luciferase with a series of carboxylic acids including amino acid, aromatic acid and monohydroxyl fatty acids was determined [131]. Very weak activities of adenylation were observed for amino and imino acids (l-proline, d-proline, l-cysteine, d-cysteine, l-phenylalanine, l-thiaproline, and thiazolidine-2-carboxylic acid). No significant activity was observed for short chain acids (acetic acid, propionic acid, n-hexanoic acid, and lipoic acid) and aromatic acids (p-coumaric acid, caffeic acid, ferulic acid, 1-naphthylacetic acid, 2-naphthylacetic acid, 2-quinolinecarboxylic acid, and benzoic acid) [132].
Fig. 13.
Determination of fatty acyl-CoA synthesis through fatty acyl adenylate from fatty acid, catalyzed by firefly luciferase. Asterisks indicate 14C-labeled position
Homologous gene of firefly luciferase in Drosophila melanogaster
To understand the genetic origin and biological function of firefly luciferase, a homology search has been performed in FlyBase for Drosophila. The homology search showed that the gene product of CG6178 (GC6178) was referred to as a “firefly luciferase”, due to the highest similarity in amino acid sequence with firefly luciferase. Also, several other homologous genes including CG4830, CG8834, CG9390, CG11407, and CG18586 were detected. However, there was no experimental evidence for the luminescence activity. The cDNA of GC6178 was isolated and expressed in E. coli cells. The recombinant CG6178 was purified and characterized [132]. As expected, the purified CG6178 showed a long fatty acyl-CoA synthetic activity, but no luminescence activity. Crude extracts of D. melanogaster adults did not show detectable luminescence activity with firefly luciferin (S. Zenno, personal communication). The long-chain fatty acids (C18–C20) are a good substrate for CG6178. The optimum pH is at 7.5–8.5, similar to firefly luciferase. High concentrations (>10 μM) of oleic acid (C18:1), linolenic acid (C18:3) and arachidonic acid (C20:4) inhibit the fatty acyl-CoA synthetic activity of CG6178 and P. pyralis luciferase, and moderate inhibition was observed in linoleic acid. The K m values of CG6178 and P. pyralis luciferase for linoleic acid are 1.88 and 13.6 μM, respectively. On the other hand, no inhibition by lauric acid (C14:0) was observed at 40 μM in CG6178 and P. pyralis luciferase. The K m values of CG6178, P. pyralis luciferase and L. cruciata luciferase for lauric acid were 1.68, 7.41, and 16.3 μM, respectively [131]. Thus, CG6178 is the first protein that was identified as a long-chain fatty acyl-CoA synthetase in Drosophila. To understand the catalytic properties, the chimeric proteins of P. pyralis luciferase and CG6178 were prepared, and only the chimeric protein of the N-terminal domain of P. pyralis luciferase and C-terminal domain of CG6178 showed the luminescence activity with ~4% activity of P. pyralis luciferase [133].
Homologous genes of firefly luciferase in luminous and non-luminous beetles are fatty acyl-CoA synthetases
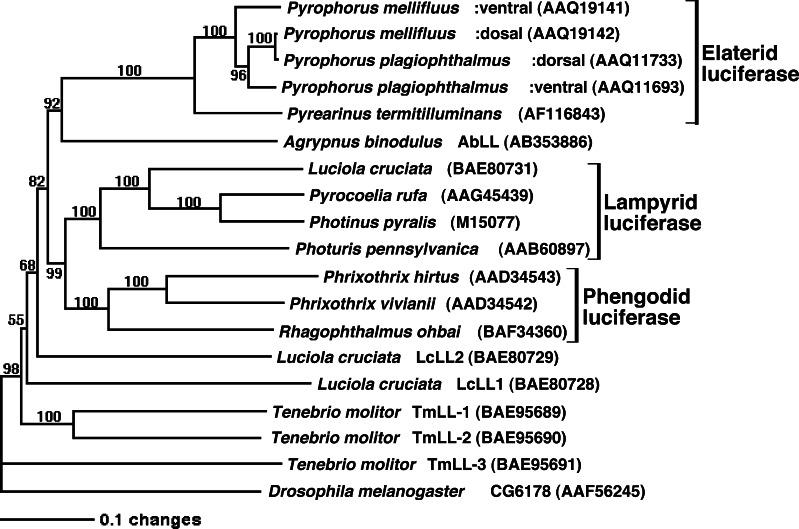
Two homologous genes of firefly luciferase, LcLL1 and LcLL2, were cloned from the Japanese firefly, L. cruciata, and characterized [134]. The gene product of LcLL1 had long-chain fatty acyl-CoA synthetic activity, but not luciferase activity. The other gene product of LcLL2 did not show enzymatic activities of acyl-CoA synthetase or luciferase. The gene expression levels of LcLL1, LcLL2 and L. cruciata luciferase in larva and adults of L. cruciata were analyzed by RT-PCR, indicating that three genes were abundant in larvae and LcLL2 was only abundant in adults. Thus, LcLL1 and LcLL2 are paralogous genes of firefly luciferase and may have been derived from gene duplication before the luminous and non-luminous insects diverged. From the non-luminous mealworm beetle, Tenebrio molitor, three orthologous genes of firefly luciferase, TmLL-1, TmLL-2 and TmLL-3, were isolated, and the recombinant proteins were purified. The gene products were identified as a fatty acyl-CoA synthetase [135], and they did not show detectable luminescence activity. Further, a homologous gene of firefly luciferase was isolated from the Japanese non-luminous click beetle, A. binodulus, and assigned AbLL (Agrypnus binodulus luciferase-like gene) [136]. The identity of the amino acid sequence deduced from AbLL with the click beetle luciferase from P. plagiophthalmus was 55%, which is higher than that between click beetle luciferase and firefly luciferase (~48%). The gene product of AbLL (AbLL) had medium- and long-chain fatty acyl-CoA synthetase activity, but not luciferase activity [136]. Thus, all homologous genes of firefly luciferase from luminous and non-luminous insects did not show the significant activity of luminescence [131–136]. As shown in Fig. 14, phylogenic analysis was performed using homologous genes from luminous and non-luminous insects. Taken together with the fatty acyl-CoA synthetic activity in these homologues, firefly luciferase (beetle luciferase) originated from a fatty acyl-CoA synthetase by gene duplication in insects [136]. The gene duplications might occur before luminous insects within Lampyridae, Phengodidae and Elateridae diverged [132–137].
Fig. 14.
A phylogenetic reconstruction of luciferases and their similar genes in insects. Numbers on the nodes indicate bootstrap values % from 10,000 replicates, and only the values over 50% are shown. Horizontal branch lengths indicate the genetic distances. GenBank accession number (Protein/Nucleotide) is shown in parentheses
Functional conversion of fatty acyl-CoA synthetase to firefly luciferase
To demonstrate the evolutionary origin of luciferase in insects, we tried functional conversion to firefly luciferase from fatty acyl-CoA synthetase. For this purpose, we focused on AbLL, which possesses 55% identity in amino acid sequence with the luminous click beetle luciferase and is the most homologous gene to firefly luciferase among non-luminous insects [136] (Fig. 15). The functional conversion of AbLL to a luciferase-like AbLL possessing luminescence activity was performed by site-directed mutagenesis [138]. The crystal structure analysis of L. cruciata luciferase-bound with an adenylate analogue of DLSA suggested that the benzothiazole ring of DLSA is in van der Waals contact with several amino acid residues and water molecules [130] (Fig. 12b). The hydroxyl group of Ser349 forms a hydrogen bond with the benzothiazole ring through a water molecule. By comparison of the amino acid residues in several luciferases with AbLL, two AbLL mutants of AbLL-L216R (Mt2) and AbLL-L345S (Mt14) were constructed (Table 2). The mutant AbLL-Mt2 did not show luminescence enhancement but AbLL-Mt14 did. This result indicated that a single substitution to Ser345 is crucial for light emission in AbLL. The amino acid sequence from 337 to 346 is Gly-Tyr-Gly-Leu-Thr-Glu-Cys-Gly-Leu-Ala in AbLL, and the three amino acids italicized are different from the corresponding sequence in luciferases: Gly-Tyr-Gly-Leu-Thr-Glu-Ser(Thr)-Thr-Ser-Ala. The mutant AbLL-Mt11 was introduced by additional mutations of Ser343 and Thr344 in AbLL-Mt14 and showed a significant increase of luminescence activity (Table 2). The luminescence enhancement of AbLL-11 might be explained by the formation of a hydrogen-bonding network, similar to the case of Met398, Thr347, Thr348, and Ser349 in L. cruciata luciferase. The interaction of the hydroxyl group of Ser345 in AbLL mutants with the benzothiazole moiety of luciferin through a water molecule is key for functional conversion from a fatty acyl-CoA to a luciferase. However, the luminescence intensity of AbLL-Mt11 was only 0.001% of that of L. cruciata luciferase. It is not clear whether this mutation affects the adenylation step or oxygenation step in the luciferase reaction. To increase the conversion efficiency of the luminescence activity, X-ray structural analysis of AbLL might suggest the amino residues for mutations. The luminescence pattern of AbLL-Mt11 was different from firefly luciferases, and showed a steady-state luminescence reaction like a general enzyme, in the presence and absence of coenzyme A. The maxima of emission spectra of AbLL mutants, Mt14, Mt17, Mt11, and Mt16, were around 610 nm at pH 7.8, different from P. pyralis and L. cruciata luciferases. Further, the CG6178 mutant at the same positions enhanced the luminescence intensity, similar to the case of AbLL [138].
Fig. 15.
Comparison of the amino acid sequences of AbLL (AbLL, AB353886), P. plagiophthalmus luciferase (PplLuc; AF543373), P. pyralis luciferase (PpyLuc; M15077) and L. cruciata luciferase (LcuLuc; M26194). Letters in the black boxes and shaded boxes indicate the identical amino acids and the similar amino acid groups with AbLL, respectively. The groups are defined as follows: A, S, T, P, and G; N, D, E, and Q; H, R, and K; M, L, I, and V; F, Y, and W. Gaps were inserted to assist in sequence alignment. The numbers on the right margin indicate the position of amino acid residues. Double arrow indicates the AMP-binding motif. Three asterisks indicate the positions of mutation in AbLL. The linker region between the N-terminal domain and C-terminal domain is shown by the sharp symbols
Table 2.
Comparison of amino acid sequence of mutated AbLL and their activity [138]
| Enzymes | Amino acid sequencesa | Relative activity (%) |
|---|---|---|
| A. binodulus AbLL | (337)-GYGLTECGLA-(346) | 0.2 |
| AbLL-Mt14 | (337)-GYGLTECGSA-(346) | 5.9 |
| AbLL-Mt17 | (337)-GYGLTECTSA-(346) | 27.0 |
| AbLL-Mt11 | (337)-GYGLTESTSA-(346) | 100.0 |
| AbLL-Mt13 | (337)-GYGLTECTLA-(346) | 0.2 |
| AbLL-Mt12 | (337)-GYGLTESGLA-(346) | 0.2 |
| AbLL-Mt16 | (337)-GYGLTESGSA-(346) | 3.9 |
| AbLL-Mt15 | (337)-GYGLTESTLA-(346) | 0.2 |
| -Enzyme | 0.1 | |
| L. cruciata luciferase | (341)-GYGLTETTSA-(350) | – |
| P. pyralis luciferase | (339)-GYGLTETTSA-(348) | – |
| P. mellifluus luciferase | (336)-GYGLTESTSA-(345) | – |
| P. termitilluminans luciferase | (336)-GYGLTESTSA-(345) | – |
aThe mutated amino acid residues in AbLL are shown in bold
Concluding remarks
Many valuable books and reviews on bioluminescence have been published in the past [1–9], and firefly luciferase is one of the well-characterized enzymes in bioluminescence. However, there are still many unsolved problems, including the light-emitting species of oxyluciferin, the catalytic process of luminescence in a luciferase molecule, and the biosynthetic pathway of luciferin in the firefly.
Acknowledgment
The author is grateful to Dr. A. Nakagawa (Osaka University, Japan) for illustrating the X-ray structure model of firefly luciferase. Also, the author thanks Dr. T. Hosoya (Tokyo Medical and Dental University, Japan) for discussion about luminescence mechanisms.
References
- 1.Harvey EN. Bioluminescence. New York: Academic; 1952. [Google Scholar]
- 2.Haneda Y, Johnson FH, editors. Bioluminescence in progress. New Jersey: Princeton University Press; 1966. [Google Scholar]
- 3.Herring PJ, editor. Bioluminescence in action. New York: Academic; 1978. [Google Scholar]
- 4.Campbell AK. Chemiluminescence: principle and applications in biology and medicine. New York: VCH; 1988. [Google Scholar]
- 5.Shimomura O. Bioluminescence: chemical principles and methods. Singapore: World Scientific; 2006. [Google Scholar]
- 6.Shimomura O. Bioluminescence in the sea: photoprotein systems. Soc Exp Biol Symp. 1985;39:351–372. [PubMed] [Google Scholar]
- 7.Hastings JW, Morin JG (1991) Bioluminescence. In: Prosser CL (ed) Neural and integrative animal physiology. Wiley, New York, pp 131–170
- 8.Meighen EA, Dunlap PV. Physiological, biochemical and genetic control of bacterial bioluminescence. Adv Microbial Physiol. 1993;34:1–67. doi: 10.1016/s0065-2911(08)60027-2. [DOI] [PubMed] [Google Scholar]
- 9.Wilson T, Hastings JW. Bioluminescence. Annu Rev Cell Dev Biol. 1998;14:197–230. doi: 10.1146/annurev.cellbio.14.1.197. [DOI] [PubMed] [Google Scholar]
- 10.Bronstein I, Fortin J, Stanley PE, Stewart GSAB, Kricka LJ. Chemiluminescent and bioluminescent reporter gene assays. Anal Biochem. 1994;219:169–181. doi: 10.1006/abio.1994.1254. [DOI] [PubMed] [Google Scholar]
- 11.Greer LF, III, Szalay AA. Imaging of light emission from the expression of luciferases in living cells and organisms: a review. Luminescence. 2002;17:43–73. doi: 10.1002/bio.676. [DOI] [PubMed] [Google Scholar]
- 12.Lloyd JE. Bioluminescence and communication in insects. Annu Rev Entomol. 1983;28:131–160. [Google Scholar]
- 13.Wood KV. The chemical mechanism and evolutionary development of beetle bioluminescence. Photochem Photobiol . 1995;62:662–673. [Google Scholar]
- 14.Viviani VR. The origin, diversity, and structure function relationships of insect luciferases. Cell Mol Life Sci. 2002;59:1833–1850. doi: 10.1007/PL00012509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Nakamura M, Niwa K, Maki S, Hirano T, Ohmiya Y, Niwa H. Conversion of a new firefly bioluminescence system using l-luciferin as substrate. Tetrahedron Lett. 2006;47:1197–1200. [Google Scholar]
- 16.McElroy WD, DeLuca M, Travis J. Molecular uniformity in biological catalyses. Science. 1967;157:150–160. doi: 10.1126/science.157.3785.150. [DOI] [PubMed] [Google Scholar]
- 17.Oba Y, Ojika M, Inouye S. Firefly luciferase is a bifunctional enzyme: ATP-dependent monooxygenase and a long chain fatty acyl-CoA synthetase. FEBS Lett. 2003;540:251–254. doi: 10.1016/s0014-5793(03)00272-2. [DOI] [PubMed] [Google Scholar]
- 18.Bitler B, McElroy WD. The preparation and properties of crystalline firefly luciferin. Arch Biochem Biophys. 1957;72:358–368. doi: 10.1016/0003-9861(57)90212-6. [DOI] [PubMed] [Google Scholar]
- 19.Kishi Y, Matsuura S, Inoue S, Shimomura O, Goto T. Luciferin and luciopterin isolated from the Japanese firefly, Luciola cruciata . Tetrahedron Lett. 1968;9:2847–2850. doi: 10.1016/s0040-4039(00)75644-9. [DOI] [PubMed] [Google Scholar]
- 20.White EH, McCapra F, Field GF, McElroy WD. The structure and synthesis of firefly luciferin. J Am Chem Soc. 1961;83:2402–2403. [Google Scholar]
- 21.White EH, McCapra F, Field GF. The structure and synthesis of firefly luciferin. J Am Chem Soc. 1963;85:337–343. [Google Scholar]
- 22.Blank GE, Pletcher J, Sax M. The molecular structure of firefly d-(−)-luciferin: a single crystal analysis. Biochem Biophys Res Commun. 1971;42:583–588. doi: 10.1016/0006-291x(71)90411-6. [DOI] [PubMed] [Google Scholar]
- 23.Branchini BR, Magyar RA, Murtiashaw MH, Porieer NC. The role of active site residue arginine 218 in firefly luciferase. Biochemistry. 2001;40:2410–2418. doi: 10.1021/bi002246m. [DOI] [PubMed] [Google Scholar]
- 24.Oba Y, Shintani T, Nakamura T, Ojika M, Inouye S. Determination of the luciferin contents in luminous and non-luminous beetles. Biochem Biosci Biotechnol. 2008;272:384–1387. doi: 10.1271/bbb.80017. [DOI] [PubMed] [Google Scholar]
- 25.Seliger HH, McElroy WD, White EH, Field GF. Stereospecificity and firefly bioluminescence, a comparison of natural and synthetic luciferins. Proc Natl Acad Sci USA. 1961;47:1129–1961. doi: 10.1073/pnas.47.8.1129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lembert N. Firefly luciferase can use l-luciferin to produce light. Biochem J. 1996;317:273–277. doi: 10.1042/bj3170273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Niwa K, Nakamura M, Ohmiya Y. Stereoisomeric bio-inversion key to biosynthesis of firefly d-luciferin. FEBS Lett. 2006;580:5283–5287. doi: 10.1016/j.febslet.2006.08.073. [DOI] [PubMed] [Google Scholar]
- 28.Rhodes WC, McElroy WD. The synthesis and function of luciferyl-adenylate and oxyluciferyl-adenylate. J Biol Chem. 1958;233:1528–1537. [PubMed] [Google Scholar]
- 29.Seliger HH, McElroy WD. Chemiluminescence of firefly luciferin without enzyme. Science. 1962;138:683–685. doi: 10.1126/science.138.3541.683. [DOI] [PubMed] [Google Scholar]
- 30.Suzuki N, Sato M, Nishikawa K, Goto T. Synthesis and spectral properties of 2-(6′-hydroxybenzothiazol-2′-yl)-4-hydroxytiazole, a possible emitting species in the firefly bioluminescence. Tetrahedron Lett. 1969;53:4683–4684. [Google Scholar]
- 31.Suzuki N, Goto T. Firefly bioluminescence II. Identification of 2-(6′-hydroxybenzothiazol-2′-yl)-4-hydroxythiazole as a product in the bioluminescence of firefly lanterns and a product in the chemiluminescence of firefly luciferin in DMSO. Tetrahedron Lett. 1971;22:2021–2024. [Google Scholar]
- 32.Suzuki N, Goto T. Synthesis of 4-thiazolone derivatives related to firefly luciferin. Agric Biol Chem. 1972;36:2213–2221. [Google Scholar]
- 33.Hopkins TA, Seliger HH, White EH, Cass MW. The chemiluminescence of firefly luciferin. A model for the bioluminescent reaction and identification if the product excited state. J Am Chem Soc. 1969;89:7148–7150. doi: 10.1021/ja01002a076. [DOI] [PubMed] [Google Scholar]
- 34.Esteves da Silva JCG, Magalhaes JMCS, Fontes T. Identification of enzyme produced firefly oxyluciferin by reverse phase HPLC. Tetrahedron Lett. 2001;42:8173–8176. [Google Scholar]
- 35.Ribeiro C, Esteves da Silva JC. Kinetics of inhibition of firefly luciferase by oxyluciferin and dehydroluciferyl-adenylate. Photochem Photobiol . 2008;7:1085–1090. doi: 10.1039/b809935a. [DOI] [PubMed] [Google Scholar]
- 36.Branchini BR, Murtiashaw MH, Magyer RA, Anderson SM. The role of lysine 529, a conserved residue of the acyl-adenylate-forming enzyme superfamily, in firefly luciferase. Biochemistry. 2000;39:5433–5440. doi: 10.1021/bi9928804. [DOI] [PubMed] [Google Scholar]
- 37.Imai K, Goto T. Improved synthesis of firefly d-luciferyl–d-adenylate—A key intermediate of firefly bioluminescence. Agric Biol Chem. 1988;52:2803–2809. [Google Scholar]
- 38.White EH, Steinmetz MG, Miano JD, Wildes PD, Morland R. Chemi- and bioluminescence of firefly luciferin. J Am Chem Soc. 1980;102:3199–3208. [Google Scholar]
- 39.Shimomura O, Goto T, Johnson FH. Source of oxygen in the CO2 produced in the bioluminescent oxidation of firefly luciferin. Proc Natl Acad Sci USA. 1977;74:2799–2802. doi: 10.1073/pnas.74.7.2799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.McCapra F (1977) Alternative mechanism of dioxetanone decomposition. J Chem Soc Chem Commun (1977) 946–948
- 41.Koo J-Y, Schmidt SP, Schuster GB. Bioluminescence of the firefly: key steps in the formation of the electronically excited state for model systems. Proc Natl Acad Sci USA. 1978;84:131–135. doi: 10.1073/pnas.75.1.30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Seliger HH, McElroy WD. Spectral emission and quantum yield of firefly bioluminescence. Arch Biochem Biophys. 1960;88:136–141. doi: 10.1016/0003-9861(60)90208-3. [DOI] [PubMed] [Google Scholar]
- 43.White EH, Roswell DF. Analogs and derivatives of firefly oxyluciferin, the light emitter in firefly bioluminescence. Photochem Photobiol . 1991;53:131–136. doi: 10.1111/j.1751-1097.1991.tb08478.x. [DOI] [PubMed] [Google Scholar]
- 44.Kajiyama N, Nakano E. Isolation and characterization of mutants of firefly luciferase which produced different color of light. Protein Eng. 1991;4:691–693. doi: 10.1093/protein/4.6.691. [DOI] [PubMed] [Google Scholar]
- 45.Branchini BR, Murtiashaw MH, Magyar RA, Protier NC, Ruggiero MC, Stroh JG. Yellow-green and red firefly bioluminescence from 5,5′-dimethoxyluciferin. J Am Chem Soc. 2002;124:2112–2113. doi: 10.1021/ja017400m. [DOI] [PubMed] [Google Scholar]
- 46.McCapra F, Gilfoyle DJ, Young DW, Church NJ, Spencer P. The chemical origin of color differences in beetle bioluminescence. In: Campbell AK, Kricka LJ, Stanley PE, editors. Bioluminescence and chemiluminescence. Fundamentals and applied aspects. Chichester, UK: Wiley; 1994. pp. 387–391. [Google Scholar]
- 47.McCapra F. Mechanism in chemiluminescence and bioluminescence—unfinished business. In: Hastings JW, Kricka LJ, Stanley PE, editors. Bioluminescence and chemiluminescence molecular reporting with photons. Chichester, UK: Wiley; 1997. pp. 7–15. [Google Scholar]
- 48.Branchini RB, Southworth TL, Murtiashaw MH, Magyar RA, Gonzalez SA, Ruggiero MC, Stroh JG. An alternative mechanism of bioluminescence color determination in firefly luciferase. Biochemistry. 2004;43:7255–7262. doi: 10.1021/bi036175d. [DOI] [PubMed] [Google Scholar]
- 49.Orlova G, Goddard JD, Brovko LY. The theoretical study of the amazing firefly bioluminescence: the formation and structures of the light emitters. J Am Chem Soc. 2003;125:6962–6971. doi: 10.1021/ja021255a. [DOI] [PubMed] [Google Scholar]
- 50.Yang T, Goddard JD. Prediction of the geometries and florescence emission energies of oxyluciferins. J Phys Chem. 2007;111:4489–4497. doi: 10.1021/jp068542b. [DOI] [PubMed] [Google Scholar]
- 51.Nakatani N, Hasegawa J, Nakatuji H. Red light in chemiluminescence and yellow-green light in bioluminescence: color-turning mechanism of firefly, Photinus pyralis, studied by the symmetry-adapted cluster-conjugation interaction method. J Am Chem Soc. 2007;129:8756–8765. doi: 10.1021/ja0611691. [DOI] [PubMed] [Google Scholar]
- 52.Hirano T, Hasumi Y, Ohtsuka K, Maki S, Niwa H, Yamaji M, Hashizume D. Spectroscopic studies of the light-color modulation mechanism of firefly (beetle) bioluminescence. J Am Chem Soc. 2009;131:2385–2396. doi: 10.1021/ja808836b. [DOI] [PubMed] [Google Scholar]
- 53.DeLuca M, McElroy WD. Purification and properties of firefly luciferase. Methods Enzymol. 1978;57:3–15. [Google Scholar]
- 54.DeLuca M, Wannlund J, McElroy WD. Factors affecting the kinetics of light emission from crude and purified firefly luciferase. Anal Biochem. 1979;95:194–198. doi: 10.1016/0003-2697(79)90204-5. [DOI] [PubMed] [Google Scholar]
- 55.Webster JJ, Chang JC, Manley ER, Spivey HO, Leach FR. Buffer effects on ATP analysis by firefly luciferase. Anal Biochem. 1980;106:7–11. doi: 10.1016/0003-2697(80)90111-6. [DOI] [PubMed] [Google Scholar]
- 56.Simpson WJ, Hammond JRM. The effect of detergents on firefly luciferase reaction. J Biolumi Chemilumi. 1991;6:97–106. doi: 10.1002/bio.1170060207. [DOI] [PubMed] [Google Scholar]
- 57.Ford SR, Chenault KH, Bunton LS, Hampton GJ, McCarthy J, Hall MS, Pangburn SJ, Buck LM, Leach FR. Use of firefly luciferase for ATP measurement: other nucleotides enhance turnover. J Biolumi Chemilumi. 1996;11:149–167. doi: 10.1002/(SICI)1099-1271(199605)11:3<149::AID-BIO411>3.0.CO;2-Q. [DOI] [PubMed] [Google Scholar]
- 58.Airth RL, Rhodes WC, McElroy WD. The function of coenzyme A in luminescence. Biochim Biophys Acta. 1958;27:519–532. doi: 10.1016/0006-3002(58)90381-0. [DOI] [PubMed] [Google Scholar]
- 59.Roda A, Guardigli M, Michelini E, Mirasoli M. Bioluminescence in analytical chemistry and in vivo imaging. Trends Anal Chem. 2009;28:307–322. [Google Scholar]
- 60.Seliger HH, McElroy WD. Quantum yield in the oxidation of firefly luciferin. Biochem Biophys Res Commun. 1959;1:21–24. [Google Scholar]
- 61.Ando Y, Niwa K, Yamada N, Enomoto T, Irie T, Kubota H, Ohmiya Y, Akiyama H. Firefly bioluminescence quantum yield and color change by pH-sensitive green emission. Nat Photon. 2008;2:44–46. [Google Scholar]
- 62.Fraga H, Esteves da Silva JCG, Fontes R. Identification of luciferyl adenylate and luciferyl coenzyme A synthesized by firefly luciferase. Chem Bio Chem. 2006;5:110–115. doi: 10.1002/cbic.200300735. [DOI] [PubMed] [Google Scholar]
- 63.Okada K, Iio H, Kubota I, Goto T. Firefly bioluminescence. III. Conversion of oxyluciferin to luciferin in firefly. Tetrahedron Lett. 1974;32:2771–2774. [Google Scholar]
- 64.Okada K, Iio H, Goto T (1976) Biosynthesis of firefly luciferin. Probable formation of benzothiazole from p-benzoquinone and cysteine. J Chem Soc Chem Comm (1976) 32
- 65.McCapra F, Razavi Z (1975) A model for firefly luciferin biosynthesis. J Chem Soc Chem Comm (1975) 42–43
- 66.McCapra F, Razavi Z (1976) Biosynthesis of luciferin in Pyroporus pellucens. J Chem Soc Chem Commun (1976) 153–154
- 67.Gomi K, Kajiyama N. Oxyluciferin, a luminescence product of firefly luciferase, is enzymatically regenerated into luciferin. J Biol Chem. 2001;276:36508–36513. doi: 10.1074/jbc.M105528200. [DOI] [PubMed] [Google Scholar]
- 68.Gomi K, Hiorkawa K, Kajiyama N. Molecular cloning and expression of the cDNAs encoding luciferin-regenerating enzyme from Luciola cruciata and Luciola lateralis . Gene. 2002;294:157–166. doi: 10.1016/s0378-1119(02)00764-3. [DOI] [PubMed] [Google Scholar]
- 69.Day JC, Tisi L, Bailey MJ. Evolution of beetle bioluminescence: the origin of beetle luciferin. Luminescence. 2004;19:8–20. doi: 10.1002/bio.749. [DOI] [PubMed] [Google Scholar]
- 70.Marques SM, Estreves da Silva JCG. Firefly bioluminescence: a mechanistic approach of luciferase catalyzed reactions. IUBMB Life. 2009;61:6–17. doi: 10.1002/iub.134. [DOI] [PubMed] [Google Scholar]
- 71.Oba Y, Kato S, Ojika M, Inouye S. Biosynthesis of luciferin in the sea firefly, Cypridina hilgendorfii: l-tryptophan is a component in Cypridina luciferin. Tetrahedron Lett. 2002;43:2389–2392. [Google Scholar]
- 72.Kato S, Oba Y, Ojika M, Inouye S. Identification of the biosynthetic units of Cypridina luciferin in Cypridina (Vargula) hilgendorfii by LC/ESI-TOF-MS. Tetrahedron. 2004;60:11427–11434. [Google Scholar]
- 73.White EH, Wörther H, Field GF, McElroy WD. Analogs of firefly luciferin. J Org Chem. 1965;30:2344–2348. doi: 10.1021/jo01343a039. [DOI] [PubMed] [Google Scholar]
- 74.White EH, Wörther H, Seliger HH, McElroy WD. Amino analogs of firefly luciferin and biological activity thereof. J Am Chem Soc. 1966;88:2015–2019. [Google Scholar]
- 75.White EH, Wörther H. Analogs of firefly luciferin. III. J Org Chem. 1966;31:1484–1488. doi: 10.1021/jo01343a039. [DOI] [PubMed] [Google Scholar]
- 76.White EH, Rapaport E, Hopkins TA, Seliger HH. Chemi- and bioluminescence of firefly luciferin. J Am Chem Soc. 1969;91:2178–2180. doi: 10.1021/ja01036a093. [DOI] [PubMed] [Google Scholar]
- 77.Seto S, Ogura K, Nishiyama Y. A convenient synthetic method of 2-carbamoyl-6-methoxybenzothiazole, one of intermediates for the synthesis of firefly luciferin. Bull Chem Soc Jpn. 1963;36:331–333. [Google Scholar]
- 78.France C, Blanchot B, Champiar D, Couble P, Declercq G, Millet J-L. Synthesis and characterization of a new substrate of Photinus pyralis luciferase: 4-methyl-d-luciferin. J Clin Chem Clin Biochem. 1990;28:471–474. doi: 10.1515/cclm.1990.28.7.471. [DOI] [PubMed] [Google Scholar]
- 79.Woodroofe CC, Shultz JW, Wood MG, Osterman J, Cali JJ, Daily WJ, Meisenheimer PL, Klaubert DH. N-Alkylated 6′-aminoluciferins are bioluminescent substrates for Ultra-Glo and QuantiLum luciferase: new potential scaffolds for bioluminescent assays. Biochemistry. 2008;47:10383–10393. doi: 10.1021/bi800505u. [DOI] [PubMed] [Google Scholar]
- 80.Branchini BR, Hayward MM, Bamford S, Brennan PM, Lajiness EJ. Naphtyl- and quinolyllucifferin: green and red light emitting firefly luciferin analogues. Photochem Photobiol . 1989;49:689–695. doi: 10.1111/j.1751-1097.1989.tb08442.x. [DOI] [PubMed] [Google Scholar]
- 81.Miska W, Geiger R. Synthesis and characterization of luciferin derivatives for use in bioluminescence enhanced enzyme immunoassays. J Clin Chem Clin Biochem. 1987;25:23–30. doi: 10.1515/cclm.1987.25.1.23. [DOI] [PubMed] [Google Scholar]
- 82.Geiger R, Miska W. Bioluminescence enhanced enzyme immunoassay. J Clin Chem Clin Biochem. 1987;25:31–38. doi: 10.1515/cclm.1987.25.1.31. [DOI] [PubMed] [Google Scholar]
- 83.Miska W, Geiger R. Luciferin derivatives in bioluminescence-enhanced enzyme immunoassays. J Biolumi Chemilumi. 1989;4:119–128. doi: 10.1002/bio.1170040118. [DOI] [PubMed] [Google Scholar]
- 84.Hauber R, Geiger R. A sensitive, bioluminescent-enhanced detection method for DNA-dot-hybridization. Nucl Acids Res. 1988;16:1213. doi: 10.1093/nar/16.3.1213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Huber R, Geiger R. A new, very sensitive, bioluminescence-enhanced detection system for protein blotting. Ultrasensitive detection systems for protein blotting and DNA hybridization, I. J Clin Chem Clin Biochem. 1987;25:511–514. doi: 10.1515/cclm.1987.25.8.511. [DOI] [PubMed] [Google Scholar]
- 86.Amess R, Baggett N, Darby PR, Goode AR, Vickers EE. Synthesis of luciferin glycoside as substrate for novel ultrasensitive enzyme assays. Carbohydrate Res. 1990;205:225–233. [Google Scholar]
- 87.Monsees T, Miska W, Geiger R. Synthesis and characterization of a bioluminogenic substrate for α-chymotrypsin. Anal Biochem. 1994;221:329–334. doi: 10.1006/abio.1994.1421. [DOI] [PubMed] [Google Scholar]
- 88.Sudhaharan T, Reddy AR. A bifunctional luminogenic substrate for two luminescent enzymes: firefly luciferase and horseradish peroxidase. Anal Biochem. 1999;271:159–167. doi: 10.1006/abio.1999.4114. [DOI] [PubMed] [Google Scholar]
- 89.Zhou W, Valley MP, Shultz J, Hawkins EM, Bernad L, Good T, Good D, Riss TL, Klaubert DH, Wood KV. New bioluminogenic substrates for monoamine oxidase assays. J Am Chem Soc. 2006;128:3122–3123. doi: 10.1021/ja058519o. [DOI] [PubMed] [Google Scholar]
- 90.Valley MP, Zhou W, Hawkins EM, Shultz J, Cali JJ, Worzella T, Bernad L, Good T, Good D, Riss TL, Klaubert DH, Wood KV. A bioluminescent assay for monoamine oxidase activity. Anal Biochem. 2006;359:236–246. doi: 10.1016/j.ab.2006.09.035. [DOI] [PubMed] [Google Scholar]
- 91.Craig FF, Simmonds AC, Watmore D, McCapra F, White MRH. Membrane-permeable luciferin esters for assay of firefly luciferase in live intact cells. Biochem J. 1991;276:637–641. doi: 10.1042/bj2760637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Yong J, Thompson DB. An easily synthesized, photolyzable luciferase substrate for in vivo luciferase activity measurement. Biotechniques. 1993;15:848–850. [PubMed] [Google Scholar]
- 93.Shinde R, Perkins J, Contag CH. Luciferin derivatives for enhanced in vitro and in vivo bioluminescence assays. Biochemistry. 2006;45:11103–11112. doi: 10.1021/bi060475o. [DOI] [PubMed] [Google Scholar]
- 94.Jones LR, Goun EA, Shinde R, Rothbard JB, Contag CH, Wender PA. Releasable luciferin-transporter conjugates: tools for the real-time analysis of cellular uptake and release. J Am Chem Soc. 2006;128:6526–6527. doi: 10.1021/ja0586283. [DOI] [PubMed] [Google Scholar]
- 95.Wender PA, Goun EA, Jones LR, Pillow TH, Rothbard JB, Shinde R, Contag CH. Real-time analysis of uptake and bioactivatable cleavage of luciferin-transporter conjugates in transgenic reporter mice. Proc Natl Acad Sci USA. 2007;104:10340–10345. doi: 10.1073/pnas.0703919104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Chandran SS, Williams SA, Denmeade SR. Extended-release PEG-luciferin allows for long-term imaging of firefly luciferase activity in vivo. Luminescence. 2008;24:35–38. doi: 10.1002/bio.1060. [DOI] [PubMed] [Google Scholar]
- 97.Gates BJ, DeLuca M. The production of oxyluciferin during the firefly luciferase light reaction. Arch Biochem Biophys. 1975;169:616–621. doi: 10.1016/0003-9861(75)90205-2. [DOI] [PubMed] [Google Scholar]
- 98.McElroy WD, Hastings JW, Coulombre J, Sonnenfeld V. The mechanism of action of pyrophosphate in firefly luminescence. Arch Biochem Biophys. 1953;46:399–416. doi: 10.1016/0003-9861(53)90211-2. [DOI] [PubMed] [Google Scholar]
- 99.Lee RT, McElroy WD. Isolation and partial characterization of a peptide derived from the luciferin binding site of firefly luciferase. Arch Biochem Biophys. 1971;146:551–556. doi: 10.1016/0003-9861(71)90160-3. [DOI] [PubMed] [Google Scholar]
- 100.Lee RT, Denburg TL, McElroy WD. Substrate-binding properties of firefly luciferase. Arch Biochem Biophys. 1970;141:38–52. doi: 10.1016/0003-9861(70)90103-7. [DOI] [PubMed] [Google Scholar]
- 101.Ueda I. Effects of diethylether and halothane on firefly luciferin bioluminescence. Anesthesiology. 1965;26:603606. doi: 10.1097/00000542-196509000-00003. [DOI] [PubMed] [Google Scholar]
- 102.Ueda I, Suzuki A. Is there a specific receptor for anesthetics? Contrary effects of alcohols and fatty acids on phase transition and bioluminescence of firefly luciferase. Biophys J. 1998;75:1052–1057. doi: 10.1016/S0006-3495(98)77594-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Ueda I, Suzuki A. Irreversible phase transition of firefly luciferase: contrasting effects of volatile anesthetics and myristic acid. Biochim Biophys Acta. 1998;1380:313–319. doi: 10.1016/s0304-4165(97)00159-1. [DOI] [PubMed] [Google Scholar]
- 104.Rocha S, Campbell KJ, Roche KC, Perkins ND. The p53-inhibitor pifithrin-α inhibits firefly luciferase activity in vivo and in vitro. BMC Mol Biol. 2003;4:1–8. doi: 10.1186/1471-2199-4-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Niwa K, Ohmiya Y. Inhibitory effect of lipoic acid on firefly luciferase bioluminescence. Biochem Biophys Res Commun. 2004;323:625–629. doi: 10.1016/j.bbrc.2004.08.137. [DOI] [PubMed] [Google Scholar]
- 106.Heitman LH, van Veldhoven JP, Zweemer AM, Ye K, Brussee J, IJzerman AP. False positives in a reporter gene assay: Identification and synthesis of substituted N-pyridin-2-ylbenzamides as competitive inhibitors of firefly luciferase. J Med Chem. 2008;51:4724–4729. doi: 10.1021/jm8004509. [DOI] [PubMed] [Google Scholar]
- 107.Auld DS, Zhang YQ, Southall NT, Rai G, Landsman M, Maclure J, Langevin D, Thomas CJ, Austin CP, Inglese J. A basis for reduced chemical library inhibition of firefly luciferase obtained from directed evolution. J Med Chem. 2009;52:1450–1458. doi: 10.1021/jm8014525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Auld DS, Southall NT, Jadhav A, Johnson RL, Diller DJ, Simeonov A, Austin CP, Inglese J. Characterization of chemical libraries for luciferase inhibitory activity. J Med Chem. 2008;5:2372–2386. doi: 10.1021/jm701302v. [DOI] [PubMed] [Google Scholar]
- 109.Auld DS, Thorne N, Nguyen D-T, Inglese J. A specific mechanism for nonspecific activation in reporter-gene assays. ACS Chem Biol. 2008;3:463–470. doi: 10.1021/cb8000793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Green AA, McElroy WD. Crystalline firefly luciferase. Biochim Biophys Acta. 1956;20:170–176. doi: 10.1016/0006-3002(56)90275-x. [DOI] [PubMed] [Google Scholar]
- 111.Branchini BR, Marschner TM, Montemurro AM. A convenient affinity chromatography-based purification of firefly luciferase. Anal Biochem. 1980;104:386–396. doi: 10.1016/0003-2697(80)90089-5. [DOI] [PubMed] [Google Scholar]
- 112.Rajgopal S, Vijayalakshmi M. Purification of luciferase by affinity elution chromatography on Blue Dextran columns: comparison of Sepharose and silica as support matrices. J Chromatogr. 1982;243:164–167. [Google Scholar]
- 113.Belinga HF, Steghens JP, Collombel C. Firefly luciferase purification using polyethlene glycol and Dyematrex Orange A. J Chromatogr A. 1995;695:33–40. [Google Scholar]
- 114.Kajiyama N, Masuda T, Tatsumi H, Nakano E. Purification and characterization of luciferases from fireflies, Luciola cruciata and Luciola lateralis . Biochim Biophys Acta. 1992;1120:228–232. doi: 10.1016/0167-4838(92)90275-i. [DOI] [PubMed] [Google Scholar]
- 115.Devine JH, Kutuzova GD, Green VA, Ugarova NN, Baldwin TO. Luciferase from the East European firefly Luciola mingrelica: cloning and nucleotide sequence of the cDNA, overexpression in Escherichia coli and purification of the enzyme. Biochim Biophys Acta. 1993;1173:121–132. doi: 10.1016/0167-4781(93)90172-a. [DOI] [PubMed] [Google Scholar]
- 116.Branchini BR, Magyar RA, Murtiashaw MH, Anderson SM, Zimmer M. Site-directed mutagenesis of histidine 245 in firefly luciferase: a proposed model of the active site. Biochemistry. 1998;37:15311–15319. doi: 10.1021/bi981150d. [DOI] [PubMed] [Google Scholar]
- 117.Inouye S, Sahara Y. Soluble protein expression in E. coli cells using IgG binding domain of protein A as a solubilizing partner in the cold induced system. Biochem Biophys Res Commun. 2008;376:448–453. doi: 10.1016/j.bbrc.2008.08.149. [DOI] [PubMed] [Google Scholar]
- 118.de Wet JR, Wood KV, Helinski DR, DeLuca M. Cloning of firefly luciferase cDNA and the expression of active luciferase in Escherichia coli . Proc Natl Acad Sci USA. 1985;82:7870–7873. doi: 10.1073/pnas.82.23.7870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.de Wet JR, Wood KV, DeLuca M, Helinski DR, Subramani S. Firefly luciferase gene: structure and expression in mammalian cells. Mol Cell Biol. 1987;7:725–737. doi: 10.1128/mcb.7.2.725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Keller GA, Gould S, DeLuca M, Subramani S. Firefly luciferase is targeted to peroxisomes in mammalian cells. Proc Natl Acad Sci USA. 1987;84:3264–3268. doi: 10.1073/pnas.84.10.3264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Cho KH, Lee JS, Choi YD, Boo KS. Structural polymorphism of the luciferase gene in the firefly, Luciola lateralis . Insect Mol Biol. 1999;8:193–200. doi: 10.1046/j.1365-2583.1999.820193.x. [DOI] [PubMed] [Google Scholar]
- 122.Schröder J. Protein sequence homology between plant 4-coumarate:CoA ligase and firefly luciferase. Nucl Acids Res. 1989;17:460. doi: 10.1093/nar/17.1.460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Suzuki H, Kawarabayasi Y, Kondo J, Abe T, Nishikawa K, Kimura S, Hashimoto T, Yamamoto T. Structure and regulation of rat long-chain acyl-CoA synthetase. J Biol Chem. 1990;265:8681–8685. [PubMed] [Google Scholar]
- 124.Toh H. Sequence analysis of firefly luciferase family reveals a conservative sequence motif. Protein Seq Data Anal. 1991;4:111–117. [PubMed] [Google Scholar]
- 125.Ye L, Buck LM, Schaeffer HJ, Leach FR. Cloning and sequencing of a cDNA for firefly luciferase from Photuris pennsylvanica . Biochim Biophys Acta. 1997;1339:39–52. doi: 10.1016/s0167-4838(96)00211-7. [DOI] [PubMed] [Google Scholar]
- 126.Choi YS, Bae JS, Lee KS, Kim SR, Kim I, Kim JG, Kim KY, Kim SE, Suzuki H, Lee SM, Sohn HD, Jin BR. Genomic structure of the luciferase gene and phylogenetic analysis in the Hotaria-group fireflies. Comp Biochem Physiol . 2003;134B:199–214. doi: 10.1016/s1096-4959(02)00249-x. [DOI] [PubMed] [Google Scholar]
- 127.Conti E, Franks NP, Brick P. Crystal structure of firefly luciferase throws light on superfamily of adenylate-forming enzymes. Structure. 1996;4:287–298. doi: 10.1016/s0969-2126(96)00033-0. [DOI] [PubMed] [Google Scholar]
- 128.Franks NP, Jenkins JE, Conti E, Lieb WR, Brick P. Structural basis for the inhibition of firefly luciferase by a general anesthetic. Biophys J. 1998;75:2205–2211. doi: 10.1016/S0006-3495(98)77664-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Branchini BR, Murtiashaw MH, Carmody JN, Mygatt EE, Southworth TL. Synthesis of an N-acyl sulfamate analog of luciferyl-AMP: a stable and potent inhibitor of firefly luciferase. Bioorg Med Chem Lett. 2005;15:3860–3864. doi: 10.1016/j.bmcl.2005.05.115. [DOI] [PubMed] [Google Scholar]
- 130.Nakatsu T, Ichiyama S, Hiratake J, Saldanha A, Kobashi N, Sakata K, Kato H. Structural basis for the spectral difference in luciferase bioluminescence. Nature. 2006;440:372–376. doi: 10.1038/nature04542. [DOI] [PubMed] [Google Scholar]
- 131.Oba Y, Sato M, Ojika M, Inouye S. Enzymatic and genetic characterization of firefly luciferase and Drosophila CG6178 as a fatty acyl-CoA synthetase. Biosci Biotechnol Biochem. 2005;69:819–828. doi: 10.1271/bbb.69.819. [DOI] [PubMed] [Google Scholar]
- 132.Oba Y, Ojika M, Inouye S. Characterization of CG6178 gene product with high sequence similarity to firefly luciferase in Drosophila melanogaster . Gene. 2004;329:137–145. doi: 10.1016/j.gene.2003.12.026. [DOI] [PubMed] [Google Scholar]
- 133.Oba Y, Tanaka K, Inouye S. Catalytic properties of domain-exchanged chimeric Proteins between firefly luciferase and Drosophila fatty acyl-CoA synthetase CG6178. Biochem Biocsci Biotechnol. 2006;70:2739–2744. doi: 10.1271/bbb.60364. [DOI] [PubMed] [Google Scholar]
- 134.Oba Y, Sato M, Ohta Y, Inouye S. Identification of paralogous genes of firefly luciferase in the Japanese firefly, Luciola cruciata . Gene. 2006;368:53–60. doi: 10.1016/j.gene.2005.10.023. [DOI] [PubMed] [Google Scholar]
- 135.Oba Y, Sato M, Inouye S. Cloning and characterization of the homologous genes of firefly luciferase in the mealworm beetle, Tenebrio molitor . Insect Mol Biol. 2006;15:293–299. doi: 10.1111/j.1365-2583.2006.00646.x. [DOI] [PubMed] [Google Scholar]
- 136.Oba Y, Iida K, Ojika M, Inouye S. Orthologous gene of beetle luciferase in non-luminous click beetle, Agrypnus binodulus (Elateridae), encodes a fatty acyl-CoA synthetase. Gene. 2008;407:169–175. doi: 10.1016/j.gene.2007.10.013. [DOI] [PubMed] [Google Scholar]
- 137.Oba Y. On the origin of beetle luminescence. In: Meyer-Rochow VB, editor. Bioluminescence in focus. India: Res Signpost; 2009. pp. 277–290. [Google Scholar]
- 138.Oba Y, Iida K, Inouye S. Functional conversion of fatty acyl-CoA synthetase to firefly luciferase by site-directed mutagenesis: a key substitution responsible for luminescence activity. FEBS Lett. 2009;583:2004–2008. doi: 10.1016/j.febslet.2009.05.018. [DOI] [PubMed] [Google Scholar]
- 139.Masuda T, Tatsumi H, Nakano E. Cloning and sequence analysis of cDNA for luciferase of a Japanese firefly, Luciola cruciata . Gene. 1989;77:265–270. doi: 10.1016/0378-1119(89)90074-7. [DOI] [PubMed] [Google Scholar]
- 140.Tatsumi H, Kajiyama N, Nakano E. Molecular cloning and expression in Escherichia coli of a cDNA clone encoding luciferase of a firefly, Luciola lateralis . Biochim Biophys Acta. 1992;1131:161–165. doi: 10.1016/0167-4781(92)90071-7. [DOI] [PubMed] [Google Scholar]
- 141.Zenno S, Shiraishi S, Inouye S, Saigo K (1993) Photuris firefly luciferase gene. Jpn Pat Appl 5-119050
- 142.Ohmiya Y, Ohba N, Toh H, Tsuji FI. Cloning, expression and sequence analysis of cDNA for the luciferases from the Japanese fireflies, Pyrocoelia miyako and Hataria parvula . Photochem Photobiol . 1995;62:309–313. doi: 10.1111/j.1751-1097.1995.tb05273.x. [DOI] [PubMed] [Google Scholar]
- 143.Lee KS, Park HJ, Bae JS, Goo TW, Kim I, Sohn HD, Jin BR. Molecular cloning and expression of a cDNA encoding the luciferase from the firefly, Pyrocoelia rufa . J Biotechnol. 2001;92:9–19. doi: 10.1016/s0168-1656(01)00323-6. [DOI] [PubMed] [Google Scholar]
- 144.Choi YS, Lee KS, Bae JS, Lee KM, Kim SR, Kim I, Lee SM, Sohn HD, Jin BR. Molecular cloning and expression of a cDNA encoding the luciferase from the firefly, Hotaria unmunsana . Comp Biochem Physiol B Biochem Mol Biol. 2002;132:661–670. doi: 10.1016/s1096-4959(02)00085-4. [DOI] [PubMed] [Google Scholar]
- 145.Sala-Newby GB, Thomson CM, Campbell AK. Sequence and biochemical similarities between the luciferases of the glow-worm Lampyris noctiluca and the firefly Photinus pyralis . Biochem J. 1996;313:761–767. doi: 10.1042/bj3130761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Alipour BS, Hosseinkhani S, Nikkhah M, Naderi-Manesh H, Chaichi MJ, Osaloo SK. Molecular cloning, sequence analysis, and expression of a cDNA encoding the luciferase from the glow-worm, Lampyris turkestanicus . Biochem Biophys Res Commun. 2004;325:215–222. doi: 10.1016/j.bbrc.2004.10.022. [DOI] [PubMed] [Google Scholar]
- 147.Viviani VR, Arnoldi FG, Brochetto-Braga M, Ohmiya Y. Cloning and characterization of the cDNA for the Brazilian Cratomorphus distinctus larval firefly luciferase: similarities with European Lampyris noctiluca and Asiatic Pyrocoelia luciferases. Comp Biochem Physiol B Biochem Mol Biol. 2004;139:151–156. doi: 10.1016/j.cbpc.2004.05.012. [DOI] [PubMed] [Google Scholar]
- 148.Emamzadeh AR, Hosseinkhani S, Sadeghizadeh M, Nikkhah M, Chaichi MJ, Mortazavi M. cDNA cloning, expression and homology modeling of a luciferase from the firefly Lampyroidea maculata . J Biochem Mol Biol. 2006;39:578–585. doi: 10.5483/bmbrep.2006.39.5.578. [DOI] [PubMed] [Google Scholar]
- 149.Wood KV, Lam YA, Seliger HH, McElroy WD. Complementary DNA coding click beetle luciferases can elicit bioluminescence of different colors. Science. 1989;244:700–702. doi: 10.1126/science.2655091. [DOI] [PubMed] [Google Scholar]
- 150.Viviani VR, Silva AC, Perez GL, Santelli RV, Bechara EJ, Reinach FC. Cloning and molecular characterization of the cDNA for the Brazilian larval click-beetle Pyrearinus termitilluminans luciferase. Photochem Photobiol . 1999;70:254–260. doi: 10.1562/0031-8655(1999)070<0254:camcot>2.3.co;2. [DOI] [PubMed] [Google Scholar]
- 151.Stolz U, Velez S, Wood KV, Wood M, Feder JL. Darwinian natural selection for orange bioluminescent color in a Jamaican click beetle. Proc Natl Acad Sci USA. 2003;100:14955–14959. doi: 10.1073/pnas.2432563100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Viviani VR, Bechara EJ, Ohmiya Y. Cloning, sequence analysis, and expression of active Phrixothrix railroad-worms luciferases: relationship between bioluminescence spectra and primary structures. Biochemistry. 1999;38:8271–8279. doi: 10.1021/bi9900830. [DOI] [PubMed] [Google Scholar]