Abstract
Acronyms, abbreviations, and symbols play a significant role in clinical notes. Acronym and symbol sense disambiguation are crucial natural language processing (NLP) tasks that ensure the clarity and consistency of clinical notes and downstream NLP processing. Previous studies using traditional machine learning methods have been relatively successful in tackling this issue. In our research, we conducted an evaluation of large language models (LLMs), including ChatGPT 3.5 and 4, as well as other open LLMs, and BERT-based models, across three NLP tasks: acronym and symbol sense disambiguation, semantic similarity, and relatedness. Our findings emphasize ChatGPT’s remarkable ability to distinguish between senses with minimal or zero-shot training. Additionally, open source LLM Mixtrial-8x7B exhibited high accuracy for acronyms with fewer senses, and moderate accuracy for symbol sense accuracy. BERT-based models outperformed previous machine learning approaches, achieving an impressive accuracy rate of over 95%, showcasing their effectiveness in addressing the challenge of acronym and symbol sense disambiguation. Furthermore, ChatGPT exhibited a strong correlation, surpassing 70%, with human gold standards when evaluating similarity and relatedness.
Introduction
Word Sense Disambiguation (WSD) is a natural language processing (NLP) task that aims to determine the correct sense of a word in a given context. Many words in the English language have multiple meanings, and WSD helps NLP systems select the appropriate sense based on the surrounding words and context. Acronym and non-alphanumeric symbol sense disambiguation is a similar concept applied to acronyms and symbols. Doctors use a lot of acronyms and symbols in clinical notes for efficiency, precision, and standardization. These shorthand notations help healthcare professionals document patient information quickly, saving time in fast-paced medical settings.
However, ambiguity in acronym senses can introduce confusion and challenges when interpreting clinical notes or biomedical texts. For example, ‘AB’ could mean Abortion, Blood Group in ABO System, or other 10 senses. The plus symbol ‘+’ could mean Plus, or Reflexes etc. Our earlier study employed supervised machine learning methods1,2 to solve the acronyms and symbol disambiguation issues. In recent years, pre-trained language models (PLM) and Large Language Models (LLM) have gained popularity to solve traditional NLP problems due to their pretrained knowledge. The goal of this study aims to assess the performance of PLM (i.e., Bert-based models) and LLM (i.e..,ChatGPT 3.5 and 4) on acronym and symbol sense disambiguation.
Background
Traditional, automatically disambiguating words in biomedical text methods can be categorized into four groups3: supervised, semi-supervised, unsupervised, and knowledge-based. Supervised4–7 and semi-supervised8 techniques employ machine learning algorithms to assign senses to ambiguous words based on annotated training data. This requires creating training data for each target word, which is often impractical on a large scale. Unsupervised6,9 and knowledge-based3,10,11 methods, on the other hand, utilize external knowledge sources and sense inventories, bypassing the need for training data.
McInnes et al.3 creates UMLS::SenseRelate. It is an open-source Perl package designed for biomedical text analysis, specifically to disambiguate ambiguous terms by assigning UMLS concepts to them. This method assesses each potential sense of a word by calculating a score based on the similarity between that sense and the terms in the context window around the ambiguous word. The sense with the highest score is then assigned to the target word. To identify the terms surrounding the target word, the algorithm relies on the SPECIALIST Lexicon12, treating sequences of words that match lexicon terms as single entities. Once the terms are determined, the algorithm employs the open-source Perl package UMLS::Similarity13 to compute the similarity or relatedness between the potential sense of the target word and each of the identified surrounding terms, facilitating effective word sense disambiguation in biomedical texts.
Our previous supervised machine learning study1,2 examined into the Clinical Acronym Sense Inventory (CASI) dataset and emphasizes the window size, especially the left side of words within a context window around the target acronym or symbol that holds more disambiguation-relevant information than the right side. In recent years, with the advent of deep learning and neural network-based approaches, traditional WSD methods have been complemented and sometimes replaced by more advanced techniques. Neural networks and advanced NLP algorithms14 have ushered in a transformative era for WSD. Neural network-based approaches, such as transformer models like BERT and ELMo15, excel at capturing intricate contextual information, enabling them to discern word senses based on surrounding words.16 Agrawal et al. leveraged GPT-3 with zero-shot learning to work with the CASI dataset, achieving notable accuracy in sense disambiguation.17 With the release of ChatGPT 3.5 and 4, we aim to build upon this research and assess their performance. Our contributions encompass the evaluation of CASI and symbol sense disambiguation using ChatGPT 3.5 and 4, along with other LLMs (e.g., Mixtral-8x7B) and BERT-based models. Moreover, we examined the computational cost considerations related to using ChatGPT and the annotation process. Additionally, we discussed how to strike a balance between computational resources and human labeling efforts.
Methods
Data
CASI Dataset18: The study utilized clinical notes from Fairview Health Services spanning 2004 to 2008, originating from four metropolitan hospitals in the Twin Cities. These clinical notes, totaling 604,944, were primarily generated through voice dictation and transcription and encompassed various types, including admission notes, inpatient consult notes, operative notes, and discharge summaries. To identify relevant clinical acronyms and abbreviations, a hybrid technique combining heuristic rule-based and statistical methods was employed. Acronyms and abbreviations composed of capital letters (with or without numbers and specific symbols) and occurring over 500 times in the corpus were selected. For each of these acronyms or abbreviations, 500 random occurrences, along with the surrounding 12 preceding and subsequent word tokens, were manually annotated by two physicians to determine their meanings. These manual annotations served as the gold standard. The original CASI dataset has 75 acronyms, with a majority sense of less than or equal to 95%, totalling 37,500 de-identified clinical notes. For example, ‘CVA,’ which had two distinct senses: “cerebrovascular accident” (constituting 55.6% of samples) and “costovertebral angle” (44.4% of samples). We randomly selected 5 acronyms and visualized their sense distribution among 500 occurrences (Figure 1). Adams et al. chose 41 acronyms by removing noisy data and anonymizing the notes.19 In the ChatGPT experiments, we used this version of data to prompt ChatGPT for privacy consideration.
Figure 1.
Acronyms examples and their senses distributions.
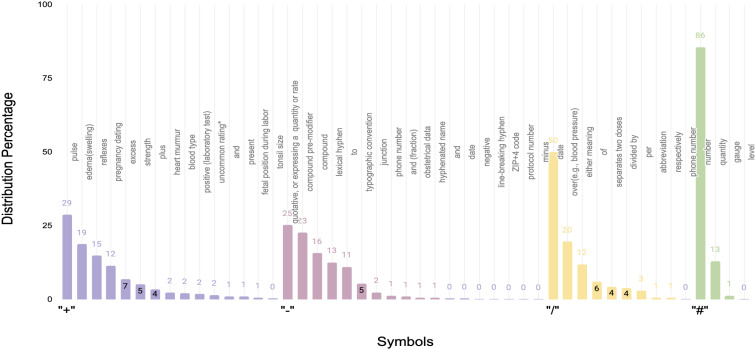
Symbol Dataset20: The researchers initially constructed a sense inventory for the target symbols (‘+’, ‘–’, ‘/’, and ‘#’) by drawing upon two textbooks, Speech and Language Processing21 and Foundations of Statistical Natural Language Processing22. Additionally, several medical references were consulted to obtain senses for these symbols. These medical references encompassed a medical dictionary, Stedman’s Medical Abbreviations, Acronyms & Symbols23, a medical terminological reference24, Medical Terminology and references of approved symbols25, and references from clinical literature, including Abbreviations and acronyms in healthcare26. Subsequently, the symbol sense inventory was refined by eliminating unclear senses and incorporating missing senses. The resulting senses and their distribution are visualized in Figure 2. The clinical notes for this dataset were anonymized and adhered to IRB protocols.
Figure 2.
“+”, “-”, “/”, and “#” symbol senses and distributions.
Similarity and Relatedness dataset27: This collection consists of reference standards used to test computer methods for measuring how similar or related medical terms are. Each dataset contains pairs of terms, systematically assessed by healthcare experts such as medical coders, residents, and clinicians, to determine their degree of closeness. MayoSRS.csv is a set of 101 medical concept pairs manually rater by medical coders for semantic relatedness. MiniMayoSRS.csv is a subset of 29 medical concept pairs manually rater with high inter-rater agreement.
ChatGPT
ChatGPT is a specialized variant of the GPT (Generative Pre-trained Transformer) model developed by OpenAI, tailored for NLP and generation in conversational contexts. It assists in various NLP tasks by generating coherent and contextually relevant responses in text-based conversations. ChatGPT’s applications include building chatbots, customer support, content generation, and language translation etc. Before we conducted the acronym sense disambiguation, we first tested the similarity/relatedness scoring by ChatGPT. Similarity and relatedness serve as the fundamental basis for traditional methods in word sense disambiguation3. Our aim is to understand how ChatGPT’s similarity scoring compared to human judgments and gold standards. A prompt example is shown in Figure 3 for the relatedness and similarity test.
Figure 3.
A ChatGPT prompt example for relatedness or similarity test.
A prompt is a specific instruction provided to ChatGPT to generate a desired response. When training or fine-tuning ChatGPT, prompts play a crucial role in guiding the model’s behavior. In Figures 4 and 5, we provided a zero-shot and one-shot prompt example, respectively, of ChatGPT to test acronym and symbol sense disambiguation.
Figure 4.
A ChatGPT zero-shot prompt example for acronym AMA, and ChatGPT gave the correct sense antimitochondrial antibody.
Figure 5.
A ChatGPT one-shot prompt example for symbol #, and ChatGPT gave the correct sense number.
Other LLM models
We also examined the open source LLM Mistral 8x7B28. The reason we chose this open source model is because of their impressive performance. Mixtral outperformed Llama 2 70B on most benchmarks, and it matches or outperforms GPT3.5 on most standard benchmarks. We randomly selected 3 notes for each selected acronym, and used zero-shot to measure the accuracy. Mistral 8x7B demands approximately 35 GB of RAM for execution. We refrained from fine-tuning it further with additional examples due to the substantial computing resources required, especially when multiple pipelines are concurrently active at the local computer center.
BERT-Based Models
Bidirectional Encoder Representations from Transformers (BERT)29 based models have revolutionized how we approach text classification. BERT learns contextualized word embeddings by considering the entire input text bidirectionally. One way to frame the acronym and symbol sense disambiguation task as a multi-class classification problem is to treat each possible sense of a word as a distinct class. In order to assess the impact of different training dataset sizes, we explored scenarios using the entire dataset, 50% of it, and just 25%, allowing us to determine the most suitable training size for testing purposes. For instance, if a symbol had 1000 examples, to evaluate 100 training sets, we first randomly selected 125 examples and allocated 80% (=100) for training and 20% (=25) for testing. We used the BioBERT30.
Results
Similarity and Relatedness
In the provided example in Figure 3, ChatGPT 3.5 assigned a similarity/relatedness score of 7.5 to “difficulty walking” and “antalgic gait,” whereas the human evaluator assigned a score of 6.69. We conducted these assessments through three rounds (temperature 0.5) of calculations for 101 concept pairs from the MayoSRS dataset. Subsequently, we employed the Pearson correlation coefficient to measure the correlation between ChatGPT’s results and human-established gold standards. We also replicated these experiments using the 29 pairs from the MiniMayoSRS dataset, and the resulting Pearson correlation coefficients are presented in Table 1. The findings revealed a strong association between similarity scores generated by ChatGPT and those determined by humans.
Table 1.
Pearson correlation of medical concepts similarity and relatedness experiments by ChatGPT.
| Round | MayoSRS | MiniMayoSRS |
|---|---|---|
| 1st | 71% | 78% |
| 2nd | 68% | 73% |
| 3rd | 67% | 70% |
ChatGPT Prompt Experiment Results
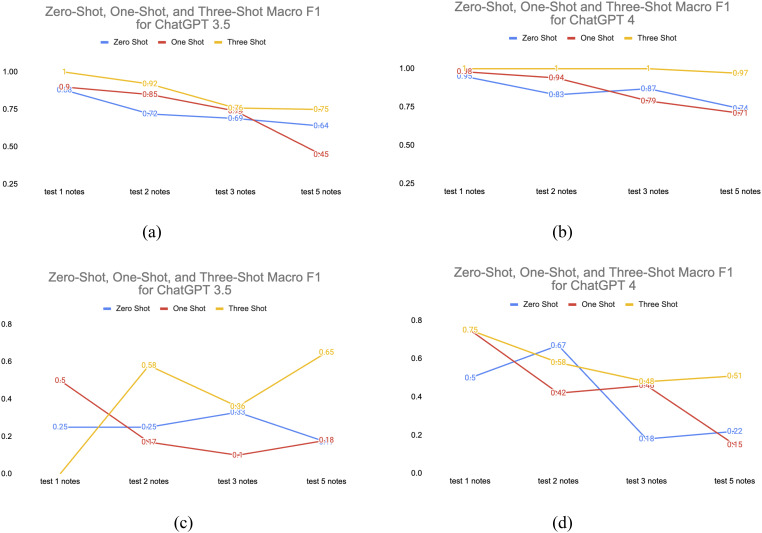
The ChatGPT experiment results are listed in Table 2. Average accuracies and Macro F1 were measured by 1 to 5 random tests for each acronym. Due to the budget limit to use ChatGPT API, we cannot test the whole clinical notes, and the accuracy and Macro F1 are the average results for the 41 acronyms with 1 to 5 randomly selected notes. Figure 6 shows the avg Macro F1 changes for different shots and tests. We set the temperature to 0 for the experiments. Compared to previous research by Agrawal et al.17, the zero shot Macro F1 of ChatGPT 4 has increased from 0.69 to 0.75 (5 notes).
Table 2.
Zero-, One-, and Three-Shot for ChatGPT 3.5 and 4 for the acronym and symbol sense disambiguation.
| Acronym | Symbol | |||||||
|---|---|---|---|---|---|---|---|---|
| ChatGPT 3.5 | ChatGPT 4 | ChatGPT 3.5 | ChatGPT 4 | |||||
| Zero-Shot | ||||||||
| Accuracy | Macro F1 | Accuracy | Macro F1 | Accuracy | Macro F1 | Accuracy | Macro F1 | |
| 1 notes | 0.88 | 0.88 | 0.95 | 0.95 | 0.25 | 0.25 | 0.50 | 0.05 |
| 2 notes | 0.77 | 0.72 | 0.87 | 0.83 | 0.38 | 0.25 | 0.75 | 0.67 |
| 3 notes | 0.79 | 0.69 | 0.91 | 0.87 | 0.42 | 0.33 | 0.25 | 0.18 |
| 5 notes | 0.75 | 0.64 | 0.85 | 0.74 | 0.25 | 0.17 | 0.35 | 0.22 |
| One-Shot | ||||||||
| 1 notes | 0.90 | 0.90 | 0.98 | 0.98 | 0.50 | 0.50 | 0.75 | 0.75 |
| 2 notes | 0.88 | 0.85 | 0.95 | 0.94 | 0.25 | 0.17 | 0.50 | 0.42 |
| 3 notes | 0.83 | 0.74 | 0.86 | 0.79 | 0.17 | 0.10 | 0.50 | 0.46 |
| 5 notes | 0.83 | 0.45 | 0.83 | 0.71 | 0.15 | 0.18 | 0.35 | 0.15 |
| Three-Shot | ||||||||
| 1 notes | 1.00 | 1.00 | 1.00 | 1.00 | 0.00 | 0.00 | 0.75 | 0.75 |
| 2 notes | 0.94 | 0.92 | 1.00 | 1.00 | 0.63 | 0.58 | 0.63 | 0.58 |
| 3 notes | 0.84 | 0.76 | 1.00 | 1.00 | 0.42 | 0.36 | 0.58 | 0.48 |
| 5 notes | 0.84 | 0.75 | 0.98 | 0.97 | 0.75 | 0.65 | 0.65 | 0.51 |
Figure 6.
(a) ChatGPT 3.5 and (b) ChatGPT 4 average Macro F1 for acronym sense disambiguation. (c) ChatGPT 3.5 and (d) ChatGPT 4 average Macro F1 scores for symbol sense disambiguation.
From the acronym experiments we could see the decrease in the average macro F1 score while the average accuracy remains stable for ChatGPT 3.5. This indicaties of class imbalance issues in this multiclass classification problems. The CASI dataset is highly imbalanced data. The decrease in macro F1 score suggests that the model’s performance on minority senses was worsening across different multiclasses. Macro F1 gives equal weight to each class, and if some classes are underrepresented or poorly predicted, it can result in a decrease in macro F1. Accuracy is less sensitive to class imbalances, especially when one or more classes dominate the dataset. If the majority class has high accuracy, it can offset lower accuracy in minority classes, leading to a stable overall accuracy. Nonetheless, the average Macro F1 score had no sudden drop and remained consistent for ChatGPT 4, especially for three-shot, suggesting its continued robustness for handling imbalanced datasets. Additionally, Oh et al. conducted a separate study about surgical education and training that also demonstrated improved accuracy in ChatGPT 4.31
For the symbol disambiguation tests, the analysis of individual tests reveals fluctuations in model performance for both ChatGPT 3.5 and 4. The zero-shot scenario generally demonstrates lower accuracy and Macro F1 scores, while the one-shot and three-shot scenarios show improved performance, with variations among individual tests. The number of examples provided to the model appears to impact its ability to classify samples accurately and maintain a balance between precision and recall. These results suggest that providing more training examples or shots can enhance the model’s ability to classify samples accurately and maintain a better balance between precision and recall. Since the dataset has only 4 symbols, the results can vary when ChatGPT doesn’t provide a correct answer for one of the symbols. Overall, ChatGPT 4 consistently outperforms ChatGPT 3.5 across all scenarios.
Another important factor to take into account is the length of the prompt. ChatGPT-3.5 or 4 has a context limit of 4,096 tokens, roughly equivalent to 3,072 words. The length of the prompt is primarily constrained by the length of the clinical notes. The average and maximum prompt length for three-shot is 825 and 2130. ChatGPT 3.5 has no token per day limit (TPD), while ChatGPT 4 is limited to 50,000 TPD as of the date we submitted the paper. The length of the prompts is a crucial consideration when designing experiments.
Open Source LLMs and BERT-Based Model Experiment Results
In prior research, Moon et al. conducted a comparison of different machine learning algorithms, including SVM, Naive Bayes, and decision trees. The study also examined the consideration and discussion of window and training sizes.1,2 We opted for the BioBERT model due to its impressive ability to comprehend contextual information and relationships within sentences. The bidirectional architecture of BERT allows it to capture the context of words effectively in a sentence. Our approach also involved utilizing open-source LLMs solutions locally. We chose the Mixtral 8x7B28 model with zero-shot test by randomly selecting three notes. The results of these experiments are presented in Tables 3. We can observe that with the increase of the training sizes, the accuracy increased, especially for acronyms with less senses.
Table 3.
Use Mixtral-8x7B zero shot and BioBERT with different training sizes 100, 200, and 400 for acronym sense disambiguation.
| Acronym Sense Disambiguation Accuracy | |||||
|---|---|---|---|---|---|
| Mixtral-8x7B Zero-Shot | BioBERT | ||||
| Acronym | #Sense | Accuracy | 100 training | 200 training | 400 training |
| PD | 15 | 33% | 82% | 92% | 92% |
| AB | 12 | 67% | 90% | 97% | 96% |
| LE | 7 | 100% | 92% | 92% | 92% |
| OP | 6 | 33% | 88% | 93% | 97% |
| PDA | 3 | 100% | 92% | 96% | 100% |
| VBG | 2 | 100% | 100% | 100% | 100% |
| Symbol Sense Disambiguation | |||||
| Symbol | #Sense | Accuracy | 100 training | 400 training | 800 training |
| - | 18 | 67% | 75% | 95% | 98% |
| + | 15 | 67% | 80% | 92% | 98% |
| / | 10 | 34% | 83% | 96% | 98% |
| # | 4 | 67% | 92% | 98% | 99% |
Cost
At the time of our paper submission, Table 4 provides the pricing details for ChatGPT. Notably, the price of gpt-4-0613 is approximately 30 times higher than that of gpt-3.5-turbo-1106. As shown in the accuracy and F1 comparison in Table 2, the performance of three-shot ChatGPT 3.5 rivals that of zero or one-shot ChatGPT 4, while being significantly more cost-effective. Considering our task’s requirement for extensive input and minimal output from ChatGPT, opting for cost-effective input models like gpt-4-1106-preview could lead to significant cost savings. The cost of annotation by human annotators can fluctuate between $0.001 and $0.13 per unit of text in Q3 2023, contingent on the annotation units and complexity involved. A study conducted by Microsoft’s Wang et al. analyzed nine datasets and proposed that labeling data using fine tuned GPT-3 for downstream NLP tasks could result in cost savings ranging from 50% to 96%.32 These nine datasets didn’t include any medical dataset. Our experiments have shown that ChatGPT can effectively separate the correct sense of the medical acronyms, which support the possibility for automatically labeling medical information by ChatGPT. In addition to being costly, human labeling is also extremely time-consuming. The original CASI dataset took months to be annotated. In our experiments, the total cost for ChatGPT 4 is $28 in total for the CASI dataset and $3 for the symbol dataset, and the majority of tasks can be completed in under 10 minutes.
Table 4.
The pricing of the ChatGPT models used in experiments.
| Model | Input Price | Output Price |
|---|---|---|
| ChatGPT 3.5 gpt-3.5-turbo-1106 | $0.0010/1k token | $0.0020/1k token |
| ChatGPT 4 gpt-4-0613 | $0.03 / 1K tokens | $0.06 / 1K tokens |
| ChatGPT 4 gpt-4-1106-preview | $0.01 / 1K tokens | $0.03 / 1K tokens |
Discussion
The dataset for disambiguating 41 acronyms and 4 symbols is small in scale. Nevertheless, the advantage of a modest dataset lies in its ability to offer us a closer look at ChatGPT’s performance. During our evaluation of ChatGPT 3.5 and 4, we observed that significant performance improvements were achieved for ChatGPT 4 for CASI dataset. ChatGPT 4 has 1.76 trillion parameters which is 10 times of ChatGPT 3.5. ChatGPT exhibits the ability to retain memory and engage in self-training.3432 Another reason we need to remember is the CASI and symbol dataset are both highly unbalanced. When a random note was selected, the probability to select a majority sense is very high. Another thing we need to notice is the memory of LLMs. However, as of now, we have not discovered a method to eliminate previous training from the memory of LLMs from the end user side. Conducting multiple training iterations on these acronyms leads to performance improvements.
The BioBERT model demonstrates consistent and resilient performance when supplied with additional training examples for both acronym and symbol datasets. It’s worth noting that BERT-based models, being encoders, tend to benefit from a larger number of training examples. On the other hand, GPT-based models, as decoders, can achieve impressive accuracy with just three shots, as evidenced by the acronym dataset. The choice of model should be made judiciously, taking into consideration the specific dataset and tasks at hand.
As LLMs gain widespread popularity, an increasing number of researchers are eager to assess their capabilities, particularly with open-source LLMs. This surge in demand can potentially strain our local servers’ memory capacity. Our research heavily relies on a vast dataset of clinical notes, but due to privacy and safety concerns, we cannot utilize ChatGPT for processing them. In such scenarios, smaller yet more accurate LLMs become highly valuable for our research. This also raises the question of optimizing the trade-off between human labeling and automatic labeling through ChatGPT, especially when dealing with projects that have limited budgets and timeframes.
We also conducted evaluations using BioGPT,33 which was trained on biomedical literature, and ClinicalGPT,32 trained on Chinese clinical notes, on the CASI dataset. However, their performance on the CASI dataset did not yield satisfactory results. In comparison, Mixtral 8x7B28 outperformed the other two models, albeit with a high memory requirement of approximately 35 GB RAM. This posed challenges for our local computing center, especially when multiple pipelines were running concurrently, resulting in reduced processing speed. Consequently, we conducted zero-shot testing with a limited selection of randomly chosen notes. Our future work will explore the possibility of fine-tuning smaller GPT models using local clinical notes. This endeavor would necessitate the availability of a high-quality question-answer dataset extracted from our local clinical notes. We will consider leveraging previously annotated datasets and open-source clinical datasets such as MIMIC III34 to facilitate this task.
Conclusion
We leveraged LLMs to address the intricate task of disambiguating acronyms and symbols within clinical notes. In the face of the intricate nuances of clinical language and significant data imbalances, ChatGPT 3.5 and 4 showcased impressive and promising outcomes. Additionally, we conducted experiments with open-source GPTs and BERT-based models. These discoveries led us to explore the related expenses, computing resource demands, and human resource aspects, offering valuable insights for researchers in the domain to contemplate.
Acknowledgements
This work was supported by the National Institutes of Health under grant number 2R01AT009457, 1R01AG078154, and 1R01CA287413. The content is solely the responsibility of the authors and does not represent the official views of the National Institutes of Health. The authors would like to acknowledge support from the Center for Learning Health System Sciences, a partnership between the Medical School and School of Public Health at the University of Minnesota.
Figures & Table
References
- 1.Moon S, Pakhomov S, Melton GB. Automated disambiguation of acronyms and abbreviations in clinical texts: window and training size considerations. AMIA Annu Symp Proc AMIA Symp. 2012;2012:1310–9. [PMC free article] [PubMed] [Google Scholar]
- 2.Moon S, Pakhomov S, Ryan J, Melton GB. Automated non-alphanumeric symbol resolution in clinical texts. AMIA Annu Symp Proc AMIA Symp. 2011;2011:979–86. [PMC free article] [PubMed] [Google Scholar]
- 3.McInnes BT, Pedersen T, Liu Y, Melton GB, Pakhomov SV. Knowledge-based method for determining the meaning of ambiguous biomedical terms using information content measures of similarity. AMIA Annu Symp Proc AMIA Symp. 2011;2011:895–904. [PMC free article] [PubMed] [Google Scholar]
- 4.Liu H. A Multi-aspect Comparison Study of Supervised Word Sense Disambiguation. J Am Med Inform Assoc. 2004 Apr 2;11(4):320–31. doi: 10.1197/jamia.M1533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Leroy G, Rindflesch TC. Effects of information and machine learning algorithms on word sense disambiguation with small datasets. Int J Med Inf. 2005 Aug;74(7–8):573–85. doi: 10.1016/j.ijmedinf.2005.03.013. [DOI] [PubMed] [Google Scholar]
- 6.McInnes BT, Pedersen T, Carlis J. Using UMLS Concept Unique Identifiers (CUIs) for word sense disambiguation in the biomedical domain. AMIA Annu Symp Proc AMIA Symp. 2007 Oct 11;2007:533–7. [PMC free article] [PubMed] [Google Scholar]
- 7.Stevenson M, Guo Y, Gaizauskas R, Martinez D. Disambiguation of biomedical text using diverse sources of information. BMC Bioinformatics. 2008 Dec;9(S11):S7. doi: 10.1186/1471-2105-9-S11-S7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Fan JW, Friedman C. Word sense disambiguation via semantic type classification. AMIA Annu Symp Proc AMIA Symp. 2008 Nov 6;2008:177–81. [PMC free article] [PubMed] [Google Scholar]
- 9.Pedersen T. In: Proceedings of the 1st ACM International Health Informatics Symposium [Internet] Arlington Virginia USA: ACM; 2010 [cited 2023 Sep 13]. Pedersen effect of different context representations on word sense discrimination in biomedical texts; pp. 56–65. Available from: https://dl.acm.org/doi/10.1145/1882992.1883003. [Google Scholar]
- 10.Humphrey SM, Rogers WJ, Kilicoglu H, Demner-Fushman D, Rindflesch TC. Word sense disambiguation by selecting the best semantic type based on Journal Descriptor Indexing: Preliminary experiment. J Am Soc Inf Sci Technol. 2006 Jan 1;57(1):96–113. doi: 10.1002/asi.20257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Alexopoulou D, Andreopoulos B, Dietze H, Doms A, Gandon F, Hakenberg J, et al. Biomedical word sense disambiguation with ontologies and metadata: automation meets accuracy. BMC Bioinformatics. 2009 Dec;10(1):28. doi: 10.1186/1471-2105-10-28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lu CJ, Payne A, Mork JG. The Unified Medical Language System SPECIALIST Lexicon and Lexical Tools: Development and applications. J Am Med Inform Assoc. 2020 Oct 1;27(10):1600–5. doi: 10.1093/jamia/ocaa056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.McInnes BT, Pedersen T, Pakhomov SVS. UMLS-Interface and UMLS-Similarity : open source software for measuring paths and semantic similarity. AMIA Annu Symp Proc AMIA Symp. 2009 Nov 14;2009:431–5. [PMC free article] [PubMed] [Google Scholar]
- 14.Popov A. Popov Network Models for Word Sense Disambiguation: An Overview. Cybern Inf Technol. 2018 Mar 1;18(1):139–51. [Google Scholar]
- 15.Peng Y, Yan S, Lu Z. Transfer Learning in Biomedical Natural Language Processing: An Evaluation of BERT and ELMo on Ten Benchmarking Datasets. 2019 [cited 2023 Sep 13]. Available from: https://arxiv.org/abs/1906.05474.
- 16.Wagh A, Khanna M. Clinical Abbreviation Disambiguation Using Clinical Variants of BERT. In: Morusupalli R, Dandibhotla TS, Atluri VV, Windridge D, Lingras P, Komati VR, editors. Multi-disciplinary Trends in Artificial Intelligence [Internet] Cham: Springer Nature Switzerland; 2023 [cited 2024 Jan 23]. pp. 214–24. Available from: https://link.springer.com/10.1007/978-3-031-36402-0_19. [Google Scholar]
- 17.Agrawal M, Hegselmann S, Lang H, Kim Y, Sontag D. Large Language Models are Few-Shot Clinical Information Extractors. 2022 [cited 2024 Jan 16]. Available from: https://arxiv.org/abs/2205.12689.
- 18.Moon S, Pakhomov S, Liu N, Ryan JO, Melton GB. A sense inventory for clinical abbreviations and acronyms created using clinical notes and medical dictionary resources. J Am Med Inform Assoc. 2014 Mar;21(2):299–307. doi: 10.1136/amiajnl-2012-001506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Adams G, Ketenci M, Bhave S, Perotte A, Elhadad N. Zero-Shot Clinical Acronym Expansion via Latent Meaning Cells. Proc Mach Learn Res. 2020 Dec;136:12–40. [PMC free article] [PubMed] [Google Scholar]
- 20.Moon S, Pakhomov S, Melton G. Clinical Symbol Sense Inventory [Internet] 2012 [cited 2023 Sep 13]. Available from: http://conservancy.umn.edu/handle/11299/137704.
- 21.Jurafsky D, Martin JH. Speech and language processing: an introduction to natural language processing, computational linguistics, and speech recognition. Upper Saddle River, N.J: Prentice Hall. 2000;934 p. (Prentice Hall series in artificial intelligence) [Google Scholar]
- 22.Manning CD, Schütze H. Cambridge, Mass: MIT Press; 1999. Foundations of statistical natural language processing; p. 680. [Google Scholar]
- 23.Stedman TL, editor. 5th ed. Philadelphia: Wolters Kluwer Health/Lippincott Williams & Wilkins; 2013. Stedman’s medical abbreviations, acronyms & symbols; p. 1. [Google Scholar]
- 24.Yumpu.com. SHC Approved Abbreviations, Acronyms and Symbols - Stanford ... [Internet] yumpu.com. [cited 2023 Sep 13]. Available from: https://www.yumpu.com/en/document/view/20706571/shc-approved-abbreviations-acronyms-and-symbols-stanford-
- 25.Approved and Unapproved Abbreviations For Medical Records | PDF | Glaucoma | Human Eye [Internet] Scribd. [cited 2023 Sep 13]. Available from: https://www.scribd.com/document/111543918/Approved-and-Unapproved-Abbreviations-for-Medical-Records.
- 26.Kuhn IF. Abbreviations and acronyms in healthcare: when shorter isn’t sweeter. Pediatr Nurs. 2007;33(5):392–8. [PubMed] [Google Scholar]
- 27.Pakhomov S. Pakhomov Relatedness and Similarity Reference Standards for Medical Terms [Internet] 2018 [cited 2023 Sep 13]. Available from: http://conservancy.umn.edu/handle/11299/196265.
- 28.Jiang AQ, Sablayrolles A, Roux A, Mensch A, Savary B, Bamford C, et al. Mixtral of Experts. 2024 [cited 2024 Jan 19]. Available from: https://arxiv.org/abs/2401.04088.
- 29.Devlin J, Chang MW, Lee K, Toutanova K. BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding [Internet] arXiv. 2019 [cited 2022 Nov 15]. Available from: http://arxiv.org/abs/1810.04805.
- 30.Lee J, Yoon W, Kim S, Kim D, Kim S, So CH, et al. BioBERT: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics. 2020 Feb 15;36(4):1234–40. doi: 10.1093/bioinformatics/btz682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Oh N, Choi GS, Lee WY. ChatGPT goes to the operating room: evaluating GPT-4 performance and its potential in surgical education and training in the era of large language models. Ann Surg Treat Res. 2023 May;104(5):269–73. doi: 10.4174/astr.2023.104.5.269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wang G, Yang G, Du Z, Fan L, Li X. ClinicalGPT: Large Language Models Finetuned with Diverse Medical Data and Comprehensive Evaluation. 2023 [cited 2024 Jan 19]. Available from: https://arxiv.org/abs/2306.09968.
- 33.Luo R, Sun L, Xia Y, Qin T, Zhang S, Poon H, et al. BioGPT: generative pre-trained transformer for biomedical text generation and mining. Brief Bioinform. 2022 Nov 19;23(6):bbac409. doi: 10.1093/bib/bbac409. [DOI] [PubMed] [Google Scholar]
- 34.Wang S, McDermott MBA, Chauhan G, Ghassemi M, Hughes MC, Naumann T. Proceedings of the ACM Conference on Health, Inference, and Learning [Internet] Toronto Ontario Canada: ACM; 2020 [cited 2024 Jan 19]. MIMIC-Extract: a data extraction, preprocessing, and representation pipeline for MIMIC-III; pp. 222–35. Available from: https://dl.acm.org/doi/10.1145/3368555.3384469. [Google Scholar]